Abstract
The IRE1α-XBP1 pathway, a key component of the endoplasmic reticulum (ER) stress response, is considered to be a critical regulator for survival of multiple myeloma (MM) cells. Therefore, the availability of small-molecule inhibitors targeting this pathway would offer a new chemotherapeutic strategy for MM. Here, we screened small-molecule inhibitors of ER stress-induced XBP1 activation, and identified toyocamycin from a culture broth of an Actinomycete strain. Toyocamycin was shown to suppress thapsigargin-, tunicamycin- and 2-deoxyglucose-induced XBP1 mRNA splicing in HeLa cells without affecting activating transcription factor 6 (ATF6) and PKR-like ER kinase (PERK) activation. Furthermore, although toyocamycin was unable to inhibit IRE1α phosphorylation, it prevented IRE1α-induced XBP1 mRNA cleavage in vitro. Thus, toyocamycin is an inhibitor of IRE1α-induced XBP1 mRNA cleavage. Toyocamycin inhibited not only ER stress-induced but also constitutive activation of XBP1 expression in MM lines as well as primary samples from patients. It showed synergistic effects with bortezomib, and induced apoptosis of MM cells including bortezomib-resistant cells at nanomolar levels in a dose-dependent manner. It also inhibited growth of xenografts in an in vivo model of human MM. Taken together, our results suggest toyocamycin as a lead compound for developing anti-MM therapy and XBP1 as an appropriate molecular target for anti-MM therapy.
Keywords: multiple myeloma, ER stress, IRE1α, XBP1, toyocamycin, adenosine analog
Introduction
Multiple myeloma (MM) is a hematological malignancy characterized by the accumulation of clonogenic mature plasma cells in bone marrow. Recently, bortezomib (BTZ), a proteasome inhibitor, was approved by the Food and Drug Administration for the treatment of MM. However, BTZ treatment often achieves only very short-duration responses and drug resistance tends to develop rapidly.1, 2 Therefore, there is a need to develop novel molecular targets for new therapeutic approaches.
Recent studies have suggested that XBP1, a basic region/leucine zipper (bZIP) transcription factor of the CREB-ATF family, has an important role in the survival of MM cells. XBP1 is required for the terminal differentiation of B lymphocytes to plasma cells and is essential for immunoglobulin secretion.3, 4 Abundant or deregulated expression of XBP1 has been detected in MM cells5, 6 and in hepatocellular carcinomas.7, 8 Because of the production of abundant immunoglobulins and cytokines, MM cells must be able to survive under conditions of chronic endoplasmic reticulum (ER) stress. This involves the unfolded protein response (UPR) including activation of the IRE1α-XBP1 pathway. In addition, MM cells are located in the bone marrow milieu, which is usually considered hypoxic compared with other organs.9, 10 Therefore, MM cells have to survive and grow even at low oxygen, with poor nutrition and adverse pH in vivo. Thus, MM cells need to possess mechanisms to protect against ER stress. Among the UPR in MM cells, the IRE1α-XBP1 pathway has been implicated in the proliferation and survival of MM cells to a greater extent than in monoclonal gammopathy of undetermined significance or normal plasma cells.11 It has been reported to be a prognostic factor12 and could be a target for immunotherapy13 or chemotherapy.14 Based on previous reports, it is proposed that an inhibitor of IRE1α-XBP1 activation should be a potent therapeutic agent for MM.
The transcriptional activity of XBP1 is regulated by ER-located transmembrane kinase/endoribonuclease (RNase) protein IRE1α. Recent studies have proposed a model of ER stress-induced IRE1α activation and subsequent XBP1 activation as follows: (1) accumulation of unfolded proteins triggers oligomerization of luminal domains of IRE1α.15 (2) Oligomerization of IRE1α causes trans-auto-phosphorylation of the kinase activation loop domain, which leads to a conformational change.16, 17, 18, 19 (3) This conformational change permits cofactor (ADP) binding,16, 20 promoting back-to-back dimer configuration of cytosolic domains.21 (4) The oligomerization of cytosolic domains activates the RNase activity of IRE1α which subsequently cleaves XBP1 mRNA at two sites to initiate an unconventional splicing reaction.21 (5) IRE1α-induced cleavage of XBP1 mRNA results in the removal of a 26-nucleotide intron and the 5′ and 3′ fragments are subsequently joined by RNA ligase activity. This unconventional splicing reaction creates a translational frame shift to produce an active XBP1 transcription factor.22, 23
Previously, we reported a novel screening system for inhibitors of XBP1 activation, using luciferase reporter signals in HeLa/XBP1-luc cells.24 In the present study, we identified toyocamycin25 as an XBP1 inhibitor in the culture broth of an Actinomycete strain using this screening system (Figure 1a). We observed that toyocamycin inhibited IRE1α-induced ATP-dependent XBP1 mRNA cleavage in vitro without affecting IRE1α auto-phosphorylation. Moreover, this compound markedly inhibited not only ER stress-induced but also constitutively activated IRE1α-XBP1 pathway both in MM cell lines and primary MM cells, resulting in strong cytotoxic activity.
Figure 1.
Toyocamycin suppressed thapsigargin, tunicamycin or 2-deoxyglucose-induced XBP1 mRNA splicing. (a) Structure of toyocamycin, sagivamycin and tubercidin. (b) Effect of toyocamycin on thapsigargin-induced XBP1 activation. HeLa/XBP1-luc cells were treated with the indicated concentration of toyocamycin in the presence of 0.1 μℳ of thapsigargin. After 24 h, the cells were lysed and subjected to luciferase assay. Data are the fold change+s.d. of a thapsigargin-induced luciferase activity in the presence or absence of various concentration of toyocamycin. All experiments were performed in triplicate. (c) Toyocamycin inhibition of thapsigargin-, tunicamycin- or 2-deoxyglucose (2DG)-induced endogenous XBP1 mRNA splicing. HeLa cells were treated with the indicated concentration of toyocamycin in the presence or absence of 0.1 μℳ of thapsigargin, 10 μg/ml of tunicamycin or 1 mℳ of 2DG for 4 h. The cells were collected and RNA extracted. Spliced- or unspliced-XBP1 mRNA was detected as described in Materials and methods. (d) Effect of toyocamycin or actinomycin D on [3H]-uridine incorporation into acid-insoluble fractions of HeLa cells. HeLa cells were incubated with the indicated concentration of toyocamycin or actinomycin D in the presence of 1 μCi/ml [3H]-uridine for 1 h. The reaction was stopped by addition of 10% TCA, and the acid-insoluble fractions were collected. Data represent the mean of three experiments. (e) Effect of actinomycin D on thapsigargin-induced endogenous XBP1 mRNA splicing assessed by RT-PCR. HeLa cells were treated with the indicated concentration of actinomycin D in the presence or absence of 0.1 μℳ of thapsigargin for 4 h. The cells were collected and RNA extracted. Spliced- or unspliced-XBP1 mRNA was detected as described in Materials and Methods.
Materials and methods
Cell culture and reagents
Human epithelial adenocarcinoma HeLa cells and previously generated HeLa/XBP1-luc cells24 were cultured in DMEM supplemented with 10% FBS. Human MM and other hematological cell lines were cultured in RPMI-1640 supplemented with 10% FBS. Human fibrosarcoma HT1080 was cultured in EMEM supplemented with 2 mℳ glutamine, 1% non-essential amino acids and 10% FBS. A BTZ-resistant MM cell lines, KMS-11/BTZ and OPM-2/BTZ, were established from the parental line, KMS-11 and OPM-2, respectively, under continuous exposure to BTZ over a half year.26 Toyocamycin, sangivamycin, tubercidin, tunicamycin, 2-deoxyglucose and 5-fluorouracil were purchased from Sigma-Aldrich (St Louis, MO, USA). Thapsigargin was purchased from Santa Cruz Biotechnology (Santa Cruz, CA, USA). BTZ was purchased from Toronto Research Chemicals (North York, ON, Canada).
Primary MM specimens
Nine primary MM specimens derived from eight patients with symptomatic MM were obtained after written informed consent at Nagoya City University Hospital. The assay protocols using patient samples were approved by the Institutional Ethical Committee. MM cells were purified from the marrow mononuclear cell fraction or pleural effusion using anti-CD138 antibody-coated beads with the aid of an automatic magnetic cell sorting system (Miltenyi Biotec, Auburn, CA, USA).26
Preparation of toyocamycin
The culture broth (3 l) of Streptmyces sp. 1893-56 was extracted with EtOAc, filtered and concentrated in vacuo. This suspension was extracted with hexane, and the insoluble fraction concentrated. The active fractions were then collected and further isolated by silica gel column chromatography (Silica gel 60, 60–230 μm; Merck, Darmstadt, Germany) using a CHCl3-MeOH stepwise system. As a result, we obtained 5 mg of active compound. The UV spectrum, HRESI-MS measurement, and 1H NMR spectra of this active compound confirmed its identity as toyocamycin.
Luciferase assay
As previously reported,24 HeLa/XBP1-luc cells were seeded into 96-well plates at 2 × 104 cells/well, and then incubated with 0.1 μℳ of thapsigargin together with or without test compounds. After 24 h of incubation, the cells were lysed in Passive lysis buffer (Promega, Madison, WI, USA), and luciferase activity measured using the luciferase assay system (Promega) and a luminometer (Wallac, PerkinElmer, Waltham, MA, USA). IC50 values were determined from the dose–response curves of the inhibition of XBP1-luciferase activity, setting the result of thapsigargin treatment as 100%.
Reverse transcriptase (RT)-PCR and real-time PCR analysis
HeLa cells and MM cell lines were incubated with test compounds for 4 or 6 h together with the ER stress inducers thapsigargin, tunicamycin, or 2-deoxyglucose. Briefly, total RNA was extracted from HeLa cells using TRIzol reagent (Invitrogen, Carlsbad, CA, USA). Aliquots of 2 μg of total RNA were treated with M-MLV reverse transcriptase (Promega) to produce first-strand complementary DNA (cDNA). For RT-PCR analysis, this first-strand cDNA was subjected to PCR with KOD plus polymerase (Toyobo, Osaka, Japan) using a pair of primers corresponding to nucleotides 505–525 and 609–629 of XBP1 cDNA. The amplified products were separated by electrophoresis on an 8% polyacrylamide gel and visualized by ethidium bromide staining.
For real-time RT-PCR analysis, the synthesized first-strand cDNA was amplified in triplicate using SYBR Premix ExTaq (TaKaBaBio, Shiga, Japan), and the products were detected on a MiniOpticon system (Bio-Rad, San Diego, CA, USA). PCRs were incubated for 10 s at 95 °C followed by 45 amplification cycles with 3 s denaturing at 95 °C, 10 s annealing at 60 °C and 10 s extension at 72 °C. The primers designed for quantitative real-time RT-PCR analysis were as follows: for GRP78, 5′-GCGCATGAAGGAGAAAGAAC-3′ and 5′-TCACCATTCGGTCAATCAGA-3′ for ERdj4, 5′-AAAATAAGAGCCCGGATGCT-3′ and 5′-CGCTTCTTGGATCCAGTGTT-3′ for EDEM, 5′-TGGACTGCAGGTGCTGATAG-3′ and 5′-GGATTCTTGGTTGCCTGGTA-3′ and for GAPDH, 5′-AGGTCGGAGTCAACGGATTT-3′ and 5′-TAGTTGAGGTCAATGAAGGG-3′. Specificity of the PCR was evaluated by analyzing melting curves and sequences of the amplicons.
For the detection of spliced XBP1 isoform or all forms (spliced and unspliced), primer sets and Taqman probes were purchased from Applied Biosystems (Foster City, CA, USA). Quantitative PCR was performed using Taqman Gene Expression Assays and a StepOnePlus real-time PCR instrument according to the manufacturer's instructions.
Western blotting
Western blotting was performed as described.27 Primary antibodies used were: anti-KDEL from Enzo LifeScience (Farmingdale, NY, USA); anti-ATF6, anti-actin and anti-XBP1 from Santa Cruz Biotechnology; anti-phospho-eIF2α (Ser51) and anti-eIF2α and anti-IRE1α from Cell Signaling (Boston, MA, USA); anti-tubulin and anti-FLAG (M2) from Sigma (St Louis, MO, USA); anti-phospho-Ser724-IRE1α from Novus Biologicals (Littleton, CO, USA).
In vitro XBP1 mRNA cleavage assay
In vitro XBP1 mRNA cleavage assays were performed as described previously.28 Briefly, 337-nucleotide RNA substrate (XBP1(266-602) RNA) consisting of the XBP1 intron (26 nucleotides) flanked on both sides by truncated exon sequences (228 nucleotides on the 5′ side and 83 nucleotides on the 3′ side), which contained the minimum sequence for ER stress-induced XBP1 splicing, was prepared by in vitro transcription using T7 RNA polymerase. N-terminally FLAG-tagged human IRE1α(467-977) was prepared by immunoprecipitation with anti-FLAG antibody from 293T cells transiently transfected with pCAX-FLAG-IRE1α(467-977) plasmid. The IRE1α(467-977)-induced XBP1(206-602) mRNA cleavage reaction was performed in the presence of 100 μℳ ATP. RNA fragments were resolved on a 7 ℳ urea 6% PAGE gel and stained with ethidium bromide.
Apoptosis and cell proliferation assays
After exposure to toyocamycin or BTZ, apoptotic cells were evaluated using the Annexin V-FITC Apoptosis Detection Kit I (BD Pharmingen, Franklin Lakes, NJ, USA). The cell proliferation assay has been described previously.26 The percentage of specific apoptosis was calculated as follows: % specific apoptosis=(% AnnexinV-positive cells−% spontaneous AnnexinV-positive cells)/(100−% spontaneous positive cells) × 100.
Cell proliferation assays of the MM cell lines, primary MM cells from patients and peripheral blood mononuclear cell (PBMC) from healthy individuals exposed to various concentrations of toyocamycin for 24 h were performed using the CEllTiter 96 AQueous One Solution Cell Proliferation Assay Kit (Promega) as described previously.26 The mean of three determinations at each concentration of BTZ was calculated, and IC50 values were obtained using XLfit 4.2 curve fitting software for Excel (ID Business Solutions Inc., Alameda, CA, USA).
Animals and the murine xenograft model
Animal studies were performed in conformity with the UK Coordinating Committee on Cancer Research Guidelines for the Welfare of Animals in Experimental Neoplasia (second edition). The study method was described previously.29 Briefly, 0.5 × 107 RPMI8226 cells were inoculated subcutaneously into SCID mice previously administered with rabbit antiasialo-GM1 intraperitoneally (Wako Pure Chemical Industries, Osaka, Japan) 1 day before tumor inoculation. At 10 days after tumor inoculation, the tumor-bearing mice were divided into four groups of five mice each, such that the mean tumor volumes were approximately equal in the four groups. Tumor volume was calculated by the following formula: tumor volume (mm3)=0.5 × (major diameter) × (minor diameter)2. Mice were treated by intraperitoneal injection of 0.5 mg/kg toyocamycin twice weekly, 1.0 mg/kg toyocamycin once weekly, or 1.0 mg/kg BTZ twice weekly for 2 weeks. Volumes of tumors in toyocamycin-treated mice were compared with untreated or BTZ-treated animals during the treatment period.
Statistical analysis
The significance of differences in tumor volume between toyocamycin-treated mice and others were examined with the Mann–Whitney U-test. Data were analyzed with the aid of StatView software ver. 5.0 (SAS Institute, Cary, NC, USA). In this study, P<0.05 was considered significant.
Results
Identification of toyocamycin as an inhibitor of ER stress-induced XBP1 mRNA splicing
Previously, we established a screening system for inhibitors of ER stress-induced XBP1 activation and identified the new active small molecules trierixin and quinotrierixin.24, 30 In the course of further screening for an inhibitor of ER stress-induced XBP1 activation, we isolated toyocamycin25 from a culture broth of an Actinomycete strain (Figure 1a). As shown in Figure 1b, 0.1 μℳ thapsigargin, an inhibitor of the ER calcium pump (SERCA), elevated XBP1-luciferase activity about 2.5-fold more than the control in HeLa/XBP1-luc cells. Toyocamycin suppressed thapsigargin-induced XBP1-luciferase activation in a dose-dependent manner with an IC50 value of 0.08 μℳ. To examine whether toyocamycin also inhibited thapsigargin-induced endogenous XBP1 mRNA splicing in HeLa cells, we next performed RT-PCR analysis of RNA isolated from toyocamycin-treated or -untreated HeLa cells. As shown in Figure 1c, toyocamycin suppressed thapsigargin-induced XBP1 mRNA splicing in a dose-dependent manner with an IC50 value of 0.18 μℳ. Furthermore, toyocamycin also suppressed tunicamycin, a N-glycosylation inhibitor, or 2-deoxyglucose, a hypoglycemia-mimicking agent, -induced XBP1 mRNA splicing with an IC50 value of 0.13 μℳ and 0.11 μℳ, respectively (Figure 1c). These results show that toyocamycin inhibits ER stress-induced XBP1 mRNA splicing.
The inhibitory activity of toyocamycin on RNA synthesis is not responsible for inhibition of ER stress-induced XBP1 mRNA splicing
Because previous studies have reported that toyocamycin inhibits RNA synthesis in mammalian cells,31, 32 we tested whether its inhibitory activity on ER stress-induced XBP1 mRNA splicing was due to the inhibition of RNA synthesis. As shown in Figure 1d, toyocamycin suppressed incorporation of [3H]-uridine into the macromolecular fraction of HeLa cells in a dose-dependent manner with an IC50 value of 12 μℳ without affecting incorporation of [3H]-thymidine or [3H]-leucine. This indicates that toyocamycin inhibits RNA synthesis in HeLa cells. However, the IC50 value of toyocamycin for RNA synthesis is much higher than for ER stress-induced XBP1 mRNA splicing. On the other hand, actinomycin D, a well-known RNA synthesis inhibitor, also blocked the incorporation of [3H]-uridine into HeLa cells in a dose-dependent manner with an IC50 value of 0.06 μℳ. However, as shown in Figure 1e, actinomycin D did not suppress thapsigargin-induced XBP1 mRNA splicing even at 0.1 μℳ (which caused 80% inhibition of RNA synthesis). These results indicate that actinomycin D does not inhibit ER stress-induced XBP1 mRNA splicing. Therefore, we conclude that the inhibitory activity of toyocamycin on ER stress-induced XBP1 mRNA splicing is not due to the inhibition of RNA synthesis.
The adenosine moiety of toyocamycin is important for inhibition of XBP1 mRNA splicing
We next investigated whether sangivamycin or tubercidin, a small molecule structurally related to toyocamycin (Figure 1a), also inhibited ER stress-induced XBP1 activation. As shown in Table 1, sangivamycin and tubercidin inhibited thapsigargin-induced XBP1-luciferase activation in a dose-dependent manner with IC50 values of 0.5 and 0.34 μℳ, respectively. Furthermore, sangivamycin and tubercidin also inhibited thapsigargin-induced endogenous XBP1 mRNA splicing evaluated by RT-PCR analysis (data not shown). On the other hand, 5-Aza-2-deoxycytidine (5Aza-C), a cytidine analog, or 5-fluorouridine, a uridine analog, neither inhibited thapsigargin-induced XBP1-luciferase activation nor endogenous XBP1 mRNA splicing even at 100 μℳ (Table 1 and data not shown). These results suggest that the adenine moiety of toyocamycin is important for inhibition of XBP1 activation.
Table 1. The IC50 of inhibition of XBP1-luciferase activation when exposed to each drug.
| Small-molecule inhibitor | XBP1-luciferase inhibition (μM) |
|---|---|
| Toyocamycin | 0.08 |
| Sangivamycin | 0.5 |
| Tubecidin | 0.34 |
| 5-aza-2-deoxycytidine | >100 |
| 5-fluoro-uridine | >100 |
IC50 values were determined from the dose–response curves of the inhibition of XBP1-luciferase activity.
Toyocamycin selectively inhibits the IRE1α-XBP1 pathway
We next examined the effect of toyocamycin on the transcriptional activity of XBP1. Consistent with the regulation of transcription of EDEM and ERdj4 by the IRE1-XBP1 pathway,33, 34 toyocamycin suppressed tunicamycin-increased EDEM and ERdj4 mRNA with an IC50 of 0.079 and 0.172 μℳ, respectively (Figure 2a). These results suggest that toyocamycin might be inhibiting the transcriptional activity of XBP1 due to inhibition of ER stress-induced XBP1 mRNA splicing. On the other hand, because ER stress activates the three ER transmembrane proteins, IRE1, ATF6 and PERK, we also investigated whether toyocamycin inhibited ER stress-induced activation of ATF6 and PERK. ATF6 is constitutively synthesized as a type-II transmembrane protein in the ER.35 This membrane-bounded precursor form, designated as pATF6(P), is transported to the Golgi apparatus in response to ER stress, where it is cleaved by the sequential actions of Site-1 and Site-2 proteases.36, 37, 38, 39 The cytoplasmic region of ATF6 thus liberated from the membrane is translocated into the nucleus, where it functions as an active transcription factor. PERK is a serine–threonine protein kinase that phosphorylates eukaryotic initiation factor-2α (eIF2α) on Ser51.40, 41 Therefore, we assessed ATF6 activation and PERK activation by measuring the amount of the precursor form of ATF6 (pATF6(P)) and the phosphorylation of eIF2α on Ser51, respectively. As shown in Figure 2b, tunicamycin decreased the 90 kDa membrane-bound precursor form of ATF6 (pATF6(P)) and phosphorylated eIF2α on Ser51, indicating that it activated both ATF6 and PERK. However, toyocamycin inhibited neither the tunicamycin-induced decrease of pATF6(P) nor phosphorylation of eIF2α on Ser51. These results indicate that toyocamycin selectively inhibits the ER stress-induced activation of the IRE1α-XBP1 pathway.
Figure 2.
Toyocamycin selectively inhibits ER stress-induced activation of the IRE1α-XBP1 pathway and IRE1α-induced XBP1 mRNA cleavage. (a) Toyocamycin suppresses tunicamycin-induced EDEM and ERdj4 mRNA expression in HeLa cells. HeLa cells were incubated with the indicated concentration of toyocamycin in the presence or absence of 10 μg/ml tunicamycin for 4 h. Thereafter, the cells were collected and expression of EDEM and ERdj4 mRNA quantified by real-time RT-PCR as described in Materials and methods. Data represent the mean±s.d. of three experiments. (b) Toyocamycin suppresses neither tunicamycin-induced ATF6 nor PERK activation. HeLa cells were treated with the indicated concentration of toyocamycin in the presence or absence of 10 μg/ml of tunicamycin for 4 h. Thereafter, cells were collected and subjected to western blotting using anti-ATF6, anti-phospho-eIF2α, anti-eIF2α or anti-tubulin antibody. (c) Toyocamycin does not inhibit IRE1α phosphorylation on Ser724. HEK293T cells were transiently transfected with 1 μg of pcDNA3-IRE1α-flag plasmid. After 24 h, the cells were treated with the indicated concentration of toyocamycin for 4 h. The cells were collected and subjected to western blotting using anti-FLAG, anti-Ser724-IRE1α antibodies. Another set of the cells was collected and RNA was extracted. Spliced- or unspliced-XBP1 mRNA was detected as described in Materials and Methods. (d) Toyocamycin inhibits IRE1α-induced XBP1 mRNA cleavage in vitro. A 337-nucleotide RNA substrate (XBP1 (266-602) RNA) containing the XBP1 intron (26 nucleotides) was mixed with N-terminally FLAG-tagged human IRE1α(467-977) in the presence of 100 μℳ of ATP. RNA fragments were resolved on a 7 ℳ urea 6% PAGE gel and stained with ethidium bromide.
Toyocamycin inhibits IRE1α-induced XBP1 mRNA cleavage
We next investigated the mechanism by which toyocamycin inhibits ER stress-induced activation of the IRE1α-XBP1 pathway. Overexpression of IRE1α has been reported to lead to IRE1α homo-oligomerization, which occurs without accumulation of unfolded proteins in the ER or GRP78 dissociation from IRE1α.42, 43 Indeed, as shown in Figure 2c, overexpression of IRE1α induced Ser724 phosphorylation and XBP1 mRNA splicing even under normal conditions in 293T cells transiently transfected with pcDNA3-IRE1α-FLAG plasmid. Although toyocamycin did not affect the Ser724 phosphorylation of IRE1α, it inhibited IRE1α overexpression-induced XBP1 mRNA splicing in a dose-dependent manner. On the other hand, toyocamycin inhibited IRE1α (467-977)-induced XBP1 mRNA cleavage in vitro (Figure 2d). Thus, these results indicate that toyocamycin inhibits IRE1α-induced XBP1 mRNA cleavage.
Toyocamycin inhibits constitutive activation of XBP1 in MM cell lines
As shown in Figure 3a, most MM cell lines have activated XBP1 protein expression, represented as the overexpression of spliced XBP1 isoform, whereas non-MM cells including other hematological malignant and solid tumor cells have little activation of XBP1. This suggests that the IRE1α-XBP1 pathway is constitutively active in MM cells but not in non-MM cells. Among 10 MM cell lines, 7 MM cells including FR4, RPMI8226, XG7, OPM-2, ILKM2, ILKM8 and AMO1 express high levels of spliced-XBP1, whereas other 3 including U266, KMS-11 and SKMM1 express relatively lower levels.
Figure 3.
Toyocamycin inhibits constitutive activation of XBP1 in MM cells, and triggers marked apoptosis of MM cell lines. (a) XBP1 expression consisting of spliced and unspliced form was assessed by immunoblot analysis in 10 MM and 4 non-MM cell lines including 2 of hematological and 2 of solid tumor origin. (b) Reduced expression of spliced XBP1 isoform induced by various concentrations of toyocamycin in RMPI8226 cells detected by immunoblot. (c) RPMI8226 cells were treated with 0.1 μℳ of toyocamycin and the altered expression of IRE1α-XBP1 were evaluated by RT-PCR and immunoblot analysis. (d) The kinetic change of constitutive XBP1 expression in two MM cell lines during exposure to 0.1 μℳ toyocamycin. (e) Toyocamycin induced apoptosis in three MM cell lines, RPMI8226, XG7 and U266. Cells were treated with toyocamycin at the indicated concentration for 24 h and apoptotic cells were evaluated by double staining with Annexin V and PI. Data represent the mean+s.d. of three independent experiments.
We next tested the influence of toyocamycin on the constitutive activation of XBP1 in MM cell lines. After treatment with 10 nℳ or higher concentrations of toyocamycin, the levels of spliced isoform of XBP1 protein in RPMI8226 cells declined and resulted in caspase activation (Figure 3b). The level of spliced-XBP1 mRNA in RPMI8226 cells decreased at 6 h after exposure to 100 nℳ toyocamycin (Figure 3c). The protein levels of spliced-XBP1 also decreased by toyocamycin treatment, whereas phosphorylated levels of IRE1α remained unchanged. A similar reduction in the levels of spliced-XBP1 following toyocamycin treatment was also confirmed in two other MM cell lines, such as XG7 and U266, harboring high or low active XBP1 expression at baseline, respectively (Figure 3d). Furthermore, toyocamycin inhibited thapsigargin-induced expression of spliced XBP1 protein without affecting IRE1α phosphorylation on Ser724 (Supplementary Figure 3).
Toyocamycin induces marked apoptosis of MM cell lines harboring high levels of spliced-XBP1
Two MM cell lines with highly expressed spliced-XBP1, including RPMI8226 and XG7, showed robust dose-dependent apoptosis after exposure to various concentrations of toyocamycin for 24 h, as assessed by the number of Annexin V-positive cells (Figure 3e). These cells showed marked apoptosis at 30 nℳ or higher concentrations of toyocamycin, whereas U266 cells with relatively low spliced-XBP1 expression showed mild apoptosis compared with that of RPMI8226 and XG7.
MM cells with high spliced-XBP1 expression show marked growth inhibition compared with those with low spliced-XBP1 expression and to non-MM cells by toyocamycin treatment
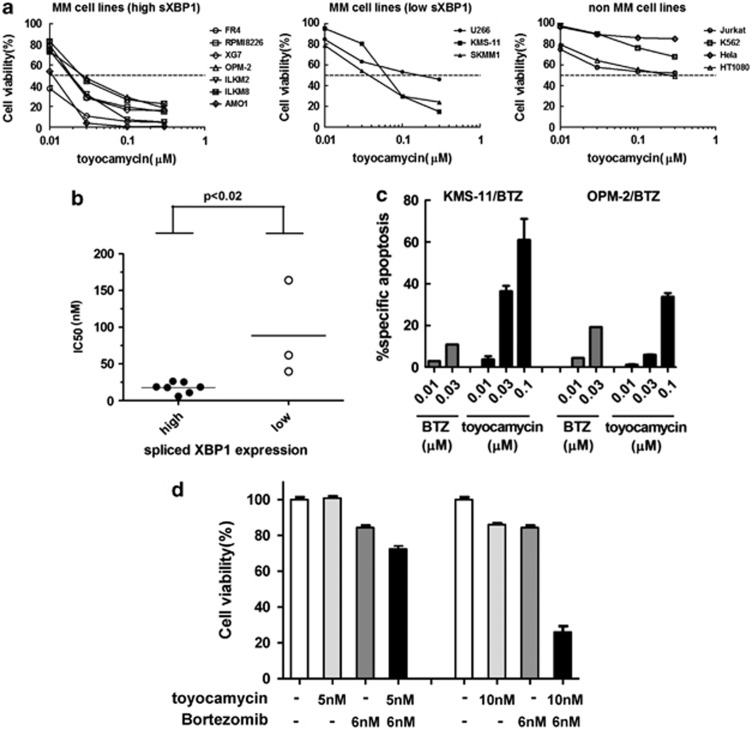
We then evaluated the growth inhibitory effect of toyocamycin on seven MM cell lines with high spliced-XBP1 expression, three MM cell lines with low spliced-XBP1 expression and four non-MM cell lines. All MM cells with high spliced-XBP1 expression showed remarkable decline in cellular viability at 30 nℳ or higher concentrations of toyocamycin (Figure 4a, left) than the other MM cells with low spliced-XBP1 expression (Figure 4a, middle). Most non-MM cells showed the subtle reduction in cellular viability even at the higher concentrations of toyocamycin (Figure 4a, right). The mean IC50 values of toyocamycin on two MM cell groups, each having high or low spliced-XBP1 expression were 17.69±2.78 and 88.57±38.31 nℳ, respectively, and this difference was considered statistically significant (P=0.016) (Figure 4b).
Figure 4.
Growth inhibitory effect of toyocamycin on MM cells with high or low expression of spliced XBP1 and on non-MM cells. (a) Growth inhibitory effect of toyocamycin exposure for 24 h on two groups of MM cells with high or low expression of spliced XBP1 and on non-MM cells. Data represent the mean of three independent experiments. (b) The IC50 values of each MM cell line after 24 h exposure to toyocamycin are calculated and plotted according to the high or low expression of XBP1. (c) Toyocamycin induced marked apoptosis of the BTZ-resistant MM cell lines, KMS-11/BTZ and OPM-2/BTZ. These two MM cells were treated, respectively, with BTZ or toyocamycin alone at the indicated concentrations for 24 h. Each column represents the mean+s.d. of three independent experiments. (d) RPMI8226 cells were exposed to either 5 or 10 nℳ of toyocamycin in the presence or absence of bortezomib (6 nℳ) for 72 h. Cell viability was measured by MTS assay. Data represent the mean±s.d. of three independent experiments.
Toyocamycin also induces marked apoptosis on BTZ-resistant MM cells
Two BTZ-resistant MM cell lines, KMS-11/BTZ and OPM-2/BTZ showed a little apoptosis progression even in the presence of 10 or 30 nℳ of BTZ, at which most MM cell lines including parental cells of these resistant ones were induced to marked apoptosis. However, they showed remarkable apoptosis in the presence of 30 or 100 nℳ of toyocamycin (Figure 4c).
Toyocamycin cooperates with BTZ to mediate anti-tumor effects on MM cells
At suboptimal concentrations toyocamycin mediated a little apoptosis induction (Figure 3e) in MM cells. However, treatment with 5 or 10 nℳ toyocamycin together with 6 nℳ BTZ-mediated additive or synergistic cytotoxicity in RPMI8226 cells (Figure 4d).
Toyocamycin is also cytotoxic for primary MM cells, associated with inhibition of IRE1α-XBP1 activation
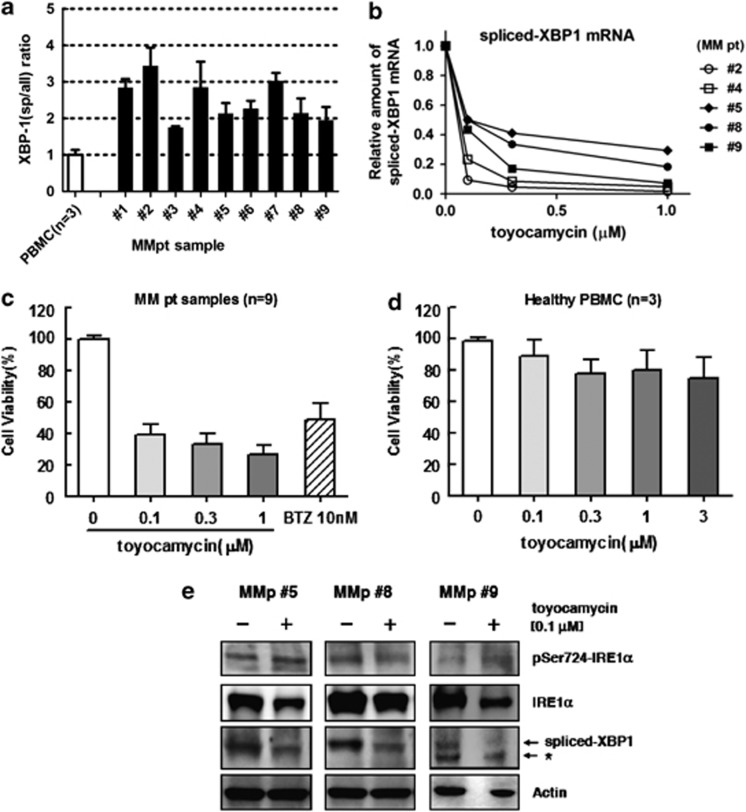
Most freshly prepared primary MM cells had activated XBP1 expression, represented as a high ratio of XBP1-spliced isoform: all isoforms (1.74:3.43; median 2.48), although the mRNA levels are similar to those in healthy donor control PBMC (Figure 5a). The relative amount of spliced-XBP1 mRNA in five MM samples (#2, #4, #5, #8 and #9), which was quantitated by real-time PCR method, decreased after 6 h exposure to toyocamycin (Figure 5b). Most of the primary MM cells were sensitive to 10 nℳ BTZ (17.1–106.8; median 48.8) and they also showed dose-dependent reduced viability following toyocamycin treatment (Figure 5c). Of nine MM samples, sample no. 4 was derived from the pleural effusion of a patient with MM refractory to BTZ; these tumor cells were resistant to BTZ in vitro. Nonetheless, toyocamycin was also cytotoxic for these cells (Supplementary Figure 4). In contrast, toyocamycin showed little cytotoxicity against healthy PBMC even at higher concentrations for 24 h (Figure 5d). Three primary MM samples (#5, #8 and #9) with sufficient cell numbers for protein analysis, showed reduction in activated (spliced) XBP1 protein after 6 h treatment with toyocamycin (Figure 5e). As in the MM cell lines, the phosphorylation status of IRE1α was not affected by toyocamycin.
Figure 5.
Sensitivity of primary MM cells to toyocamycin treatment. (a) The ratio of the spliced form of XBP1 to all forms (spliced and non-spliced) in nine primary MM cells and three healthy donors' PBMC. To compare the ratios among the 12 samples, the ratio in healthy PBMC was defined as the control. Each column represents the mean+s.d. of three independent experiments. (b) Altered amount of spliced-XBP1 mRNA after exposure to toyocamycin for 6 h at the indicated concentrations. The amount was quantitated by real-time PCR method. Data represent the mean of three independent experiments. (c) Analysis of cellular viability of primary MM cells (n=9) after exposure to toyocamycin or BTZ for 24 h at the indicated concentrations. The primary MM cells derived from eight patients were evaluated with mean+s.d. being plotted. (d) Analysis of cellular viability of PBMC (n=3) derived from healthy donors after exposure to toyocamycin for 24 h at the indicated concentrations. Each column represents the mean+s.d. of three independent experiments. (e) Altered expression levels of IRE1-XBP1 pathway-related molecules in primary MM cells on exposure to toyocamycin. The primary MM cells (#5, #8 and #9) were cultured in the presence of 0.1 μℳ toyocamycin for 6 h. Expression levels of spliced forms of XBP1, IRE1 and pSer724-phosphorylated IRE1 were evaluated by immunoblot analysis. *Depicts non-specific band.
In vivo anti-tumor activity of toyocamycin alone and in combination with BTZ in a human MM xenograft model
To evaluate in vivo efficacy of toyocamycin on MM cells, SCID mice subcutaneously inoculated with RPMI8226 were treated with twice- or once-weekly intraperitoneal toyocamycin at either 0.5 or 1.0 mg/kg. In addition, the combination treatment of toyocamycin with BTZ was tested. Toyocamycin alone showed robust anti-tumor activity resulting in smaller tumor volumes compared with controls on day 15. This was similar to the effect of BTZ (Figures 6a and b). No obvious difference in tumor inhibitory effect was seen on twice- or once-weekly injection of toyocamycin.
Figure 6.
Toyocamycin exerts anti-tumor activity in an in vivo MM model. (a) Average tumor volume in untreated mice (5% Traubenzucker alone) and mice treated with bortezomib alone, toyocamycin alone or with combination of bortezomib and toyocamycin at the indicated dose. (b) Representative images and mean values of tumor size observed at day 15 in each group; closed square, untreated; open square, bortezomib-treated (1.0 mg/kg) twice a week; open triangle, toyocamycin-treated (1.0 mg/kg) once a week; open circle, toyocamycin-treated (0.5 mg/kg) twice a week; closed triangle, toyocamycin (1.0 mg/kg, once a week)+bortezomib (1.0 mg/kg, twice a week)-treated; and closed circle, toyocamycin (0.5 mg/kg, twice a week)+ bortezomib (1.0 mg/kg, twice a week)-treated. *Represents statistically significant.
The combination treatment of BTZ with toyocamycin, either at 0.5 mg/kg or 1.0 mg/kg, showed a trend toward enhancing anti-tumor activity represented as smaller tumor volumes when compared with BTZ or toyocamycin alone (Figures 6a and b).
Discussion
Several lines of evidence suggest the importance of the IRE1α-XBP1 pathway in tumor progression, adaptation to the hypoxic tumor microenvironment, as a prognostic marker and a potential therapeutic target in both solid tumors and MM. Here, we identified toyocamycin as an inhibitor of both ER stress-induced and constitutive activation of the IRE1α-XBP1 pathway in MM cells, and showed that it exerted synergistic or at least additive anti-tumor effects with BTZ. Furthermore, it induced marked apoptosis of primary MM cells as well as MM cell lines without showing any cytotoxicity to PBMCs from healthy donors. This anti-tumor effect was also confirmed in a mouse model in vivo.
Previously, toyocamycin was reported to inhibit RNA synthesis and ribozyme function.31, 32, 44 This raised the possibility that it inhibited ER stress-induced XBP1 mRNA splicing and transcription of EDEM and ERdj4 genes through RNA synthesis inhibition. However, comparing the IC50 values of toyocamycin on RNA synthesis, ER stress-induced XBP1 mRNA splicing and transcription of UPR target genes revealed that its inhibitory activity on ER stress-induced XBP1 mRNA splicing and transcription of EDEM and ERdj4 genes was much stronger than the RNA synthesis blockade effect. Furthermore, although 100 nℳ actinomycin D completely inhibited both RNA synthesis (Figure 1d) and ER stress-induced transcription of GRP78 (Supplementary Figure 1), it did not inhibit ER stress-induced XBP1 mRNA splicing (Figure 2b). These results support the notion that the inhibitory activity of toyocamycin on RNA synthesis is not responsible for inhibition of XBP1 mRNA splicing and transcription of EDEM and ERdj4 genes. On the other hand, toyocamycin has also been reported to inhibit kinase activities, such as PKC,45 cdc246 or PI4K.47 Considering the structure of toyocamycin and its analogs, it was also predicted that it would inhibit IRE1 auto-phosphorylation. However, it inhibited IRE1 phosphorylation on Ser724 not only in IRE1-overexpressing 293T cells (Figure 2c) but also in MM cell lines (Figures 3c and 5e). Recent studies have suggested that the trigger for IRE1 endoribonuclease activity is not phosphorylation but a conformational change in the kinase domain induced by cofactor (ATP or ADP) binding. Therefore, it is likely that toyocamycin inhibits IRE1α-induced XBP1 mRNA cleavage through a cofactor-induced conformational change of IRE1α rather than inhibition of IRE1α auto-phosphorylation.
Previous studies have suggested that the IRE1α-XBP1 pathway has a critical role in ER stress-induced cytoprotection. 1NM-PP1, a small molecule selectively activating the IRE1α(I642G) mutant,18 protected cells from tunicamycin- or thapsigargin-induced cell death.48, 49 In contrast, overexpression of dominant-negative XBP1 or knockdown of XBP1 has been reported to enhance tunicamycin-induced apoptosis.50 Consistent with previous reports, we also found that toyocamycin synergistically induced cell death in ER-stressed HeLa, HT29 and HCT116 cells (Supplementary Figure 2 and data not shown). More recently, as with toyocamycin, STF-083010 was shown to inhibit IRE1 ndonuclease activity without affecting its kinase activity in vitro.14 However, these compounds mediate their inhibitory activity at ⩾60 μℳ, and show little MM cell apoptosis induction as single agents. STF-083010 also shows anti-tumor activity in human MM xenograft models. However, while toyocamycin suppressed tumor volume to around 50% at 1 mg/kg by once-weekly injection, the STF-083010 dose needed to be 30 mg/kg by once-weekly injection.14 In our study, adenosine analogs showed potent inhibition of ER stress-induced IRE1α-XBP1 activation at the nanomolar level. Compared with these, toyocamycin induced marked apoptosis in ER-stressed tumor cells and MM cells at much lower concentrations. In addition, it also inhibited the constitutive activation of XBP1 in MM cells even at suboptimal concentrations such as 10 nℳ. However, the mechanism by which toyocamycin mediates dose-dependent apoptosis remains unknown. Although three MM cell lines with low active XBP1 expression showed lower sensitivity to toyocamycin treatment than other seven MM cells with high active XBP1 expression, these three cell lines still demonstrated sensitivity to the drug at the nM concentration. It may be speculated that toyocamycin also triggers another stress-inducing factor at higher concentrations, which may induce strong apoptosis under IRE1α-XBP1-suppressed conditions. Further analysis needs be conducted to elucidate the whole picture of its mechanisms of action on MM cells.
An earlier phase I toyocamycin single-agent study also testing possible anti-tumor effects in patients with advanced solid tumors has been reported.51 However, because no apparent clinical responses were observed in that study, further clinical evaluation was not planned. In that study, toyocamycin showed no systemic side effects, such as organ dysfunction and cytopenia, and only local necrosis at the site of infusion was reported to occur when the drug was delivered into the soft tissues. This suggests that toyocamycin adverse events could be manageable if it is infused through central venous catheters. In addition, this study does not exclude potential clinical efficacy of toyocamycin against solid tumors, because it was a phase I trial lacking evaluation of stable disease often applied in more recent clinical trials of molecular-targeting therapies.
In conclusion, we demonstrated that the adenosine analog toyocamycin has a potent IRE1-XBP1 inhibitory effect on ER-stressed tumors and MM cells, as well as triggering dose-dependent apoptosis in these cells. These results provide a preclinical rationale for clinical trials of toyocamycin and other adenosine analogs alone and in combination with BTZ for treating MM.
Acknowledgments
We thank Dr Masayuki Igarashi, Dr Masaki Hatano, Mrs Naoko Kinoshita and Dr Yoshio Nishimura (Institute of Microbial Chemistry, Tokyo, Japan) for fermentation of Actinomycete strain MK653-101F13, and Ms Chiori Fukuyama (Nagoya City University) for her skillful technical assistance. This study was partly supported by grants from Takeda Science Foundation. This work was partly supported by a Grant-in-Aid for Scientific Research from the Ministry of Education, Culture, Sports, Science and Technology (23791805) and a Grant-in-Aid for Cancer Research from the Ministry of Health, Labor and Welfare (21-8-5), Japan.
Y Shiotsu is an employee of Kyowa Hakko Kirin Co., Ltd, Japan. S Iida received research funding from Kyowa Hakko Kirin and Chugai Pharmaceutical Co., Ltd. S Iida declares honoraria from Janssen Pharmaceutical KK. The other authors declare no conflict of interest.
Footnotes
Supplementary Information accompanies the paper on Blood Cancer Journal website (http://www.nature.com/bcj)
Supplementary Material
References
- Kumar S, Rajkumar SV. Many facets of bortezomib resistance/susceptibility. Blood. 2008;112:2177–2178. doi: 10.1182/blood-2008-07-167767. [DOI] [PubMed] [Google Scholar]
- Shah JJ, Orlowski RZ. Proteasome inhibitors in the treatment of multiple myeloma. Leukemia. 2009;23:1964–1979. doi: 10.1038/leu.2009.173. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reimold AM, Iwakoshi NN, Manis J, Vallabhajosyula P, Szomolanyi-Tsuda E, Gravallese EM, et al. Plasma cell differentiation requires the transcription factor XBP-1. Nature. 2001;412:300–307. doi: 10.1038/35085509. [DOI] [PubMed] [Google Scholar]
- Shaffer AL, Lin KI, Kuo TC, Yu X, Hurt EM, Rosenwald A, et al. Blimp-1 orchestrates plasma cell differentiation by extinguishing the mature B cell gene expression program. Immunity. 2002;17:51–62. doi: 10.1016/s1074-7613(02)00335-7. [DOI] [PubMed] [Google Scholar]
- Munshi NC, Hideshima T, Carrasco D, Shammas M, Auclair D, Davies F, et al. Identification of genes modulated in multiple myeloma using genetically identical twin samples. Blood. 2004;103:1799–1806. doi: 10.1182/blood-2003-02-0402. [DOI] [PubMed] [Google Scholar]
- Carrasco DR, Sukhdeo K, Protopopova M, Sinha R, Enos M, Carrasco DE, et al. The differentiation and stress response factor XBP-1 drives multiple myeloma pathogenesis. Cancer Cell. 2007;11:349–360. doi: 10.1016/j.ccr.2007.02.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shuda M, Kondoh N, Imazeki N, Tanaka K, Okada T, Mori K, et al. Activation of the ATF6, XBP1 and grp78 genes in human hepatocellular carcinoma: a possible involvement of the ER stress pathway in hepatocarcinogenesis. J Hepatol. 2003;38:605–614. doi: 10.1016/s0168-8278(03)00029-1. [DOI] [PubMed] [Google Scholar]
- Lee K, Tirasophon W, Shen X, Michalak M, Prywes R, Okada T, et al. IRE1-mediated unconventional mRNA splicing and S2P-mediated ATF6 cleavage merge to regulate XBP1 in signaling the unfolded protein response. Genes Dev. 2002;16:452–466. doi: 10.1101/gad.964702. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Asosingh K, De Raeve H, de Ridder M, Storme GA, Willems A, Van Riet I, et al. Role of the hypoxic bone marrow microenvironment in 5T2MM murine myeloma tumor progression. Haematologica. 2005;90:810–817. [PubMed] [Google Scholar]
- Martin SK, Diamond P, Gronthos S, Peet DJ, Zannettino AC. The emerging role of hypoxia, HIF-1 and HIF-2 in multiple myeloma. Leukemia. 2011;25:1533–1542. doi: 10.1038/leu.2011.122. [DOI] [PubMed] [Google Scholar]
- Smith KD, Wrenshall LE, Nicosia RF, Pichler R, Marsh CL, Alpers CE, et al. Delayed graft function and cast nephropathy associated with tacrolimus plus rapamycin use. J Am Soc Nephrol. 2003;14:1037–1045. doi: 10.1097/01.asn.0000057542.86377.5a. [DOI] [PubMed] [Google Scholar]
- Bagratuni T, Wu P, Gonzalez de Castro D, Davenport EL, Dickens NJ, Walker BA, et al. XBP1s levels are implicated in the biology and outcome of myeloma mediating different clinical outcomes to thalidomide-based treatments. Blood. 2010;116:250–253. doi: 10.1182/blood-2010-01-263236. [DOI] [PubMed] [Google Scholar]
- Bae J, Carrasco R, Lee AH, Prabhala R, Tai YT, Anderson KC, et al. Identification of novel myeloma-specific XBP1 peptides able to generate cytotoxic T lymphocytes: a potential therapeutic application in multiple myeloma. Leukemia. 2011;25:1610–1619. doi: 10.1038/leu.2011.120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Papandreou I, Denko NC, Olson M, Van Melckebeke H, Lust S, Tam A, et al. Identification of an Ire1alpha endonuclease specific inhibitor with cytotoxic activity against human multiple myeloma. Blood. 2011;117:1311–1314. doi: 10.1182/blood-2010-08-303099. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bertolotti A, Zhang Y, Hendershot LM, Harding HP, Ron D. Dynamic interaction of BiP and ER stress transducers in the unfolded-protein response. Nat Cell Biol. 2000;2:326–332. doi: 10.1038/35014014. [DOI] [PubMed] [Google Scholar]
- Korennykh AV, Egea PF, Korostelev AA, Finer-Moore J, Zhang C, Shokat KM, et al. The unfolded protein response signals through high-order assembly of Ire1. Nature. 2009;457:687–693. doi: 10.1038/nature07661. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shamu CE, Walter P. Oligomerization and phosphorylation of the Ire1p kinase during intracellular signaling from the endoplasmic reticulum to the nucleus. EMBO J. 1996;15:3028–3039. [PMC free article] [PubMed] [Google Scholar]
- Papa FR, Zhang C, Shokat K, Walter P. Bypassing a kinase activity with an ATP-competitive drug. Science. 2003;302:1533–1537. doi: 10.1126/science.1090031. [DOI] [PubMed] [Google Scholar]
- Aragon T, van Anken E, Pincus D, Serafimova IM, Korennykh AV, Rubio CA, et al. Messenger RNA targeting to endoplasmic reticulum stress signalling sites. Nature. 2009;457:736–740. doi: 10.1038/nature07641. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sidrauski C, Walter P. The transmembrane kinase Ire1p is a site-specific endonuclease that initiates mRNA splicing in the unfolded protein response. Cell. 1997;90:1031–1039. doi: 10.1016/s0092-8674(00)80369-4. [DOI] [PubMed] [Google Scholar]
- Lee KP, Dey M, Neculai D, Cao C, Dever TE, Sicheri F. Structure of the dual enzyme Ire1 reveals the basis for catalysis and regulation in nonconventional RNA splicing. Cell. 2008;132:89–100. doi: 10.1016/j.cell.2007.10.057. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shen X, Ellis RE, Lee K, Liu CY, Yang K, Solomon A, et al. Complementary signaling pathways regulate the unfolded protein response and are required for C. elegans development. Cell. 2001;107:893–903. doi: 10.1016/s0092-8674(01)00612-2. [DOI] [PubMed] [Google Scholar]
- Yoshida H, Matsui T, Yamamoto A, Okada T, Mori K. XBP1 mRNA is induced by ATF6 and spliced by IRE1 in response to ER stress to produce a highly active transcription factor. Cell. 2001;107:881–891. doi: 10.1016/s0092-8674(01)00611-0. [DOI] [PubMed] [Google Scholar]
- Tashiro E, Hironiwa N, Kitagawa M, Futamura Y, Suzuki S, Nishio M, et al. Trierixin, a novel Inhibitor of ER stress-induced XBP1 activation from Streptomyces sp. 1. Taxonomy, fermentation, isolation and biological activities. J Antibiot. 2007;60:547–553. doi: 10.1038/ja.2007.69. [DOI] [PubMed] [Google Scholar]
- Nishimura H, Katagiri K, Sato K, Mayama M, Shimaoka N. Toyocamycin, a new anti-candida antibiotics. J Antibiot. 1956;9:60–62. [PubMed] [Google Scholar]
- Ri M, Iida S, Nakashima T, Miyazaki H, Mori F, Ito A, et al. Bortezomib-resistant myeloma cell lines: a role for mutated PSMB5 in preventing the accumulation of unfolded proteins and fatal ER stress. Leukemia. 2010;24:1506–1512. doi: 10.1038/leu.2010.137. [DOI] [PubMed] [Google Scholar]
- Tashiro E, Maruki H, Minato Y, Doki Y, Weinstein IB, Imoto M. Overexpression of cyclin D1 contributes to malignancy by up-regulation of fibroblast growth factor receptor 1 via the pRB/E2F pathway. Cancer Res. 2003;63:424–431. [PubMed] [Google Scholar]
- Iwawaki T, Akai R. Analysis of the XBP1 splicing mechanism using endoplasmic reticulum stress-indicators. Biochem Biophys Res Commun. 2006;350:709–715. doi: 10.1016/j.bbrc.2006.09.100. [DOI] [PubMed] [Google Scholar]
- Yano H, Kayukawa S, Iida S, Nakagawa C, Oguri T, Sanda T, et al. Overexpression of carboxylesterase-2 results in enhanced efficacy of topoisomerase I inhibitor, irinotecan (CPT-11), for multiple myeloma. Cancer Sci. 2008;99:2309–2314. doi: 10.1111/j.1349-7006.2008.00936.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kawamura T, Tashiro E, Yamamoto K, Shindo K, Imoto M. SAR study of a novel triene-ansamycin group compound, quinotrierixin, and related compounds, as inhibitors of ER stress-induced XBP1 activation. J Antibiot. 2008;61:303–311. doi: 10.1038/ja.2008.43. [DOI] [PubMed] [Google Scholar]
- Suhadolnik RJ, Uematsu T, Uematsu H. Toyocamycin: phosphorylation and incorporation into RNA and DNA and the biochemical properties of the triphosphate. Biochim Biophys Acta. 1967;149:41–49. doi: 10.1016/0005-2787(67)90689-2. [DOI] [PubMed] [Google Scholar]
- Tavitian A, Uretsky SC, Acs G. The effect of toyocamycin on cellular RNA synthesis. Biochim Biophys Acta. 1969;179:50–57. doi: 10.1016/0005-2787(69)90121-x. [DOI] [PubMed] [Google Scholar]
- Yoshida H, Matsui T, Hosokawa N, Kaufman RJ, Nagata K, Mori K. A time-dependent phase shift in the mammalian unfolded protein response. Dev Cell. 2003;4:265–271. doi: 10.1016/s1534-5807(03)00022-4. [DOI] [PubMed] [Google Scholar]
- Lee AH, Iwakoshi NN, Glimcher LH. XBP-1 regulates a subset of endoplasmic reticulum resident chaperone genes in the unfolded protein response. Mol Cell Biol. 2003;23:7448–7459. doi: 10.1128/MCB.23.21.7448-7459.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Haze K, Yoshida H, Yanagi H, Yura T, Mori K. Mammalian transcription factor ATF6 is synthesized as a transmembrane protein and activated by proteolysis in response to endoplasmic reticulum stress. Mol Biol Cell. 1999;10:3787–3799. doi: 10.1091/mbc.10.11.3787. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ye J, Rawson RB, Komuro R, Chen X, Dave UP, Prywes R, et al. ER stress induces cleavage of membrane-bound ATF6 by the same proteases that process SREBPs. Mol Cell. 2000;6:1355–1364. doi: 10.1016/s1097-2765(00)00133-7. [DOI] [PubMed] [Google Scholar]
- Shen J, Chen X, Hendershot L, Prywes RER. stress regulation of ATF6 localization by dissociation of BiP/GRP78 binding and unmasking of Golgi localization signals. Dev Cell. 2002;3:99–111. doi: 10.1016/s1534-5807(02)00203-4. [DOI] [PubMed] [Google Scholar]
- Okada T, Haze K, Nadanaka S, Yoshida H, Seidah NG, Hirano Y, et al. A serine protease inhibitor prevents endoplasmic reticulum stress-induced cleavage but not transport of the membrane-bound transcription factor ATF6. J Biol Chem. 2003;278:31024–31032. doi: 10.1074/jbc.M300923200. [DOI] [PubMed] [Google Scholar]
- Nadanaka S, Yoshida H, Kano F, Murata M, Mori K. Activation of mammalian unfolded protein response is compatible with the quality control system operating in the endoplasmic reticulum. Mol Biol Cell. 2004;15:2537–2548. doi: 10.1091/mbc.E03-09-0693. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harding HP, Novoa I, Zhang Y, Zeng H, Wek R, Schapira M, et al. Regulated translation initiation controls stress-induced gene expression in mammalian cells. Mol Cell. 2000;6:1099–1108. doi: 10.1016/s1097-2765(00)00108-8. [DOI] [PubMed] [Google Scholar]
- Harding HP, Zhang Y, Ron D. Protein translation and folding are coupled by an endoplasmic-reticulum-resident kinase. Nature. 1999;397:271–274. doi: 10.1038/16729. [DOI] [PubMed] [Google Scholar]
- Tirasophon W, Lee K, Callaghan B, Welihinda A, Kaufman RJ. The endoribonuclease activity of mammalian IRE1 autoregulates its mRNA and is required for the unfolded protein response. Genes Dev. 2000;14:2725–2736. doi: 10.1101/gad.839400. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhou J, Liu CY, Back SH, Clark RL, Peisach D, Xu Z, et al. The crystal structure of human IRE1 luminal domain reveals a conserved dimerization interface required for activation of the unfolded protein response. Proc Natl Acad Sci USA. 2006;103:14343–14348. doi: 10.1073/pnas.0606480103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yen L, Svendsen J, Lee JS, Gray JT, Magnier M, Baba T, et al. Exogenous control of mammalian gene expression through modulation of RNA self-cleavage. Nature. 2004;431:471–476. doi: 10.1038/nature02844. [DOI] [PubMed] [Google Scholar]
- Osada H, Sonoda T, Tsunoda K, Isono K. A new biological role of sangivamycin; inhibition of protein kinases. J Antibiot. 1989;42:102–106. doi: 10.7164/antibiotics.42.102. [DOI] [PubMed] [Google Scholar]
- Osada H, Cui CB, Onose R, Hanaoka F. Screening of cell cycle inhibitors from microbial metabolites by a bioassay using a mouse cdc2 mutant cell line, tsFT210. Bioorg Med Chem. 1997;5:193–203. doi: 10.1016/s0968-0896(96)00207-6. [DOI] [PubMed] [Google Scholar]
- Nishioka H, Sawa T, Hamada M, Shimura N, Imoto M, Umezawa K. Inhibition of phosphatidylinositol kinase by toyocamycin. J Antibiot. 1990;43:1586–1589. doi: 10.7164/antibiotics.43.1586. [DOI] [PubMed] [Google Scholar]
- Lin JH, Li H, Yasumura D, Cohen HR, Zhang C, Panning B, et al. IRE1 signaling affects cell fate during the unfolded protein response. Science. 2007;318:944–949. doi: 10.1126/science.1146361. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Han D, Upton JP, Hagen A, Callahan J, Oakes SA, Papa FR. A kinase inhibitor activates the IRE1alpha RNase to confer cytoprotection against ER stress. Biochem Biophys Res Commun. 2008;365:777–783. doi: 10.1016/j.bbrc.2007.11.040. [DOI] [PubMed] [Google Scholar]
- Lee AH, Iwakoshi NN, Anderson KC, Glimcher LH. Proteasome inhibitors disrupt the unfolded protein response in myeloma cells. Proc Natl Acad Sci USA. 2003;100:9946–9951. doi: 10.1073/pnas.1334037100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wilson WL. Phase I study with toyocamycin (NSC-63701) Cancer Chemother Rep. 1968;52:301–303. [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.