Abstract
The mechanisms of the age-associated exponential increase in the incidence of leukemia are not known in detail. Leukemia as well as aging are initiated and regulated in multi-factorial fashion by cell-intrinsic and extrinsic factors. The role of aging of the microenvironment for leukemia initiation/progression has not been investigated in great detail so far. Clonality in hematopoiesis is tightly linked to the initiation of leukemia. Based on a retroviral-insertion mutagenesis approach to generate primitive hematopoietic cells with an intrinsic potential for clonal expansion, we determined clonality of transduced hematopoietic progenitor cells (HPCs) exposed to a young or aged microenvironment in vivo. While HPCs displayed primarily oligo-clonality within a young microenvironment, aged animals transplanted with identical pool of cells displayed reduced clonality within transduced HPCs. Our data show that an aged niche exerts a distinct selection pressure on dominant HPC-clones thus facilitating the transition to mono-clonality, which might be one underlying cause for the increased age-associated incidence of leukemia.
Introduction
Aging is a major risk factor for leukemia. The mechanisms of the age-associated exponential increase in the incidence of leukemia are not known in detail, but it is widely accepted that the combination of leukemia initiating cell-intrinsic as well as extrinsic mechanisms drive this process. [1]–[5] While the contribution of hematopoietic cell intrinsic mechanisms to the increase in leukemia with age have been described [6], [7], whether aging of the bone marrow (BM) microenvironment also influences pre-leukemic states or leukemia has not been studied in great detail so far. [8], [9] The marrow microenvironment is able to influence differentiation [10], drug resistance [11] and proliferation of leukemic cells in diverse mouse leukemia-models [12], [13] while it undergoes multiple changes upon aging as demonstrated by decreased bone-remodeling [14], enhanced adipogenesis and changes in ECM components [15], [16].
Clonality is tightly linked to initiation of leukemia. [17] We consequently tested whether the age of the microenvironment influences in vivo clonality among dominant hematopoietic progenitor cells obtained via retroviral insertional mutagenesis. In summary our data support the novel concept that an aged niche exerts a distinct selection pressure on dominant HPC-clones that eases the transition to monoclonality, which might be one underlying cause for the increased age-associated incidence of leukemia.
Results and Discussion
To test whether an aged microenvironment influences in vivo clonality, BM depleted of mature cells (lin-) were transduced with the SF91/IRES-eGFP replication incompetent retrovirus, a set-up similar to insertional mutagenesis screens. The SF91 virus confers a high potential to generate in vivo dominant HPSCs by activation of proto-oncogenes in the neighborhood of the retroviral integration site (RIS), while the frequency of malignant transformation remains low [18], [19].
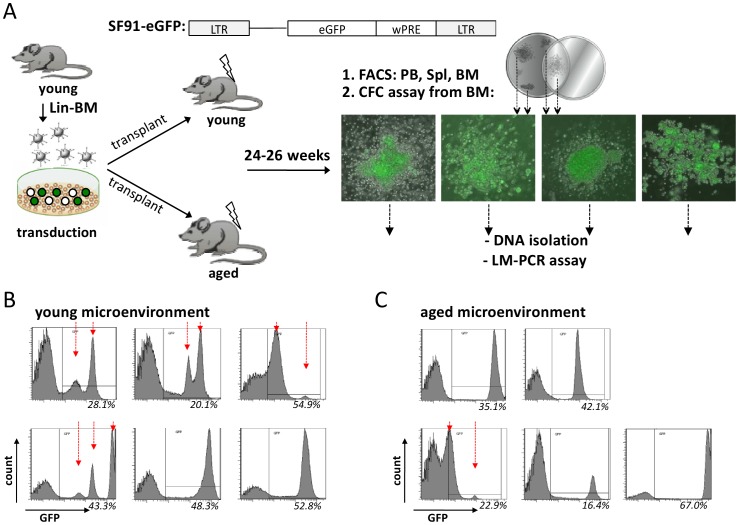
Pools of cells from the same transduction procedure were subsequently transplanted into young (2 month) and aged (18–19 months) recipient mice and GFP chimerism in peripheral blood was monitored by flow cytometry to follow the fate of the transduced cell population in vivo. 24–26 weeks post transplantation, when hematopoiesis is driven by the transplanted hematopoietic stem cells, GFP+ (transduced) cells in PB, spleen and BM were analyzed in detail (Figure 1A). This experimental set-up combined two goals: (1) The determination of clonality was possible as HSCs were individually marked by the random nature of the genomic site of the retroviral-integration. (2) Integration of this vector elicits retroviral insertional mutagenesis, and the transduced cells are prone to form dominant clones, serving as a model system for the initiation of leukemogenesis.
Figure 1. The clonal composition of the hematopoiesis is different in old compared to young microenvironment. (A).
Schematic representation of the gammaretroviral SF91/IRES-eGFP vector used for insertional mutagenesis and the experimental setup (LTR: long terminal repeat with strong enhancer element, wPRE: woodchuck hepatitis virus posttranscriptional regulatory element). Lineage depleted (lin-) BM cells from young C57BL/6 mice were pre-stimulated with a cytokine cocktail and transduced. Cells were subsequently transplanted in equal amounts into young and aged recipient mice. (The graft contained 51.6%, 30.7%, 32.4% GFP+ cells in the 3 transduction/transplantation.) 24–26 week post-transplantation recipients were sacrificed and PB, spleen and BM analyzed by flow cytometry for lineage differentiation markers as well as the number of primitive L-S+K+ cells. At the same time CFC-assays on BM cells were performed and DNA from GFP+ colonies (representative pictures) isolated for LM-PCR. GFP expression among BM cells of young (B) and aged (C) recipient mice. (young mice n = 6, aged mice n = 5, from 3 independent experiments).
The BM of the recipient mice showed distinct contribution of GFP+ cells, which might be due to the distinct influence of the viral integration site on HSCs survival as well as the contribution of the niche and microenvironment on HSCs behavior. In order to analyze the effect of a young or aged BM niche on clonality within the transduced and transplanted cells, we focused on animals with more than 15% GFP+ chimerism in bone marrow to perform ligation-mediated PCR (LM-PCR) to determine both integration sites as well as clonality. The 15% threshold for eGFP+ cells was set because at this level of chimerism mutagenesis potentially resulted in a survival advantage of the transduced HSC leading to proliferation/expansion as the first step of pre-leukemic stage, thus fulfilling a basic requirement for our model.
Although the GFP profile of the BM sample has the limitation that it represents differentiated cells and not the progenitor/stem cell population, it can serve as at least an indicator of clonality within differentiated cell populations in hematopoiesis. The GFP expression profile of transduced cells harvested from recipient’s BM revealed that the majority of young recipients presented with multiple GFP peaks (Figure 1B) while transduced cells in the aged microenvironment displayed primarily only one GFP peak (Figure 1C), suggesting differences in clonality within hematopoiesis in an old compared to a young microenvironment.
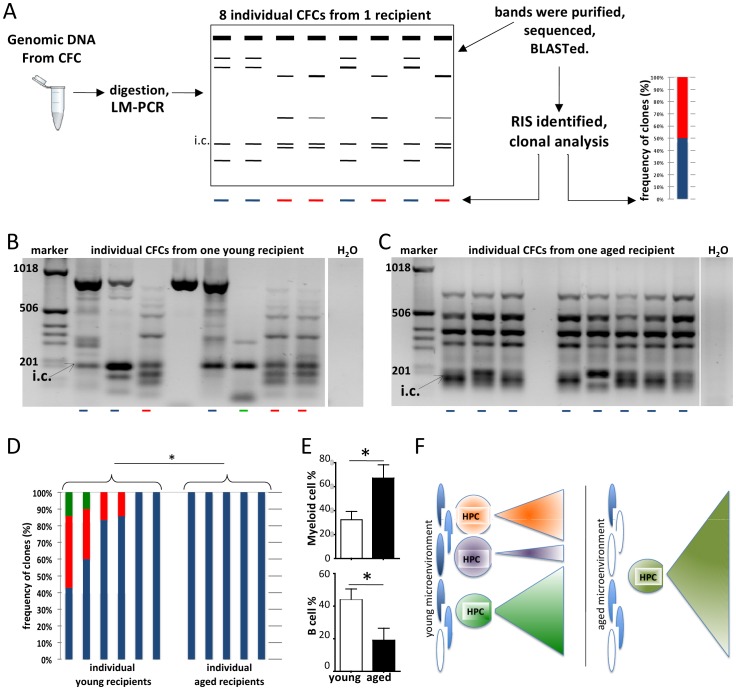
To further investigate clonality in more primitive hematopoietic cells exposed to a young or aged microenvironment, retroviral insertion sites (RISs) from individual GFP+ colony forming cells (CFCs) from BM were amplified by ligation-mediated PCR and genomic location of RISs were determined (Figure 1A). The gel-band pattern of the LM-PCR reaction, in combination with the sequence information from distinct bands were used to determine the frequency of distinct clones in the myeloid progenitor-cell compartment (Figure 2A). As the same pool of transduced graft cells were transplanted into young and aged animals, differences in the two experimental groups in hematopoiesis result primarily from the distinct age of the microenvironment. In young mice, hematopoiesis within the GFP+CFCs remained in the majority oligoclonal as indicated by distinct banding patterns among individual CFC-clones and confirmed by sequencing (Figure 2B,D). GFP+CFCs from individual aged recipients presented under the LM-PCR protocol used without exception a monoclonal pattern (Figure 2C,D). The difference in the clonality pattern among the myeloid stem/progenitor population between young and aged animals was significant (p = 0.035). The LM-PCR method was sensitive enough to demonstrate minor clone contributing down to least 10% of the transduced population (Figure 2D), while the restriction enzyme used for LM-PCR analysis will cover at least 85% of all possible integration sites [20], implying that the observed mono-clonality in CFC colonies might represent in a good number of cases true monoclonality in vivo.
Figure 2. Coexistence of balanced HSC populations is characteristic for young microenvironment whereas an aged microenvironment favors the expansion of single dominant hematopoietic clone.
(A) Schematic representation of the retroviral insertional mutagenesis screen. DNA was extracted from individual CFCs (average: 7.3±1.8 GFP+CFCs/mouse were isolated) and digested with four-cutter enzymes. After LM-PCR, the recovered bands were isolated and sequenced. The sequences were aligned to the mouse genome with NCBI/BLAST program. The RISs were identified and the closest genes (within a ±100 kb window) were listed. Based on the LM-PCR band-pattern and the identified RISs, clonal analyses were performed. (B) Representative agarose gel analysis of LM-PCR performed on methylcellulose colonies isolated from BM of one representative young transplanted mouse. Clonality was identified based on the band pattern and DNA sequence information on individual bands. Distinct colors represent distinct clones. (C) Insertion sites recovered by LM-PCR from CFCs of one representative aged recipient mouse. The red arrows depict distinct GFP peaks. i.c.: internal control bands represent PCR products amplified from the viral genome. (D) Distribution of clonality within the GFP+ CFC population of young and aged recipient mice determined by the LM-PCR pattern and sequence information. (E) Flow cytometric analyses of PB of young and aged recipient mice (same mice were used in clonal analysis and RISs identification). Myeloid cells were identified based on Gr1 and Mac1 expression and B220 was used as B cell marker. (young mice n = 6, aged mice n = 5, from 3 independent experiments), * = p<0.05 (F) Proposed model for clonality of hematopoiesis in young and aged microenvironment.
All animals transplanted in this study remained leukemia-free at least up to 6 month post-transplantation as indicated by normal spleen weight (Figure S1) and normal PB white blood cell counts (data not shown) in both age, supporting that the frequency of malignant transformation in response to transduction with SF91 is low, while at the same time generating dominant clones. As the chimerism for GFP positivity in PB, in lineage+ BM (differentiated) cells as well as in the primitive hematopoietic cell compartment (L-S+K+) and the GFP+CFC% in methylcellulose culture were similar in young and aged recipients, and the overall total number of L-S+K+ (GFP+ and GFP-) cells was similar in an aged or young niche (Figure S1), the difference in clonality between young and aged recipients can not be a consequence of an assumed distinct levels of chimerism of transduced/untransduced cells in young and aged recipients. Our data indicates that the same pool of cells, when exposed to an aged instead of a young microenvironment resulted in significantly reduced clonality, ultimately approaching mono-clonality in an aged microenvironment.
The genomic locations of RISs and the nearest neighboring genes within 100 kb of the RIS in the individual CFCs are listed in Table 1 and Table S1. Overall, more RISs were recovered from the oligoclonal young recipient group, because more clones were present in this group than in the monoclonal aged recipient group. As depicted in figure 2A and B, some clones contained more than one RIS and our technique was able to distinguish which RISs belong to individual clones. Genes in close vicinity of the RISs, which are frequently involved in the formation of dominant clones [19] were distinct between CFCs from young and aged recipients, and DAVID gene ontology [21] clustering also revealed distinct GO categories for RISs listed in a young or an aged microenvironment. These data further support that distinct set of cells/CFCs were selected in the two different (young/aged) microenvironments.
Table 1. Insertion sites in CFCs isolated from young and aged recipient mice.
| In young microenvironment | In old microenvironment | ||||
| Gene ID | Official gene symbol | Gene ID | Official gene symbol | ||
| Endocytosis GO cluster: | Regulation of transcription GO cluster: | ||||
| 20403 | Itsn2 | 14013 | Mecom, Evi1 | ||
| 20910 | Stxbp1 | 17863 | Myb | ||
| 227738 | Lrsam1 | 52915 | Zmiz2 | ||
| Genes with miscellaneous function: | Genes with miscellaneous function: | ||||
| 11534 | Adk | 12857 | Cox4i1 | ||
| 12014 | Bach2/Evi59 * | 17909 | Myo10 | ||
| 12540 | Cdc42 | 19252 | Dusp1 | ||
| 12561 | Cdh4 | 22379 | Fmnl3 | ||
| 14297 | Fxn | 52906 | Ahi1 | ||
| 15403 | Hoxa6 | 54614 | Prpf40b | ||
| 15404 | Hoxa7 | 67561 | Wdr48 | ||
| 16800 | Arhgef2 | 68581 | Tmed 10 | ||
| 17977 | Ncoa1 | 74498 | Gorasp1 | ||
| 18263 | Odc1 | 75740 | Egfem1 | ||
| 21873 | Tjp2 | 110213 | Tmbim6 | ||
| 23984 | Pde10a | 217718 | Nek9 | ||
| 53602 | Hpcal1 | 268373 | Ppi A | ||
| 54169 | Myst4 | Genes with unknown function: | |||
| 56459 | Sae1 | 432939 | Gm5468 | ||
| 57915 | Tbc1d1 | 545510 | Gm10258 | ||
| 71801 | Plekhf2 | 666668 | Gm8225 | ||
| 140858 | Wdr5 | 100417604 | Gm18709 | ||
| 227737 | Fam129b | 100502810 | Gm19388 | ||
| 242093 | Rxfp4 | 100504283 | Gm20149 | ||
| 320150 | ZDHHC17 | ||||
| 330474 | Zc3h4 | ||||
| 381409 | Cdh26 | ||||
| 414084 | Tnip3 | ||||
| 435391 | Dupd1 | ||||
| Genes with unknown function: | |||||
| 66931 | 1700010I14Rik | ||||
| 67791 | 6530411M01Rik | ||||
| 68169 | A930038C07Rik | ||||
| 72628 | 2700086A05Rik | ||||
| 76947 | 2310030N02Rik | ||||
| 432502 | Gm5428 | ||||
| 595140 | Gm6044 | ||||
| 665533 | Gm13004 | ||||
| 100038651 | D130062J21Rik* | ||||
| 100188958 | Gm15824 | ||||
| 100417345 | Gm18548 | ||||
| 100417946 | Gm18913 | ||||
| 100503024 | Gm19512 | ||||
| 100503065 | LOC100503065 | ||||
| 100504144 | Gm13421 | ||||
RISs isolated from CFC mice with a chimerism higher than 15% (also displayed in Figure 2/D, see also Material and Methods S1) are listed. Gene categories were defined by the DAVID gene ontology (GO)21, using high classification stringency option by clustering. For Endocytosis GO group, the Fisher Exact test gave a P-value of 4.0E−2 and for Regulation of transcription GO group P-value = 2.9E−1.
= same RISs were found in 2 different young recipient mice (transplanted with two different pools of transduced cells), all the other RISs were unique within the experiments.
Aging in hematopoiesis is associated with a shift in lineage differentiation, in which B-lymphoid output is diminished and myeloid output enhanced, which has been primarily linked to stem cell intrinsic changes with age. [22] Interestingly, and in agreement with recently published data [23], the output of transduced BM cells exposed to aged microenvironment was significantly skewed towards the myeloid lineage with a diminished B-lymphoid contribution (Figure 2E), supporting as another novel concept that also niches, and not only stem cell intrinsic mechanisms are able to induce a lineage shift associated with aging under our dominant clone conditions.
In summary our data support a model in which an aged microenvironment, in contrast to a young one, selects for a few or only a single dominant hematopoietic clone and thus paving the way to mono-clonality (Figure 2F), which might be a contributing factor to the elevated incidence of leukemia in the elderly.
Materials and Methods
Retroviral Transduction and Transplantation Conditions
Lineage depleted (lin-) [24] and pre-stimulated young bone marrow (BM) cells were transduced overnight on retronectin-coated (TaKaRa, Otsu, Japan) plates with cell-free supernatants containing SF91/IRES-eGFP retrovirus as described in ref. [18]. See Material and Methods S1. Prior to transplantation, young (2 month) and aged (18–19 month) C57Bl/6 mice were non-lethally irradiated (total-body, 9.5 gray) and approximately 106 cells/recipient were injected. The number of young transplanted mice was 15 and 11 aged mice along with three independent transductions. See in Material and Methods S1.
Analysis of Recipient Mice
At 24–26 week post-transplant, when hematopoiesis is supported by long-term HSCs, we analyzed all of the 26 recipients for CFC formation among BM cells and also subjected then to FACS analysis. All animals initially transplanted were sacrificed, and PB, BM and spleen were analyzed for lineage markers (CD3, B220, Gr-1, Mac1) and lin−/c-Kit+/Sca1+ cells by flow cytometry as described in ref. [24]. For CFC-assays, 4×105 BM cells from each mice were seeded in methylcellulose culture (M3534, StemCell Technologies) for 9 days, GFP+ colonies were picked and genomic DNA from individual colonies were isolated (Qiagen) for LM-PCR. Only recipients with more than 15% eGFP+ cell frequency (similar frequency in young and aged recipients, see Figure S1 and Figure 1B,C) were further analyzed, as it was anticipated that in these animals the initial formation of dominant clones was successful, while they also gave rise to sufficient numbers of eGFP+CFCs within the numbers plated for further analysis. We then analyzed the eGFP+CFCs from these recipients that provided higher than 15% chimerism by LM-PCR.
Statistics
T-test was performed to determine the significance of the difference between means of two groups. Values were considered significant when p<0.05.
Ligation-mediated PCR (LM-PCR) and Retroviral Insertion Sites (RISs) Identification
LM-PCR was performed as described in ref. [25] and Material and Methods S1. PCR fragments from gels were isolated, sequenced (GATC, Konstanz, Germany) and sequences obtained were analyzed with following websites/databases: BLAST(http://blast.ncbi.nlm.nih.gov), Ensembl (http://www.ensembl.org), DAVID [21], RTCGD [26] and iDDb [19]. See in Material and Methods S1.
Ethics Statement
Animal experiments were carried out in University Ulm accordance with Tierschutzgesetz Paragraph8 Abs.1 and 3, were approved by the “Regierungspräsidium Tübingen” (Az:35/9185.81 protocol number 957). The study was performed in cooperation with the Comprehensive Mouse and Cancer Core in the Division of Experimental Hematology, Cincinnati Children’s Hospital Medical Center in accordance with the approved animal handling protocols (Nr. 9D04039 and 8D10089) provided by the Institutional Animal Care and Use Committee (IACUC) Cincinnati.
Supporting Information
Transplanted aged and young mice display normal hematopoiesis. (A) Spleen weight indicates normal spleen size (up to 120 mg). (B) GFP+ cell contribution in PB, differentiated BM (lineage marker positive: lin+) and lin−/c-kit+/Sca1+ (L-S+K+) cell-population 24–26 weeks post-transplant in young and aged recipient mice. The similar GFP levels in the different cell population indicate that transduced cells do not show a differentiation arrest. (C) GFP+CFC frequency among the total CFCs in methylcellulose culture of BM cells. (D) The size of the primitive cell compartment (LSK cell number) remained in normal range and did not differ from young and aged transplanted mice, which indicates that this cell pool remained under the regulatory control of the niche. (young transplanted mice n = 15, aged transplanted mice n = 11, from a total of 3 independent biological repeats, bars represent the mean ±SEM). Based on the analyzed PB and BM samples, the recipient mice did not show malengraftment.
(TIF)
Detailed list of integration sites isolated from young (A) and aged (B) recipient mice after 24–26 weeks of transplantation. The genomic locations of RISs and the nearest neighboring genes within 100 kb are listed. The RISs isolated from to the same recipient were grouped together to show that more RISs were recovered from young mice with oligoclonal GFP+population and less insertion sites from the monoclonal GFP+ population of the aged recipients. Figure 2/D was created based on sequencing results in combination with LM-PCR band patterns. The range of integrations per clone was calculated. The RIS/cell ranged from 1 to 4). The listed RISs were compared with two databases (Retroviral Tagged Cancer Gene Database (RTCGD), Insertional Dominance Database (IDDb)) to strengthen the validity of the applied methods. Within the RTCGD, retroviral integration sites associated with candidate cancer genes have been collected (ref. 22). The IDDb listed RISs related to dominant clone formation (ref. 23). Overall, 27.9% of the recovered RISs from young mice and 27.2% from old were found in the RTCGD, and 3 genes from young recipients and 1 gene from old recipients were found in IDDb. This efficiency to recover already described viral integration sites supports the validity of our approach. * = insertion sites belong to the dominant clone.
(DOCX)
Detailed description of the performed experiments
(XLSX)
Acknowledgments
We thank the Tierforschungszentrum of University of Ulm for supporting our animal work.
Funding Statement
This work was supported by grants from the Deutsche Forschungsgemeinschaft SFB497, TP10 and KFO142, GE2063/1 and from National Institute of Health, HL076604, DK077762 and AG040118 to HG. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
- 1. Iwasaki H, Suda T (2009) Cancer stem cells and their niche. Cancer Sci 100: 1166–1172. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Raaijmakers MHGP (2011) Niche contributions to oncogenesis: emerging concepts and implications for the hematopoietic system. Haematologica 96: 1041–1048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Henry CJ, Marusyk A, Degregori J (2011) Aging-associated changes in hematopoiesis and leukemogenesis: what’s the connection? Aging (Albany NY) 3: 643–656. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Dick JE, Lapidot T (2005) Biology of normal and acute myeloid leukemia stem cells. Int J Hematol 82: 389–396. [DOI] [PubMed] [Google Scholar]
- 5. Geiger H, Rudolph KL (2009) Aging in the lympho-hematopoietic stem cell compartment. Trends Immunol 30: 360–365. [DOI] [PubMed] [Google Scholar]
- 6. Rossi DJ, Bryder D, Seita J, Nussenzweig A, Hoeijmakers J, et al. (2007) Deficiencies in DNA damage repair limit the function of haematopoietic stem cells with age. Nature 447: 725–729. [DOI] [PubMed] [Google Scholar]
- 7. Signer RAJ, Montecino-Rodriguez E, Witte ON, Dorshkind K (2008) Aging and cancer resistance in lymphoid progenitors are linked processes conferred by p16ink4a and arf. Genes & Development 22: 3115–3120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Lane SW, Scadden DT, Gilliland DG (2009) The leukemic stem cell niche: current concepts and therapeutic opportunities. Blood 114: 1150–1157. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Carlesso N, Cardoso AA (2010) Stem cell regulatory niches and their role in normal and malignant hematopoiesis. Curr Opin Hematol 17: 281–286. [DOI] [PubMed] [Google Scholar]
- 10. Wei J, Wunderlich M, Fox C, Alvarez S, Cigudosa JC, et al. (2008) Microenvironment determines lineage fate in a human model of mll-af9 leukemia. Cancer Cell 13: 483–495. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Vianello F, Villanova F, Tisato V, Lymperi S, Ho KK, et al. (2010) Bone marrow mesenchymal stromal cells non-selectively protect chronic myeloid leukemia cells from imatinib-induced apoptosis via the CXCR4/CXCL12 axis. Haematologica 95: 1081–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Colmone A, Amorim M, Pontier AL, Wang S, Jablonski E, et al. (2008) Leukemic cells create bone marrow niches that disrupt the behavior of normal hematopoietic progenitor cells. Science 322: 1861–1865. [DOI] [PubMed] [Google Scholar]
- 13. Walkley CR, Olsen GH, Dworkin S, Fabb SA, Swann J, et al. (2007) A microenvironment-induced myeloproliferative syndrome caused by rarγ deficiency. Cell 129: 1097–1110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Freemont AJ, Hoyland JA (2007) Morphology, mechanisms and pathology of musculoskeletal ageing. J Pathol 211: 252–259. [DOI] [PubMed] [Google Scholar]
- 15. Wagner W, Horn P, Bork S, Ho AD (2008) Aging of hematopoietic stem cells is regulated by the stem cell niche. Experimental Gerontology 43: 974–980. [DOI] [PubMed] [Google Scholar]
- 16. Bellantuono I, Aldahmash A, Kassem M (2009) Aging of marrow stromal (skeletal) stem cells and their contribution to age-related bone loss. Biochim Biophys Acta 1792: 364–370. [DOI] [PubMed] [Google Scholar]
- 17. Bagby GC, Fleischman AG (2011) The stem cell fitness landscape and pathways of molecular leukemogenesis. Front Biosci (Schol Ed) 3: 487–500. [DOI] [PubMed] [Google Scholar]
- 18. Li Z, Schwieger M, Lange C, Kraunus J, Sun H, et al. (2003) Predictable and efficient retroviral gene transfer into murine bone marrow repopulating cells using a defined vector dose. Experimental Hematology 31: 1206–1214. [DOI] [PubMed] [Google Scholar]
- 19. Kustikova OS, Geiger H, Li Z, Brugman MH, Chambers SM, et al. (2007) Retroviral vector insertion sites associated with dominant hematopoietic clones mark “stemness” pathways. Blood 109: 1897–1907. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Bystrykh LV, Verovskaya E, Zwart E, Broekhuis M, de Haan G (2012) Counting stem cells: methodological constraints. Nat Methods 9: 567–574. [DOI] [PubMed] [Google Scholar]
- 21. Huang DW, Sherman BT, Lempicki RA (2009) Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 4: 44–57. [DOI] [PubMed] [Google Scholar]
- 22. Beerman I, Bhattacharya D, Zandi S, Sigvardsson M, Weissman IL, et al. (2010) Functionally distinct hematopoietic stem cells modulate hematopoietic lineage potential during aging by a mechanism of clonal expansion. Proc Natl Acad Sci U S A 107: 5465–5470. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Ergen AV, Boles NC, Goodell MA (2012) Rantes/Ccl5 influences hematopoietic stem cell subtypes and causes myeloid skewing. Blood 119: 2500–2509. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Daria D, Filippi M-D, Knudsen ES, Faccio R, Li Z, et al. (2008) The retinoblastoma tumor suppressor is a critical intrinsic regulator for hematopoietic stem and progenitor cells under stress. Blood 111: 1894–1902. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Kustikova OS, Baum C, Fehse B (2008) Retroviral integration site analysis in hematopoietic stem cells. Hematopoietic Stem Cell Protocols 430: 255–267. [DOI] [PubMed] [Google Scholar]
- 26. Akagi K (2004) RTCGD: retroviral tagged cancer gene database. Nucleic Acids Research 32: 523D–527. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Transplanted aged and young mice display normal hematopoiesis. (A) Spleen weight indicates normal spleen size (up to 120 mg). (B) GFP+ cell contribution in PB, differentiated BM (lineage marker positive: lin+) and lin−/c-kit+/Sca1+ (L-S+K+) cell-population 24–26 weeks post-transplant in young and aged recipient mice. The similar GFP levels in the different cell population indicate that transduced cells do not show a differentiation arrest. (C) GFP+CFC frequency among the total CFCs in methylcellulose culture of BM cells. (D) The size of the primitive cell compartment (LSK cell number) remained in normal range and did not differ from young and aged transplanted mice, which indicates that this cell pool remained under the regulatory control of the niche. (young transplanted mice n = 15, aged transplanted mice n = 11, from a total of 3 independent biological repeats, bars represent the mean ±SEM). Based on the analyzed PB and BM samples, the recipient mice did not show malengraftment.
(TIF)
Detailed list of integration sites isolated from young (A) and aged (B) recipient mice after 24–26 weeks of transplantation. The genomic locations of RISs and the nearest neighboring genes within 100 kb are listed. The RISs isolated from to the same recipient were grouped together to show that more RISs were recovered from young mice with oligoclonal GFP+population and less insertion sites from the monoclonal GFP+ population of the aged recipients. Figure 2/D was created based on sequencing results in combination with LM-PCR band patterns. The range of integrations per clone was calculated. The RIS/cell ranged from 1 to 4). The listed RISs were compared with two databases (Retroviral Tagged Cancer Gene Database (RTCGD), Insertional Dominance Database (IDDb)) to strengthen the validity of the applied methods. Within the RTCGD, retroviral integration sites associated with candidate cancer genes have been collected (ref. 22). The IDDb listed RISs related to dominant clone formation (ref. 23). Overall, 27.9% of the recovered RISs from young mice and 27.2% from old were found in the RTCGD, and 3 genes from young recipients and 1 gene from old recipients were found in IDDb. This efficiency to recover already described viral integration sites supports the validity of our approach. * = insertion sites belong to the dominant clone.
(DOCX)
Detailed description of the performed experiments
(XLSX)