Abstract
Aims
Recent studies have suggested that increased α-smooth muscle-actin-positive myofibroblastic cells (α-SMA-positive CAFs) in the desmoplastic stroma may relate to a more aggressive cancer and worse survival outcomes for intrahepatic cholangiocarcinoma (ICC) patients. To facilitate investigating cellular and molecular interactions between α-SMA-positive CAFs and cholangiocarcinoma cells related to ICC progression, we developed a novel 3-dimensional (3-D) organotypic culture model of cholangiocarcinoma that more accurately mimics the stromal microenvironment, gene expression profile, and select pathophysiological characteristics of desmoplastic ICC in vivo.
Methods
This unique model was established by co-culturing within a type I collagen gel matrix, a strain of cholangiocarcinoma cells (derived from an ICC formed in syngeneic rat liver following bile duct inoculation of spontaneously-transformed rat cholangiocytes) with varying numbers of clonal α-SMA-positive CAFs established from the same tumor type.
Results
Cholangiocarcinoma cells and α-SMA-positive CAFs in monoculture each exhibited cell specific biomarker gene expression profiles characteristic of stromal myofibroblastic cell versus malignant cholangiocyte cell types. In comparison, the gene expression profile and histopathological characteristics exhibited by the organotypic co-culture closely resembled those of whole tissue samples of the parent orthotopic ICC. We further showed α-SMA-positive CAFs to significantly enhance cholangiocarcinoma cell “ductal-like” growth and cancer cell migration/invasiveness in vitro, as well as to promote up-regulated expression of select genes known to be associated with ICC invasion.
Conclusions
This novel organotypic model provides an important new resource for studying the effects of microenvironment on cholangiocarcinoma progression in vitro and may have potential as a preclinical model for identifying molecularly targeted therapies.
Introduction
Cancer-associated fibroblastic cells (CAFs) immunoreactive for α-smooth muscle actin (α-SMA) are a major cellular component of the desmoplastic stroma of primary cholangiocarcinomas formed in liver (1–3). Although largely descriptive, there is accumulating evidence to suggest that these myofibroblastic-like cells may have a crucial role in accelerating intrahepatic cholangiocarcinoma (ICC) progression (4, 5).
In humans, increased α-SMA-positive CAFs in the stroma of ICCs has been shown to correlate with larger tumor size and/or shorter patient survival times when compared with ICCs low in positive α-SMA stromal cell immunoreactivity (6, 7). Moreover, non-contact co-culturing of human cholangiocarcinoma cell lines (HuCCT1 or MEC) with either the immortalized myofibroblastic cell line LX-2 derived from human hepatic stellate cells (8) or the LI90 myofibroblastic cell line derived from a human hepatic mesenchymal tumor (9) was shown to significantly enhance cholangiocarcinoma cell proliferation and migration (invasion) in vitro over that exhibited by cholangiocarcinoma cells assayed in the absence of LX-2 or LI90 cells (6). Conditioned medium (CM) from LI90 cell cultures also significantly increased human HuCCT1 and RBE cholangiocarcinoma cell proliferation and invasion in vitro (10). Of further interest, CM from primary culture α-SMA-positive CAFs derived from a human cholangiocarcinoma was demonstrated in 2-dimensional (2-D) culture to stimulate a significantly greater cell proliferative response in cultured human KKU cholangiocarcinoma cell lines of varying states of differentiation, as well as in non-tumorigenic SV-40 immortalized human H-69 cholangiocytes, than that elicited by CM from primary cultures of normal skin fibroblasts or primary fibroblasts from non-tumorigenic liver (7).
While the results of the cell culture studies described above are clearly suggestive that interactions between α-SMA-positive CAFs and cholangiocarcinoma cells may be critically important to promoting ICC growth and progression, they are limited in their relevance to the in vivo situation by the fact that they were performed largely under 2-D culture conditions using longstanding cholangiocarcinoma cell lines of various biliary tumor cell origins, and in the case of LX-2 and LI90 cells, with long-term myofibroblastic-like cells derived from non-cholangiocarcinoma tissues. In an effort to more accurately reproduce in vitro the tissue architecture and complex interactive relationships between cholangiocarcinoma cells and stromal α-SMA-positive CAFs in a type I collagen matrix microenvironment reflective of desmoplastic ICC, we have now developed novel 3-dimensional (3-D) organotypic co-culture models of rat cholangiocarcinoma that closely mimic relevant histopathological, molecular, and progressive features of in vivo rat and human ICC. This unique in vitro cholangiocarcinoma model employs a 3-D gel matrix based on rat type I collagen gel to support co-culturing of a clonal strain of cholangiocarcinoma cells with a clonal α-SMA-positive CAF strain, both of which were derived from orthotopic ICC (BDEsp ICC) formed in syngeneic rat liver following bile duct inoculation of spontaneously-transformed tumorigenic rat cholangiocytes (11, 12).
Materials and Methods
Cell Lines and 3-D Organotypic Culturing
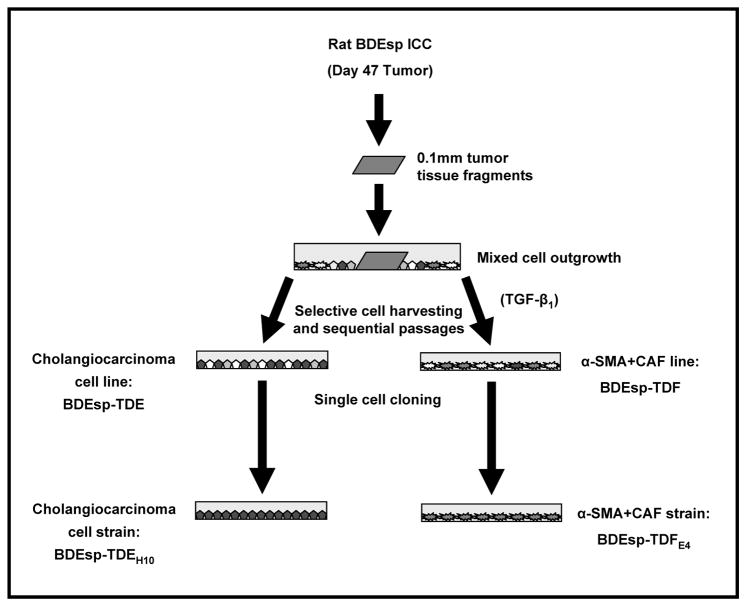
The animal experiments used to generate the orthotopic cholangiocarcinoma tumors employed in this study were performed in accordance with and approved by Virginia Commonwealth University Institutional Animal Care and Use Committee. Phenotypically distinct rat cholangiocarcinoma and desmoplastic tumor stromal-associated α-SMA-positive CAF cell lines were each obtained through selective cell harvesting and serial in vitro passaging (2–4 passages), using various cell enrichment conditions, of primary mixed cell outgrowths from 0.1mm tissue fragments isolated from orthotopic BDEsp ICC, as previously described (12) and as schematically depicted in Figure 1. The cholangiocarcinoma cell line was designated as BDEsp-TDE and the corresponding tumor-derived α-SMA-positive CAF cell line as BDEsp-TDF. Single cell cloning was then utilized to produce the cholangiocarcinoma (BDEsp-TDEH10) and α-SMA-positive CAF (BDEsp-TDFE4) cell strains, which were used to establish the 3-D organotypic cholangiocarcinoma co-culture model described in this study. Specifically, this model was developed by co-culturing cholangiocarcinoma cells at an initial plating density of 1 × 105 cells/ml and at cell viabilities of ≥ 95% with α-SMA-positive CAFs at various initial plating densities ranging between 0-to-8 × 105 cells/ml (also with cell viabilities of ≥ 95%) mixed within a gel matrix of rat tail type I collagen (BD Biosciences, Bedford, MA). Cholangiocarcinoma and α-SMA-positive CAF cell strains at in vitro passage numbers ≤ 8 were used in the co-culture experiments. Unless otherwise specified, the cultures were maintained with our standard medium (13), for experimental periods of up to 10 days.
Figure 1.
Scheme outlining the method used to concomitantly establish the BDEsp-TDEH10 cholangiocarcinoma celland BDEsp-TDFE4 cancer-associated fibroblastic (CAF) cellstrains derived from an orthotopic BDEsp ICC obtained from syngeneic rat liver at day 47 after bile duct inoculation of the tumorigenic, spontaneously-transformed rat cholangiocyte cell line (BDEsp). As an initial selection step, TGF-β1 at 5ng/ml was temporarily added to the culture medium to facilitate expansion of the CAF population.
Gene Expression Microarray Analyses
The Affymetrix® protocol utilized for microarray analyses was previously described (12) utilizing 500ng of total RNA prepared from BDEsp ICC intact tumor samples or from cell isolates harvested at day 7 from 3-D mono- or co-cultures of BDEsp-TDEH10 and BDEsp-TDFE4 cells (serum free for 24h). As per the Affymetrix® protocol, 10μg of fragmented cRNA were hybridized on the GeneChip® Rat Expression 230 2.0 Array (Affymetrix Inc., Santa Clara, CA) for 16h at 60rpm in a 45°C hybridization oven. Every chip was scanned at a high resolution on the Affymetrix® GeneChip Scanner 3000 7G as previously described (12). Overall quality of each array was assessed by monitoring the 3′/5′ ratios for the housekeeping gene, glyceraldehyde 3-phosphate dehydrogenase (Gapdh), and the percentage of “Present” genes (%P). Arrays exhibiting Gapdh 3′/5′ < 3.0 and %P > 40% were considered good quality arrays.
Real-Time Reverse-Transcriptase Polymerase Chain Reaction (QRT-PCR)
QRT-PCR was used to validate gene expression levels of selected rat genes using TaqMan® chemistry, essentially as previously described (12). Probes and primer sets specific for the detection of rat mRNA transcripts were purchased from Applied Biosystems, Foster City, CA. These included gene-specific primer-probe sets for the following biomarker genes: Krt19 (cytokeratin 19), ID# Rn01496870_g1; Muc1 (mucin 1), ID# Rn01462585_m1; Mmp7 (metalloproteinase-7, matrilysin), ID# Rn01487001_m1; Areg (amphiregulin), ID# Rn00567471_m1; Acta2 (α-SMA), ID# Rn01759928_g1; Hgf (hepatocyte growth factor), ID# Rn00566673_m1; Postn (periostin), ID# Rn1494627_m1; Sphk1 (sphingosine kinase 1), ID# Rn00682794_g1; Cxcr4 (C-X-C chemokine receptor 4), ID# Rn01483207_m1; Shh (sonic hedgehog homolog), ID# Rn00568129_m1; SMO (Smoothened), ID# Rn00563043_m1; Gli1 (glioblastoma 1), ID# Rn01504237_m1; Gli2, ID# Rn01408890_m1. PPIA (cyclophilin), ID# Rn00690933_m1 or Gapdh (glyceraldehyde 3-phosphate dehydrogenase), RefSeq: NM_017008.3 were used as controls. All QRT-PCR analyses were done in triplicate.
Histology, Immunohistochemistry, and Microscopic Imaging
Hematoxylin and eosin (H&E) staining was routinely performed on 10% formalin-fixed paraffin-embedded tissue sections of collagen gel cultures. Histochemical staining for fibrous collagen (primarily type I) was also carried out on comparably fixed and processed tissue sections from the collagen gel cultures, as well as from BDEsp-ICC using the picrosirius red staining method using reagents obtained from Polysciences, Inc. (Warrington, PA). Picrosirius red staining was viewed under a BX41 light microscope equipped with a BX-POL polarizer (Olympus Corp., Center Valley, PA). Immunohistochemical staining was performed essentially as described previously (13–15) on appropriately processed formalin-fixed, paraffin-embedded tissue sections, using the following primary antibodies: monoclonal mouse anti-human SMA (Cat. No. M0851) from DakoCytomation, Carpinteria, CA; mouse monoclonal cytokeratin 19 (CK19) (Cat. No. VP-C415) from Vector Laboratories, Burlingame, CA. Computational image analysis was used to quantify cholangiocarcinoma spheroidal or ductal-like structures formed in 3-D mono- or co-cultures using the cellSens imaging software system (Vers. 1.4.1) from Olympus Corp. This analysis was carried out on digital images of randomized microscopic fields of 10μm thick tissue sections prepared from formalin-fixed, paraffin-embedded gel cultures stained with H&E (total number of collagen gel cultures analyzed per experimental condition: 3; total number of analyzed tissue sections per individual collagen gel culture: 2-to-4).
Invasion Assay
Cholangiocarcinoma cell migration in vitro (invasion) was assayed using the BD Biocoat Matrigel™ Invasion chamber (24 well plate coated with rat type 1 collagen; Matrigel-coated inserts with 8μm pore) (BD Biosciences), similar to the method described by Ohira et al. (16). BDEsp-TDFE4 cancer-associated fibroblasts were initially plated onto the collagen-coated bottom chamber of the 24 well plates at a plating density of 4 × 104 CAFs/well. Twenty-four hours later, BDEsp-TDEH10 cholangiocarcinoma cells were plated at 1 × 105 cells/well onto the upper surface of the Matrigel-coated insert (top chamber) of the tissue culture well. After 48 hours of incubation at 37° C (under standard culture conditions), the non-invaded cells were removed from the top surface of the Matrigel-coated porous insert with cotton-tipped swabs. The cholangiocarcinoma cells that had migrated to the lower surface of the insert were fixed with 4% paraformaldehyde and stained with 0.1% crystal violet. The migrated cells were then counted and the invasion index determined according to the manufacturer’s instructions.
Statistics
For the microarray data analysis, background correction, normalization, and estimation of probe set expression summaries were performed using the log-scale robust multi-array analysis (RMA) method (17). The Student 2-tailed t test was used to determine P values, with a P ≤ 0.05 considered significant. Statistical significance for multivariate analysis to assess probe set specific false discovery rates (FDR) was performed by estimating the q-values, using the Bioconductor q-value package (18), as previously described (12).
Results
Selective gene expression profiling of BDEsp-TDFE4 and BDEsp-TDEH10
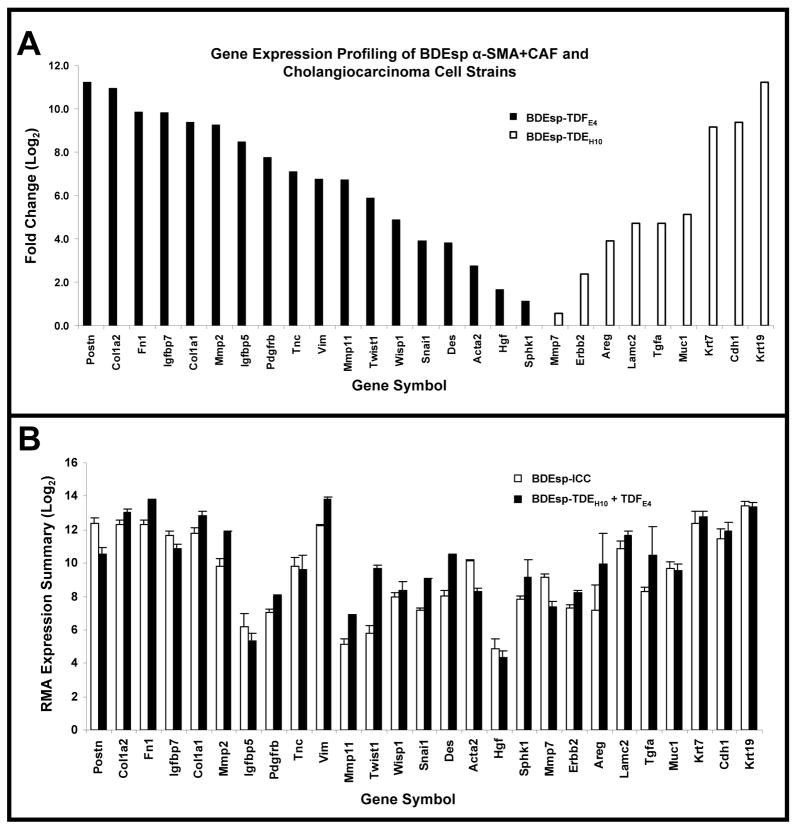
Gene expression profiles determined by microarray analysis for selected biomarker genes expressed in BDEsp-TDFE4 versus BDEsp-TDEH10 cells cultured separately or together for 7 days within rat type I collagen gel matrix are shown in Figure 2. As clearly demonstrated in Figure 2A, BDEsp-TDFE4 cells cultured alone in collagen gel matrix expressed a biomarker profile consistent with that of a cholangiocarcinoma-associated stromal myofibroblastic cell phenotype (4, 5), whereas BDEsp-TDEH10 cells cultured alone in collagen gel matrix expressed biomarker genes indicative of a neoplastic cholangiocyte origin (3, 19). Particularly noteworthy are the high levels of gene expression for extracellular matrix genes (Postn, collagen type 1a, fibronectin, and tenascin-C), as well as for Mmp2 and 11, insulin-like growth factor binding proteins 5 and 7, platelet derived growth factor receptor-β, vimentin, WNT1 inducible signaling protein 1, and desmin selectively exhibited by the cultured BDEsp-TDFE4 cells. In contrast, BDEsp-TDEH10 cells cultured alone under identical conditions were characterized by their high levels of gene expression for biliary cell Krt 7 and 19, E-cadherin, Muc1, and Areg. The gene expression pattern for specific integrins was also determined to be distinctly different for BDEsp-TDFE4 versus BDEsp-TDEH10 in monoculture in type I collagen gel matrix, with integrin β5 (Itgβ5) mRNA being predominantly expressed in the CAFs and Itgβ4 mRNA being most highly expressed in the cholangiocarcinoma cells (Supplemental Figure 1). In addition, our QRT-PCR analyses confirmed cultured BDEsp-TDEH10 cellsto selectively express mRNA for Krt19, Muc1, Mmp-7, and Areg, when compared against cultured BDEsp-TDFE4 cells, and conversely, cultured BDEsp-TDFE4 cells differentially expressed mRNA for Acta2 (α-SMA), Hgf, Postn, and Sphk1 (Supplemental Figure 2). Gene members of the Hedgehog (Hh) signaling pathway were also demonstrated by QRT-PCR to be differentially expressed in the BDEsp-TDEH10 versus BDEsp-TDFE4 cultures, with Shh being largely expressed in the cholangiocarcinoma cell strain and SMO, Gli1 and Gli2 being predominantly expressed in the CAF strain (Supplemental Figure 3).
Figure 2.
Gene expression profiling based on selected gene biomarkers of BDEsp ICC-derived cancer-associated myofibroblastic cells and cholangiocarcinoma cell strains. (A) Bar chart representation of 27 marker genes assessed by microarray analysis that showed significant differences in expression levels between BDEsp-TDFE4 and BDEsp-TDEH10 when cultured individually in a rat tail type I collagen gel matrix. (B) Robust Multiarray Analysis (RMA) expression summaries for the 27 analyzed biomarker genes shown in A, comparing orthotopic BDEsp ICC to organotypic BDEsp-TDFE4/BDEsp-TDEH10 culture. Postn, periostin; Col1a2, collagen type Ia2; Fn1, fibronectin 1; Igfbp7, insulin-like growth factor binding protein 7; Col1a1, collagen type Ia1; Mmp2, metalloproteinase-2; Igfbp5, insulin-like growth factor binding protein 5; Pdgfrb, platelet-derived growth factor receptor β; Tnc, tenascin-C; Vim, vimentin; Mmp11, metalloproteinase 11; Twist1, Twist homolog 1; Wisp1, WNT1 inducible signaling protein 1; Snail1, Snail homolog 1; Des, desmin; Acta2, smooth muscle actin; Hgf, hepatocyte growth factor; Sphk1, sphingosine kinase 1; Mmp7, metalloproteinase-7; Erbb2, c-erbB2 protooncogene receptor protein; Areg, amphiregulin; Lamc2, laminin gamma2; Tgfa, transforming growth factor α; Muc1, mucin 1; Krt7, cytokeratin 7; Cdh1, E-cadherin; Krt19, cytokeratin 19.
Organotypic 3-D co-culture of BDEsp-TDFE4 and BDEsp-TDEH10 cells in type I collagen gel matrix closely mimics desmoplastic BDEsp ICC
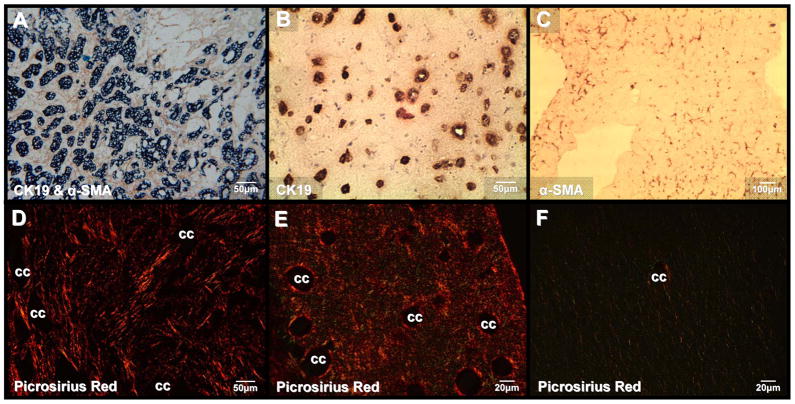
As depicted in Figure 3, 3-D organotypic co-cultures of CK19 (Krt 19)-positive BDEsp-TDEH10 cholangiocarcinoma cells and α-SMA-positive BDEsp-TDFE4 CAFs in rat type I collagen gel matrix reproduce characteristic histopathological features of desmoplastic BDEsp ICC in vivo, including enhanced histochemical staining for type I collagen secreted into the matrix (Figure 3E). Furthermore, our microarray data shown in Figure 2B revealed the organotypic α-SMA-positive CAF/cholangiocarcinoma co-cultures to exhibit an RMA expression summary for CAF and cholangiocarcinoma biomarker genes closely paralleling that of resected whole BDEsp ICC tumor tissue.
Figure 3.
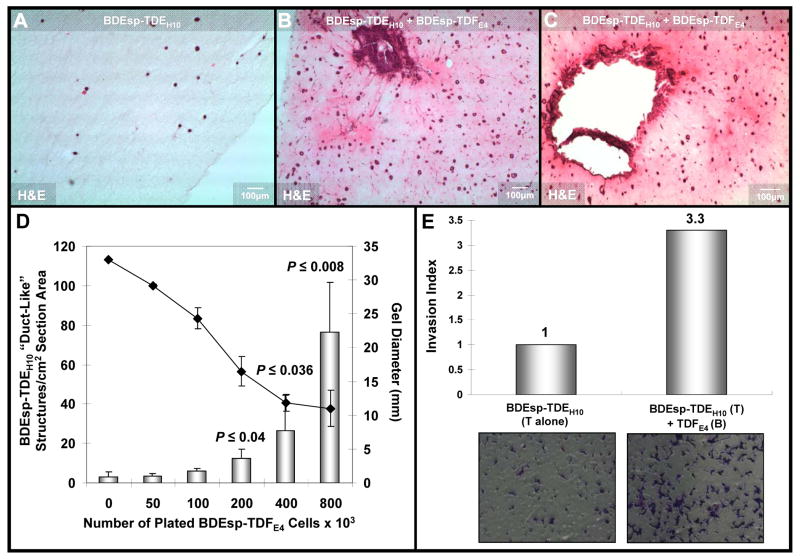
Co-culturing of BDEsp-TDEH10 cells with BDEsp-TDFE4 cells in rat tail type I collagen gel matrix reproduces characteristic histopathological features of desmoplastic BDEsp ICC in vivo. (A) Desmoplastic BDEsp ICC formed in rat liver showing selective positive immunostaining of well differentiated cholangiocarcinoma ducts for biliary cytokeratin 19 (CK19) (blue staining) and cancer-associated myofibroblastic cells positive for α-smooth muscle actin (α-SMA) (brown staining) in the surrounding tumor stroma. (B) Like BDEsp ICC, well differentiated “duct-like” structures formed from BDEsp-TDEH10 cells in organotypic co-culture exhibit selective immunoreactivity for CK19 (brown staining). (C) Also like BDEsp ICC, stromal BDEsp-TDFE4 cells in organotypic co-culture are selectively immunoreactive for α-SMA (brown staining). (D) Picrosirius red staining under polarized light of type I collagenous fibers (orange-yellow) densely populating the desmoplastic stroma of a BDEsp ICC. (E) Picrosirius red staining of a representative histological section of a BDEsp-TDE/BDEsp-TDF co-culture mimicking the strong extracellular staining reaction for type I collagen exhibited by the orthotopic ICC in D. (F) Picrosirius red staining of a BDEsp-TDEH10 control culture without BDEsp-TDF showing only background staining for type I collagen fibers in gel matrix. cc denotes representative cholangiocarcinoma ducts or “duct-like” structures.
BDEsp-TDFE4 CAFs promote BDEsp-TDEH10 cholangiocarcinoma cell growth and invasion in co-culture
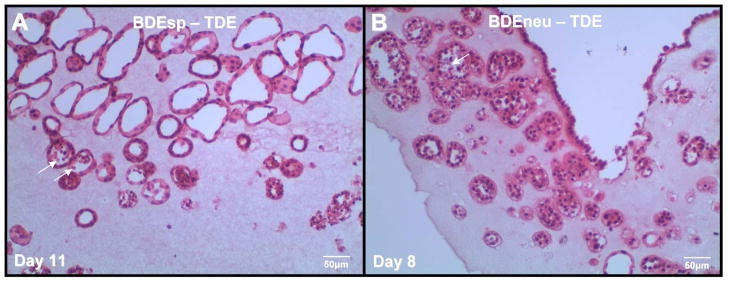
BDEsp-TDE cholangiocarcinoma cells when cultured alone in rat type 1 collagen gel matrix form into 3-D spheroid structures, which when viewed in routine H & E stained histological sections appear in cross-section as duct-like structures mimicking in their morphology the neoplastic ducts of well differentiated BDEsp ICC from which they were derived (Figure 4). As further demonstrated in Figure 5B versus 5A, and histological sections shown in Figure 6A-D, co-culturing BDEsp-TDEH10 with BDEsp-TDFE4 cells under our 3-D organotypic culture conditions profoundly increased the number of cholangiocarcinoma cell spheroid/duct-like structures formed in the collagen gel matrix over those formed in the absence of BDEsp-TDFE4 cells. Furthermore, as demonstrated in Figure 6D, the mean number of spheroid/duct-like structures formed per cm2 section area in randomly analyzed histological preparations of the gel cultures significantly increased as a function of higher initial BDEsp-TDFE4 cell plating densities when the ICC-derived cancer cells and CAFs were co-cultured for 6 days in rat type I collagen gel matrix, correlating with an increasing shrinkage of the collagen gel substratum. A more progressive cholangiocarcinoma cell phenotype also resulted from organotypic co-culturing of the BDEsp-TDFE4 cells with BDEsp-TDEH10 cells compared to mono-cultures of cholangiocarcinoma cells only (Figure 6B & Cversus 6A). Moreover, BDEsp-TDEH10 cholangiocarcinoma cells exhibited significantly increased in vitro invasiveness when assayed in the presence versus absence of BDEsp-TDFE4 CAFsusing the Matrigel invasion chamber.
Figure 4.
Representative H & E stained histological section of our rat cholangiocarcinoma cell line BDEsp-TDE at in vitro passage 3, derived from well differentiated BDEsp ICC, and cultured alone for 11 days within rat tail type I collagen gel matrix. Note that the cholangiocarcinoma cells are organized into well differentiated “duct-like” structures reminiscent of those observed in histological sections of the parent low grade BDE ICC. For comparison, the photomicrograph in (B) demonstrates a rat cholangiocarcinoma cell line designated as BDEneu-TDE that was established in our laboratory from a more highly malignant and moderately-differentiated ICC formed in syngeneic rat liver after bile duct inoculation of oncogenic-neu transformed rat cholangiocytes. As demonstrated in B, rat BDEneu-TDE cells in organotypic culture in a type I collagen gel matrix reflect the less differentiated “duct-Like” phenotype of the parent orthotopic tumor. In both A & B, arrows point to dead cells within the lumens of some of the neoplastic “duct-like” structures, indicating that cells within the central areas of formed cholangiocarcinoma cell spheroids are likely undergoing apoptotic cell death.
Figure 5.
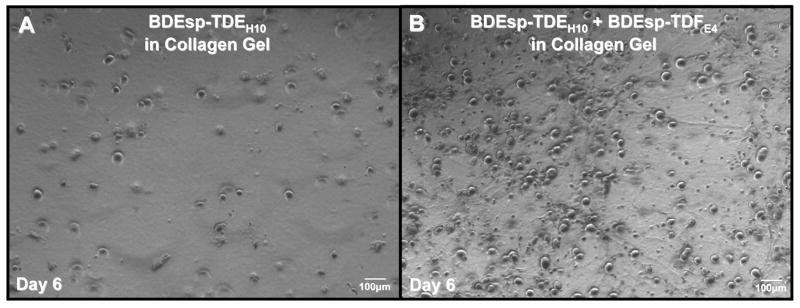
Representative phase contrast photomicrographs of (A) rat BDEsp-TDEH10 cholangiocarcinoma cells cultured alone for 6 days in rat tail type I collagen gel compared with (B) in which the BDEsp-TDEH10 cells were co-cultured under comparable conditions over the same time period with BDEsp-TDFE4 CAFs. Compare with the photomicrographs of the H & E stained histological sections shown in Figure 6, noting the marked increase in 3-dimensional cell spheroids formed in the co-culture over those observed in the cholangiocarcinoma cell monoculture.
Figure 6.
Photomicrographs of representative H & E stained histological sections of 3-dimensional organotypic cultures of BDEsp-TDEH10 cholangiocarcinomacells cultured for 6 days either in monoculture (A) or in co-culture with BDEsp-TDFE4 CAFs (B & C). Cholangiocarcinoma cells were initially plated at a cell density of 2 × 105 cells/type I collagen gel in both the mono- and co-cultures. The initial CAF cell plating density in the co-cultures was at 8 × 105 cells/gel. The histological section shown in B was obtained in the same experiment as that for A, whereas that in C was from a separate experiment essentially repeating the culture conditions used for B. Note the dramatic increase in the number of spheroidal/”duct-like” structures formed from the cholangiocarcinoma cells in the co-cultures (B & C) over those formed under comparable culture conditions in control cholangiocarcinoma cell monoculture without CAFs (A). Also observe a much more intense eosinophilic staining of the gel matrix in B & C compared to that of A, indicative of an increase in the cellular secretion of insoluble proteins into the co-culture matrix versus that produced in the cholangiocarcinoma cell monoculture. As further depicted in B & C, only the co-cultures exhibited prominent irregular clusters of anaplastic cholangiocarcinoma cells surrounded by stromal CAFs, reflective of malignant progression and apparent invasiveness. (D) Bar graph demonstrating the mean number of spheroidal/”duct-like” structures/cm2 tissue section area formed from BDEsp-TDEH10 cholangiocarcinoma cells to significantly increase as a function of higher initial BDEsp-TDFE4 plating densities when these ICC derived cell types were co-cultured for 6 days in rat type I collagen gel matrix. Each value represents the mean ± SD determined from microimaging measurements made on 10μm thick H & E stained histological sections (n = at least 2 sections/slide prepared from triplicate cultures for each CAF plating density). P values were determined against the 0 CAF culture condition. Gel contraction, measured as a reduction in mm gel diameter (
 = mean ± SD) was determined to be significantly greater at the higher initial BDEsp-TDFE4 cell plating densities. (E) Representative data demonstrating BDEsp-TDFE4 CAFsto increase the cell migration/invasiveness of BDEsp-TDEH10 cholangiocarcinoma cells in vitro when assessed in the Matrigel™ invasion bioassay system. T= top chamber coated by Matrigel; B = bottom culture well coated with rat tail type I collagen. Invasion Index was determined according to the manufacturer’s instructions.
= mean ± SD) was determined to be significantly greater at the higher initial BDEsp-TDFE4 cell plating densities. (E) Representative data demonstrating BDEsp-TDFE4 CAFsto increase the cell migration/invasiveness of BDEsp-TDEH10 cholangiocarcinoma cells in vitro when assessed in the Matrigel™ invasion bioassay system. T= top chamber coated by Matrigel; B = bottom culture well coated with rat tail type I collagen. Invasion Index was determined according to the manufacturer’s instructions.
Interaction between BDEsp-TDFE4 and BDEsp-TDEH10 cells induces up-regulation of genes linked to cholangiocarcinoma progression and/or invasion
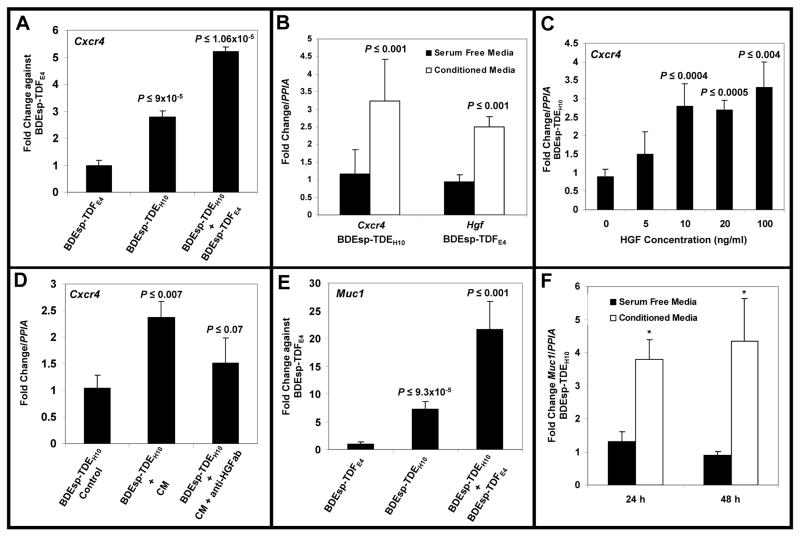
Both Muc1 and CXCR4 have been suggested to play key roles in promoting ICC progression and/or migration/invasion (3, 5, 19, 20). Here it is relevant that culturing BDEsp-TDEH10 cholangiocarcinoma cells in the presence of BDEsp-TDFE4 CAFs or conditioned medium from BDEsp-TDFE4 cultures significantly increased both Cxcr4 and Muc1 expression in the cholangiocarcinoma cells (Figure 7). In comparison, conditioned medium from BDEsp-TDEH10 cultures produced a significant up-regulation of Hgf expression in cultured BDEsp-TDFE4 cells (Figure 7B). The data shown in Figure 7C & D further suggest that HGF produced by BDEsp-TDFE4 CAFs is the likely inducer of increased Cxcr4 gene expression in the cholangiocarcinoma cells. Recombinant HGF has also been recently reported to up-regulate Cxcr4 expression in cultured human glioma cells (21). In contrast, Muc1 up-regulation in cultured BDEsp-TDEH10 cells was determined by us not to be up-regulated by HGF (data not shown).
Figure 7.
(A) TaqMan® QRT-PCR data demonstrating that co-culturing of rat BDEsp-TDEH10 cholangiocarcinoma cells in rat tail type I collagen gel matrix in the presence of BDEsp-TDFE4 CAFs grown to confluence on type I collagen-coated plastic significantly induced Cxcr4 gene expression in the cholangiocarcinoma cells over monoculture cell values. (B). QRT-PCR data showing conditioned medium from cultured BDEsp-TDFE4 cells to significantly up-regulate Cxcr4 gene expression in BDEsp-TDEH10 cholangiocarcinoma cells cultured alone on type I collagen-coated plastic, whereas conditioned medium from BDEsp-TDE H10 cells significantly induced Hgf gene expression in BDEsp-TDFE4 cells cultured alone under identical conditions. (C) QRT-PCR data showing a significant concentration-dependent increase in Cxcr4 gene expression in cultured BDEsp-TDEH10 cells following exposure over a 48-hour treatment period to recombinant human HGF. (D) QRT-PCR data showing the effect of conditioned medium from cultured BDEsp-TDFE4 CAFs on up-regulating Cxcr4 gene expression in cultured BDEsp-TDEH10 cholangiocarcinoma cells is partially blocked by the addition of neutralizing HGF antibody (anti-HGFab). (E) QRT-PCR data demonstrating that co-culturing of BDEsp-TDEH10 cholangiocarcinoma cells in the presence of BDEsp-TDFE4 CAFs under identical conditions as in A also produced a significant up-regulation of Muc1 gene expression in the cholangiocarcinoma cells over monoculture cell values. (F) Similar to B, conditioned medium from cultured BDEsp-TDFE4 CAFs induced a significant increase in Muc1 gene expression in cultured BDEsp-TDEH10 cholangiocarcinoma cells. PPIA (cyclophilin A) was used as the housekeeping gene to normalize Cxcr4, Hgf and Muc1 gene expression. Comparable QRT-PCR results were also obtained when normalized with Gapdh. Each value represents the mean ± SD, obtained from determinations made on a minimum of six cultures per experimental condition.
Discussion
Organotypic culture models have been developed for different tumor types, including breast, prostate, pancreatic, ovarian, and esophageal cancers (22–25). However, to our knowledge, there have been no previous reports describing the development of a 3-D culture model based on co-culturing of α-SMA-positive CAFs and cholangiocarcinoma cells simultaneously derived from orthotopic ICC.
Unique features distinguishing this novel 3-D co-culture model include: (1) its construction using “phenotypically pure” α-SMA-positive CAF and cholangiocarcinoma cell strains derived from desmoplastic ICC formed from spontaneously transformed rat cholangiocytes orthotopically transplanted into syngeneic rat liver; (2) α-SMA-positive CAFs and cholangiocarcinoma cells used to establish this organotypic rat cholangiocarcinoma model each exhibited characteristic gene expression profiles for select classes of genes (e.g., matricellular proteins, growth factor/chemokines and receptors, mucins, proteases, integrins, Hh family members) relevant to cholangiocarcinogenesis; (3) the identification of gene expression profiles using this novel 3-D cellular model closely resembled that of tissue samples obtained from parental BDEsp cholangiocarcinoma grown orthotopically; (4) 3-D co-culturing of rat cholangiocarcinoma cells and α-SMA-positive CAFs derived from the same rat ICC type significantly enhanced “duct-like” growth and cancer cell migration/invasion, as well as augmented type I collagen deposition into the gel matrix so as to mimic in vitro the desmoplastic stroma of the parental tumor; (5) α-SMA-positive CAF-cholangiocarcinoma cell interactions resulted in up-regulated gene expression of Hgf, Cxcr4, and Muc1, each of which has been associated with cholangiocarcinoma cell invasiveness and/or poorer survival outcomes for ICC patients following tumor resection (3–5, 16, 20, 26); and (6) HGF produced by α-SMA-positive CAFs is an inducer of significant Cxcr4 overexpression in cholangiocarcinoma cells.
Our histological data further indicate that 3-D co-culturing of rat cholangiocarcinoma cells with α-SMA-positive CAFs within a type I collagen gel matrix closely reproduces characteristic morphological features of the parental orthotopic BDEsp tumor, also resembling the histopathology of well-differentiated, desmoplastic tubular ICC in humans. Moreover, this organotypic rat cholangiocarcinoma model was found to exhibit gene expression profiles reflecting highly expressed CAF (e.g., Postn, Tnc) and cholangiocarcinoma genes (e.g., Muc1 ), whose overexpression in human ICC has been reported to be significantly correlated with human ICC progression, as well as shorter patient survival times following surgical resection (3, 5, 20, 27).
Overexpression of Postn in cholangiocarcinoma-derived myofibroblastic cells and of Muc1 in cholangiocarcinoma cells have also been observed by us to correlate with increased malignancy in our orthotopic ICC syngeneic rat model (11, 12). In addition, we have now determined that BDEsp transformants stably transfected to overexpress rat Postn or Muc1 cDNA exhibit significantly enhanced cell migration/invasion in vitro compared with that of corresponding empty vector and untransfected BDEsp controls (Sirica, A. E., et al., unpublished data). Although these latter findings are preliminary, they suggest a molecular strategy for ICC therapy based on combinational targeting of Postn and Muc1 as candidate gene pathways related to ICC invasive growth. Such a novel molecular therapeutic strategy can now be readily tested in the complementary organotypic culture and orthotopic cholangiocarcinoma syngeneic rat models that are currently available in our laboratory.
The 3-D organotypic culture model of cholangiocarcinoma described in his study appears to be very well suited to serve as a pathophysiologically relevant in vitro system to investigate key stromal/cancer cell interactions mediated by aberrantly expressed CAF or cholangiocarcinoma genes postulated to accelerate ICC malignancy (3– 5). It is also anticipated that this 3-D culture model, combined with its complementary parent orthotopic ICC, syngeneic rat model (11), will also prove to be invaluable as preclinical platforms for rapidly assessing mechanism-based therapeutic strategies targeting interactive pathways correlated with ICC progression. Future studies are now needed to validate these complementary in vitro and in vivo syngeneic models for use in identifying and testing molecular targeting strategies for ICC therapy. Moreover, our current study also suggests the feasibility of establishing an organotypic model of human desmoplastic cholangiocarcinoma potentially suitable for screening therapeutic agents that would be similarly based on 3-D culturing within type I collagen matrix of cholangiocarcinoma and α-SMA-positive CAF cell lines/stains simultaneously established from primary cell isolates derived from surgically resected human ICC tissues. In this context, establishment of primary cultures of α-SMA-positive CAF from human ICCs has been described (7, 27), and methods to generate cholangiocarcinoma cell lines from resected human ICC tissues are now well established. Thus, 3-D organotypic modeling of human desmoplastic ICC now seems plausible and should be pursued.
Supplementary Material
Differential integrin (itg) gene expression profiles assessed by microarray analysis for BDEsp-TDEH10 cholangiocarcinoma cells versus BDEsp-TDFE4 CAFs individually cultured for 7 days in rat tail type I collagen gel matrix. Bar values indicate means ± SD
TaqMan® QRT-PCR analysis of select biomarker genes differentiating BDEsp-TDEH10 cholangiocarcinoma cells from BDEsp-TDFE4 CAFs individually cultured in type I collagen gel matrix. These data validate the differential gene expression results obtained by microarray analysis (shown in Figure 2) for the cholangiocarcinoma cell markers [Krt19, Muc1, Mmp7, Areg] compared with gene markers exclusively or more highly expressed differentially in the CAFs [Acta2 (α-SMA), Hgf, Postn, Sphk1]. Bar values indicate means ± SD.
TaqMan® QT-PCR analysis demonstrating differential expression of Hedgehog (Hh) pathway genes in cultured BDEsp-TDEH10 cholangiocarcinoma cells versus BDEsp-TDFE4 CAFs. (A) Cultured BDEsp-TDEH10 cells express a significantly higher level of gene expression for the Hh ligand sonic hedgehog homolog (Shh) than that of cultured BDEsp-TDFE4 cells. (B) Conversely, mRNA for the Shh receptor Smoothened (SMO) is constitutively expressed at a significantly higher level in cultured BDEsp-TDFE4 cells over that of cultured BDEsp-TDEH10 cells. (C) Cultured BDEsp-TDFE4 cells also express significantly higher mRNA levels of both Gli1 and Gli2 (transcription factors ultimately activated by Shh binding to SMO, that regulate the transcription of Hh target genes) over those measured in the cholangiocarcinoma cell lines.
Acknowledgments
Grant Support: Supported by NIH grants R01 CA 039225 and R01 CA 083650 (A.E.S.)
Footnotes
Conflicts of Interest: None
References
- 1.Terada T, Makimoto K, Terayama N, et al. Alpha-smooth muscle actin-positive stromal cells in cholangiocarcinomas, hepatocellular carcinomas and metastatic liver carcinomas. J Hepatol. 1996;24:706–712. doi: 10.1016/s0168-8278(96)80267-4. [DOI] [PubMed] [Google Scholar]
- 2.Okamura N, Yoshida M, Shibuya A, et al. Cellular and stromal characteristics in the scirrhous hepatocellular carcinoma: comparison with hepatocellular carcinomas and intrahepatic cholangiocarcinomas. Pathol Int. 2005;55:724–731. doi: 10.1111/j.1440-1827.2005.01891.x. [DOI] [PubMed] [Google Scholar]
- 3.Sirica AE, Dumur CI, Campbell DJ, et al. Intrahepatic cholangiocarcinoma progression: prognostic factors and basic mechanisms. Clin Gastroenterol Hepatol. 2009;7:S68–S78. doi: 10.1016/j.cgh.2009.08.023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Sirica AE, Campbell DJ, Dumur CI. Cancer-associated fibroblasts in intrahepatic cholangiocarcinoma. Curr Opin Gastroenterol. 2011;27:276–284. doi: 10.1097/MOG.0b013e32834405c3. [DOI] [PubMed] [Google Scholar]
- 5.Sirica AE. The role of cancer-associated myofibroblasts in intrahepatic cholangiocarcinoma. Nat Rev Gastroenterol Hepatol. 2012;9:44–54. doi: 10.1038/nrgastro.2011.222. [DOI] [PubMed] [Google Scholar]
- 6.Okabe H, Beppu T, Hayashi H, et al. Hepatic stellate cells may relate to progression of intrahepatic cholangiocarcinoma. Ann Surg Oncol. 2009;16:2555–2564. doi: 10.1245/s10434-009-0568-4. [DOI] [PubMed] [Google Scholar]
- 7.Chuaysri C, Thuwajit P, Paupairoj A, et al. Alpha-smooth muscle actin-positive fibroblasts promote biliary cell proliferation and correlate with poor survival in cholangiocarcinoma. Oncol Rep. 2009;21:957–969. doi: 10.3892/or_00000309. [DOI] [PubMed] [Google Scholar]
- 8.Murakami K, Abe T, Miyazawa M, et al. Establishment of a new human cell line, LI90, exhibiting characteristics of hepatic Ito (fat-storing) cells. Lab Invest. 1995;72:731–739. [PubMed] [Google Scholar]
- 9.Xu L, Hui AY, Albanis E, et al. Human hepatic stellate cell lines, LX-1 and LX-2: new tools for analysis of hepatic fibrosis. Gut. 2005;54:142–151. doi: 10.1136/gut.2004.042127. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Okabe H, Beppu T, Hayashi H, et al. Hepatic stellate cells accelerate the malignant behavior of cholangiocarcinoma cells. Ann Sur Oncol. 2011;18:1175–1184. doi: 10.1245/s10434-010-1391-7. [DOI] [PubMed] [Google Scholar]
- 11.Sirica AE, Zhang Z, Lai GH, et al. A novel “patient-like” model of cholangiocarcinoma progression based on bile duct inoculation of tumorigenic rat cholangiocyte cell lines. Hepatology. 2008;47:1178–1190. doi: 10.1002/hep.22088. [DOI] [PubMed] [Google Scholar]
- 12.Dumur CI, Campbell DJ, DeWitt JL, et al. Differential gene expression profiling of cultured neu-transformed versus spontaneously-transformed rat cholangiocytes and of corresponding cholangiocarcinomas. Exp Mol Pathol. 2010;89:227–235. doi: 10.1016/j.yexmp.2010.08.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Lai GH, Zhang Z, Shen XN, et al. erbB-2/neu transformed rat cholangiocytes recapitulate key cellular and molecular features of human bile duct cancer. Gastroenterology. 2005;129:2047–2057. doi: 10.1053/j.gastro.2005.10.010. [DOI] [PubMed] [Google Scholar]
- 14.Sirica AE, Gainey TW. A new rat bile ductular epithelial cell culture model characterized by the appearance of polarized bile ducts in vitro. Hepatology. 1997;26:537–549. doi: 10.1002/hep.510260302. [DOI] [PubMed] [Google Scholar]
- 15.Lai GH, Radaeva S, Nakamura T, et al. Unique epithelial cell production of hepatocyte growth factor/scatter factor by putative precancerous intestinal metaplasias and associated “intestinal-type” biliary cancer chemically induced in rat liver. Hepatology. 2000;31:1257–1265. doi: 10.1053/jhep.2000.8108. [DOI] [PubMed] [Google Scholar]
- 16.Ohira S, Sasaki M, Harada K, et al. Possible regulation of migration of intrahepatic cholangiocarcinoma cells by interactions of CXCR4 expressed in carcinoma cells with tumor necrosis factor-α and stromal-derived factor-1 released in stroma. Am J Pathol. 2006;168:1155–1168. doi: 10.2353/ajpath.2006.050204. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Irizarry RA, Bolstad BM, Collin F, et al. Summaries of Affymetrix GeneChip probe level data. Nucleic Acids Res. 2003;31:e15. doi: 10.1093/nar/gng015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Storey JD. A direct approach to false discovery rates. JR Stat Soc B. 2002;64:479–498. [Google Scholar]
- 19.Sirica AE, Nathanson MH, Gores GJ, et al. Pathobiology of biliary epithelia and cholangiocarcinoma: proceedings of the Henry M. and Lillian Stratton Basic Research Single-Topic Conference. Hepatology. 2008;48:2040–2046. doi: 10.1002/hep.22623. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Park SY, Roh SJ, Kim YN, et al. Expression of MUC1, MUC2, MUC5AC and MUC6 in cholangiocarcinoma: prognostic impact. Oncology Reports. 2009;22:649–657. doi: 10.3892/or_00000485. [DOI] [PubMed] [Google Scholar]
- 21.Esencay M, Newcomb EW, Zagzag D. HGF upregulates CXCR4 expression in gliomas via NF-κB: implications for glioma cell migration. J Neurooncol. 2010;99:33–40. doi: 10.1007/s11060-010-0111-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Chioni A-M, Grose R. Organotypic modelling as a means of investigating epithelial-stromal interatcions during tumourigenesis. Fibrogenesis Tissue Repair. 2008;1:8. doi: 10.1186/1755-1536-1-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Froeling FEM, Mirza TA, Feakins RM, et al. Organotypic culture model of pancreatic cancer demonstrates that stromal cells modulate E-cadherin, β-catenin, and Ezrin expression in tumor cells. Am J Pathol. 2009;175:636–648. doi: 10.2353/ajpath.2009.090131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Froeling FEM, Marshall JF, Kocher HM. Pancreatic cancer organotypic cultures. J Biotechnol. 2010;148:16–23. doi: 10.1016/j.jbiotec.2010.01.008. [DOI] [PubMed] [Google Scholar]
- 25.De Wever O, Hendrix A, De Boeck A, et al. Modeling and quantification of cancer cell invasion through collagen type I matrices. Int J Dev Biol. 2010;54:887–896. doi: 10.1387/ijdb.092948ow. [DOI] [PubMed] [Google Scholar]
- 26.Matsumoto K, Nakamura T. Hepatocyte growth factor and the met system as a mediator of tumor-stromal interactions. Int J Cancer. 2006;119:477–483. doi: 10.1002/ijc.21808. [DOI] [PubMed] [Google Scholar]
- 27.Utispan K, Thuwajit P, Abiko Y, et al. Gene expression profiling of cholangiocarcinoma-derived fibroblasts reveals alterations related to tumor progression and indicates periostin as a poor prognostic marker. Mol Cancer. 2010;9:13. doi: 10.1186/1476-4598-9-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Differential integrin (itg) gene expression profiles assessed by microarray analysis for BDEsp-TDEH10 cholangiocarcinoma cells versus BDEsp-TDFE4 CAFs individually cultured for 7 days in rat tail type I collagen gel matrix. Bar values indicate means ± SD
TaqMan® QRT-PCR analysis of select biomarker genes differentiating BDEsp-TDEH10 cholangiocarcinoma cells from BDEsp-TDFE4 CAFs individually cultured in type I collagen gel matrix. These data validate the differential gene expression results obtained by microarray analysis (shown in Figure 2) for the cholangiocarcinoma cell markers [Krt19, Muc1, Mmp7, Areg] compared with gene markers exclusively or more highly expressed differentially in the CAFs [Acta2 (α-SMA), Hgf, Postn, Sphk1]. Bar values indicate means ± SD.
TaqMan® QT-PCR analysis demonstrating differential expression of Hedgehog (Hh) pathway genes in cultured BDEsp-TDEH10 cholangiocarcinoma cells versus BDEsp-TDFE4 CAFs. (A) Cultured BDEsp-TDEH10 cells express a significantly higher level of gene expression for the Hh ligand sonic hedgehog homolog (Shh) than that of cultured BDEsp-TDFE4 cells. (B) Conversely, mRNA for the Shh receptor Smoothened (SMO) is constitutively expressed at a significantly higher level in cultured BDEsp-TDFE4 cells over that of cultured BDEsp-TDEH10 cells. (C) Cultured BDEsp-TDFE4 cells also express significantly higher mRNA levels of both Gli1 and Gli2 (transcription factors ultimately activated by Shh binding to SMO, that regulate the transcription of Hh target genes) over those measured in the cholangiocarcinoma cell lines.