Abstract
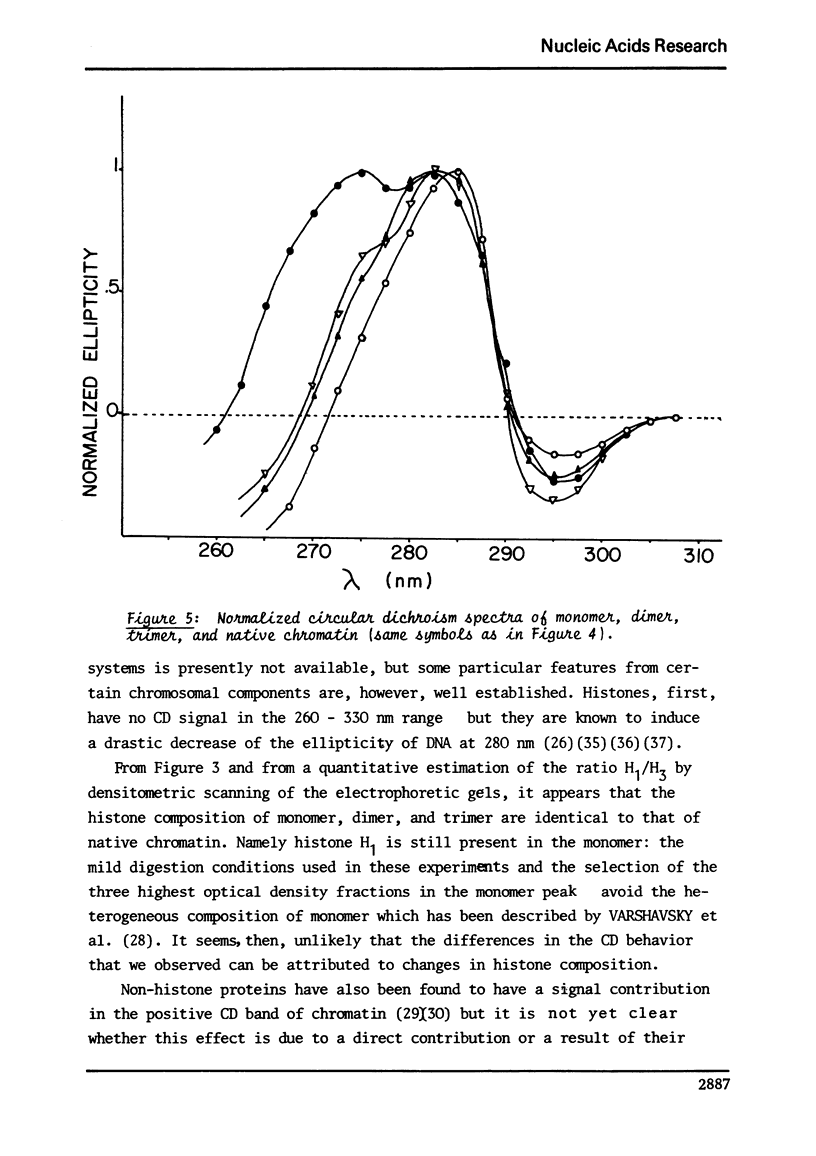
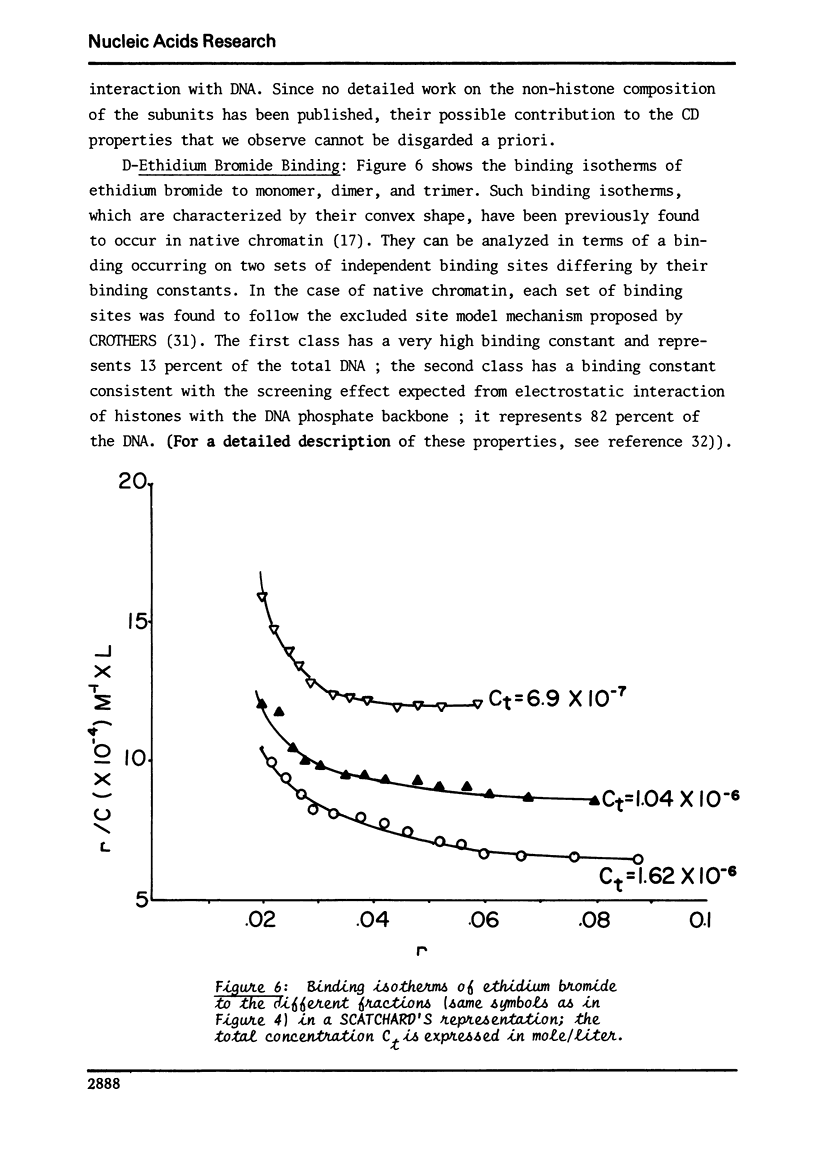
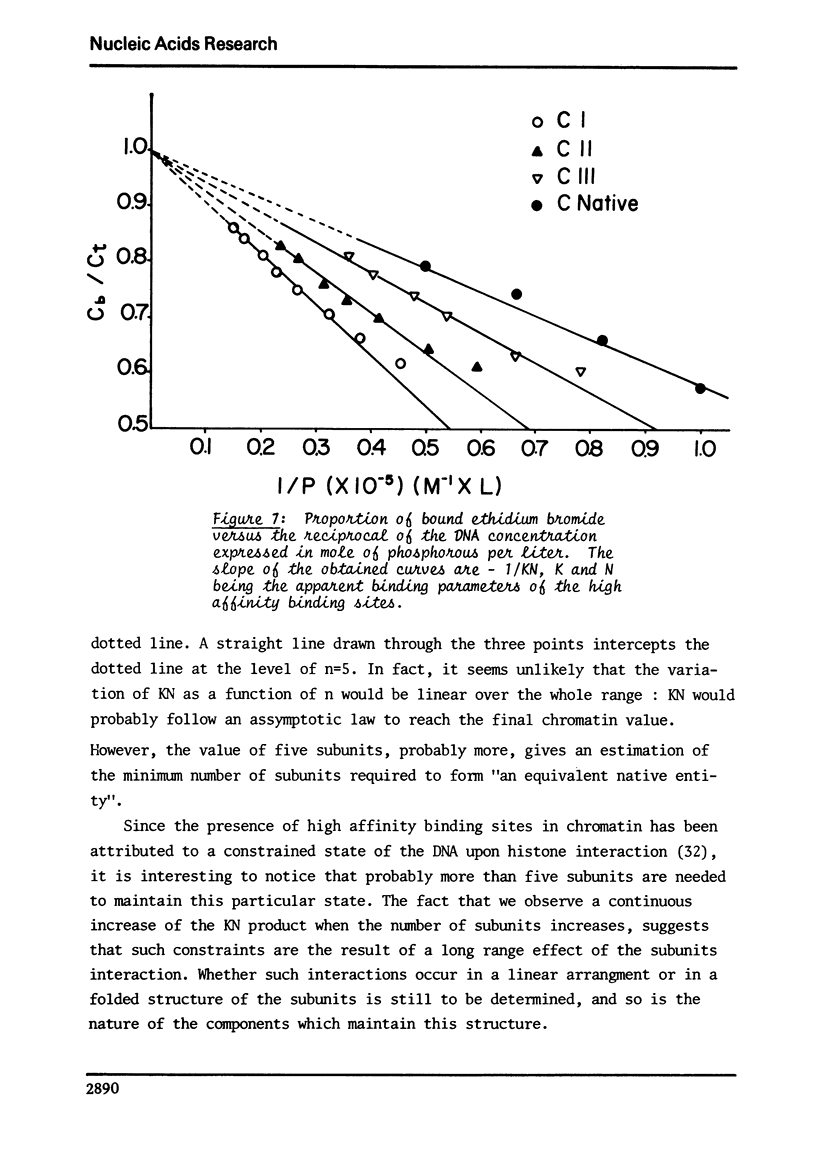
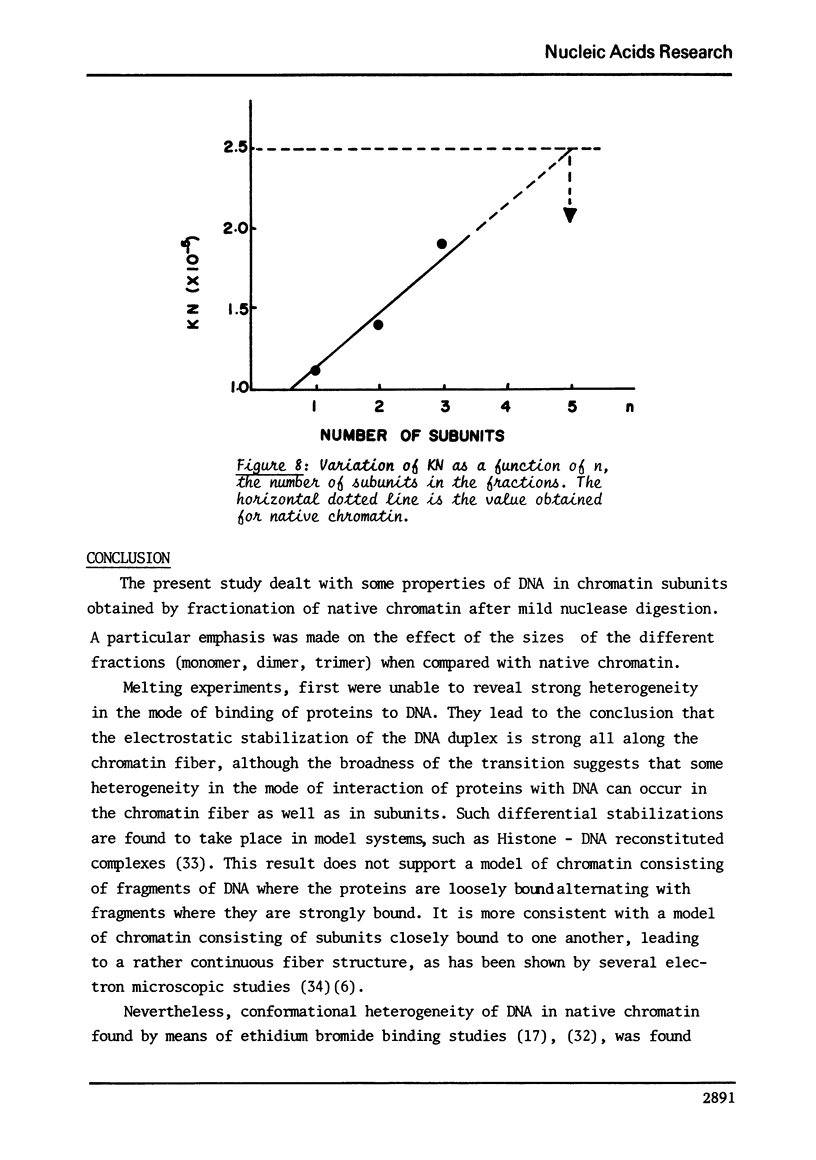
This study compares some physical properties of DNA in native chromatin and mono-, di-, trinucleosomes obtained after mild micrococcal nuclease digestion. Melting curves and derivatives are shown to be very similar from one sample to another although a shift from 79 to 82 degrees C is observed between the mainly monophasic peak of multimers and chromatin. Careful analysis of the positive band of the circular dichroism spectra shows the appearance of a shoulder at 275nm, the intensity of which increases from the mono- to the di- and trinucleosome. This shoulder is maximum for native chromatin. At the same time binding isotherms of ethidium - bromide are characterized by two highly fluorescent binding sites for all the samples but the product KN of the apparent binding constant of the higher affinity binding sites by the apparent number of those sites increases from the mono- to the di- and trinucleosome. There again the valus is maximum for native chromatin. Such results strongly suggest that the native state of chromatin requires something more than the indefinite repeat of an elementary subunit.
Full text
PDF



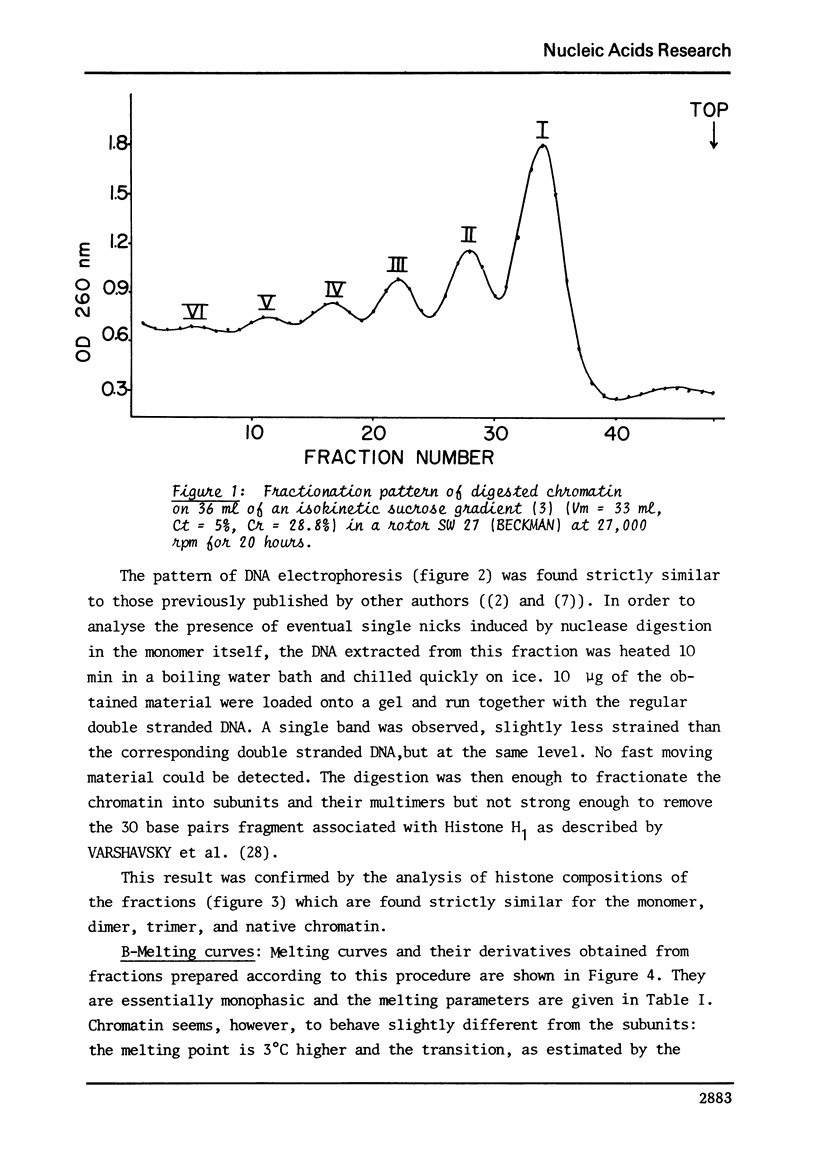
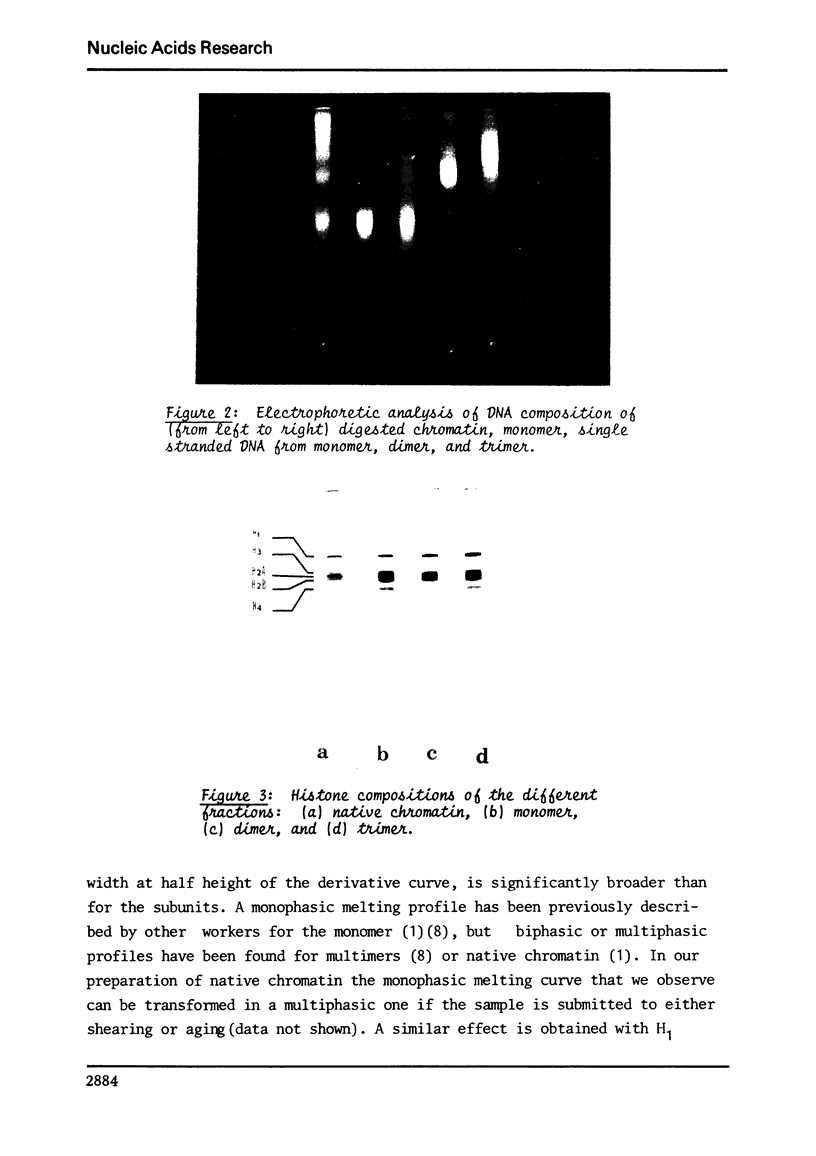
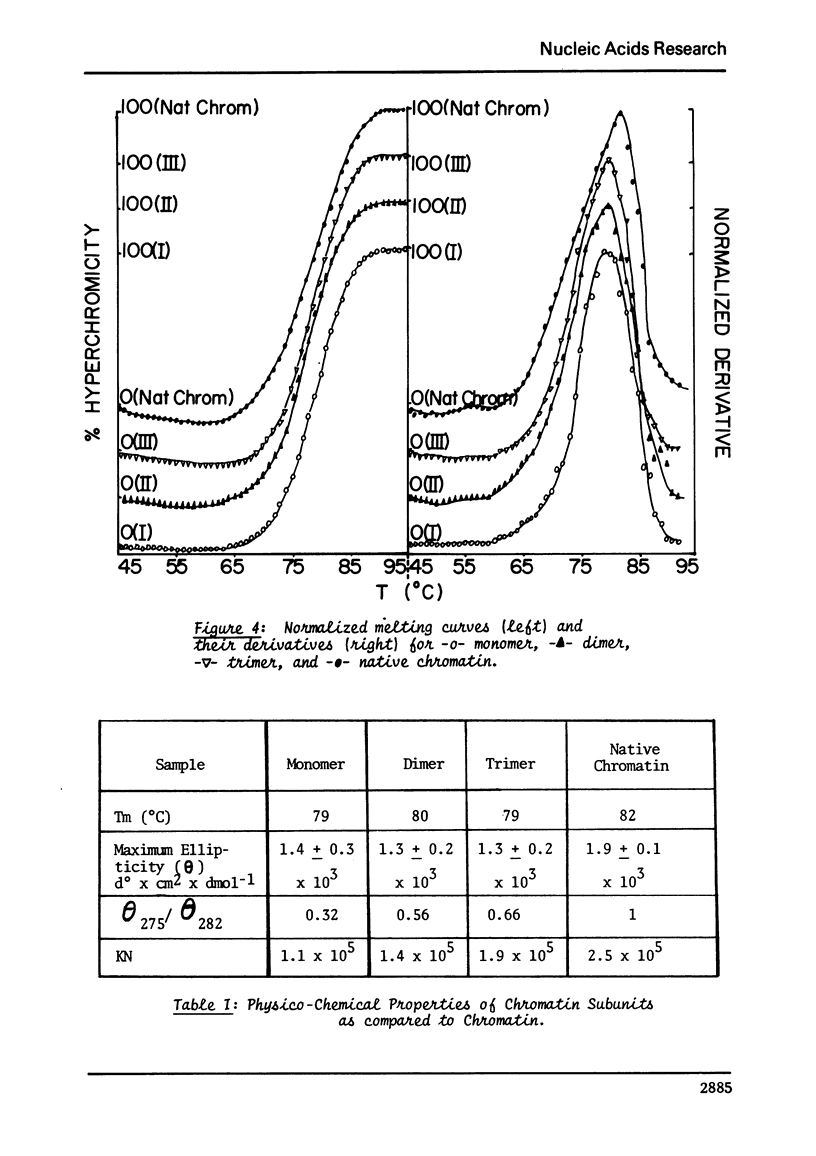




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Finch J. T., Noll M., Kornberg R. D. Electron microscopy of defined lengths of chromatin. Proc Natl Acad Sci U S A. 1975 Sep;72(9):3320–3322. doi: 10.1073/pnas.72.9.3320. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lawrence J. J., Daune M. Ethidium bromide as a probe of conformational heterogeneity of DNA in chromatin. The role of histone H1. Biochemistry. 1976 Jul 27;15(15):3301–3307. doi: 10.1021/bi00660a021. [DOI] [PubMed] [Google Scholar]
- Oudet P., Gross-Bellard M., Chambon P. Electron microscopic and biochemical evidence that chromatin structure is a repeating unit. Cell. 1975 Apr;4(4):281–300. doi: 10.1016/0092-8674(75)90149-x. [DOI] [PubMed] [Google Scholar]
- Panyim S., Chalkley R. High resolution acrylamide gel electrophoresis of histones. Arch Biochem Biophys. 1969 Mar;130(1):337–346. doi: 10.1016/0003-9861(69)90042-3. [DOI] [PubMed] [Google Scholar]
- Ris H., Kubai D. F. Chromosome structure. Annu Rev Genet. 1970;4:263–294. doi: 10.1146/annurev.ge.04.120170.001403. [DOI] [PubMed] [Google Scholar]
- Shaw B. R., Herman T. M., Kovacic R. T., Beaudreau G. S., Van Holde K. E. Analysis of subunit organization in chicken erythrocyte chromatin. Proc Natl Acad Sci U S A. 1976 Feb;73(2):505–509. doi: 10.1073/pnas.73.2.505. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Varshavsky A. J., Bakayev V. V., Georgiev G. P. Heterogeneity of chromatin subunits in vitro and location of histone H1. Nucleic Acids Res. 1976 Feb;3(2):477–492. doi: 10.1093/nar/3.2.477. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weintraub H., Palter K., Van Lente F. Histones H2a, H2b, H3, and H4 form a tetrameric complex in solutions of high salt. Cell. 1975 Sep;6(1):85–110. doi: 10.1016/0092-8674(75)90077-x. [DOI] [PubMed] [Google Scholar]
- Wilhelm F. X., Champagne M. H., Daune M. P. Conformation du DNA dans la nucléoprotéine. Eur J Biochem. 1970 Aug;15(2):321–330. doi: 10.1111/j.1432-1033.1970.tb01010.x. [DOI] [PubMed] [Google Scholar]
- Woodcock C. L., Frado L. L. Thermal denaturation of subchromosomal particles. Biochem Biophys Res Commun. 1975 Sep 2;66(1):403–410. doi: 10.1016/s0006-291x(75)80342-1. [DOI] [PubMed] [Google Scholar]
- Yu S. S., Li H. J., Shih T. Y. Interactions between arginine-rich histones and deoxyribonucleic acids. I. Thermal denaturation. Biochemistry. 1976 May 18;15(10):2027–2034. doi: 10.1021/bi00655a001. [DOI] [PubMed] [Google Scholar]