Abstract
The sensory patches in the ear of a vertebrate can be compared with the mechanosensory bristles of a fly. This comparison has led to the discovery that lateral inhibition mediated by the Notch cell–cell signaling pathway, first characterized in Drosophila and crucial for bristle development, also has a key role in controlling the pattern of sensory hair cells and supporting cells in the ear. We review the arguments for considering the sensory patches of the vertebrate ear and bristles of the insect to be homologous structures, evolved from a common ancestral mechanosensory organ, and we examine more closely the role of Notch signaling in each system. Using viral vectors to misexpress components of the Notch pathway in the chick ear, we show that a simple lateral-inhibition model based on feedback regulation of the Notch ligand Delta is inadequate for the ear just as it is for the fly bristle. The Notch ligand Serrate1, expressed in supporting cells in the ear, is regulated by lateral induction, not lateral inhibition; commitment to become a hair cell is not simply controlled by levels of expression of the Notch ligands Delta1, Serrate1, and Serrate2 in the neighbors of the nascent hair cell; and at least one factor, Numb, capable of blocking reception of lateral inhibition is concentrated in hair cells. These findings reinforce the parallels between the vertebrate ear and the fly bristle and show how study of the insect system can help us understand the vertebrate.
Almost all animals, from cnidarians (1) to mammals, have mechanosensory organs for touch and detection of vibrations and other disturbances of the air or water in which they live. This sensory capability, it seems, is as important and as universal as sensitivity to light, suggesting that the apparatus of mechanosensation, like that of photoreception (2), may have a very ancient evolutionary origin. Mechanosensory organs such as the ear may be elaborate and highly specialized according to the animal's way of life, but at their core they must always have a set of mechanosensory transducer cells to perform the fundamental task. At the level of these cells and their immediate companions one may hope to find conserved features reflecting evolution from a common prototype: homologous cell types, homologous developmental processes, and homologous molecular mechanisms.
In the vertebrate ear, the core structures are the sensory patches, consisting of hair cells (the transducers), supporting cells (which form the epithelial framework in which hair cells are held), and the adjacent cochleovestibular sensory neurons (which synapse with the hair cells). How are these cell types, especially the hair cells and supporting cells, generated in the correct pattern and proportions? Parallels with Drosophila provide a route toward an answer: the sensory patches in the vertebrate ear have a counterpart in the sensory bristles of the fly, suggesting that homologous mechanisms may operate (3, 4). This approach has revealed that lateral inhibition mediated by the Notch signaling pathway (5), a key mechanism for controlling cell diversification in fly sense organs, has a similarly crucial role in the vertebrate inner ear.
In the first part of this paper we review the published evidence on Notch signaling in the ear and see how it fits with a simple model (6) that has been proposed to explain the patterning of the ear's sensory patches (7). This model does not, however, correspond accurately to the way in which Notch signaling governs development of sensory bristles in the fly (8–10). Are the vertebrate ear and the insect bristle not so closely homologous after all, or is the simple model proposed for the ear misleading? We review the arguments for homology, and in the second half of the paper we present experimental evidence to test the role of Notch signaling in the ear. We show that Notch signaling in the ear is more complex than was originally suspected: at least three Notch ligands are at work; they are regulated in contrary and complementary ways; and, contrary to previous suggestions, the pattern of cell determination cannot simply be explained in terms of lateral inhibition rules governing Notch-ligand expression. These corrections to previous ideas, far from undermining the arguments for homology with Drosophila sensilla, make the parallels seem even closer than before.
The Hair Cells, Supporting Cells, and Neurons of a Sensory Patch Have a Common Origin in the Otic Placode.
The inner ear derives from the otic placode, a thickening of the epidermis adjacent to the hindbrain in the early embryo (11–13). This placode gives rise both to the inner ear epithelium, with its sensory patches consisting of hair cells and supporting cells, and to the sensory neurons that innervate these patches. The neuronal lineage becomes segregated from the sensory epithelial lineage at an early stage: as the otic placode invaginates and forms first a cup and then a vesicle, neuroblasts become singled out within the otic epithelium and begin to delaminate from its anteroventral portion (4). These cells will divide a variable number of times before differentiating as cochleovestibular neurons. In the epithelium, the future sensory patches become identifiable by their expression of markers such as Serrate1 (Ser1) and BMP4 (4, 14), but differentiation of hair cells and supporting cells does not begin until 3–5 days later (in birds and mammals).
Although the lineage relationship between the neuroblasts and the hair and supporting cells has not been rigorously determined, it seems that the anteroventral region of the early otocyst is a source for all three cell types, while a more posteromedial region gives rise only to hair cells and supporting cells (4). It has, however, been shown that hair cells and supporting cells have the same ancestry and frequently arise as pairs of sister cells (15–17). Given that these two cell types have a common origin and lie intermingled in the otic epithelium, sharing a common environment, what causes them to be different? How is the correct mixture of hair cells and supporting cells generated?
The Alternating Mosaic of Hair Cells and Supporting Cells Suggests a Simple Lateral Inhibition Model for Genesis of Spatial Pattern.
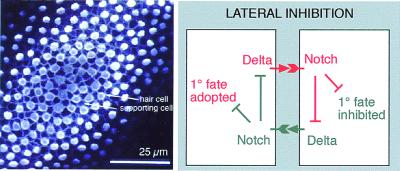
The ratio of hair cells to supporting cells in a mature sensory patch is variable from one region to another. In the auditory epithelium (the basilar papilla) of the 12-day chicken embryo, for example, it ranges from 1:1.7 at the distal end to 1:3.9 at the inferior-proximal end (18). Despite this variation, the distribution of hair cells among supporting cells seems almost everywhere to obey the same simple rule: with few exceptions, every cell that lies in contact with a hair cell is a supporting cell, and any cell that escapes all contact with hair cells is itself a hair cell. This immediately suggests a pattern-generating mechanism based on lateral inhibition (Fig. 1) (3, 19). According to this hypothesis, the cells in the undifferentiated sensory patch have a choice of two fates (hair cell or supporting cell), and the choice is governed by interactions between nearest neighbors. The default or primary fate, in the absence of interaction, is to become a hair cell, but in the interacting system each nascent hair cell delivers an inhibitory signal to the cells in contact with it, deterring these from becoming hair cells, too, and at the same time preventing them from producing inhibitory signals that would act back on the nascent hair cell. Neighbors thus compete to be hair cells, and the losers in the competition, the cells surrounding hair cells, become supporting cells.
Figure 1.
(Left) An optical section (confocal image) in the plane of the chick basilar papilla, showing the mosaic of hair cells (stained with HCA antibody, white blobs) and supporting cells (outlined by their cortical actin, stained with fluorescent phalloidin). (Reproduced from ref. 4.) (Right) Lateral inhibition with feedback: a simple model of Delta-Notch signaling that can generate spatial pattern. Only two cells are shown, interacting to adopt two different fates, but the same principles apply to a field of many cells, which by interacting in this way can in theory give rise to the type of pattern seen at Left.
Lateral inhibition of this sort is well documented in Drosophila and Caenorhabditis elegans, and studies in these species have revealed the molecular mechanism that transmits the inhibitory signal (5, 20). The receptor on the cell receiving inhibition is the transmembrane protein Notch, and the ligand on the adjacent cell that delivers inhibition is the transmembrane protein Delta. Loss-of-function mutations in the Delta–Notch signaling pathway abolish lateral inhibition and allow an excessive proportion of the population to adopt the primary fate.
On this basis, a simple formal model for pattern generation can be proposed (21–23): an increase of Delta in one cell causes increased activation of Notch in its neighbor, and Notch activation in the neighbor down-regulates Delta expression in that same cell, as well as inhibiting commitment to the primary fate (Fig. 1). The feedback control of Delta expression has the effect of amplifying contrasts between adjacent cells: a sheet of initially similar cells, with only small random differences between them, will spontaneously develop into a mosaic of alternate cell types of just the type seen in the sensory patches of the ear (6).
Lateral Inhibition in the Sensory Patches of the Ear Is Mediated by the Notch Signaling Pathway.
In the vertebrate central nervous system, or at least in the neural plate (24, 25) and retina (26), the production of neurons from neuroepithelial progenitor cells seems to be controlled in just the way that the simple lateral-inhibition-with-feedback model proposes. All of the progenitors express the Notch homolog Notch1, while nascent neurons appear within this population as scattered cells expressing the Delta homolog Delta1 (Dl1) at higher levels than their neighbors. When Dl1 is artificially overexpressed, so that all cells strongly deliver and receive the inhibitory signal, the cells all are prevented from differentiating as neurons. Conversely, when all of the cells are forced to express a dominant-negative form of Dl1, blocking Delta–Notch signaling, they all escape inhibition and differentiate as neurons prematurely. Moreover, expression of the endogenous Dl1 gene is regulated in accordance with the simple feedback model. All of the requirements for pattern generation by lateral inhibition with feedback, and all of the predictions of the model, seem to be satisfied.
To see whether the same is true in the ear, the first step is to examine the normal gene expression patterns in the developing sensory patches (4, 27, 28). Notch1 is expressed, as in the central nervous system, throughout the population of progenitor cells (and in fact even more widely in the otic epithelium). Dl1 is expressed in a scattered subset of the population, and these Dl1-expressing cells can be identified as the nascent hair cells (4, 27). These observations are backed up by similar circumstantial evidence for Notch signaling during hair-cell regeneration in birds (63).
Direct functional evidence comes from zebrafish and mice. Thus in zebrafish with a mutation in the deltaA gene—one of four zebrafish Delta homologs, all normally expressed in nascent hair cells—hair cells are produced in increased numbers, as though inhibition has been reduced (29). Likewise, there is some overproduction of hair cells in mice with a knockout mutation in a gene coding for another Notch ligand, Serrate2 (also known as Jagged2, but henceforward in this paper Serrate2 to match chick and Drosophila terminology) (28). Serrate2 (Ser2) is a homolog of the Drosophila Serrate gene, closely related to Delta and likewise coding for a protein capable of activating Notch; in the sensory patches of the ear it is expressed with Dl1 in the nascent hair cells (although more persistently).
The phenotypes of the deltaA and Ser2 mutations are relatively mild but consistent with the model. The zebrafish mind bomb mutant provides more spectacular evidence. This mutant takes its name from its neurogenic phenotype: in the central nervous system, it grossly overproduces neurons at the expense of neural progenitor cells, in a manner that is diagnostic of a failure of lateral inhibition. The excess of neurons goes with an up-regulation of Delta expression: the cells are deaf to the inhibitory signal that Delta normally delivers. In the ear, the result of this failure of Notch signaling is dramatic: each of the prospective sensory patches differentiates in its entirety into a uniform mass of hair cells, with no supporting cells (7). The hair cells are not only produced in great excess, as much as 30-fold, but also prematurely. In the absence of supporting cells to hold them in place, the hair cells all are then extruded from the epithelium and die and disappear within a day or two after their first appearance (30).
The mind bomb phenotype tells us that Notch signaling is required (i) to prevent the cells in a developing sensory patch from all differentiating alike as hair cells, and (ii) to delay production of hair cells until the proper time. It shows that lateral inhibition mediated by the Notch pathway is essential, but it falls short of proving that lateral inhibition with feedback is the mechanism that generates the normal orderly mixture of hair cells and supporting cells. Indeed, there are several reasons to be cautious in adopting this simple model mechanism, seductive as it may seem. Goodyear and Richardson (18) have shown, for example, that the orderliness in the basilar papilla of the chick depends to a large extent on cell rearrangements occurring after cell differentiation. While Dl1 and Ser2 are concentrated in nascent hair cells, there is yet another Notch ligand, Ser1, that is strongly expressed in supporting cells (4, 27, 31). Most significantly, in the sensory bristles of Drosophila, Delta expression is not regulated in the way the model postulates, even though lateral inhibition via Notch is as essential as in the ear (8, 9). Is the Drosophila bristle a misleading paradigm, or is the simple theoretical model wrong? To answer these questions, we first review the parallels between insect bristles and the sensory patches of the vertebrate ear.
The Sensory Patches of the Vertebrate Ear Resemble the Sensilla of a Fly in Function and Developmental History.
Each insect bristle is a miniature sense organ, or sensillum, consisting of a set of four cells: a neuron, a neural sheath cell, a bristle socket cell, and a bristle shaft cell. These cells normally all derive, along with a migratory glial cell (32), from a single sensory mother cell, or SMC. Lateral inhibition mediated by Notch operates repeatedly in development of the bristle, first to single out the SMC within a cluster of competent cells in the epidermis, and then at each subsequent cell division to drive the diversification of the progeny of the SMC to form the differentiated cells of the bristle (8, 10, 33, 34).
The insect bristle resembles the sensory patch in the vertebrate ear in several respects. First of all, the mechanosensory function is essentially the same, with the same kinetics of response and adaptation (35).
Second, the cell types correspond, in part at least: neuron with neuron, bristle shaft cell with hair cell, bristle socket cell with ear supporting cell. Bundles of actin filaments form the shaft of the bristle shaft cell, just as bundles of actin filaments form the stereocilia of the hair cell (36). Both these cell types have a well-defined planar polarity, essential for directional sensitivity.
Third, the developmental anatomy is similar. Just as the component cells of the bristle have a common origin in the epidermis, so do the neurons, hair cells, and supporting cells of the ear have a common origin in the otic placode ectoderm. Just as the first division of the SMC gives rise to a neuronal precursor, which delaminates from the epidermis, and an epithelial precursor, which stays behind to generate the socket and shaft cells, so also the first step in differentiation of the sensory patch is production of neuroblasts, which delaminate from the otic epithelium, and sensory epithelial precursors, which stay behind to form hair cells and supporting cells. In both systems, the final differentiated cell types are the products of a series of dichotomous cell-fate choices, and this series of choices is similar in the fly sensillum and the vertebrate ear.
Lastly, and most crucially, the molecular mechanisms underlying these choices are similar.
Corresponding Events in Development of Insect Sensillum and Vertebrate Inner Ear Are Controlled by Homologous Systems of Genes.
In fly sensilla, basic helix–loop–helix (bHLH) transcription factors of the Achaete/Scute family and the related Atonal family have a key role, both in initiating the program of sensory development (the “proneural” function) and as differentiation factors for the final cell types (22). Examples of the latter include the products of the asense and cousin of atonal (cato) genes, which drive differentiation both in bristles and chordotonal organs (a closely related type of sensillum) (37). In the ear, although bHLH genes serving the early proneural function have not been identified, it has been shown that the atonal homolog Math1 is expressed selectively in hair cells and drives hair-cell differentiation: hair cells fail to develop in a Math1 knockout mouse (38), and nonsensory cells in the cochlea differentiate into hair cells when transfected with Math1 (39). Upstream from the proneural genes in the fly, controlling their domains of expression, lie transcription factors of the Iroquois family; these, too, have vertebrate homologs that are expressed at early stages in the future sensory epithelium of the ear (40).
Components of the Notch signaling pathway lie downstream from the proneural genes and play a central and recurrent role in the subsequent development of the fly sensillum. In each of the series of cell-fate decisions in the sensillum lineage, from the singling out of the SMC onward, lateral inhibition mediated by Notch signaling is required (8, 33, 34).
The Notch signaling pathway has a similarly central and recurrent role in the development of the sensory patches of the vertebrate ear. Notch1 is expressed throughout the otic placode at the very beginning of ear development and thereafter throughout the developing sensory epithelium, persisting into adult life in the supporting cells (4, 16). Dl1 is expressed at each of the sites where cell-fate choices are being made (4). During the segregation of neural and epithelial sublineages of the ear, Dl1 RNA is seen in scattered cells in the neurogenic region of otic epithelium, apparently the nascent neuroblasts. Subsequently it is expressed in the ganglion formed by the neuroblasts, and, as we have seen, in the sensory-patch epithelium as hair cells are being generated and becoming different from supporting cells. The block of Notch signaling in the mind bomb mutant not only causes overproduction of hair cells, but also neurons (7).
Finally, the molecular data reveal additional similarities between the differentiated cell types. In particular, the Pax gene D-pax2 is expressed specifically in the bristle shaft cell and required for its correct differentiation (41); the vertebrate homolog Pax2 is expressed in the early otic epithelium and then selectively in hair cells and also is required for their correct differentiation. In the mouse Pax2 knockout, no cochlea forms (42), and in a zebrafish pax2.1 mutant, hair-cell differentiation is abnormal (29).
The parallels between insect bristles and ear sensory patches that we have summarized above add up to a persuasive argument that these mechanosensory organs are indeed homologous—that they resemble one another because they have evolved from a common ancestral prototype. There are, of course, also important differences, as one might expect after 800 million years of divergent evolution. The fly bristle is ensheathed in a semirigid cuticle, making the mechanics of stimulus delivery quite different from that in the ear. In the ear, the hair cells are the transducers and synapse with the neurons; in the bristle, there is no synapse and the dendrite of the neuron acts as the transducer. In the development of the ear, there is no step corresponding to the singling out of the SMCs from the epidermis: instead, the future sensory patch behaves like a uniform mass of contiguous SMCs. The numbers of cell divisions elapsing between one cell-fate choice and the next are different in the two systems, and the cell lineage patterns are not identical. Almost all of these developmental differences, however, correspond to plausible evolutionary variations and have parallels in differences between the various, but unmistakably related, types of sensilla in the fly (see ref. 4 for details).
The Pattern of Cell Types in the Insect Bristle Is Not Controlled Through Regulation of Delta Expression.
Notch signaling in the developing bristle shows three important departures from the simple lateral-inhibition model sketched in Fig. 1. First, recent studies have shown that Delta is not the only Notch ligand at work. Serrate is also present and must be mutated along with Delta to give the most extreme Notch-pathway loss-of-function phenotype (43). Second, Delta is not regulated in the way the simple model postulates, either at the mRNA or at the protein level. Delta expression remains high in the winning and losing cells during many of the lateral-inhibition interactions that decide cell fate (8, 9, 34), implying that some other factor(s) must create a difference by modulating the efficacy of Delta or the ability to respond to it. Third, factors acting in this way to bias Notch signaling have been identified. In particular, the intracellular protein Numb is distributed asymmetrically between the daughters and granddaughters of the SMC and makes the cells that contain it deaf to lateral inhibition (10, 44).
We now turn to our experimental observations on Notch signaling in the sensory patches of the chick ear.
Materials and Methods
Viral Constructs.
The RCAS-Dl1 and RCAS-Dl1dn replication-competent retroviral constructs were as described (26). RCAS-X-Su(H)dn was a gift from J.-C. Izpisua-Belmonte (Salk Institute, La Jolla, CA) and contains a form of the Xenopus Suppressor-of-Hairless [Su(H)] cDNA with its DNA binding mutated, as described (45). The viruses were used at a titer of 5 × 107-109 cfu/ml.
The pseudotype virus will be described in detail elsewhere (I.L.R., unpublished work). Briefly, plasmids based on the LZRSpBMN-Z plasmid (46) were prepared by inserting cDNA for the product of interest (Dl1 or Dl1dn) linked to an internal ribosome entry site followed by DNA coding for green fluorescent protein (GFP); this composite coding sequence was placed under the control of a 253-bp upstream enhancer sequence from Rous sarcoma virus, within LZRSpBMN-Z. Pseudotype virus then was generated by transiently cotransfecting 293gp packaging cells (Qiagen, Chatsworth, CA) with this construct plus a plasmid coding for vesicular stomatitis virus (VSV)-G protein. The resulting pseudotype virus particles contain RNA coding for Dl1 + GFP or Dl1dn + GFP, with Gag, Pol, and VSV-G proteins provided by the packaging cells. Virus released into the supernatant was concentrated by ultracentrifugation to a final titer of 5 × 108-109 cfu/ml.
Embryos and Chicken Injection.
Chick embryos were windowed at stage 13 [embryonic day 2 (E2)], and ≈0.5 μl of virus solution (with 0.8 μg/μl polybrene, 3% methyl cellulose, and a trace of fast green dye) was injected into the lumen of the otic cup. Embryos were fixed between E6 and E9.
In Situ Hybridization and Immunohistochemistry.
In situ hybridization was performed on 15-μm cryosections of fixed embryos as described (26), by using FastRed (Boehringer) for detection by fluorescence. For immunostaining, the cryosections were incubated overnight at 4°C in blocking solution (PBS with 3% BSA, 10% FCS, 0.1% Triton) containing the primary antibody. For Numb, sections were taken through an additional methanol fixation before the primary antibody was added. Chick anti-Numb antibody (47) was a gift from Y. Wakamatsu, Tohoku University, Japan. Ser1 was detected with a polyclonal antiserum as described (4). Dl1 and Dl1dn were detected with a rabbit polyclonal antiserum directed against amino acids 325–462 of chick Dl1 (26). In the specimens shown here, this antiserum was used at a concentration sufficient to detect the high levels of exogenous Dl1 or Dl1dn, but not endogenous Dl1. Hair cells were detected by using the hair-cell antigen (HCA) antibody (48). GFP was detected with a rabbit polyclonal antiserum (gift from D. Shima, Imperial Cancer Research Fund, London). Secondary antibodies were labeled with Alexa488 or Alexa594 (Molecular Probes). Images were collected by confocal microscopy. A total of ≈180 virus-injected embryos were serially sectioned and analyzed. Results are based on 47 embryos in which we saw informative patches of infection, i.e., patches that overlapped or touched sensory patches in the ear epithelium.
Results
Ser1 Is Regulated by Lateral Induction.
During the development of sensory patches in the chick ear, the Notch ligand Ser1 is expressed in a pattern quite different from that of Dl1 and Ser2. From a very early stage, it marks all of the cells in the prospective sensory patches, and although it is eventually down-regulated in hair cells as they differentiate, it remains high in all of the supporting cells (31). Because these are in contact with one another, this finding suggests that the expression of Ser1 is not regulated by lateral inhibition, which would tend to make levels different in neighboring cells. In the Drosophila wing margin, it has been found, however, that Notch activation also can regulate Notch ligand expression in a contrary way, by lateral induction, so that neighboring cells stimulate one another to express ligand strongly (49, 50). The pattern we see in the ear suggests that, while Dl1 and Ser2 may be regulated by lateral inhibition, Ser1 may be regulated by lateral induction.
To test this, it is necessary to interfere with Notch activation levels and see how Ser1 expression is altered. This can be achieved in the chick by infecting the cells with a viral vector that drives expression of a molecule that blocks Notch signaling. We have used two constructs, both based on the replication-competent RCAS virus (see Materials and Methods). One, RCAS-Dl1dn, contains the dominant-negative truncated form of Dl1, which makes cells that contain it deaf to Notch signaling (26). The other blocks Notch signaling by interfering with Su(H), a transcription factor through which Notch acts (5): the viral construct, RCAS-X-Su(H)dn, contains a mutated vertebrate homolog of Su(H) with a defective DNA binding domain; it thus also serves to block Notch signaling (45).
Embryos were injected with virus at 2 days and fixed 4–7 days later. Patches of infection within sensory regions of the otic epithelium were analyzed for Ser1 expression (Fig. 2). Ser1 expression was lost or clearly reduced in 8 of 10 patches of infection with RCAS-Dl1dn, and in 8 of 10 patches of infection with RCAS-X-Su(H)dn. A control for nonspecific effects of viral infection was provided by a parallel series of experiments (see below) using another virus, RCAS-Dl1, containing the full-length form of Dl1. In the majority of these cases (20 of 25 patches) no down-regulation of Ser1 was seen. Taken together, these data indicate that Ser1 is indeed positively regulated by Notch activity.
Figure 2.
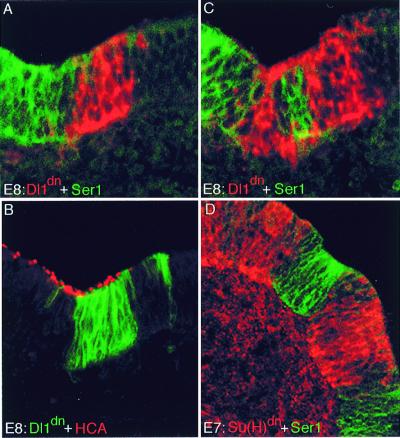
Blocking Notch signaling causes down-regulation of Ser1. (A–C) Infection with RCAS-Dl1dn virus, blocking signaling via Notch. Three consecutive sections through the utricle of an 8-day (E8) embryo are shown. The expression of Dl1dn is shown by in situ hybridization with a Dl1 probe (red) (A and C) and by antibody staining (green) (B). Ser1 expression is shown by antibody staining (green) (A and C); the distribution of hair cells is shown with the HCA antibody (detecting hair bundles, red) (B). Ser1 expression is lost at sites of infection. Presence of hair cells proves that these sites lie within a sensory patch, where normally Ser1 is expressed. (D) Infection with RCAS-XSu(H)dn. Infection was detected by antibody against viral gag protein (red) and Ser1 expression by Ser1 antibody (green). Again, the two stains are mutually exclusive: cells infected within the sensory patch down-regulate Ser1.
Effects of Dl1dn Expression on Hair Cell Differentiation Are Difficult to Decipher.
It might be expected that hair cells should be seen in excess in regions where Notch signaling was blocked with RCAS-Dl1dn or RCAS-X-Su(H)dn. We have examined hair cell production in our experiments with RCAS-Dl1dn, in which embryos were infected at 2 days of incubation and fixed 4 or 6 days later. Altogether, we found 33 potentially informative infected patches. Strikingly, just over half (17/33) of the informative patches of infection directly abutted sensory patches, without any overlap (Fig. 3 A and B): infection with the virus and sensory character appeared to be mutually exclusive. By comparison, in parallel experiments using the RCAS viral vector to misexpress full-length Dl1 (see below and Fig. 4), only 10% of informative infected patches abutted sensory patches in this way, as against 90% that overlapped with or were internal to sensory patches. At least two interpretations are possible. It could be that blockade of Notch signaling by RCAS-Dl1dn converted prospective sensory patch cells to a nonsensory character. Alternatively, the loss of Notch activity may have caused premature hair-cell differentiation as in mind bomb, and as in mind bomb this may have lead to early death and disappearance of any infected cells that lay in a prospective sensory patch (7, 30). Where an early infection partially overlapped a prospective sensory patch, the loss of cells in the region of overlap would bring infected nonsensory cells into juxtaposition with uninfected sensory cells.
Figure 3.
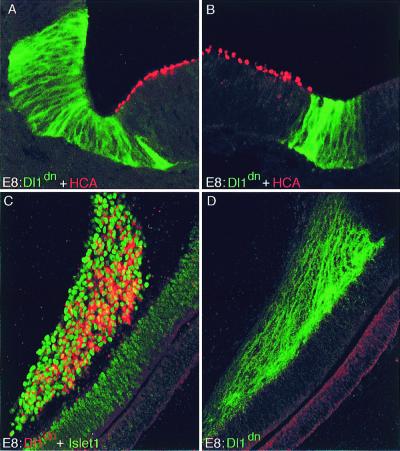
Infections with RCAS-Dl1dn virus. (A and B) Sections through utricle at E8; the Dl1dn-expressing patches abut but do not overlap sensory patches. Two examples are shown. Dl1dn expression is shown with an antibody against Dl1 (green), hair cells with HCA antibody (red). (C and D) Dl1dn-expressing cells often end up as neurons in the cochleovestibular ganglion. (C) Section of basilar papilla and underlying ganglion at E6 stained by in situ hybridization for Dl1dn (red) and with Islet1/2 antibody (green) to mark the nuclei of neurons in the cochleovestibular ganglion. (D) Adjacent section stained with Dl1 antibody, revealing the dendrites of infected neurons with Dl1dn in their membranes.
Figure 4.
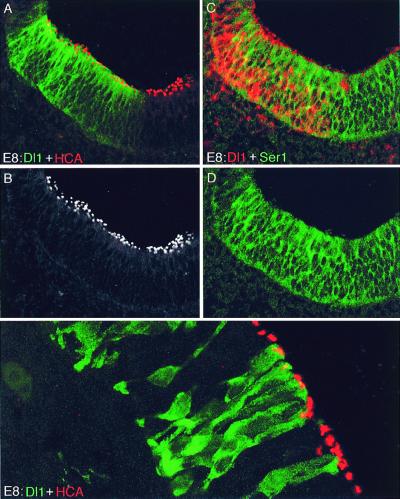
Overexpression of Dl1 in a sensory patch does not inhibit hair-cell production. (A–D) Sections of utricle at E8, infected with RCAS-Dl1 virus. (A) Infected patch stained with an antibody against Dl1 (green) and with HCA antibody (red). (B) The same scene, showing the HCA signal only, to reveal the distribution of hair cells more clearly. (C) Adjacent section stained by in situ hybridization for Dl1 (red) and with Ser1 antibody. (D) Same scene as in C, with the red fluorescence hidden to show that Dl1, in contrast with Dl1dn, does not cause down-regulation of Ser1. (E) Specimen infected with pseudotype (replication-defective) virus expressing Dl1 + GFP, and stained with antibody against GFP (green) plus HCA antibody (red). Hair cells are produced normally even within clusters of contiguous cells all expressing Dl1.
Patches of infection with RCAS-Dl1dn were seen inside sensory patches, or overlapping them, in 16 of 33 informative cases. Contrary to expectation, none of these patches showed an excessive density of hair cells (see Fig. 2B): 0.22 ± 0.07 hair cells were counted per μm length of sectioned epithelium in the infected regions, as compared with 0.23 ± 0.05 in the adjacent uninfected sensory tissue (mean ± SD, n = 11 in both cases; counts from representative sections showing patches ≥20 μm wide). Again, several interpretations are possible. Blocking Notch activity with RCAS-Dl1dn, although it affects Ser1 expression, may fail to affect cell differentiation (different Notch family members with different sensitivities to Dl1dn and different downstream actions could be involved, for example, or Ser1 expression and cell differentiation might have different thresholds of response to Notch activity). Alternatively, the cells may have been already irreversibly committed as hair cells or supporting cells before they became infected. Lastly, the cells may not have been irreversibly committed, but may only recently have become infected by the replication-competent virus as it spread through the tissue and may not have had time yet to change their phenotype. More experiments will be needed to decide between these interpretations.
A further observation is more straightforward in conformity with the expected effects of blocking Notch activity. Infected cells expressing Dl1dn often ended up as neurons in the cochleovestibular ganglion (Fig. 3 C and D). Indeed, in 6 of a set of 6 embryos fixed at E6, infected cells were found only in the ganglion. This finding suggests that the infected cells were biased toward a neuronal fate during the early phase of ear development when neuroblasts delaminate from the otic epithelium. Virus-infected nerve fibers innervating a sensory patch of epithelium are a possible source of late infection for the epithelial cells.
Ectopic Expression of Dl1 Does Not Inhibit Hair-Cell Production.
There are two ways in which a cell might escape lateral inhibition so as to become a hair cell: its neighbors might not deliver inhibitory signals, or it might be deaf to signals that it receives. In the sensory patches of the ear, all cells are in contact with neighbors expressing Ser1, suggesting that the nascent hair cells are deaf to at least this Notch ligand. They are not, however, normally exposed to Dl1, because Dl1 is not expressed by the supporting cells. To test whether forced expression of Dl1 throughout the cell population would block cell differentiation, as it does in the retina, we once again used viral vectors, this time containing the full-length Dl1 cDNA. We found 29 RCAS-Dl1-infected patches that lay within sensory patches or overlapping them (Fig. 4 A–D), and three that lay directly abutting sensory patches. We counted the numbers of hair cells per unit length of sectioned epithelium in infected as compared with adjacent uninfected sensory epithelium. No significant difference was seen. The values were respectively 0.23 ± 0.08 and 0.24 ± 0.08 hair cells per μm (mean ± SD, n = 18 in both cases; counts from representative sections showing patches ≥30 μm wide).
The observations using RCAS-Dl1 to misexpress Dl1 were confirmed by using a pseudotype replication-defective virus for the same purpose. This virus gives smaller patches of infected cells, but has the advantage that they can be assumed to have all become infected at the same early time. Again, hair cells could be seen to develop normally even where all of the cells in the neighborhood expressed Dl1 (Fig. 4E).
The cells that become hair cells therefore do so regardless of whether their neighbors express Dl1. The simple lateral-inhibition model based on regulation of Dl1 expression cannot be the correct explanation of why some cells escape inhibition to become hair cells and others do not. Some other factor must operate, either interacting with Notch ligands in the neighbors of the nascent hair cell and blocking their ability to deliver an inhibitory signal, or interacting with the Notch pathway in the nascent hair cell and blocking its ability to respond.
Numb Protein May Make Nascent Hair Cells Deaf to Notch Signaling.
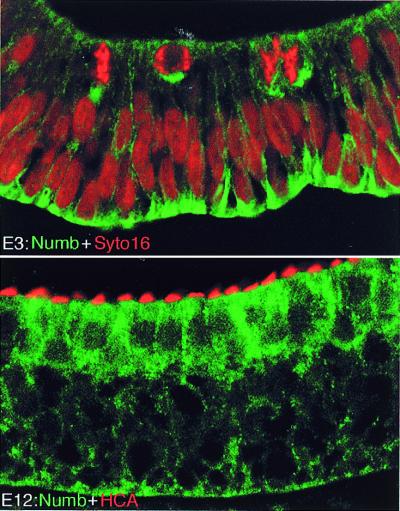
Almost nothing is known about molecules that might interact with Notch ligands in cis to prevent them delivering a signal to an adjacent cell (see Discussion). Analogies with Drosophila suggest, however, at least three factors that might act in nascent hair cells to make them insensitive to signals received. First, the hair cells might down-regulate their expression of Notch1 itself—indeed, they are known to do so, although this may occur too late to control the cell-fate decision (16). Second, Delta protein in large quantities can make cells that contain it unresponsive to signals from neighbors (51), and hair cells contain Dl1. Lastly, there are proteins such as Numb that interact with Notch to block its activity (51). We have used immunohistochemistry to look for expression of a chick Numb homolog in the inner ear. As shown in Fig. 5, the chick Numb protein is localized to the basolateral membranes of the epithelial cells at stages before hair-cell differentiation. Once hair cells have differentiated, it is seen at high concentration in hair cells. These observations are preliminary. They clearly suggest, however, that Numb protein acts in nascent hair cells to make them immune to Notch signaling.
Figure 5.
C-Numb expression during sensory patch development. (A) Section of otic epithelium at E3, stained with Numb antibody (green) and counterstained with the nuclear dye Syto16 (red); note basal localization of Numb, apparently in all cells, including those undergoing mitosis close to the lumen. (B) Section of basilar papilla at E12, stained with Numb antibody (green) and HCA (red); the hair cells preferentially contain Numb, and it is no longer basally localized.
Discussion
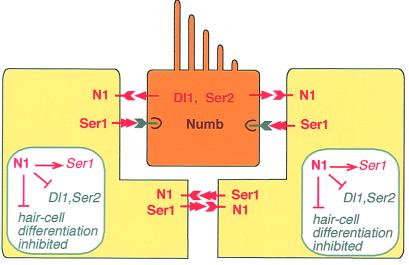
The implications of our experiments are summarized diagrammatically in Fig. 6, showing the patterns of expression of the various Notch ligands in a newly differentiated sensory patch and their regulatory interactions. Supporting cells express Ser1; hair cells express Dl1 and Ser2. Ser1 expression is regulated positively by Notch activity; Dl1 and Ser2 are regulated negatively (7). Supporting cells contact one another, so that mutual lateral inductive signals, as well as signals from hair cells, keep them all in a state of high Notch activation, which maintains high expression of Ser1 and low expression of Dl1 and Ser2. Hair cells contain Numb and down-regulate their Notch1 expression; thus despite their exposure to Ser1 from all sides, their level of Notch activation is low, their expression of Ser1 is low, and their expression of Dl1 and Ser2 is high. The cells that differentiate as hair cells do so because their level of Notch activation is low; those that differentiate as supporting cells do so because their level of Notch activation is high.
Figure 6.
Summary diagram of Notch-mediated interactions in a newly differentiated sensory patch in the chick ear. See Discussion for commentary.
This system of regulatory interactions is robust and consistent with all of the observations. Our account is, however, based on some tacit assumptions and raises several questions that need to be discussed. The first concerns Ser1.
Signaling by Ser1 May Serve to Prevent Premature Hair-Cell Production.
An attractive possibility is that Ser1, expressed throughout the prospective sensory patch from a very early stage, serves to prevent premature or ectopic hair-cell production by maintaining a high background level of Notch activation. This suggestion is supported by the recent finding that hair cells are overproduced in mouse cochlea explants when Ser1 (Jag1) production is inhibited with antisense oligonucleotides (52). Ser1 thus may be important in enforcing the long delay that occurs between neuroblast delamination and hair-cell differentiation in birds and mammals and in preventing the peripheral regions of growing sensory patches from differentiating prematurely in fish.
The supporting cells in sensory patches also express Lunatic fringe (Lfng) (4, 31), a homolog of the Fringe protein that has been shown in Drosophila to interact with Notch in cis and make Notch refractory by activation by Serrate (53, 54). This might be taken to suggest that Ser1 function normally is blocked by Lfng, in which case removal of Lfng should drastically alter the pattern of Notch activation and cell differentiation. However, ear development appears entirely normal in mice with a knockout mutation of the Lfng gene (55), and we see no abnormalities in ears overexpressing Lfng as a result of infection with RCAS-Lfng retrovirus (I.L.R., unpublished work). An effect of the Lfng knockout is seen only in mice that also have a mutation in Ser2: the Ser2 single mutants show mild overproduction of hair cells, and this effect is partially suppressed in the double Lfng; Ser2 mutants. Zhang et al. (54) interpret this in terms of an effect of Lfng as inhibitor of signaling by Dl1, although there is no precedent for such an effect. An alternative interpretation is that Lfng normally acts as a weak inhibitor of signaling by Ser1, so that loss of Lfng weakly potentiates the Notch-activating effect of Ser1; this might be expected to show up as a decrease of hair-cell production in the Ser2 mutant background, where absence of one Notch ligand makes the functions of the others more critical.
More Than One Notch Homolog May Be Involved in Ear Development.
A questionable assumption is that the various Notch ligands all act on the same receptor, Notch1. In fact, it has been reported that Notch2 and Notch3 are expressed along with Notch1 in the mouse ear (56, 57), and the different Notch family members might respond selectively to the different ligands and exert different downstream effects. Notch3, for example, may act as an antagonist of Notch1 (58). In the chick, however, only two Notch genes have been identified, and Notch1 is the only one that we have been able to detect in the ear (59).
Differing Levels of Exposure to Notch Ligands Cannot Account for the Choice Between Hair-Cell and Supporting-Cell Fate.
Regardless of these uncertainties, our data make one thing clear. The pattern of production of hair cells and supporting cells cannot be determined simply by the pattern of expression of Notch ligands, in the manner proposed by the simple model of lateral inhibition with feedback. The cells that become hair cells are not selected to do so by escape from exposure to Ser1 (they are constantly exposed), Dl1 (its ectopic expression does not change cell fate), or Ser2 (the knockout has only a mild effect). We have shown, however, that hair cells contain Numb, which can block Notch activation (47), supporting the idea that hair cells escape the inhibitory effect of Notch activation not because of lack of ligands from their neighbors, but because they are deaf to the signal delivered by the ligands (see ref. 60, however, for a contrary view of Numb function in the mouse).
We still have to answer our original question: why are hair cells and supporting cells produced in the observed ratio? This cannot be accounted for simply in terms of the rules of asymmetric inheritance of Numb. If each cell in the developing sensory patch went through a final asymmetric division, yielding one daughter that inherited Numb and one daughter that did not, the result would be a 1:1 ratio of hair cells to supporting cells, whereas the measured ratio (in chick basilar papilla) ranges from 1:1.7 to 1:3.9 (18). The level of Numb in the prospective hair cells as opposed to supporting cells may be controlled in some more complex way or through more complex sequences of cell divisions, or some molecule other than Numb and its asymmetrically located companion proteins (61, 63) may confer immunity to lateral inhibition and serve as the key determinant of cell fate.
It is also possible that some factor interacts with the Notch ligands in cis, making those produced by the prospective hair cells more powerful and those produced by the prospective sensory cells less so. If this modulatory factor were itself negatively regulated by Notch activity, we would recover the pattern-generating mechanism of the original simple model, but with the modulatory factor playing the key role instead of Delta. Other signaling pathways also may be involved. Epidermal growth factor signaling, for example, acts in conjunction with the Notch pathway to control cell fate in developing Drosophila sense organs (62).
New Insights Come from Ancient Homologies.
Our analysis has highlighted many unanswered questions about the role of Notch signaling in the development of the ear. The data leave no doubt, however, that the Notch pathway has a central role in controlling cell fate in this system. Our findings reveal a more complex picture than initially suspected, but reinforce the parallels between the sensory patches of the vertebrate ear and the sensilla of a fly, adding weight to the argument for homology. Thus in both systems, both the Delta and the Serrate subfamilies of Notch ligands are involved; cell fate choice is not simply dictated by the level of Notch ligand production in neighboring cells; and the cells that escape Notch-mediated inhibition contain high levels of Numb, a factor that can block Notch activity by binding to Notch.
Homologies between insects and vertebrates are commonplace at the molecular level. There are innumerable examples of homologous proteins serving the same molecular functions. It is also true that Notch signaling is important in many different tissues, both in vertebrates and invertebrates. But the homologies we have pointed out for the mechanosensory structures go deeper, including function, multicellular anatomy, development, and molecular controls. Indeed, there are few, if any, other multicellular structures where correspondences between insect and vertebrate seem so clear, detailed, and extensive. Of course, there are many differences: the Drosophila data can only provide us with hypotheses as to the workings of the vertebrate system, not dogmatic answers. By testing these hypotheses, as we have attempted to do in this paper, we gain a better appreciation of the evolutionary relationship, and a better understanding of the evolutionary relationship opens the way to new hypotheses and new insights into the development of the inner ear.
Acknowledgments
We are grateful to Yoshio Wakamatsu, Juan-Carlos Izpisua-Belmonte, David Shima, and Guy Richardson for reagents, Jenny Corrigan for cryosectioning, and Helen McNeill for comments on the manuscript. Above all, we are indebted to the colleagues who participated in the earlier work reviewed in this paper: Julie Adam, Catherine Haddon, Domingos Henrique, Christine Hodgetts, David Ish-Horowicz, Yun-Jin Jiang, Alastair Morrison, Anna Myat, and Tanya Whitfield. The Imperial Cancer Research Fund supported us.
Abbreviations
- Dl1
Delta1
- Ser1
Serrate1
- Ser2
Serrate2
- Su(H)
Suppressor of Hairless
- En
embryonic day n
- SMC
sensory mother cell
- GFP
green fluorescent protein
- HCA
hair cell antigen
- Lfng
Lunatic fringe.
Footnotes
This paper was presented at the National Academy of Sciences colloquium “Auditory Neuroscience: Development, Transduction, and Integration,” held May 19–21, 2000, at the Arnold and Mabel Beckman Center in Irvine, CA.
References
- 1.Watson G M, Mire P, Hudson R R. Hearing Res. 1997;107:53–66. doi: 10.1016/s0378-5955(97)00022-1. [DOI] [PubMed] [Google Scholar]
- 2.Gehring W J, Ikeo K. Trends Genet. 1999;15:371–377. doi: 10.1016/s0168-9525(99)01776-x. [DOI] [PubMed] [Google Scholar]
- 3.Lewis J. Regeneration of Vertebrate Sensory Cells/ , Ciba Symp. 1991;160:25–39. [Google Scholar]
- 4.Adam J, Myat A, Le Roux I, Eddison M, Henrique D, Ish-Horowicz D, Lewis J. Development (Cambridge, UK) 1998;125:4645–4654. doi: 10.1242/dev.125.23.4645. [DOI] [PubMed] [Google Scholar]
- 5.Artavanis-Tsakonas S, Rand M D, Lake R J. Science. 1999;284:770–776. doi: 10.1126/science.284.5415.770. [DOI] [PubMed] [Google Scholar]
- 6.Collier J R, Monk N A M, Maini P K, Lewis J H. J Theoret Biol. 1996;183:429–446. doi: 10.1006/jtbi.1996.0233. [DOI] [PubMed] [Google Scholar]
- 7.Haddon C, Jiang Y-J, Smithers L, Lewis J. Development (Cambridge, UK) 1998;125:4637–4644. doi: 10.1242/dev.125.23.4637. [DOI] [PubMed] [Google Scholar]
- 8.Parks A L, Huppert S S, Muskavitch M A. Mech Dev. 1997;63:61–74. doi: 10.1016/s0925-4773(97)00675-8. [DOI] [PubMed] [Google Scholar]
- 9.Baker N E. BioEssays. 2000;22:264–273. doi: 10.1002/(SICI)1521-1878(200003)22:3<264::AID-BIES8>3.0.CO;2-M. [DOI] [PubMed] [Google Scholar]
- 10.Guo M, Jan L Y, Jan Y N. Neuron. 1996;17:27–41. doi: 10.1016/s0896-6273(00)80278-0. [DOI] [PubMed] [Google Scholar]
- 11.Fekete D M. Curr Opin Neurobiol. 1996;6:533–541. doi: 10.1016/s0959-4388(96)80061-4. [DOI] [PubMed] [Google Scholar]
- 12.Haddon C, Lewis J. J Comp Neurol. 1996;365:113–128. doi: 10.1002/(SICI)1096-9861(19960129)365:1<113::AID-CNE9>3.0.CO;2-6. [DOI] [PubMed] [Google Scholar]
- 13.Whitfield T, Haddon C, Lewis J. Semin Cell Dev Biol. 1997;8:239–247. doi: 10.1006/scdb.1997.0143. [DOI] [PubMed] [Google Scholar]
- 14.Wu D K, Oh S H. J Neurosci. 1996;16:6454–6462. doi: 10.1523/JNEUROSCI.16-20-06454.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Fekete D M, Muthukumar S, Karagogeos D. J Neurosci. 1998;18:7811–7821. doi: 10.1523/JNEUROSCI.18-19-07811.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Stone J S, Rubel E W. Development (Cambridge, UK) 1999;126:961–973. doi: 10.1242/dev.126.5.961. [DOI] [PubMed] [Google Scholar]
- 17.Stone J S, Rubel E W. J Comp Neurol. 2000;417:1–16. doi: 10.1002/(sici)1096-9861(20000131)417:1<1::aid-cne1>3.0.co;2-e. [DOI] [PubMed] [Google Scholar]
- 18.Goodyear R, Richardson G. J Neurosci. 1997;17:6289–6301. doi: 10.1523/JNEUROSCI.17-16-06289.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Corwin J T, Jones J E, Katayama A, Kelley M W, Warchol M E. Ciba Found Symp. 1991;160:103–130. doi: 10.1002/9780470514122.ch6. [DOI] [PubMed] [Google Scholar]
- 20.Kimble J, Simpson P. Annu Rev Cell Dev Biol. 1997;13:333–361. doi: 10.1146/annurev.cellbio.13.1.333. [DOI] [PubMed] [Google Scholar]
- 21.Heitzler P, Simpson P. Cell. 1991;64:1083–1092. doi: 10.1016/0092-8674(91)90263-x. [DOI] [PubMed] [Google Scholar]
- 22.Ghysen A, Dambly-Chaudière C, Jan L Y, Jan Y-N. Genes Dev. 1993;7:723–733. doi: 10.1101/gad.7.5.723. [DOI] [PubMed] [Google Scholar]
- 23.Lewis J. Curr Opin Neurobiol. 1996;6:3–10. doi: 10.1016/s0959-4388(96)80002-x. [DOI] [PubMed] [Google Scholar]
- 24.Chitnis A, Henrique D, Lewis J, Ish-Horowicz D, Kintner C. Nature (London) 1995;375:761–766. doi: 10.1038/375761a0. [DOI] [PubMed] [Google Scholar]
- 25.Haddon C, Smithers L, Schneider-Maunoury S, Coche T, Henrique D, Lewis J. Development (Cambridge, UK) 1998;125:359–370. doi: 10.1242/dev.125.3.359. [DOI] [PubMed] [Google Scholar]
- 26.Henrique D, Hirsinger E, Adam J, Le Roux I, Pourquié O, Ish-Horowicz D, Lewis J. Curr Biol. 1997;7:661–670. doi: 10.1016/s0960-9822(06)00293-4. [DOI] [PubMed] [Google Scholar]
- 27.Morrison A, Hodgetts C, Gossler A, Hrabe de Angelis M, Lewis J. Mech Dev. 1999;84:169–172. doi: 10.1016/s0925-4773(99)00066-0. [DOI] [PubMed] [Google Scholar]
- 28.Lanford P J, Lan Y, Jiang R, Lindsell C, Weinmaster G, Gridley T, Kelley M W. Nat Genet. 1999;21:289–292. doi: 10.1038/6804. [DOI] [PubMed] [Google Scholar]
- 29.Riley B B, Chiang M, Farmer L, Heck R. Development (Cambridge, UK) 1999;126:5669–5678. doi: 10.1242/dev.126.24.5669. [DOI] [PubMed] [Google Scholar]
- 30.Haddon C, Mowbray C M, Whitfield T, Jones D, Gschmeissner S, Lewis J. J Neurocytol. 1999;28:837–850. doi: 10.1023/a:1007013904913. [DOI] [PubMed] [Google Scholar]
- 31.Cole L, Le Roux I, Nunes F, Laufer E, Lewis J, Wu D K. J Comp Neurol. 2000;424:509–520. doi: 10.1002/1096-9861(20000828)424:3<509::aid-cne8>3.0.co;2-q. [DOI] [PubMed] [Google Scholar]
- 32.Gho M, Bellaiche Y, Schweisguth F. Development (Cambridge, UK) 1999;126:3573–3584. doi: 10.1242/dev.126.16.3573. [DOI] [PubMed] [Google Scholar]
- 33.Hartenstein V, Posakony J W. Dev Biol. 1990;142:13–30. doi: 10.1016/0012-1606(90)90147-b. [DOI] [PubMed] [Google Scholar]
- 34.Parks A L, Muskavitch M A. Dev Biol. 1993;157:484–496. doi: 10.1006/dbio.1993.1151. [DOI] [PubMed] [Google Scholar]
- 35.Walker R G, Willingham A T, Zuker C S. Science. 2000;287:2229–2234. doi: 10.1126/science.287.5461.2229. [DOI] [PubMed] [Google Scholar]
- 36.Tilney L G, Connelly P, Smith S, Guild G M. J Cell Biol. 1996;135:1291–1308. doi: 10.1083/jcb.135.5.1291. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Goulding S E, White N M, Jarman A P. Dev Biol. 2000;221:120–131. doi: 10.1006/dbio.2000.9677. [DOI] [PubMed] [Google Scholar]
- 38.Bermingham N A, Hassan B A, Price S D, Vollrath M A, Ben-Arie N, Eatock R A, Bellen H J, Lysakowski A, Zoghbi H Y. Science. 1999;284:1837–1841. doi: 10.1126/science.284.5421.1837. [DOI] [PubMed] [Google Scholar]
- 39.Zheng J L, Gao W Q. Nat Neurosci. 2000;3:580–586. doi: 10.1038/75753. [DOI] [PubMed] [Google Scholar]
- 40.Bosse A, Zulch A, Becker M B, Torres M, Gomez-Skarmeta J L, Modolell J, Gruss P. Mech Dev. 1997;69:169–181. doi: 10.1016/s0925-4773(97)00165-2. [DOI] [PubMed] [Google Scholar]
- 41.Kavaler J, Fu W, Duan H, Noll M, Posakony J W. Development (Cambridge, UK) 1999;126:2261–2272. doi: 10.1242/dev.126.10.2261. [DOI] [PubMed] [Google Scholar]
- 42.Torres M, Gomez-Pardo E, Gruss P. Development (Cambridge, UK) 1996;122:3381–3391. doi: 10.1242/dev.122.11.3381. [DOI] [PubMed] [Google Scholar]
- 43.Zeng C, Younger-Shepherd S, Jan L Y, Jan Y N. Genes Dev. 1998;12:1086–1091. doi: 10.1101/gad.12.8.1086. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Rhyu M S, Jan L Y, Jan Y N. Cell. 1994;76:477–491. doi: 10.1016/0092-8674(94)90112-0. [DOI] [PubMed] [Google Scholar]
- 45.Wettstein D A, Turner D L, Kintner C. Development (Cambridge, UK) 1997;124:693–702. doi: 10.1242/dev.124.3.693. [DOI] [PubMed] [Google Scholar]
- 46.Kinsella T M, Nolan G P. Hum Gene Ther. 1996;7:1405–1413. doi: 10.1089/hum.1996.7.12-1405. [DOI] [PubMed] [Google Scholar]
- 47.Wakamatsu Y, Maynard T M, Jones S U, Weston J A. Neuron. 1999;23:71–81. doi: 10.1016/s0896-6273(00)80754-0. [DOI] [PubMed] [Google Scholar]
- 48.Bartolami S, Goodyear R, Richardson G. J Comp Neurol. 1991;314:777–788. doi: 10.1002/cne.903140410. [DOI] [PubMed] [Google Scholar]
- 49.de Celis J F, Bray S. Development (Cambridge, UK) 1997;124:3241–3251. doi: 10.1242/dev.124.17.3241. [DOI] [PubMed] [Google Scholar]
- 50.Lewis J. Semin Cell Dev Biol. 1998;9:583–589. doi: 10.1006/scdb.1998.0266. [DOI] [PubMed] [Google Scholar]
- 51.Jacobsen T L, Brennan K, Arias A M, Muskavitch M A. Development (Cambridge, UK) 1998;125:4531–4540. doi: 10.1242/dev.125.22.4531. [DOI] [PubMed] [Google Scholar]
- 52.Zine A, Van De Water T R, de Ribaupierre F. Development (Cambridge, UK) 2000;127:3373–3383. doi: 10.1242/dev.127.15.3373. [DOI] [PubMed] [Google Scholar]
- 53.Irvine K D. Curr Opin Genet Dev. 1999;9:434–441. doi: 10.1016/S0959-437X(99)80066-5. [DOI] [PubMed] [Google Scholar]
- 54.Ju B G, Jeong S, Bae E, Hyun S, Carroll S B, Yim J, Kim J. Nature (London) 2000;405:191–195. doi: 10.1038/35012090. [DOI] [PubMed] [Google Scholar]
- 55.Zhang N, Martin G V, Kelley M W, Gridley T. Curr Biol. 2000;10:659–662. doi: 10.1016/s0960-9822(00)00522-4. [DOI] [PubMed] [Google Scholar]
- 56.Lardelli M, Dahlstrand J, Lendahl U. Mech Dev. 1994;46:123–136. doi: 10.1016/0925-4773(94)90081-7. [DOI] [PubMed] [Google Scholar]
- 57.Hamada Y, Kadokawa Y, Okabe M, Ikawa M, Coleman J R, Tsujimoto Y. Development (Cambridge, UK) 1999;126:3415–3424. doi: 10.1242/dev.126.15.3415. [DOI] [PubMed] [Google Scholar]
- 58.Beatus P, Lundkvist J, Berg C, Lendahl U. Development (Cambridge, UK) 1999;126:3925–3935. doi: 10.1242/dev.126.17.3925. [DOI] [PubMed] [Google Scholar]
- 59.Myat A, Henrique D, Ish-Horowicz D, Lewis J. Dev Biol. 1996;174:233–247. doi: 10.1006/dbio.1996.0069. [DOI] [PubMed] [Google Scholar]
- 60.Zhong W, Jiang M M, Schonemann M D, Meneses J J, Pedersen R A, Jan L Y, Jan Y N. Proc Natl Acad Sci USA. 2000;97:6844–6849. doi: 10.1073/pnas.97.12.6844. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Jan Y N, Jan L Y. Cell. 2000;100:599–602. doi: 10.1016/s0092-8674(00)80695-9. [DOI] [PubMed] [Google Scholar]
- 62.zur Lage P, Jarman A P. Development (Cambridge, UK) 1999;126:3149–3157. doi: 10.1242/dev.126.14.3149. [DOI] [PubMed] [Google Scholar]
- 63.Stone J S, Rubel E W. Proc Natl Acad Sci USA. 2000;97:11714–11721. doi: 10.1073/pnas.97.22.11714. [DOI] [PMC free article] [PubMed] [Google Scholar]