Abstract
The nonobese diabetic (NOD) mouse is a classical animal model for autoimmune type 1 diabetes (T1D), closely mimicking features of human T1D. Thus, the NOD mouse presents an opportunity to test the effectiveness of induced pluripotent stem cells (iPSCs) as a therapeutic modality for T1D. Here, we demonstrate a proof of concept for cellular therapy using NOD mouse-derived iPSCs (NOD-iPSCs). We generated iPSCs from NOD mouse embryonic fibroblasts or NOD mouse pancreas-derived epithelial cells (NPEs), and applied directed differentiation protocols to differentiate the NOD-iPSCs toward functional pancreatic beta cells. Finally, we investigated whether the NPE-iPSC-derived insulin-producing cells could normalize hyperglycemia in transplanted diabetic mice. The NOD-iPSCs showed typical embryonic stem cell-like characteristics such as expression of markers for pluripotency, in vitro differentiation, teratoma formation, and generation of chimeric mice. We developed a method for stepwise differentiation of NOD-iPSCs into insulin-producing cells, and found that NPE-iPSCs differentiate more readily into insulin-producing cells. The differentiated NPE-iPSCs expressed diverse pancreatic beta cell markers and released insulin in response to glucose and KCl stimulation. Transplantation of the differentiated NPE-iPSCs into diabetic mice resulted in kidney engraftment. The engrafted cells responded to glucose by secreting insulin, thereby normalizing blood glucose levels. We propose that NOD-iPSCs will provide a useful tool for investigating genetic susceptibility to autoimmune diseases and generating a cellular interaction model of T1D, paving the way for the potential application of patient-derived iPSCs in autologous beta cell transplantation for treating diabetes.
Introduction
Type 1 diabetes (T1D), a polygenic autoimmune disease, has been linked to more than 25 genetic loci [1] and is caused by the destruction of pancreatic beta cells through insulitis, which involves the infiltration of leukocytes into pancreatic islets [2]. Currently, pancreatic islet transplantation appears a promising treatment option for T1D patients [3,4]. However, cell replacement treatment for T1D needs a source of glucose-responsive insulin-secreting cells [5]. Results achieved following transplantation of pancreatic islets of Langerhans or cadaver-derived pancreatic tissue are encouraging, but this therapy is not widely used due to shortage of donor islets as well as severe immune rejection [6–8]. Although embryonic stem cells (ESCs) have been reported to differentiate into pancreatic beta-like insulin-producing cells [9,10], cells derived from immunologically unmatched ESCs could be the targets of both allograft reactions and the autoimmune response, resulting in cell destruction. In the case of mesenchymal stem cells (MSCs), which are self-renewable multipotent progenitor cells [11], profound immunomodulatory effect was reported both in vitro and in vivo and a variety of clinical trials were conducted in aiming at reducing the burden of immune-mediated disease [12,13]. However, the precise mechanisms underlying the immunomodulatory effects of MSC remain largely unknown and their immunogenicity following transplantation into allogeneic recipients is unclear [14,15]. An alternative source of insulin-producing cells may be patient-derived induced pluripotent stem cells (iPSCs). Such iPSCs can be generated from cells taken from humans of all ages with any genetic disease, and used for future cell therapy [16]. Such an autologous approach would eliminate the possibility of alloimmune rejection of transplanted cells. We propose that a combination of cellular reprogramming and differentiation techniques might be used for the generation of patient-specific iPSCs and their differentiation into pancreatic beta-like cells, and that such cells might provide a promising resource for cell therapy to treat diabetes without requiring high doses of immune-suppressive drugs [17–19].
Nonobese diabetic (NOD) mice spontaneously develop autoimmune T1D, which has many similarities to human autoimmune diabetes, and is a valuable model for investigating the pathogenesis of and genetic susceptibility to T1D [20]. A number of researchers have identified strategies for the treatment and prevention of diabetes in NOD mice [21–33]. For example, anti-CD3 treatment in NOD mice formed the basis for a clinical trial of anti-CD3 monoclonal antibody therapy in human T1D models. Other promising avenues of investigation include finding ways to reverse established autoimmunity and identifying new potential therapeutic targets. Given the similarity to human TID, we chose this model to test the effectiveness of a cell therapy approach.
We established NOD-iPSC lines and applied a chemically defined stepwise method to differentiate the NOD mouse-derived iPSCs (NOD-iPSCs) into functional pancreatic beta-like insulin-producing cells. The differentiated NOD-iPSCs expressed diverse markers of pancreatic beta cells and released insulin in response to glucose and KCl stimulation. Moreover, when transplanted into diabetic mice, the NOD-iPSC-derived insulin-producing cells ameliorated hyperglycemia.
Materials and Methods
Mice
NOD/ShiJcl and NOD/SCID mice were purchased from the Korean Research Institute of Bioscience and Biotechnology. All animal experiments conformed to the Guide for the Care and Use of Laboratory Animals and were approved by the Institutional Animal Care and Use Committee (IACUC) of Konkuk University (KU10069 and KU10070).
Preparation of NOD mouse embryonic fibroblasts and NOD pancreas-derived epithelial cells
NOD mouse embryonic fibroblasts (NMs) were isolated from 14-day-old NOD mouse embryos (n=7) [34]. NOD mouse pancreas-derived epithelial cells (NPEs) were isolated from the pancreatic tissue from an 8-week-old NOD mouse, as previously described [35]. Isolated NMs and NPEs were cultured in 60-mm dishes containing the Dulbecco's modified Eagle's medium (DMEM; Sigma, St. Louis, MO), 10% fetal bovine serum (FBS; HyClone, Logan, UT), and 100 U/mL of penicillin/streptomycin (Invitrogen, Carlsbad, CA).
Plasmid preparation and magnet-based nanofection
For generation of exogenous DNA-free iPS cells, magnet-based nanofection was conducted according to the previous reports [17,36]. Briefly, the plasmid DNAs containing Octamer-binding transcription factor 4 (OCT4), SRY-box containing gene 2 (SOX2), Krüppel-like factor 4 (KLF4), and c-MYC were mixed with PolyMAG (chemicell, GmbH, Berlin, Germany). For transfection, cells were seeded into each well of 24-well plates (1×105 cells/well) 1 day before transfection and the cells were transfected by the Magnetofection system.
Lentiviral infection and iPS cell generation
NMs and NPEs were infected with concentrated self-inactivating human immunodeficiency virus type-1-based lentiviral vectors encoding OCT4, SOX2, KLF4, and c-MYC under the control of the elongation factor-1 alpha (EF-1 alpha) promoter. G4-2 ESCs [37,38] and NOD-iPSCs were maintained on feeder layers of mitomycin C-treated mouse embryonic fibroblast (MEF) cells in DMEM supplemented with 20% FBS, 2 mM l-glutamine (Sigma), 0.1 mM β-mercaptoethanol (Invitrogen), and leukemia inhibitory factor (1,000 U/mL of ESGRO; Chemicon, Temecula, CA) at 37°C in 5% CO2.
Bisulfite DNA-sequencing analysis
Genomic DNA was purified from ESCs, NOD-iPSCs, NPEs, and NMs using the Genomic DNA Prep kit (Qiagen, Hilden, Germany) and 2 μg of DNA was utilized for bisulfite conversion using the EpiTect Bisulfite kit (Qiagen), according to the manufacturer's protocol. Bisulfite-treated DNA was amplified by polymerase chain reaction (PCR), cloned into the pGEMT-easy vectors (Promega, Fitchburg, WI), and then sequenced using T7 forward and SP6 reverse primers. Primer sequences are listed in Supplementary Table S1 (Supplementary Data are available online at www.liebertpub.com/scd).
Genotyping analysis
The genotypes of established NOD-iPSCs were determined using microsatellite markers (Mouse Genome Informatics). Of these markers, we selected D11Mit320 and D3Mit21, which distinguish the NOD genotype from the C57BL/6 genotype. Genomic DNA was isolated using the Genomic DNA Prep kit (Qiagen) and specific DNA regions were amplified from the purified DNA using specific primers (Supplementary Table S1). Amplified products were analyzed on a 4% agarose gel.
Flow cytometry analysis
Single-cell suspensions of differentiating NOD-iPSCs and G4-2 ESCs were obtained by dissociating cells using TrypLE (Invitrogen). Differentiated cells were fixed with 4% paraformaldehyde (PFA; Sigma), permeabilized with 0.5% Triton X-100 (Sigma), and immunostained with specific antibodies. The primary and secondary antibodies used were anti-SOX17 (Santa Cruz Biotechnology, Santa Cruz, CA) and Cy 3-conjugated goat anti-rabbit (Jackson ImmunoResearch labs, Inc, PA), respectively. Flow cytometry data were analyzed on 10,000 cells by using a FACScan flow cytometer and Cell Quest software (Becton, Dickinson and Company, Franklin Lakes, NJ). For multicolor flow cytometry analysis, cells were obtained by dissociating cells using TrypLE (Invitrogen), washed twice in PBS, and then stained using the Multicolor Flow Cytometry kit (R&D Systems, Minneapolis, MN), according to the manufacturer's protocol.
Quantitative real-time reverse transcription–polymerase chain reaction
Total RNA was isolated using the TRI reagent (Sigma), according to the manufacturer's protocol. One microgram of total RNA was used for cDNA synthesis (ImProm-II reverse transcriptase; Promega), and cDNA was subjected to PCR amplification with specific primers. Primer sequences are listed in Supplementary Table S1. Quantitative real-time PCR was carried out using the SsoFast EvaGreen Supermix (Bio-Rad, Hercules, CA). Gene expression was quantified only in the linear range of each primer pair. The ΔΔCT method [39] was used to quantify changes in the expression of each specific gene, using the housekeeping gene GAPDH as the normalizer.
Spontaneous differentiation
To induce differentiation into the 3 germ layers, cells were cultured in suspension to form embryoid bodies (EBs) in low-attachment dishes. The cells were subjected to an 8-day induction procedure (4−/4+) that consisted of 4 days of culture as aggregates without 0.1 μM retinoic acid (RA; Sigma), followed by 4 days of culture with RA. EBs were cultured in the differentiation medium (DMEM-GlutaMax-1; Invitrogen) containing 1% nonessential amino acids (NEAA; Sigma), 0.1 mM β-mercaptoethanol, and 10% FBS. EBs were cultured for 2 weeks on plates coated with 0.1% gelatin (Sigma).
Teratoma formation assay
One million NOD-iPSCs were injected intramuscularly into the hind leg of NOD/SCID mice. Teratomas were excised 5 weeks after injection and processed according to standard procedures for paraffin embedding and hematoxylin/eosin staining.
Eight-cell stage injection and embryo transfer
B6D2F1 (C57BL/6×DBA/2) and imprinting control region (ICR) male and female mice were used to produce in vitro fertilized 8-cell embryos. Embryo injections were performed on 8-cell embryos before obvious compaction using an Olympus micromanipulator (Olympus, Center Valley, PA). Eight-cell embryos were held using a holding pipette, and a perforation in the zona pellucida was created by the XYClone laser system (Hamilton Thorne Biosciences, Beverly, MA). Twenty NOD-iPSCs or ESCs were injected into each 8-cell embryo using 20 μM injection needles. Injected embryos were cultured for 24 h and transferred into the uteri of recipient ICR female mice at 2.5 dpc.
Differentiation of NOD-iPSCs into pancreatic beta-like cells
Directed differentiation was conducted with slight modifications of previously described protocols [40,41]. ESCs and NOD-iPSCs were plated onto dishes coated with Matrigel (1:50; BD Biosciences, Franklin Lakes, NJ) and incubated for 3 days with X-Vivo 10 (Cambrex, East Rutherford, NJ) supplemented with 0.1% bovine serum albumin (BSA) (Sigma) containing 50 ng/mL activin A (R&D Systems). Then, the differentiated cells were cultured for 2 days in the F12/Iscove's modified Dulbecco's medium (1:1; Invitrogen) supplemented with 0.5% BSA and 1% insulin-transferrin-selenium (ITS; Stemcell Technologies, Vancouver, Canada) along with 2 μM RA. After RA induction, the cells were cultured in the DMEM (low glucose; Invitrogen) supplemented with 0.5% BSA and 1% ITS, together with 10 ng/mL basic fibroblast growth factor (bFGF; R&D Systems) and 20 ng/mL epidermal growth factor (EGF; Sigma) for 3 days, and for another 3–5 days in the DMEM/F12 medium (Invitrogen) with ITS, 10 ng/mL bFGF and 10 mM nicotinamide (Sigma) for maturation.
Immunofluorescence
For immunofluorescence staining, ESCs, NOD-iPSCs, and differentiated cells were fixed with 4% PFA (Sigma), permeabilized with 0.5% Triton X-100 (Sigma), and immunostained with specific antibodies. To block any nonspecific binding, cells were incubated first in 10% normal goat serum (Vector Laboratories, Burlingame, CA) for 1 h and then incubated with the primary antibody at 4°C overnight. After washing with PBS, cells were incubated with the secondary antibody for 1 h and nuclei were stained with 5 μg/mL DAPI. The primary antibodies used included those against OCT4, stage-specific embryonic antigen-1 (SSEA-1), and paired box gene 6 (PAX6), pancreatic and duodenal homeobox 1 (PDX1), SOX 17, Islet-1 (ISL-1), Somatostatin, beta-tubulin III, and Brachyury (all from Santa Cruz Biotechnology), Nestin (Abcam, Cambridge, United Kingdom), alpha-fetoprotein (AFP; R&D Systems), forkhead box A2 (FOXA2), C-peptide, and Glucagon (Cell Signaling, Danvers, MA), and Insulin (Cell Signaling and Sigma). The secondary antibodies used were Cy 3-conjugated goat anti-mouse or rabbit (Jackson ImmunoResearch labs) and DyLight 649-conjugated goat anti-mouse or rabbit (Jackson ImmunoResearch labs) antibodies. All images were captured on a Carl Zeiss laser scanning (LSM-710) confocal microscope (Carl Zeiss, Oberkochen, Germany).
Insulin release ELISA assay
Primary adult mouse islets were isolated according to a standard procedure as described [42]. Insulin release was measured by incubating the cells in the Krebs-Ringer buffer. After preincubating with the Krebs-Ringer buffer at 37°C for 90 min, the differentiated cells were further incubated at 37°C for 60 min with the Krebs-Ringer buffer containing 2.5 mM d-glucose or 27.5 mM d-glucose or 30 mM KCl (Sigma). The insulin levels in culture supernatants were measured using the rat/mouse ELISA kit (EZRMI-13K; Linco, Weldon Spring, MO). The total protein content was detected using a Bradford protein assay kit (Bio-Rad).
Transplantation of NPE-iPSC-derived insulin-producing cells
Streptozotocin (STZ) was injected intraperitoneally for 3 days at 50 mg/kg into 6- to 8-week-old NOD/SCID mice. When nonfasting blood glucose levels were above 13.9 mM on 2 consecutive days, 5×106 differentiated cells were transplanted into the left subcapsular renal space. Blood glucose levels were measured every 2 days between 9:00 and 10:00 am, under nonfasting conditions, using a portable glucometer (ACCU-CHEK Active; Roche, Basel, Switzerland). To quantify beta cell proliferation, bromodeoxyuridine (BrdU, Sigma) incorporation into beta cells was evaluated 12 h after an intraperitoneal injection of 100 mg/kg BrdU by double immunofluorescence staining for BrdU, insulin, and nestin. The primary antibodies used included those against BrdU (Santa Cruz Biotechnology), insulin (Cell Signaling), and nestin (Abcam, Cambridge, United Kingdom). The secondary antibodies used were Alexa Fluor 488 goat anti-rabbit (Invitrogen) and DyLight 649-conjugated goat anti-mouse (Jackson ImmunoResearch labs) antibodies as described [43]. We conducted left nephrectomy (removal of both left kidney and the graft) at 5 weeks post-transplantation and analyzed by hematoxylin/eosin staining or immunohistochemistry [immunofluorescence or diaminobenzidine (DAB)-nickel reactions].
Statistical analysis
Data were expressed as in terms of mean and standard error. Statistical significance was determined by the unpaired Student's t-test. Statistical analysis was performed on data using Origin software (OriginLab Corporation, Northampton, MA). All results were derived from at least 3 independent experiments.
Results
Establishment and characterization of NOD-iPSC lines
We prepared MEFs or pancreas-derived epithelial cells (PEs) from NOD mice and used quantitative reverse transcription (RT)-PCR to analyze the level of pancreas-specific gene expression, including the expression of hepatocyte nuclear factor 3 beta (HNF 3 beta), PDX1, and PAX6. Pancreas-specific genes were expressed to a greater extent in NPEs than in NMs and control ESCs (G4-2) (Fig. 1A). At first, as we previously established a magnet-based nanofection technique for generation of exogenous DNA-free iPS cells, this transient expression technique was applied for generation of NOD-iPSCs. However, we could not stably establish the NOD-iPSCs without constitutive expression of exogenous factors in the NOD background [44]. Next, we infected the cultured NMs and NPEs with a combination of lentiviruses that encoded OCT4, SOX2, KLF4, and c-MYC under the control of the EF-1 alpha promoter, and morphological transformations of the infected cells were examined. Colonies were selected 14 days after infection, based on their morphological resemblance to mouse ESC colonies (Fig. 1B). NOD-iPSCs formed compact colonies, as observed by phase-contrast microscopy, and were positive for alkaline phosphatase activity (Fig. 1C). Reprogramming efficiency was defined as the number of GFP positive colonies per 10,000 NMs or NPEs seeded. We observed a higher reprogramming efficiency for NPEs (0.08%) than for NMs (0.04%). Genotype analysis of the established NOD-iPSC lines confirmed a pure NOD genetic background (Fig. 1D). Additionally, all the NOD-iPSC lines acquired the characteristic stem cell-cycle signature of ESCs (Supplementary Fig. S1). The expression levels of the endogenous pluripotency markers OCT4, SOX2, KLF4, and c-MYC were similar in NOD-iPSCs and G4-2 ESCs. Moreover, additional ESC markers (Nanog, FGF4, and Cripto) were also detected in NOD-iPSCs, and their expression levels were similar to those seen in ESCs (Fig. 1E and Supplementary Fig. S2). Both NM-iPSCs and NPE-iPSCs were stained with antibodies against OCT4 and SSEA-1, similar to G4-2 ESCs (Fig. 1F and Supplementary Fig. S3). Bisulfite genomic-sequencing analyses revealed similarities between NOD-iPSCs and G4-2 ESCs. The promoter region of the OCT4 gene was highly unmethylated in NOD-iPSCs, whereas CpG dinucleotides in this region were highly methylated in NMs and NPEs (Fig. 1G). Moreover, using fluorescence-activated cell-sorting analysis, we confirmed high-level expression of OCT4, SSEA-1, and SOX2 markers in both NM-iPSCs and NPE-iPSCs (Fig. 1H and Supplementary Fig. S4).
FIG. 1.
Generation of NOD-iPSCs from a NOD mouse. (A) Quantitative real-time PCR analysis of expression of pancreatic-related genes (HNF-3 beta, PDX1, and PAX6). Relative expression levels were normalized to the average expression of the control ESC line G4-2. (B) Time schedule of NOD-iPSC generation by infection with lentiviruses encoding OCT4, SOX2, KLF4, and c-MYC. (C) Typical images of ESC-like colonies. NOD-iPSC lines were established from NMs and NPEs. Representative images of colonies after 20 passages, and detection of alkaline phosphatase activity. (D) Genotyping of NOD-iPSCs generated from NMs and NPEs. Two microsatellite markers corresponding to the Idd locus were examined in NOD-iPSCs. Note that all of the NOD-iPSCs examined exhibited the NOD genetic background and differed from that of the C57/BL6 mice. (E) RT-PCR analysis of expression of ESC marker genes in NOD-iPSC lines. Primers for OCT4, SOX2, KLF4, and c-MYC specifically detected transcripts from endogenous genes. (F) Immunostaining for the expression of pluripotency markers OCT4, and SSEA-1. Nuclei were stained with DAPI; G4-2 ESCs were used as positive controls. (G) Bisulfite genomic sequencing of the OCT4 promoter region. Open and closed circles indicate unmethylated and methylated CpGs dinucleotides, respectively. Six representative sequenced clones from NMEF, G4-2 ESC, NMEF-iPS1, and NPE-iPS1 are shown. (H) Representative fluorescence-activated cell-sorting analysis of the expression profiles of OCT4, SOX2, and SSEA-1 in NOD-iPSCs and G4-2 ESCs. Scale bars, 200 μm (Day 14 colony; B), 500 μm (Phase and AP; B), and 50 μm (OCT4 and SSEA-1; E). AP, alkaline phosphatase; ESC, embryonic stem cell; HNF 3 beta, hepatocyte nuclear factor 3 beta; KLF4, Krüppel-like factor 4; NOD, nonobese diabetic; iPSCs, induced pluripotent stem cells; NOD-iPSCs, NOD mouse-derived iPSCs; NMs, NOD mouse embryonic fibroblasts; NPEs, NOD mouse pancreas-derived epithelial cells; OCT4, octamer-binding transcription factor 4; PAX6, paired box gene 6; PDX1, pancreatic and duodenal homeobox 1; SOX2, SRY-box containing the gene 2; SSEA-1, stage-specific embryonic antigen-1; RT-PCR, reverse transcription-polymerase chain reaction. *p<0.05. Color images available online at www.liebertonline.com/scd
NOD-iPSCs spontaneously differentiate into 3-germ-layer cell types
To determine their pluripotency, we studied whether NOD-iPSCs could be spontaneously differentiated into 3-germ-layer cells. EB formation (Fig. 2A) was induced by culturing G4-2 ESCs and NOD-iPSCs in a differentiation medium (20% DMEM without LIF) on low-attachment plates, followed by plating onto gelatin-coated dishes for additional culture. We confirmed the differentiation of NOD-iPSCs into diverse cell types from the 3 germ layers by checking the expression of specific markers for ectodermal (nestin and beta tubulin III), endodermal (AFP and Gata4), and mesodermal (brachyury and BMP4) lineages. RT-PCR (Fig. 2B) and immunocytochemical analysis (Fig. 2C and Supplementary Fig. S5) confirmed that differentiated NOD-iPSCs (both NM-iPSCs and NPE-iPSCs) expressed markers for all 3 germ layers. To examine the in vivo developmental pluripotency of the NOD-iPSCs, we performed a teratoma formation assay. When injected into NOD/SCID mice, both NM-iPSCs and NPE-iPSCs formed teratomas, including derivatives of the endoderm (gut-like epithelium and respiratory epithelium), mesoderm (smooth muscle and cartilage), and ectoderm (nerve fibers and skin) (Fig. 2D). We successfully produced chimeric mice by injecting NM-iPSCs or NPE-iPSCs into B6D2F2 8-cell embryos, and compared them with ICR 8-cell embryos that were injected with 129 ESCs (G4-2; Fig. 2E). Thus, NOD-iPSCs were successfully generated from NOD mice.
FIG. 2.
In vitro and in vivo differentiation of NOD-iPSCs into the 3 germ layers. (A) Phase-contrast images of embryoid bodies derived from NOD-iPSCs. (B) RT-PCR analyses of differentiation markers for the 3 germ layers (endoderm: AFP and GATA4; mesoderm: brachyury and BMP4; ectoderm: nestin and beta tubulin III). (C) Immunocytochemistry of NOD-iPSC lines (NM-iPS1 and NPE-iPS1) using markers for the 3 germ layers (endoderm: AFP; mesoderm: brachyury; ectoderm: nestin and beta tubulin III). (D) Hematoxylin/eosin staining of teratoma sections. Cells were transplanted intramuscularly into the hind leg of individual NOD/SCID mice. Hematoxylin/eosin staining of teratoma sections revealed smooth muscle (M), cartilage (C), nerve fibers (N), skin (S), gut-like epithelium (G), and respiratory epithelium (R). (E) Production of a chimeric mouse derived from NOD-iPSCs. NOD-iPSCs and control ESCs (G4-2) were injected into the space between the zona pellucida and blastomeres of host 8-cell embryos through a perforation created by a XYClone laser. Chimeric mice generated from NOD-iPSCs. Red arrows indicate that the chimerism originated from the NPE-iPSCs. Scale bars, 500 μm (A), and 100 μm (C, D). AFP, alpha-fetoprotein. Color images available online at www.liebertonline.com/scd
Highly efficient differentiation of NPE-iPSCs into insulin-producing cells
To confirm the transitions in the specific marker gene expression patterns of each cell (NM-iPSCs, NPE-iPSCs, and G4-2 ESCs) during direct differentiation, quantitative RT-PCR was performed at each step throughout differentiation for genes involved in pancreas development (Fig. 3A, C). After formation of the induced definitive endoderm, the specific marker gene expression patterns in the endoderm were monitored and SOX17 expression peaks were observed at step 1 (Fig. 3C), at a time when the OCT4 expression level rapidly decreased (Fig. 3B). The NPE-iPSC-derived definitive endoderm exhibited higher SOX17 peak expression than did the NM-iPSCs and G4-2 ESCs. Furthermore, the increased SOX17 expression level in the NPE-derived definitive endoderm was confirmed by flow cytometry. NPE-derived definitive endoderm differentiated cells were found to have a higher percentage of SOX17-positive cells, that is, >87.23%, than did NM-iPSCs (72.55%) and G4-2 ESCs (74.75%) in a 3-day culture (Fig. 3D). In step 2, the cells differentiated into PDX1-positive pancreatic progenitors and we were able to detect upregulated PDX1 gene expression on day 6 (Fig. 3C). After the induction process (steps 3 and 4), the expression of pancreatic beta cell-specific genes, ISL-1 and insulin, was detected in the differentiated cells. In particular, our data showed that the expression of pancreatic beta cell-specific genes was 2-fold higher in NPE-iPSC-derived, than in NM-iPSC- and G4-2 ESC-derived differentiated cells. These results suggest that NPE-iPSCs can be more efficiently differentiated into pancreatic lineage cells than NM-iPSCs or G4-2 ESC-derived cells can.
FIG. 3.
Dynamic expression patterns of pancreatic lineage genes during direct pancreatic differentiation from NOD-iPSCs. (A) The flow chart of the differentiation protocol. NOD-iPSCs and G4-2 ESCs were differentiated into the definitive endoderm (step 1; 3 days), pancreatic progenitor cells (step 2; 4 days), and expanded progenitor cells (step 3; 4 days). (B) and (C) Dynamic gene expression patterns of the differentiated NOD-iPSCs were analyzed at each step of induction in 3 separate experiments by quantitative RT-PCR (qRT-PCR) analysis. (B) The expression level of OCT4, an ES cell pluripotency marker, rapidly decreases around step 1. (C) The differentiated cell samples were collected at steps 1, 2, 3, and 4 and analyzed by qRT-PCR for SOX17, PDX1, ISL-1, and Insulin. (D) Flow cytometry analysis of the differentiated NOD-iPSCs showed the presence of SOX17-positive cells after 3-day induction with activin A and wort. For each sample, gene expression was normalized to that of GAPDH. *P< 0.05. ISL-1, Islet-1.
NOD-iPSCs can be induced to differentiate into insulin-producing cells
We used immunocytochemistry and quantitative RT-PCR to characterize the expression of differentiation state-specific genes after each of the 4 steps in our differentiation process. In step 1, consisting of 3 days of treatment with activin A and wortmannin, the endoderm-specific expression markers SOX17, HNF-3 beta, CXCR4, GATA4, and FOXA2 were expressed in the differentiating cells. In step 2, further differentiation toward PDX1-positive pancreatic progenitors was achieved by treating the cells with RA, Noggin, and FGF7 for 3 days. In step 3, to facilitate the proliferation of pancreatic progenitors, differentiated cells in the pancreatic progenitor state were treated with EGF for 4 days. After EGF treatment, the expression of PAX6, NeuroD1, Neurogenin3, and ISL-1 was detected in the differentiated cells (Fig. 4A, B and Supplementary Fig. S6). Insulin-producing cells differentiated from NPE-iPSCs expressed pancreatic lineage-related genes to a greater degree than those differentiated from either NM-iPSCs or G4-2 ESCs (Fig. 4B). In the final stage (step 4), we induced the differentiation of NOD-iPSC-derived pancreatic progenitors into cells of endocrine lineage, which were positive for insulin, C-peptide, and PDX1 (Fig. 5A and Supplementary Fig. S7). We observed the apparent co-expression of insulin/PDX1 or C-peptide/PDX1, and some of the insulin-positive cells were costained for Glucagon or Somatostatin, suggesting that while these cells synthesized insulin, they were still immature [45]. We also confirmed the expression of pancreatic beta cell-specific genes, including IAPP, insulin, and GLUT2, in the differentiated cells. Additionally, we confirmed that the differentiated cells expressed both insulin 1 and insulin 2 in the final stage (Supplementary Fig. S8). We observed higher expression of these genes in cells differentiated from NPE-iPSCs than in those from NM-iPSCs and G4-2 ESCs (Fig. 5B). To determine whether insulin release by NOD-iPSC-derived insulin-producing cells was responsive to glucose or other physiological stimulation, we exposed these cells to low concentrations of glucose, high concentrations of glucose, KCl, or 100 μM carbachol and then used ELISA to analyze the quantity of insulin released into the culture medium. Upon stimulation with high levels of glucose (25 mM), KCl (30 mM), or 100 μM carbachol the amount of insulin released was at least 2-fold the amount released when cells were stimulated with low levels of glucose (2.5 mM; Fig. 5C). We conducted more experiments to compare insulin release from iPSC-derived beta cells and that from natural beta cells using primary islets from NOD/scid mice. Although the amount of released insulin was lower compared to primary islets, this finding indicates that these cells secrete insulin in response to glucose, KCl, and carbachol stimulation, in a manner mimicking that of adult mouse islets (50.15±0.16 vs. 33.82±0.35 ng/mg protein at 25 mM glucose; 62.8±0.3 ng/mg protein at 30 mM KCl; 55.59±0.32 ng/mg protein at 100 μM carbachol, Supplementary Fig. S9). In particular, NPE-iPSC-derived cells released more insulin when stimulated with high levels of glucose stimulation (12.22±0.32 ng/mg protein at 25 mM glucose; 12.56±0.24 ng/mg protein at 30 mM KCl; 12.6±0.35 ng/mg protein at 100 μM carbachol, *P<0.05) than did either NM-iPSC- or G4-2 ESC-derived cells.
FIG. 4.
Differentiation of NOD-iPSCs into insulin-producing cells by a stepwise direct differentiation protocol. (A) Immunocytochemical staining confirming stepwise differentiation. NOD-iPSCs and G4-2 ESCs were differentiated into definitive endoderm (Step 1; 3 days; markers, SOX17 and FOXA2); pancreatic progenitor (Step 2; 4 days; marker, PDX1); expanded progenitor (Step 3; 4 days; markers, PAX6 and ISL-1). (B) Quantitative real-time PCR (Step 1 markers: HNF-3 beta, CXCR4, GATA4, and SOX17; Step 2 markers: PDX1, SOX9, and HLXB9; and Step 3 markers: PAX6, ISL-1, NeuroD1, and Neurogenin). Gene expression was normalized to GAPDH. *P<0.05 versus G4-2 control. Scale bars, 100 μm (SOX17, FOXA2, and PDX1; A), and 50 μm (PAX6 and ISL-1; A). The insets are a higher magnification. FOXA2, forkhead box A2. Color images available online at www.liebertonline.com/scd
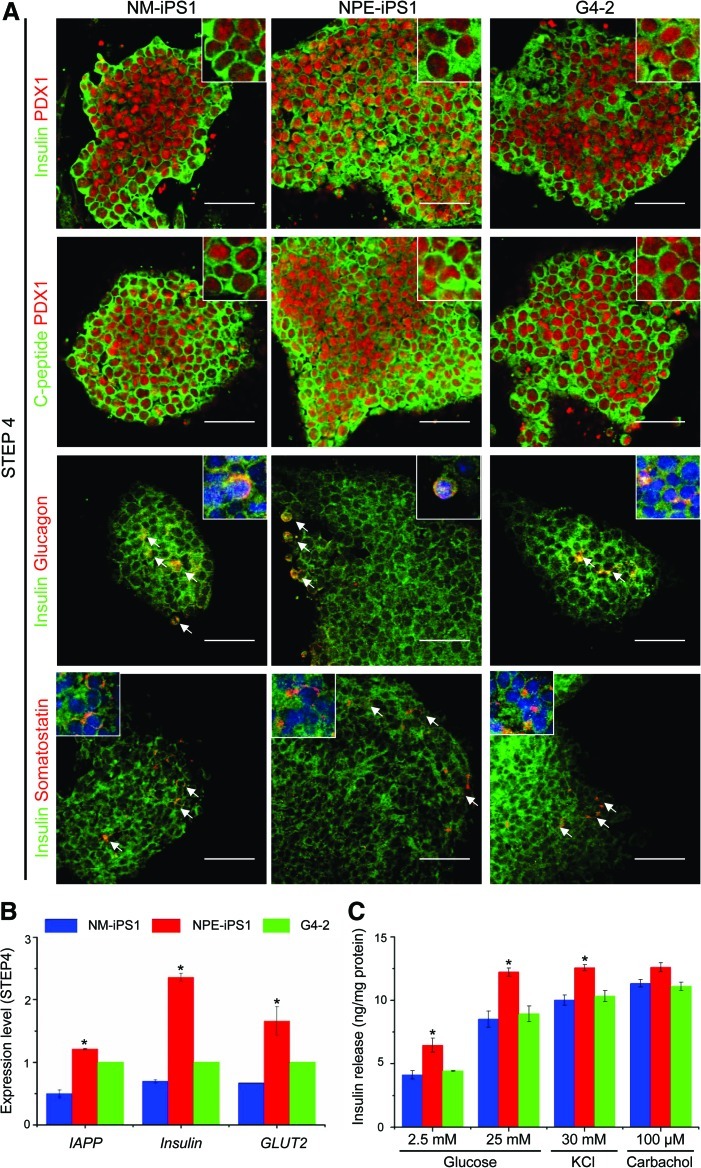
FIG. 5.
Differentiation of NOD-iPSCs into pancreatic insulin-producing cells. (A) Immunocytochemical staining reveals that 2 NOD-iPSC lines (NM-iPS1 and NPE-iPS1), and control ESCs (G4-2), differentiate into pancreatic beta-like cells that express PDX1, Insulin, C-peptide, Glucagon, and Somatostatin. (B) Quantitative real-time PCR analyses indicates that these 2 NOD-iPSC lines could be induced to express islet cell-specific marker genes including IAPP, Insulin, and GLUT2 at the final induction stage. Gene expression was normalized to GAPDH. *P<0.05 versus G4-2 control. (C) Analysis of insulin release from NOD-iPSC-derived insulin-producing cells. NOD-iPSC-derived populations were stimulated with 2.5 mM d-glucose, 25 mM d-glucose, 30 mM KCl, or 100 μM carbachol, and the amount of insulin released into the culture supernatant was analyzed by ELISA. NM-iPSCs, NPE iPSCs, and G4-2 ESCs showed significant differences in insulin release (*P<0.01) when stimulated with 2.5 mM versus 25 mM d-glucose, 30 mM KCl, and 100 μM carbachol. Scale bars, 50 μm (A). The insets are a higher magnification and the white arrows represent Glucagon or Somatostatin. Color images available online at www.liebertonline.com/scd
Transplantation of NPE-iPSC-derived glucose-responsive insulin-secreting cells reverses hyperglycemia in diabetic mice
To determine whether the induced cells could function in vivo, the ability of NPE-iPSC-derived glucose-responsive insulin-producing cells to reverse hyperglycemia was examined in vivo using NOD/SCID mice, which were rendered diabetic by the STZ treatment. Blood glucose levels that were elevated after STZ administration were significantly lower after transplantation of NPE-iPSC-derived cells (n=8; Fig. 6A), suggesting that these cells can normalize hyperglycemia in diabetic mice. Normalization of blood glucose levels occurred within 2–4 days after transplantation. In contrast, sham-transplanted control diabetic mice (n=3) remained hyperglycemic, with glucose levels above 18.3 mM. After transplantation of NPE-iPSC-derived insulin-producing cells, normoglycemia (7.91±0.26 mM) was maintained up to 35 days after transplantation in the NPE-iPSC group, while the sham-transplanted control group continued to exhibit glucose levels above 18.3 mM at 35 days after transplantation. The body weights of the sham-transplanted control and NPE-iPSC-derived cell–transplanted group were also monitored. Consistent with the previous results, only the mice grafted with NPE-iPSC-derived cells transplanted group showed attenuated weight loss and continued to gain body weight over time (Supplementary Fig. S10). Serum insulin level after transplantation of NPE-iPSC-derived cells was 0.17±0.008 ng/mL, which was significantly higher than that in sham-transplanted control diabetic mice (∼0.05±0.01 ng/mL; Fig. 6B), while lower than that in normal mice (1.98±0.021 ng/mL). The amelioration of blood glucose level in mice by the graft of NPE-iPSC-derived cells was not related to spontaneous regeneration of beta cells because we observed no evidence of proliferating beta cells in STZ-treated mice by double immunofluorescence-using anti-BrdU antibody together with -insulin or -nestin antibody (Supplementary Fig. S11). Forty days after transplantation, we removed the graft-bearing kidney and found that removal of grafts from those mice transplanted with NPE-derived cells resulted in a rapid return to hyperglycemia, again confirming that the graft may be responsible for the restoration of glucose levels (Fig. 6A). Hematoxylin/eosin staining after nephrectomy revealed that the transplanted cells were able to engraft under the kidney capsule (Fig. 7A, B). Moreover, DAB staining revealed insulin- and C-peptide-positive cells in the grafts (Fig. 7C). These results indicate that NPE-iPSC-derived glucose-responsive insulin-producing cells, when transplanted into the left subcapsular renal space, were able to secrete insulin in vivo and to lower blood glucose levels in STZ-treated diabetic mice.
FIG. 6.
Transplantation of NPE-iPSC-derived insulin-producing cells into STZ-induced diabetic NOD/SCID mice. (A) Blood glucose levels in transplanted STZ-induced diabetic mice. NPE-iPSC-derived insulin-producing cells were transplanted into STZ-induced diabetic mice. Shown in the graph are sham-transplanted control STZ-induced mice (PBS treated group; n=3; black squares), and STZ-induced mice transplanted with NPE-iPSC-derived insulin producing cells (n=8; gray circles). (B) Circulating insulin levels in transplanted STZ-induced diabetic mice. Insulin in blood from untreated normal mice (n=3), sham-transplanted diabetic mice (n=3), and diabetic mice transplanted with NPE-iPSC-derived insulin-producing cells (n=8), after 3 weeks. *P<0.05 versus sham-transplanted control. STZ, streptozotocin.
FIG. 7.
Analysis of a grafted kidney. (A, B) H&E staining of the grafted kidney. Grafts were removed 5 weeks after transplantation and analyzed by hematoxylin/eosin staining of either the nontransplanted kidney (A) or the NPE-iPSC-grafted kidney (B). The black arrows represent the site of the kidney capsule injection. K, kidney; E, engrafted cells. (C) Expression of insulin, and C-peptide in the graft. Brown diaminobenzidine staining was positive. Sections were counterstained with hematoxylin (blue). Scale bars, 500 μm (A-B), 50 μm (C). H&E, hematoxylin/eosin. Color images available online at www.liebertonline.com/scd
Discussion
Recent studies suggest that cell therapy has potential for treating T1D, which is caused by T-cell-mediated autoimmune destruction of pancreatic beta cells in genetically predisposed individuals [1]. Islet transplantation has been regarded as a potential treatment for T1D for the last 2 decades, following a report of excellent clinical outcomes by researchers in Edmonton [4,6]. However, there are 2 major drawbacks in the application of islet transplantation for the treatment of human T1D. First, the regimen of immunosuppression required to prevent rejection of the allotransplant drastically impacts the lifestyle of the patient, making it inappropriate for younger aged diabetics [2–8]. Second and more importantly, adequate quantities of islet tissue are not available to treat the increasing population of diabetic patients. Thus, one of the most productive areas of stem cell research is cell replacement therapy for TID; however, no one has succeeded in creating a fully functional and mature insulin-producing pancreatic beta cell [46]. Although stem cell-derived beta cells are less efficient than normal beta cells, they are expected to be possibly efficient enough to help a person with diabetes. Our data suggest that as an alternative to the Edmonton protocol, a combination of cell reprogramming and differentiation techniques could be used to generate genetically matched patient-specific pancreatic beta cells. This would create an unlimited source of beta cells and reduce the requirement for immune-suppressive drugs.
NOD mice have been of great value for research in the broad field of T1D [20] and in the development of humanized mice [47]. The NOD mouse strain is considered a nonpermissive strain for ESC derivation [48], because ESC derivation requires specific exogenous factors to maintain the ICM-like pluripotent state [44]. In our study, when the magnet-based nanofection technique was used for generation of exogenous DNA-free iPS cells, we could not stably establish the NOD-iPSCs. However, we were able to efficiently establish NOD-iPSC lines from both NMs and NPEs using a lentiviral expression system. In this system, the OCT4, SOX2, KLF4, and c-MYC genes are expressed under the control of the EF-1 promoter. Our data show that the efficiency of generating iPSCs from NPEs was at least 2-fold that of iPSC generation from NMs. We detected the apparent expression of Nanog and Cripto in NPEs, but not in NMs, suggesting the possibility that the endogenous expression of Nanog and Cripto in NPEs may accelerate reprogramming [49]. Analysis of gene expression patterns revealed that iPSCs established from both NMs and NPEs have pluripotency similar to that of mouse G4-2 ESCs. Additionally, the ability of these cells to spontaneously differentiate into the 3 germ layers was investigated using specific markers, such as endoderm (AFP; hepatocyte and GATA4; zinc-finger transcription factors), mesoderm (brachyury; T-box family of transcription factors and BMP4; a member of the bone morphogenetic protein), and ectoderm (nestin; a type VI intermediate filament protein and beta tubulin III; neuron-specific beta tubulin). These cells were also almost the same as that of control G4-2 ESCs. However, NPE-iPSCs differentiated more efficiently into pancreatic beta-like insulin-producing cells than did NM-iPSCs and G4-2 ESCs. We developed an optimized stepwise differentiation protocol, based on several different direct differentiation methods [40,41], that led to the successful differentiation of NOD-iPSCs into insulin-producing cells. Interestingly, we found differences between the gene expression patterns of differentiated NM-iPSCs and those of NPE-iPSCs (Figs. 4 and 5). Our data confirm that the expression of pancreatic-specific genes in differentiated NPE-iPSCs was higher than in differentiated NM-iPSCs or G4-2 ESCs. This substantiates previous reports that suggested that genetically matched iPSCs retain a transient transcriptional and epigenetic memory of their cellular origin affecting their potential to differentiate into specific cell types [50–52].
Our data, particularly the transplantation results, suggest that cells differentiated from patient-derived iPSCs may be an effective therapy in the treatment for T1D; however, further studies are needed to address the long-term safety and effects of these disease model-derived iPSCs. In particular, tumor formation may be a problem associated with the graft of differentiated ESCs or iPSCs. While we did not detect teratoma formation in the grafts in our study, long-term data are needed to show that NPE-iPSC-derived cells do not form tumors. We expect that NOD-iPSCs will be useful for investigations of genetic susceptibility to autoimmune diseases, and for generating a cellular-interaction model of autoimmune diseases. NOD mice spontaneously develop autoimmune responses in a variety of tissues, including salivary, lacrimal, thyroid, parathyroid, adrenal, testis, large bowel, and red blood cells, in addition to diabetes [28,53]. NOD mice can also be used as models to experimentally induce autoimmune diseases, such as experimental autoimmune thyroiditis, encephalomyelitis, and systemic lupus erythematosus [54]. Therefore, further research with NOD-iPSCs should provide a way to examine the onset and progression of autoimmune diseases in vitro or in reconstituted animal models. These in vitro and in vivo systems will also be useful for testing preventive and therapeutic strategies. In this investigation, we used immunodeficient NOD/SCID mice rendered diabetic by the STZ treatment. One of the advantages of using iPSC-derived beta cells for cell therapy is the avoidance of alloimmunity. However, in the case of autoimmune T1D, autoimmunity that persists after establishment of the disease could pose serious practical and theoretical problems for the application of iPSC-derived beta cell therapy. Thus, further studies using immunocompetent NOD mice are necessary to show that iPSC-derived beta cells, in conjunction with immunosuppression to inhibit autoimmune destruction of grafted iPSC-derived beta cells, can treat established T1D. In addition to the problem of ongoing autoimmunity, a recent study suggests that there may also be immune rejection of even autologous iPSCs [55]. It is therefore necessary to determine if the immunogenicity of transplanted iPSC-derived cells leads to immune rejection. Our data support the potential use of patient-derived iPSCs in autologous beta cell transplantation for the treatment of diabetes, although additional measures may be necessary to avoid the immunogenicity that is intrinsically associated with the production of iPSCs and their differentiation into beta cells.
In summary, we have successfully established NOD mouse iPSC lines and applied a chemically defined stepwise method to differentiate NOD-iPSCs into functional pancreatic beta cells. NPE-iPSCs more efficiently differentiated into this cell type than the NM-iPSCs and even ESCs, suggesting that epigenetic memory may predispose. The differentiated NOD-iPSCs expressed diverse markers for pancreatic beta cells and released insulin in response to glucose and KCl stimulation. Furthermore, we found that the NOD-iPSC-derived insulin-producing cells could ameliorate hyperglycemia in diabetic mice.
Supplementary Material
Acknowledgments
We thank Dr. Natalie DeWitt (Institut Pasteur Korea, Korea), Dr. Kyung-Mi Lee (Korea University, Korea), Dr. Andre Veillette (Clinical Research Institute of Montreal, Canada), Dr. Cassian Yee (Dept. Medicine, University of Washington), and Dr. Ofer Mandelboim (Hebrew University, Israel) for helpful suggestions and critical reading of the article. This work was supported by a grant from the Korea Health 21 R&D Project, Ministry for Health, Welfare and Family Affairs, Republic of Korea (A08-4065) and by a National Research Foundation (NRF) grant funded by the Korea government (MEST) (No. 2010-0020348). We acknowledge the graduate fellowship provided by the Ministry of Education and Human Resources Development through the Brain Korea 21 project, Republic of Korea.
Author Disclosure Statement
None of the authors has any conflict of interest to disclose.
References
- 1.Deruytter N. Boulard O. Garchon HJ. Mapping non-class II H2-linked loci for type 1 diabetes in nonobese diabetic mice. Diabetes. 2004;53:3323–3327. doi: 10.2337/diabetes.53.12.3323. [DOI] [PubMed] [Google Scholar]
- 2.Report of the expert committee on the diagnosis and classification of diabetes mellitus. Diabetes Care. 1997;20:1183–1197. doi: 10.2337/diacare.20.7.1183. [DOI] [PubMed] [Google Scholar]
- 3.Naftanel MA. Harlan DM. Pancreatic islet transplantation. PLoS Med. 2004;1:e58. doi: 10.1371/journal.pmed.0010058. quiz e75. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Lakey JR. Mirbolooki M. Shapiro AM. Current status of clinical islet cell transplantation. Methods Mol Biol. 2006;333:47–104. doi: 10.1385/1-59745-049-9:47. [DOI] [PubMed] [Google Scholar]
- 5.Robertson RP. Holohan TV. Genuth S. Therapeutic controversy: pancreas transplantation for type I diabetes. J Clin Endocrinol Metab. 1998;83:1868–1874. doi: 10.1210/jcem.83.6.4869. [DOI] [PubMed] [Google Scholar]
- 6.Shapiro AM. Lakey JR. Ryan EA. Korbutt GS. Toth E. Warnock GL. Kneteman NM. Rajotte RV. Islet transplantation in seven patients with type 1 diabetes mellitus using a glucocorticoid-free immunosuppressive regimen. N Engl J Med. 2000;343:230–238. doi: 10.1056/NEJM200007273430401. [DOI] [PubMed] [Google Scholar]
- 7.Hering BJ. Wijkstrom M. Graham ML. Hardstedt M. Aasheim TC. Jie T. Ansite JD. Nakano M. Cheng J, et al. Prolonged diabetes reversal after intraportal xenotransplantation of wild-type porcine islets in immunosuppressed nonhuman primates. Nat Med. 2006;12:301–303. doi: 10.1038/nm1369. [DOI] [PubMed] [Google Scholar]
- 8.Hecht G. Eventov-Friedman S. Rosen C. Shezen E. Tchorsh D. Aronovich A. Freud E. Golan H. El-Hasid R, et al. Embryonic pig pancreatic tissue for the treatment of diabetes in a nonhuman primate model. Proc Natl Acad Sci U S A. 2009;106:8659–8664. doi: 10.1073/pnas.0812253106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.D'Amour KA. Bang AG. Eliazer S. Kelly OG. Agulnick AD. Smart NG. Moorman MA. Kroon E. Carpenter MK. Baetge EE. Production of pancreatic hormone-expressing endocrine cells from human embryonic stem cells. Nat Biotechnol. 2006;24:1392–1401. doi: 10.1038/nbt1259. [DOI] [PubMed] [Google Scholar]
- 10.Kroon E. Martinson LA. Kadoya K. Bang AG. Kelly OG. Eliazer S. Young H. Richardson M. Smart NG, et al. Pancreatic endoderm derived from human embryonic stem cells generates glucose-responsive insulin-secreting cells in vivo. Nat Biotechnol. 2008;26:443–452. doi: 10.1038/nbt1393. [DOI] [PubMed] [Google Scholar]
- 11.Tyndall A. Walker UA. Cope A. Dazzi F. De Bari C. Fibbe W. Guiducci S. Jones S. Jorgensen C, et al. Immunomodulatory properties of mesenchymal stem cells: a review based on an interdisciplinary meeting held at the Kennedy Institute of Rheumatology Division, London, UK, 31 October 2005. Arthritis Res Ther. 2007;9:301. doi: 10.1186/ar2103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Morigi M. Imberti B. Zoja C. Corna D. Tomasoni S. Abbate M. Rottoli D. Angioletti S. Benigni A, et al. Mesenchymal stem cells are renotropic, helping to repair the kidney and improve function in acute renal failure. J Am Soc Nephrol. 2004;15:1794–1804. doi: 10.1097/01.asn.0000128974.07460.34. [DOI] [PubMed] [Google Scholar]
- 13.Duffield JS. Park KM. Hsiao LL. Kelley VR. Scadden DT. Ichimura T. Bonventre JV. Restoration of tubular epithelial cells during repair of the postischemic kidney occurs independently of bone marrow-derived stem cells. J Clin Invest. 2005;115:1743–1755. doi: 10.1172/JCI22593. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Abdi R. Fiorina P. Adra CN. Atkinson M. Sayegh MH. Immunomodulation by mesenchymal stem cells: a potential therapeutic strategy for type 1 diabetes. Diabetes. 2008;57:1759–1767. doi: 10.2337/db08-0180. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Camp DM. Loeffler DA. Farrah DM. Borneman JN. LeWitt PA. Cellular immune response to intrastriatally implanted allogeneic bone marrow stromal cells in a rat model of Parkinson's disease. J Neuroinflammation. 2009;6:17. doi: 10.1186/1742-2094-6-17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Colman A. Dreesen O. Pluripotent stem cells and disease modeling. Cell Stem Cell. 2009;5:244–247. doi: 10.1016/j.stem.2009.08.010. [DOI] [PubMed] [Google Scholar]
- 17.Lee CH. Kim EY. Jeon K. Tae JC. Lee KS. Kim YO. Jeong MY. Yun CW. Jeong DK, et al. Simple, efficient, and reproducible gene transfection of mouse embryonic stem cells by magnetofection. Stem Cells Dev. 2008;17:133–141. doi: 10.1089/scd.2007.0064. [DOI] [PubMed] [Google Scholar]
- 18.Takahashi K. Yamanaka S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell. 2006;126:663–676. doi: 10.1016/j.cell.2006.07.024. [DOI] [PubMed] [Google Scholar]
- 19.Rubin LL. Stem cells and drug discovery: the beginning of a new era? Cell. 2008;132:549–552. doi: 10.1016/j.cell.2008.02.010. [DOI] [PubMed] [Google Scholar]
- 20.Anderson MS. Bluestone JA. The NOD mouse: a model of immune dysregulation. Annu Rev Immunol. 2005;23:447–485. doi: 10.1146/annurev.immunol.23.021704.115643. [DOI] [PubMed] [Google Scholar]
- 21.Wicker LS. Clark J. Fraser HI. Garner VE. Gonzalez-Munoz A. Healy B. Howlett S. Hunter K. Rainbow D, et al. Type 1 diabetes genes and pathways shared by humans and NOD mice. J Autoimmun. 2005;25(Suppl):29–33. doi: 10.1016/j.jaut.2005.09.009. [DOI] [PubMed] [Google Scholar]
- 22.Atkinson MA. Leiter EH. The NOD mouse model of type 1 diabetes: as good as it gets? Nat Med. 1999;5:601–604. doi: 10.1038/9442. [DOI] [PubMed] [Google Scholar]
- 23.Delovitch TL. Singh B. The nonobese diabetic mouse as a model of autoimmune diabetes: immune dysregulation gets the NOD. Immunity. 1997;7:727–738. doi: 10.1016/s1074-7613(00)80392-1. [DOI] [PubMed] [Google Scholar]
- 24.Lakey JR. Singh B. Warnock GL. Rajotte RV. BCG immunotherapy prevents recurrence of diabetes in islet grafts transplanted into spontaneously diabetic NOD mice. Transplantation. 1994;57:1213–1217. doi: 10.1097/00007890-199404270-00013. [DOI] [PubMed] [Google Scholar]
- 25.Chatenoud L. Primo J. Bach JF. CD3 antibody-induced dominant self tolerance in overtly diabetic NOD mice. J Immunol. 1997;158:2947–2954. [PubMed] [Google Scholar]
- 26.Wang T. Singh B. Warnock GL. Rajotte RV. Prevention of recurrence of IDDM in islet-transplanted diabetic NOD mice by adjuvant immunotherapy. Diabetes. 1992;41:114–117. doi: 10.2337/diab.41.1.114. [DOI] [PubMed] [Google Scholar]
- 27.Ryu S. Kodama S. Ryu K. Schoenfeld DA. Faustman DL. Reversal of established autoimmune diabetes by restoration of endogenous beta cell function. J Clin Invest. 2001;108:63–72. doi: 10.1172/JCI12335. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Rabinovitch A. Suarez-Pinzon WL. Shapiro AM. Rajotte RV. Power R. Combination therapy with sirolimus and interleukin-2 prevents spontaneous and recurrent autoimmune diabetes in NOD mice. Diabetes. 2002;51:638–645. doi: 10.2337/diabetes.51.3.638. [DOI] [PubMed] [Google Scholar]
- 29.Wu Q. Salomon B. Chen M. Wang Y. Hoffman LM. Bluestone JA. Fu YX. Reversal of spontaneous autoimmune insulitis in nonobese diabetic mice by soluble lymphotoxin receptor. J Exp Med. 2001;193:1327–1332. doi: 10.1084/jem.193.11.1327. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Gregori S. Giarratana N. Smiroldo S. Uskokovic M. Adorini L. A 1alpha,25-dihydroxyvitamin D(3) analog enhances regulatory T-cells and arrests autoimmune diabetes in NOD mice. Diabetes. 2002;51:1367–1374. doi: 10.2337/diabetes.51.5.1367. [DOI] [PubMed] [Google Scholar]
- 31.Bertry-Coussot L. Lucas B. Danel C. Halbwachs-Mecarelli L. Bach JF. Chatenoud L. Lemarchand P. Long-term reversal of established autoimmunity upon transient blockade of the LFA-1/intercellular adhesion molecule-1 pathway. J Immunol. 2002;168:3641–3648. doi: 10.4049/jimmunol.168.7.3641. [DOI] [PubMed] [Google Scholar]
- 32.Rayat GR. Singh B. Korbutt GS. Rajotte RV. Single injection of insulin delays the recurrence of diabetes in syngeneic islet-transplanted diabetic NOD mice. Transplantation. 2000;70:976–979. doi: 10.1097/00007890-200009270-00016. [DOI] [PubMed] [Google Scholar]
- 33.Sandberg JO. Eizirik DL. Sandler S. IL-1 receptor antagonist inhibits recurrence of disease after syngeneic pancreatic islet transplantation to spontaneously diabetic non-obese diabetic (NOD) mice. Clin Exp Immunol. 1997;108:314–317. doi: 10.1046/j.1365-2249.1997.3771275.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Parrinello S. Samper E. Krtolica A. Goldstein J. Melov S. Campisi J. Oxygen sensitivity severely limits the replicative lifespan of murine fibroblasts. Nat Cell Biol. 2003;5:741–747. doi: 10.1038/ncb1024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Bolton WE. Terrell SP. Andrews KL. Boyd AE. Preparation of primary monolayer cultures of mouse pancreatic epithelial cells. Methods Cell Sci. 1982;7:39–42. [Google Scholar]
- 36.Lee CH. Kim JH. Lee HJ. Jeon K. Lim H. Choi H. Lee ER. Park SH. Park JY, et al. The generation of iPS cells using non-viral magnetic nanoparticle based transfection. Biomaterials. 2011;32:6683–6691. doi: 10.1016/j.biomaterials.2011.05.070. [DOI] [PubMed] [Google Scholar]
- 37.Yamada T. Yoshikawa M. Kanda S. Kato Y. Nakajima Y. Ishizaka S. Tsunoda Y. In vitro differentiation of embryonic stem cells into hepatocyte-like cells identified by cellular uptake of indocyanine green. Stem Cells. 2002;20:146–154. doi: 10.1634/stemcells.20-2-146. [DOI] [PubMed] [Google Scholar]
- 38.Niwa H. Miyazaki J. Smith AG. Quantitative expression of Oct-3/4 defines differentiation, dedifferentiation or self-renewal of ES cells. Nat Genet. 2000;24:372–376. doi: 10.1038/74199. [DOI] [PubMed] [Google Scholar]
- 39.Livak KJ. Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods. 2001;25:402–408. doi: 10.1006/meth.2001.1262. [DOI] [PubMed] [Google Scholar]
- 40.Maehr R. Chen S. Snitow M. Ludwig T. Yagasaki L. Goland R. Leibel RL. Melton DA. Generation of pluripotent stem cells from patients with type 1 diabetes. Proc Natl Acad Sci U S A. 2009;106:15768–15773. doi: 10.1073/pnas.0906894106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Zhang D. Jiang W. Liu M. Sui X. Yin X. Chen S. Shi Y. Deng H. Highly efficient differentiation of human ES cells and iPS cells into mature pancreatic insulin-producing cells. Cell Res. 2009;19:429–438. doi: 10.1038/cr.2009.28. [DOI] [PubMed] [Google Scholar]
- 42.Szot GL. Koudria P. Bluestone JA. Murine pancreatic islet isolation. J Vis Exp. 2007:255. doi: 10.3791/255. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Kim YH. Kim S. Kim KA. Yagita H. Kayagaki N. Kim KW. Lee MS. Apoptosis of pancreatic beta-cells detected in accelerated diabetes of NOD mice: no role of Fas-Fas ligand interaction in autoimmune diabetes. Eur J Immunol. 1999;29:455–465. doi: 10.1002/(SICI)1521-4141(199902)29:02<455::AID-IMMU455>3.0.CO;2-A. [DOI] [PubMed] [Google Scholar]
- 44.Hanna J. Markoulaki S. Mitalipova M. Cheng AW. Cassady JP. Staerk J. Carey BW. Lengner CJ. Foreman R, et al. Metastable pluripotent states in NOD-mouse-derived ESCs. Cell Stem Cell. 2009;4:513–524. doi: 10.1016/j.stem.2009.04.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Teitelman G. Alpert S. Polak JM. Martinez A. Hanahan D. Precursor cells of mouse endocrine pancreas coexpress insulin, glucagon and the neuronal proteins tyrosine hydroxylase and neuropeptide Y, but not pancreatic polypeptide. Development. 1993;118:1031–1039. doi: 10.1242/dev.118.4.1031. [DOI] [PubMed] [Google Scholar]
- 46.Hayden EC. Stem cells: the growing pains of pluripotency. Nature. 2011;473:272–274. doi: 10.1038/473272a. [DOI] [PubMed] [Google Scholar]
- 47.Shultz LD. Ishikawa F. Greiner DL. Humanized mice in translational biomedical research. Nat Rev Immunol. 2007;7:118–130. doi: 10.1038/nri2017. [DOI] [PubMed] [Google Scholar]
- 48.Brook FA. Evans EP. Lord CJ. Lyons PA. Rainbow DB. Howlett SK. Wicker LS. Todd JA. Gardner RL. The derivation of highly germline-competent embryonic stem cells containing NOD-derived genome. Diabetes. 2003;52:205–208. doi: 10.2337/diabetes.52.1.205. [DOI] [PubMed] [Google Scholar]
- 49.Theunissen TW. van Oosten AL. Castelo-Branco G. Hall J. Smith A. Silva JC. Nanog overcomes reprogramming barriers and induces pluripotency in minimal conditions. Curr Biol. 2011;21:65–71. doi: 10.1016/j.cub.2010.11.074. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Polo JM. Liu S. Figueroa ME. Kulalert W. Eminli S. Tan KY. Apostolou E. Stadtfeld M. Li Y, et al. Cell type of origin influences the molecular and functional properties of mouse induced pluripotent stem cells. Nat Biotechnol. 2010;28:848–855. doi: 10.1038/nbt.1667. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Kim K. Doi A. Wen B. Ng K. Zhao R. Cahan P. Kim J. Aryee MJ. Ji H, et al. Epigenetic memory in induced pluripotent stem cells. Nature. 2010;467:285–290. doi: 10.1038/nature09342. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Bar-Nur O. Russ HA. Efrat S. Benvenisty N. Epigenetic memory and preferential lineage-specific differentiation in induced pluripotent stem cells derived from human pancreatic islet Beta cells. Cell Stem Cell. 2011;9:17–23. doi: 10.1016/j.stem.2011.06.007. [DOI] [PubMed] [Google Scholar]
- 53.Alyanakian MA. You S. Damotte D. Gouarin C. Esling A. Garcia C. Havouis S. Chatenoud L. Bach JF. Diversity of regulatory CD4+ T cells controlling distinct organ-specific autoimmune diseases. Proc Natl Acad Sci U S A. 2003;100:15806–15811. doi: 10.1073/pnas.2636971100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Wu O. Chen GP. Chen H. Li XP. Xu JH. Zhao SS. Sheng J. Feng JB. Cai J, et al. The expressions of Toll-like receptor 9 and T-bet in circulating B and T cells in newly diagnosed, untreated systemic lupus erythematosus and correlations with disease activity and laboratory data in a Chinese population. Immunobiology. 2009;214:392–402. doi: 10.1016/j.imbio.2008.10.004. [DOI] [PubMed] [Google Scholar]
- 55.Zhao T. Zhang ZN. Rong Z. Xu Y. Immunogenicity of induced pluripotent stem cells. Nature. 2011;474:212–215. doi: 10.1038/nature10135. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.