Abstract
A large number of proteins were tested for the property of intrinsic phosphorescence in deoxygenated aqueous solution at room temperature. The majority of proteins exhibit phosphorescence under normal solution conditions. Phosphorescence lifetimes from 0.5 millisecond to 2 seconds were observed in three-fourths of the proteins tested. The lifetime appears to correlate with relative isolation of the tryptophan indole side chain from solvent. With few exceptions, proteins in general can be expected to display a phosphorescence lifetime greater than 30 microseconds. This widespread characteristic of proteins has been largely overlooked because long-lived phosphorescence is highly sensitive to quenching by low levels of dissolved oxygen in solution. Protein phosphorescence offers a new time domain and a far wider dynamic range than has been used before for photoluminescence experimentation.
Experimental approaches based on the measurement of intrinsic protein fluorescence have proven immensely useful for a broad range of problems in macromolecular science. However, tryptophan fluorescence, in which photon emission occurs on the nanosecond time scale, is only one of the possible consequences of photoexcitation. Intersystem crossing to the triplet state is another, and this can result in long-lived phosphorescence emission. The ability to measure protein phosphorescence in liquid solutions would greatly expand the potential for photoluminescence studies of protein interactions and dynamics on a longer and often more pertinent time scale. However, this possibility has received little attention from the biochemists, since phosphorescence is thought to be largely a property of the solid state. The results reported here demonstrate that this is not the case.
Debye and Edwards (1) reported in 1952 that phosphorescence can be observed from proteins at cryogenic temperatures. Under these conditions phosphorescence lifetimes for a variety of proteins attain a fairly standard value of 5.5 seconds. Twenty years later, Saviotti and Galley (2) found that two proteins, alcohol dehydrogenase from horse liver and alkaline phosphatase from Escherichia coli, phosphoresce in aqueous solution at room temperature with lifetimes on the order of 1 second. Kai and Imakubo (3) found three additional proteins with lifetimes between 1 and 20 msec.
In studies of protein photoluminescence we showed that oxygen quenches the phosphorescence of even the extraordinarily well-protected tryptophans of alcohol dehydrogenase and alkaline phosphatase with a rate constant of 109M−1 sec−1 (4). Thus when oxygen is present at normal solution concentration (250 μM in air-exposed water), phosphorescence emission with an intrinsic lifetime as long as 5 seconds will be reduced in intensity by a factor of 106 and exhibit a lifetime of only 4 μsec. This explains why the phosphorescence of proteins in solution has escaped detection. The original discovery of long-lived protein phosphorescence by Saviotti and Galley (2) was connected with a fortunate accident in this regard. Molecular oxygen in the protein samples was photolytically removed by the intense xenon-arc beam and the long irradiation times used in that work (4, 5).
To obtain the results listed in Table 1, we reduced oxygen concentrations to subnanomolar levels by using the methods described in (4) and (6). Phosphorescence was measured in the time-resolved mode, and the more intense fluorescence emission, which occurs on a faster time scale, was removed by gating the data acquisition to start at 1 msec after the excitation flash. For single tryptophan proteins, phosphorescence decay is accurately exponential (unlike fluorescence), but the multitryptophan proteins produce multiexponential decay curves. Table 1 lists the slowest emissive decay observed for each protein. Most of the proteins exhibited measurable phosphorescence emission with lifetimes greater than 0.5 msec. Cytochrome c did not; the excited singlet state is internally quenched by resonance energy transfer to the nearby heme, so that the triplet state is minimally populated. For the 38 non-heme proteins tested (Table 1), 25 exhibited phosphorescence lifetimes of 0.5 msec or more. Four more proteins (Table 1) showed detectable phosphorescence but with lifetimes somewhat less than 0.5 msec, which was too short to be resolved by our instrumentation. Only 9 of the 38 non-heme proteins tested failed to show phosphorescence that extended into the millisecond range. It is possible to measure the excited triplet state even for free tryptophan in solution. Pepmiller et al. (7) used triplet state absorption methods to measure a lifetime of 29 μsec for 1-methylindole in aqueous solution. It can thus be expected that almost all protein tryptophans will be found to exhibit measurable phosphorescence with lifetimes that span the range from 30 μsec to 5 seconds.
Table 1.
Phosphorescence from proteins at 20°C. Data from other studies are also shown parenthetically; discrepancies between some of our lifetime values and those previously measured by others are largely due to the better removal of oxygen in our work. The following proteins had detectable phosphorescence but lifetimes less than 0.5 msec: collagenase, monellin, mellitin, and lipoamide dehydrogenase. No phosphorescence was detected from carboxypeptidase A, carboxypeptidase B-DPF, cytochrome c (horse and tuna), elastase, lactalbumin, glucose-6-phosphate, parvalbumin without calcium, thermolysin, lysozyme, and chymotrypsin. The samples in this work contained approximately 2 mg of protein per milliliter in 0.01M sodium phosphate and 0.1M sodium chloride at pH 7.0. The deoxygenating system included glucose oxidase (80 nM), catalase (16 nM), and 0.3% glucose. After incubation at room temperature for 10 minutes, 5 μl of a 17 mM sodium dithionite solution was added to produce a final concentration of 80 μM; the sample was then overlaid with mineral oil and argon, and the cuvette was closed with a glass stopper; see also (6). Azurin was prepared by a modification of the preparation of Ambler and Brown (8). Parvalbumin was prepared from cod with the procedure of Calhoun et al. (9). All other proteins were obtained from Sigma Chemical Company, St. Louis, Missouri
| Protein | Lifetime (msec) |
|---|---|
| Actin, rabbit muscle (10) | (15) |
| Albumin, bovine serum | 0.9 |
| Albumin, human serum (11) | 0.9 (0.5) |
| Albumin, chicken egg | 15 |
| Alcohol dehydrogenase, horse liver (2, 4) | (300, 134) |
| Alcohol dehydrogenase* | 550 |
| Aldolase, rabbit muscle (3) | 45 (20) |
| Alkaline phosphatase, Escherichia coli (2) | 1500 (800) |
| Asparaginase, Escherichia coli | 50 |
| Azurin, Pseudomonas aeruginosa | 400 |
| Cellulase, Aspergillus niger | 200 |
| Edestin, hemp seed | 500 |
| Glucose oxidase, Aspergillus niger | 120 |
| Keratin, sheep wool (12)† | (1400) |
| Lactic dehydrogenase, rabbit muscle | 25 |
| Beta-lactoglobulin | 15 |
| Myosin, rabbit muscle | 100 |
| Nuclease, micrococcal | 400 |
| Parvalbumin (with calcium), cod | 5 |
| Protease (acid), Aspergillus saitoi | 100 |
| Protease Streptomyces griseus | 600 |
| Protease (subtilisin Carlsberg), Bacillus subtilis | 10 |
| Ribonuclease T1, Aspergillus oryzae (3) | 14 (4) |
| Streptokinase, streptococcus | 50 |
| Trypsin, bovine pancrease (3) | (1.4) |
D2O solvent
No solvent.
The bases of this great spread in rates, that is, the specific factors that promote, protect, and destroy the tryptophan triplet state within proteins, are not known. For the single tryptophan proteins studied here, a correlation exists between phosphorescence lifetime and the relative blue shift of the fluorescence emission (Table 2), which is related to the degree of isolation of the indole ring system from solvent water. This suggests that the longer lived phosphors are more fully buried. Phosphorescence emission showed no such correlation but peaked consistently at 440 nm, at least for the fairly long-lived cases we have measured.
Table 2.
Correlation of fluorescence emission maximum and phosphorescence lifetime in single tryptophan proteins
| Protein | Fluorescence maximum(nm) | Phosphorescence lifetime(msec) |
|---|---|---|
| Azurin | 305 | 400 |
| Protease (subtilin Carlsberg) | 310 | 10 |
| Parvalbumin (calcium) | 320 | 5 |
| Ribonuclease T1 | 325 | 14 |
| Mellitin | 340 | < 0.5 |
| Monellin | 345 | < 0.5 |
| Parvalbumin (no calcium) | 350 | < 0.5 |
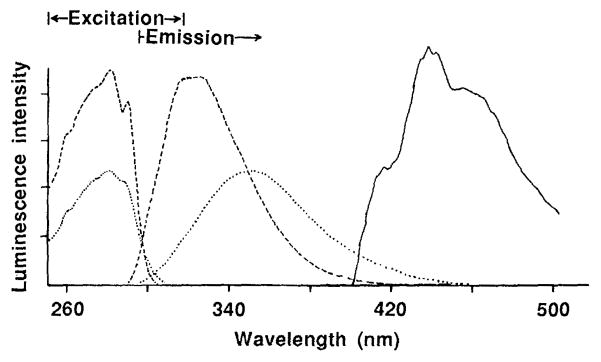
Figure 1 shows typical fluorescence and phosphorescence excitation and emission spectra for calcium-containing parvalbumin. When calcium was removed, the fluorescence emission maximum shifted from 320 to 350 nm, which indicates increased exposure to solvent. The phosphorescence lifetime, which was initially 5 msec, decreased to less than 1 msec.
Fig. 1.
Excitation and emission spectra of cod parvalbumin. The samples contained 2 mg of cod parvalbumin per milliliter in 0.01M sodium phosphate and 0.1M sodium chloride at pH 7.0. For phosphorescence measurements the deoxygenating system described in Table 1 was included. The ordinate for each spectrum is separately scaled for ease of visualization. Fluorescence spectra were taken with protein-bound calcium (- - -) and after addition of 3 mM EDTA to remove calcium (…). A detection wavelength of 330 nm was used with the excitation spectrum and an excitation wavelength of 280 nm was used with the emission spectrum. The phosphorescence emission spectrum (–––) was taken with an excitation wavelength of 280 nm.
Some of the phosphorescence lifetimes could be significantly increased in D2O (Table 1), perhaps because of an effect of the indole NH to ND exchange on the indole vibronic manifold and its coupling to the lattice. Unlike tryptophan fluorescence, phosphorescence emission spectra showed some partially resolved vibrational structure [see also (2) and (4) for other examples].
Protein phosphorescence is more difficult to measure than fluorescence. The problem of collisional quenching by dissolved oxygen can be easily solved. Englander et al. (6) have described methods that allow the preparation and handling of essentially oxygen-free samples in the open air. The relatively low quantum yield for protein phosphorescence puts some demands on instrumentation, but this problem can also be overcome. The widespread occurrence of protein phosphorescence provides many opportunities for extending photoluminescence studies of intra- and intermolecular dynamics into the wide phosphorescence time domain.
Acknowledgments
Supported by NIH grant GM 34448, GM 36393, and AM 11295.
REFERENCES AND NOTES
- 1.Debye P, Edwards JO. Science. 1952;116:143. doi: 10.1126/science.116.3006.143. [DOI] [PubMed] [Google Scholar]
- 2.Saviotti ML, Galley WC. Proc Natl Acad Sci USA. 1974;71:4154. doi: 10.1073/pnas.71.10.4154. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Kai Y, Imakubo K. Photochem Photobiol. 1979;29:261. [Google Scholar]
- 4.Calhoun DB, Vanderkooi JM, Woodrow GV, III, Englander SW. Biochemistry. 1983;22:1526. doi: 10.1021/bi00276a002. [DOI] [PubMed] [Google Scholar]
- 5.Strambini GB. Biophys J. 1983;43:127. doi: 10.1016/S0006-3495(83)84331-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Englander SW, Calhoun DB, Englander JJ. Anal Biochem. in press. [Google Scholar]
- 7.Pepmiller C, Bedwell E, Kuntz RR, Ghiron CA. Photochem Photobiol. 1983;38:273. [Google Scholar]
- 8.Ambler RP, Brown LH. Biochem J. 1967;104:784. doi: 10.1042/bj1040784. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Calhoun DB, Vanderkooi JM, Holtom GR, Englander SW. Proteins Struct Funct Genet. 1986;1:109. doi: 10.1002/prot.340010202. [DOI] [PubMed] [Google Scholar]
- 10.Horie T, Vanderkooi JM. FEBS Lett. 1982;147:69. doi: 10.1016/0014-5793(82)81013-2. [DOI] [PubMed] [Google Scholar]
- 11.Hicks B, White M, Ghiron CA, Kuntz RR, Volkert WA. Proc Natl Acad Sci USA. 1978;75:1172. doi: 10.1073/pnas.75.3.1172. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Leaver IH. Photochem Photobiol. 1979;27:439. [Google Scholar]