Abstract
Background
Next-Generation Sequencing (NGS) technologies and Genome-Wide Association Studies (GWAS) generate millions of reads and hundreds of datasets, and there is an urgent need for a better way to accurately interpret and distill such large amounts of data. Extensive pathway and network analysis allow for the discovery of highly significant pathways from a set of disease vs. healthy samples in the NGS and GWAS. Knowledge of activation of these processes will lead to elucidation of the complex biological pathways affected by drug treatment, to patient stratification studies of new and existing drug treatments, and to understanding the underlying anti-cancer drug effects. There are approximately 141 biological human pathway resources as of Jan 2012 according to the Pathguide database. However, most currently available resources do not contain disease, drug or organ specificity information such as disease-pathway, drug-pathway, and organ-pathway associations. Systematically integrating pathway, disease, drug and organ specificity together becomes increasingly crucial for understanding the interrelationships between signaling, metabolic and regulatory pathway, drug action, disease susceptibility, and organ specificity from high-throughput omics data (genomics, transcriptomics, proteomics and metabolomics).
Results
We designed the Integrated Pathway Analysis Database for Systematic Enrichment Analysis (IPAD, http://bioinfo.hsc.unt.edu/ipad), defining inter-association between pathway, disease, drug and organ specificity, based on six criteria: 1) comprehensive pathway coverage; 2) gene/protein to pathway/disease/drug/organ association; 3) inter-association between pathway, disease, drug, and organ; 4) multiple and quantitative measurement of enrichment and inter-association; 5) assessment of enrichment and inter-association analysis with the context of the existing biological knowledge and a "gold standard" constructed from reputable and reliable sources; and 6) cross-linking of multiple available data sources.
IPAD is a comprehensive database covering about 22,498 genes, 25,469 proteins, 1956 pathways, 6704 diseases, 5615 drugs, and 52 organs integrated from databases including the BioCarta, KEGG, NCI-Nature curated, Reactome, CTD, PharmGKB, DrugBank, PharmGKB, and HOMER. The database has a web-based user interface that allows users to perform enrichment analysis from genes/proteins/molecules and inter-association analysis from a pathway, disease, drug, and organ.
Moreover, the quality of the database was validated with the context of the existing biological knowledge and a "gold standard" constructed from reputable and reliable sources. Two case studies were also presented to demonstrate: 1) self-validation of enrichment analysis and inter-association analysis on brain-specific markers, and 2) identification of previously undiscovered components by the enrichment analysis from a prostate cancer study.
Conclusions
IPAD is a new resource for analyzing, identifying, and validating pathway, disease, drug, organ specificity and their inter-associations. The statistical method we developed for enrichment and similarity measurement and the two criteria we described for setting the threshold parameters can be extended to other enrichment applications. Enriched pathways, diseases, drugs, organs and their inter-associations can be searched, displayed, and downloaded from our online user interface. The current IPAD database can help users address a wide range of biological pathway related, disease susceptibility related, drug target related and organ specificity related questions in human disease studies.
Background
With the age of big data approaching [1], bioinformatics for Next-Generation Sequencing (NGS) and Genome-Wide Association Studies (GWAS) will be one of the biggest areas of disruptive innovation in life science tools over the next few years [2]. Next-Generation Sequencing technologies and Genome-Wide Association Studies generate millions of reads and hundreds of datasets, and there is an urgent need for a better way to accurately interpret and distill such large amounts of data. The use of large scale gene expression analysis has been proven to be useful in identifying differentially expressed genes to classify and predict various disease subtypes. However, it is often difficult to assign biological significance to a large number of genes that are implicated. This problem persists even when users are able to reduce the number of differentially expressed genes substantially via hierarchical clustering methods.
As more information is revealed through large-scale "omics" techniques, it is becoming increasingly apparent that genes do not function alone but through complex biological pathways. Unraveling these intricate pathways is essential to understanding biological mechanisms, disease states, and the function of drugs that transform them. Extensive pathway and network analysis allow for the discovery of highly significant pathways from a set of disease vs. healthy samples in the NGS and GWAS. Knowledge of activation of these processes will lead to elucidation of the complex biological pathways affected by drug treatment, to patient stratification studies of new and existing drug treatments, and to understanding the underlying anti-cancer drug effects.
Pathway databases serve as repositories of current knowledge on cell signaling, enzymatic reaction, and genetic regulation. There are more than 300 pathway repositories listed in Pathguide resource http://www.pathguide.org[3], from which over 141 are specialized on reactions in human as of Jan 2012, for example, BioCarta http://www.biocarta.com[4], KEGG http://www.genome.jp/kegg/[5], NCI-Nature curated http://pid.nci.nih.gov/PID/index.shtml[6], Reactome http://www.reactome.org[7], and Wikipathways http://www.wikipathways.org/[8]. However, these resources have several limitations. First, most currently available resources do not contain disease, drug or organ specificity information such as disease-pathway, drug-pathway, and organ-pathway associations. Next, these resources have been developed with variable degrees of data coverage, quality, and utility [9]. In addition, only half of them provide pathways and reactions in computer-readable formats needed for automatic retrieval and processing [10]. Lastly, many pathway databases are in distinct formats [11].
Systematic collection of pathway information not only in the form of pathway databases but also including inter-association between pathway, disease, drug, and organ specificity is crucial, because 1) it provides a bridge between pathway, disease, drug and organ, and 2) this bridge can not only capture relevant biological pathways but also provide disease, drug target, and organ specificity information. For "inter-association", we refer to a biological connection between two or more biological components on basis of intermediary genes (dotted lines in Figure 1).
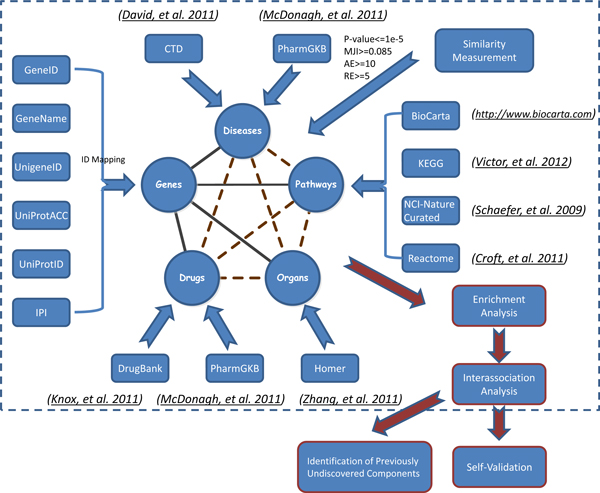
Figure 1.
Data Integration Process. The whole data integration process was divided into three steps: 1) associations of molecule-pathway, molecule-disease, molecule-drug, molecule-organ; 2) inter-association analysis between pathway, disease, drug and organ; and 3) enrichment analysis and inter-association analysis: 3a) self-validation and 3b) identification of previously undiscovered components by the enrichment analysis.
A component is a biomedical concept such as pathway, disease, drug and organ (nodes in Figure 1). Some pilot studies about this kind of connections have been done in the past. For example, Li et al. investigated disease relationships based on their shared pathways [12]. First, they extracted disease associated genes by literature mining. Then, they connected diseases to biological pathways through overlapping genes. Lastly, they built a disease network by connecting diseases sharing common pathways. Smith et al. combined pathway analysis and drug analysis to identify common biological pathways and drug targets across multiple respiratory viruses based on human host gene expression analysis. Their study suggested that multiple and diverse respiratory viruses invoked several common host response pathways [13]. One study found that disease candidate genes were functionally related in the form of protein complexes or biological pathways and complex disease ensued from the malfunction of one or a few specific signaling pathways [14]. Another study aimed to explore complex relationships among diseases, drugs, genes, and target proteins altogether [15] and found that mapping the polypharmacology network onto the human disease-gene network revealed not only that drugs commonly acted on multiple targets but also that drug targets were often involved with multiple diseases. Berger and Iyengar also discussed how analysis of biological networks had contributed to the genesis of systems pharmacology and how these studies had improved global understanding of drug targets [16]. They described that an emerging area of pharmacology, systems pharmacology, which utilizes biological network analysis of drug action as one of its approaches, is becoming increasingly important in: providing new approaches for drug discovery for complex diseases; considering drug actions and side effects in the context of the regulatory networks within which the drug targets and disease gene products function; understanding the relationships between drug action and disease susceptibility genes; and increasing knowledge of the mechanisms underlying the multiple actions of drugs [16].
Therefore we created the Integrated Pathway Analysis Database for Systematic Enrichment Analysis (IPAD) for users to query information about genes, diseases, drugs, organ specificity, and signaling and metabolic pathways. First, we integrated data from four kinds of sources: 1) pathway databases from BioCarta [4], KEGG [5], NCI-Nature curated [6], and Reactome [7], 2) disease databases from CTD http://ctdbase.org/[17] and PharmGKB http://www.pharmgkb.org[18], 3) drug databases from DrugBank httP://www.drugbank.ca[19] and PharmGKB [18], and 4) organ-specific genes/proteins from HOMER http://discern.uits.iu.edu:8340/Homer/index.html[20]. Next, we created inter-association between pathway, disease, drug, and organ specificity. Then, we built a web interface for users to perform 1) enrichment analysis from genes/proteins/molecules, and 2) inter-association analysis from a pathway, disease, drug and organ. Lastly, we presented three case studies: 1) breast cancer related markers, 2) brain-specific markers, and 3) prostate cancer model to demonstrate that the IPAD can enable users to analyze enrichment and inter-association between pathway, disease, drug and organ, to discover previously undiscovered pathway, disease, drug and organ, and to validate the enrichments.
The Integrated Pathway Analysis Database for Systematic Enrichment Analysis (IPAD), located at http://bioinfo.hsc.unt.edu/ipad is a comprehensive database covering about 22,498 genes, 25,469 proteins, 1956 pathways, 6704 diseases, 5615 drugs, and 52 organs integrated from databases including the BioCarta [4], KEGG [5], NCI-Nature curated [6], Reactome [7], CTD [17], PharmGKB [18], DrugBank [19], PharmGKB [18], and HOMER [20].
It is the first comprehensive database that can be used to analyze, discover, and validate enrichment and inter-association between pathway, disease, drug and organ. The inter-associations allow further identification of enriched pathways, diseases, drugs and organs. The quality of the database is validated on a "gold standard" constructed from reputable and reliable sources. The ability to choose multiple quantitative parameters (p-value, Absolute Enrichment Value (AE), Relative Enrichment Value (RE), and Mean Jaccard Index (MJI)) provides us with powerful statistics and computation to accurately calculate enrichment and inter-association. And the cross-linking of multiple data sources enables subsequent validation.
Results
Database content statistics
By integrating pathway, disease, drug, and organ specificity databases including BioCarta [4], KEGG [5], NCI-Nature curated [6], Reactome [7], CTD [17], PharmGKB [18], DrugBank [19], and Homer [20], we have developed IPAD, the Integrated Pathway Analysis Database for systematic enrichment analysis. As of the current release (May 2012), IPAD contains 25,469 proteins (IPI IDs), 22,498 genes (gene IDs), 1956 pathways covering 11663 genes, 6,704 diseases covering 17925 genes, 5,615 drugs covering 3735 genes, and 52 organs covering 5599 genes (Table 1). A comparison of pathways in IPAD against several common pathway data sources is shown in Table 2.
Table 1.
Current Statistics of Database
| Total Number | Count |
|---|---|
| genes | 22,498 GeneIDs |
| proteins | 25,469 UniProtIDs |
| Pathways | 1956 (BioCarta:310,KEGG:247, NCI-Nature curated:222, Reactome:1177) |
| Molecules in Pathway | 11663 |
| Diseases | 6704(CTD:5892, PharmGKB:812) |
| Molecules in Disease | 17925 |
| Drugs | 5615(DrugBank:4604, PharmGKB:1011) |
| Molecules in Drug | 3735 |
| Organs | 52 |
| Molecules in Organ | 5599 |
Table 2.
A Comparison of Human Pathways in IPAD against Several Common Pathway Data Sources
| BioCarta[4] | KEGG[5] | NCI-Nature curated[6] | Reactome[7] | IPAD | |
|---|---|---|---|---|---|
| Pathway coverage | 310 | 247 | 222 | 1177 | 1956 |
| Molecule coverage | 1372 | 9238 | 2561 | 5668 | 11663 |
| Last Updated | 2010 | Mar 2012 | July 2010 | Jan 2011 | Mar 2012 |
| Curation Type | Manual | Manual | Manual | Manual | Integrated |
| Disease Association | No | Yes | No | No | Yes |
| Drug Association | No | Yes | No | No | Yes |
| Organ Specificity Association | No | No | No | No | Yes |
| Inter-associations Quantitative | No | No | No | No | Yes |
| Enrichment Score Quantitative | No | No | No | No | Yes |
| Similarity | No | No | No | No | Yes |
P-value distribution of inter-association
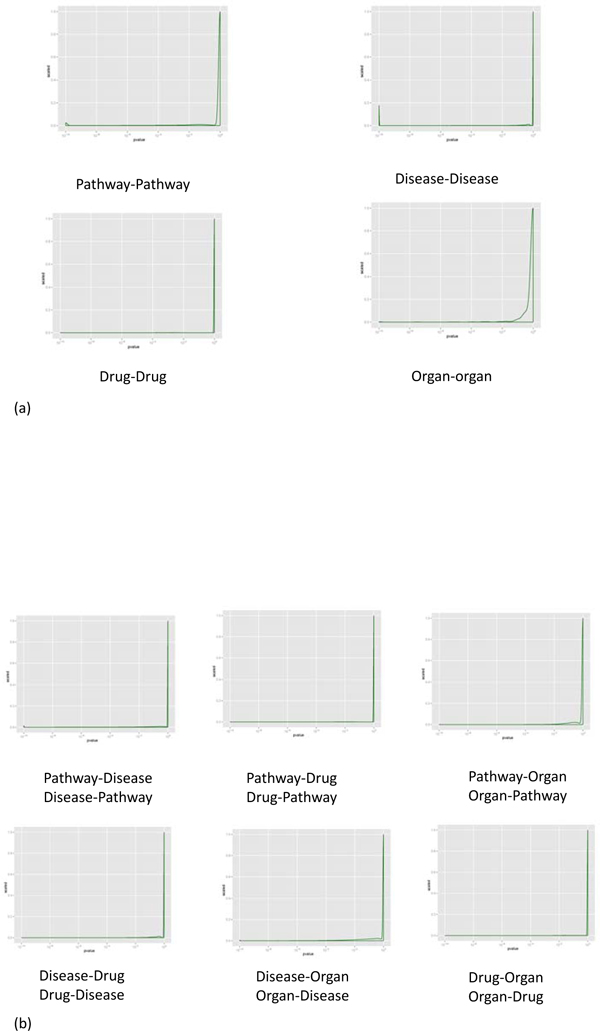
We performed statistical testing using p-value described in the method section to describe the inter-association between pathway, disease, drug and organ in IPAD (Figure 2a and Figure 2b). Although the majority of associations are not significant (p-value close to 1), there are still some which are significant (p-value ≤ 10-5). Component similarity can also be measured by Absolute Expression Value (AE), Relative Expression Value (RE) and Mean Jaccard Index (MJI). The four measurements (p-value, AE, RE, MJI) can complement each other and compensate for the weaknesses inherent in each alone to create better criteria for enrichment analysis.
Figure 2.
p-value Distribution of Inter-association. The y-axis is the scaled density of p-value which is calculated by Fisher Exact test. (a) p-value Distribution of Inter-association between pathway-pathway, disease-disease, drug-drug, and organ-organ. (b) p-value Distribution of Inter-association between pathway, disease, drug, and organ.
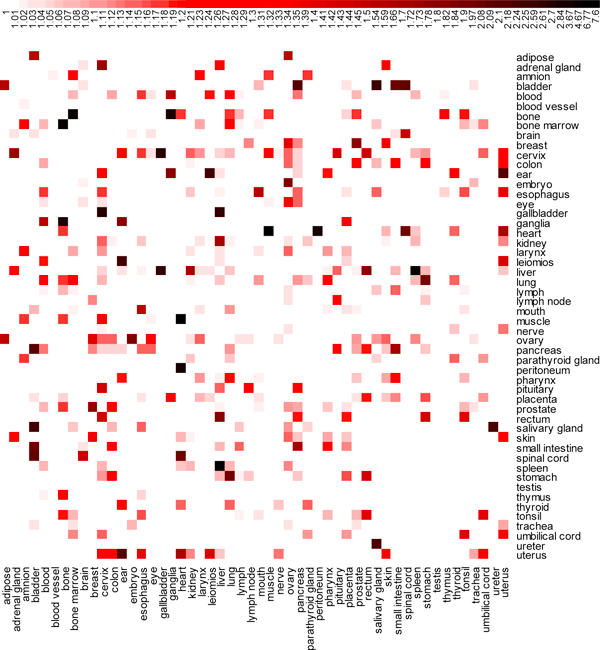
The inter-association between the 52 organs in Figure 3 shows that the heart and muscle have strongest association with a smallest p-value:2.51e-7 (1-log10p-value = 7.6) and 14 genes in common. The other strong associations occur between spleen and liver (20 genes in common, p-value = 1.69e-6, and 1-log10p-value = 6.77), and bone marrow and bone (7 genes in common, p-value = 2.15e-4, and 1-log10p-value = 4.67).
Figure 3.
Heatmap of Inter-associations between 52 Organs. x-axis and y-axis are both 52 organs. The degrees of redness and blackness in each cell represent increase of association between organs. The legend above the heatmap indicates the range of association between organs. The association between organs is expressed by 1 minus log10 of p-value. It is nonlinear color scale from white to red to black, correspondingly, indicating the value of 1-log10(p-value) scales from 1 to 7.6.
General online features
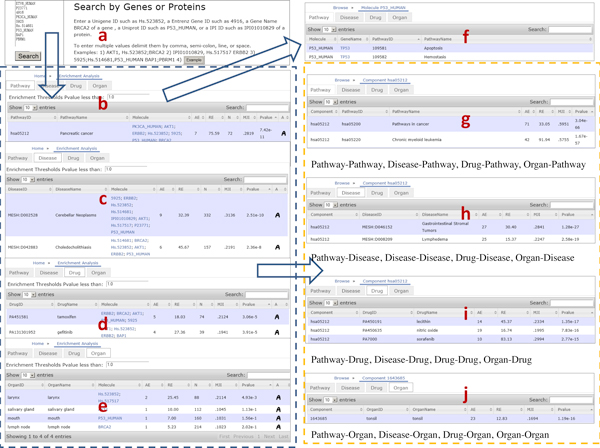
In Figure 4, we show the user interfaces of the web-based online version of IPAD. It supports standard and powerful search options that allow users to specify a list of genes/proteins as the query input. Some interesting features of IPAD include the ability to browse for pathway, disease, drug, and organ with tabs in one page, search by keyword in the Search Box over the table, and set the p-value cutoff in the enrichment threshold box to select enriched pathway sets, disease sets, drug sets and organ sets.
Figure 4.
Web Interface Structure. a) Query by genes or proteins. For example, UniGene IDs, Entrez gene IDs, gene names, UniProt IDs, UniProt Accessions or IPI IDs are all supported. To enter multiple values, delimit them by comma, semi-colon, line or space. b,c,d,e) search result. In the enrichment analysis table, it shows Pathway ID (Disease ID, Drug ID, Organ ID), Pathway Name (Disease Name, Drug Name, Organ Name), Molecule, significance (AE, RE, N, MJI, p-value), and inter-association. For each enriched component, users can further browse the inter-association between its molecule and pathway, disease, drug and organ by clicking on the link in the column of molecule, and its inter-association between pathway, disease, drug and organ by clicking on the inter-association icon in the last column. f) molecule inter-association. It shows molecule, Gene Symbol, Pathway ID (Disease ID, Drug ID, Organ ID), and Pathway Name (Disease Name, Drug Name, Organ Name). g,h,i,j) inter-association between pathway, disease, drug and organ. It shows associations between Pathway-Pathway, Pathway-Disease, Pathway-Drug, Pathway-Organ, Disease-Pathway, Disease-Disease, Disease-Drug, Disease-Organ, Drug-Pathway, Drug-Disease, Drug-Drug, Drug-Organ, Organ-Pathway, Organ-Disease, Organ-Drug, Organ-Organ, and their significance (AE, RE, MJI, p-value).
In response to the query input, IPAD can retrieve a list of related components (pathways, diseases, drugs, and organs) in a highly flexible table, with which users can further explore details about inter-association between the components. For example, users can browse the inter-association between each component's molecule and pathway, disease, drug and organ by clicking on the link in the column of molecule, and look through the component-related inter-association between pathway, disease, drug and organ by clicking on the inter-association icon in the last column. There are totally sixteen types of inter-associations between pathway, disease, drug and organ in IPAD: Pathway-Pathway, Pathway-Disease, Pathway-Drug, Pathway-Organ, Disease-Pathway, Disease-Disease, Disease-Drug, Disease-Organ, Drug-Pathway, Drug-Disease, Drug-Drug, Drug-Organ, Organ-Pathway, Organ-Disease, Organ-Drug, and Organ-Organ. User queried inter-association pathway/disease/drug/organ data stored in IPAD can also be freely downloaded as tab-delimited text files using links below each enrichment or inter-association table.
Assessment of IPAD
Assessing the capabilities of any pathway/disease/drug/organ enrichment analysis in real experiments is a challenge in itself because the full truth of what really occurred between the components and how they are actually inter-associated, if at all, may never be known. In the absence of a "gold standard" - a reference standard against which to establish the performance of the filter, the best alternative is to analyze the results of the enrichment analysis method in the context of the existing biological knowledge [21]. We first used two identified studies to illustrate how well the significant pathways/diseases/drugs/organs identified by the enrichment analysis and inter-association analysis of IPAD fit with the existing biological knowledge. Then we constructed a "gold standard" of 30161 known associations and used it to assess the inter-association analysis of IPAD.
Assessment of enrichment analysis
The absence of a definitive answer regarding the involvement of a particular pathway/disease/drug/organ in a given condition makes it impossible to calculate exact values for sensitivity, specificity, ROCs, etc. Therefore, we compared the result of IPAD's enrichment analysis and inter-association analysis and tested whether the significant pathways/diseases/drugs/organs fit with the existing biological context. This type of assessment is the current best practice in this area of enrichment analysis [22].
In the first dataset, we assessed the features of IPAD by testing the inter-association between breast cancer markers related pathway, disease, drug and organ. Breast cancer is a cancer that starts in the tissues of the breast. We first downloaded the 15 breast cancer related genes from the Cancer Gene Census [23]: AKT1, BAP1, BRCA2, CCND1, CDH1, EP300, ERBB2, ETV6, GATA3, MAP2K4, NTRK3, PBRM1, PIK3CA, RB1, and TP53. The top 5 associated drugs (p-value ≤ 9.9 × 10-3, AE ≥ 2.57, RE ≥ 13.51 and MJI ≥ 0.154; PA451581 tamoxifen, PA131301952 gefitinib, PA152241907 lapatinib, PA449509 estrogens, and PA449383 docetaxel) we identified using IPAD are all reportedly linked to breast cancer by previously published papers (Table 3). For example, most women with estrogen-sensitive breast cancer benefit from the drug tamoxifen [24]. This drug blocks the effects of estrogen, which can help breast cancer cells survive and grow. Green et al. tested whether Gefinitib as an orally active selective EGFR inhibitor might benefit advanced breast cancer (ABC) patients either with acquired hormone resistance or with hormone receptor (HR)-negative tumors. They concluded that at a dose of 500 mg/day, gefitinib monotherapy resulted in a low Clinical Benefit Rate (CBR) and no tumor response was identified [25]. Lapatinib is used as a treatment for treatment-naive women with breast cancer, ER+/EGFR+/HER2+ breast cancer patients (now often called "triple positive") and patients who have HER2-positive advanced breast cancer that has progressed after previous treatment with other chemotherapeutic agents, such as anthracycline, taxane-derived drugs, or trastuzumab [26]. Estrogen is a hormone that is necessary for the normal development and growth of the breasts and organs important for childbearing. For example, several weeks after a study suggested that women who take estrogen-only hormone replacement to treat menopause symptoms may be at lower risk for developing breast cancer, another, much-larger study found that when used for longer than 10 years, estrogen-only regimens actually raise a woman's long-term risk for breast cancer [27]. Docetaxel (given with doxorubicin and cyclophosphamide) is recommended as a possible adjuvant treatment for women with early node-positive breast cancer. For example, Martin et al. compared docetaxel plus doxorubicin and cyclophosphamide (TAC) with fluorouracil plus doxorubicin and cyclophosphamide (FAC) as adjuvant chemotherapy for operable node-positive breast cancer and found that adjuvant chemotherapy with TAC significantly improves the rates of disease-free and overall survival among women with operable node-positive breast cancer [28].
Table 3.
Enrichment Analysis of Breast Cancer Related Markers
| PathwayID | PathwayName | N | P-value | AE | RE | MJI |
|---|---|---|---|---|---|---|
| hsa05212 | Pancreatic cancer | 72 | 3.74E-08 | 7 | 75.59 | 0.2819 |
| hsa05213 | Endometrial cancer | 53 | 1.07E-07 | 6 | 88.02 | 0.2566 |
| hsa05215 | Prostate cancer | 100 | 1.07E-07 | 7 | 54.43 | 0.2683 |
| hsa05223 | Non-small cell lung cancer | 60 | 1.60E-07 | 6 | 77.75 | 0.25 |
| hsa05218 | Melanoma | 72 | 3.55E-07 | 6 | 64.79 | 0.2417 |
| hsa05200 | Pathways in cancer | 348 | 1.02E-06 | 9 | 20.11 | 0.3129 |
| hsa05219 | Bladder cancer | 42 | 1.02E-06 | 5 | 92.56 | 0.2262 |
| h_RacCycDPathway | Influence of Ras and Rho proteins on G1 to S Transition | 26 | 6.66E-06 | 4 | 119.62 | 0.2103 |
| hsa05214 | Glioma | 69 | 6.66E-06 | 5 | 56.34 | 0.2029 |
| hsa05220 | Chronic myeloid leukemia | 74 | 7.91E-06 | 5 | 52.54 | 0.2005 |
| hsa05166 | HTLV-I infection | 272 | 1.71E-05 | 7 | 20.01 | 0.2462 |
| hsa05222 | Small cell lung cancer | 90 | 1.71E-05 | 5 | 43.2 | 0.1944 |
| 200124 | E-cadherin signaling in the nascent adherens junction | 39 | 2.10E-05 | 4 | 79.75 | 0.1846 |
| 200141 | FOXM1 transcription factor network | 41 | 2.37E-05 | 4 | 75.86 | 0.1821 |
| 200190 | a6b1 and a6b4 Integrin signaling | 46 | 3.40E-05 | 4 | 67.61 | 0.1768 |
| hsa04110 | Cell cycle | 124 | 5.80E-05 | 5 | 31.35 | 0.1868 |
| hsa05210 | Colorectal cancer | 62 | 8.23E-05 | 4 | 50.16 | 0.1656 |
| 200041 | Signaling events mediated by Hepatocyte Growth Factor Receptor (c-Met) | 80 | 1.67E-05 | 4 | 38.88 | 0.1583 |
| DiseaseID | DiseaseName | N | P-value | AE | RE | MJI |
| MESH:D002528 | Cerebellar Neoplasms | 332 | 6.34E-07 | 9 | 32.39 | 0.3136 |
| MESH:D020967 | Myotonic Disorders | 278 | 1.87E-05 | 7 | 30.09 | 0.2459 |
| MESH:D042883 | Choledocholithiasis | 157 | 1.87E-05 | 6 | 45.67 | 0.2191 |
| MESH:D002282 | Adenocarcinoma, Bronchiolo-Alveolar | 339 | 3.44E-05 | 7 | 24.68 | 0.2437 |
| MESH:D009134 | Muscular Atrophy, Spinal | 1119 | 3.44E-05 | 11 | 11.75 | 0.3716 |
| MESH:D016510 | Corneal Neovascularization | 669 | 3.44E-05 | 9 | 16.08 | 0.3067 |
| MESH:D044483 | Intestinal Polyposis | 120 | 4.42E-05 | 5 | 49.79 | 0.1875 |
| PA443756 | Colonic Neoplasms | 122 | 4.42E-05 | 5 | 48.98 | 0.1872 |
| PA445062 | Neoplasms | 237 | 4.42E-05 | 6 | 30.25 | 0.2127 |
| MESH:D007972 | Leukoplakia, Oral | 238 | 4.42E-05 | 6 | 30.13 | 0.2126 |
| MESH:D003123 | Colorectal Neoplasms, Hereditary Nonpolyposis | 126 | 4.42E-05 | 5 | 47.42 | 0.1865 |
| MESH:D046152 | Gastrointestinal Stromal Tumors | 148 | 8.01E-05 | 5 | 40.37 | 0.1836 |
| DrugID | DrugName | N | P-value | AE | RE | MJI |
| PA451581 | tamoxifen | 74 | 1.83E-03 | 5 | 18.03 | 0.2124 |
| PA131301952 | gefitinib | 39 | 1.83E-03 | 4 | 27.36 | 0.1941 |
| PA152241907 | lapatinib | 14 | 1.83E-03 | 3 | 57.17 | 0.2143 |
| PA449383 | docetaxel | 77 | 9.81E-03 | 4 | 13.86 | 0.1688 |
| PA449509 | estrogens | 79 | 9.89E-03 | 4 | 13.51 | 0.1682 |
| OrganID | OrganName | N | P-value | AE | RE | MJI |
| larynx | larynx | 88 | 1.97E-2 | 2 | 25.45 | 0.2114 |
By the pathway analysis (p-value ≤ 1.69 × 10-4, AE ≥ 3.03, RE ≥ 20.01 and MJI ≥ 0.158), we identified 18 associated pathways of which most are linked with cancer such as hsa05212 Pancreatic cancer, hsa05213 Endometrial cancer, hsa05215 Prostate cancer, hsa05223 Non-small cell lung cancer, hsa05218 Melanoma, hsa05219 Bladder cancer, hsa05200 Pathways in cancer, hsa05214 Glioma, hsa05220 Chronic myeloid leukemia, hsa05222 Small cell lung cancer, and hsa05210 Colorectal cancer (Table 3). We also discovered 107 diseases (p-value ≤ 1.59 × 10-4, AE ≥ 4.35, RE ≥ 6.31 and MJI ≥ 0.17, Table 3, the top 12 diseases were shown due to space limitation). Most of them are linked with cancer such as MESH:D002528 Cerebellar Neoplasms, MESH:D016510 Corneal Neovascularization, MESH:D002282 Adenocarcinoma, Bronchiolo-Alveolar, MESH:D044483 Intestinal Polyposis, PA443756 Colonic Neoplasms, PA445062 Neoplasms, MESH:D003123 Colorectal Neoplasms, Hereditary Nonpolyposis, and MESH:D046152 Gastrointestinal Stromal Tumors.
By the inter-association, we found that the number 1 pathway (hsa05212, pancreatic cancer) we identified from the enrichment analysis is also highly associated with the pathway (hsa05200, pathways in cancer, p-value = 3.04 × 10-66, 46 orders of magnitude more significant than the pathway-pathway p-value threshold 2.13 × 10-19), disease (MESH:D046152 Gastrointestinal Stromal Tumors, p-value = 1.89 × 10-32, 25 orders of magnitude more significant than the pathway-disease p-value threshold 1.28 × 10-6), and drug (PA450191 lecithin, p-value = 4.55 × 10-11, 7 orders of magnitude more significant than the pathway-drug p-value threshold 5.73 × 10-4). Highly is measured by p-value. When the individual p-values are at least three orders of magnitude lower than current used p-value threshold, they are called "highly significant."
The pathway "hsa05200, pathways in cancer" and disease "MESH:D046152 Gastrointestinal Stromal Tumors" are already included in our previous enrichment analysis and were validated by the inter-association analysis. The drug PA450191 lecithin was filtered out in the enrichment analysis due to its insignificant measurement (p-value = 0.0472, AE = 2, RE = 9.04, MJI = 0.0884) and was discovered by the inter-association analysis as a previously undiscovered drug (p-value = 4.55 × 10-11, AE = 14, RE = 14.53, MJI = 0.2334). Similarly, the number 1 disease (MESH:D002528 Cerebellar Neoplasms) was found to be inter-associated with hsa05200 Pathways in cancer (validated, p-value = 6.86 × 10-42, AE = 79, RE = 9.39, MJI = 0.2536), MESH:D016410 Lymphoma, T-Cell, Cutaneous (previously undiscovered, p-value = 3.76 × 10-100, AE = 320, RE = 6.15, MJI = 0.5389), and PA449780 glutathione (previously undiscovered, p-value = 4.41 × 10-18, AE = 37, RE = 8.20, MJI = 0.3173); and the number 1 drug (PA451581 tamoxifen) was found to be inter-associated with 211859 Biological oxidations (previously undiscovered, p-value = 9.31 × 10-25, AE = 24, RE = 30.06, MJI = 0.2654), PA443560 Breast Neoplasms (previously undiscovered, p-value = 3.26 × 10-50, AE = 49, RE = 35.43, MJI = 0.4042), and PA449503 estradiol (previously undiscovered, p-value = 1.2 × 10-21, AE = 30, RE = 15.45, MJI = 0.3558).
Another dataset we used to assess the enrichment analysis is with the "self-validation" in Case Study 1. The self-validation makes the result of enrichment analysis more reliable and meaningful and consistent with the existing biological context. If a result of enrichment analysis can be validated by its subsequent inter-association analysis, it is also validated that the enrichment analysis and inter-association analysis are consistent and are both somewhat reliable.
Assessment of inter-associated analysis
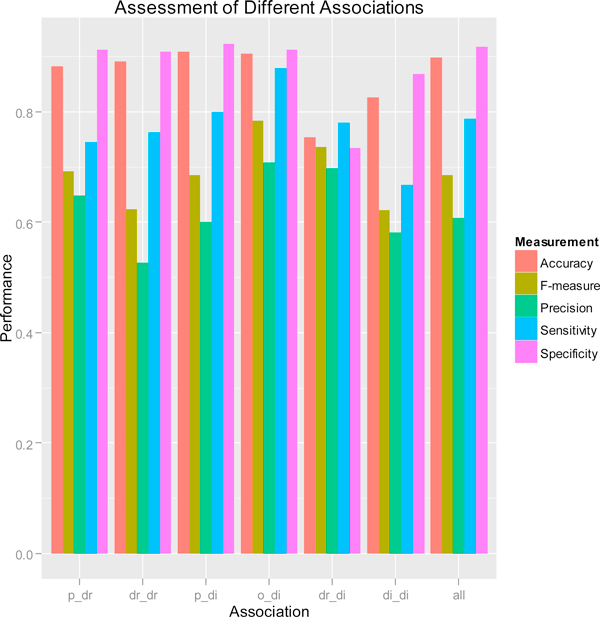
We constructed a "gold standard" of 30161 inter-associations (247 Pathway-Drug; 274 Drug-Drug; 23659 Pathway-Disease; 405 Organ-Disease; 2826 Drug-Disease; 2750 Disease-Disease) from KEGG [5], CTD [17], PharmGKB [18], DrugBank [19], and Disease Ontology http://do-wiki.nubic.northwestern.edu/do-wiki/index.php/Main_Page[29]. We evaluated the performance of inter-association analysis method for the above six types of inter-associations (Figure 5). Pathway-Disease inter-associations have the highest specificity (92.4%), and Organ-Disease inter-associations have the highest sensitivity (87.9%) and F_measure (78.4%).
Figure 5.
Assessment of Different Associations. The bar plot shows sensitivity, specificity, prediction, accuracy, and F_measure of pathway-drug, drug-drug, pathway-disease, organ-disease, drug-disease, disease-disease, and all associations as a whole.
Compared to sensitivity, specificity and accuracy, the prediction rates are relatively low because the size of testing set are much larger than that of the "gold standard" set. When more "gold standards" of inter-associations become available in the future, the prediction rates and F_measure can be improved because the currently unpredicted pairs will be able to be predicted correctly. Figure 5 also gives a global evaluation for all 30161 inter-associations (Precision 60.73%, Accuracy 89.90%, Sensitivity 78.69%, Specificity 91.72%, F_measure 68.56%). Overall, the balanced F_measure (68.56%) shows our inter-association analysis method is reliable and can be used for further enrichment analysis.
Case Studies
We show two case studies of increasing complexity and biological significance to achieve two goals: 1) to demonstrate the IPAD's ability to self-validate by using it to perform enrichment analysis and inter-association analysis on the 369 brain-specific markers, and 2) to demonstrate the ability of IPAD to identify previously undiscovered components by the enrichment analysis based on differentially expressed genes identified from a prostate cancer study.
Case Study 1: Self-validation with inter-association analysis
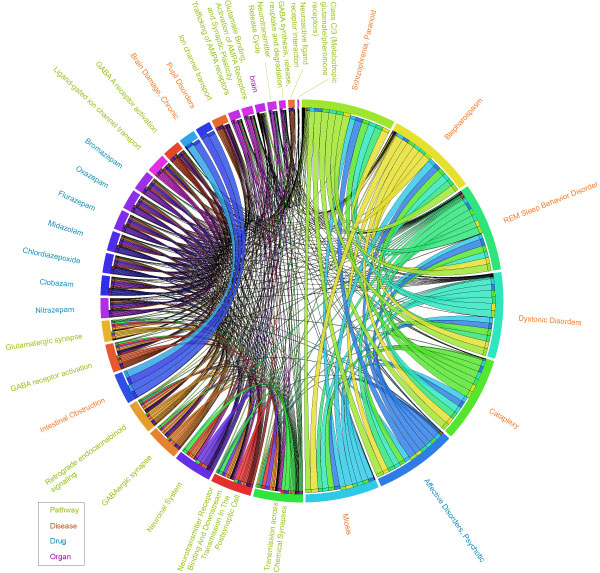
The highly associated relationships between pathway, disease, drug and organ can be used to validate the identified enriched pathway, disease, drug and organ candidates. The more dense and complex the inter-association between the four components, the more reliable and robust the identified candidates. In order to demonstrate the self-validation of IPAD, we first performed enrichment analysis on the 369 brain-specific markers we extracted from Homer [20] and then used the inter-association analysis in IPAD to validate the traditional enrichment analysis. We identified 16 enriched pathways (p-value ≤ 5.67 × 1056, AE ≥ 4.86, RE ≥ 7.42 and MJI ≥ 0. 107), 92 enriched diseases (p-value ≤ 4.52 × 10-7, AE ≥ 28.55, RE ≥ 1.31 and MJI ≥ 0.10), 7 enriched drugs (p-value ≤ 2.49 × 10-7, AE ≥ 8.06, RE ≥ 26.98 and MJI ≥ 0.30), and 1 enriched organ (p-value ≤ 0.05, AE ≥ 260, RE ≥ 15.42 and MJI ≥ 0.85) (Table 4, only 10 diseases are shown due to space limitation). All components were validated by the inter-association analysis except that only 88 out of 92 diseases were validated. Due to space limitation, we selected the top 10 diseases and other components to draw a circular view. The circular view of the 16 pathways, 7 drugs, 1 organ and top 10 diseases shows that all these 34 components are inter-associated with at least one other component (Figure 6). The table visualization was created by the tableviewer utility script, which is included in Circos [30]. We set the four text colors: palegreen, chocolate, royalblue, and magenta which stand for the four components: pathway, disease, drug, and organ, respectively. We transformed the extent of association between two components by using 1 minus log10 of p-value and set links with variable thickness representing the extent of inter-associations. The direction of association (A->B) is represented by a ribbon's end touching A and its other end not touching B.
Table 4.
Enrichment Analysis of Brain-Specific Markers
| Rank | PathwayID | PathwayName | N | Pvalue | AE | RE | MJI |
|---|---|---|---|---|---|---|---|
| 1 | 112315 | Transmission across Chemical Synapses | 190 | 0 | 27 | 17.26 | 0.2117 |
| 2 | 112316 | Neuronal System | 283 | 0 | 32 | 13.74 | 0.2232 |
| 3 | hsa04723 | Retrograde endocannabinoid signaling | 116 | 4.68E-14 | 18 | 18.85 | 0.1713 |
| 4 | 112314 | Neurotransmitter Receptor Binding And Downstream Transmission In The Postsynaptic Cell | 136 | 4.48E-13 | 18 | 16.08 | 0.1599 |
| 5 | hsa04727 | GABAergic synapse | 98 | 5.48E-13 | 16 | 19.84 | 0.165 |
| 6 | hsa04080 | Neuroactive ligand-receptor interaction | 401 | 4.42E-12 | 26 | 7.88 | 0.1678 |
| 7 | 977441 | GABA A receptor activation | 12 | 7.38E-09 | 7 | 70.87 | 0.3281 |
| 8 | 975298 | Ligand-gated ion channel transport | 25 | 1.41E-08 | 8 | 38.88 | 0.2017 |
| 9 | 977443 | GABA receptor activation | 53 | 1.30E-07 | 9 | 20.63 | 0.1318 |
| 10 | hsa04724 | Glutamatergic synapse | 134 | 2.62E-07 | 12 | 10.88 | 0.1073 |
| 11 | 983712 | Ion channel transport | 61 | 3.20E-07 | 9 | 17.92 | 0.1206 |
| 12 | 420499 | Class C/3 (Metabotropic glutamate/pheromone receptors) | 15 | 5.19E-07 | 6 | 48.6 | 0.2313 |
| 13 | 888590 | GABA synthesis, release, reuptake and degradation | 19 | 1.52E-06 | 6 | 38.37 | 0.1891 |
| 14 | 399719 | Trafficking of AMPA receptors | 30 | 1.33E-05 | 6 | 24.3 | 0.1313 |
| 15 | 399721 | Glutamate Binding, Activation of AMPA Receptors and Synaptic Plasticity | 30 | 1.33E-05 | 6 | 24.3 | 0.1313 |
| 16 | 112310 | Neurotransmitter Release Cycle | 36 | 3.03E-05 | 6 | 20.25 | 0.1146 |
| Rank | DiseaseID | DiseaseName | N | Pvalue | AE | RE | MJI |
| 1 | MESH:D001764 | Blepharospasm | 699 | 0 | 45 | 5.52 | 0.1398 |
| 2 | MESH:D012563 | Schizophrenia, Paranoid | 649 | 3.18E-12 | 40 | 5.29 | 0.1265 |
| 3 | MESH:D002385 | Cataplexy | 723 | 1.33E-11 | 41 | 4.86 | 0.1264 |
| 4 | MESH:D020187 | REM Sleep Behavior Disorder | 506 | 1.44E-11 | 34 | 5.76 | 0.1149 |
| 5 | MESH:D020821 | Dystonic Disorders | 837 | 4.52E-10 | 41 | 4.2 | 0.1226 |
| 6 | MESH:D015877 | Miosis | 1000 | 1.94E-09 | 44 | 3.77 | 0.1273 |
| 7 | MESH:D001925 | Brain Damage, Chronic | 1732 | 1.20E-08 | 59 | 2.92 | 0.1582 |
| 8 | MESH:D000341 | Affective Disorders, Psychotic | 700 | 1.50E-08 | 34 | 4.17 | 0.1056 |
| 9 | MESH:D007415 | Intestinal Obstruction | 1293 | 2.33E-08 | 48 | 3.18 | 0.1334 |
| 10 | MESH:D011681 | Pupil Disorders | 1612 | 2.61E-08 | 55 | 2.93 | 0.1486 |
| Rank | DrugID | DrugName | N | Pvalue | AE | RE | MJI |
| 1 | DB01595 | Nitrazepam | 20 | 2.92E-08 | 10 | 29.64 | 0.3294 |
| 2 | DB00349 | Clobazam | 19 | 4.26E-08 | 9 | 28.08 | 0.3083 |
| 3 | DB00475 | Chlordiazepoxide | 19 | 4.26E-08 | 9 | 28.08 | 0.3083 |
| 4 | DB00683 | Midazolam | 19 | 4.26E-08 | 9 | 28.08 | 0.3083 |
| 5 | DB00690 | Flurazepam | 19 | 4.26E-08 | 9 | 28.08 | 0.3083 |
| 6 | DB00842 | Oxazepam | 19 | 4.26E-08 | 9 | 28.08 | 0.3083 |
| 7 | DB01558 | Bromazepam | 19 | 4.26E-08 | 9 | 28.08 | 0.3083 |
| Rank | OrganID | OrganName | N | Pvalue | AE | RE | MJI |
| 1 | brain | brain | 363 | 0 | 260 | 15.42 | 0.8581 |
Figure 6.
A Circular View of the Inter-association Analysis of 369 Brain-Specific Markers. The text colors for the four components: pathway, disease, drug, and organ are palegreen, Chocolate, royalblue, and magenta, respectively. Links with variable thickness represent the extent of association between two components which is 1 minus log10 of p-value. The direction of association (A->B) is represented by a ribbon's end touching A and its other end not touching B.
The 10 identified diseases: 1) MESH:D001764, Blepharospasm, 2) MESH:D012563, Schizophrenia, Paranoid, 3) MESH:D002385, Cataplexy, 4) MESH:D020187, REM Sleep Behavior Disorder, 5) MESH:D020821, Dystonic Disorders, 6) MESH:D015877, Miosis, 7) MESH:D001925, Brain Damage, Chronic, 8) MESH:D000341, Affective Disorders, Psychotic, 9) MESH:D007415, Intestinal Obstruction, and 10) MESH:D011681, Pupil Disorders, have on average 766 inter-associations between pathway, disease, drug and organ, which shows a strong association with those 369 brain-specific markers.
A blepharospasm is any abnormal contraction or twitch of the eyelid. There have been several important advances in understanding the brain mechanisms associated with blepharospasm. Baker et al. identified blinking-induced functional magnetic resonance imaging (fMRI) activation patterns in five benign essential blepharospasm (BEB) patients and five age-matched control subjects and concluded that the activations observed might represent a hyperactive cortical circuit linking visual cortex, limbic system, supplementary motor cortex, cerebellum, and supranuclear motor pathways innervating the periorbital muscles [31]. Antal et al. examined whether magnetic or electrical stimulation of the brain could improve the involuntary closure of the eyelids in patients with blepharospasm or Meige syndrome [32].
Schizophrenia is a brain disorder that affects the way a person acts, thinks, and sees the world. People with schizophrenia have an altered perception of reality, often a significant loss of contact with reality. Chen et al. utilized a multivariate approach to identify genomic risk components associated with brain function abnormalities in schizophrenia [33]. They first derived 5157 candidate single nucleotide polymorphisms (SNPs) from genome-wide array based on their possible connections with schizophrenia and further investigated for their associations with brain activations captured with functional magnetic resonance imaging (fMRI) during a sensorimotor task. Then, they identified 222 SNPs which showed significant difference between 92 schizophrenia patients and 116 healthy controls. Their further pathway analysis showed that the genes associated with the identified SNPs participated in four neurotransmitter pathways: GABA receptor signaling, dopamine receptor signaling, neuregulin signaling and glutamate receptor signaling. Their finding is consistent with our inter-association analysis from the 369 brain-specific markers.
Our 16 pathways identified by inter-association analysis using IPAD contains 1) Neurotransmitter Receptor Binding And Downstream Transmission In The Postsynaptic Cell, 2) Neuroactive ligand-receptor interaction, 3) GABAergic synapse, 4) GABA receptor activation, 5) Glutamate Binding, Activation of AMPA Receptors and Synaptic Plasticity, 6) Neurotransmitter Release Cycle, 7) GABA synthesis, release, reuptake and degradation, 8) Class C/3 (Metabotropic glutamate/pheromone receptors), and 9) GABA A receptor activation etc.
The other 7 diseases (except Intestinal Obstruction) also show strong links with brain, such as Cataplexy [34], REM Sleep Behavior Disorder [35], Dystonic Disorders [36], Miosis [37], Brain Damage [38], Chronic [39], Affective Disorders [40], Psychotic [41], and Pupil Disorders [42].
The 7 identified drugs: 1) DB00349, Clobazam, 2) DB00475, Chlordiazepoxide, 3) DB00683, Midazolam, 4) DB00690, Flurazepam, 5) DB00842, Oxazepam, 6) DB01558, Bromazepam, and 7) DB01595, Nitrazepam have on average 63 inter-associations between pathway, disease, drug and organ. They show strong links with brain, such as 1) Clobazam [43], 2) Chlordiazepoxide [44], 3) Midazolam [45], 4) Flurazepam [46], 5) Oxazepam [47], 6) Bromazepam [48], and 7) Nitrazepam [49].
In conclusion, this case study shows that the self-validation of IPAD is an innovation of traditional enrichment analysis and can be useful for validating any pathways, diseases, drugs or organs that users identify with their own data and methods.
Case Study 2: Identification of previously undiscovered components by IPAD
RNA-seq is an emerging technology for surveying gene expression and transcriptome content by directly sequencing the mRNA molecules in a sample. RNA-seq can provide gene expression measurements and is regarded as an attractive approach to analyze a transcriptome in an unbiased and comprehensive manner. In this case study, we demonstrate the use of IPAD to identify previously undiscovered components by the enrichment analysis based on differentially expressed genes identified from the transcriptional profiling sequencing data [50]. The original purpose is to provide a general guide for analysis of gene expression and alternative splicing by deep sequencing. In the prostate cancer study, the prostate cancer cell line LNCap was treated with androgen/DHT. Mock-treated and androgen-stimulated LNCap cells were sequenced using the Illumina 1G Genome Analyzer. For the mock-treated cells, there were four lanes totaling ~10 million reads. For the DHT-treated cells, there were three lanes totaling ~7 million reads. All replicates were technical replicates. Samples labeled s1 through s4 are from mock-treated cells. Samples labeled s5, s6, and s8 are from DHT-treated cells. The read sequences are stored in FASTA files. The sequence IDs break down as follows: seq_(unique sequence id)_(number of times this sequence was seen in this lane). We first downloaded the publicly available transcriptional profiling sequencing data from the author's Web Site at http://yeolab.ucsd.edu/yeolab/Papers.html and computed the digital gene expression, next identified 278 differentially expressed genes in RNA-seq data from hormone treated prostate cancer cell line samples, then performed the enrichment analysis of the 278 genes with IPAD, and lastly carried out the inter-association analysis for these enriched components with IPAD.
In total, we identified 11 enriched pathways (p-value ≤ 5 × 10-2,AE ≥ 3.45, RE ≥ 1.95 and MJI ≥ 0.040), 100 diseases(p-value ≤ 1.6 × 10-3, AE ≥ 68.35, RE ≥ 1.30 and MJI ≥ 0.147), and 2 organs (p-value ≤ 1.9 × 10-2,AE ≥ 4.38, RE ≥ 5.45 and MJI ≥ 0.080) for the 278 genes. And the further inter-association analysis of IPAD identified 10 pathways, 8 diseases, 2 drugs and 1 organs which are not previously discovered by the enrichment analysis of IPAD (Table 5).
Table 5.
Identification of Previously Undiscovered Components by IPAD
| PathwayID | PathwayName | p-value | AE | RE | MJI | C |
|---|---|---|---|---|---|---|
| 1430728 | Metabolism | 4.32E-35 | 525 | 2.19 | 0.34 | 93 |
| 556833 | Metabolism of lipids and lipoproteins | 1.00E-13 | 175 | 2.25 | 0.27 | 91 |
| 453279 | Mitotic G1-G1/S phases | 2.41E-42 | 47 | 27.44 | 0.33 | 87 |
| 200137 | AP-1 transcription factor network | 2.36E-06 | 44 | 2.97 | 0.32 | 87 |
| 453279 | Mitotic G1-G1/S phases | 2.77E-09 | 78 | 2.73 | 0.30 | 87 |
| 200120 | Direct p53 effectors | 6.09E-07 | 70 | 2.4 | 0.27 | 85 |
| 69278 | Cell Cycle, Mitotic | 1.81E-14 | 166 | 2.37 | 0.28 | 82 |
| 1640170 | Cell Cycle | 1.51E-12 | 183 | 2.11 | 0.26 | 80 |
| 535734 | Fatty acid, triacylglycerol, and ketone body metabolism | 1.38E-07 | 81 | 2.35 | 0.26 | 79 |
| 71291 | Metabolism of amino acids and derivatives | 7.54E-15 | 120 | 2.9 | 0.33 | 76 |
| DiseaseID | DiseaseName | p-value | AE | RE | MJI | C |
| MESH:D015228 | Hypertriglyceridemia | 1.60E-192 | 4573 | 2.09 | 0.77 | 79 |
| MESH:D009468 | Neuromuscular Diseases | 4.34E-07 | 117 | 2.04 | 0.41 | 74 |
| MESH:D009468 | Neuromuscular Diseases | 2.16E-293 | 4573 | 2.52 | 0.82 | 74 |
| MESH:D052016 | Mucositis | 4.99E-199 | 4572 | 2.11 | 0.77 | 72 |
| MESH:D002543 | Cerebral Hemorrhage | 7.21E-128 | 4572 | 1.8 | 0.73 | 72 |
| MESH:D006463 | Hemolytic-Uremic Syndrome | 3.65E-215 | 4570 | 2.18 | 0.78 | 72 |
| MESH:D020246 | Venous Thrombosis | 6.59E-106 | 4573 | 1.71 | 0.72 | 70 |
| MESH:D013923 | Thromboembolism | 1.78E-123 | 4573 | 1.79 | 0.73 | 70 |
| DrugID | DrugName | p-value | AE | RE | MJI | C |
| PA449383 | Docetaxel | 3.08E-02 | 52 | 1.89 | 0.36 | 24 |
| PA449780 | Glutathione | 1.74E-10 | 20 | 10.48 | 0.24 | 13 |
| PA131301952 | Gefitinib | 8.52E-17 | 29.88 | 10.84 | 0.395 | 8 |
| PA451283 | Rosiglitazone | 7.73E-22 | 41.4 | 10.58 | 0.386 | 5 |
| PA448803 | Carboplatin | 2.57E-15 | 27 | 11.36 | 0.397 | 5 |
| OrganID | OrganName | p-value | AE | RE | MJI | C |
| liver | Liver | 1.82E-17 | 179 | 2.72 | 0.40 | 84 |
We found that some of these components that were previously undiscovered but identified by inter-association analysis still showed strong association with prostate cancer. For example, previous studies reported that the top 5 drugs we identified with inter-association analysis: docetaxel, glutathione, gefitinib, rosiglitazone, and carboplatin were all associated with prostate cancer. Docetaxel is a drug used in men whose prostate cancer no longer responds to hormone therapy. Tannock et al. compared docetaxel plus prednisone in men with advanced, hormone-refractory prostate cancer with mitoxantrone plus prednisone. They found that treatment with docetaxel every three weeks led to superior survival and improved rates of response in terms of pain, serum PSA level, and quality of life, as compared with mitoxantrone plus prednisone, when given with prednisone [51]. The deficiency in the glutathione enzyme system has been proposed to increase the likelihood of developing both an enlarged prostate and prostate cancer. Nelson discovered a genetic defect in prostate cancer cell prevents the body from producing glutathione S-transferase (GST), an enzyme needed by the liver to detoxify harmful chemicals [52]. The function of a particular glutathione enzyme glutathione-S-transferase-pi-i (GSTP1) is almost universally lost in both cancerous and pre-cancerous prostate cells. The inactivation of this glutathione enzyme is an early event in the development of prostate cancer. Many studies have linked the loss of GSTP 1 to malignant transformation of prostatic tissues [52].
One study found that gefitinib and bicalutamide showed synergistic effects in primary cultures of prostate cancer derived from androgen-dependent naive patients [53]. Another study discovered that gefitinib-trastuzumab combination showed promising clinical activity in hormone refractory prostate cancer [54]. Smith et al. assessed the biological activity of rosiglitazone, a peroxisome proliferator-activated receptor gamma agonist that has been approved to treat type 2 diabetes, in men with recurrent prostate carcinoma using change in prostate specific antigen (PSA) doubling time (PSADT) as the primary outcome variable and concluded that Rosiglitazone did not increase PSADT or prolong the time to disease progression more than placebo in men with a rising PSA level after radical prostatectomy and/or radiation therapy [55]. But Rosiglitazone was found to suppress human lung carcinoma cell growth through PPARγ-dependent and PPARγ-independent signal pathways [56]. The number 3 drug, Carboplatin is a chemotherapy agent used for treatment of many types of cancer. Some studies examined the efficacy of carboplatin as a second line chemotherapy agent (after the failure of taxotere) as well as along with taxotere therapy for men with advanced prostate cancer [57,58]. A phase II study assessed the outcome and predictive factors for prognosis and toxicity following intermittent chemotherapy with docetaxel, estramustine phosphate, and carboplatin (DEC) in patients with castrate resistant prostate cancer (CRPC) and found that combination chemotherapy with DEC has a potential effect on CRPC with acceptable toxicity [59]. Jeske et al. conducted a retrospective, bi-institutional review of patients with advanced CRPC treated with carboplatin plus paclitaxel after docetaxel and concluded that Carboplatin/paclitaxel chemotherapy following docetaxel in metastatic CRPC is well tolerated with favorable PSA response rates and survival and the combination is a viable option after progression on docetaxel-based therapy [60].
This case study shows that compared to traditional enrichment analysis, the IPAD's inter-association analysis can be more powerful and useful in identification of previously undiscovered pathways, diseases, drugs or organ specification.
Conclusion
We developed IPAD as an integrated database system to analyze, identify, and validate pathway, disease, drug, organ specificity and their inter-associations. IPAD integrates many different types of pathway, disease, drug and organ-specificity information: pathway gene relationship from the BioCarta [4], KEGG [5], NCI-Nature curated [6], and Reactome [7] database; disease gene relationship from the CTD [17] and PharmGKB [18] database; drug gene relationship from the DrugBank [19] and PharmGKB [18] database; and organ-specific genes/proteins from the HOMER [20] databases.
Enriched pathways, diseases, drugs, organs and their inter-associations can be searched, displayed, and downloaded from our online user interface. The current IPAD database can help users address a wide range of pathway related, disease related, drug related and organ specificity related questions in human disease studies. We also developed a statistical method for similarity measurement and statistics and described two criteria for setting the threshold parameters, which can be extended to other enrichment applications. Lastly, our database was evaluated by comparison to other known databases, a constructed "gold standard" of 30161 known associations, and two case studies.
Discussion
In this paper, we have demonstrated that IPAD can be used to discover, analyze, and validate pathway, disease, drug, and organ specificity from experimental data. We illustrated the features of IPAD by testing the inter-association between breast cancer markers related pathway, disease, drug and organ. In Case Study 1, we demonstrated the IPAD's ability to self-validate by using it to perform enrichment analysis and inter-association analysis on the 369 brain-specific markers. In Case Study 2, we further demonstrated the ability of IPAD to identify previously undiscovered components by the enrichment analysis based on differentially expressed genes identified from a prostate cancer study.
Selecting the appropriate statistical parameters for enrichment analysis and inter-association analysis is important. We presented a novel algorithm to measure relationships among the annotation terms based on p-value, Absolute Expression Value (AE), Relative Expression Value (RE) and Mean Jaccard Index (MJI). We also described the two criteria for setting the threshold parameters: 1) p-value below the 5% quantile and 2) 1 sigma lower control limits for AE, RE and MJI. However, defining each threshold parameter and implementing them effectively can be still challenging. Because the gene list size affects the enrichment score and the sizes of four types of component are largely different (Table 1, 11663 molecules in 1956 Pathways, 17925 molecules in 6704 diseases, 3735 molecules in 5615 drugs, and 5599 molecules in 52 organs).
In our website we provide all results for users to cut off according to the specificity of their input data. The number of enriched component sets depends on the structure of the data and the problem space. If no enriched component sets or a very large number of enriched component sets pass the thresholds, users first check whether too few or too many genes are loaded. If there are no such issues, users can tighten up the thresholds for too many significant component sets and relax them for no significant component sets.
In this paper, we introduced organ-pathway, organ-disease, organ-drug, organ-organ inter-associations for the first time. An organ actually means organ specificity in the paper. An organ is a group of tissues that perform a specific function or group of functions. Organ specificity is referred as the specificity of level of expression of a gene or protein in a certain type of organ. Identification of the association of organ-gene, organ-pathway, organ-disease, organ-drug, and organ-organ can be helpful in the discovery potentially therapeutic genes related to specific organs, measuring and understanding the function and characteristics of cells and tissues in an organ from the perspective of cooperative network, disease diagnosis, and drug target, indicating important clues about gene function, network signaling, disease treatment and drug target, and monitoring organ integrity both during preclinical toxicological assessment and clinical safety testing of investigational drugs.
Methods
Data sources
We show an overview of the data integration process in Figure 1. Pathway data in IPAD were collected from the four most commonly used sources, i.e., BioCarta [4], KEGG [5], NCI-Nature curated [6], and Reactome [7].
The BioCarta [4] includes expert-curated interactive graphic models of many pathways from diverse fields like apoptosis, cell cycle, cell signaling, development, immunology, neuroscience, adhesion, and metabolism. BioCarta data from June 2004 was imported from its website.
The KEGG [5] pathway is a collection of manually drawn pathway maps containing the knowledge on the molecular interaction and reaction networks in Metabolism, Genetic Information Processing, Environmental Information Processing, Cellular Processes, Organismal Systems, Human Diseases, and Drug Development. The KEGG data was downloaded from its ftp site.
The NCI-Nature curated [6] are created by Nature Publishing Group editors and reviewed by experts in the field. Biomolecules are annotated with UniProt protein identifiers and relevant post-translational modifications. Interactions are annotated with evidence codes and references. The NCI-Nature curated data was downloaded from its website.
Reactome [7] is an expert-authored, peer-reviewed knowledgebase of human reactions and pathways that provides infrastructure for computation and data mining across the biologic reaction network. Human pathways from Reactome were downloaded from its website.
Disease data in IPAD was downloaded from two different sources: CTD [17] and PharmGKB [18]. The Comparative Toxicogenomics Database CTD [17] is a public website and research tool that curates scientific data describing relationships between chemicals, genes, and human diseases. The Pharmacogenetics Knowledge Base (PharmGKB) [18] is curate knowledgebase about the impact of genetic variation on drug response with focus on clinical interpretation of variants associated with drug response, drug dosing guidelines and genetic tests, drug-centered pathways, important PGx gene summaries, and relationships among genes, drugs and diseases.
Drug data in IPAD were downloaded from two different sources, DrugBank [19] and PharmGKB [18]. The DrugBank database [19] is a unique bioinformatics and cheminformatics resource that combines detailed drug (i.e. chemical, pharmacological and pharmaceutical) data with comprehensive drug target (i.e. sequence, structure, and pathway) information.
The organ specificity in IPAD was downloaded from HOMER [20]. HOMER [20] is an integrated Human Organ-specific Molecular Electronic Repository, defining human organ-specific genes/proteins and covering about 22,598 proteins, 52 organs, and 4,290 diseases integrated and filtered from organ-specific proteins/genes and disease databases like dbEST [61], TiSGeD [62], HPA [63], CTD [17], and Disease Ontology [29].
We used PERL to parse the text data we downloaded and a light-weight implementation of the Document Object Model interface in Python 2.7.l [64], xml.dom.minidom to parse the XML format data.
Similarity measure for the inter-association analysis
The Jaccard Index measures similarity between pathways, diseases, drugs and organs, and is defined as the size of the intersection divided by the size of the union of the component sets. The component similarity measure can be defined as the extent of overlaps, e.g., common number of genes/proteins, shared between two different components [65]. In IPAD, we have four types of components: pathway, disease, drug and organ.
The component-component similarity score JIi,j is defined as Jaccard Index,
where, N, M denotes total number of components. Pi and Pj denote two different components, Pi and Pj can be the same or different type, while |Pi| and |Pj| are the numbers of molecules in these two components. Their intersection Pi ∩Pj is the set of all molecules that appear in both Pi and Pj, while their union Pi ∪Pj is the set of all molecules either appearing in the Pi or in the Pj. Duplicates are eliminated in the intersection set and union set.
Similarly, we define the left component-component similarity score LJIi,j as Left Jaccard Index,
the right component-component similarity score RJIi,j as Right Jaccard Index,
and the mean component-component similarity score MJIi,j as Mean Jaccard Index,
With the equations above, we can calculate similarity scores (Jaccard Index, Left Jaccard Index, Right Jaccard Index, and Mean Jaccard Index) for pathway-pathway, disease-disease, drug-drug, organ-organ, pathway-disease, pathway-drug, pathway-organ, disease-drug, disease-organ, and drug-organ associations.
Statistics for the inter-association analysis
In addition to similarity scores, we developed a statistic model based on Fisher Exact test [66,67] and number of genes involved in a component for systematic enrichment analysis. When members of two independent groups can fall into one of two mutually exclusive categories, Fisher Exact test [66,67] is used to determine whether the proportions of those falling into each category differs by group. In IPAD enrichment system, Fisher Exact test is adopted to measure the gene-enrichment in annotation terms and the enrichment between components. Given p to be the probability of success in a Bernoulli trial where one gene in component i falls in component j, the probability of x successes is
Where L is the total number of genes in component i, M is the total number of genes in component j, N is the total number of genes in the type of component, p = M/N, x is the number of genes corresponding to component i in component j, and is the number of possible combinations of x genes from a set of L genes.
The p-value for component i in component j is the probability of obtaining a test statistic at least as extreme as the one observed, given that the null hypothesis that there is no enrichment between component i and component j is true, and calculated according to the following formula
To prevent multiple testing problem from happening, IPAD adjust the p-value by Benjamini & Hochberg method [68].
The absolute enrichment value (AE) of component i in component j is defined as x, the number of genes corresponding to component i in component j. The expected enrichment value (EE) of component i in component j is defined as the expected number of genes of component i in component j under the null hypothesis that the component i and component j are independent of each other.
The relative enrichment value (RE) of component i in component j is defined as AE/EE.
We define inter-associations as enriched ones if they satisfy the thresholds in table 6 (i.e. for Pathway-Pathway association: p-value ≤ 2.13-19, RE ≥ 3.131, AE ≥ 9 and MJI ≥ 0.328; and so on). We determine the parameters based on the following two criteria: 1) Associations with p-value below the 5% quantile are chosen as enriched associations based on the p-value distribution of inter-association in the Figure 2 and the comparison of the five quantile thresholds in Table 7. 2) 1-sigma limits (1-standard error) are used to set the lower control limits for AE, RE and MJI. There are no upper control limits for AE, RE and MJI. AE, RE and MJI must be greater than or equal to one standard deviation from their means. Associations falling below the lower control limits are considered to be not stably enriched.
Table 6.
Thresholds for Inter-association Analysis in IPAD
| Typea | Typeb | p-value≤ | AE≥ | RE≥ | MJI≥ |
|---|---|---|---|---|---|
| pathway | Pathway | 2.13E-19 | 9.000 | 3.131 | 0.328 |
| Pathway | Disease | 1.28E-06 | 3.000 | 1.268 | 0.127 |
| Pathway | Disease | 5.73E-04 | 2.168 | 2.133 | 0.193 |
| Pathway | Organ | 5.00E-02 | 1.132 | 1.970 | 0.109 |
| Disease | Pathway | 1.02E-05 | 3.000 | 1.254 | 0.140 |
| Disease | Disease | 4.19E-72 | 73.538 | 1.370 | 0.393 |
| Disease | Drug | 5.00E-02 | 2.000 | 1.422 | 0.171 |
| Disease | Organ | 5.00E-02 | 1.000 | 1.313 | 0.121 |
| Drug | Pathway | 1.60E-05 | 2.666 | 2.468 | 0.141 |
| Drug | Disease | 6.51E-03 | 1.000 | 1.358 | 0.133 |
| Drug | Drug | 1.59E-05 | 3.000 | 3.391 | 0.333 |
| Drug | Organ | 5.00E-02 | 2.678 | 3.856 | 0.201 |
| Organ | Pathway | 4.01E-02 | 1.000 | 2.018 | 0.056 |
| Organ | Disease | 6.27E-03 | 2.000 | 1.384 | 0.085 |
| Organ | Drug | 5.00E-02 | 2.206 | 3.093 | 0.155 |
| Organ | Organ | 5.00E-02 | 7.000 | 4.279 | 0.095 |
Table 7.
A Comparison of the Five Quantile Thresholds
| # Associations In Pathway | #Associations In Disease | #Associations In Drug | #Associations In Organ | #total | F_measure | |
|---|---|---|---|---|---|---|
| Quantile 3% | 111374 | 594647 | 119170 | 3627 | 828818 | 60.75% |
| Quantile 4% | 148455 | 786699 | 124467 | 4471 | 1064092 | 68.58% |
| Quantile 5% | 185474 | 984366 | 130029 | 4471 | 1304340 | 68.56% |
| Quantile 6% | 222175 | 1176166 | 135915 | 4471 | 1538727 | 68.18% |
| Quantile 7% | 259592 | 1367923 | 143947 | 4471 | 1775933 | 66.92% |
P-value below the 5% quantile performs better than other p-value thresholds with a balanced F_measure and an appropriate total number of inter-associations (Table 7). First, the threshold (p-value ≤ Quantile 3%) is too strict. It filters out about half of the inter-associations that are identified by the threshold (p-value ≤ Quantile 7%). Secondly, the thresholds (p-value ≤ Quantile 6%) and (p-value ≤ Quantile 7%) cannot perform better in F_measure than the threshold (p-value ≤ Quantile 5%). Finally, we choose (p-value ≤ Quantile 5%) as the best threshold because we can identify 23% more inter-associations with (p-value ≤ Quantile 5%) than with (p-value ≤ Quantile 4%), although the F_measure of the threshold (p-value ≤ Quantile 4%) is a little bit higher than that of the threshold (p-value ≤ Quantile 5%).
Further comparison between four sigma thresholds (Table 8) shows that 1-sigma threshold to set the lower control limits for AE, RE and MJI can have the better prediction performance than other sigma thresholds.
Table 8.
A Comparison of the Four Sigma Thresholds
| # Associations In Pathway | #Associations In Disease | #Associations In Drug | #Associations In Organ | #total | F_measure | |
|---|---|---|---|---|---|---|
| 0.5 Sigma | 117535 | 644957 | 58579 | 2222 | 823293 | 60.58% |
| 1 Sigma | 185474 | 984366 | 130029 | 4471 | 1304340 | 68.56% |
| 2 Sigma | 223000 | 1215652 | 156337 | 5329 | 1600318 | 67.78% |
| 3 Sigma | 223000 | 1215652 | 156337 | 5329 | 1600318 | 67.78% |
Similarity measure and statistics for the enrichment analysis
If a user's gene list is treated as a component, then the similarity measures and the statistics for genes-pathway, genes-disease, genes-drug and genes-organ can be similarly computed with the equations in the sections: "Similarity Measure for the Inter-association Analysis" and "Statistics for the Inter-association Analysis".
Performance measurements
A "gold standard" of 30161 inter-associations (247 Pathway-Drug; 274 Drug-Drug; 23659 Pathway-Disease; 405 Organ-Disease; 2826 Drug-Disease; 2750 Disease-Disease) was constructed from KEGG [5], CTD [17], PharmGKB [18], DrugBank [19], and Disease Ontology [29] for performance evaluation purpose only. The following measurements were involved in our evaluation. (1) Sensitivity (also called recall) is the proportion of actual positive pairs which are correctly identified; (2) Specificity measures the proportion of negative pairs which are correctly identified; (3) Precision is the probability of correct positive prediction; (4) F_measure is the harmonic mean of precision and recall; (5) Accuracy is the proportion of correctly predicted pairs.
Online IPAD server design
The online version of IPAD database is a typical 3-tier web application [69], with an SQL Server2008R2 database at the backend database service layer, Apache/PHP server scripts to the middleware application web server layer, and CSS-driven web pages presented on the browser.
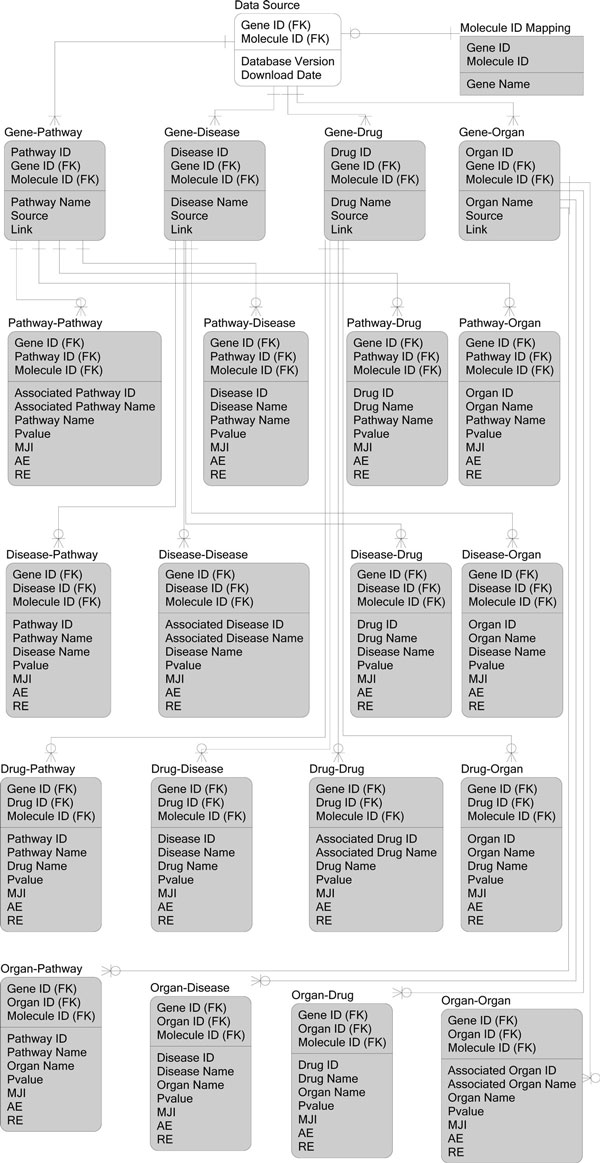
The result tables derived from the data generation steps were imported into the SQL Server2008R2 database (Figure 7). The pathway-gene, disease-gene, drug-gene, organ-gene, pathway-disease, pathway-drug, pathway-organ, disease-drug, organ-disease, organ-drug tables enable users to query the database with different IDs.
Figure 7.
Relational Metadata Model. The datasets derived by the data generation pipeline are filled in gray.
Competing interests
The authors declare that they have no competing interests.
Authors' contributions
RD conceived the initial work and designed the method for the database construction. FZ generated the datasets, developed the statistics method, the database backend and the web-based interface, and performed the statistical analyses of the case studies. All authors are involved in the drafting and revisions of the manuscript.
Contributor Information
Fan Zhang, Email: fan.zhang@unthsc.edu.
Renee Drabier, Email: Renee.Drabier@unthsc.edu.
Acknowledgements
We thank Brian Denton, Woody Hagar, Anthony Tissera, and Lynley Dungan for help with database design and web development. We also thank three anonymous reviewers for comments that helped us improve this manuscript.
This article has been published as part of BMC Bioinformatics Volume 13 Supplement 15, 2012: Proceedings of the Ninth Annual MCBIOS Conference. Dealing with the Omics Data Deluge. The full contents of the supplement are available online at http://www.biomedcentral.com/bmcbioinformatics/supplements/13/S15
References
- LOHR S. The Age of Big Data. New York Times; 2012. [Google Scholar]
- Pollack A. DNA Sequencing Caught in Deluge of Data. New York Times; 2011. [Google Scholar]
- Bader GD, Cary MP, Sander C. Pathguide: a pathway resource list. Nucleic Acids Res. 2006;34:D504–506. doi: 10.1093/nar/gkj126. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Luis CA, Abdullah L, Paris D, Quadros A, Mullan M, Mouzon B, Ait-Ghezala G, Crawford F. Serum beta-amyloid correlates with neuropsychological impairment. Neuropsychol Dev Cogn B Aging Neuropsychol Cogn. 2009;16(2):203–218. doi: 10.1080/13825580802411766. [DOI] [PubMed] [Google Scholar]
- Victor KG, Rady JM, Cross JV, Templeton DJ. Proteomic Profile of Reversible Protein Oxidation Using PROP, Purification of Reversibly Oxidized Proteins. PloS one. 2012;7(2):e32527. doi: 10.1371/journal.pone.0032527. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schaefer CF, Anthony K, Krupa S, Buchoff J, Day M, Hannay T, Buetow KH. PID: the Pathway Interaction Database. Nucleic Acids Res. 2009;37:D674–679. doi: 10.1093/nar/gkn653. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Croft D, O'Kelly G, Wu G, Haw R, Gillespie M, Matthews L, Caudy M, Garapati P, Gopinath G, Jassal B. et al. Reactome: a database of reactions, pathways and biological processes. Nucleic Acids Res. 2011;39:D691–697. doi: 10.1093/nar/gkq1018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kelder T, van Iersel MP, Hanspers K, Kutmon M, Conklin BR, Evelo CT, Pico AR. WikiPathways: building research communities on biological pathways. Nucleic Acids Res. 2012;40:D1301–1307. doi: 10.1093/nar/gkr1074. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chowbina SR, Wu X, Zhang F, Li PM, Pandey R, Kasamsetty HN, Chen JY. HPD: an online integrated human pathway database enabling systems biology studies. BMC Bioinformatics. 2009;10(Suppl 11):S5. doi: 10.1186/1471-2105-10-S11-S5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bauer-Mehren A, Furlong LI, Sanz F. Pathway databases and tools for their exploitation: benefits, current limitations and challenges. Mol Syst Biol. 2009;5:290. doi: 10.1038/msb.2009.47. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yu N, Seo J, Rho K, Jang Y, Park J, Kim WK, Lee S. hiPathDB: a human-integrated pathway database with facile visualization. Nucleic Acids Res. 2012;40:D797–802. doi: 10.1093/nar/gkr1127. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li Y, Agarwal P. A pathway-based view of human diseases and disease relationships. PloS one. 2009;4(2):e4346. doi: 10.1371/journal.pone.0004346. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smith SB, Dampier W, Tozeren A, Brown JR, Magid-Slav M. Identification of common biological pathways and drug targets across multiple respiratory viruses based on human host gene expression analysis. PloS one. 2012;7(3):e33174. doi: 10.1371/journal.pone.0033174. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oti M, Brunner HG. The modular nature of genetic diseases. Clin Genet. 2007;71(1):1–11. doi: 10.1111/j.1399-0004.2006.00708.x. [DOI] [PubMed] [Google Scholar]
- Hopkins AL. Network pharmacology. Nat Biotechnol. 2007;25(10):1110–1111. doi: 10.1038/nbt1007-1110. [DOI] [PubMed] [Google Scholar]
- Berger SI, Iyengar R. Network analyses in systems pharmacology. Bioinformatics. 2009;25(19):2466–2472. doi: 10.1093/bioinformatics/btp465. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Davis AP, King BL, Mockus S, Murphy CG, Saraceni-Richards C, Rosenstein M, Wiegers T, Mattingly CJ. The Comparative Toxicogenomics Database: update 2011. Nucleic Acids Res. 2011;39:D1067–1072. doi: 10.1093/nar/gkq813. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McDonagh EM, Whirl-Carrillo M, Garten Y, Altman RB, Klein TE. From pharmacogenomic knowledge acquisition to clinical applications: the PharmGKB as a clinical pharmacogenomic biomarker resource. Biomark Med. 2011;5(6):795–806. doi: 10.2217/bmm.11.94. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Knox C, Law V, Jewison T, Liu P, Ly S, Frolkis A, Pon A, Banco K, Mak C, Neveu V. et al. DrugBank 3.0: a comprehensive resource for 'omics' research on drugs. Nucleic Acids Res. 2011;39:D1035–1041. doi: 10.1093/nar/gkq1126. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang F, Chen JY. HOMER: a human organ-specific molecular electronic repository. BMC Bioinformatics. 2011;12(Suppl 10):S4. doi: 10.1186/1471-2105-12-S10-S4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tarca AL, Draghici S, Khatri P, Hassan SS, Mittal P, Kim JS, Kim CJ, Kusanovic JP, Romero R. A novel signaling pathway impact analysis. Bioinformatics. 2009;25(1):75–82. doi: 10.1093/bioinformatics/btn577. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, Paulovich A, Pomeroy SL, Golub TR, Lander ES. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA. 2005;102(43):15545–15550. doi: 10.1073/pnas.0506580102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Futreal PA, Coin L, Marshall M, Down T, Hubbard T, Wooster R, Rahman N, Stratton MR. A census of human cancer genes. Nat Rev Cancer. 2004;4(3):177–183. doi: 10.1038/nrc1299. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fisher B, Costantino JP, Wickerham DL, Redmond CK, Kavanah M, Cronin WM, Vogel V, Robidoux A, Dimitrov N, Atkins J. et al. Tamoxifen for prevention of breast cancer: report of the National Surgical Adjuvant Breast and Bowel Project P-1 Study. J Natl Cancer Inst. 1998;90(18):1371–1388. doi: 10.1093/jnci/90.18.1371. [DOI] [PubMed] [Google Scholar]
- Green MD, Francis PA, Gebski V, Harvey V, Karapetis C, Chan A, Snyder R, Fong A, Basser R, Forbes JF. Gefitinib treatment in hormone-resistant and hormone receptor-negative advanced breast cancer. Ann Oncol. 2009;20(11):1813–1817. doi: 10.1093/annonc/mdp202. [DOI] [PubMed] [Google Scholar]
- Moreira C, Kaklamani V. Lapatinib and breast cancer: current indications and outlook for the future. Expert Rev Anticancer Ther. 2010;10(8):1171–1182. doi: 10.1586/era.10.113. [DOI] [PubMed] [Google Scholar]
- Chen WY. Postmenopausal hormone therapy and breast cancer risk: current status and unanswered questions. Endocrinology and metabolism clinics of North America. 2011;40(3):509–518. doi: 10.1016/j.ecl.2011.05.006. viii. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martin M, Pienkowski T, Mackey J, Pawlicki M, Guastalla J-P, Weaver C, Tomiak E, Al-Tweigeri T, Chap L, Juhos E. et al. Adjuvant Docetaxel for Node-Positive Breast Cancer. New England Journal of Medicine. 2005;352(22):2302–2313. doi: 10.1056/NEJMoa043681. [DOI] [PubMed] [Google Scholar]
- Osborne JD, Flatow J, Holko M, Lin SM, Kibbe WA, Zhu LJ, Danila MI, Feng G, Chisholm RL. Annotating the human genome with Disease Ontology. BMC Genomics. 2009;10(Suppl 1):S6. doi: 10.1186/1471-2164-10-S1-S6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krzywinski M, Schein J, Birol I, Connors J, Gascoyne R, Horsman D, Jones SJ, Marra MA. Circos: an information aesthetic for comparative genomics. Genome Res. 2009;19(9):1639–1645. doi: 10.1101/gr.092759.109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baker RS, Andersen AH, Morecraft RJ, Smith CD. A functional magnetic resonance imaging study in patients with benign essential blepharospasm. J Neuroophthalmol. 2003;23(1):11–15. doi: 10.1097/00041327-200303000-00003. [DOI] [PubMed] [Google Scholar]
- Antal A, Kincses TZ, Nitsche MA, Bartfai O, Paulus W. Excitability changes induced in the human primary visual cortex by transcranial direct current stimulation: direct electrophysiological evidence. Invest Ophthalmol Vis Sci. 2004;45(2):702–707. doi: 10.1167/iovs.03-0688. [DOI] [PubMed] [Google Scholar]
- Chen J, Calhoun VD, Pearlson GD, Ehrlich S, Turner JA, Ho BC, Wassink TH, Michael AM, Liu J. Multifaceted genomic risk for brain function in schizophrenia. Neuroimage. 2012;61:866–875. doi: 10.1016/j.neuroimage.2012.03.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dauvilliers Y, Comte F, Bayard S, Carlander B, Zanca M, Touchon J. A brain PET study in patients with narcolepsy-cataplexy. J Neurol Neurosurg Psychiatry. 2010;81(3):344–348. doi: 10.1136/jnnp.2009.175786. [DOI] [PubMed] [Google Scholar]
- Hanyu H, Inoue Y, Sakurai H, Kanetaka H, Nakamura M, Miyamoto T, Sasai T, Iwamoto T. Voxel-based magnetic resonance imaging study of structural brain changes in patients with idiopathic REM sleep behavior disorder. Parkinsonism Relat Disord. 2012;18(2):136–139. doi: 10.1016/j.parkreldis.2011.08.023. [DOI] [PubMed] [Google Scholar]
- Walcott BP, Nahed BV, Kahle KT, Duhaime AC, Sharma N, Eskandar EN. Deep brain stimulation for medically refractory life-threatening status dystonicus in children. J Neurosurg Pediatr. 2012;9(1):99–102. doi: 10.3171/2011.10.PEDS11360. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sadek AR, Kirkham F, Barker S, Gray WP, Allen D. Seizure-induced miosis. Epilepsia. 2011;52(12):e199–203. doi: 10.1111/j.1528-1167.2011.03310.x. [DOI] [PubMed] [Google Scholar]
- Balduini W, Carloni S, Perrone S, Bertrando S, Tataranno ML, Negro S, Proietti F, Longini M, Buonocore G. The use of melatonin in hypoxic-ischemic brain damage: an experimental study. J Matern Fetal Neonatal Med. 2012;25(Suppl 1):119–124. doi: 10.3109/14767058.2012.663232. [DOI] [PubMed] [Google Scholar]
- Breggin PR. Psychiatric drug-induced Chronic Brain Impairment (CBI): implications for long-term treatment with psychiatric medication. Int J Risk Saf Med. 2011;23(4):193–200. doi: 10.3233/JRS-2011-0542. [DOI] [PubMed] [Google Scholar]
- Zhang H, Ozbay F, Lappalainen J, Kranzler HR, van Dyck CH, Charney DS, Price LH, Southwick S, Yang BZ, Rasmussen A. et al. Brain derived neurotrophic factor (BDNF) gene variants and Alzheimer's disease, affective disorders, posttraumatic stress disorder, schizophrenia, and substance dependence. Am J Med Genet B Neuropsychiatr Genet. 2006;141B(4):387–393. doi: 10.1002/ajmg.b.30332. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Garrett A, Kelly R, Gomez R, Keller J, Schatzberg AF, Reiss AL. Aberrant brain activation during a working memory task in psychotic major depression. Am J Psychiatry. 2011;168(2):173–182. doi: 10.1176/appi.ajp.2010.09121718. [DOI] [PubMed] [Google Scholar]
- Wilhelm H. Disorders of the pupil. Handb Clin Neurol. 2011;102:427–466. doi: 10.1016/B978-0-444-52903-9.00022-4. [DOI] [PubMed] [Google Scholar]
- Sobaniec W, Kulak W, Sobaniec H, Farbiszewski R, Drozdowski W. Effects of clobazam and vitamin E on the lipid peroxidation in the rat brain after electroconvulsive shock. Rocz Akad Med Bialymst. 1999;44:134–140. [PubMed] [Google Scholar]
- Shaw D, Norwood K, Leslie JC. Chlordiazepoxide and lavender oil alter unconditioned anxiety-induced c-fos expression in the rat brain. Behav Brain Res. 2011;224(1):1–7. doi: 10.1016/j.bbr.2011.05.014. [DOI] [PubMed] [Google Scholar]
- Tanguy M, Seguin P, Laviolle B, Bleichner JP, Morandi X, Malledant Y. Cerebral Microdialysis Effects of Propofol versus Midazolam in Severe Traumatic Brain Injury. J Neurotrauma. 2012;29:1105–1110. doi: 10.1089/neu.2011.1817. [DOI] [PubMed] [Google Scholar]
- Zhang LP, Wang L. Changes of brain neuropeptide Y and its receptors in rats with flurazepam tolerance and dependence. Acta Pharmacol Sin. 2005;26(11):1290–1296. doi: 10.1111/j.1745-7254.2005.00179.x. [DOI] [PubMed] [Google Scholar]
- van Leeuwen TH, Verbaten MN, Koelega HS, Slangen JL, van der Gugten J, Camfferman G. Effects of oxazepam on event-related brain potentials, EEG frequency bands, and vigilance performance. Psychopharmacology (Berl) 1995;122(3):244–262. doi: 10.1007/BF02246546. [DOI] [PubMed] [Google Scholar]
- Saad SF, Agha AM, Amrin Ael N. Effect of bromazepam on stress-induced gastric ulcer in rats and its relation to brain neurotransmitters. Pharmacol Res. 2001;44(6):495–501. doi: 10.1006/phrs.2001.0894. [DOI] [PubMed] [Google Scholar]
- Das S, Dey S, Datta SC, Guin AK, Sengupta D. In vitro effects of diazepam, desmethyldiazepam & nitrazepam on ATPase activity of human fetal brain. Indian J Exp Biol. 1979;17(9):895–898. [PubMed] [Google Scholar]
- Li H, Lovci MT, Kwon YS, Rosenfeld MG, Fu XD, Yeo GW. Determination of tag density required for digital transcriptome analysis: application to an androgen-sensitive prostate cancer model. Proc Natl Acad Sci USA. 2008;105(51):20179–20184. doi: 10.1073/pnas.0807121105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tannock IF, de Wit R, Berry WR, Horti J, Pluzanska A, Chi KN, Oudard S, Théodore C, James ND, Turesson I. et al. Docetaxel plus Prednisone or Mitoxantrone plus Prednisone for Advanced Prostate Cancer. New England Journal of Medicine. 2004;351(15):1502–1512. doi: 10.1056/NEJMoa040720. [DOI] [PubMed] [Google Scholar]
- Vaughn MP, Biswal Shinohara D, Castagna N, Hicks JL, Netto G, De Marzo AM, Speed TJ, Reichert ZR, Kwabi-Addo B, Henderson CJ. et al. Humanizing pi-class glutathione S-transferase regulation in a mouse model alters liver toxicity in response to acetaminophen overdose. PloS one. 2011;6(10):e25707. doi: 10.1371/journal.pone.0025707. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Festuccia C, Gravina GL, Muzi P, Biordi L, Ronchi P, Martella O, Vicentini C, Bologna M. Gefitinib and bicalutamide show synergistic effects in primary cultures of prostate cancer derived from androgen-dependent naive patients. Oncol Rep. 2007;18(5):1321–1327. [PubMed] [Google Scholar]
- Formento P, Hannoun-Levi JM, Gerard F, Mazeau C, Fischel JL, Etienne-Grimaldi MC, Gugenheim J, Milano G. Gefitinib-trastuzumab combination on hormone-refractory prostate cancer xenograft. Eur J Cancer. 2005;41(10):1467–1473. doi: 10.1016/j.ejca.2005.03.021. [DOI] [PubMed] [Google Scholar]
- Smith MR, Manola J, Kaufman DS, George D, Oh WK, Mueller E, Slovin S, Spiegelman B, Small E, Kantoff PW. Rosiglitazone versus placebo for men with prostate carcinoma and a rising serum prostate-specific antigen level after radical prostatectomy and/or radiation therapy. Cancer. 2004;101(7):1569–1574. doi: 10.1002/cncr.20493. [DOI] [PubMed] [Google Scholar]
- Han S, Roman J. Rosiglitazone suppresses human lung carcinoma cell growth through PPARgamma-dependent and PPARgamma-independent signal pathways. Mol Cancer Ther. 2006;5(2):430–437. doi: 10.1158/1535-7163.MCT-05-0347. [DOI] [PubMed] [Google Scholar]
- Oh WK, George DJ, Tay MH. Response to docetaxel/carboplatin in patients with hormone-refractory prostate cancer not responding to taxane-based chemotherapy. Clin Prostate Cancer. 2005;4(1):61–64. doi: 10.3816/cgc.2005.n.014. [DOI] [PubMed] [Google Scholar]
- Jungi WF, Bernhard J, Hurny C, Schmitz SF, Hanselmann S, Gusset H, Pestalozzi D, Goldhirsch A. Effect of carboplatin on response and palliation in hormone-refractory prostate cancer. Swiss Group for Clinical Cancer Research (SAKK) Support Care Cancer. 1998;6(5):462–468. doi: 10.1007/s005200050195. [DOI] [PubMed] [Google Scholar]
- Narita S, Tsuchiya N, Yuasa T, Maita S, Obara T, Numakura K, Tsuruta H, Saito M, Inoue T, Horikawa Y. et al. Outcome, clinical prognostic factors and genetic predictors of adverse reactions of intermittent combination chemotherapy with docetaxel, estramustine phosphate and carboplatin for castration-resistant prostate cancer. Int J Clin Oncol. 2012;17:204–211. doi: 10.1007/s10147-011-0275-6. [DOI] [PubMed] [Google Scholar]
- Jeske S, Tagawa ST, Olowokure O, Selzer J, Giannakakou P, Nanus DM. Carboplatin plus paclitaxel therapy after docetaxel in men with metastatic castrate resistant prostate cancer. Urol Oncol. 2011;29(6):676–681. doi: 10.1016/j.urolonc.2009.12.023. [DOI] [PubMed] [Google Scholar]
- Boguski MS, Tolstoshev CM, Bassett DE Jr. Gene discovery in dbEST. Science. 1994;265(5181):1993–1994. doi: 10.1126/science.8091218. [DOI] [PubMed] [Google Scholar]
- Xiao SJ, Zhang C, Zou Q, Ji ZL. TiSGeD: a database for tissue-specific genes. Bioinformatics. 2010;26(9):1273–1275. doi: 10.1093/bioinformatics/btq109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Uhlen M, Oksvold P, Fagerberg L, Lundberg E, Jonasson K, Forsberg M, Zwahlen M, Kampf C, Wester K, Hober S. et al. Towards a knowledge-based Human Protein Atlas. Nat Biotechnol. 2010;28(12):1248–1250. doi: 10.1038/nbt1210-1248. [DOI] [PubMed] [Google Scholar]
- Myers CR, Sethna JP. Python for Education: Computational Methods for Nonlinear Systems. Computing in Science and Engg. 2007;9(3):75–79. [Google Scholar]
- Wu X, Chowbina SR, Li PM, Pandey R, Kasamsetty HN, Chen JY. Characterizing Mergeability of Human Molecular Pathways. in press .
- FISHER RA. A New Test for 2 × 2 Tables. Nature. 1945;156:388. [Google Scholar]
- Huang da W, Sherman BT, Lempicki RA. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 2009;37(1):1–13. doi: 10.1093/nar/gkn923. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Benjamini Y, Hochberg Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. Journal of the Royal Statistical Society Series B (Methodological) 1995;57(1):289–300. [Google Scholar]
- Zhou A, Zhang F, Chen JY. PEPPI: a peptidomic database of human protein isoforms for proteomics experiments. BMC Bioinformatics. 2010;11(Suppl 6):S7. doi: 10.1186/1471-2105-11-S6-S7. [DOI] [PMC free article] [PubMed] [Google Scholar]