Abstract
Variation in host resistance and in the ability of pathogens to infect and grow (i.e. pathogenicity) is important as it provides the raw material for antagonistic (co)evolution, and therefore underlies risks of disease spread, disease evolution, and host shifts. Moreover, the distribution of this variation in space and time may inform us about the mode of coevolutionary selection (arms race vs. fluctuating selection dynamics) and the relative roles of GxG interactions, gene flow, selection and genetic drift in shaping coevolutionary processes. While variation in host resistance has recently been reviewed, little is known about overall patterns in the frequency and scale of variation in pathogenicity, particularly in natural systems. Using 48 studies from 30 distinct host-pathogen systems, this review demonstrates that variation in pathogenicity is ubiquitous across multiple spatial and temporal scales. Quantitative analysis of a subset of extensively studied plant-pathogen systemsshows that the magnitude of within-population variation in pathogenicity is large relative to among-population variation, and that the distribution of pathogenicity partly mirrors the distribution of host resistance. At least part of the variation in pathogenicity found at a given spatial scale is adaptive, as evidenced by studies that have examined local adaptation at scales ranging from single hosts through metapopulations to entire continents, and – to a lesser extent - by comparisons of pathogenicity with neutral genetic variation. Together these results support coevolutionary selection through fluctuating selection dynamics. We end by outlining several promising directions for future research.
Keywords: aggressiveness, coevolution, epidemiology, infectivity, local adaptation, pathogenicity, spatial scale, temporal scale, virulence
Introduction
The maintenance of diversity in host resistance and pathogenicity (i.e., the ability to infect and grow; see section Definitions) of pathogens has intrigued empiricists and theoreticians for decades (Haldane, 1949, Bergelson et al., 2001, Brown & Tellier, 2011). Importantly, genetic variation in patterns of host susceptibility and pathogen infectivity and aggressiveness are essential underlying factors influencing disease epidemiology(Wolfe, 1985, Garrett & Mundt, 1999, Thrall & Burdon, 2000, Mundt, 2002)and the emergence and spread of new diseases(Parker & Gilbert, 2004, Friesen et al., 2006, Gomez et al., 2008, Fisher et al., 2012). However, we still have little empirical data on how ecological and evolutionary processes interact toinfluence the generation and maintenance of spatial and temporal variation in natural host-pathogen interactions.
Our current understanding of the maintenance of variation in resistance and pathogenicity is largely based on theoretical predictions. Antagonistic coevolution between hosts and pathogens has been invoked as a key driver of biological diversity, but only negative frequency-dependent selection (or fluctuating selection dynamics: FSD) can stably maintain within-population genetic diversity. Diversity results as the most common host genotypes are also those most susceptible to infection, as the pathogen genotypes that specialize on those host genotypes dominate. This leads to parasites driving continual selection against common genotypes, favouring rare genotypes and promoting diversity through time (Lively, 2001). Coevolutionary dynamics can be viewed as a continuum, with escalation of defence and counter-defence (directional arms race dynamics; ARD) at one extreme, and selection for rare host and parasite genotypes (FSD) at the other extreme. Crucially, as a result of mutational limitations (Lenski, 1984) or costs associated with defence and counter-defence (Frank, 1994, Sasaki & Godfray, 1999, Sasaki, 2000), ARD are proposed to be short-lived, with coevolution either stopping or giving way to FSD. To date, the best empirical support for ARD comes from microcosm studies of bacteria-phage coevolution (Buckling & Rainey, 2002), while the few available investigations of natural associations lend support for FSD (Decaestecker et al., 2007, Gómez & Buckling, 2011, Thrall et al., 2012). Although the mode of selection is the central tenet of coevolutionary research, few systems have the biological properties or the resources available to undertake studies that actually track coevolutionary dynamics through space and time (Gaba & Ebert, 2009). Hence, assessing patterns of variation in host-parasite interactions can help us understand the relative importance of different modes of selection (FSD or ARD).
Furthermore, numerical simulations show that these coevolutionary dynamics yield very different patterns of adaptation over time (Gandon et al., 2008, Gandon & Day, 2009). Under the hypothesis that coevolutionary dynamics across multiple populations are not synchronized, the dynamics over space are very similar to the dynamics over time under FSD, leading to pathogen local adaptation (Gandon et al., 2008). Because evolution is directional under ARD, expected patterns of adaptation are less clear as the emergence of local adaptation requires some differentiation among populations. In this case, for local adaption to occur different populations need to reach adaptation through different routes (i.e. different genes involved in local adaptation; Gandon et al., 2008). Hence, detecting local adaptation according to the metric of higher pathogen performance with foreign vs. local host genotypes (Kawecki & Ebert, 2004) may be less likely under ARD than it is under FSD.
Although FSD can maintain diversity, these predictions are complicated by variation in the assumed genetic architectures underlying these key traits. On one hand, the majority of scientists working on plant-pathogen interactions have –stemming from the pioneering experimental work of Flor (1956) – largely focused on the maintenance of diversity when hosts and pathogens interact via major gene-for-gene mechanisms (Leonard, 1977, Bergelson et al., 2001). Here, many plant pathogens (including fungi, bacteria and viruses; Thompson & Burdon, 1992), insects (Hatchett & Gallun, 1970, Bangham et al., 2007), and microbial systems (Buckling & Rainey, 2002, Forde et al., 2004) have been shown to conform to the gene-for-gene (GFG) model. According to GFG, each host resistance gene produces a receptor that can recognize a particular pathogen infectivity effector. A key feature of this model is that a pathogen can become ‘universally infective’, and a cost of infectivity is usually required to maintain variation in host resistance and pathogenicity (Bergelson et al., 2001, but see e.g. Thrall & Burdon, 2002 and Damgaard, 1999). In contrast, biologists studying host-parasite interactions in animal and human populations have frequently adopted the matching-allele model (MAM), which is based on the self /non-self recognition mechanism in invertebrates (Agrawal & Lively, 2002). Notably, empirical support for the matching-allele model is only recently emerging, possibly because studies of the genetics of the interactions are hampered by difficulties in generating genetically pure parasite isolates (see Box A). In this model, resistance and infection are a result of specific matches between host and pathogen genotypes, and no ‘universally infective’ pathogen can emerge. Hence, there is no need to invoke costs of resistance and infectivity to maintain variation.
Box A. Variation in infectivity and aggressiveness in animal-pathogen systems.
Studies on spatial variation in infectivity and aggressiveness in animal-parasite systems are relatively infrequent as compared to plant-pathogen studies (see Table Box A). This may partly be due to the long-standing experimental work with purified pathogen genotypes in plant studies (Barrus, 1911, Flor, 1942), whereas studies on animal pathogens have often used inoculation material potentially consisting of a mixture of multiple pathogen genotypes. However, several recent advancements allow for a first comparison of plant-pathogen and animal-pathogen interactions.
For example, a recent study using single clones of the bacterial parasite Pasteuria ramosa in Daphnia magnarevealed strong genotype-by-genotype interactions among host and pathogen (Luijckx et al., 2011). This study then suggests that strong genotype-by-genotype interactions are a more general feature of host-pathogen interactions than previously envisioned. However, while strong genotype-by-genotype interactions may be prevalent in both animal and plant hosts, the interaction type is not necessarily the same. While plant-pathogen interactions frequently confer to the gene-for-gene interaction, the first evidence for animal-pathogen interactions for Pasteuria ramosa in Daphnia magnaindicates that a matching allele system is more likely (Luijckx et al., 2012).
Given these first characterizations of genotype-by-genotype interactions in animal pathosystems, we hope that future studies in this field will explore the spatial and temporal dimensions of variation in pathogenicity using animal pathogens. Importantly, the first evidence suggests that variation in pathogenicity in animal-pathogen systems is comparable with that of plant-pathogen systems, where variation in pathogenicity is common both within and among populations (Table Box A). Moreover, a study by Carius et al (2001) concluded – as based on high-within population variation in pathogenicity – that within-population processes are dominant. Future studies may then provide further insights of the differences and similarities of plant and animal pathosystems.
Table Box A.
A list of studies assessing the spatial scale of variation in pathogenicity in wild animal-pathogen interactions
| Reference | Species | No. pathogen populations | Spatial scale (min. and max. distance separating pathogen populations, km) | How pathogenicity is measured | At what scale variation in pathogenicity is detected |
|---|---|---|---|---|---|
| Altizer 2001 | Danaus plexippus (monarch butterfly) ~ Ophryocystis elektroscirrha (neogragarine protozoan parasite) | 3 | ca. 2000 – 3000 km | Inoculations | Between populations (within population not tested) |
| Carius et al. 2001 | Daphnia magna (planktonic crustacean) ~ Pasteuria ramosa (bacterial endoparasite) | 2 | Single population (for each of two populations) | Inoculations | Within population |
| Ebert 1994 | Daphnia magna (planktonic crustacean) ~ Glugoides intestinalis (microsporidium) | 3 | 0.5 – 1.5 km | Inoculations | Between populations (within population not tested) |
| Ebert, Zschokke-Rohringer &Carius 1998 | Daphnia magna (planktonic crustacean) ~ Pasteuria ramosa(bacterial endoparasite) | 3 | 50 – 2500 km | Inoculations | Between populations (within population not tested) |
| Imhoof & Schmid-Hempel 1998 | Bombus terrestris (bumblebee) ~ Crithidia bombi (trypanosome intestinal parasite) | 3 (local scale); 3 (regional scale) | 18 – 37 km (local scale) / ca. 80 – 200 km (regional scale) | Inoculations | Between regions and between populations |
| Oppliger, Vernet & Baez 1999 | Gallotia galloti (Canarian lizard) ~ haemogregarine genus (blood parasite) | 3 | 20 – 30 km | Inoculations | Between populations |
Despite this dichotomy in modelling approaches, theoreticians now generally agree that both theoretical frameworks can explain cyclic dynamics of allele frequencies, and thereby maintain diversity in resistance and pathogenicity (Agrawal & Lively, 2002, Dybdahl & Storfer, 2003, Laine & Tellier, 2008). The matching-allele model is prone to produce cycles (and thereby conserve polymorphism) under a wide range of parameter values. In contrast, models of gene-for-gene systems have to invoke a certain level of natural realism – like spatial population structure – in order to promote the maintenance of polymorphisms (Thrall & Burdon, 2002, Laine & Tellier, 2008, Brown & Tellier, 2011). In gene-for-gene models stabilization of polymorphisms is mainly related to factors that uncouple host and pathogen life cycles in time or space (resulting in negative direct frequency-dependent selection; Tellier & Brown, 2007), like spatial heterogeneity in selection pressure (e.g. due to spatial variation in disease severity, or in the cost of resistance and pathogenicity; Laine & Tellier, 2008, Wolinska & King, 2009, Mostowy & Engelstädter, 2011, Tellier & Brown, 2011). Factors that promote the persistence of dynamic polymorphisms in time (rather than creating stable equilibria) are linked to genetic complexity (e.g. the number of genes involved) and, again, spatial structuring of host and pathogen populations.
In addition to local selection, there is potential for various nonselective processes to influence the distribution of variation among populations. Isolation by distance processes promote asynchrony among populations, and hence, localized dispersal allows the maintenance of spatial variation in genotype frequencies through the emergence of fixed or moving spatial patterns(Gandon, 2002, Sasaki et al., 2002). A predominance of neutral variation is likely to occur when selection is weak relative to mutation, genetic drift and the homogenizing effect of gene flow (Slatkin, 1987, Hedrick, 2000). As the balance between genetic drift, mutation, selection and gene flow varies with spatial scale, the degree of pathogen adaptation may likewise vary across spatial scales. Importantly, as the majority of wild plants, animals and pathogens are patchily distributed across landscapes (e.g. Burdon, 1993, Hanski, 1999), spatial structure is likely to play a critical role in maintaining polymorphism in host resistance and pathogenicity in a natural setting (Frank, 1991, Thrall & Antonovics, 1995, Sasaki, 2000, Sasaki et al., 2002, Thrall & Burdon, 2002, Thrall et al., 2002, Gavrilets & Michalakis, 2008, Laine & Tellier, 2008, Tellier & Brown, 2011). Characterization of spatial and temporal patterns in the distribution of pathogenicity and host resistance can hence be used to dissect the mode of coevolutionary selection, relative importance of gene flow, mutation and genetic drift, and provide an empirical base, and future directions, for theoretical studies. While the spatial distribution of variation in host resistance has been recently reviewed (Salvaudon et al., 2008, Laine et al., 2010), there has been no consolidated effort to review and quantify levels of variation in pathogenicity present in natural pathogen populations, and how such variation is distributed in either space or time (but see Stukenbrock & McDonald, 2008 for several agricultural examples).
Here we characterize patterns of variation in pathogenicity in wild host-pathogen associations across a range of spatial scales: within individual hosts, within host populations, among host populations (i.e. metapopulations), and among broader geographic regions. We also examine at what spatial scale(s) pathogenicity varies through time. More specifically: 1) Given the difficulties of maintaining polymorphism in pathogenicity in gene-for-gene models under empirically realistic parameter values (Bergelson et al., 2001, Brown & Tellier, 2011), we first evaluate the frequency of pathogenic variation at multiple spatial scales across 45 published studies on wild host-pathogen interactions. Importantly, variation in pathogenicity across spatial and temporal scales in a large fraction of study systems will indicate that factors that increase the maintenance of pathogenicity, such as spatial structure and environmental heterogeneity (Sasaki et al., 2002, Brown & Tellier, 2011, Tellier & Brown, 2011), are crucial for our understanding of coevolutionary dynamics; 2) We then quantify the relative amount of variation in pathogenicity and host resistance present at spatial and temporal scales across seven wild host-pathogen systems. Using these patterns, we evaluate a) whether the observed patterns of diversity yield support for ARD or FSD mode of coevolutionary selection, whereby high levels of within population diversity would suggest FSD and low levels ARD, b) whether spatial, temporal and spatiotemporal variation are important in safeguarding variation in pathogenicity, as suggested by a significant fraction of variation and level of asynchrony in pathogenicity among populations, and c) whether variation in host diversity and pathogen diversity are interrelated, as would be expected based on local coevolutionary interactions (Dybdahl & Storfer, 2003); 3) Finally, to understand whether variation in pathogenicity is due to selectively neutral or adaptive forces (Slatkin, 1987, Thrall & Burdon, 2002), we explore patterns of pathogen local adaptation, and differences between neutral and pathogenic variation (the latter is discussed in more detail in the section Molecular and genetic variation).
Literature search
We focused our literature review search efforts on microbial pathogens (thereby excluding animal macro-parasites such as nematodes and helminths). For this purpose, we searched the Web of Science for relevant studies using combinations of search terms ‘plant’, ‘animal’, ‘pathogen’, ‘virulence’, ‘infectivity’, ‘pathogenicity’, ‘aggressiveness’, ‘local adaptation’, ‘resistance’, ‘spatial scale’ or ‘spatial structure’. From each paper we scanned references to other relevant studies, and we performed forward searches on a number of key papers. This search resulted in 48 studies covering plant (n=42) and animal (n=6) pathogens. Given the paucity of information on the spatial distribution of pathogenicity for animal pathogens, we focus in the main text on plant pathogens (see Box A for a comparison with animal pathogens).
Definitions
The vocabulary and definitions used in plant pathology (Vanderplank, 1968, Agrios, 2005) and animal pathology and evolutionary ecology (e.g. Read, 1994) are at least partly contradictory (cf. Sacristán & García-Arenal, 2008). To prevent confusion, we explicitly spell out our definitions. We define infectivity as the ability of the pathogen to overcome host resistance and infect a given host individual (often referred to as ‘virulence’ in plant pathology); aggressiveness as the extent of within-host growth; and virulence as the extent of damage to the host. As we are interested in spatial and temporal variation in both infectivity and aggressiveness, we use the term pathogenicity in those instances where we want to encompass both the qualitative (infectivity) and quantitative (aggressiveness) aspects of pathogen fitness.
How is variation in pathogenicity measured?
Reciprocal transplant experiments
These are characterised by planting a range of host genotypes from a number of different locations into all those locations (i.e. all sympatric and allopatric combinations), recognizing that each location is likely to be characterized by a distinct pathogen community. Interpretation of the results of reciprocal transplant experiments can be confounded by among-location environmental differences in addition to variation in the structure of the pathogen population. However, an advantage of field transplant experiments is that estimates of pathogenicity reflect both natural encounter rates and infection outcomes between specific host and pathogen genotypes (Burdon, 1987, Nuismer & Gandon, 2008). Hence, experimental results may be more likely to reflect host-pathogen dynamics that play out in the field compared to results obtained from controlled inoculation experiments (see below).
Cross-inoculations
In many plant pathogen studies, a set of host lines known to differ in resistance phenotype (a ‘differential set’) is inoculated separately with a series of different pathogen isolates in a common environment. As the host lines are specifically selected for their differential response to individual pathogen isolates, such an approach is likely to detect a large number of pathotypes with different infectivity patterns. However, as the host lines in the differential set are frequently collected from a much broader geographical area than the pathogen, the host-pathogen interaction may at least partly be taken out of its ecological and evolutionary context, and the variation observed may be functionally irrelevant or hard to interpret. For example, Antonovics et al.2011 showed that the interaction outcome (resistance, partial resistance, susceptible) of hosts inoculated with local (sympatric) pathogens differs from those inoculated with allopatric pathogens, with a larger fraction of hosts showing partial resistance to local pathogens.
Unfortunately, results from common garden and cross-inoculation approaches can be hard to reconcile. As such, a combination of laboratory inoculation experiments and common garden field experiments may be useful in attempting to disentangle the contribution of encounter rate and encounter outcome in natural settings. Using these two approaches, Alexander and colleagues (1989, 1993) showed that infection of Silene alba by Microbotryum violaceum is dependent on both floral phenology (encounter frequency) and genotype-dependent interactions (encounter outcome). Likewise, other studies involving the same host-pathogen interaction have shown that plant height and floral abundance (Alexander et al., 1984, Thrall & Jarosz, 1994) correlate with the likelihood of infection. It is likely that less studied features, such as intraspecific variation in pathogen traits (e.g. the timing of sporulation), may also affect the frequency of interactions between specific host and pathogen genotypes.
Another challenge in these studies is that the pathogen may simultaneously evolve in response to the host population, the environment, and possibly the interaction between host genotype and the environment (Laine, 2008, Wolinska & King, 2009). In such cases, measuring local adaptation in a common environment (like the laboratory or greenhouse) may not reflect adaptation measured in the field (Laine, 2007, Ridenhour & Nuismer, 2007).
Molecular and genetic variation
The genetic structure of many pathogen populations has been elucidated by targeting underlying neutral genetic variation. In addition to using neutral genetic markers, several recent studies have also targeted genetic variation in genes involved in the infection process (‘resistance’ and ‘avirulence’ genes; e.g. Barrett et al., 2009b). In combination with phenotypic resistance and infectivity data, such molecular data can provide powerful insight into the forces structuring both host and pathogen populations. For example, the spatial structure of neutral genetic variation – as compared to variation in phenotypes or functional genes – can give insight into the strength and sign of selective forces (Merilä & Crnokrak, 2001, Jorgensen et al., 2006, Barrett et al., 2008) or into the mating system of the pathogen (Barrett et al., 2008). Moreover, analysis of patterns of variation in coding sequences (e.g. the ratio of synonymous vs. non-synonymous mutations) underlying variation in resistance or pathogenicity can provide powerful insight into long term patterns of evolution in populations [e.g. the importance of genetic arms races vs. balancing selection (Barrett et al., 2009b)].
The spatial scale of variation in pathogenicity
Strikingly, our review of the literature (42 studies involving 25 distinct plant-pathogen associations) unambiguously demonstrates the existence of variation in pathogenicity in virtually all study systems, and at spatial scales ranging from single host individuals to entire regions spanning hundreds of kilometres (Table 1; see Box A for a comparison with animal-pathogen studies). Variation in infectivity among pathogen isolates was omnipresent, where each study system contained multiple pathogen strains that varied in their ability to infect different host plant genotypes.
Table 1.
A list of studies assessing the spatial scale of variation in pathogenicity in wild plant-pathogen interactions
| Reference | Species | No. pathogen populations | Spatial scale (min. and max. distance separating pathogen populations, km) | How pathogenicity is measured | At what scale variation in pathogenicity is detected |
|---|---|---|---|---|---|
| Alexander 1989 | Silene latifolia (white campion) ~ Microbotryum violaceum (anther-smut fungus) | 1 | Single population | Inoculations | Within population |
| Alexander, Antonovics & Kelly 1993 | Silene latifolia (white campion) ~ Microbotryum violaceum (anther-smut fungus) | 1 | Single population | Inoculations | No variation within population |
| Barrett et al. 2009b | Linum marginale (wild flax) ~ Melampsora lini (rust fungus) | 10 | 0.5 – 320 km | Avr genes | Within and between populations |
| Bevan, Crute & Clarke 1993b | Senecio vulgaris (groundsel) ~ Erysiphefischeri (powdery mildew fungus) | 2 | 480 km | Inoculations | Within and between populations |
| Burdon & Jarosz 1991 | Linum marginale (wild flax) ~ Melampsora lini (rust fungus) | 1 | Single population | Inoculations | Within population |
| Burdon & Jarosz 1992 | Linum marginale (wild flax) ~ Melampsora lini (rust fungus) | 9 | 0.3 – 110 km | Inoculations | Within and between populations |
| Burdon, Thrall & Brown 1999 | Linum marginale (wild flax) ~ Melampsora lini (rust fungus) | 4 | 0.2 – 265 km | Inoculations | Within and between populations and between metapopulations |
| Burdon, Thrall & Lawrence 2002 | Linum marginale (wild flax) ~ Melampsora lini (rust fungus) | 37 | 4 – 3310 km | Inoculations | Continental |
| Capelle & Neema, 2005 | Phaseolus vulgaris (common bean) ~ Colletotrichum lindemuthianum (fungal pathogen) | 1 | Single population (3 – 500 m) | Inoculations | Within host and Within population |
| Carlsson-Granér 1997 | Silene dioica (red campion) – Microbotryum violaceum (anther-smut fungus) | 3 | < 10 km | Common gardens | Between populations (within population not tested) |
| Carlsson-Granér, Burdon and Thrall 1999 | Linum marginale (wild flax) ~ Melampsora lini (rust fungus) | 3 | 0.1 km | Inoculations | Within and between populations |
| Davelos, Alexander & Slade 1996 | Spartina pectinata ~ Puccinia seymouriana and Puccinia sparganioides (rust fungi) | 3 | 0.2 – 4.5 km | Inoculations and common gardens | Between populations (within population not tested) |
| De Nooij & van Damme 1988b | Plantago lanceolata (ribwort plantain) ~ Phomopsis subordinaria (fungal pathogen) | 3 | 40 – 110 km | Inoculations | Within hosts, within and between populations |
| Ennos & McConnell 1995 | Pinus sylvestris (Scots pine) ~ Crumenulopsis sororia (fungal canker) | 3 | 60 –120 km | Inoculations | Between populations (within population not tested) |
| Ericson, Burdon & Müller 2002 | Filipendula ulmaria (meadowsweet) ~ Triphragmium ulmariae (rust fungus) | 4 | 3 –350 km | Inoculations | Between populations (within population not tested) |
| Espiau et al. 1998 | Chondrilla juncea (skeletonweed) ~ Puccinia chondrillina (rust fungus) | 1 | Single population | Inoculations | Within population |
| French & Manion 1975 | Populus tremuloides (quaking aspen) ~ Hypoxylon mammatum (fungal canker) | 1 | Single population | Inoculations | Within population |
| Geffroy et al. 1999 | Phaseolus vulgaris (common bean) ~ Colletotrichum lindemuthianum (fungal pathogen) | 3 | ca. 3000 – 6500 km | Inoculations | Continental |
| Gerard et al. 2006 | Populus nigra (black poplar) ~ Melampsora larici-populina (rust fungus) | 5 | 8 – 93 km | Inoculations | Within and between populations |
| Goss & Bergelson, 2006 | Arabidopsis thaliana ~ Pseudomonas viridiflava (foliar bacterial pathogen) | 5 | 8 – 95 km | Inoculations | Within and between populations |
| Jarosz & Burdon 1991 | Linum marginale (wild flax) ~ Melampsora lini (rust fungus) | 10 | 0.3 – 100 km | Inoculations | Within and between populations |
| Kaltz & Shykoff 2002 | Silene latifolia (white campion) ~ Microbotryum violaceum (anther-smut fungus) | 4 | 17 – 114 km | Inoculations | Within and between populations |
| Kaltz et al 1999 | Silene latifolia (white campion) ~ Microbotryum violaceum (anther-smut fungus) | 14 | 2 – 166 km | Inoculations | Within and between populations |
| Kinloch & Stonecypher 1969 | Pinus taeda (loblolly pine) ~ Cronartium fusiforme (fusiform rust) | 6 | 0 – 3.6 km | Common gardens | Between populations (within population not tested) |
| Laine 2005 | Plantago lanceolata (ribwort plantain) ~ Podosphaera plantaginis (fungal pathogen) | 4 | 13 – 43 km | Inoculations | Between populations |
| Lebeda and Petrželová 2004 | Lactuca serriola (prickly lettuce) ~ Bremia lactucae (lettuce downy mildew) | 78 | ca. 0.1 – 300 km | Inoculations | Within and between populations |
| Lenné & Burdon 1990 | Stylosanthes guianensis (common stylo) ~ Colletotrichum gloeosporioides (anthracnose) | 5 | Single population (for each of five populations) | Inoculations | Within population |
| Meyer, Nelson & Clement 2001 | Bromus tectorum (cheatgrass) ~ Ustilago bullata (head smut fungus) | 4 | ca. 40 – 600 km | Inoculations | Within and between populations |
| Meyer et al. 2005 | Bromus tectorum (cheatgrass) ~ Ustilago bullata (head smut fungus) | 4 | ca. 40 – 600 km | Inoculations | Within and between populations |
| Miles & Lenné 1984 | Stylosanthes guianensis (common stylo) ~ Colletotrichum gloeosporioides (anthracnose) | 1 | Single population | Inoculations | Within population |
| Niemi et al. 2006 | Salix triandra (almond willow) – Melampsora amygdalinae (rust fungus) | 4 | 355 – 907 km | Inoculations | Within and between populations |
| Oates, Burdon & Brouwer 1983 | Avena barbata, A. fatua and A. ludoviciana ~ P. coronata and P. graminis | 248 | Across the state of New South Wales, Australia | Inoculations | Regional |
| Parker 1988 | Amphicarpaea bracteata (hog peanut) ~ Synchytrium decipiens (fungal pathogen) | 1 | Single population | Inoculations | Within population |
| Parker 1989 | Podophyllum peltatum (mayapple) ~ Puccinia podophylli (rust fungus) | 6 | ca. 0.7 – 48 km | Reciprocal common gardens | Between populations |
| Roslin, Laine & Gripenberg 2007 | Quercus robur (pedunculate oak) ~ Erysiphe alphitoides (oak powdery mildew) | 4 | 0.3 – 1.8 km | Inoculations | Within population |
| Roy 1998 | Arabis holboellii (rockcress) ~ Puccinia monoica and P. thlaspeos (rust fungi) | 3 | ca. 25 – 40 km | Reciprocal common gardens | Between populations |
| Sicard et al. 1997a | Phaseolus vulgaris (common bean) ~ Colletotrichum lindemuthianum (fungal pathogen) | 5 | 40 – 385 km | Inoculations & ITS / RAPD | Within and between populations |
| Sicard et al. 1997b | Phaseolus vulgaris (common bean) ~ Colletotrichum lindemuthianum (fungal pathogen) | 3 | ca. 3000 – 6500 km | Molecular (RAPD; ITS) and inoculations | Continental |
| Sicard et al. 2007 | Phaseolus vulgaris (common bean) and P. coccineus ~ Colletotrichum lindemuthianum (fungal pathogen) | 5 | 5 – 25 km | Inoculations | Within and between populations |
| Springer 2007 | Hesperolinon californicum (California dwarf flax) ~ Melampsora lini (rust fungus) | 10 | 1 – 80 km | Inoculations | Regional cline |
| Thrall, Burdon & Bever 2002 | Linum marginale (wild flax) ~ Melampsora lini (rust fungus) | 6 | 0.2 – 10 km | Inoculations | Within and between populations |
| Thrall, Burdon & Young 2001 | Linum marginale (wild flax) ~ Melampsora lini (rust fungus) | 16 | 0.1 – 10 km | Inoculations | Within and between populations |
Variation in pathogenicity within hosts
The issue of multiple infections (i.e. superinfection) within single host individuals has been a topic of considerable interest for studies of human and animal-parasite interactions (Anderson et al., 1995, Torres, 1996, Al-Yaman et al., 1997, Butto et al., 1997, Huo et al., 1997, Ebert, 1998, Read & Taylor, 2001, Theron et al., 2004, Pisoni et al., 2007), in part because of the implications for how within-host competition among pathogen strains might influence the evolution of virulence (Lipsitch & Moxon, 1997). While this issue has only recently gained widespread interest among plant pathologists, the few studies to date suggest that co-infection is also common in natural plant pathosystems (de Nooij & van Damme, 1988b, Wille et al., 2002, Hood, 2003, Capelle & Neema, 2005, Ganz & Washburn, 2006, López-Villavicencio et al., 2007). Likewise, studies in agricultural systems frequently show a high rate of co-infection (Keller et al., 1997, McDonald et al., 1999, Linde et al., 2002). While co-infection is hardly surprising, it has potentially pronounced consequences for host-parasite dynamics, with predicted effects on the evolution of virulence and transmission ability (e.g. Bull, 1994, Nowak & May, 1994, Frank, 1996, Brown et al., 2002, de Roode et al., 2005, Alizon & Lion, 2011, Laine, 2011).
Importantly, the presence of different strains infecting a single host increases the likelihood for the evolution and emergence of new variation in pathogenicity. For some plant pathogens, co-infection is the prerequisite of sexual reproduction, and even among asexual pathogens co-infection promotes exchange of genetic information (e.g. by somatic exchange of genetic information; Burdon et al., 1982). Furthermore, theory predicts that the existence of multiple infections may facilitate the maintenance of polymorphism in pathogenicity within local populations via competitive interactions (Levin & Pimentel, 1981, Anderson & May, 1982, Bonhoeffer & Nowak, 1994, Nowak & May, 1994, Gandon et al., 2002). For example, Wille et al. (2002) demonstrated that the outcome of competition among endophyte genotypes can vary depending on host plant genotype, thereby demonstrating a mechanism by which host plant genetic diversity can maintain pathogen diversity. Likewise, coexistence of pathogens could be mediated by a trade-off between within-host competitive ability and other traits like dispersal ability (Leibold & Miller, 2004) and off-host survival (Abang et al., 2006, Sommerhalder et al., 2011), or based on the classic hypothesis that more virulent pathogens have less potential for dispersal due to a shorter life-time of the host, but are better within-host competitors (Anderson & May, 1982). Moreover, ‘cheater’ genotypes may emerge that perform better in mixed infections than in single infections (Barrett et al., 2011). Finally, genetic diversity of the pathogen population may also influence the frequency at which co-infections occur. López-Villavicencio et al. (2007) observed that high genetic diversity within pathogen populations may result in less co-infections, seemingly due to higher within-host competitive exclusion among unrelated strains.
Variation in pathogenicity within host populations
Table 1 demonstrates that variation in pathogenicity within populations is widespread (97% or 28 out of 29 of the studies). As the interaction between host and pathogen is frequently genotype-specific (i.e. the interaction outcome depends on both host and pathogen genotype), this strongly supports frequency-dependent maintenance of resistance and pathogenic polymorphisms within local populations. Here, the well-established theory of negative frequency-dependent selection states that rare alleles are advantageous, and that fitness declines when the frequency of the allele increases (Haldane, 1949, Jayakar, 1970, Leonard, 1997, Tellier & Brown, 2007).
While the role of negative frequency-dependent selection in maintaining diversity within pathogen populations has proven difficult to quantify unequivocally, there are a range of studies that show patterns consistent with such dynamics. For example, Chaboudez and Burdon (1995) provide convincing evidence for negative frequency-dependent dynamics, by demonstrating that locally common clones of Chondrilla juncea occurring in their home range in Turkey were more likely to be infected by the rust fungus Puccinia chondrillina. In animal host-pathogen systems, a classic example from a natural system involves a longitudinal study of macroparasitic Microphallus trematodes preferentially infecting locally common snail genotypes of the species Potamopyrgus antipodarum (Dybdahl & Lively, 1998, Lively & Dybdahl, 2000). More recently, a 6-year study of the interaction between native flax (Linum marginale) and its associated rust (Melampsora lini) strongly supports a role for reciprocal coevolution in maintaining resistance and infectivity polymorphisms via negative frequency dependence, as well as a role for trade-offs between infectivity and aggressiveness(Thrall et al., 2012).
Nevertheless, convincing demonstrations of negative frequency-dependent selection still remain rare in wild host-pathogen interactions (Barrett, 1988, Roy, 1998, Decaestecker et al., 2007, Meyer et al., 2010). While this lack of evidence may be due to the absence or weakness of negative frequency-dependent selection in wild host-pathogen interactions, it is equally if not more likely, that this reflects a shortage of long-term studies in natural systems (Gandon et al., 2008). Moreover, it is possible that the same system may switch between ARD and FSD, as ARD is considered difficult to maintain over time due to fitness costs and mutation limitations (Buckling & Rainey, 2002, Gómez & Buckling, 2011). Furthermore, when multiple loci are involved in the interaction, some loci may evolve according to FSD while others follow ARD (Gandon et al., 2008). Broad-scale spatio-temporal data of natural host-pathogen interactions is required given that the effects of gene flow, genetic drift and mutation, and local extinction-recolonisation processes can potentially obscure evolutionary patterns. These patterns may be further complicated given the potential for the strength and direction of selection to vary through space and time (Thompson, 2005, Smith et al., 2011). Hence, an explicit consideration of the spatial structure of multiple interacting populations will be paramount in developing a quantitative understanding of the processes that drive the evolution and maintenance of diversity in pathogenicity and resistance in host-pathogen interactions (Thrall et al., 2012).
Variation in pathogenicity among host populations
Variation in pathogenicity among populations is as pervasive as it is within pathogen populations (100%, or 27 out of 27 studies; Table 1). As most natural host-pathogen associations (including those listed in Table 1) are characterized by migration and gene flow among neighbouring populations within a metapopulation context (e.g. Jarosz & Burdon, 1991, Laine, 2005), assessing the role of among-population processes is critical for understanding the coevolutionary dynamics of local host-pathogen interactions. For example, given substantial variation in pathogenicity among populations, local coevolutionary outcomes will not only depend on fine-scale processes generating genetic variation (e.g. mutation, sexual recombination), but will be simultaneously fuelled by the arrival of potentially novel pathogenicity and resistance genes from neighbouring populations (Thrall & Burdon, 1997). A focus on a single population will clearly leave the researcher unable to trace the origin of new variation (e.g. whether due to local mutation or immigration); however, even when immigrant status can be attached to a new pathotype its spatial origin (near or far) may still be undeterminable. For example, in the Australian flax – flax rust system, a 12-year annual survey of M. lini pathogenicity in the Kiandra population detected the appearance of a novel pathotype. Over a 3-year period the frequency of this pathotype increased from <1% to >28% before suddenly disappearing entirely. While its novelty in the Kiandra population was confirmed by its unique isozyme signature its spatial origin was never identified (J.J. Burdon, unpublished data). While examples from natural systems are few, studies of agricultural pathogens provide clear-cut examples of new mutations arising in response to new resistant host plant varieties, and the subsequent spread of such pathogens across large spatial scales (McDonald & Linde, 2002).
As discussed below, variation in pathogenicity among locations may be either due to neutral (e.g. due to mutation, genetic drift) or adaptive (where selection varies among populations) processes. We note that such spatially heterogeneous selection may be due to biotic (i.e. adaptation to spatially varying host plant genotypes or other members of the community) or abiotic factors. Notably, when the host evolves faster than the pathogen, variation in pathogenicity may be maladaptive (Gandon et al., 1996, Kaltz & Shykoff, 1998). Moreover, immigration may result in the inflow of maladaptive genes into the local population.
Variation in pathogenicity among metapopulations (and larger spatial scales)
Given that variation among pathogen populations within meta populations is omnipresent, it is not surprising that variation in pathogenicity is also universal at larger spatial scales (ranging from among geographic regions to intercontinental scales;100%, or 6 out of 6 studies; Table 1). For example, Burdon and colleagues have shown that pathogenic variation in wild flax rust exists across the entire Australian continent, as well as among environmentally distinct plains and mountain regions in southeastern Australia (Burdon et al., 1999, Burdon et al., 2002). Likewise, pathogenicity of the fungal pathogen Colletotrichum lindemuthianum varies among three geographic areas within the range of the common bean Phaseolus vulgaris in Latin America (Sicard et al., 1997b). These studies support the idea that coevolutionary interactions may vary across broad geographical areas (Thompson, 2005). In addition, the work of Barrett et al. (2008) further suggests that environmental and climatic differences between regions are likely to play a significant role with regard to maintaining variation in host and pathogen life history traits that can markedly influence levels of diversity.
Spatial and temporal structure of variation in pathogenicity and resistance – a quantitative analysis
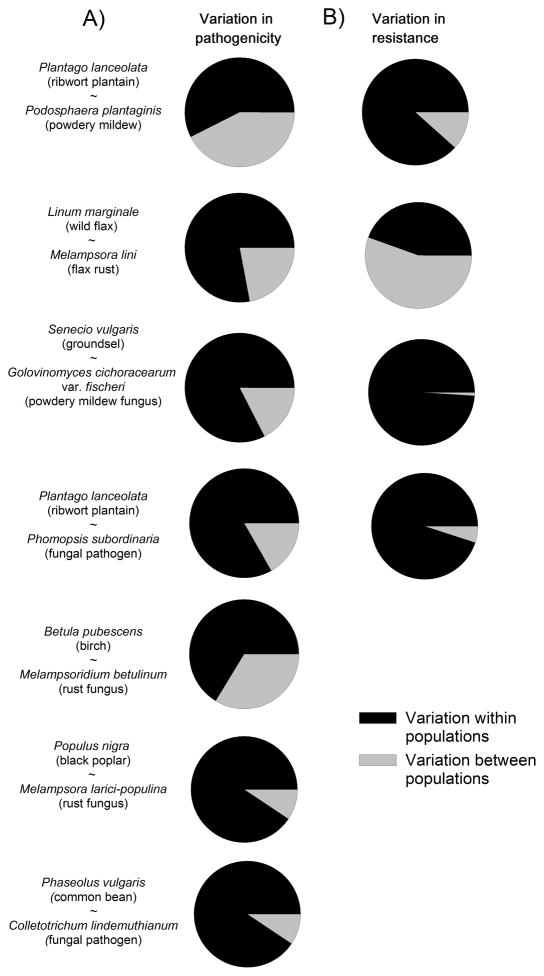
While variation in pathogenicity exists both within and among populations (Table 1), a quantitative comparison of seven wild pathosystems that have been studied in detail reveals that the majority of pathogenic variation can be attributed to variation within populations (Fig. 1A). Low levels of between population variation relative to strong genetic variation within populations suggest that local interactions are dominant. Traits under strong frequency dependent selection are expected to show such patterns according to models by Schierup et al.(2000a, 2000b). This pattern may further be shaped by the interplay between balancing selection, locally variable selection generated by G × (G ×) E -interactions, replenishment of variation by frequent gene flow, and possibly uniform selection across the landscape. As a consequence, genetic drift may only infrequently result in the loss of pathogenic variation within populations. Disentangling the relative importance of these factors is likely to be crucial in understanding the spatial distribution and maintenance of pathogenic polymorphisms.
Figure 1.
The relative amount of variation in A) pathogenicity and B) resistance within and among populations for seven pathosystems. Multivariate analyses were conducted using pathogenicity and resistance (either binary or quantitative) as the response variable and the fixed variable Population as the explanatory variable (note that the residual error term reflects the within-population variation). Analyses were implemented with the function adonis in package vegan (version 1.17-6) in R (Oksanen et al., 2010), a method related to the AMOVA procedure implemented by Excoffier (1992). Data sources (and for more detailed information): Plantago lanceolata – Podosphaera plantaginis from Laine (2005); Linum marginale ~ Melampsora linifrom Barrett et al (2009b and unpublished data; data shown are from the ‘Plains’ metapopulation); Senecio vulgaris ~ Golovinomyces cichoracearum from Bevan et al.(1993a, 1993b); Plantago lanceolata ~ Phomopsis subordinaria from de Nooij and van Damme(1988a, 1988b); Betula pubescens ~ Melampsoridium betulinum from Ericson and Burdon (2009); Populus nigra ~ Melampsora larici-populina from the AMOVA table given in Gérard et al (2006); Phaseolus vulgaris ~ Colletotrichum lindemuthianum from the AMOVA table given in Sicard et al (1997a).
Unfortunately, the role of gene flow in homogenizing populations is hard to evaluate given the lack of data across a wide range of pathosystems: while populations were generally separated by a few to hundreds of kilometres, each of the pathogens we were able to evaluate disperses aerially, likely resulting in frequent long-distance dispersal. Broadening our ability to conduct comparative studies including other disease transmission modes such as vector-dispersed (e.g. insect-transmitted floral smuts) or soil-borne pathogens will be essential if we are to further our understanding of the role of gene flow and other aspects of life-history in determining the spatial scale of variation in pathogenicity.
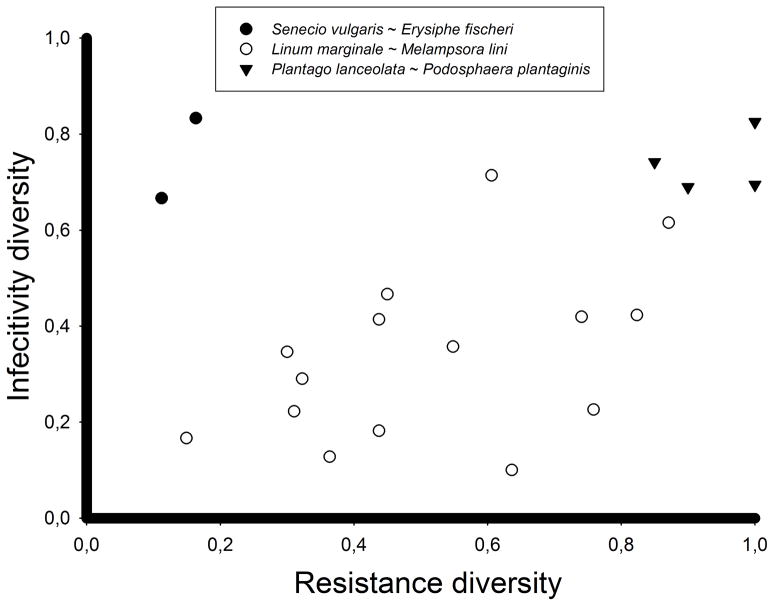
Variation in host resistance – and its distribution within and among host populations - is likely to impact heavily on the distribution of variation in pathogenicity (and vice versa). For example, if patches consist of single clones, local selection is likely to be directional and may result in low within-patch pathogenic diversity. On the other hand, large variation in host resistance within populations may select for a high number of associated pathogen genotypes within host populations. Indeed, Figure 1 shows that the distribution of host resistance shows a pattern roughly comparable to the distribution in pathogenicity, consistent with the idea that host resistance diversity is the driving force behind variation in pathogenicity. Moreover, within three natural host-pathogen study systems, there is a positive relationship between within-population diversity in infectivity and diversity in host resistance (Fig. 2). A related pattern emerges in the Australian flax – flax rust and the almond willow – rust systems, wherewithin-population mean infectivity and mean resistance are highly correlated(Thrall & Burdon, 2003, Niemi et al., 2006, but see Springer, 2007). Likewise, agricultural studies have shown a clear-cut relationship between host plant resistance and variation in pathogenicity. For example, the diversity and identity of host plant genotypes in agricultural fields may affect infection intensity, pathogen diversity, and the genotypic composition of the pathogen population (Wolfe, 1985, Zhu et al., 2000, Mundt, 2002, Goyeau et al., 2012).
Figure 2.
Within-population diversity in pathogen infectivity and host resistance diversity are positively associated for three study systems (ANCOVA; F1,17= 4.37; P= 0.04, R2=0.09). Additional variation is explained by differences among study systems in the mean diversity in infectivity (F2,17= 14.36; P= 0.002, R2=0.57).
Nevertheless, the results show some level of discrepancy between the distribution of resistance and pathogenicity: for example, the host plant L. marginale shows strong variation in resistance among local populations, whereas its associated rust M. lini is less differentiated among locations (Fig 1; Thrall et al., 2001). This pattern may be explained by the large dispersal range of the pathogen as compared to the host plant (Thrall & Burdon, 1999), and is mirrored by evidence for stronger pathogen adaptation at regional as opposed to local spatial scales (Thrall et al., 2002). In striking contrast, for the ribwort plantain Plantago lanceolata(de Nooij & van Damme, 1988b, de Nooij & van Damme, 1988a) and the groundsel Senecio vulgaris(Bevan et al., 1993a, Bevan et al., 1993b) systems, the majority of variation in host resistance is present within populations, whereas their pathogens vary considerably more among locations (Fig. 1). While these first quantitative studies of the spatial structure of variation in pathogenicity and host resistance provide novel insights, there is a clear need for studies of many more wild systems to generalize these patterns.
The temporal scale of variation in pathogenicity
For several decades host-pathogen models have emphasized that variation in pathogenicity and host resistance can be maintained by (frequency-dependent) changes in allele/gene-frequencies through time (Brown & Tellier, 2011). However, most empirical studies have replicated sampling across space rather than through time, based on the argument that assessment across multiple populations in space provides a reasonable surrogate for variation through time (Burdon et al., 1990). Nevertheless, recent work suggests this may not always be the case. For example, Gomez & Buckling (2011) showed that while phages perform better on local than foreign bacterial hosts, it is the bacteria that are best adapted to the contemporaneous as compared to past phages. In a similar vein, the bacterial endoparasite Pasteuria ramosa shows no spatial adaptation (Ebert et al., 1998), but it is temporally adapted to its host Daphnia magna(Decaestecker et al., 2007). A time-shift assay of changes in resistance and infectivity within populations of L. marginale-M. lini demonstrated striking shifts in interaction traits at just two-year intervals, suggesting that any assessments will be highly sensitive to the point in time the host-pathogen interaction is sampled on(Lively, 1999, Gandon et al., 2008, Thrall et al., 2012).
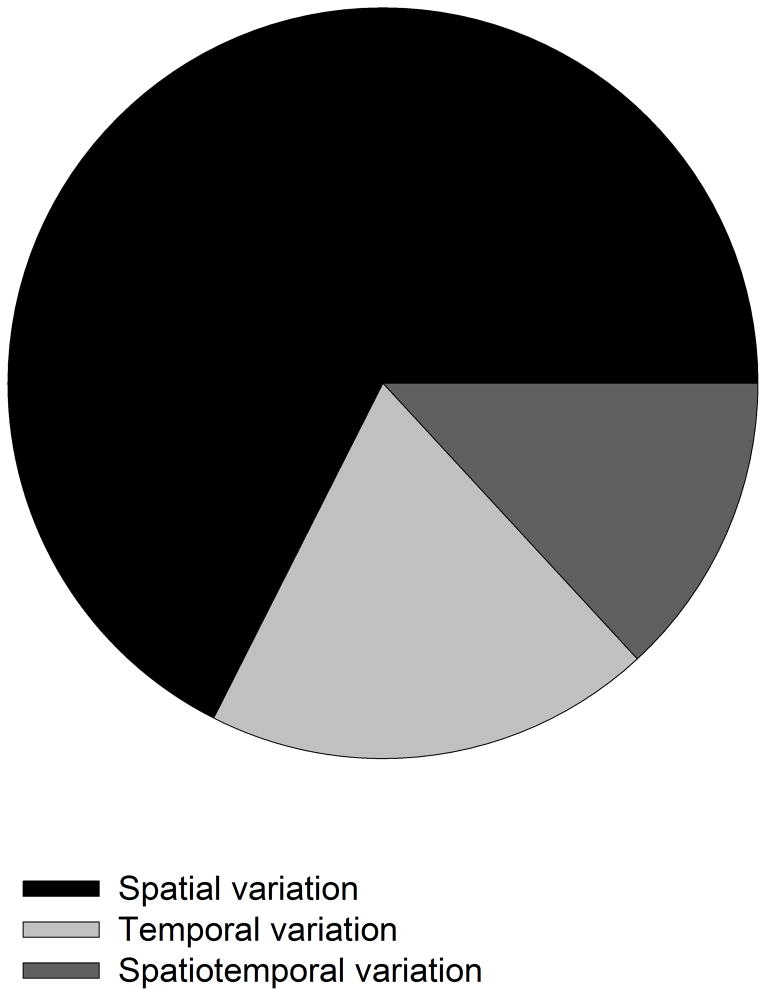
Due to the lack of published results on spatio-temporal variation in pathogenicity in natural systems, we here use a body of unpublished data generated in a long-term study of the flax rust, M. lini, infecting the wild flax L. marginale in Australia (Burdon & Jarosz, 1992, Burdon & Thompson, 1995). This twelve-year sampling program across three populations illustrates clearly that spatial variation (68% of total variation), temporal variation (19% of total variation) and the interaction between space and time (13% of total variation) all contribute to the maintenance of pathogenic polymorphisms (Fig. 3). Such spatio-temporal variation is a signature of asynchrony among populations hence promoting the potential for pathogen local adaptation. While comparative data are lacking, we expect that some study systems will harbour even more temporal and spatio-temporal variation, as the host populations included in the current study represented two different ecotypes (i.e. ‘bog’ and ‘hill’ sites; Carlsson-Granér et al., 1999).
Figure 3.
Relative amount of spatial, temporal, and spatiotemporal variation in pathogenicity in the wild flax – flax rust pathosystem (Linum marginale ~ Melampsora lini). Data is based on pathogen samples (n=659) randomly collected from three populations across twelve years (1986–1997; Burdon et al., unpublished data). Populations are Kiandra, P1, and P2 in Kosciuszko National Park, Australia (see Jarosz & Burdon, 1991 for a map of the populations). Pathogens were tested on a standard set of eleven Linum marginale host lines described by Jarosz & Burdon (1991). A multivariate analysis explained the infectivity (0/1) of the pathogen across all host lines by the variables Population, Year, and the Population x Year interaction. Analyses were implemented with the function adonis in package Vegan (version 1.17-6) in R (Oksanen et al., 2010).
We note that another interesting approach would be to follow both phenotypic and genotypic changes in pathogen population structure over the course of a single epidemic during a host growing season. For example, selection for more aggressive or infective genotypes may be predicted to be an important agent of temporal change within pathogen populations. However, the exact evolutionary trajectory may depend on the existence or strength of a trade-off between aggressiveness and infectivity (Thrall & Burdon, 2003), the frequency of re-colonisations and immigration, and the diversity and average resistance of hosts.
Evidence for selection in shaping variation in pathogenicity
If variation in pathogenicity – as detected across several spatial scales (Table 1) – is adaptive, we would expect local adaptation (i.e. a better performance on the local versus foreign host) of the pathogen to occur across each of these spatial scales. Indeed, a review of the literature shows that local adaptation is common in host-pathogen systems (Table A1 in Appendix A; see also Kaltz & Shykoff, 1998, Greischar & Koskella, 2007, Hoeksema & Forde, 2008). Most importantly, local adaptation is frequently detected at each spatial scale: individual hosts (100%; 3 of 3 studies); populations (52%; 14 of 27 studies), and among regions (67%; 2 of 3 studies). Moreover, for three of the studies included in Fig. 1A, among-population variation in pathogenicity has been shown to be at least partly adaptive at multiple spatial scales (L. marginale ~ M. lini; Plantago lanceolata ~ Podosphaera plantaginis; and P. vulgaris~ C. lindemuthianum). For example, Thrall and colleagues (2002) detected local adaptation of M. lini at both population and metapopulation levels. Likewise, local adaptation of the fungal pathogen C. lindemuthianum to the common bean P. vulgaris has been convincingly demonstrated at the level of the individual host plant (Capelle & Neema, 2005), population (Sicard et al., 1997a) and regional scale (Sicard et al., 1997b). This strongly suggests that the variation detected at each of several spatial scales may be adaptive – emphasizing the potential role for natural selection in shaping variation in pathogenicity within and among populations. Moreover, the frequent detection of local adaptation suggests that FSD may be more common than ARD.
Nevertheless, we note that a lack of local adaptation is alsoa common finding (Kaltz et al., 1999, Goss & Bergelson, 2006). While some studies may have failed to detect local adaptation because of inappropriate experimental designs (Kawecki & Ebert, 2004), the absence of local adaptation should be expected in many situations given the potentially cyclical nature of coevolution where hosts may gain the upper hand over their pathogens occasionally (Kaltz & Shykoff, 1998). Indeed, the occurrence and extent of local adaptation is likely to depend on the strength of selection, frequency of local extinctions (where high extinction rates preclude local adaptation), and the frequency of gene flow (where high gene flow may swamp local adaptation; Slatkin, 1987, Tack & Roslin, 2010). Moreover, pathogen and host life history features (such as dispersal ability, host specialization and generation time) will play a crucial role in which species will be ahead in the coevolutionary race (Kaltz & Shykoff, 1998, Lajeunesse & Forbes, 2002, Greischar & Koskella, 2007, Hoeksema & Forde, 2008).
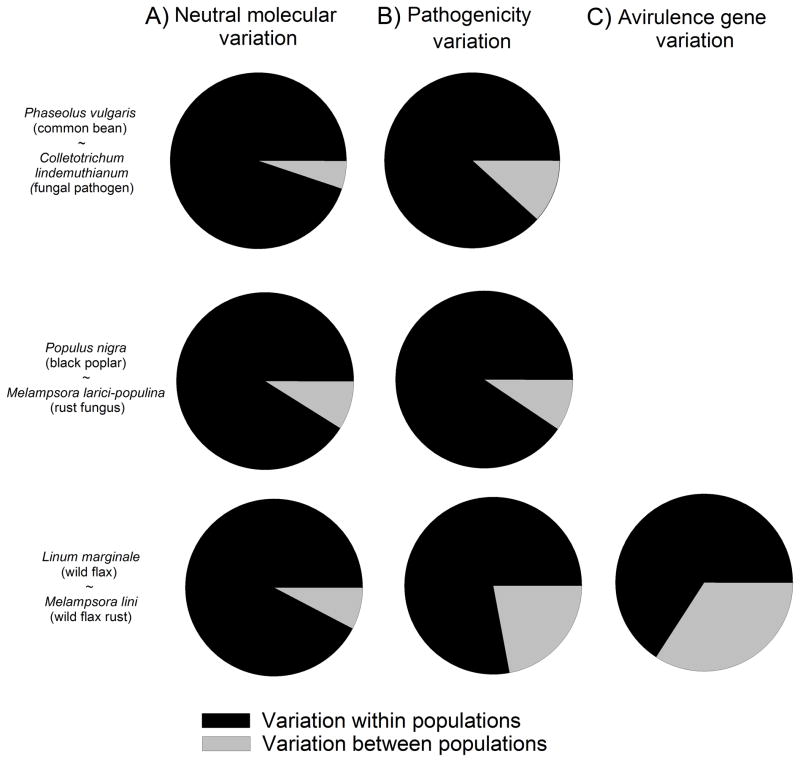
The above examples are based on cross-infection and reciprocal transplant studies. The sign and strength of selection may – for sexually reproducing species -also be inferred from a comparison of patterns of neutral and pathogenic variation among populations. For example, if variation in pathogenicity among populations is driven largely by neutral processes (i.e. mutation and genetic drift), we may expect a match in the amount of differentiation among populations as measured by putatively neutral genetic (e.g. microsatellite markers) and phenotypic variation in pathogenicity (Merilä & Crnokrak, 2001). In contrast, discrepancies between the degree of pathogenic and neutral differentiation may reflect either homogeneous selection across populations (neutral variation among populations > pathogenic variation among populations) or divergent selection (neutral variation among populations <pathogenic variation among populations). Strikingly, two of the three studies in which this can be assessed support the occurrence of divergent host-mediated selection (Figure 4). Thus, Sicard et al. (1997), in a study of the fungal pathogen C. lindemuthianum attacking common bean P. vulgaris, detected increased differentiation at pathogenic compared to neutral genetic markers among populations, suggesting that local selection pressures are important to structure pathogenicity across the metapopulation. Likewise, populations of flax rust M. lini attacking wild flax L. marginale in Australia show generally more divergence in pathogenicity traits and avirulence gene polymorphisms compared to neutral genetic variation (AFLP markers; Barrett et al., 2009b; Figure 4).
Figure 4.
Spatial scale of variation in the pathogen: a) neutral genetic variation, b) pathogenicity, and c) genetic variation in avirulence genes. Multivariate analyses were conducted using the response variables pathogenicity (0/1), RAFLP markers (0/1), and avr-alleles (0/1). The explanatory variable was Population, and the residual error term reflects the within-population variation. Analyses were implemented with the function adonis in package Vegan (version 1.17-6) in R (Oksanen et al., 2010). Data sources(and for more detailed information): Phaseolus vulgaris ~ Colletotrichum lindemuthianum from AMOVA table given in Sicard et al (1997a); Populus nigra ~Melampsora larici-populina from AMOVA table given in Gérard et al (2006); Linum marginale ~ Melampsora lini from Barrett et al.(2009b).
Discussion and future directions
This review on variation in pathogenicity in wild host pathogen systems highlights several general conclusions. First, we show that variation in pathogenicity is ubiquitous across space, ranging from single hosts through patches and metapopulations to entire continents (Table 1). Importantly, this suggests that factors promoting the maintenance of pathogenic polymorphisms - like spatial structure, environmental heterogeneity and gene complexity - play a crucial role in host-pathogen interactions. Second, we demonstrate that much variation in pathogenicity and host resistance is present within populations, thereby indicating the importance of local interactions and FSD. Nevertheless, a significant fraction of variation is present among populations and through time (Figs 1 and 3), likely playing a crucial role in safeguarding the maintenance of genetic variation within the pathosystem. Moreover, the interrelationship between host resistance and pathogenicity suggest that coevolutionary interactions are important in the maintenance of polymorphisms. Finally, at least some part of the variation in pathogenicity at each spatial scale is –in a large fraction of the study systems – adaptive, as illustrated by the frequent detection of local adaptation at each spatial scale (Table A1) and, to a lesser extent, a comparison of neutral and pathogenic variation (Fig. 4). The consistent maintenance of variation within populations, and frequent local adaptation provide some evidence that FSD may be more common in wild host-pathogen systems than ARD. At the same time, we see several gaps and promising directions for future research.
First, environmental variation may be a crucial factor in maintaining the variation in resistance and pathogenicity that we observed, as several recent studies have shown how spatial (Gavrilets & Michalakis, 2008, Laine & Tellier, 2008, Tellier & Brown, 2011) and temporal (Mostowy & Engelstädter, 2011) heterogeneity may maintain polymorphisms. The effect of spatial and temporal heterogeneity on pathogen prevalence is indeed well-described (Schnathorst, 1965, Duniway, 1979, Burdon, 1987). Moreover, recent experiments have indicated the occurrence of genotype-by-environment interactions (i.e. Ghostx E, Gpathogenx E, or Ghostx Gpathogenx E) in host-parasite systems(Laine, 2008, Vale & Little, 2009, Wolinska & King, 2009, Bryner & Rigling, 2011, Sadd, 2011, Hall & Ebert, 2012). Hence, while many of the studies in Table 1 have conducted experiments in a single controlled environment, interactions may be more variable across environments than previously realized. Moreover, environmentally mediated trade-offs between different fitness traits (e.g. infectivity, aggressiveness, survival between epidemics and transmission ability; Koskela et al., 2000, Dybdahl & Storfer, 2003, Hatcher et al., 2005, Abang et al., 2006, Refardt & Ebert, 2007, Wolinska & King, 2009, Barrett et al., 2011, Sommerhalder et al., 2011), or spatially varying costs in resistance and pathogenicity (Gavrilets & Michalakis, 2008, Tellier & Brown, 2011), may well maintain population-level variation in infectivity and aggressiveness.
Second, the current focus on specialist and aerially dispersed pathogens (see Table 1 and Fig. 1) hinders any strong generalizations across the notoriously diverse array of host and parasite life-histories. Most strikingly, appropriate data on soil-borne and vector-transmitted pathogens, generalist pathogens (Barrett et al., 2009a), and viruses (Fraile & García-Arenal, 2010)are lacking (or scarce) for natural pathosystems. New data across a broad suite of study systems would help unravel the contributions of different host and pathogen life-history traits to generating patterns of variation in resistance and pathogenicity at different spatial and temporal scales. Moreover, recent developments in animal pathogen studies provide a promising starting point for future comparisons across a wide range of organisms (Box A).
Third, much is to be gained from integrating the rapidly advancing understanding of molecular interactions governing pathogen infectivity and aggressiveness with studies of ecological genetics in wild-pathogen systems (Burdon & Thrall, 2009, Fraile & García-Arenal, 2010). For example, work on the interaction between flax rust M. liniand its domesticated host (Linum ussitissimum) revealed divergent selection on avirulence genes (as based on a high ratio of nonsynonymous to synonymous mutations; Dodds et al., 2006). Similar patterns have been observed in preliminary studies of the wild host, L. marginale(Barrett et al., 2008). Interestingly, the application of this molecular knowledge and approach to the dynamics of the flax rust – wild flax interaction revealed temporal variation in allele frequencies within populations, with evidence for negative frequency-dependent selection varying across populations and years (Thrall et al., 2012).
Finally, the overwhelming evidence from natural systems is that hosts are attacked by a whole array of parasite species – either at the same time or physically separated in space and time (Agrios, 2005). Crucially, several studies suggest that the community members do not act in isolation, thereby necessitating a community genetics perspective to understand host-parasite interactions (Thrall et al., 2007, Telfer et al., 2010).
Supplementary Material
Acknowledgments
This study was supported by the Academy of Finland (grants 133499, 136393 and 250444) to ALL, by the National Institutes of Health (NIH grant 5RO1 GM074265-01A2) to JJB and PHT, and the Australian Research Council grant (DP1097256) to LGB.
References
- Abang MM, Baum M, Ceccarelli S, Grando S, Linde CC, Yahyaoui A, et al. Differential selection on Rhynchosporium secalis during parasitic and saprophytic phases in the barley scald disease cycle. Phytopathology. 2006;96:1214–1222. doi: 10.1094/PHYTO-96-1214. [DOI] [PubMed] [Google Scholar]
- Agrawal A, Lively CM. Infection genetics: gene-for-gene versus matching-alleles models and all points in between. Evol Ecol Res. 2002;4:79–90. [Google Scholar]
- Agrios GN. Plant pathology. 5. Academic press; New York, USA: 2005. [Google Scholar]
- Al-Yaman F, Genton B, Reeder JC, Anders RF, Smith T, Alpers MP. Reduced risk of clinical malaria in children infected with multiple clones of Plasmodium falciparum in a highly endemic area: a prospective community study. Trans R Soc Trop Med Hyg. 1997;91:602–605. doi: 10.1016/s0035-9203(97)90046-8. [DOI] [PubMed] [Google Scholar]
- Alexander HM. An experimental field study of anther-smut disease of Silene alba caused by Ustilago violacea: genotypic variation and disease incidence. Evolution. 1989;43:835–847. doi: 10.1111/j.1558-5646.1989.tb05181.x. [DOI] [PubMed] [Google Scholar]
- Alexander HM, Antonovics J, Kelly AW. Genotypic variation in plant disease resistance – physiological resistance in relation to field disease transmission. J Ecol. 1993;81:325–333. [Google Scholar]
- Alexander HM, Antonovics J, Rausher MD. Relationship of phenotypic and genetic variation in Plantago lanceolata to disease caused by Fusarium moniliforme var. subglutinans. Oecologia. 1984;65:89–93. doi: 10.1007/BF00384468. [DOI] [PubMed] [Google Scholar]
- Alizon S, Lion S. Within-host parasite cooperation and the evolution of virulence. Proceedings of the Royal Society B: Biological Sciences. 2011;278:3738–3747. doi: 10.1098/rspb.2011.0471. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Altizer SM. Migratory behaviour and host-parasite co-evolution in natural populations of monarch butterflies infected with a protozoan parasite. Evol Ecol Res. 2001;3:611–632. [Google Scholar]
- Anderson RM, May RM. Coevolution of hosts and parasites. Parasitology. 1982;85:411–426. doi: 10.1017/s0031182000055360. [DOI] [PubMed] [Google Scholar]
- Anderson TJC, Romero-Abal ME, Jaenike J. Mitochondrial DNA and Ascaris microepidemiology: the composition of parasite populations from individual hosts, families and villages. Parasitology. 1995;110:221–229. doi: 10.1017/s003118200006399x. [DOI] [PubMed] [Google Scholar]
- Antonovics J, Thrall PH, Burdon JJ, Laine AL. Partial resistance in the Linum-Melampsora host-pathogen system: does partial resistance make the red queen run slower? Evolution. 2011;65:512–522. doi: 10.1111/j.1558-5646.2010.01146.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bangham J, Obbard DJ, Kim KW, Haddrill PR, Jiggins FM. The age and evolution of an antiviral resistance mutation in Drosophila melanogaster. Proceedings of the Royal Society B: Biological Sciences. 2007;274:2027–2034. doi: 10.1098/rspb.2007.0611. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barrett JA. Frequency-dependent selection in plant-fungal interactions. Philos Trans R Soc Lond B Biol Sci. 1988;319:473–483. [Google Scholar]
- Barrett LG, Bell T, Dwyer G, Bergelson J. Cheating, trade-offs and the evolution of aggressiveness in a natural pathogen population. Ecol Lett. 2011;14:1149–1157. doi: 10.1111/j.1461-0248.2011.01687.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barrett LG, Kniskern JM, Bodenhausen N, Zhang W, Bergelson J. Continua of specificity and virulence in plant host–pathogen interactions: causes and consequences. New Phytol. 2009a;183:513–529. doi: 10.1111/j.1469-8137.2009.02927.x. [DOI] [PubMed] [Google Scholar]
- Barrett LG, Thrall PH, Burdon JJ, Nicotra AB, Linde CC. Population structure and diversity in sexual and asexual populations of the pathogenic fungus Melampsora lini. Mol Ecol. 2008;17:3401–3415. doi: 10.1111/j.1365-294X.2008.03843.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barrett LG, Thrall PH, Dodds PN, van der Merwe M, Linde CC, Lawrence GJ, et al. Diversity and evolution of effector loci in natural populations of the plant pathogen Melampsora lini. Mol Biol Evol. 2009b;26:2499–2513. doi: 10.1093/molbev/msp166. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barrus MF. Variation of varieties of beans in their susceptibility to anthracnose. Phytopathology. 1911;1:190–195. [Google Scholar]
- Bergelson J, Dwyer G, Emerson JJ. Models and data on plant-enemy coevolution. Annu Rev Genet. 2001;35:469–499. doi: 10.1146/annurev.genet.35.102401.090954. [DOI] [PubMed] [Google Scholar]
- Bevan JR, Clarke DD, Crute IR. Resistance to Erysiphe fischeri in two populations of Senecio vulgaris. Plant Pathology. 1993a;42:636–646. [Google Scholar]
- Bevan JR, Crute IR, Clarke DD. Variation for virulence in Erysiphe fischeri from Senecio vulgaris. Plant Pathology. 1993b;42:622–635. [Google Scholar]
- Bonhoeffer S, Nowak MA. Intra-host versus inter-host selection: viral strategies of immune function impairment. Proceedings of the National Academy of Sciences. 1994;91:8062–8066. doi: 10.1073/pnas.91.17.8062. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brown JKM, Tellier A. Plant-parasite coevolution: bridging the gap between genetics and ecology. Annu Rev Phytopathol. 2011;49:345–367. doi: 10.1146/annurev-phyto-072910-095301. [DOI] [PubMed] [Google Scholar]
- Brown SP, Hochberg ME, Grenfell BT. Does multiple infection select for raised virulence? Trends Microbiol. 2002;10:401–405. doi: 10.1016/s0966-842x(02)02413-7. [DOI] [PubMed] [Google Scholar]
- Bryner SF, Rigling D. Temperature-dependent genotype-by-genotype interaction between a pathogenic fungus and its hyperparasitic virus. The American Naturalist. 2011;177:65–74. doi: 10.1086/657620. [DOI] [PubMed] [Google Scholar]
- Buckling A, Rainey PB. Antagonistic coevolution between a bacterium and a bacteriophage. Proceedings of the Royal Society of London Series B: Biological Sciences. 2002;269:931–936. doi: 10.1098/rspb.2001.1945. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bull JJ. Virulence. Evolution. 1994;48:1423–1437. doi: 10.1111/j.1558-5646.1994.tb02185.x. [DOI] [PubMed] [Google Scholar]
- Burdon JJ. Diseases and plant population biology. Cambridge University Press; Cambridge, UK: 1987. [Google Scholar]
- Burdon JJ. The structure of pathogen populations in natural plant communities. Annu Rev Phytopathol. 1993;31:305–323. [Google Scholar]
- Burdon JJ, Brown AHD, Jarosz AM. The spatial scale of genetic interactions in host-pathogen coevolved systems. In: Burdon JJ, Leather SR, editors. Pests, pathogens and plant communities. Blackwell; Oxford, UK: 1990. pp. 233–247. [Google Scholar]
- Burdon JJ, Jarosz AM. Host-pathogen interactions in natural populations of Linum marginale and Melampsora lini: I. Patterns of resistance and racial variation in a large host population. Evolution. 1991;45:205–217. doi: 10.1111/j.1558-5646.1991.tb05278.x. [DOI] [PubMed] [Google Scholar]
- Burdon JJ, Jarosz AM. Temporal variation in the racial structure of flax rust (Melampsora lini) populations growing on natural stands of wild flax (Linum marginale): local versus metapopulation dynamics. Plant Pathology. 1992;41:165–179. [Google Scholar]
- Burdon JJ, Marshall DR, Luig NH, Gow DJS. Isozyme studies on the origin and evolution of Puccinia graminis f. sp tritici in Australia. Aust J Biol Sci. 1982;35:231–238. [Google Scholar]
- Burdon JJ, Thompson JN. Changed patterns of resistance in a population of Linum marginale attacked by the rust pathogen Melampsora Lini. J Ecol. 1995;83:199–206. [Google Scholar]
- Burdon JJ, Thrall PH. Coevolution of plants and their pathogens in natural habitats. Science. 2009;324:755–756. doi: 10.1126/science.1171663. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Burdon JJ, Thrall PH, Brown AHD. Resistance and virulence structure in two Linum marginale-Melampsora lini host-pathogen metapopulations with different mating systems. Evolution. 1999;53:704–716. doi: 10.1111/j.1558-5646.1999.tb05365.x. [DOI] [PubMed] [Google Scholar]
- Burdon JJ, Thrall PH, Lawrence GJ. Coevolutionary patterns in the Linum marginale - Melampsora lini association at a continental scale. Can J Bot. 2002;80:288–296. [Google Scholar]
- Butto S, Argentini C, Mazzella AM, Iannotti MP, Leone P, Nicolosi A, et al. Dual infection with different strains of the same HIV-1 subtype. AIDS (London, England) 1997;11:694–696. [PubMed] [Google Scholar]
- Capelle J, Neema C. Local adaptation and population structure at a micro-geographical scale of a fungal parasite on its host plant. J Evol Biol. 2005;18:1445–1454. doi: 10.1111/j.1420-9101.2005.00951.x. [DOI] [PubMed] [Google Scholar]
- Carius HJ, Little TJ, Ebert D. Genetic variation in a host-parasite association: potential for coevolution and frequency-dependent selection. Evolution. 2001;55:1136–1145. doi: 10.1111/j.0014-3820.2001.tb00633.x. [DOI] [PubMed] [Google Scholar]
- Carlsson-Granér U. Anther-smut disease in Silene dioica: variation in susceptibility among genotypes and populations, and patterns of disease within populations. Evolution. 1997;51:1416–1426. doi: 10.1111/j.1558-5646.1997.tb01465.x. [DOI] [PubMed] [Google Scholar]
- Carlsson-Granér U, Burdon JJ, Thrall PH. Host resistance and pathogen virulence across a plant hybrid zone. Oecologia. 1999;121:339–347. doi: 10.1007/s004420050937. [DOI] [PubMed] [Google Scholar]
- Chaboudez P, Burdon JJ. Frequency-dependent selection in a wild plant-pathogen system. Oecologia. 1995;102:490–493. doi: 10.1007/BF00341361. [DOI] [PubMed] [Google Scholar]
- Damgaard C. Coevolution of a plant host–pathogen gene-for-gene system in a metapopulation model without cost of resistance or cost of virulence. J Theor Biol. 1999;201:1–12. doi: 10.1006/jtbi.1999.1007. [DOI] [PubMed] [Google Scholar]
- Davelos AL, Alexander HM, Slade NA. Ecological genetic interactions between a clonal host plant (Spartina pectinata) and associated rust fungi (Puccinia seymouriana and Puccinia sparganioides) Oecologia. 1996;105:205–213. doi: 10.1007/BF00328548. [DOI] [PubMed] [Google Scholar]
- de Nooij MP, van Damme JMM. Variation in host susceptibility among and within populations of Plantago lanceolata L. infected by the fungus Phomopsis subordinaria (Desm) Trav. Oecologia. 1988a;75:535–538. doi: 10.1007/BF00776417. [DOI] [PubMed] [Google Scholar]
- de Nooij MP, van Damme JMM. Variation in pathogenicity among and within populations of the fungus Phomopsis subordinaria infecting Plantago lanceolata. Evolution. 1988b;42:1166–1171. doi: 10.1111/j.1558-5646.1988.tb04177.x. [DOI] [PubMed] [Google Scholar]
- de Roode JC, Helinski MEH, Anwar MA, Read AF. Dynamics of multiple infection and within-host competition in genetically diverse malaria infections. The American Naturalist. 2005;166:531–542. doi: 10.1086/491659. [DOI] [PubMed] [Google Scholar]
- Decaestecker E, Gaba S, Raeymaekers JAM, Stoks R, van Kerckhoven L, Ebert D, et al. Host-parasite ‘Red Queen’ dynamics archived in pond sediment. Nature. 2007;450:870–873. doi: 10.1038/nature06291. [DOI] [PubMed] [Google Scholar]
- Dodds PN, Lawrence GJ, Catanzariti AM, Teh T, Wang CIA, Ayliffe MA, et al. Direct protein interaction underlies gene-for-gene specificity and coevolution of the flax resistance genes and flax rust avirulence genes. Proceedings of the National Academy of Sciences. 2006;103:8888–8893. doi: 10.1073/pnas.0602577103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Duniway JM. Water relations of water molds. Annu Rev Phytopathol. 1979;17:431–460. [Google Scholar]
- Dybdahl MF, Lively CM. Host-parasite coevolution: evidence for rare advantage and time-lagged selection in a natural population. Evolution. 1998;52:1057–1066. doi: 10.1111/j.1558-5646.1998.tb01833.x. [DOI] [PubMed] [Google Scholar]
- Dybdahl MF, Storfer A. Parasite local adaptation: Red Queen versus Suicide King. Trends Ecol Evol. 2003;18:523–530. [Google Scholar]
- Ebert D. Virulence and local adaptation of a horizontally transmitted parasite. Science. 1994;265:1084–1086. doi: 10.1126/science.265.5175.1084. [DOI] [PubMed] [Google Scholar]
- Ebert D. Experimental evolution of parasites. Science. 1998;282:1432–1435. doi: 10.1126/science.282.5393.1432. [DOI] [PubMed] [Google Scholar]
- Ebert D, Zschokke-Rohringer CD, Carius HJ. Within– and between–population variation for resistance of Daphnia magna to the bacterial endoparasite Pasteuria ramosa. Proceedings of the Royal Society of London Series B: Biological Sciences. 1998;265:2127–2134. [Google Scholar]
- Ennos RA, McConnell KC. Using genetic markers to investigate natural selection in fungal populations. Can J Bot. 1995;73:S302–S310. [Google Scholar]
- Ericson L, Burdon JJ. Linking field epidemiological and individual plant resistance patterns in the Betula pubescens–Melampsoridium betulinum host–pathogen interaction. Oikos. 2009;118:225–232. [Google Scholar]
- Ericson L, Burdon JJ, Müller WJ. The rust pathogen Triphragmium ulmariae as a selective force affecting its host, Filipendula ulmaria. J Ecol. 2002;90:167–178. [Google Scholar]
- Espiau C, Riviere D, Burdon JJ, Gartner S, Daclinat B, Hasan S, et al. Host-pathogen diversity in a wild system: Chondrilla juncea – Puccinia chondrillina. Oecologia. 1998;113:133–139. doi: 10.1007/s004420050361. [DOI] [PubMed] [Google Scholar]
- Excoffier L, Smouse PE, Quattro JM. Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics. 1992;131:479–491. doi: 10.1093/genetics/131.2.479. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fisher MC, Henk DA, Briggs CJ, Brownstein JS, Madoff LC, McCraw SL, et al. Emerging fungal threats to animal, plant and ecosystem health. Nature. 2012;484:186–194. doi: 10.1038/nature10947. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Flor HH. Inheritance of pathogenicity in Melampsora lini. Phytopathology. 1942;32:653–669. [Google Scholar]
- Flor HH. The complementary genic systems in flax and flax rust. Adv Genet. 1956;8:29–54. [Google Scholar]
- Forde SE, Thompson JN, Bohannan BJM. Adaptation varies through space and time in a coevolving host-parasitoid interaction. Nature. 2004;431:841–844. doi: 10.1038/nature02906. [DOI] [PubMed] [Google Scholar]
- Fraile A, García-Arenal F. The coevolution of plants and viruses: resistance and pathogenicity. Adv Virus Res. 2010;76:1–32. doi: 10.1016/S0065-3527(10)76001-2. [DOI] [PubMed] [Google Scholar]
- Frank SA. Spatial variation in coevolutionary dynamics. Evol Ecol. 1991;5:193–217. [Google Scholar]
- Frank SA. Coevolutionary genetics of hosts and parasites with quantitative inheritance. Evol Ecol. 1994;8:74–94. [Google Scholar]
- Frank SA. Models of parasite virulence. The Quarterly Review of Biology. 1996;71:37–78. doi: 10.1086/419267. [DOI] [PubMed] [Google Scholar]
- French JR, Manion PD. Variability of host and pathogen in Hypoxylon canker of aspen. Can J Bot. 1975;53:2740–2744. [Google Scholar]
- Friesen TL, Stukenbrock EH, Liu ZH, Meinhardt S, Ling H, Faris JD, et al. Emergence of a new disease as a result of interspecific virulence gene transfer. Nat Genet. 2006;38:953–956. doi: 10.1038/ng1839. [DOI] [PubMed] [Google Scholar]
- Gaba S, Ebert D. Time-shift experiments as a tool to study antagonistic coevolution. Trends in Ecology and Evolution. 2009;24:226–232. doi: 10.1016/j.tree.2008.11.005. [DOI] [PubMed] [Google Scholar]
- Gandon S. Local adaptation and the geometry of host-parasite coevolution. Ecol Lett. 2002;5:246–256. [Google Scholar]
- Gandon S, Buckling A, Decaestecker E, Day T. Host–parasite coevolution and patterns of adaptation across time and space. J Evol Biol. 2008;21:1861–1866. doi: 10.1111/j.1420-9101.2008.01598.x. [DOI] [PubMed] [Google Scholar]
- Gandon S, Capowiez Y, Dubois Y, Michalakis Y, Olivieri I. Local adaptation and gene-for-gene coevolution in a metapopulation model. Proc R Soc Lond Ser B-Biol Sci. 1996;263:1003–1009. [Google Scholar]
- Gandon S, Day T. Evolutionary epidemiology and the dynamics of adaptation. Evolution. 2009;63:826–838. doi: 10.1111/j.1558-5646.2009.00609.x. [DOI] [PubMed] [Google Scholar]
- Gandon S, van Baalen M, Jansen VAA. The evolution of parasite virulence, superinfection, and host resistance. The American Naturalist. 2002;159:658–669. doi: 10.1086/339993. [DOI] [PubMed] [Google Scholar]
- Ganz HH, Washburn JO. Relative migration rates and local adaptation in a mosquito-protozoan interaction. J Evol Biol. 2006;19:816–824. doi: 10.1111/j.1420-9101.2005.01047.x. [DOI] [PubMed] [Google Scholar]
- Garrett KA, Mundt CC. Epidemiology in mixed host populations. Phytopathology. 1999;89:984–990. doi: 10.1094/PHYTO.1999.89.11.984. [DOI] [PubMed] [Google Scholar]
- Gavrilets S, Michalakis Y. Effects of environmental heterogeneity on victim-exploiter coevolution. Evolution. 2008;62:3100–3116. doi: 10.1111/j.1558-5646.2008.00513.x. [DOI] [PubMed] [Google Scholar]
- Geffroy V, Sicard D, de Oliveira JCF, Sévignac M, Cohen S, Gepts P, et al. Identification of an ancestral resistance gene cluster involved in the coevolution process between Phaseolus vulgaris and its fungal pathogen Colletotrichum lindemuthianum. Mol Plant-Microbe Interact. 1999;12:774–784. doi: 10.1094/MPMI.1999.12.9.774. [DOI] [PubMed] [Google Scholar]
- Gérard PR, Husson C, Pinon J, Frey P. Comparison of genetic and virulence diversity of Melampsora larici-populina populations on wild and cultivated poplar and influence of the alternate host. Phytopathology. 2006;96:1027–1036. doi: 10.1094/PHYTO-96-1027. [DOI] [PubMed] [Google Scholar]
- Gomez DR, Evans KJ, Baker J, Harvey PR, Scott ES. Dynamics of introduced populations of Phragmidium violaceum and implications for biological control of European blackberry in Australia. Appl Environ Microbiol. 2008;74:5504–5510. doi: 10.1128/AEM.02885-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gómez P, Buckling A. Bacteria-phage antagonistic coevolution in soil. Science. 2011;332:106–109. doi: 10.1126/science.1198767. [DOI] [PubMed] [Google Scholar]
- Goss EM, Bergelson J. Variation in resistance and virulence in the interaction between Arabidopsis thaliana and a bacterial pathogen. Evolution. 2006;60:1562–1573. [PubMed] [Google Scholar]
- Goyeau H, Berder J, Czerepak C, Gautier A, Lanen C, Lannou C. Low diversity and fast evolution in the population of Puccinia triticina causing durum wheat leaf rust in France from 1999 to 2009, as revealed by an adapted differential set. Plant Pathology. 2012 doi: 10.1111/j.1365–3059.2011.02554.x. [DOI] [Google Scholar]
- Greischar MA, Koskella B. A synthesis of experimental work on parasite local adaptation. Ecol Lett. 2007;10:418–434. doi: 10.1111/j.1461-0248.2007.01028.x. [DOI] [PubMed] [Google Scholar]
- Haldane JBS. Disease and evolution. La Ricerca Scientifica Supplemento. 1949;19:1–11. [Google Scholar]
- Hall MD, Ebert D. Disentangling the influence of parasite genotype, host genotype and maternal environment on different stages of bacterial infection in Daphnia magna. Proceedings of the Royal Society B: Biological Sciences; 2012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hanski I. Metapopulation ecology. Oxford University Press; Oxford, UK: 1999. [Google Scholar]
- Hatcher MJ, Hogg JC, Dunn AM. Local adaptation and enhanced virulence of Nosema granulosis artificially introduced into novel populations of its crustacean host, Gammarus duebeni. Int J Parasitol. 2005;35:265–274. doi: 10.1016/j.ijpara.2004.12.004. [DOI] [PubMed] [Google Scholar]
- Hatchett JH, Gallun RL. Genetics of the ability of the Hessian Fly, Mayetiola destructor, to survive on wheats having different genes for resistance. Ann Entomol Soc Am. 1970;63:1400–1407. [Google Scholar]
- Hedrick PW. Genetics of populations. Jones & Bartlett; Sudbury, MA, USA: 2000. [Google Scholar]
- Hoeksema JD, Forde SE. A meta-analysis of factors affecting local adaptation between interacting species. Am Nat. 2008;171:275–290. doi: 10.1086/527496. [DOI] [PubMed] [Google Scholar]
- Hood ME. Dynamics of multiple infection and within-host competition by the anther-smut pathogen. The American Naturalist. 2003;162:122–133. doi: 10.1086/375539. [DOI] [PubMed] [Google Scholar]
- Huo TI, Wu JC, Lin RY, Sheng WY, Chang FY, Lee SD. Decreasing hepatitis D virus infection in Taiwan: an analysis of contributory factors. J Gastroenterol Hepatol. 1997;12:747–751. doi: 10.1111/j.1440-1746.1997.tb00364.x. [DOI] [PubMed] [Google Scholar]
- Imhoof B, Schmid-Hempel P. Patterns of local adaptation of a protozoan parasite to its bumblebee host. Oikos. 1998;82:59–65. [Google Scholar]
- Jarosz AM, Burdon JJ. Host-pathogen interactions in natural populations of Linum marginale and Melampsora lini: II. Local and regional variation in patterns of resistance and racial structure. Evolution. 1991;45:1618–1627. doi: 10.1111/j.1558-5646.1991.tb02667.x. [DOI] [PubMed] [Google Scholar]
- Jayakar SD. A mathematical model for interaction of gene frequencies in a parasite and its host. Theor Popul Biol. 1970;1:140–164. doi: 10.1016/0040-5809(70)90032-8. [DOI] [PubMed] [Google Scholar]
- Jorgensen TH, Richardson DS, Andersson S. Comparative analyses of population structure in two subspecies of Nigella degenii: evidence for diversifying selection on pollen-color dimorphisms. Evolution. 2006;60:518–528. [PubMed] [Google Scholar]
- Kaltz O, Gandon S, Michalakis Y, Shykoff JA. Local maladaptation in the anther-smut fungus Microbotryum violaceum to its host plant Silene latifolia: evidence from a cross-inoculation experiment. Evolution. 1999;53:395–407. doi: 10.1111/j.1558-5646.1999.tb03775.x. [DOI] [PubMed] [Google Scholar]
- Kaltz O, Shykoff JA. Local adaptation in host-parasite systems. Heredity. 1998;81:361–370. [Google Scholar]
- Kaltz O, Shykoff JA. Within- and among-population variation in infectivity, latency and spore production in a host–pathogen system. J Evol Biol. 2002;15:850–860. [Google Scholar]
- Kawecki TJ, Ebert D. Conceptual issues in local adaptation. Ecol Lett. 2004;7:1225–1241. [Google Scholar]
- Keller SM, McDermott JM, Pettway RE, Wolfe MS, McDonald BA. Gene flow and sexual reproduction in the wheat glume blotch pathogen Phaeosphaeria nodorum (anamorph Stagonospora nodorum) Phytopathology. 1997;87:353–358. doi: 10.1094/PHYTO.1997.87.3.353. [DOI] [PubMed] [Google Scholar]
- Kinloch BB, Stonecypher RW. Genetic variation in susceptibility to fusiform rust in seedlings from a wild population of loblolly pine. Phytopathology. 1969;59:1246–55. [Google Scholar]
- Koskela T, Salonen V, Mutikainen P. Local adaptation of a holoparasitic plant, Cuscuta europaea: variation among populations. J Evol Biol. 2000;13:749–755. [Google Scholar]
- Laine AL. Spatial scale of local adaptation in a plant-pathogen metapopulation. J Evol Biol. 2005;18:930–938. doi: 10.1111/j.1420-9101.2005.00933.x. [DOI] [PubMed] [Google Scholar]
- Laine AL. Detecting local adaptation in a natural plant - pathogen metapopulation: a laboratory vs. field transplant approach. J Evol Biol. 2007;20:1665–1673. doi: 10.1111/j.1420-9101.2007.01359.x. [DOI] [PubMed] [Google Scholar]
- Laine AL. Temperature-mediated patterns of local adaptation in a natural plant-pathogen metapopulation. Ecol Lett. 2008;11:327–337. doi: 10.1111/j.1461-0248.2007.01146.x. [DOI] [PubMed] [Google Scholar]
- Laine AL. Context-dependent effects of induced resistance under co-infection in a plant–pathogen interaction. Evolutionary Applications. 2011;4:696–707. doi: 10.1111/j.1752-4571.2011.00194.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Laine AL, Burdon JJ, Dodds PN, Thrall PH. Spatial variation in disease resistance: from molecules to metapopulations. J Ecol. 2010;99:96–112. doi: 10.1111/j.1365-2745.2010.01738.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Laine AL, Tellier A. Heterogeneous selection promotes maintenance of polymorphism in host–parasite interactions. Oikos. 2008;117:1281–1288. [Google Scholar]
- Lajeunesse MJ, Forbes MR. Host range and local parasite adaptation. Proc R Soc Lond Ser B-Biol Sci. 2002;269:703–710. doi: 10.1098/rspb.2001.1943. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lebeda A, Petrželová I. Variation and distribution of virulence phenotypes of Bremia lactucae in natural populations of Lactuca serriola. Plant Pathology. 2004;53:316–324. [Google Scholar]
- Leibold M, Miller TE. From metapopulations to metacommunities. In: Hanski I, Gaggiotti OE, editors. Ecology, genetics and evolution of metapopulations. Elsevier Academic Press; London, UK: 2004. pp. 133–150. [Google Scholar]
- Lenné JM, Burdon JJ. Preliminary study of virulence and isozymic variation in natural populations of Colletotrichum gloeosporioides from Stylosanthes guianensis. Phytopathology. 1990;80:728–731. [Google Scholar]
- Lenski RE. Two-step resistance by Escheria coli B to bacteriophage T2. Genetics. 1984;107:1–7. doi: 10.1093/genetics/107.1.1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leonard KJ. Selection pressures and plant pathogens. Ann N Y Acad Sci. 1977;287:207–222. [Google Scholar]
- Leonard KJ. Modeling gene frequency dynamics. In: Crute IR, Holub EB, Burdon JJ, editors. The gene-for-gene relationship in plant-parasite interactions. CAB International; London, UK: 1997. pp. 245–262. [Google Scholar]
- Levin S, Pimentel D. Selection of intermediate rates of increase in parasite-host systems. The American Naturalist. 1981;117:308–315. [Google Scholar]
- Linde CC, Zhan J, McDonald BA. Population structure of Mycosphaerella graminicola: from lesions to continents. Phytopathology. 2002;92:946–955. doi: 10.1094/PHYTO.2002.92.9.946. [DOI] [PubMed] [Google Scholar]
- Lipsitch M, Moxon ER. Virulence and transmissibility of pathogens: what is the relationship? Trends Microbiol. 1997;5:31–37. doi: 10.1016/S0966-842X(97)81772-6. [DOI] [PubMed] [Google Scholar]
- Lively CM. Migration, virulence, and the geographic mosaic of adaptation by parasites. The American Naturalist. 1999;153:S34–S47. doi: 10.1086/303210. [DOI] [PubMed] [Google Scholar]
- Lively CM. Parasite-host interactions. In: Fox CW, Roff DA, Fairbairn DJ, editors. Evolutionary ecology: concepts and case studies. Oxford University Press; Oxford, UK: 2001. pp. 331–343. [Google Scholar]
- Lively CM, Dybdahl MF. Parasite adaptation to locally common host genotypes. Nature. 2000;405:679–681. doi: 10.1038/35015069. [DOI] [PubMed] [Google Scholar]
- López-Villavicencio M, Jonot O, Coantic A, Hood ME, Enjalbert J, Giraud T. Multiple infections by the anther smut pathogen are frequent and involve related strains. PLoS pathogens. 2007;3:e176. doi: 10.1371/journal.ppat.0030176. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Luijckx P, Ben-Ami F, Mouton L, du Pasquier L, Ebert D. Cloning of the unculturable parasite Pasteuria ramosa and its Daphnia host reveals extreme genotype–genotype interactions. Ecol Lett. 2011;14:125–131. doi: 10.1111/j.1461-0248.2010.01561.x. [DOI] [PubMed] [Google Scholar]
- Luijckx P, Fienberg H, Duneau D, Ebert D. Resistance to a bacterial parasite in the crustacean Daphnia magna shows Mendelian segregation with dominance. Heredity. 2012;108:547–551. doi: 10.1038/hdy.2011.122. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McDonald BA, Linde C. Pathogen population genetics, evolutionary potential, and durable resistance. Annu Rev Phytopathol. 2002;40:349–379. doi: 10.1146/annurev.phyto.40.120501.101443. [DOI] [PubMed] [Google Scholar]
- McDonald BA, Zhan J, Burdon JJ. Genetic structure of Rhynchosporium secalis in Australia. Phytopathology. 1999;89:639–645. doi: 10.1094/PHYTO.1999.89.8.639. [DOI] [PubMed] [Google Scholar]
- Merilä J, Crnokrak P. Comparison of genetic differentiation at marker loci and quantitative traits. J Evol Biol. 2001;14:892–903. [Google Scholar]
- Meyer SE, Nelson DL, Clement S. Evidence for resistance polymorphism in the Bromus tectorum - Ustilago bullata pathosystem: implications for biocontrol. Can J Plant Pathol. 2001;23:19–27. [Google Scholar]
- Meyer SE, Nelson DL, Clement S, Ramakrishnan A. Ecological genetics of the Bromus tectorum (Poaceae) - Ustilago bullata (Ustilaginaceae) pathosystem: a role for frequency-dependent selection? Am J Bot. 2010;97:1304–1312. doi: 10.3732/ajb.0900261. [DOI] [PubMed] [Google Scholar]
- Meyer SE, Nelson DL, Clement S, Waters J, Stevens M, Fairbanks D. Genetic variation in Ustilago bullata: molecular genetic markers and virulence on Bromus tectorum host lines. Int J Plant Sci. 2005;166:105–115. [Google Scholar]
- Miles JW, Lenné JM. Genetic variation within a natural Stylosanthes guianensis, Colletotrichum gloeosporioides host-pathogen population. Aust J Agric Res. 1984;35:211–218. [Google Scholar]
- Mostowy R, Engelstädter J. The impact of environmental change on host-parasite coevolutionary dynamics. Proc R Soc B-Biol Sci. 2011;278:2283–2292. doi: 10.1098/rspb.2010.2359. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mundt CC. Use of multiline cultivars and cultivar mixtures for disease management. Annu Rev Phytopathol. 2002;40:381–410. doi: 10.1146/annurev.phyto.40.011402.113723. [DOI] [PubMed] [Google Scholar]
- Niemi L, Wennström A, Hjältén J, Waldmann P, Ericson L. Spatial variation in resistance and virulence in the host–pathogen system Salix triandra–Melampsora amygdalinae. J Ecol. 2006;94:915–921. [Google Scholar]
- Nowak MA, May RM. Superinfection and the evolution of parasite virulence. Proceedings of the Royal Society of London Series B: Biological Sciences. 1994;255:81–89. doi: 10.1098/rspb.1994.0012. [DOI] [PubMed] [Google Scholar]
- Nuismer SL, Gandon S. Moving beyond common garden and transplant designs: insight into the causes of local adaptation in species interactions. The American Naturalist. 2008;171:658–668. doi: 10.1086/587077. [DOI] [PubMed] [Google Scholar]
- Oates JD, Burdon JJ, Brouwer JB. Interactions between Avena and Puccinia species. II The pathogens: Puccinia coronata CDA and P graminis Pers f sp avenae Eriks & Henn. J Appl Ecol. 1983;20:585–596. [Google Scholar]
- Oksanen J, Blanchet FG, Kindt R, Legendre P, O’Hara RB, Simpson GL, et al. vegan: Community Ecology Package. 2010 http://CRAN.R-project.org/package=vegan.pp.
- Oppliger A, Vernet R, Baez M. Parasite local maladaptation in the Canarian lizard Gallotia galloti (Reptilia: Lacertidae) parasitized by haemogregarian blood parasite. J Evol Biol. 1999;12:951–955. [Google Scholar]
- Parker IM, Gilbert GS. The evolutionary ecology of novel plant-pathogen interactions. Ann Rev Ecol Evol Syst. 2004;35:675–700. [Google Scholar]
- Parker MA. Local population differentiation for compatibility in an annual legume and its host-specific fungal pathogen. Evolution. 1985;39:713–723. doi: 10.1111/j.1558-5646.1985.tb00414.x. [DOI] [PubMed] [Google Scholar]
- Parker MA. Polymorphism for disease resistance in the annual legume Amphicarpaea bracteata. Heredity. 1988;60:27–31. [Google Scholar]
- Parker MA. Disease impact and local genetic diversity in the clonal plant Podophyllum peltatum. Evolution. 1989;43:540–547. doi: 10.1111/j.1558-5646.1989.tb04250.x. [DOI] [PubMed] [Google Scholar]
- Pisoni G, Bertoni G, Puricelli M, Maccalli M, Moroni P. Demonstration of coinfection with and recombination by caprine arthritis-encephalitis virus and maedi-visna virus in naturally infected goats. J Virol. 2007;81:4948–4955. doi: 10.1128/JVI.00126-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Read AF. The evolution of virulence. Trends Microbiol. 1994;2:73–76. doi: 10.1016/0966-842x(94)90537-1. [DOI] [PubMed] [Google Scholar]
- Read AF, Taylor LH. The ecology of genetically diverse infections. Science. 2001;292:1099–1102. doi: 10.1126/science.1059410. [DOI] [PubMed] [Google Scholar]
- Refardt D, Ebert D. Inference of parasite local adaptation using two different fitness components. J Evol Biol. 2007;20:921–929. doi: 10.1111/j.1420-9101.2007.01307.x. [DOI] [PubMed] [Google Scholar]
- Ridenhour BJ, Nuismer SL. Polygenic traits and parasite local adaptation. Evolution. 2007;61:368–376. doi: 10.1111/j.1558-5646.2007.00029.x. [DOI] [PubMed] [Google Scholar]
- Roslin T, Laine AL, Gripenberg S. Spatial population structure in an obligate plant pathogen colonizing oak Quercus robur. Funct Ecol. 2007;21:1168–1177. [Google Scholar]
- Roy BA. Differentiating the effects of origin and frequency in reciprocal transplant experiments used to test negative frequency-dependent selection hypotheses. Oecologia. 1998;115:73–83. doi: 10.1007/s004420050493. [DOI] [PubMed] [Google Scholar]
- Sacristán S, García-Arenal F. The evolution of virulence and pathogenicity in plant pathogen populations. Mol Plant Pathol. 2008;9:369–384. doi: 10.1111/j.1364-3703.2007.00460.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sadd BM. Food-environment mediates the outcome of specific interactions between a bumblebee and its trypanosome parasite. Evolution. 2011;65:2995–3001. doi: 10.1111/j.1558-5646.2011.01345.x. [DOI] [PubMed] [Google Scholar]
- Salvaudon L, Giraud T, Shykoff JA. Genetic diversity in natural populations: a fundamental component of plant-microbe interactions. Curr Opin Plant Biol. 2008;11:135–143. doi: 10.1016/j.pbi.2008.02.002. [DOI] [PubMed] [Google Scholar]
- Sasaki A. Host-parasite coevolution in a multilocus gene-for-gene system. Proc R Soc Lond Ser B-Biol Sci. 2000;267:2183–2188. doi: 10.1098/rspb.2000.1267. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sasaki A, Godfray HCJ. A model for the coevolution of resistance and virulence in coupled host–parasitoid interactions. Proceedings of the Royal Society of London Series B: Biological Sciences. 1999;266:455–463. [Google Scholar]
- Sasaki A, Hamilton WD, Ubeda F. Clone mixtures and a pacemaker: new facets of Red-Queen theory and ecology. Proceedings of the Royal Society of London Series B: Biological Sciences. 2002;269:761–772. doi: 10.1098/rspb.2001.1837. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schierup MH, Charlesworth D, Vekemans X. The effect of hitch-hiking on genes linked to a balanced polymorphism in a subdivided population. Genetics Research. 2000a;76:63–73. doi: 10.1017/s0016672300004547. [DOI] [PubMed] [Google Scholar]
- Schierup MH, Vekemans X, Charlesworth D. The effect of subdivision on variation at multi-allelic loci under balancing selection. Genetics Research. 2000b;76:51–62. doi: 10.1017/s0016672300004535. [DOI] [PubMed] [Google Scholar]
- Schnathorst WC. Environmental relationships in the powdery mildews. Annu Rev Phytopathol. 1965;3:343–366. [Google Scholar]
- Sicard D, Buchet S, Michalakis Y, Neema C. Genetic variability of Colletotrichum lindemuthianum in wild populations of common bean. Plant Pathology. 1997a;46:355–365. [Google Scholar]
- Sicard D, Michalakis Y, Dron M, Neema C. Genetic diversity and pathogenic variation of Colletotrichum lindemuthianum in the three centers of diversity of its host, Phaseolus vulgaris. Phytopathology. 1997b;87:807–813. doi: 10.1094/PHYTO.1997.87.8.807. [DOI] [PubMed] [Google Scholar]
- Sicard D, Pennings PS, Grandclément C, Acosta J, Kaltz O, Shykoff JA. Specialization and local adaptation of a fungal parasite on two host plant species as revealed by two fitness traits. Evolution. 2007;61:27–41. doi: 10.1111/j.1558-5646.2007.00003.x. [DOI] [PubMed] [Google Scholar]
- Slatkin M. Gene flow and the geographic structure of natural populations. Science. 1987;236:787–792. doi: 10.1126/science.3576198. [DOI] [PubMed] [Google Scholar]
- Smith DL, Ericson L, Burdon JJ. Co-evolutionary hot and cold spots of selective pressure move in space and time. J Ecol. 2011;99:634–641. [Google Scholar]
- Sommerhalder RJ, McDonald BA, Mascher F, Zhan J. Effect of hosts on competition among clones and evidence of differential selection between pathogenic and saprophytic phases in experimental populations of the wheat pathogen Phaeosphaeria nodorum. BMC Evol Biol. 2011;11:188. doi: 10.1186/1471-2148-11-188. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Springer YP. Clinal resistance structure and pathogen local adaptation in a serpentine flax-flax rust interaction. Evolution. 2007;61:1812–1822. doi: 10.1111/j.1558-5646.2007.00156.x. [DOI] [PubMed] [Google Scholar]
- Stukenbrock EH, McDonald BA. The origins of plant pathogens in agro-ecosystems. Annu Rev Phytopathol. 2008;46:75–100. doi: 10.1146/annurev.phyto.010708.154114. [DOI] [PubMed] [Google Scholar]
- Tack AJM, Roslin T. Overrun by the neighbors: Landscape context affects strength and sign of local adaptation. Ecology. 2010;91:2253–2260. doi: 10.1890/09-0080.1. [DOI] [PubMed] [Google Scholar]
- Telfer S, Lambin X, Birtles R, Beldomenico P, Burthe S, Paterson S, et al. Species interactions in a parasite community drive infection risk in a wildlife population. Science. 2010;330:243–246. doi: 10.1126/science.1190333. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tellier A, Brown JKM. Stability of genetic polymorphism in host–parasite interactions. Proceedings of the Royal Society B: Biological Sciences. 2007;274:809–817. doi: 10.1098/rspb.2006.0281. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tellier A, Brown JKM. Spatial heterogeneity, frequency-dependent selection and polymorphism in host-parasite interactions. BMC Evol Biol. 2011;11:319. doi: 10.1186/1471-2148-11-319. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Theron A, Sire C, Rognon A, Prugnolle F, Durand P. Molecular ecology of Schistosoma mansoni transmission inferred from the genetic composition of larval and adult infrapopulations within intermediate and definitive hosts. Parasitology. 2004;129:571–585. doi: 10.1017/s0031182004005943. [DOI] [PubMed] [Google Scholar]
- Thompson JN. The geographic mosaic of coevolution. University of Chicago Press; Chicago, USA: 2005. [Google Scholar]
- Thompson JN, Burdon JJ. Gene-for-gene coevolution between plants and parasites. Nature. 1992;360:121–125. [Google Scholar]
- Thrall PH, Antonovics J. Theoretical and empirical studies of metapopulations: population and genetic dynamics of the Silene–Ustilago system. Can J Bot. 1995;73:1249–1258. [Google Scholar]
- Thrall PH, Burdon JJ. Host-pathogen dynamics in a metapopulation context: the ecological and evolutionary consequences of being spatial. J Ecol. 1997;85:743–753. [Google Scholar]
- Thrall PH, Burdon JJ. The spatial scale of pathogen dispersal: consequences for disease dynamics and persistence. Evol Ecol Res. 1999;1:681–701. [Google Scholar]
- Thrall PH, Burdon JJ. Effect of resistance variation in a natural plant host-pathogen metapopulation on disease dynamics. Plant Pathol. 2000;49:767–773. [Google Scholar]
- Thrall PH, Burdon JJ. Evolution of gene-for-gene systems in metapopulations: the effect of spatial scale of host and pathogen dispersal. Plant Pathology. 2002;51:169–184. [Google Scholar]
- Thrall PH, Burdon JJ. Evolution of virulence in a plant host-pathogen metapopulation. Science. 2003;299:1735–1737. doi: 10.1126/science.1080070. [DOI] [PubMed] [Google Scholar]
- Thrall PH, Burdon JJ, Bever JD. Local adaptation in the Linum marginale - Melampsora lini host-pathogen interaction. Evolution. 2002;56:1340–1351. doi: 10.1111/j.0014-3820.2002.tb01448.x. [DOI] [PubMed] [Google Scholar]
- Thrall PH, Burdon JJ, Young A. Variation in resistance and virulence among demes of a plant host–pathogen metapopulation. J Ecol. 2001;89:736–748. [Google Scholar]
- Thrall PH, Hochberg ME, Burdon JJ, Bever JD. Coevolution of symbiotic mutualists and parasites in a community context. Trends in Ecology and Evolution. 2007;22:120–126. doi: 10.1016/j.tree.2006.11.007. [DOI] [PubMed] [Google Scholar]
- Thrall PH, Jarosz AM. Host-pathogen dynamics in experimental populations of Silene alba and Ustilago violacea I Ecological and genetic determinants of disease spread. J Ecol. 1994;82:549–559. [Google Scholar]
- Thrall PH, Laine AL, Ravensdale M, Nemri A, Dodds PN, Barrett LG, et al. Rapid genetic change underpins antagonistic coevolution in a natural host-pathogen metapopulation. Ecol Lett. 2012;15:425–435. doi: 10.1111/j.1461-0248.2012.01749.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Torres JR. Hepatitis B and hepatitis delta virus infection in South America. Gut. 1996;38:S48–S55. doi: 10.1136/gut.38.suppl_2.s48. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vale PF, Little TJ. Measuring parasite fitness under genetic and thermal variation. Heredity. 2009;103:102–109. doi: 10.1038/hdy.2009.54. [DOI] [PubMed] [Google Scholar]
- Vanderplank JE. Disease resistance in plants. Academic Press; New York, USA: 1968. [Google Scholar]
- Wennström A, Ericson L. The effect and transmission of one isolate of the rust Puccinia minussensis on five clones of Lactuca sibirica. Oecologia. 1994;97:407–411. doi: 10.1007/BF00317332. [DOI] [PubMed] [Google Scholar]
- Wille P, Boller T, Kaltz O. Mixed inoculation alters infection success of strains of the endophyte Epichloë bromicola on its grass host Bromus erectus. Proceedings of the Royal Society of London Series B: Biological Sciences. 2002;269:397–402. doi: 10.1098/rspb.2001.1889. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wolfe MS. The current status and prospects of multiline cultivars and variety mixtures for disease resistance. Annu Rev Phytopathol. 1985;23:251–273. [Google Scholar]
- Wolinska J, King KC. Environment can alter selection in host-parasite interactions. Trends Parasitol. 2009;25:236–244. doi: 10.1016/j.pt.2009.02.004. [DOI] [PubMed] [Google Scholar]
- Zhu Y, Chen H, Fan J, Wang Y, Li Y, Chen J, et al. Genetic diversity and disease control in rice. Nature. 2000;406:718–722. doi: 10.1038/35021046. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.