Abstract
Background
Anopheles gambiae sensu stricto (s.s.) is a primary vector of Plasmodium falciparum in sub-Saharan Africa. Although some physiological differences among molecular and chromosomal forms of this species have been demonstrated, the relative susceptibility to malaria parasite infection among them has not been unequivocally shown. The objective of this study was to investigate P. falciparum circumsporozoite protein infection (CSP) positivity among An. gambiae s.s. chromosomal and molecular forms.
Methods
Wild An. gambiae from two sites Kela (n = 464) and Sidarebougou (n = 266) in Mali were screened for the presence of P. falciparum CSP using an enzyme-linked immunosorbent assay (ELISA). Samples were then identified to molecular form using multiple PCR diagnostics (n = 713) and chromosomal form using chromosomal karyotyping (n = 419).
Results
Of 730 An. gambiae sensu lato (s.l.) mosquitoes, 89 (12.2%) were CSP ELISA positive. The percentage of positive mosquitoes varied by site: 52 (11.2%) in Kela and 37 (13.9%) in Sidarebougou. Eighty-seven of the positive mosquitoes were identified to molecular form and they consisted of nine Anopheles arabiensis (21.4%), 46 S (10.9%), 31 M (12.8%), and one MS hybrid (14.3%). Sixty of the positive mosquitoes were identified to chromosomal form and they consisted of five An. arabiensis (20.0%), 21 Savanna (15.1%), 21 Mopti (30.4%), 11 Bamako (9.2%), and two hybrids (20.0%).
Discussion
In this collection, the prevalence of P. falciparum infection in the M form was equivalent to infection in the S form (no molecular form differential infection). There was a significant differential infection by chromosomal form such that, P. falciparum infection was more prevalent in the Mopti chromosomal forms than in the Bamako or Savanna forms; the Mopti form was also the most underrepresented in the collection. Continued research on the differential P. falciparum infection of An. gambiae s.s. chromosomal and molecular forms may suggest that Plasmodium – An. gambiae interactions play a role in malaria transmission.
Keywords: Anopheles gambiae, Mali, Malaria, Plasmodium falciparum, Molecular form, Chromosomal form
Background
Several members of the Anopheles gambiae complex are vectors of human malaria parasites, including Plasmodium falciparum, the species of greatest public health importance in Sub-Saharan Africa. The An. gambiae complex consists of at least seven member species and subspecies [1-3]. Anopheles gambiae sensu stricto (s.s.) is further divided into chromosomal and molecular forms [3]. Molecular studies of the ribosomal DNA region on the X chromosome revealed a fixed difference between populations of An. gambiae that is the basis of the sub-division into the M and S molecular forms [4]. The M and S forms are assortatively mating discreet forms [5-7] that are hypothesized to be undergoing ecotypic speciation due to different larval habitat adaptation [8-10]. Anopheles gambiae are also divided into chromosomal forms based on the arrangements of 5 paracentric inversions on 2R (j, b,c, u and d) and one on 2 L (a), which define Mopti from Savanna from Bamako forms [3,11]. In Mali, the M form generally associates with the Mopti chromosomal form and the S form with the Savanna and Bamako forms [12]. In some locations in Africa, including Mali, the presence of significant deficiencies in certain inversion heterozygotes suggests there are barriers to gene flow among the chromosomal forms [11,13-16].
Genetic polymorphisms within An. gambiae have been associated with adaptation to different local environments [17-21]. For example, a number of genetic polymorphisms are associated with tolerance of arid conditions [22]. There is also significant interest in genotypic variation associated with malaria parasite infectivity. Plasmodium falciparum circumsporozoite protein (CSP) positivity in An. gambiae was significantly greater in mosquitoes with isozyme allotypes Mpi130/130 and Acp110/100[23]. In addition, 2La/a individuals were nearly twice as likely to be positive than 2La-/a-[24]. Quantitative trait loci associated with resistance to parasite development have also been identified on chromosome 2 L [25]. Chromosome 2 L includes APL1[26], and chromosome 3 L includes TEP1r[27] both genes are thought to have anti-parasitic properties. Genome-wide transcriptome analyses of An. gambiae have identified transcript expression patterns that are significantly associated with malaria parasite and bacterial infection [28-31]. Further, some transcription patterns and infection-associated sequence polymorphisms have been associated with An. gambiae laboratory and field-collected M and S molecular forms [32,33]. Three single nucleotide polymorphisms (SNPs) in immune signaling genes of Malian An. gambiae were significantly associated with natural P. falciparum infection [33]. These SNPs are predicted to alter the structure and function of the encoded proteins and, therefore, alter refractoriness and susceptibility to P. falciparum infection [33]. Additionally, population-specific SNPs associated with either the M or S molecular forms were associated with P. falciparum infection, indicating potential differential immune responses of the two molecular forms to parasite infection [33]. Recently, a cryptic subgroup of An. gambiae was identified in the Sudan-Savanna zone as susceptible to P. falciparum[34].
Reports of genetic, proteomic, and genomic differences in An. gambiae s.s. suggest, by extension, that susceptibility to P. falciparum infection varies among different molecular forms and populations of An. gambiae[25,27,33,35]. This study examined in more detail the hypothesis that P. falciparum infection prevalence of An. gambiae s.s. differs among molecular and chromosomal forms in Mali. Natural P. falciparum infection levels were compared among An. gambiae molecular and chromosomal forms in two villages in Mali where these forms occur in sympatry. Inversion-specific correlations between P. falciparum infection and standard, heterozygous, and homozygous chromosomal arrangements [3] were also investigated.
Methods
Mosquito collections
Adult An. gambiae mosquitoes were collected in October 2009 from the villages of Kela (11.88683 N, -8.44744 W), and Sidarebougou (11.4568 N,–5.7323 W) joined with Kolayerebougou (11.4563 N,–5.746 W) in Mali. Resting mosquitoes were collected in the morning via mouth aspirators from inside homes. Mosquitoes were held in cups until they had reached the half-gravid stage. Mosquitoes morphologically identified as An. gambiae s.l. were dissected and separated into head/thorax for P. falciparum CSP ELISA, abdomen/wings/legs for molecular form identification via polymerase chain reaction (PCR), and ovaries for chromosomal form identification via karyotyping. Head/thorax samples were stored in 100% ethanol, while abdomens/legs/wings were stored in 70% ethanol and half-gravid extracted ovaries were stored in modified Carnoy’s solution (3:1 ethanol to glacial acetic acid).
ELISA identification of P. falciparum infection
Lysates of head/thorax samples were assayed using a P. falciparum CSP ELISA [36,37] according to protocols provided by the Centers for Disease Control and Prevention (Atlanta, Georgia, USA) to identify the sporozoite stage (not gametocyte) and that the P. falciparum protozoan had disseminated across the midgut. The head and thorax, stored in 100% ethanol, were dried prior to tissue lysis. For each ELISA plate, a minimum of two colony-reared An. gambiae mosquitoes (e.g., negative controls) and serial dilutions of P. falciparum monoclonal antibodies (i.e., sensitivity positive controls) were used. The positive control P. falciparum CSP was serially diluted (i.e., 100 pg to 1.5 pg of antigen per 50 μl of blocking buffer) to quantify CSP in field-collected mosquitoes. Samples with absorbance values greater than three times the standard deviations from the mean of the negative control samples on each ELISA plate were designated as “positive” for P. falciparum infection [38]. CSP ELISAs were conducted instead of PCR for molecular detection of P. falciparum to ensure the protozoan had disseminated the midgut and that the protozoan was in the ‘infective’ sporozoite phase.
Identification of species and molecular forms
Abdomen, legs and wings stored in ethanol were ground using a TissueLyser (Qiagen, Valencia, CA, USA), after which DNA was extracted using the BioSprint 96 Bloodkit and automated workstation (Qiagen, Valencia CA, USA). Mosquitoes morphologically identified as An. gambiae s.l. were identified to species [39] and An. gambiae molecular form identifications were performed on each mosquito [40-42].
Cytogenetics
Polytene chromosome spreads were prepared from ovarian nurse cells [43], except that spreads were not stained with lacto-orcein prior to examination. Chromosome banding patterns were visualized using an Olympus BX-50 phase contrast microscope. Species identification and paracentric inversion scoring were accomplished using the polytene chromosome maps for An. gambiae complex and chromosomal forms [3,11].
Statistical analyses
Summary statistics, relative abundance of forms, Fisher’s exact tests and two-tailed T-tests were performed in Excel 2007 to determine differences within populations [44]. Where appropriate, we adjusted p-values for multiple comparisons using the Bonferroni correction for an α of 0.05. For the molecular form comparisons, five Chi-square comparisons were conducted (form, village, form x site) that generated a significant p-value less than 0.010. Six comparisons were performed with the chromosomal form data (form, village, form x village) and after the Bonferroni correction, Chi-square comparisons were considered significant if the p-value was less than 0.008. Eighteen comparisons were performed with the karyotype data (arrangement, village, arrangement x village) and, after the Bonferroni correction was applied, Chi-square comparisons were considered significant if the p-value was less than 0.003.
Results
In total, 730 An. gambiae s.l. were analysed, of which 42 (5.8%) were identified as An. arabiensis[3,39]. Data from Kolayerebougou and Sidarebougou were combined (hereafter Sidarebougou) because they are located within 1.5 km of one another, have similar habitats, and likely represent a single Mendelian population. In Kela, 21.9% (7/25) of Anopheles arabiensis were CSP ELISA positive and 20% (2/10) were CSP ELISA positive in Sidarebougou. Seventeen of the mosquitoes were not identified to molecular form or karyotyped, and two were positive, both from Kela. The remaining 671 mosquitoes were identified as An. gambiae s.s., of which 78 (11.6%) were P. falciparum CSP ELISA positive. Infection prevalence differed for the two collection sites, from 13.9% (35/251) in Sidarebougou and 10.2% (43/420) in Kela, but they were not significantly different (X2 = 2.101, df = 1, P = 0.1472).
The frequencies of molecular and chromosomal forms of An. gambiae s.s are presented in Figure 1. Of the 713 mosquitoes identified, 671 were identified to An. gambiae s.s. molecular form (94.1%) and 372 were identified to An. gambiae s.s. chromosomal form (52.1%). Both molecular and chromosomal form identifications were performed for 395 mosquitoes of which 370 were An. gambiae s.s. and 25 were An. arabiensis, separate analyses were conducted for each.
Figure 1.
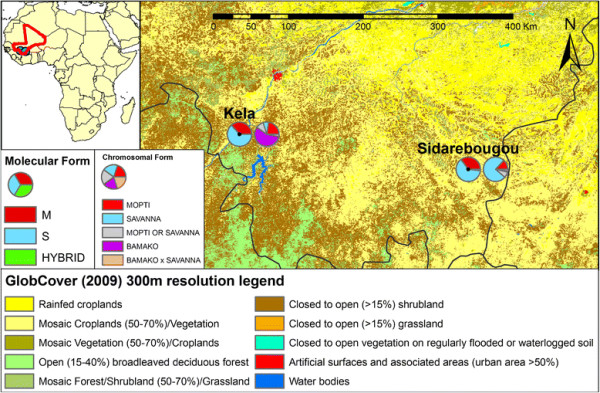
Molecular (left) and chromosomal (right) form frequencies inAnopheles gambiae s.s.collected from Kela and Sidarebougou in western Mali during October 2009. Mopti/Savanna refers to karyotypes could be classified as either Mopti or Savanna (e.g., bcu heterozygotes). Black point in the middle of the molecular graph identifies the site of the village.
Molecular form and P. Falciparum infection
Infectivity and molecular form association analyses were conducted on 671 An. gambiae s.s. This sample was comprised of 62.9% S, 36.1% M and 1.0% M/S molecular form hybrids, but the relative abundance of each molecular form and infection prevalence varied between sites (Figure 1A, Table 1). Within Kela, 63.1% were S, 36.4% were M and 0.5% were M/S hybrids whereas within Sidarebougou 62.5% were S, 35.5% were M and 2.0% were M/S hybrids. Of the 671 mosquitoes identified to the An. gambiae s.s. molecular forms 78 (11.6%) were P. falciparum CSP ELISA positive, of which, 31 were M, 46 were S, and 1 was a M/S molecular hybrid. Contingency tests on the number of positive to negative samples for each An. gambiae molecular form were not significantly different (X2 = 0.595; df = 2; P = 0.743: Bonferroni adjusted p- value = 0.0102) (Table 1). Contingency tests revealed that the number of CSP positive and negative S molecular forms (X2 =0.870; df = 1; P = 0.351) or M molecular forms (X2 = 1.07; df = 1; P = 0.300) were not significantly different at these collection sites. At Sidarebougou (X2 = 0.427; df = 1; P = 0.514) and Kela (X2 = 0.178; df = 1; P = 0.673), infection prevalence among molecular forms was not significantly different.
Table 1.
The M molecular form ofAnopheles gambiae s.s.was associated with greaterP. falciparumCSP positivity than were other molecular forms in October 2009
| Molecular Form | No. Screened | No. Pos. (% Pos.) |
|---|---|---|
|
Kela | ||
| M |
153 |
17 (11.1%) |
| S |
265 |
26 (9.8%) |
| M/S |
2 |
0 (0.0%) |
| Total |
420 |
43 (10.2%) |
|
Sidarebougou | ||
| M |
89 |
14 (15.7%) |
| S |
157 |
20 (12.7%) |
| M/S |
5 |
1 (20.0%) |
| Total |
251 |
35 (13.9%) |
|
Total in Mali | ||
| M |
242 |
31 (12.8%) |
| S |
422 |
46 (10.9%) |
| M/S |
7 |
1 (14.3%) |
| Total | 671 | 78 (11.6%) |
Chromosomal form and P. Falciparum infection
Of 372 karyotyped An. gambiae s.s., 37.4% were Savanna, 32.3% were Bamako, 18.5% were Mopti, 9.1% were Mopti-Savanna, and 2.7% were Bamako-Savanna. Of the 207 samples karyotyped from Kela, 57.0% were Bamako, 22.7% were Mopti, 7.2% were Savanna, 10.1% were Mopti-Savanna, and 2.9% were Bamako-Savanna (Figure 1B). In Sidarebougou, the frequency of each chromosomal form was 75.2% Savanna, 13.3% Mopti, 7.9% Mopti or Savanna, 2.4% Bamako-Savanna, and 1.2% Bamako (Figure 1B). A total of 55 (14.8%) of the 372 karyotyped An. gambiae s.s. were CSP positive (Table 2). Those mosquitoes that were positive for P. falciparum CSP were distributed among chromosomal forms as follows: 21 were Savanna, 21 were Mopti, 11 were Bamako, and two were Bamako-Savanna (Table 2). None of the 34 undifferentiated (2R bcu/+ or bc/u or b/cu) Mopti or Savanna forms were positive. Among the three main chromosomal forms, Mopti had the highest CSP positivity (30.4%), followed by Savanna (15.1%) and Bamako (9.2%), and this trend was significant after Bonferroni correction (X2 = 14.8; df = 2; P value = 0.001: Bonferroni adjusted P value = 0.009) (Table 2). Overall, Mopti chromosomal forms had nearly twice as many positive specimens (compared to negative specimens) than the Savanna or Bamako forms although fewer Mopti specimens were collected (Table 2).
Table 2.
The Mopti chromosomal form ofAnopheles gambiae s.s.was associated with greaterP. falciparumCSP positivity than were other chromosomal forms or hybrids in October 2009
|
Species |
No. Screened |
No. Pos. (% Pos.) |
|
Kela | ||
| Bamako |
118 |
11 (9.3%) |
| Savanna |
15 |
5 (33.3%) |
| Mopti |
47 |
14 (29.8%) |
| Bamako x Savanna |
6 |
1 (16.7%) |
| Mopti or Savanna |
21 |
0 (0%) |
| Total |
207 |
31 (15.0%) |
|
Sidarebougou | ||
| Bamako |
2 |
0 (0.0%) |
| Savanna |
124 |
16 (12.9%) |
| Mopti |
22 |
7 (31.8%) |
| Bamako x Savanna |
4 |
1 (25.0%) |
| Mopti or Savanna |
13 |
0 (0.0%) |
| Total |
165 |
24 (14.5%) |
|
Mali | ||
| Bamako |
120 |
11 (9.2%) |
| Savanna |
139 |
21 (15.1%) |
| Mopti |
69 |
21 (30.4%) |
| Bamako x Savanna |
10 |
2 (20.0%) |
| Mopti or Savanna |
34 |
0 (0.0%) |
| Total | 372 | 55 (14.8%) |
Based on site, there were no significant differences between the number of CSP positive and negative Bamako (X2 = 0.050; df = 1; P = 0.651), Savanna (X2 = 4.35; df = 1; P = 0.037), or Mopti chromosomal forms (X2 = 0.029; df = 1; P = 0.864). Within Kela, Savanna and Mopti chromosomal forms were significantly more likely to be positive than the Bamako form (X2 = 13.4; df = 2; P = 0.001: Bonferroni adjusted P value = 0.0085; Table 2). There were no significant differences in infection prevalence based on chromosomal forms within Sidarebougou (X2 = 5.47; df = 2; P = 0.065; Table 2). The trend that the most prevalent form from each village (Bamako in Kela and Savanna in Sidarebougou) was least likely to be infected, was noted.
Chromosomal inversions and P. Falciparum infection
There were no significant associations of individual inversions and infection status (Table 3). None of the mosquitoes collected had the standard 2La arrangement, and only 16 were heterozygous for the inversion.
Table 3.
Chromosomal inversions inAnopheles gambiae s.s.were not significantly associated withP. falciparumCSP positivity in October 2009 after the Bonferroni correction (P < 0.003)
| Inversion |
No. Positive / No. Screened (% Positive) |
||
|---|---|---|---|
|
Kela |
Sidarebougou |
Total |
|
| 32 / 215 (14.9%) | 27 / 179 (15.1%) | 59 / 335 (15.0%) | |
|
2La Inversion | |||
| Standard |
0 / 0 (0%) |
0 / 0 (0%) |
0 / 0 (0%) |
| Heterozygous |
1 / 2 (50%) |
1 / 14 (7.1%) |
2/ 16 (12.5%) |
| Inversion |
31 / 213 (14.6% |
26 / 165 (15.8%) |
57 / 378 (15.1%) |
| Statistic |
NAa |
NA |
X2 = 0.080; df = 1; P = 0.777 |
|
2Rb Inversion | |||
| Standard |
8 / 83 (9.6%) |
1 / 9 (11.1%) |
9 / 92 (9.8%) |
| Heterozygous |
14 / 90 (15.6%) |
7 / 47 (14.9%) |
21 / 137 (15.3%) |
| Inversion |
10 / 42 (23.8%) |
19 / 123 (15.4%) |
29 / 165 (17.6%) |
| Statistic |
X2 = 4.48; df = 2; P = 0.107 |
X2 = 0.125; df = 2; P = 0.939 |
X2 = 2.84; df = 2; P = 0.242 |
|
2Rc Inversion | |||
| Standard |
11 / 46 (23.9%) |
19 / 139 (13.7%) |
30 / 185 (16.2%) |
| Heterozygous |
9 / 46 (19.6%) |
5 / 30 (16.7%) |
14 / 76 (18.4%) |
| Inversion |
12 / 123 (9.8%) |
3 / 10 (30.0%) |
15 / 133 (11.3%) |
| Statistic |
X2 = 6.31; df = 2; P = 0.043 |
X2 = 2.01; df = 2; P = 0.366 |
X2 = 2.36; df = 2; P = 0.307 |
|
2Rd Inversion | |||
| Standard |
31 / 210 (14.8%) |
24 / 166 (14.5%) |
55 / 376 (14.6%) |
| Heterozygous |
1 / 5 (20.0%) |
3 / 12 (25.0%) |
4 / 17 (23.5%) |
| Inversion |
0 / 0 (0%) |
0 / 1 (0.0%) |
0 / 1 (0.0%) |
| Statistic |
NA |
X2 = 0.967; df = 1; P = 0.326 |
X2 = 1.01; df = 1; P = 0.315 |
|
2Rj Inversion | |||
| Standard |
20 / 93(21.5%) |
27 / 176 (15.3%) |
47 / 269 (17.5%) |
| Heterozygous |
0 / 0 (0.0%) |
0 / 1 (0%) |
0 / 1 (0.0%) |
| Inversion |
12 / 122 (9.8%) |
0 / 2 (0%) |
12 / 124 (9.7%) |
| Statistic |
X2 = 5.67; df = 1; P = 0.017 |
NA |
X2 = 4.04; df = 1; P = 0.044 |
|
2Ru Inversion | |||
| Standard |
9 / 40 (22.5%) |
24 / 153 (15.7%) |
33 / 193 (17.1%) |
| Heterozygous |
12 / 52 (23.1%) |
3 / 24 (12.5%) |
15 / 76 (19.7%) |
| Inversion |
11 / 123 (8.9%) |
0 / 2 (0%) |
11 / 125 (8.8%) |
| Statistic |
X2 = 8.01; df = 2; P = 0.018 |
X2 = 0.163; df = 1; P = 0.686 |
X2 = 5.78; df = 2; P = 0.056 |
a No calculations were conducted because sample size was too small (n < 5).
Molecular and chromosomal forms and P. Falciparum infection
There were 370 An. gambiae s.s. identified to both molecular and chromosomal form. Of the 206 typed from Kela and 165 typed from Sidarebougou, 30 and 24 were CSP positive, respectively. The literature suggests that in Mali, most M molecular forms correlate with the Mopti chromosomal form, and the S molecular form correlates with the Bamako and the Savanna chromosomal forms [12] (Table 4). This was largely the case with these data: 57 of 62 M molecular form samples were identified as Mopti chromosomal form and 118 Savanna and 119 Bamako chromosomal forms were identified as S molecular. Two S molecular forms were identified as Mopti chromosomal forms, whereas 19 M molecular forms were identified as Savanna chromosomal forms. Only 21 of 315 individuals (6.7%) did not follow this trend. Of the five MS molecular hybrids, one was identified as a Bamako chromosomal form and four were identified as Savanna chromosomal forms. In Kela, there were 31 M and one S molecular form mosquitoes identified as MoptixSavanna chromosomal hybrids; six were positive. In Sidarebougou, there were 15 M (one positive) and three S (one positive) identified as MoptixSavanna chromosomal hybrids. Discordant associations (e.g., Savanna chromosomal form with M molecular form or S molecular form with Mopti chromosomal) resulted in these discrepancies. The basis for these discrepancies may stem from the molecular form diagnostics overestimating the number of hybrids in the samples [45].
Table 4.
P. falciparumCSP positivity ofAnopheles gambiae s.s.identified to both molecular and chromosomal form. Status as M molecular form and Mopti chromosomal form was significantly associated with CSP positivity in October 2009 (P = 0.001)
|
No. CSP Pos. / No. Screened (% Pos.) | ||||
|---|---|---|---|---|
| Chromosomal Form |
Molecular Form |
Total | ||
| M | S | M/S | ||
|
Kela | ||||
|
Bamako |
0/0 (0%) |
11/117 (9.4%) |
0/1 (0%) |
11/118 (9.3%) |
|
Savanna |
2/6 (33.3%) |
5/10 (50.0%) |
0/0 (0%) |
7/16 (43.8%) |
|
Mopti |
6/38 (15.8%) |
0/1 (0.0%) |
0/0 (0%) |
6/39 (15.4%) |
|
Total |
8/44 (18.2%) |
16/128 (12.5%) |
0/1 (0%) |
24/173 (13.9%) |
|
Sidarebougou | ||||
|
Bamako |
0/0 (0.0%) |
0/2 (0%) |
0/0 (0.0%) |
0/2 (0.0%) |
|
Savanna |
1/13 (7.7%) |
15/108 (13.9%) |
1/4 (25.0%) |
17/125 (13.6%) |
|
Mopti |
5/19 (26.3%) |
0/1 (0%) |
0/0 (0.0%) |
5/20 (25.0%) |
|
Total |
6/32 (18.8%) |
15/111 (13.5%) |
1/4 (25.0%) |
22/147 (15.0%) |
|
Mali | ||||
|
Bamako |
0/0 (0.0%) |
11/119 (9.2%) |
0/1 (0.0%) |
11/120 (9.2%) |
|
Savanna |
3/19 (15.8%) |
20/118 (16.9%) |
1/4 (25.0%) |
24/141 (17.0%) |
|
Mopti |
11/57 (19.3%) |
0/2 (0.0%) |
0/0 (0.0%) |
11/59 (18.6%) |
| Total | 14/76 (18.4%) | 31/239 (13.0%) | 1/5 (20.0%) | 46/320 (14.4%) |
Among mosquitoes identified as Mopti chromosomal and M molecular forms, 15.8% (6/38) and 26.3% (5/19) were CSP positive in Kela and Sidarebougou, respectively. Among mosquitoes identified as Bamako chromosomal and S molecular forms in Kela, 9.4% (11/106) were CSP positive, whereas none (0/2) with this chromosomal and molecular form combination were CSP positive in Sidarebougou. Among mosquitoes identified as Savanna chromosomal and S molecular forms in Kela, 50.0% (5/10) were CSP positive, while 13.9% (15/108) of this form combination were CSP positive in Sidarebougou. These differences were significant in Kela (X2 = 13.4; df = 2; P < 0.001), but not in Sidarebougou (X2 = 1.88; df = 1; P = 0.170). On average across villages, 19.3% (11/57) of Mopti M forms were CSP positive, 16.9% (20/118) of Savanna S forms were CSP positive, and 9.2% (11/119) Bamako S forms were positive and these patterns were not significant (X2 =4.32, df = 2, P = 0.115).
Discussion
An average of 12.2% of An. gambiae s.s. resting indoors were P. falciparum CSP positive from southern Mali in October 2009. Both molecular forms were CSP ELISA positive and there was no differential infection rate among molecular forms. Significantly more Mopti chromosomal forms (30.4%) were positive than were the Savanna (15.1%) and Bamako (9.2%) chromosomal forms. As the Mopti chromosomal form corresponds to M molecular form in Mali in most cases [12], finding that both M molecular and Mopti chromosomal forms were significantly associated with P. falciparum infection is not surprising. Site-specific differences in the number of CSP positive chromosomal form infection between Kela and Sidarebougou were also observed. In particular, the most common chromosomal form in each village, Bamako in Kela and Savanna in Sidarebougou, was least likely to be CSP positive.
The insignificant infection prevalence in the M molecular form in southern Mali corroborates with other studies from Cameroon and Senegal that reported no differences in P. falciparum infection between M and S molecular forms [6,46]. A recent P. falciparum susceptibility assay among An. gambiae s.s. molecular forms from Senegal found significantly higher numbers of P. falciparum oocysts and sporozoites in the S molecular form than in the M form [47]. This study analysed field-collected specimens of an unknown-age structure that were naturally infected with P. falciparum, whereas the Senegal study [47] collected eggs from the field and allowed the surviving adults to feed directly on a membrane with P. falciparum to standardize age and potential for infection. These conflicting findings may result from the origin of field collected samples (Mali vs. Senegal), to different techniques (natural vs. artificial infection), or to a differences in the age structure of the samples. Field studies from multiple sites and over multiple sampling periods are necessary to confirm the observed patterns.
Cryptic genetic differences in An. gambiae s.s. among sample sites can also limit comparisons among the previous studies [6,46] and the present study. Genetic subdivisions beyond the M and S form designations have been reported and it is possible that differences among these subdivisions include genes associated with differential response to parasite infection. For example, recent studies in Cameroon demonstrated a subdivision in the M molecular form into discrete Forest-M, characterized as M molecular form and Forest chromosomal form (fixed for standard gene arrangement) and Mopti-M populations with typical Mopti karyotypes [12,21]. In analyses using SNPs from immune signaling genes, three genetically distinct An. gambiae s.s. populations were observed in Mali: the M molecular form, the S molecular form (S1), and a subdivided S Pimperena form (S2) [33]. Further, SNPs associated with P. falciparum infection were differentially distributed among M, S1, and S2 populations [33]. Of interest, data presented here is similar to that reported in Riehle et al[34] where a cryptic subgroup of An. gambiae, indistinguishable in molecular form but distinguishable via microsatellites amplified from chromosome 3, was susceptible to P. falciparum[34].
Differences in the local environment may likewise affect associations between An. gambiae forms and P. falciparum infection. For example, Dolo et al[48] demonstrated that irrigated zones of Mali allowed for constant CSP positivity across seasons along with low human blood feeding and sporozoites indices, whereas in the non-irrigated zones, CSP positivity fluctuated seasonally, being high in the wet season and low in the dry season. Dolo et al[48] hypothesized that malaria prevalence in villages adjacent to irrigated rice fields is consistently low in this environment because adult density is inversely related to blood feeding due to high mosquito densities driving villagers to protect themselves with repellants and bed nets [49,50]. The Bamako chromosomal and S molecular forms were dominant in Kela (~3 km to a river), whereas the Savanna chromosomal form and S molecular form predominated in Sidarebougou (~10 km to agriculture fields). Kela is located close to a river that has the ability to flood and create additional oviposition sites not in the dry season likely increasing mosquito densities (11.2% prevalence), whereas in Sidarebougou mosquitoes densities are likely dependent on the wet season (13.9% prevalence). These habitat differences could contribute to the genetic variation (and potentially phenotypic variation) observed at different locations and, as Dolo et al[48] hypothesized, habitat may play a role in the vector ecology of An. gambiae. Collectively, these studies highlight the importance of both genetic and environmental determinants of susceptibility to infection.
There were statistically significant P. falciparum differential infection rates among chromosomal forms and trends among chromosome inversions. Mopti had the highest CSP positivity (30.4%), followed by Savanna (15.1%) and Bamako (9.2%) forms. The data presented here indicated that Mopti chromosomal forms (2Rbc/u) were more likely to be positive for P. falciparum CSP. A previous study in Kenya identified a significant association between the total number of inversions and a decreased likelihood for P. falciparum infection [24]. Data presented here did not show this specific association to be statistically significant, but standard chromosomal arrangements tended to increase the likelihood of being CSP positive. In particular, in Kela where the Bamako form was dominant and least likely to be positive, mosquitoes with standard or heterozygous arrangements were more likely to be CSP positive than mosquitoes homozygous for 2Rjcu and 2Rjbcu (Bamako form) (Table 3).
A number of studies have examined associations of chromosomal polymorphisms with malaria infection. A small region of chromosome 2 L has been associated with infection susceptibility, regardless of P. falciparum genotype [25]. Genes within this region encode for melanization or parasite encapsulation [25]. Within the 2La region, the APL1 gene, which encodes for natural resistance to P. falciparum, exhibited extremely low genetic diversity within the M molecular form, but high diversity in the S molecular form that may have arisen from larval infection [26,35]. Alternatively, higher diversity at the APL1 locus in the S molecular form may be associated with a more diverse array of responses to P. falciparum and reduced susceptibility to infection. Similar results were identified within the anti-parasite and anti-bacterial gene TEP1r[27]. Specifically, TEP1r was diverged between the M and S molecular forms and one variant showed a strong association with resistance to infection with a rodent malaria parasite and with P. falciparum[27]. Additional studies comparing An. gambiae molecular and chromosomal forms with P. falciparum infection that incorporate An. gambiae speciation, genetic diversity at immune loci (e.g., TEP1r, APL1, Toll5B) as well as larger temporal and spatial scales may help to extend findings reported in this study.
Conclusion
Whilst the correlation between M and Mopti forms was expected, higher infection prevalences in the Mopti chromosomal form have not been demonstrated previously. In general, significant differences in P. falciparum infection prevalence at geographic locations with multiple molecular and chromosomal forms are likely due to discrepancies in the relative abundance of forms, genetic diversity in immune signaling genes within different forms, age structure of field collections, and local environmental variations that influence infection and transmission success.
Competing interests
The authors declare we have no competing interests with the work presented in this manuscript.
Authors’ contributions
AJC, GCL, SL, and YL conceived the study and designed the experiments. YL, AJC, AF, and SFT conducted the work in Mali. YL, CSN, and RTF conducted the laboratory work. RTF analysed the data and wrote the manuscript. All authors have read and approved the final manuscript.
Contributor Information
Rebecca T Trout Fryxell, Email: RFryxell@utk.edu.
Catelyn C Nieman, Email: ccsingh@ucdavis.edu.
Abdrahamane Fofana, Email: afofana@icermali.org.
Yoosook Lee, Email: yoslee@ucdavis.edu.
Sekou F Traoré, Email: cheick@mrtcbko.org.
Anthony J Cornel, Email: Cornel@uckac.edu.
Shirley Luckhart, Email: sluckhart@ucdavis.edu.
Gregory C Lanzaro, Email: gclanzaro@ucdavis.edu.
Acknowledgements
We are grateful to the field collectors and the Mali Malaria Research and Training Center (MRTC-Bamako) for support and assistance during field collections. We would like to thank Drs. Clare Marsden and Michelle Sanford (Dept. Pathology, Microbiology, and Immunology, School of Veterinary Medicine, University of California Davis) for edits in the initial drafts of the manuscript. We would like to acknowledge Sarah Han, Saman Mahmood, and Allison Weakley, (Dept. Pathology, Microbiology, and Immunology, School of Veterinary Medicine, University of California Davis) for helping with additional molecular form identifications. This research was funded by NIH 1R01 AI078183 to SL and GCL. RTF is supported by the NIH T32 AI074550 Training Grant.
References
- Ribeiro H, Da Cunha Ramos H, Alves Pires C, Antunes Capela R. Description and biometric study ofAnopheles (Cellia) quadriannulatus davidsonissp. N., a seventh member of theAnopheles gambiaeGiles complex(Diptera: Culicidae)endemic to the Cape Verde archipelago. Sep Garcia de Orta Sér Zool. 1979;8:75–88. [Google Scholar]
- Brunhes J, Le Goff G, Geoffroy B. Anophéles afro-tropicaux. I. Desciptions d’Espéces nouveller et changements de status taxonomiques (Diptera: Culicidae) Ann Soc Entomol Fr. 1997;33:173–183. [Google Scholar]
- Coluzzi M, Sabatini A. della Torre A, Di Deco MA, Petrarca V: A polytene chromosome analysis of theAnopheles gambiaespecies complex. Science. 2002;298:1415–1418. doi: 10.1126/science.1077769. [DOI] [PubMed] [Google Scholar]
- Favia G, Della Torre A, Bagayoko M, Lanfrancotti A, Sagnon NF, Touré YT, Coluzzi M. Molecular identification of sympatric chromosomal forms ofAnopheles gambiaeand further evidence of their reproductive isolation. Insect Mol Biol. 1997;6:377–383. doi: 10.1046/j.1365-2583.1997.00189.x. [DOI] [PubMed] [Google Scholar]
- Lanzaro GC, Tripet F. In: Ecological aspects for the application of genetically modified mosquitoes. Takken W, Scott TW, editor. Frontis Press, Wageningen, The Netherlands; 2003. Gene flow among populations ofAnopheles gambiae: a critical review; pp. 109–132. [Google Scholar]
- Wondji C, Simard F, Petrarca V, Etang J, Santolamazza F, Della Torre A, Fontenille D. Species and populations of theAnopheles gambiaecomplex in Cameroon with special emphasis on chromosomal and molecular forms ofAnopheles gambiaes.s. J Med Entomol. 2005;42:998–1005. doi: 10.1603/0022-2585(2005)042[0998:SAPOTA]2.0.CO;2. [DOI] [PubMed] [Google Scholar]
- White BJ, Cheng C, Simard F, Constantini C, Besansky NJ. Genetic association of physically unlinked islands of genomic divergence in incipient species ofAnopheles gambiae. Molec Ecol. 2010;19:925–939. doi: 10.1111/j.1365-294X.2010.04531.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Manoukis NC, Powell JR, Toure MB, Sacko A, Edillo FE, Coulibaly MB, Traore SF, Taylor CE, Besansky NJ. A test of the chromosomal theory of ecotypic speciation inAnopheles gambiae. Proc Natl Acad Sci USA. 2007;105:2940–2945. doi: 10.1073/pnas.0709806105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lehmann T, Diabate A. The molecular forms ofAnopheles gambiae:A phenotypic perspective. Infect Genet Evol. 2008;8:737–746. doi: 10.1016/j.meegid.2008.06.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gimonneau G, Bouyer J, Morand S, Besansky NJ, Diabate A, Simard F. A behavioral mechanism underlying ecological divergence in the malaria mosquitoAnopheles gambiae. Behavioral Ecol. 2010;21:1087–1092. doi: 10.1093/beheco/arq114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Toure YT, Petrarca V, Traore SF, Coulibaly A, Maiga HM, Sankare O, Sow M, Di Deco MA, Coluzzi M. The distribution and inversion polymorphism of chromosomally recognized taxa of theAnopheles gambiaecomplex in Mali, West Africa. Parassitologia. 1998;40:477–511. [PubMed] [Google Scholar]
- Lee Y, Cornel AJ, Meneses CR, Fofana A, Andrianarivo AG, McAbee R, Fondjo E, Traore SF, Lanzaro GC. Ecological and genetic relationships of theForest-Mform among chromosomal and molecular forms of the malaria vectorAnopheles gambiae sensu stricto. Malar J. 2009;8:75. doi: 10.1186/1475-2875-8-75. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Taylor CE, Toure YT, Carnahan J, Norris DE, Dolo G, Traore SF, Edillo FE, Lanzaro GC. Gene flow among populations of the malaria vector,Anopheles gambiae, in Mali, West Africa. Genetics. 2001;157:743–750. doi: 10.1093/genetics/157.2.743. [DOI] [PMC free article] [PubMed] [Google Scholar]
- della Torre A, Costantini C, Besansky NJ, Caccone A, Petrarca V, Powell JR, Coluzzi M. Speciation withinAnopheles gambiae- the glass is half full. Science. 2002;298:115–117. doi: 10.1126/science.1078170. [DOI] [PubMed] [Google Scholar]
- Costantini C, Ayala D, Guelbeogo WM, Pombi M, Some CY, Bassole IHN, Ose K, Fotsin JM, Sagnon N, Fontenille D, Besansky NJ, Simard F. Living at the edge: biogeographic patterns of habitat segregation conform to speciation by niche expansion inAnopheles gambiae. BMC Ecol. 2009;9:16. doi: 10.1186/1472-6785-9-16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Simard F, Ayala D, Kamdem GC, Pombi M, Etouna J, Ose K, Fotsing JM, Fontenille D, Besansky NJ, Costantini C. Ecological niche portioning betweenAnopheles gambiaemolecular forms in Cameroon: the ecological side of speciation. BMC Ecol. 2009;9:17. doi: 10.1186/1472-6785-9-17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lehmann T, Hawley WA, Kamau L, Fontenilles D, Simard F, Collins FH. Genetic differentiation ofAnopheles gambiaepopulations from East and West Africa: comparison of microsatellite and allozyme loci. Heredity. 1996;77:192–208. doi: 10.1038/hdy.1996.124. [DOI] [PubMed] [Google Scholar]
- Lehmann T, Licht M, Elissa N, Maega BTA, Chimumbwa JM, Watsenga FT, Wondji CS, Simard F, Hawley WA. Population structure ofAnopheles gambiaein Africa. J Heredity. 2003;94:133–147. doi: 10.1093/jhered/esg024. [DOI] [PubMed] [Google Scholar]
- Tripet F, Wright J, Cornel A, Fofana A, McAbee R, Meneses C, Reimer L, Slotman M, Thiemann T, Dolo G, Traore S, Lanzaro GC. Longitudinal survey of knockdown resistance to pyrethroid (kdr) in Mali, West Africa, and evidence of its emergence in the Bamako form of Anopheles gambiae s.s. AmJTrop Med Hyg. 2007;76:81–87. [PubMed] [Google Scholar]
- Oliviera E, Salgueiro P, Palsson K, Vicente JL, Arez AP, Jaenson TG, Caccone A, Pinto J. High levels of hybridization between molecular forms ofAnopheles gambiaefrom Guinea Bissau. J Med Entomol. 2008;45:1057–1063. doi: 10.1603/0022-2585(2008)45[1057:HLOHBM]2.0.CO;2. [DOI] [PubMed] [Google Scholar]
- Slotman MA, Parmakelis A, Marshall JC, Awono-Ambene PH, Antonio-Nkondjo C, Simard F, Caccone A, Powell JR. Patterns of selection in anti-malarial immune genes in malaria vectors: evidence for adaptive evolution inLRIM1inAnopheles arabiensis. PLoS One. 2007;8:e793. doi: 10.1371/journal.pone.0000793. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee Y, Meneses CR, Fofana A, Lanzaro G. Desiccation resistance among subpopulations ofAnopheles gambiae s.s.from Selinkenyi, Mali. J Med Entomol. 2009;46:316–320. doi: 10.1603/033.046.0216. [DOI] [PubMed] [Google Scholar]
- Coosemans M, Smits A, Roelants P. Intraspecific isozyme polymorphism ofAnopheles gambiaein relation to environment, behavior, and malaria transmission in southwestern Burkina Faso. AmJTrop Med Hyg. 1998;58:70–74. doi: 10.4269/ajtmh.1998.58.70. [DOI] [PubMed] [Google Scholar]
- Petrarca V, Beier JC. Intraspecific chromosomal polymorphism in theAnopheles gambiaecomplex as a factor affecting malaria transmission in the Kisumu area of Kenya. AmJTrop Med Hyg. 1992;46:229–237. doi: 10.4269/ajtmh.1992.46.229. [DOI] [PubMed] [Google Scholar]
- Riehle MM, Markianos K, Niare O, Xu J, Li J, Toure AM, Podiougou B, Oduol F, Diawara S, Diallo M, Coulibaly B, Ouatara A, Kruglyak L, Traore SF, Vernick KD. Natural malaria infection inAnopheles gambiaeis regulated by a single genomic control region. Science. 2006;312:577–579. doi: 10.1126/science.1124153. [DOI] [PubMed] [Google Scholar]
- Rottschaefer SM, Riehle MM, Coulibaly B, Sacko M, Niare O, Morlai I, Traore SF, Vernick KD, Lazzaro BP. Exceptional diversity, maintenance of polymorphism, and recent directional selection on theAPL1malaria resistance genes ofAnopheles gambiae. PLoS Biol. 2011;9:e1000600. doi: 10.1371/journal.pbio.1000600. [DOI] [PMC free article] [PubMed] [Google Scholar]
- White BJ, Lawniczak MKN, Cheng C, Coulibaly MB, Wilson MD, Sagnon N, White BJ, Lawniczak MKN, Cheng C, Coulibaly MB, Wilson MD, Sagnon N, Costantini C, Simard F, Christophides GK, Besansky NJ. Adaptive divergence between incipient species ofAnopheles gambiaeincreases resistance toPlasmodium. Proc Natl Acad Sci USA. 2011;108:224–249. doi: 10.1073/pnas.1013648108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Waterhouse RM, Kriventseva EV, Meister S, Xi Z, Alvarez KS, Bartholomay LC, Barillas-Mury C, Bian G, Blandin S, Christensen BM, Dong Y, Jiang H, Kanost MR, Koutsos AC, Levashina EA, Li J, Ligoxygakis P, MacCallum RM, Mayhew GF, Mendes A, Michel K, Osta MA, Paskewitz S, Shin SW, Vlachou D, Wang L, Wei W, Zheng L, Zou Z, Severson DW, Raikhel AS, Kafatos FC, Dimopoulos G, Zdobnov EM, Christophides GK. Evolutionary dynamics of immune-related genes and pathways in disease-vector mosquitoes. Science. 2007;316:1738–1743. doi: 10.1126/science.1139862. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baton LA, Robertson AN, Warr E, Strand MR, Dimopoulos G. Genome-wide transcriptomic profiling ofAnopheles gambiaehemocytes reveals pathogen-specific signatures upon bacterial challenge andPlasmodium bergheiinfection. BMC Genomics. 2009;10:257. doi: 10.1186/1471-2164-10-257. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pinto SB, Lombardo F, Koutsos AC, Waterhouse RM, McKay K, An C, Ramakrishnan C, Kafatos FC, Michel K. Discovery ofPlasmodiummodulators by genome-wide analysis of circulating hemocytes inAnopheles gambiae. Proc Natl Acad Sci USA. 2009;106:21270–21275. doi: 10.1073/pnas.0909463106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cirimotich CM, Dong Y, Garver LS, Sim S, Dimopoulos G. Mosquito immune defenses againstPlasmodiuminfection. Dev Comp Immunol. 2010;34:387–395. doi: 10.1016/j.dci.2009.12.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aguilar R, Simard F, Kamdem C, Shields T, Glass GE, Garver LS, Dimopoulos G. Genome-wide analysis of transcriptomic divergence between laboratory colony and fieldAnopheles gambiaemosquitoes of the M and S molecular forms. Insect Molec Biol. 2010;19:695–705. doi: 10.1111/j.1365-2583.2010.01031.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Horton AA, Lee Y, Coulibaly CA, Rashbrook VK, Cornel AJ, Lanzaro GC, Luckhart S. Identification of three signle nucleotide polymorphisms inAnopheles gambiaeimmune signaling genes that are associated with naturalPlasmodium falciparuminfection. Malar J. 2010;9:160. doi: 10.1186/1475-2875-9-160. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Riehle MM, Guelbeogo WM, Gneme A, Eiglmeier K, Holm I, Bischoff E, Garnier T, Snyder GM, Li X, Markianos K, Sagnon N, Vernick KD. A cryptic subgroup of Anopheles gambiae is highly susceptible to human malaria parasites. Science. 2011;331:596–598. doi: 10.1126/science.1196759. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rottschaefer SM, Riehle MM, Coulibaly B, Sacko M, Niare O, Morlai I, Traore SF, Vernick KD, Lazzaro BP. Exceptional diversity, maintenance of polymorphism, and recent directional selection on theAPL1malaria resistance genes ofAnopheles gambiae. PLoS Biol. 2010;9:e1000600. doi: 10.1371/journal.pbio.1000600. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Burkot TR, Williams JL, Schneider I. Identification ofPlasmodium falciparum-infected mosquitoes by a double antibody enzyme-linked immunosorbent assay. AmJTrop Med Hyg. 1984;33:783–788. doi: 10.4269/ajtmh.1984.33.783. [DOI] [PubMed] [Google Scholar]
- Wirtz RA, Zavala F, Charoenvit Y, Campbell GH, Burkot TR, Schneider I, Esser KM, Beaudoin RL, Andre RG. Comparative testing of monoclonal antibodies againstPlasmodium falciparumsporozoites for ELISA development. Bull World Health Organ. 1987;65:39–45. [PMC free article] [PubMed] [Google Scholar]
- Beier JC, Perkins PV, Wirtz RA, Whitmire RE, Mugambi M, Hockmeyer WT. Field evaluation of an enzyme-linked immunosorbent assay (ELISA) forPlasmodium falciparumsporozoite detection in Anopheline mosquitoes from Kenya. AmJTrop Med Hyg. 1997;36:459–468. doi: 10.4269/ajtmh.1987.36.459. [DOI] [PubMed] [Google Scholar]
- Scott JA, Brogdon WG, Collins FH. Identification of single specimens of theAnopheles gambiaecomplex by the polymerase chain reaction. AmJTrop Med Hyg. 1993;49:520–529. doi: 10.4269/ajtmh.1993.49.520. [DOI] [PubMed] [Google Scholar]
- Fanello C, Santolamazza F. della Torre A: Simultaneous identification of species and molecular forms of theAnopheles gambiaecomplex by PCR-RFLP. Med Vet Entomol. 2002;16:461–464. doi: 10.1046/j.1365-2915.2002.00393.x. [DOI] [PubMed] [Google Scholar]
- Favia G, Lanfrancotti A, Spanos L, Siden-Kiamos I, Louis C. Molecular characterization of ribosomal DNA polymorphisms discriminating among chromosomal forms ofAnopheles gambiae s.s. Insect Molec Biol. 2001;10:19–23. doi: 10.1046/j.1365-2583.2001.00236.x. [DOI] [PubMed] [Google Scholar]
- Santolamazza F, Mancini E, Simard F, Qi Y, Tu Z, Della Torre A. Insertion polymorphisms ofSINE200retrotransposons within speciation islands ofAnopheles gambiaemolecular forms. Malar J. 2008;7:163. doi: 10.1186/1475-2875-7-163. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hunt RH. A cytological technique for the study ofAnopheles gambiaecomplex. Parassitologia. 1973;15:137–139. [PubMed] [Google Scholar]
- Dodge M, Stinson C. Microsoft Office Excel 2007 inside out. Microsoft Press; What’s new in Microsoft Office Excel 2007. [Google Scholar]
- Santolamazza F, Caputo B, Calzetta M, Vicente JL, Mancini E, Petrarca V, Pinto J, della Torre A. Comparative analyses reveal discrepancies among results of commonly used methods forAnopheles gambiaemolecular form identification. Malar J. 2011;10:215. doi: 10.1186/1475-2875-10-215. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ndiath MO, Brengues C, Konate L, Sokhna C, Boudin C, Trape JF, Fontenille D. Dynamics of transmission ofPlasmodium falciparumbyAnopheles arabiensisand the molecular forms M and S ofAnopheles gambiaein Dielmo. Senegal. Malar J. 2008;7:136. doi: 10.1186/1475-2875-7-136. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ndiath MO, Cohuet A, Gaye A, Konate L, Mazenot C, Faye O, Boudin C, Sokhna C, Trape JF. Comparative susceptibility toPlasmodium falciparumof the molecular forms M and S ofAnopheles gambiaeandAnopheles arabiensis. Malar J. 2011;10:269. doi: 10.1186/1475-2875-10-269. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dolo G, Briet OJT, Dao A, Traore SF, Bouare M, Sogoba N, Niare O, Bagayogo M, Sangare D, Teuscher T, Toure YT. Malaria transmission in relation to rice cultivation in the irrigated Sahel of Mali. Acta Trop. 2004;89:147–159. doi: 10.1016/j.actatropica.2003.10.014. [DOI] [PubMed] [Google Scholar]
- Thomson MC, D’ Alessandro U, Bennett S, Connor SJ, Langerock P, Jawara M, Todd J, Greenwood BM. Malaria prevalence is inversely related to vector density in the Gambia, West Africa. Trans R Soc Trop Med Hyg. 1994;88:638–643. doi: 10.1016/0035-9203(94)90204-6. [DOI] [PubMed] [Google Scholar]
- Diurk-Wasser MA, Toure MB, Dolo G, Bagayoko M, Sogoba N, Traore SF, Manoukis N, Taylor CE. Vector abundance and malaria transmission in rice-growing villages in Mali. AmJTrop Med Hyg. 2005;72:725–731. [PMC free article] [PubMed] [Google Scholar]


