Abstract
The latency-associated nuclear antigen (LANA) encoded by Kaposi's sarcoma-associated herpesvirus (KSHV) plays a major role in maintaining latency and is critical for the perpetual segregation of viral episomes to the progeny nuclei of newly divided cells. LANA binds to KSHV terminal repeat (TR) DNA and tethers the viral episomes to host chromosomes through the association of chromatin-bound cellular proteins. TR elements serve as potential origin sites of KSHV replication and have been shown to play important roles in latent DNA replication and transcription of adjacent genes. Affinity chromatography and proteomics analysis using KSHV TR DNA and the LANA binding site as the affinity column identified topoisomerase IIβ (TopoIIβ) as a LANA-interacting protein. Here, we show that TopoIIβ forms complexes with LANA that colocalize as punctuate bodies in the nucleus of KSHV-infected cells. The specific TopoIIβ binding region of LANA has been identified to its N terminus and the first 32 amino acid residues containing the nucleosome-binding region crucial for binding. Moreover, this region could also act as a dominant negative to disrupt association of TopoIIβ with LANA. TopoIIβ plays an important role in LANA-dependent latent DNA replication, as addition of ellipticine, a selective inhibitor of TopoII, negatively regulated replication mediated by the TR. DNA break labeling and chromatin immunoprecipitation assay using biotin-16-dUTP and terminal deoxynucleotide transferase showed that TopoIIβ mediates a transient DNA break on viral DNA. These studies confirm that LANA recruits TopoIIβ at the origins of latent replication to unwind the DNA for replication.
INTRODUCTION
Kaposi's sarcoma-associated herpesvirus (KSHV), also called human herpesvirus 8 (HHV-8), is linked to Kaposi's sarcoma, primary effusion lymphomas (PELs), and multicentric Castleman's disease (MCD) (40, 41, 64). KSHV predominantly causes tumors in individuals that are immunocompromised either by HIV infection or by immunosuppressive drug therapies and is among the leading cause of AIDS-related deaths (12). Like other herpesviruses, KSHV exhibits latent as well as lytic modes of infection and persists predominantly in the latent form, wherein only a subset of proteins are expressed, including the latency-associated nuclear antigen (LANA) (16, 24, 63, 69). LANA is consistently expressed in all forms of KSHV-positive tissues and cell lines (14, 38, 45, 64). However, a small fraction (1 to 5%) of infected cells spontaneously undergo lytic replication (reactivation), which is likely to be essential for maintaining the population of newly infected cells and the development of viral pathogenesis (10, 20, 46, 66). LANA, encoded by open reading frame 73 (ORF73), is a large nuclear protein (222 to 234 kDa) that regulates transcription, cellular signaling, viral DNA replication, and genome maintenance (44, 63). In its lifelong latent state, KSHV genomic DNA exists as a closed circular episome tethered to host chromosomal DNA and is packaged onto nucleosomes with cellular histones (2, 6, 14, 63). This maintenance function is mediated by direct and indirect binding of LANA to the viral DNA and host chromosomes (3, 6, 8, 33, 54).
LANA is a multifunctional protein that plays a central role in maintenance of latency, segregation of episomes, and oncogenesis (26, 63). LANA has been shown to modulate cellular transcription by altering various cellular and viral promoters and transcription factors (1, 4, 8, 51, 62, 65). LANA has also been shown to regulate various proto-oncogene and tumor suppressors at the posttranscriptional level (9, 13, 17, 43, 49, 52, 63). Several of these interactions have crucial effects on proliferation and survival of the infected cells. LANA has been shown to induce chromosome instability and Survivin (a cellular inhibitor of apoptosis) expression to enhance proliferation of KSHV-infected cells (35, 52). LANA interacts with K-bZIP and suppresses lytic origin (ori-Lyt)-dependent DNA synthesis (48). LANA also interacts with Bub-1 and CENP-F to promote long-term persistence of KSHV episome in the infected cell (68). Further, LANA can deregulate host-cell interactions with the immune system and attenuate the antiviral response (29) and inhibits interleukin-4 (IL-4)-mediated STAT6 phosphorylation to regulate apoptosis and maintain latency (7). In addition, LANA maintains KSHV latency by repressing the transcriptional activity of viral immediate-early gene, rta (ORF50), which activates the switch from latency to lytic replication (28, 32).
In addition to modulating the transcription of viral and cellular genes, LANA recruits a number of molecules to regulate replication of the viral episome and the segregation of the newly synthesized genome copies to daughter nuclei by tethering to the host chromosomes (18, 30, 31, 50, 51, 59). LANA has three distinct domains: a proline-rich N-terminal region, important for binding with host chromosomes; a long glutamic acid-rich internal repeat domain; and a carboxy-terminal domain (63). LANA mediates tethering of the KSHV genome by binding to the terminal repeats through its carboxy terminus and associating with components of the human chromatin at its amino terminus, which includes histones and MeCP2 (3, 6, 14, 18, 37). The LANA C-terminal domain binds directly to two LANA-binding sites (LBS) in the KSHV terminal repeats (TR) adjacent to the replication element (RE), which confers DNA replication origin of the TR (3, 18, 22, 23, 55).
The long-term persistence of KSHV depends on its effective interaction with the host cellular machinery. Genome replication and viral gene transcription are consistently dependent on the involvement of a number of cellular processes and appear to be synchronized with the host cell cycle (4, 53, 63). KSHV genomes replicate once per cell cycle during latency and are partitioned perpetually into daughter cells along with host chromosomes during mitosis (3, 4, 63). KSHV-infected PEL cells maintain between 50 and 100 copies of episomes per cell, and the copy number appears to be retained at the same number over time after multiple rounds of cell division (2, 11, 42, 57). Since LANA has no detectable polymerase or helicase activity required for DNA replication, this strongly suggests that replication of the KSHV genome is dependent on enzymes that contain these activities and core components of the cellular replication machinery (42). Association of topoisomerase II (TopoIIβ) with the KSHV TR region was identified by DNA affinity chromatography and proteomics analysis using KSHV TR DNA or ori-Lyt DNA as an affinity ligand and was furthermore demonstrated to be essential for KSHV lytic DNA replication (19, 53, 67).
TopoIIβ is an enzyme that controls and alters the topologic state of DNA during transcription and replication. TopoIIβ has been shown to induce double-stranded (ds) breaks required for regulated transcription/replication (15, 25, 56). Our results show that LANA interacts with TopoIIβ and colocalizes with TopoIIβ as punctuate bodies in the nuclei of KSHV-infected BCBL-1 and JSC-1 cells. The binding domain of LANA to TopoIIβ mapped to its amino-terminal chromosome-binding region. TopoIIβ was shown to play an essential role in LANA-dependent latent DNA replication of TR-containing plasmids, since cells treated with ellipticine, a selective inhibitor of TopoII, negatively regulated replication mediated by the TR. Additionally, we show that TopoIIβ mediates transient DNA breaks on KSHV DNA in order to initiate replication. These studies confirm that LANA recruits TopoIIβ at the origins of latent replication to unwind the DNA for replication.
MATERIALS AND METHODS
Plasmids, antibodies, and cell lines.
pA3F-LANA, pA3F-LANA deletion constructs carrying the Flag-tagged ORF73 amino-terminal domain (amino acids [aa] 1 to 340) and carboxy-terminal domain (aa 940 to 1162), and KSHV TR-containing plasmids were described earlier (58, 59, 61). The GFP-LANA deletion constructs and their mutants carrying Myc-tagged ORF73 aa 1 to 340 and aa 1 to 32 were constructed by PCR amplification from LANA constructs and inserted into pEGFP-myc vector. Lentiviral construct pLVX-LANA-YFP-Flag was constructed similarly by PCR amplification from LANA constructs and inserted into pLVX-AcYFP-C1-Flag vector. Alanine substitutions in green fluorescent protein (GFP) LANA aa 1 to 32 were introduced by PCR mutagenesis with the oligonucleotides described earlier (6). KSHV ori-Lyt plasmid was obtained from the Greg Pari laboratory (University of Nevada, Reno). GFP-TopoIIβ plasmid expressing full-length TopoIIβ isoform fused to GFP in the pEGFP-C3 vector was a generous gift from William T. Beck, University of Illinois at Chicago (39). Myc-tagged proteins were detected using mouse hybridoma 9E10. The following commercially available antibodies were used: rabbit anti-TopoIIβ-H286 (Santa Cruz Biotechnology Inc., CA), rat anti-LANA (Advanced Biotechnologies, Inc.), mouse anti-GAPDH (US Biological), and mouse anti-Flag (Sigma-Aldrich).
The KSHV-negative cell line BJAB and the KSHV-positive cell lines BCBL-1 and JSC-1 were cultured in RPMI 1640 medium supplemented with 10% fetal bovine serum, 2 mM l-glutamine, and penicillin-streptomycin (5 U/ml and 5 μg/ml, respectively). Human embryonic kidney 293 (HEK 293) cells, mouse embryonic fibroblast (MEF) wild-type cells, and TopoIIβ knockdown (TopoIIβ−/−) MEF cells (36) (gift from Yi Lisa Lyu, UMDNJ-Robert Wood Johnson Medical School) were cultured in Dulbecco's modified Eagle's medium (DMEM) supplemented with 10% fetal bovine serum, 2 mM l-glutamine, and penicillin-streptomycin (5 U/ml and 5 μg/ml, respectively). All cell lines were grown at 37°C in a humidified environment supplemented with 5% CO2.
Indirect immunofluorescence microscopy.
BCBL-1 and JSC-1 cells were washed with phosphate-buffered saline (PBS) and spread evenly on coverslips before air drying. Vero cells containing KSHV bacterial artificial chromosome (BAC) (BAC36) were grown on coverslips for 24 h in order to attach and spread. The cells were fixed for 10 min at room temperature with 4% paraformaldehyde followed by permeabilization with 0.2% Triton X-100 in PBS for 10 min at room temperature. For blocking, cells were incubated with PBS containing 0.4% fish skin gelatin and 0.05% Triton X-100. Fixed cells were then incubated with primary antibodies for 1 h at room temperature, washed with PBS, incubated with Alexa Fluor secondary antibodies (Molecular Probes) for 45 min at room temperature, and washed with PBS. Nuclear stain TO-PRO 3 (Molecular Probes) was used to counterstain the nucleus. Images were obtained using a laser scanning confocal microscope (Carl Zeiss, Inc.).
Immunoprecipitation.
For immunoprecipitation, cells were washed with PBS and lysed in NP-40 buffer containing 50 mM Tris-HCl (pH 7.5), 150 mM NaCl, 1 mM EDTA, and 1% NP-40 supplemented with protease inhibitors (1 mM phenylmethylsulfonyl fluoride, 10 μg of pepstatin/ml, 10 μg of leupeptin/ml, and 10 μg of aprotinin/ml). The lysates were centrifuged at high speed to remove the cell debris. The lysates were incubated with protein A beads for 1 h at 4°C to preclear before incubation with specific antibodies. The precleared lysates were then incubated with anti-Flag or anti-LANA antibody overnight at 4°C with rotation followed by capture of the immune complex with protein A and G Sepharose beads at 4°C for 1 h. The resulting immunoprecipitates were collected by centrifugation at 2,000 × g for 3 min at 4°C. The beads were washed four times with 1 ml of ice-cold NP-40 buffer to remove loosely bound proteins. The immunoprecipitated pellets were resuspended in 30 μl of sodium dodecyl sulfate (SDS) protein sample buffer followed by resolving of the protein and Western transfer using standard protocols (Bio-Rad Laboratories). Proteins of interest were detected using specific antibodies followed by incubation with appropriate infrared-dye-tagged (IR680 and IR800) secondary antibodies and scanning with an Odyssey infrared scanner (LI-COR Biosciences, Lincoln, NE).
In vitro binding assay.
Escherichia coli BL21 expressing glutathione S-transferase (GST) fusion proteins was harvested and stored at −80°C until use. Cell pellets were resuspended in binding buffer (50 mM Tris [pH 7.5], 150 mM NaCl, 1 mM MgCl2, 1 mM EDTA [pH 8.0], 1% Nonidet P-40) and lysed with sonication. After removal of cell debris, the supernatants were incubated with glutathione-Sepharose 4B beads (GE Healthcare Life Sciences, Inc.) at 4°C for 1 h. The resin was washed three times with binding buffer, and lysates containing overexpressed protein from 293T cells prepared using the same binding buffer were added after preclearing with protein A Sepharose 4B beads (GE Healthcare Life Sciences, Inc.). The resin was washed four times with binding buffer after 3 h of incubation at 4°C, and the bound proteins were analyzed by sodium dodecyl sulfate-polyacrylamide gel electrophoresis and Western blotting.
Transient replication assay.
Transient replication assay was done as described earlier (60). Briefly, 293L cells in 100-mm dishes were cotransfected with 20 μg of KSHV TR-containing plasmid with either 20 μg of KSHV LANA expression plasmid or with empty vector pA3F/pA3M as filler DNA. At ∼96 h posttransfection, 293L cells were collected by centrifugation (∼5 × 106 cells per sample) for DNA extraction using modified Hirt's method, described earlier (61). Similarly, BCBL-1 cells were transfected with 30 μg of KSHV TR-containing plasmid by electroporation using a Bio-Rad gene pulser at 975-μF capacitance and 210 V. Following electroporation, cells were induced with 20 μg/ml tetradecanoyl phorbol acetate (TPA) and 1 mM sodium butyrate for ∼96 h to induce lytic replication. Cells were collected by centrifugation (∼107 cells per sample) and washed twice with phosphate-buffered saline followed by extraction of DNA using modified Hirt's lysis method (21). Extracted DNA was dissolved in 50 μl of distilled water containing RNase. Ten percent of the extracted DNA from ∼5 × 106 293L cells and ∼107 BCBL-1 cells was digested with EcoRI and the remainder with DpnI and EcoRI to remove the nonreplicated DNA. Digested DNA was separated on 0.8% agarose gel followed by Southern transfer on Hybond N+ membrane (GE Healthcare) and hybridized with 32P-labeled TR probes. Probes specific for the KSHV TR region were synthesized with a New England Biolabs (NEB) random prime kit and signals kit, and signals were detected using a PhosphorImager according to the manufacturer's instructions (Molecular Dynamics, Inc.). Signals were quantified using ImageQuant software (Molecular Dynamics, Inc.). Replicated DNA was determined by analyzing the relative densities of the DpnI-resistant band and normalizing with the respective EcoRI bands in the input DNA.
DNA break labeling and chIP.
In order to detect TopoIIβ-mediated transient-DNA break formation on KSHV TR DNA, we applied a DNA break detection method using biotin-16-dUTP (Roche Applied Science) and terminal deoxynucleotidyl transferase (TdT; Promega, Inc.) in the nucleus (25). Briefly, wild-type (TopoIIβ+/+) and TopoIIβ null (TopoIIβ−/−) MEF NIH 3T3 and HEK293L cells transfected with KSHV TR plasmid were fixed with Streck tissue fixative (STF) (Streck Laboratories) in the presence of 10 mM EDTA for 20 min at room temperature. STF does not cause any DNA damage during the processing (25). The cells were washed with cold Tris-buffered saline (TBS) twice and subsequently resuspended in buffer A (0.25% Triton X-100, 10 mM EDTA, 10 mM HEPES [pH 6.5]) followed by suspension in buffer B (200 mM NaCl, 1 mM EDTA, 10 mM HEPES [pH 6.5]). The nuclei were permeabilized with buffer C (100 mM Tris-HCl [pH 7.4], 50 mM EDTA, 1% Triton X-100) for 30 min at 4°C. The nuclei were sequentially washed with cold PBS, deionized water, and 1× TdT reaction buffer. The DNA breaks were labeled with biotin-16-dUTP using TdT for 30 min at 37°C. After washing the residual biotin-16-dUTP with buffer D (100 mM Tris-HCl [pH 7.4], 150 mM NaCl), nuclei were fixed again with 1% formaldehyde. The nuclear pellet was then resuspended in 100 μl digestion buffer (50 mM Tris-HCl [pH 7.5], 15 mM NaCl, 5 mM KCl, 3 mM MgCl2, 1 mM CaCl2, 10 mM NaHSO4, 0.25 M sucrose, 0.15 mM spermine, 0.5 mM spermidine, and 0.15 mM β-mercaptoethanol). Chromatin immunoprecipitation (ChIP) was performed without sonication on MEFs and with sonication on HEK 293L cells using streptavidin-conjugated magnetic Sepharose beads (GE Healthcare Life Sciences, Inc.).
Flow cytometry.
HEK 293 cells were transfected with KSHV TR plasmid along with either pA3F LANA or the empty vector pA3F. Cells were pretreated with ellipticine for 1 h before transfection, and the treatments continued posttransfection. After 24 h, cells were harvested and fixed in cold 70% ethanol for 30 to 60 min. The fixed cells were washed twice by 1× PBS and stained with propidium iodide (PI) as described previously (34, 47). Data were acquired on FACSCalibur equipped with CellQuest Pro software and analyzed using FlowJo software.
Real-time PCR.
Quantitative real-time PCR was performed in a total volume of 20 μl, including 10 μl of SYBR green PCR 2× Master mix (Applied Biosystems) and 0.5 μM each KSHV TR primer (forward, 5′-GGGGGACCCCGGGCAGCGAG-3, and reverse, 5-GGCTCCCCCAAACAGGCTCA-3) flanking TR nucleotides 677 to 766. The ampicillin gene region was amplified with forward (5′-GTAGATAACTACGATACGGGAGGG-3′) and reverse (5′-GCGAACTACTTACTCTAGCTTCCC-3′) primers. Purified DNA samples of the ChIP fraction and the input DNA samples were amplified on an ABI StepOne plus real-time PCR machine (Applied Biosystems). Relative copies of immunoprecipitated TR were calculated by the ΔCT method.
RESULTS
KSHV LANA associates with TopoIIβ in KSHV-positive cells.
KSHV establishes a lifelong latent infection after primary infection in the target cells. LANA, a nuclear protein, is expressed consistently in all the infected cells in relatively large amounts and is solely responsible for maintaining the viral genome into the dividing tumor cells. Besides tethering, LANA recruits the host cellular replication complex to replicate the terminal repeat-containing plasmids (55, 59, 63). DNA affinity column as well as LANA pulldown assays identified topoisomerase IIβ as one of the LANA-interacting proteins (26, 58). To identify whether LANA interacts with TopoIIβ in KSHV-positive cells, a coimmunoprecipitation (co-IP) assay was performed using the KSHV-positive cell lines BCBL-1 and JSC-1. Immunoprecipitation with anti-LANA antibody and subsequent detection with anti-TopoIIβ antibody showed that LANA precipitated TopoIIβ from the KSHV-positive BCBL-1 and JSC-1 cells (Fig. 1A and C, respectively). To further analyze the specificity of this interaction, a reverse co-IP assay was performed using anti-TopoIIβ antibody on the KSHV-positive cell lines BCBL-1 and JSC-1. Immunoprecipitation and subsequent detection with anti-TopoIIβ and LANA antibodies showed that LANA precipitated with TopoIIβ from the KSHV-positive cells BCBL-1 and JSC-1 (Fig. 1B and D), respectively. Similarly, co-IP analysis with anti-Flag antibody from BJAB cells expressing either yellow fluorescent protein (YFP)-Flag or YFP-LANA-Flag showed coimmunoprecipitation of TopoIIβ from the BJAB cells expressing YFP-LANA-Flag (Fig. 1E, lane 4) but not from YFP-Flag (Fig. 1E, lane 3). Expressions and immunoprecipitations of LANA and YFP were detected with anti-Flag antibody. YFP-LANA-Flag also showed a band corresponding to the YFP-Flag due to cleavage of YFP from the fusion protein (Fig. 1E, lanes 2 and 4). These results show that TopoIIβ forms a complex with LANA in KSHV-infected cells as well as in cells expressing exogenous LANA.
Fig 1.

KSHV LANA associates with TopoIIβ in KSHV-positive cells. (A to D) Coimmunoprecipitation assays were performed using 25 million KSHV-positive cells (BCBL-1) with anti-LANA antibody (A); using 25 million BCBL-1 cells with anti-TopoIIβ antibody and subsequent detection with anti-TopoIIβ and LANA antibodies (lane 4) (B); using 25 million KSHV-positive cells (JSC-1) with anti-LANA antibody (C); using 25 million JSC-1 cells with anti-TopoIIβ antibody and subsequent detection with anti-TopoIIβ and LANA antibodies (lane 4) (D). (E) Co-IP analysis with anti-FLAG antibody from 25 million BJAB cells expressing either YFP-Flag (Y-Flag) or LANA-YFP-Flag (Y-LFlag) and subsequent detection with anti-TopoIIβ antibody. TopoIIβ was found to coimmunoprecipitate with exogenously supplied LANA (lane 4).
LANA colocalizes with TopoIIβ in KSHV-positive cells.
To further confirm the association of LANA and TopoIIβ, we performed an immunofluorescence assay (IFA) on KSHV-positive BCBL-1 and JSC-1 cells. These cells were stained with rat anti-LANA and rabbit anti-TopoIIβ antibodies followed by detection of LANA with goat anti-rat Alexa Fluor 488 (green) and chicken anti-rabbit Alexa Fluor 594 (red). LANA showed a distinct punctate pattern in both of the PEL cells as detected earlier (14). TopoIIβ proteins, shown in red, localized primarily in the nuclei of the infected cells and were in the same nuclear compartment as LANA, thus suggesting colocalization (Fig. 2A, detected as a yellow signal in the merge panels). Nuclei detected by TO-PRO 3 staining showed that the colocalization signals were in the nuclei. A differential interference contrast (DIC) image showed that the cells were healthy and had distinct nuclei. Additionally, IFA for the localization of LANA and TopoIIβ on Vero cells harboring KSHV BAC36 showed punctate LANA staining and colocalization with TopoIIβ in the nucleus (Fig. 2B). Since Vero BAC36 cells have GFP, we were unable to use TO-PRO 3 to localize the nucleus, but a DIC image showed that colocalization of LANA and TopoIIβ was in the nucleus (Fig. 2B). These localization assays confirm that LANA and TopoIIβ are in the same nuclear compartment of the infected cells and may have a role in latent replication of the viral genome.
Fig 2.

LANA colocalizes with TopoIIβ in KSHV-positive cells. KSHV-positive cells BCBL-1 and JSC-1 (A) and Bac36 Vero cells (B) were stained with rat anti-LANA and rabbit anti-TopoIIβ antibodies. LANA is shown in green and TopoIIβ in red. Nuclear stain TO-PRO 3 is shown in blue. LANA and TopoIIβ colocalize in the nucleus as punctate bodies. DIC images were used to show the cells' morphology.
The amino terminus of LANA interacts with TopoIIβ.
To identify the distinct domain of LANA responsible for TopoIIβ interaction, we transiently expressed either full-length LANA (LANA-FL) or truncation mutants, one expressing the amino terminus and the other the carboxy terminus tagged with Flag epitope, along with TopoIIβ in HEK 293T cells. Immunoprecipitation analysis with anti-Flag antibody and subsequent detection with anti-TopoIIβ antibody showed that TopoIIβ was coimmunoprecipitated with full-length LANA as well as the amino terminus of LANA but not with the carboxy terminus (Fig. 3A, lanes 6 and 7, and B, lane 5). Relative binding of LANA-FL and LANA-N terminus with TopoIIβ showed that LANA-FL had stronger binding affinity, which could possibly be due to the involvement of additional factors recruited with the full-length LANA (Fig. 3A, compare lanes 6 and 7). Empty flag vector with TopoIIβ did not show any precipitation of TopoIIβ, confirming the specificity of the interaction (Fig. 3A, lanes 1 and 5, and B, lanes 1 and 4). Expression of LANA and its truncation mutants in the lysates (input) and immunoprecipitated lanes are marked with red arrows. These interactions were consistently observed with endogenous TopoIIβ. HEK 293T cells were transfected with either full-length LANA or truncation mutants expressing amino terminus, LANA amino acid residues 1 to 32, and the carboxy terminus tagged with Flag epitope. Immunoprecipitation analysis with anti-Flag antibody and subsequent detection with anti-TopoIIβ and LANA antibodies showed that endogenous TopoIIβ coimmunoprecipitated with full-length LANA as well as the amino terminus and aa 1 to 32 of LANA but not with the carboxy terminus (Fig. 3C, lanes 7, 8, and 9). Further to ensure the specificity of this interaction, cell lysates were pretreated with micrococcal nuclease prior to immunoprecipitation analysis with anti-Flag antibody. Results showed that TopoIIβ interacts specifically with full-length LANA and its amino terminus irrespective of nuclease treatment (Fig. 3D, lanes 5 and 6). These results confirmed that LANA directly interacts with TopoIIβ and that the internucleosomal DNA does not mediate their association.
Fig 3.

The amino terminus of LANA interacts with TopoIIβ. (A) Twenty million HEK 293T cells were transfected with flag epitope-tagged pA3F empty vector, pA3F LANA, pA3F LANA-N, and pA3F LANA-C along with GFP-TopoIIβ. At 36 h posttransfection, cells were harvested and immunoprecipitated with anti-Flag antibody and subsequently detected with anti-TopoIIβ antibody (lanes 6 and 7). (B) Similarly, HEK 293T cells were transfected with pA3F empty, pA3F LANA-N, and pA3F LANA-C along with GFP-TopoIIβ. At 36 h posttransfection, cells were harvested and immunoprecipitated with anti-Flag antibody and subsequently detected with anti-TopoIIβ antibody. TopoIIβ specifically interacts with full-length LANA and the N-terminal region of LANA (panel A, lane 7, and panel B, lane 5). (C) HEK 293T cells were transfected with pA3F empty, pA3F LANA, pA3F LANA-N, LANA 1 to 32, and pA3F LANA-C. At 36 h posttransfection, cells were harvested and immunoprecipitated with anti-Flag antibody and subsequently detected with anti-TopoIIβ and LANA antibodies. Endogenous TopoIIβ specifically interacts with full-length LANA and the N-terminal region of LANA (lanes 7, 8, and 9). (D) HEK 293T cells were transfected with pA3F empty, pA3F LANA, and pA3F LANA-N along with GFP-TopoIIβ. At 36 h posttransfection, cells were harvested and the lysate was incubated with 2,000 U micrococcal nuclease (NEB) for 30 min at 37°C and immunoprecipitated with anti-Flag antibody and subsequently detected with anti-TopoIIβ antibody. TopoIIβ interacts specifically with full-length LANA and its amino terminus irrespective of nuclease treatment (lanes 5 and 6). (E) Micrococcal nuclease-digested DNA on ethidium bromide (EtBr)-stained agarose gel.
The residues localized between aa 1 and 32 of the amino terminus of LANA mediate association with TopoIIβ.
To determine the domains of LANA responsible for TopoIIβ interaction, we made further truncations of the amino terminus of LANA into expression constructs containing aa 1 to 32, 1 to 150, 1 to 250, 33 to 275, and 33 to 340, tagged with Myc epitope (Fig. 4A). Transient expression of these constructs along with TopoIIβ in HEK 293T cells followed by immunoprecipitation analysis with anti-Myc antibody and subsequent detection with anti-TopoIIβ antibody showed that both LANA truncation constructs aa 1 to 150 and aa 1 to 250 were able to bind with TopoIIβ (Fig. 4B, lanes 6 and 7). LANA mutants lacking aa 1 to 32 (constructs aa 33 to 275 and aa 33 to 340) were unable to precipitate TopoIIβ (Fig. 4B, lanes 5 and 8) suggesting that LANA binding domain to TopoIIβ lies between amino acid residues 1 and 32. In order to determine the binding of TopoIIβ with LANA aa 1 to 32, we cloned aa 1 to 32 in frame with GFP and Myc epitope tag and generated a stably expressing cell line in BJAB. An immunoprecipitation assay with anti-Myc antibody to precipitate LANA aa 1 to 32 showed coprecipitation of TopoIIβ (Fig. 4C, lane 5). Vector control with GFP-Myc did not show precipitation of TopoIIβ, suggesting specific association of LANA aa 1 to 32 with TopoIIβ (Fig. 4C, lane 4). In an attempt to identify the specific residues of LANA aa 1 to 32 associating with TopoIIβ, we used LANA alanine substitution mutants between residues 5 and 15, which were shown to associate with host chromatin and be important for replication (5, 6). BJAB, stably expressing GFP-Myc LANA aa 1 to 32 with substitution mutation at aa 5 to 15 (M5), did not show precipitation of TopoIIβ, suggesting that residues 5 to 15 are critical for TopoIIβ recruitment (Fig. 4C, lane 6).
Fig 4.

The aa 1 to 32 region at the amino terminus of LANA is responsible for the association with TopoIIβ. (A) Schematic showing the truncations of LANA N-terminal region. (B) Twenty million HEK 293T cells were transfected with pEGP-Myc LANA aa 33 to 275, pEGFP-Myc LANA aa 1 to 250, pEGFP-Myc LANA aa 1 to 150, and pEGFP-Myc LANA aa 33 to 340 with GFP-TopoIIβ. At 36 h posttransfection, cells were harvested and immunoprecipitated with anti-Myc antibody (9E10) and subsequently detected with anti-TopoIIβ antibody (lanes 6 and 7). (C) Twenty-five million BJAB cells stably expressing pEGP-Myc empty, pEGP-Myc LANA aa 1 to 32, and pEGP-Myc LANA aa 1 to 32 with substitutions at aa 5 to 15 (5aa15) were harvested and immunoprecipitated with anti-Myc antibody (9E10) and subsequently detected with anti-TopoIIβ antibody (lane 5). (D) HEK 293 T cells were transfected with pEGFP-Myc empty, pEGFP-Myc LANA aa 1 to 32, pEGFP-Myc LANA aa 1 to 32 5aa15, pEGFP-Myc LANA aa 1 to 340, and pEGFP-Myc LANA aa 1 to 340 5aa15 with GFP-TopoIIβ. At 36 h posttransfection, cells were harvested and immunoprecipitated with anti-Myc antibody (9E10) and subsequently detected with anti-TopoIIβ antibody (lanes 7 and 9). (E) In vitro GST binding: GST, GST-LANA aa 1 to 32, and GST-LANA aa 1 to 32 5aa15 fusion proteins were expressed in E. coli, purified with glutathione-Sepharose beads, and incubated with TopoIIβ cell lysate prepared from HEK 293T cells transfected with GFP-TopoIIβ. The aa 1 to 32 region of LANA interacted with TopoIIβ (lane 2).
We further confirmed the association of TopoIIβ with LANA through residues 5 to 15 in the context of the entire amino terminus region (aa 1 to 340) and its alanine substitution mutant (aa 1 to 340 with mutations at aa 5 to 15) along with the aa 1 to 32 region of LANA. Immunoprecipitation with anti-Myc antibody to precipitate LANA and its mutants showed that TopoIIβ coimmunoprecipitated with LANA aa 1 to 340 and aa 1 to 32 but not with alanine substitution mutants of both of these truncations (Fig. 4D, compare lanes 7 and 9 with 8 and 10). These experiments indicated that the residues 5 to 15 of LANA are required for recruiting TopoIIβ.
The interaction of LANA residues 1 to 32 with TopoIIβ was further confirmed by an in vitro binding assay. Bacterially expressed GST-tagged LANA aa 1 to 32 and its alanine substitution mutant M5 (where residues 5 to 15 were replaced by alanine) were used for the binding assay with transiently overexpressed TopoIIβ protein from HEK 293T cells. GST protein prepared from empty vector served as control. As shown in Fig. 4E, bacterially expressed GST-LANA aa 1 to 32 efficiently precipitated TopoIIβ, whereas LANA mutant M5 did not show any binding to TopoIIβ and showed results similar to those of the GST control lane (Fig. 4E).
LANA aa 1 to 32 have been previously shown to be responsible for the association of LANA with host nucleosome through interaction with cellular histones (6). Therefore, we wanted to map the residues of aa 5 to 15 specifically involved in the TopoIIβ interaction with LANA. To this end, we made alanine substitution mutations of LANA in sets of three amino acids in both aa 1 to 32 and aa 1 to 340 LANA truncation mutants (Fig. 5A), as follows: 5GMR7 changed to AAA (M1), 8LRS10 to AAA (M2), 11GRS13 to AAA (M3), 14TG15 to AA (M4), and all residues between 5 and 15 changed to alanine (M5). Transient expression of these LANA mutants along with TopoIIβ in HEK 293 cells followed by immunoprecipitation analysis with anti-Myc to immunoprecipitate LANA mutants identified residues 8 to 15 as being essential for binding with TopoIIβ, since mutants M2 through M4 (8LRS10 to AAA, 11GRS13 to AAA, and 14TG15 to AA) failed to coimmunoprecipitate TopoIIβ (Fig. 5B and C). These results were consistent with both LANA truncation (aa 1 to 32 and aa 1 to 340) mutants.
Fig 5.

The aa 1 to 32 region of the amino terminus of LANA is responsible for TopoIIβ association. (A) Schematic showing the alanine substitution mutations of aa 1 to 32 of the LANA N-terminal region. (B) Twenty million HEK 293T cells were transfected with pEGFP-Myc empty, pEGFP-Myc LANA aa 1 to 32, pEGFP-Myc LANA aa 1 to 340, pEGFP-Myc LANA aa 1 to 340 M1, pEGFP-Myc LANA aa 1 to 340 M2, pEGFP-Myc LANA aa 1 to 340 M3, pEGFP-Myc LANA aa 1 to 340 M4, and pEGFP-Myc LANA aa 1 to 340 M5 with GFP-TopoIIβ. At 36 h posttransfection, cells were harvested and immunoprecipitated with anti-Myc antibody (9E10) and subsequently detected with anti-TopoIIβ antibody. (C) Similarly, HEK 293T cells were transfected with pEGFP-Myc empty, pEGFP-Myc LANA aa 1 to 32, pEGFP-Myc LANA aa 1 to 340, pEGFP-Myc LANA aa 1 to 32 M1, pEGFP-Myc LANA aa 1 to 32 M2, pEGFP-Myc LANA aa 1 to 32 M3, pEGFP-Myc LANA aa 1 to 32 M4, pEGFP-Myc LANA aa 1 to 32 M5 with GFP-TopoIIβ. At 36 h posttransfection, cells were harvested and immunoprecipitated with anti-Myc antibody (9E10) and subsequently detected with anti-TopoIIβ antibody. The residues 8 to 15 of LANA are crucial for binding TopoIIβ (B and C).
TopoIIβ is required for KSHV latent DNA replication.
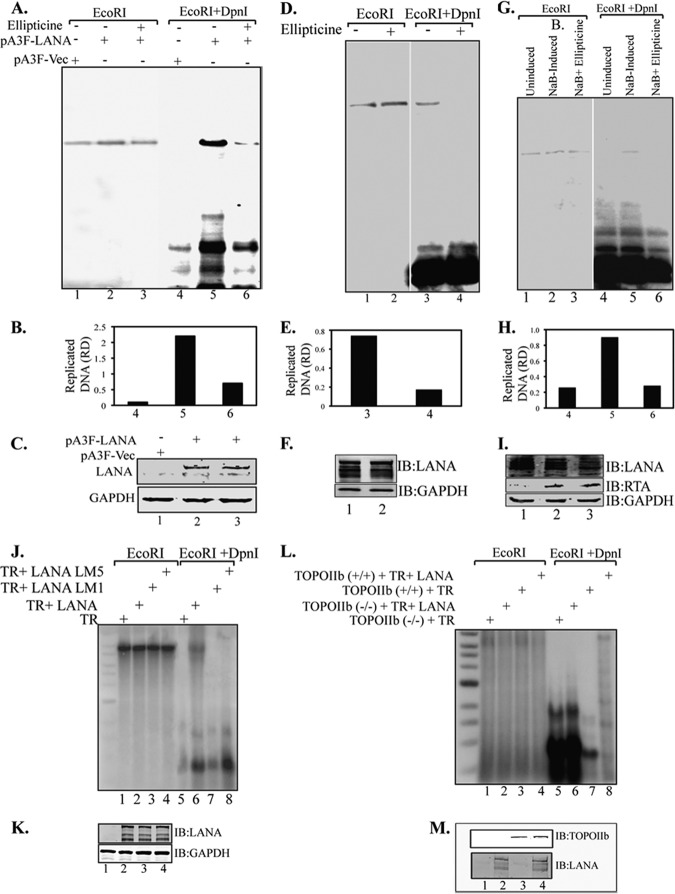
To determine whether TopoIIβ is important for the latent replication of KSHV DNA, a transient replication assay with the KSHV TR was conducted in the presence of ellipticine, a selective inhibitor of TopoIIβ. HEK 293L cells were transfected with TR plasmid along with Flag epitope-tagged KSHV-LANA, pA3F-LANA, or empty vector pA3F. Ninety-six hours posttransfection, cells were harvested and subjected for DNA extraction by modified Hirt's procedure. Extracted DNA was subjected to restriction digestion with either EcoRI alone (to linearize) or with EcoRI plus DpnI (to remove nonreplicated plasmid DNA) followed by the detection of replicated DNA in a Southern blot assay using 32P-labeled TR probe. Hybridization signals were detected by PhosphorImager and analyzed using ImageQuant software (Molecular Dynamics, Inc.). Results showed that KSHV latent DNA replication is TopoIIβ dependent, since treatment of cells with ellipticine effectively reduced latent DNA replication (Fig. 6A). LANA-expressing cells treated with ellipticine showed a faint DpnI-resistant band, whereas the untreated cells showed significantly larger amounts of replicated DNA (Fig. 6A, compare lanes 5 and 6). Relative amounts of the replicated DNA determined based on the densities of the DpnI-resistant band normalized with input lanes suggest effective inhibition of replication by ellipticine (Fig. 6B). Cells without LANA expression did not show any DpnI-resistant band, as expected. These results suggest that TopoIIβ is essential for latent DNA replication. The levels of LANA were determined by anti-Flag Western blotting in ellipticine-treated as well as nontreated cells to ensure that ellipticine did not have any adverse effect on LANA expression (Fig. 6C).
Fig 6.
TopoIIβ is required for KSHV latent DNA replication. (A) HEK 293L cells were transfected with TR plasmid along with pA3F-LANA or empty vector pA3F. Cells were treated with ellipticine 24 h posttransfection. Ninety-six hours posttransfection, cells were harvested, and the DNA extracted by Hirt's procedure was subjected to Southern blotting with a TR probe after digestion. Cells expressing LANA without ellipticine treatment show a prominent DpnI-resistant replicated DNA (lane 5), whereas cells treated with ellipticine showed a faint band (lane 6). (B) Quantitation of the replicated DNA based on the relative density (RD). (C) Expression of LANA and GAPDH. (D) BCBL-1 cells transfected with TR plasmid were treated with ellipticine 24 h posttransfection. Ninety-six hours posttransfection, cells were harvested, and the DNA extracted by Hirt's procedure was subjected to Southern blotting with TR probe after digestion. Ellipticine effectively blocked KSHV latent DNA replication in KSHV-positive cells (lane 4). (E) Quantitation of the replicated DNA. (F) Expression levels of LANA and GAPDH. (G) BCBL-1 cells transfected with ori-Lyt plasmid were first induced with 3 mM sodium butyrate and 20 μg/ml TPA followed by treatment with ellipticine 24 h posttransfection. Ninety-six hours posttransfection, cells were harvested, and the DNA extracted by Hirt's procedure was subjected to Southern blotting with ori-Lyt probe after digestion. Ellipticine blocked KSHV lytic DNA replication (lane 6). (H) Quantitation of the replicated DNA. (I) Western blots showing the expression of LANA, RTA, and GAPDH. (J) LANA mutants M1 and M5 do not support KSHV latent DNA replication (lanes 7 and 8): HEK 293L cells were transfected with TR plasmid along with pA3M-LANA, pA3M-LANA-M1, or pA3M-LANA-M5. Ninety-six hours posttransfection, cells were harvested, and the DNA extracted by Hirt's procedure was subjected to Southern blotting with TR probe after digestion. (K) Expression of LANA and GAPDH. (L) TopoIIβ is required for KSHV latent DNA replication: wild-type (TopoIIβ+/+) and TopoIIβ knockout (TopoIIβ−/−) MEF NIH 3T3 cells were transfected with TR plasmid along with empty vector pA3F or pA3F-LANA. Ninety-six hours posttransfection, cells were harvested, and the DNA extracted by Hirt's procedure was subjected to Southern blotting with TR probe after digestion. (M) Expression of LANA and TopoIIβ in the cells used for replication assays.
Similar results were observed in transient replication assays conducted on KSHV-positive BCBL-1 cells in the presence of ellipticine (Fig. 6D). BCBL-1 cells were transfected with KSHV TR plasmids, and replication assays were performed in the presence and absence of ellipticine to determine whether ellipticine was able to block replication of TR plasmids in KSHV-infected cells. Ninety-six hours posttransfection, cells were harvested, followed by the extraction of DNA using modified Hirt's procedure. Extracted DNA was digested with either EcoRI (to linearize) or with DpnI and EcoRI to identify the replicated plasmids after Southern transfer as mentioned above. Cells treated with ellipticine did not show any DpnI-resistant band, whereas the untreated cells showed a prominent DpnI-resistant band, confirming that ellipticine can effectively block TR DNA replication (Fig. 6D and E, compare lanes 1 and 2). Expression of LANA was not affected by ellipticine treatment (Fig. 6F, IB: LANA). As a control, a similar transient replication assay was conducted on induced BCBL-1 cells transfected with ori-Lyt DNA. Induced BCBL-1 cells, containing ori-Lyt DNA, were treated with ellipticine following the extraction of DNA using modified Hirt's procedure and detection of replicated DNA copies in a Southern blot. An ori-Lyt-specific 32P-labeled probe was used for the detection of replicated plasmid. Consistent with the previous report, ori-Lyt plasmid did not replicate in uninduced cells (Fig. 6G, lane 1). Induction of these cells with sodium butyrate showed replication of ori-Lyt plasmids, which was blocked by ellipticine treatment as reported earlier (Fig. 6G and H, compare lanes 2 and 3). This suggests that TopoIIβ is required for both latent and lytic KSHV DNA replication. The levels of LANA and RTA were determined in a Western blot assay to show that the treatment with sodium butyrate induced RTA expression, which is required for the lytic DNA replication (Fig. 6I).
Additionally, transient replication assays with the KSHV TR along with full-length LANA and its mutants, LANA 5GMR7-AAA and LANA with residues 5 to 15 replaced by alanine, showed that the residues 5 to 15 are crucial for KSHV latent DNA replication, which may be due to the loss of TopoIIβ binding to LANA. This was done by transfecting 293L cells with equal amounts of TR plasmids with either full-length Myc-tagged LANA (pA3M-LANA) or its mutants, LANA 5GMR7-AAA and LANA with residues 5 to 15 changed to alanine. Ninety-six hours posttransfection, cells were harvested and the DNA extracted by Hirt's procedure was digested with either EcoRI or DpnI and EcoRI to detect the replicated copies as mentioned above. Extracted DNA from cells expressing pA3M LANA showed a prominent DpnI-resistant band indicating replicated DNA (Fig. 6J, lane 6), whereas the mutants LANA 5GMR7-AAA and LANA with residues 5 to 15 changed to alanine were not able to support replication and hence failed to show any DpnI-resistant band (Fig. 6J, lanes 7 and 8). Expression of LANA detected in the Western blot showed comparable levels of protein expression (Fig. 6K).
To further substantiate the requirement of TopoIIβ for KSHV latent DNA replication, we conducted a transient replication assay with the KSHV TR on TopoIIβ knockout mouse embryonic fibroblast cells (TopoIIβ−/−) MEF NIH 3T3. TopoIIβ knockout MEF NIH 3T3 and wild-type cells were transfected with KSHV TR plasmid along with LANA expression plasmid. Ninety-six hours posttransfection, cells were harvested, and the DNA extracted by Hirt's procedure was digested with either EcoRI or DpnI and EcoRI to detect replicated DNA copies using 32P-labeled TR probe as mentioned above. TopoIIβ knockout MEF NIH 3T3 cells did not show any detectable DpnI-resistant band in the presence of LANA (Fig. 6L, lane 6). However, MEF NIH 3T3 wild-type cells expressing LANA showed a DpnI-resistant band, indicating replication of transfected plasmid (Fig. 6L, lane 8). These data suggest that TopoIIβ is essential for a LANA-dependent KSHV latent DNA replication. Expression of LANA detected in Western blotting showed comparable levels of protein expression (Fig. 6M).
The minimal binding region of LANA can act as a dominant negative and disrupts LANA-TopoIIβ interaction.
To further understand the biological significance of TopoIIβ recruitment by LANA, we used the minimal TopoIIβ binding region of LANA (construct 1 to 32) as a dominant negative in both immunoprecipitation and transient replication assays. To study the effect of LANA 1 to 32 as a dominant negative in coimmunoprecipitation assay, HEK 293T cells were transfected with GFP-tagged TopoIIβ and the amino terminus of LANA aa 1 to 340 tagged with Flag epitope along with two different amounts (20 μg and 40 μg) of LANA aa 1 to 32 (Myc tagged) as dominant negative. Cells were transfected with TopoIIβ along with empty vector, pA3F, pA3F-LANA C terminus, LANA aa 1 to 32, the M5 mutant (alanine substitutions at aa 5 to 15), full-length LANA (pA3F-LANA), and pA3F-LANA N terminus (aa 1 to 340) as negative and positive controls. Immunoprecipitation analysis with anti-Flag antibody showed coimmunoprecipitation of TopoIIβ with LANA full-length and amino-terminal domain as seen earlier (Fig. 3A, lanes 2 and 4). Interestingly, overexpression of LANA aa 1 to 32 was able to adversely affect the interaction of LANA-N with TopoIIβ and thus acted as dominant negative (Fig. 7A, lanes 5 and 6). Cells expressing increased amounts of LANA aa 1 to 32 showed a dose-dependent response in abolishing the association of TopoIIβ with LANA-N (Fig. 7A, compare lanes 4, 5, and 6). However, overexpression of LANA aa 1 to 32 with alanine substitutions at aa 5 to 15 had no effect on the association of TopoIIβ with LANA-N and was similar to LANA-FL (Fig. 7A, lanes 2 and 7). Overexpression of LANA aa 1 to 32 may have slightly reduced the expression of LANA-N (Fig. 7A, compare lanes 4 and 5); however, the immunoprecipitation of LANA-N was not affected (Fig. 7A, IB: FLAG, LANA-N). Subsequent increase of LANA aa 1 to 32 expression did not affect LANA-N expression but drastically reduced TopoIIβ association with LANA-N.
Fig 7.
The minimal binding region of LANA acts as a dominant negative to disrupt LANA-TopoIIβ interaction and replication. (A) Twenty million HEK 293T cells were transfected with GFP-TopoIIβ along with pA3F, pA3F-LANA, pA3F-LANA C terminus, pA3F-LANA N, and pA3F-LANA N with two different amounts (20 μg and 40 μg) of GFP-Myc-LANA aa 1 to 32 and 40 μg GFP-Myc-LANA aa 1 to 32 mutant M5 (with alanine substitutions at aa 5 to 15). At 36 h posttransfection, cells were harvested and immunoprecipitated with anti-flag antibody to precipitate LANA and its mutants, followed by detection of TopoIIβ as the coimmunoprecipitated proteins. LANA aa 1 to 32 was found to interfere with LANA TopoIIβ interaction (lanes 5 and 6), whereas transfection of LANA aa 1 to 32 mutant M5 did not interfere with LANA and TopoIIβ interaction (lane 7). Expression of increased amounts of LANA aa 1 to 32 further decreased TopoIIβ binding (lane 6). Expressions of LANA aa 1 to 32 were detected by anti-Myc immunoblot (asterisk). (B) HEK 293L cells were transfected with TR plasmid along with GFP-Myc-LANA 1 to 32, pA3F-LANA, or pA3F-LANA with two different amounts (20 and 40 μg) of GFP-Myc-LANA 1 to 32 and pA3F-LANA along with 40 μg of GFP-Myc-LANA 1 to 32 M5. Ninety-six hours posttransfection, cells were harvested, and the DNA extracted by Hirt's procedure was subjected to Southern blotting with TR probe after digestion. LANA aa 1 to 32 interfere with KSHV latent DNA replication as detected by a reduced level of DpnI-resistant band (compare lanes 9 and 10). Increasing expression of LANA aa 1 to 32 further reduced the replication of TR plasmid (compare lanes 11 and 10), however; mutant 5 (M5) of LANA aa 1 to 32 was unable to suppress replication (compare lanes 11 and 12). (C) Quantitation of the replicated DNA. (D) Expression levels of LANA and GFP-Myc-LANA aa 1 to 32 and GFP-Myc-LANA aa 1 to 32 M5. (E) Schematic showing DNA break labeling and ChIP assay procedure. Wild-type (TopoIIβ+/+) and TopoIIβ knockdown (TopoIIβ−/−) MEF NIH 3T3 cells were transfected with KSHV TR plasmid either with pA3F-LANA or with empty vector pA3F. The nuclei were subsequently labeled with biotin-16-dUTP using terminal deoxynucleotidyl transferase (TdT) and subjected to chromatin immunoprecipitation (ChIP). (F) Quantitative real-time PCR was done on the input and ChIP DNA samples using KSHV TR-specific primers. Relative copies of the dUTP-labeled ChIP DNA were determined as the ratio of cells without LANA to cells with LANA in TopoIIβ−/− and TopoIIβ+/+ cells to determine the effect of LANA on dsDNA breaks. TopoIIβ+/+ cells showed significant increase in the presence of LANA. (G) DNA break labeling and ChIP assay was performed on HEK 293L cells transfected with KSHV TR-containing plasmids with either pA3F-LANA or the empty vector pA3F. One set of cells with LANA with TR was treated with 5 μM ellipticine. dUTP-labeled DNAs were determined in a real-time PCR using primer for TR (black bar) and vector backbone (gray bar). An increase in dUTP-labeled ChIP DNA in the TR region but not in the ampicillin region (vector) in LANA-expressing cells suggests the existence of a dsDNA break in the TR region. Ellipticine treatment blocked dUTP incorporation, suggesting a TopoII-mediated dsDNA break. (H) At 24 h posttransfection, 293L cells were harvested for flow cytometry and cell cycle analysis.
Similarly, transient replication assays using LANA aa 1 to 32 also revealed that the expression of LANA aa 1 to 32 as a dominant negative could effectively reduce KSHV latent DNA replication (Fig. 7B). For the transient replication assay, HEK 293L cells were transfected with equal amounts of KSHV TR plasmid with either empty vector, LANA aa 1 to 32, full-length LANA, or full-length LANA with two different amounts (20 μg and 40 μg) of LANA aa 1 to 32 as dominant negative. As a control, 40 μg of LANA aa 1 to 32 mutant M5 (alanine substitutions) was transfected with TR and full-length LANA. Ninety-six hours posttransfection, cells were harvested, and the DNA extracted by Hirt's procedure was subjected to either EcoRI or DpnI and EcoRI to detect replicated TR plasmid copies in a Southern blot assay using a 32P-labeled TR probe. Cells transfected with TR with full-length LANA showed a prominent DpnI-resistant band as expected (Fig. 7B, lane 9). LANA aa 1 to 32 did not show any detectable replication, suggesting that additional factors besides TopoIIβ are required for replication (Fig. 7B, lane 8). Interestingly, cells transfected with TR and full-length LANA along with LANA aa 1 to 32 showed decreased replication of TR plasmids (Fig. 7B, compare lanes 9 and 10). Additionally, increasing amounts of LANA aa 1 to 32 progressively reduced LANA-dependent KSHV latent DNA replication (Fig. 7B, compare lanes 10 and 11). Interestingly, cells transfected with M5 of LANA aa 1 to 32 with TR and full-length LANA did not show any reduction in the replication of TR plasmid (Fig. 7B, lane 12), confirming that residues 5 to 15 are critical for recruiting TopoIIβ at the site of replication initiation. A quantitation of the replicated DNA, based on the relative density normalized to the respective EcoRI band, showed a dose-dependent response of LANA aa 1 to 32 in blocking DNA replication (Fig. 7C). These data clearly show that LANA aa 1 to 32 could act as a dominant negative, specifically disrupting the association of LANA with TopoIIβ. Moreover, LANA aa 1 to 32 is crucial for the association of LANA with TopoIIβ and LANA-dependent KSHV latent DNA replication. Expression of LANA detected in a Western blot showed comparable levels of protein expression (Fig. 7D).
TopoIIβ makes dsDNA breaks on the KSHV genome.
TopoIIβ plays a central role in altering the degree of supercoiling of double-stranded DNA molecules (15). TopoIIβ cleaves both strands of the DNA helix simultaneously and is essential in the separation of intertwined and supercoiled DNA strands during replication and transcription (25). It has been shown that TopoIIβ-mediated dsDNA breaks are also required for transcription (15, 25). We performed a DNA break labeling and ChIP assay, shown in a schematic diagram (Fig. 7E), to detect whether TopoIIβ could mediate nicks on KSHV TR DNA. Wild-type (TopoIIβ+/+) and TopoIIβ knockdown (TopoIIβ−/−) MEF NIH 3T3 cells were transfected with TR-containing plasmid either with pA3F-LANA or with empty vector pA3F. The nuclei were subsequently labeled with biotin-16-dUTP using terminal deoxynucleotidyl transferase (TdT) to label the ends of nicked DNA and subjected to chromatin immunoprecipitation using streptavidin. Quantitative real-time PCR was done on the input and on ChIP DNA samples using KSHV TR-specific primers to calculate the ratios of immunoprecipitated TR copies in the presence and absence of LANA expression in both TopoIIβ−/− and TopoIIβ+/+ cells. Fold change analysis, based on the ΔCT values, showed significant increase in copies of ChIP DNA in TopoIIβ sufficient (TopoIIβ+/+) cells in the presence of LANA, suggesting LANA was able to increase dsDNA breaks in TopoIIβ+/+ cells (Fig. 7F). TopoIIβ−/− cells showed minimal TR copies with biotin ChIP, which did not show any change with LANA expression, suggesting background levels of biotin incorporation in those cells (Fig. 7F). To further evaluate the role of TopoIIβ in KSHV latent DNA replication, DNA break labeling and ChIP assays were performed on HEK 293L cells transfected with KSHV TR-containing plasmid either with pA3F-LANA or with empty vector pA3F. One set of the transfected cells was treated with a 5 μM concentration of ellipticine. After 24 h, cells were harvested and a portion (1 million) of the cells were used for cell cycle analysis after propidium iodide (PI) staining and flow cytometry. The remaining cells were fixed with Streck tissue fixative and used subsequently to label with biotin-16-dUTP followed by ChIP and quantitative PCR as described earlier. The flow cytometry data showed that the transient overexpression of LANA slightly affected the cell cycle and pushed the cells into S phase as expected, whereas the treatment with ellipticine appeared to arrest the cells at S phase (Fig. 7H). Furthermore, the quantitative PCR data from biotin-16-dUTP ChIP showed that transient expression of LANA indeed increased the incorporation of biotin-16-dUTP, suggesting increased incidence of double-stranded breaks in LANA-expressing cells (Fig. 7G). To determine whether the dsDNA break was limited to the TR region, we compared the copies of TR and the vector backbone (ampicillin gene) in sonicated and biotin ChIP DNA, which showed increased copies of the TR but not the vector backbone (Fig. 7G, compare black and gray bars in −LANA and +LANA). Interestingly, a treatment with ellipticine effectively reduced the incorporation of biotin-16-dUTP, strongly suggesting a role for TopoIIβ in mediating double-stranded breaks in the KSHV TR DNA. These data suggest that TopoIIβ mediates nicks on KSHV TR DNA to unwind the supercoiled DNA and thus assists in latent DNA replication.
DISCUSSION
Kaposi's sarcoma-associated herpesvirus is tightly associated with multiple human malignancies including Kaposi's sarcoma (KS), primary effusion lymphomas (PELs) and multicentric Castleman's disease (MCD) (63). KSHV establishes a lifelong latent infection following primary infection and propagates into dividing tumor cells following duplication of the episomes along with the human chromosomes (68). During latency, protein expression is tightly regulated by transcriptional repression and is limited to a few crucial oncogenic proteins (63). LANA, encoded by ORF73, is consistently expressed in KS lesions and is crucial for the maintenance of viral episomes in proliferating cells (4, 63, 64, 69). LANA not only modulates the transcription of viral and cellular genes but also recruits a number of molecules to regulate the replication of the viral episome and the segregation of the newly synthesized genome copies to daughter progeny nuclei by tethering to the host chromosomes (3, 4, 63).
Others and we have shown that LANA is critical for the replication of TR-containing plasmids, but the molecular mechanism of replication initiation at the TR is poorly understood (4, 8, 42). DNA affinity and coimmunoprecipitation analysis using KSHV, TR DNA, or ori-Lyt DNA as the affinity ligand identified topoisomerase II (TopoIIβ) as an interacting protein (53, 67). KSHV and the other herpesviruses studied so far do not encode topoisomerases. Thus, it could be assumed that these viruses use cellular topoisomerases for the viral DNA replication (19, 67). Indeed, the importance of TopoIIβ in KSHV lytic DNA replication was recently demonstrated (67). It has been shown that gene silencing of TopoI and TopoII with specific short hairpin RNA (shRNA) or use of ellipticine, a specific inhibitor of TopoII, could abolish KSHV lytic DNA replication (19). Additionally, it has been shown that a wide variety of potential topoisomerase inhibitors effectively block KSHV lytic DNA replication and could be further screened as therapeutic drugs (19).
Since TopoIIβ was identified as a LANA-interacting protein in a DNA affinity column and proteomics analysis, it was suggested that TopoIIβ may have a role in DNA replication at the latent origins of the TR (53). In this study, we show that LANA recruits the cellular protein topoisomerase IIβ for KSHV latent DNA replication. TopoIIβ proteins form complexes with LANA and colocalize as punctuate bodies in the nuclei of KSHV-infected BCBL-1 and JSC-1 cells. TopoIIβ is an enzyme that controls and alters the topological state of DNA during transcription and replication and has been shown to induce double-stranded breaks required for regulated transcription/replication (15, 25).
The amino terminus of LANA is identified as the TopoIIβ binding domain; more specifically, the first 32 amino acids, containing the nucleosome-binding region, seem to be crucial for TopoIIβ binding. Additionally, this region of aa 1 to 32 acted as a dominant negative and disrupted association of TopoIIβ with LANA when expressed in excess, suggesting that TopoIIβ interacts only through aa 1 to 32 (Fig. 7A and B). Since the region encompassing aa 1 to 32 of LANA has also been shown to be the region of nucleosomal attachment, one could argue that the DNA-linking nucleosomes may have a role in associating TopoIIβ with LANA (6, 14). Nuclease treatment of the lysates prior to immunoprecipitation excludes the possibility of DNA facilitating their interaction. Additionally, the binding analysis with bacterially expressed protein confirmed that the association of LANA aa 1 to 32 with TopoIIβ is more likely a direct interaction. Alanine substitution mutations within LANA aa 1 to 32 and subsequent co-IP assays showed that the TopoIIβ binding region lies very close to the chromatin-binding region identified previously (6). Amino-terminal regions of LANA carrying mutations 8LRS10 to AAA, 11GRS13 to AAA, and 14TG15 to AA and mutant M5, where residues between aa 5 and 15 were mutated to alanine, failed to bind to TopoIIβ (Fig. 5B and C). These studies suggest that amino acid residues 8 to 15 are essential for TopoIIβ binding (Fig. 5B and C). Interestingly, aa 5 to 15 of LANA are also shown to bind with histones for chromosome tethering and are required for episome replication (5, 6, 27). This suggests that these residues may serve as binding sites for other cellular proteins along with the histones. The C-terminal domain of LANA has been shown to bind directly to the LANA-binding sites (LBS) in the KSHV TR, and the binding is required for origin firing in the RE region of the TR (3, 4, 59, 63). Also, the amino and carboxy termini of LANA have been shown to associate with each other (26). Thus, it could be hypothesized that LANA brings TopoIIβ in close proximity to the LANA binding site through its amino terminus to initiate replication in the RE region of TR.
The requirement of TopoIIβ in latent DNA replication of TR was confirmed in a transient replication assay on TopoIIβ knockout MEF NIH 3T3 cells (TopoIIβ−/−). Only wild-type (TopoIIβ+/+) MEF NIH 3T3 cells expressing LANA supported KSHV latent DNA replication, which strongly suggests that TopoIIβ is required for latent replication. The role of TopoIIβ in replication was also confirmed by treating the cells with ellipticine, a selective inhibitor of TopoII, which abolished latent DNA replication mediated by TR in BCBL-1 cells. Ellipticine, which has previously been shown to block DNA replication mediated by the lytic origin (ori-Lyt), was used as a positive control in our replication assays to demonstrate that ellipticine treatment and the replication assay were working in our hands (67). These data suggest that TopoIIβ plays an important role in KSHV latent DNA replication.
The role of specific residues of LANA important for binding with TopoIIβ was further confirmed in replication assays to determine whether the alanine substitution mutants, which were unable to bind to TopoIIβ, can support DNA replication. LANA mutants with mutations at aa 5 to 7 (GMR-AAA) and aa 5 to 15 were unable to support replication of TR plasmid, suggesting that TopoIIβ may be essential for DNA replication. Alternatively, tethering of the TR plasmids to the host chromosome may also be critical for replication, which may bring the cellular replication machinery at the TR origin. Dominant negative aa 1 to 32 may have disrupted the association of TopoIIβ along with other cellular factors including histones in order to block DNA replication. This is evidenced by the fact that the LANA mutant with substitutions at aa 5 to 7 (GMR) bound with TopoIIβ but was unable to support replication, therefore suggesting the involvement of additional factors for replication initiation at the TR. Previous studies have shown that LANA mutants lacking chromosome binding (5GMR7, 8LRS10, and 11GRS13) were unable to support replication, but the mechanisms were not determined (5). Here, we show that TopoIIβ may be the one factor required for LANA-mediated replication, as the mutants lacking TopoIIβ binding were unable to support replication. Also, our dominant negative data whereby the association of TopoIIβ with LANA was inhibited by expressing an excess amount of LANA aa 1 to 32, which reduced replication, confirmed that the recruitment of TopoIIβ by LANA is required for replication. In other words, close proximity of TopoIIβ to the site of replication initiation is important for replication. Since the replication element (RE) is adjacent to the LANA binding site (23), recruitment of TopoIIβ by LANA may bring the TopoIIβ protein at the RE site to initiate replication (Fig. 8).
Fig 8.

Schematic model showing the association of LANA with TopoIIβ. LANA recruits TopoIIβ to the sites of latent origin of replication. TopoIIβ in turn mediates double-stranded breaks required for viral DNA replication, further facilitated by the cellular replication machinery.
TopoIIβ controls and alters the topologic state of DNA during transcription and replication by changing the degree of supercoiling of double-stranded DNA molecules (15, 25). TopoIIβ has been shown to induce double-stranded breaks required for regulated transcription/replication (25). Our DNA break labeling and ChIP assay to detect nicks in the KSHV TR DNA suggests that TopoIIβ mediates transient DNA breaks on KSHV DNA in a LANA-dependent manner (Fig. 7E). Moreover, ellipticine was also found to effectively interfere with the incorporation of biotin-16-dUTP in the double-stranded nick assay, strongly suggesting a role for TopoIIβ in mediating double-stranded breaks on KSHV TR DNA (Fig. 7G). Therefore, we propose that LANA recruits TopoIIβ at the latent replication origins of TR to create a nick in order to unwind the complex supercoiled DNA for the facilitation of DNA replication (Fig. 8).
ACKNOWLEDGMENTS
We thank Erle S. Robertson, University of Pennsylvania, for providing the cell lines and LANA expression plasmids.
This work was supported by K99/R00, CA126182 to S.C.V.
Footnotes
Published ahead of print 3 July 2012
REFERENCES
- 1. An FQ, et al. 2005. The latency-associated nuclear antigen of Kaposi's sarcoma-associated herpesvirus modulates cellular gene expression and protects lymphoid cells from p16 INK4A-induced cell cycle arrest. J. Biol. Chem. 280:3862–3874 [DOI] [PubMed] [Google Scholar]
- 2. Ballestas ME, Chatis PA, Kaye KM. 1999. Efficient persistence of extrachromosomal KSHV DNA mediated by latency-associated nuclear antigen. Science 284:641–644 [DOI] [PubMed] [Google Scholar]
- 3. Ballestas ME, Kaye KM. 2001. Kaposi's sarcoma-associated herpesvirus latency-associated nuclear antigen 1 mediates episome persistence through cis-acting terminal repeat (TR) sequence and specifically binds TR DNA. J. Virol. 75:3250–3258 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Ballestas ME, Kaye KM. 2011. The latency-associated nuclear antigen, a multifunctional protein central to Kaposi's sarcoma-associated herpesvirus latency. Future Microbiol. 6:1399–1413 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Barbera AJ, Ballestas ME, Kaye KM. 2004. The Kaposi's sarcoma-associated herpesvirus latency-associated nuclear antigen 1 N terminus is essential for chromosome association, DNA replication, and episome persistence. J. Virol. 78:294–301 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Barbera AJ, et al. 2006. The nucleosomal surface as a docking station for Kaposi's sarcoma herpesvirus LANA. Science 311:856–861 [DOI] [PubMed] [Google Scholar]
- 7. Cai Q, Verma SC, Choi JY, Ma M, Robertson ES. 2010. Kaposi's sarcoma-associated herpesvirus inhibits interleukin-4-mediated STAT6 phosphorylation to regulate apoptosis and maintain latency. J. Virol. 84:11134–11144 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Cai Q, Verma SC, Lu J, Robertson ES. 2010. Molecular biology of Kaposi's sarcoma-associated herpesvirus and related oncogenesis. Adv. Virus Res. 78:87–142 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Cai Q, et al. 2012. Kaposi's sarcoma herpesvirus upregulates aurora A expression to promote p53 phosphorylation and ubiquitylation. PLoS Pathog. 8:e1002566 doi:10.1371/journal.ppat.1002566 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Cannon JS, et al. 2000. A new primary effusion lymphoma-derived cell line yields a highly infectious Kaposi's sarcoma herpesvirus-containing supernatant. J. Virol. 74:10187–10193 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Cesarman E, Chang Y, Moore PS, Said JW, Knowles DM. 1995. Kaposi's sarcoma-associated herpesvirus-like DNA sequences in AIDS-related body-cavity-based lymphomas. N. Engl. J. Med. 332:1186–1191 [DOI] [PubMed] [Google Scholar]
- 12. Chang Y, et al. 1994. Identification of herpesvirus-like DNA sequences in AIDS-associated Kaposi's sarcoma. Science 266:1865–1869 [DOI] [PubMed] [Google Scholar]
- 13. Chen W, Hilton IB, Staudt MR, Burd CE, Dittmer DP. 2010. Distinct p53, p53:LANA, and LANA complexes in Kaposi's s-associated herpesvirus lymphomas. J. Virol. 84:3898–3908 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Cotter MA, II, Robertson ES. 1999. The latency-associated nuclear antigen tethers the Kaposi's sarcoma-associated herpesvirus genome to host chromosomes in body cavity-based lymphoma cells. Virology 264:254–264 [DOI] [PubMed] [Google Scholar]
- 15. Cowell IG, Papageorgiou N, Padget K, Watters GP, Austin CA. 2011. Histone deacetylase inhibition redistributes topoisomerase IIbeta from heterochromatin to euchromatin. Nucleus 2:61–71 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Fakhari FD, Dittmer DP. 2002. Charting latency transcripts in Kaposi's sarcoma-associated herpesvirus by whole-genome real-time quantitative PCR. J. Virol. 76:6213–6223 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Friborg J, Jr, Kong W, Hottiger MO, Nabel GJ. 1999. p53 inhibition by the LANA protein of KSHV protects against cell death. Nature 402:889–894 [DOI] [PubMed] [Google Scholar]
- 18. Garber AC, Shu MA, Hu J, Renne R. 2001. DNA binding and modulation of gene expression by the latency-associated nuclear antigen of Kaposi's sarcoma-associated herpesvirus. J. Virol. 75:7882–7892 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Gonzalez-Molleda L, Wang Y, Yuan Y. 2012. Potent antiviral activity of topoisomerase I and II inhibitors against Kaposi's sarcoma-associated herpesvirus. Antimicrob. Agents Chemother. 56:893–902 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Grundhoff A, Ganem D. 2004. Inefficient establishment of KSHV latency suggests an additional role for continued lytic replication in Kaposi sarcoma pathogenesis. J. Clin. Invest. 113:124–136 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Hirt B. 1967. Selective extraction of polyoma DNA from infected mouse cell cultures. J. Mol. Biol. 26:365–369 [DOI] [PubMed] [Google Scholar]
- 22. Hu J, Garber AC, Renne R. 2002. The latency-associated nuclear antigen of Kaposi's sarcoma-associated herpesvirus supports latent DNA replication in dividing cells. J. Virol. 76:11677–11687 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Hu J, Renne R. 2005. Characterization of the minimal replicator of Kaposi's sarcoma-associated herpesvirus latent origin. J. Virol. 79:2637–2642 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Jenner RG, Alba MM, Boshoff C, Kellam P. 2001. Kaposi's sarcoma-associated herpesvirus latent and lytic gene expression as revealed by DNA arrays. J. Virol. 75:891–902 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Ju BG, et al. 2006. A topoisomerase IIbeta-mediated dsDNA break required for regulated transcription. Science 312:1798–1802 [DOI] [PubMed] [Google Scholar]
- 26. Kaul R, Verma SC, Robertson ES. 2007. Protein complexes associated with the Kaposi's sarcoma-associated herpesvirus-encoded LANA. Virology 364:317–329 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Komatsu T, Ballestas ME, Barbera AJ, Kelley-Clarke B, Kaye KM. 2004. KSHV LANA1 binds DNA as an oligomer and residues N-terminal to the oligomerization domain are essential for DNA binding, replication, and episome persistence. Virology 319:225–236 [DOI] [PubMed] [Google Scholar]
- 28. Lan K, et al. 2005. Induction of Kaposi's sarcoma-associated herpesvirus latency-associated nuclear antigen by the lytic transactivator RTA: a novel mechanism for establishment of latency. J. Virol. 79:7453–7465 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Lee HR, Lee S, Chaudhary PM, Gill P, Jung JU. 2010. Immune evasion by Kaposi's sarcoma-associated herpesvirus. Future Microbiol. 5:1349–1365 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Lim C, Sohn H, Gwack Y, Choe J. 2000. Latency-associated nuclear antigen of Kaposi's sarcoma-associated herpesvirus (human herpesvirus-8) binds ATF4/CREB2 and inhibits its transcriptional activation activity. J. Gen. Virol. 81:2645–2652 [DOI] [PubMed] [Google Scholar]
- 31. Lim C, Sohn H, Lee D, Gwack Y, Choe J. 2002. Functional dissection of latency-associated nuclear antigen 1 of Kaposi's sarcoma-associated herpesvirus involved in latent DNA replication and transcription of terminal repeats of the viral genome. J. Virol. 76:10320–10331 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Lu F, Day L, Gao SJ, Lieberman PM. 2006. Acetylation of the latency-associated nuclear antigen regulates repression of Kaposi's sarcoma-associated herpesvirus lytic transcription. J. Virol. 80:5273–5282 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Lu F, et al. 2012. Identification of host-chromosome binding sites and candidate gene targets for Kaposi's sarcoma-associated herpesvirus LANA. J. Virol. 86:5752–5762 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Lu J, et al. 2012. The RBP-Jkappa binding sites within the RTA promoter regulate KSHV latent infection and cell proliferation. PLoS Pathog. 8:e1002479 doi:10.1371/journal.ppat.1002479 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Lu J, et al. 2009. Latency-associated nuclear antigen of Kaposi's sarcoma-associated herpesvirus (KSHV) upregulates survivin expression in KSHV-associated B-lymphoma cells and contributes to their proliferation. J. Virol. 83:7129–7141 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Lyu YL, et al. 2007. Topoisomerase IIbeta mediated DNA double-strand breaks: implications in doxorubicin cardiotoxicity and prevention by dexrazoxane. Cancer Res. 67:8839–8846 [DOI] [PubMed] [Google Scholar]
- 37. Matsumura S, Persson LM, Wong L, Wilson AC. 2010. The latency-associated nuclear antigen interacts with MeCP2 and nucleosomes through separate domains. J. Virol. 84:2318–2330 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Mesri EA, Cesarman E, Boshoff C. 2010. Kaposi's sarcoma and its associated herpesvirus. Nat. Rev. Cancer 10:707–719 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Mirski SE, et al. 2007. Topoisomerase II binds importin alpha isoforms and exportin/CRM1 but does not shuttle between the nucleus and cytoplasm in proliferating cells. Exp. Cell Res. 313:627–637 [DOI] [PubMed] [Google Scholar]
- 40. Moore PS, Chang Y. 2010. Why do viruses cause cancer? Highlights of the first century of human tumour virology. Nat. Rev. Cancer 10:878–889 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Moore PS, Chang Y. 2003. Kaposi's sarcoma-associated herpesvirus immunoevasion and tumorigenesis: two sides of the same coin? Annu. Rev. Microbiol. 57:609–639 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Ohsaki E, Ueda K. 2012. Kaposi's sarcoma-associated herpesvirus genome replication, partitioning, and maintenance in latency. Front. Microbiol. 3:7 doi:10.3389/fmicb.2012.00007 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Radkov SA, Kellam P, Boshoff C. 2000. The latent nuclear antigen of Kaposi sarcoma-associated herpesvirus targets the retinoblastoma-E2F pathway and with the oncogene Hras transforms primary rat cells. Nat. Med. 6:1121–1127 [DOI] [PubMed] [Google Scholar]
- 44. Rainbow L, et al. 1997. The 222- to 234-kilodalton latent nuclear protein (LNA) of Kaposi's sarcoma-associated herpesvirus (human herpesvirus 8) is encoded by orf73 and is a component of the latency-associated nuclear antigen. J. Virol. 71:5915–5921 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Renne R, et al. 2001. Modulation of cellular and viral gene expression by the latency-associated nuclear antigen of Kaposi's sarcoma-associated herpesvirus. J. Virol. 75:458–468 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Renne R, et al. 1996. Lytic growth of Kaposi's sarcoma-associated herpesvirus (human herpesvirus 8) in culture. Nat. Med. 2:342–346 [DOI] [PubMed] [Google Scholar]
- 47. Riccardi C, Nicoletti I. 2006. Analysis of apoptosis by propidium iodide staining and flow cytometry. Nat. Protoc. 1:1458–1461 [DOI] [PubMed] [Google Scholar]
- 48. Rossetto C, Yamboliev I, Pari GS. 2009. Kaposi's sarcoma-associated herpesvirus/human herpesvirus 8 K-bZIP modulates latency-associated nuclear protein-mediated suppression of lytic origin-dependent DNA synthesis. J. Virol. 83:8492–8501 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49. Saha A, Kaul R, Murakami M, Robertson ES. 2010. Tumor viruses and cancer biology: modulating signaling pathways for therapeutic intervention. Cancer Biol. Ther. 10:961–978 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50. Sakakibara S, et al. 2004. Accumulation of heterochromatin components on the terminal repeat sequence of Kaposi's sarcoma-associated herpesvirus mediated by the latency-associated nuclear antigen. J. Virol. 78:7299–7310 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51. Schwam DR, Luciano RL, Mahajan SS, Wong L, Wilson AC. 2000. Carboxy terminus of human herpesvirus 8 latency-associated nuclear antigen mediates dimerization, transcriptional repression, and targeting to nuclear bodies. J. Virol. 74:8532–8540 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52. Si H, Robertson ES. 2006. Kaposi's sarcoma-associated herpesvirus-encoded latency-associated nuclear antigen induces chromosomal instability through inhibition of p53 function. J. Virol. 80:697–709 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53. Si H, Verma SC, Robertson ES. 2006. Proteomic analysis of the Kaposi's sarcoma-associated herpesvirus terminal repeat element binding proteins. J. Virol. 80:9017–9030 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54. Skalsky RL, Hu J, Renne R. 2007. Analysis of viral cis elements conferring Kaposi's sarcoma-associated herpesvirus episome partitioning and maintenance. J. Virol. 81:9825–9837 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55. Stedman W, Deng Z, Lu F, Lieberman PM. 2004. ORC, MCM, and histone hyperacetylation at the Kaposi's sarcoma-associated herpesvirus latent replication origin. J. Virol. 78:12566–12575 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56. Tsutsui K, Sano K, Kikuchi A, Tokunaga A. 2001. Involvement of DNA topoisomerase IIbeta in neuronal differentiation. J. Biol. Chem. 276:5769–5778 [DOI] [PubMed] [Google Scholar]
- 57. Ueda K, Sakakibara S, Ohsaki E, Yada K. 2006. Lack of a mechanism for faithful partition and maintenance of the KSHV genome. Virus Res. 122:85–94 [DOI] [PubMed] [Google Scholar]
- 58. Verma SC, et al. 2006. Latency-associated nuclear antigen of Kaposi's sarcoma-associated herpesvirus recruits uracil DNA glycosylase 2 at the terminal repeats and is important for latent persistence of the virus. J. Virol. 80:11178–11190 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59. Verma SC, Choudhuri T, Kaul R, Robertson ES. 2006. Latency-associated nuclear antigen (LANA) of Kaposi's sarcoma-associated herpesvirus interacts with origin recognition complexes at the LANA binding sequence within the terminal repeats. J. Virol. 80:2243–2256 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60. Verma SC, Choudhuri T, Robertson ES. 2007. The minimal replicator element of the Kaposi's sarcoma-associated herpesvirus terminal repeat supports replication in a semiconservative and cell-cycle-dependent manner. J. Virol. 81:3402–3413 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61. Verma SC, Lan K, Choudhuri T, Cotter MA, Robertson ES. 2007. An autonomous replicating element within the KSHV genome. Cell Host Microbe 2:106–118 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62. Verma SC, Lan K, Choudhuri T, Robertson ES. 2006. Kaposi's sarcoma-associated herpesvirus-encoded latency-associated nuclear antigen modulates K1 expression through its cis-acting elements within the terminal repeats. J. Virol. 80:3445–3458 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63. Verma SC, Lan K, Robertson E. 2007. Structure and function of latency-associated nuclear antigen. Curr. Top. Microbiol. Immunol. 312:101–136 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64. Verma SC, Robertson ES. 2003. Molecular biology and pathogenesis of Kaposi sarcoma-associated herpesvirus. FEMS Microbiol. Lett. 222:155–163 [DOI] [PubMed] [Google Scholar]
- 65. Viejo-Borbolla A, et al. 2005. Brd2/RING3 interacts with a chromatin-binding domain in the Kaposi's sarcoma-associated herpesvirus latency-associated nuclear antigen 1 (LANA-1) that is required for multiple functions of LANA-1. J. Virol. 79:13618–13629 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66. Wang CY, Sugden B. 2004. New viruses shake old paradigms. J. Clin. Invest. 113:21–23 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67. Wang Y, Li H, Tang Q, Maul GG, Yuan Y. 2008. Kaposi's sarcoma-associated herpesvirus ori-Lyt-dependent DNA replication: involvement of host cellular factors. J. Virol. 82:2867–2882 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68. Xiao B, et al. 2010. Bub1 and CENP-F can contribute to Kaposi's sarcoma-associated herpesvirus genome persistence by targeting LANA to kinetochores. J. Virol. 84:9718–9732 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69. Zhong W, Wang H, Herndier B, Ganem D. 1996. Restricted expression of Kaposi sarcoma-associated herpesvirus (human herpesvirus 8) genes in Kaposi sarcoma. Proc. Natl. Acad. Sci. U. S. A. 93:6641–6646 [DOI] [PMC free article] [PubMed] [Google Scholar]