Abstract
A method is described for cutting DNA at specific sites that are 8 and 10 base pairs long. The DNA is first treated with a specific methylase, either the restriction-modification enzyme M. Taq I, which converts the 4-base sequence T-C-G-A to T-C-G-mA, or the similar enzyme M. Cla I, which converts the 6-base sequence A-T-C-G-A-T to A-T-C-G-mA-T. The DNA is then cleaved with Dpn I, a restriction endonuclease that recognizes the sequence G-mA-T-C. Dpn I is unique in that it cuts only DNA that is methylated at adenine in both strands of its recognition sequence. In DNAs that are not otherwise methylated at adenine in both strands of the sequence G-A-T-C, cleavage by Dpn I occurs only at the following sequences: in the case of M. Taq I methylation, 5' T-C-G-mA - T-C-G-mA 3' 3' mA-G-C - T-mA-G-C - T 5'; in the case of M. Cla I methylation, 5' A - T-C-G-mA - T-C-G-mA-T 3' 3' T-mA-G-C - T-mA-G-C - T-A 5'. Specific cutting and cloning at these methylase/Dpn I-generated sites is shown experimentally. Further, we describe how the above technique can be extended to generate Dpn I cleavage sites of up to 12 base pairs. In DNA that contains equal amounts of each base distributed at random, 8- and 10-base-pair recognition sequences occur, on the average, approximately once every 65,000 and 1,000,000 base pairs, respectively. Potential applications, including the development of cloning vectors and a rapid method for chromosome walking, are discussed.
Full text
PDF

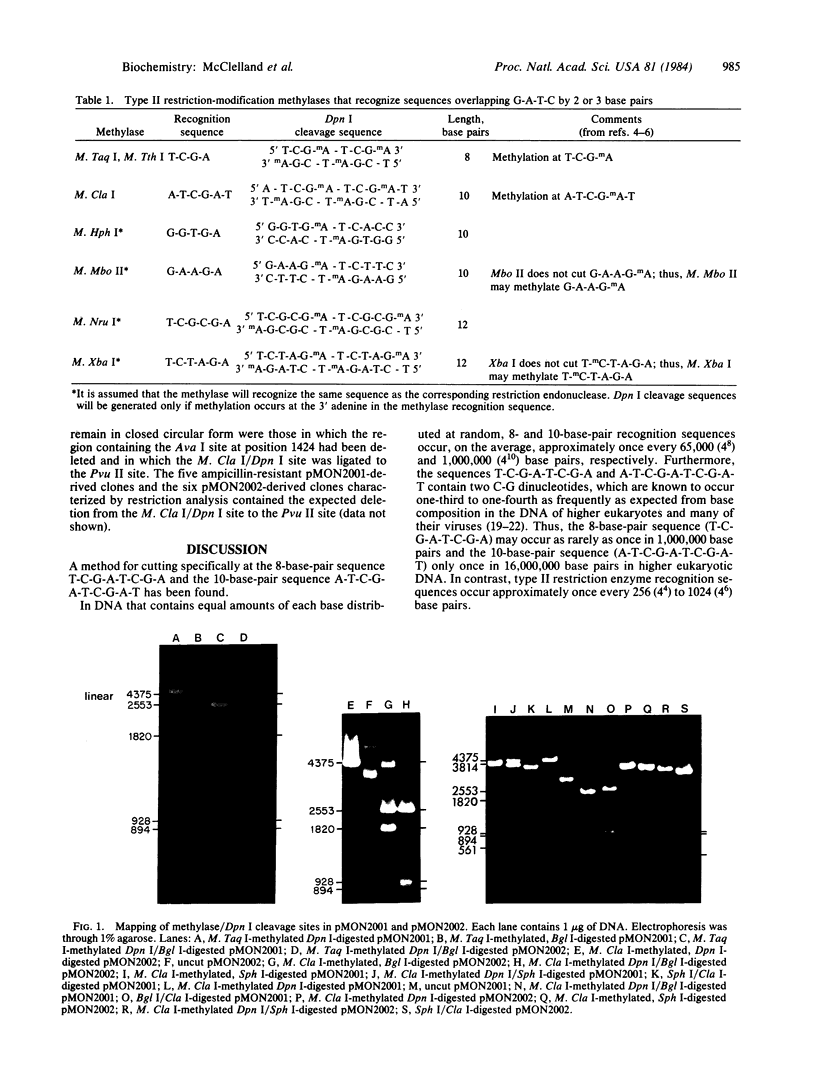
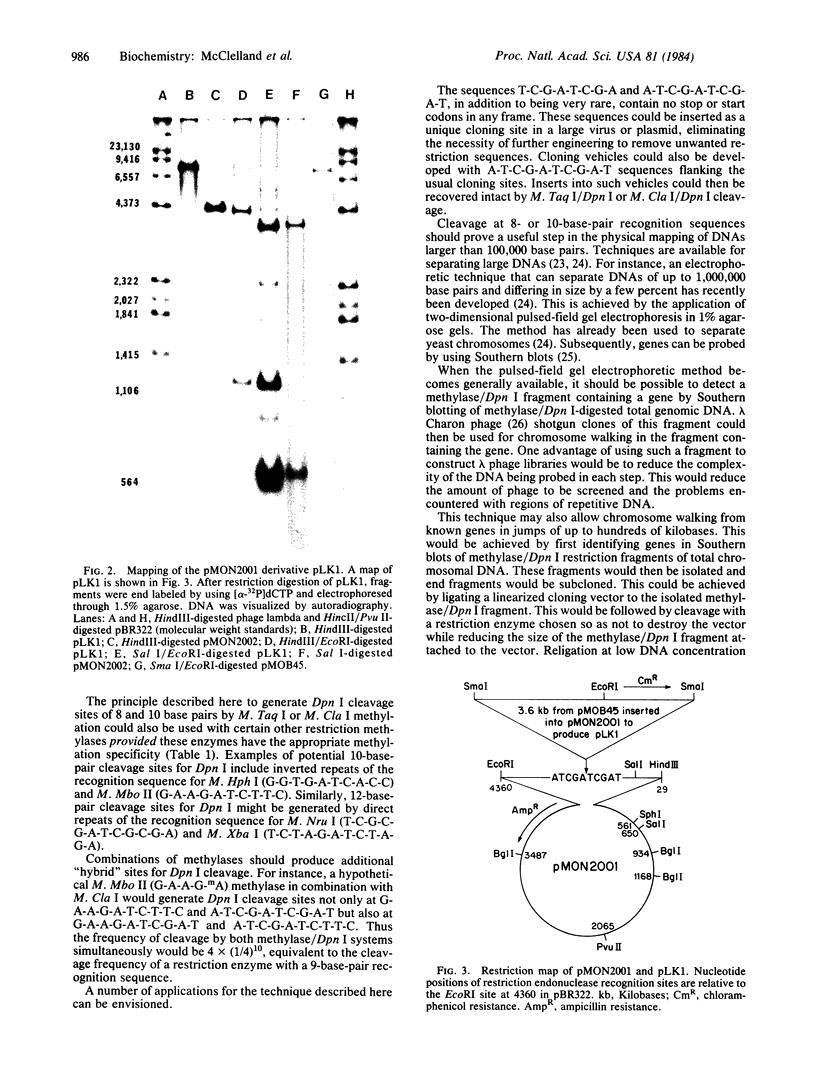

Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bittner M., Vapnek D. Versatile cloning vectors derived from the runaway-replication plasmid pKN402. Gene. 1981 Dec;15(4):319–329. doi: 10.1016/0378-1119(81)90175-x. [DOI] [PubMed] [Google Scholar]
- Dagert M., Ehrlich S. D. Prolonged incubation in calcium chloride improves the competence of Escherichia coli cells. Gene. 1979 May;6(1):23–28. doi: 10.1016/0378-1119(79)90082-9. [DOI] [PubMed] [Google Scholar]
- Dreiseikelmann B., Eichenlaub R., Wackernagel W. The effect of differential methylation by Escherichia coli of plasmid DNA and phage T7 and lambda DNA on the cleavage by restriction endonuclease MboI from Moraxella bovis. Biochim Biophys Acta. 1979 May 24;562(3):418–428. doi: 10.1016/0005-2787(79)90105-9. [DOI] [PubMed] [Google Scholar]
- Fangman W. L. Separation of very large DNA molecules by gel electrophoresis. Nucleic Acids Res. 1978 Mar;5(3):653–665. doi: 10.1093/nar/5.3.653. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Geier G. E., Modrich P. Recognition sequence of the dam methylase of Escherichia coli K12 and mode of cleavage of Dpn I endonuclease. J Biol Chem. 1979 Feb 25;254(4):1408–1413. [PubMed] [Google Scholar]
- Ish-Horowicz D., Burke J. F. Rapid and efficient cosmid cloning. Nucleic Acids Res. 1981 Jul 10;9(13):2989–2998. doi: 10.1093/nar/9.13.2989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lacks S., Greenberg B. Complementary specificity of restriction endonucleases of Diplococcus pneumoniae with respect to DNA methylation. J Mol Biol. 1977 Jul;114(1):153–168. doi: 10.1016/0022-2836(77)90289-3. [DOI] [PubMed] [Google Scholar]
- Lansman R. A., Shade R. O., Shapira J. F., Avise J. C. The use of restriction endonucleases to measure mitochondrial DNA sequence relatedness in natural populations. III. Techniques and potential applications. J Mol Evol. 1981;17(4):214–226. doi: 10.1007/BF01732759. [DOI] [PubMed] [Google Scholar]
- Marinus M. G., Morris N. R. Isolation of deoxyribonucleic acid methylase mutants of Escherichia coli K-12. J Bacteriol. 1973 Jun;114(3):1143–1150. doi: 10.1128/jb.114.3.1143-1150.1973. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McClelland M., Ivarie R. Asymmetrical distribution of CpG in an 'average' mammalian gene. Nucleic Acids Res. 1982 Dec 11;10(23):7865–7877. doi: 10.1093/nar/10.23.7865. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McClelland M. Purification and characterization of two new modification methylases: MClaI from Caryophanon latum L and MTaqI from Thermus aquaticus YTI. Nucleic Acids Res. 1981 Dec 21;9(24):6795–6804. doi: 10.1093/nar/9.24.6795. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McClelland M. The effect of sequence specific DNA methylation on restriction endonuclease cleavage. Nucleic Acids Res. 1981 Nov 25;9(22):5859–5866. doi: 10.1093/nar/9.22.5859. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McClelland M. The effect of site specific methylation on restriction endonuclease cleavage (update). Nucleic Acids Res. 1983 Jan 11;11(1):r169–r173. doi: 10.1093/nar/11.1.235-c. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McClelland M. The frequency and distribution of methylatable DNA sequences in leguminous plant protein coding genes. J Mol Evol. 1983;19(5):346–354. doi: 10.1007/BF02101638. [DOI] [PubMed] [Google Scholar]
- Messing J., Vieira J. A new pair of M13 vectors for selecting either DNA strand of double-digest restriction fragments. Gene. 1982 Oct;19(3):269–276. doi: 10.1016/0378-1119(82)90016-6. [DOI] [PubMed] [Google Scholar]
- Modrich P. Structures and mechanisms of DNA restriction and modification enzymes. Q Rev Biophys. 1979 Aug;12(3):315–369. doi: 10.1017/s0033583500005461. [DOI] [PubMed] [Google Scholar]
- Roberts R. J. Restriction and modification enzymes and their recognition sequences. Nucleic Acids Res. 1983 Jan 11;11(1):r135–r167. [PMC free article] [PubMed] [Google Scholar]
- Sanger F., Coulson A. R., Barrell B. G., Smith A. J., Roe B. A. Cloning in single-stranded bacteriophage as an aid to rapid DNA sequencing. J Mol Biol. 1980 Oct 25;143(2):161–178. doi: 10.1016/0022-2836(80)90196-5. [DOI] [PubMed] [Google Scholar]
- Sato S., Nakazawa K., Shinomiya T. A DNA methylase from Thermus thermophilus HB8. J Biochem. 1980 Sep;88(3):737–747. doi: 10.1093/oxfordjournals.jbchem.a133026. [DOI] [PubMed] [Google Scholar]
- Schwartz D. C., Saffran W., Welsh J., Haas R., Goldenberg M., Cantor C. R. New techniques for purifying large DNAs and studying their properties and packaging. Cold Spring Harb Symp Quant Biol. 1983;47(Pt 1):189–195. doi: 10.1101/sqb.1983.047.01.024. [DOI] [PubMed] [Google Scholar]
- Smith H. O., Nathans D. Letter: A suggested nomenclature for bacterial host modification and restriction systems and their enzymes. J Mol Biol. 1973 Dec 15;81(3):419–423. doi: 10.1016/0022-2836(73)90152-6. [DOI] [PubMed] [Google Scholar]
- Southern E. M. Detection of specific sequences among DNA fragments separated by gel electrophoresis. J Mol Biol. 1975 Nov 5;98(3):503–517. doi: 10.1016/s0022-2836(75)80083-0. [DOI] [PubMed] [Google Scholar]