Abstract
Aims: The human LRRK2 gene has been identified as the most common causative gene of autosomal-dominantly inherited and idiopathic Parkinson disease (PD). The G2019S substitution is the most common mutation in LRRK2. The R1441C mutation also occurs in cases of familial PD, but is not as prevalent. Some cases of LRRK2-based PD exhibit Tau pathology, which suggests that alterations on LRRK2 activity affect the pathophysiology of Tau. To investigate how LRRK2 might affect Tau and the pathophysiology of PD, we generated lines of C. elegans expressing human LRRK2 [wild-type (WT) or mutated (G2019S or R1441C)] with and without V337M Tau. Expression and redox proteomics were used to identify the effects of LRRK2 (WT and mutant) on protein expression and oxidative modifications. Results: Co-expression of WT LRRK2 and Tau led to increased expression of numerous proteins, including several 60S ribosomal proteins, mitochondrial proteins, and the V-type proton ATPase, which is associated with autophagy. C. elegans expressing mutant LRRK2 showed similar changes, but also showed increased protein oxidation and lipid peroxidation, the latter indexed as increased protein-bound 4-hydroxy-2-nonenal (HNE). Innovation: Our study brings new knowledge about the possible alterations induced by LRRK2 (WT and mutated) and Tau interactions, suggesting the involvement of G2019S and R1441C in Tau-dependent neurodegenerative processes. Conclusion: These results suggest that changes in LRRK2 expression or activity lead to corresponding changes in mitochondrial function, autophagy, and protein translation. These findings are discussed with reference to the pathophysiology of PD. Antioxid. Redox Signal. 17, 1490–1506.
Introduction
Parkinson disease (PD) is the second most prevalent degenerative disease of the nervous system. Functional studies of PD-related genes implicate dysfunction of mitochondria, autophagy, and the stress response in the pathophysiology of PD (11, 42, 47).
At least nine different genes are known to cause familial PD. Mutations in the α-synuclein, parkin, microtubule associated protein tau (MAPT), ubiquitin carboxy-terminal hydrolase L1 (UCH-L1), DJ-1, PTEN-induced kinase 1 (PINK1), and leucine-rich repeat kinase 2 (LRRK2) genes have been implicated in hereditary PD (16, 19).
The LRRK2 gene has been identified as the most common causative gene of autosomal-dominant inherited and idiopathic PD (54). The LRRK2 gene encodes a complex multidomain protein (2527 amino acids; 286 kDa,). The LRRK2 gene has specific domains including N-terminal ankyrin repeats, a leucine-rich repeat (LRR) region, a catalytic core, featuring a functional ROC (Ras Complex proteins), a GTPase domain, and a kinase domain (mitogen-activated protein kinase kinase kinase, MAPKKK) linked by the C- terminus of ROC (COR), followed by a C-terminal WD40 domain (37). In vitro studies demonstrated that LRRK2 is both a functional kinase and a GTPase, able to undergo autophosphorylation and perform phosphorylation of generic and putative physiological substrates (20).
Innovation.
Our proteomics study is the first of its kind to identify altered expression and altered oxidatively modified proteins in transgenic C. elegans models associated with PD. Our study brings new knowledge on the possible alteration induced by LRRK2 (WT and mutated) and Tau interactions, identified as impaired autophagy, energy metabolism, proteasomal function, and cellular structure among others, suggesting the involvement of G2019S and R1441C in Tau-dependent neurodegenerative processes.
Experimental evidence suggests LRRK2 may play a role in neuritic outgrowth and branching (8), and the presence of both a ROC/GTPase and a kinase domain suggests that LRRK2 plays a role in intracellular signaling. Overexpression of wild-type LRRK2 induces neuronal cell death, neurite shortening, protein aggregation, oxidative stress-induced cell death, and increased levels of intracellular reactive oxygen species (ROS) (34, 51).
LRRK2 is present in the cytoplasm but also associates with numerous organelles, such as mitochondria, endoplasmic reticulum, trans-Golgi, and also with the plasma membrane (24, 50). To date, R1441C/G, Y1699C, G2019S, and I2020T mutations have proven to be pathogenic (11, 48). The clinical symptoms of LRRK2 mutation carriers are similar to those of idiopathic PD patients, whereas the related neuropathology is pleomorphic, including α-synucleinopathy, tauopathy, and ubiquitin deposits or nigral neuronal loss solely (11, 43, 61). Among identified mutations of LRRK2, the amino acid substitution G2019S has been regarded as the most common cause of dominantly inherited as well as sporadic PD (14, 40). Previous studies showed that G2019S augments LRRK2 autophosphorylation and kinase activity, which cause neurite degeneration and neuronal cell death (34, 51, 57), suggesting a mechanism of toxic gain-of-function, probably related to deregulation of LRRK2 kinase activity. The R1441C mutation is another frequent LRRK2 mutation. This mutation decreases the GTPase activity of LRRK2 and could destabilize the interaction between monomers, which might be the mechanism of enzymatic dysfunction (28–30).
The interconnection between PD and Tau is suggested by the identification of mutations in the Tau-encoding MAPT gene that are linked to frontotemporal dementia with Parkinsonism (59). Mutations in LRRK2 generate PD that is sometimes associated with Tau pathology (19). Based on this correlation, we hypothesize that LRRK2 kinase dysfunction leads to deregulation of the post-translational state of Tau (31, 33), which in turn controls its compartmentalization and finally its functional roles in neuronal maintenance.
We previously generated lines of Caenorhabditis. elegans expressing human wild-type (WT) LRRK2 and the mutants, G2019S and R1441C (47). In the current study, we have crossed these lines with a line of C. elegans expressing V337M Tau to investigate how mutations in LRRK2 might contribute to disease. We explored these questions using expression proteomics and redox proteomics methods (13).
Results
Characterization of LRRK2 C. elegans strains
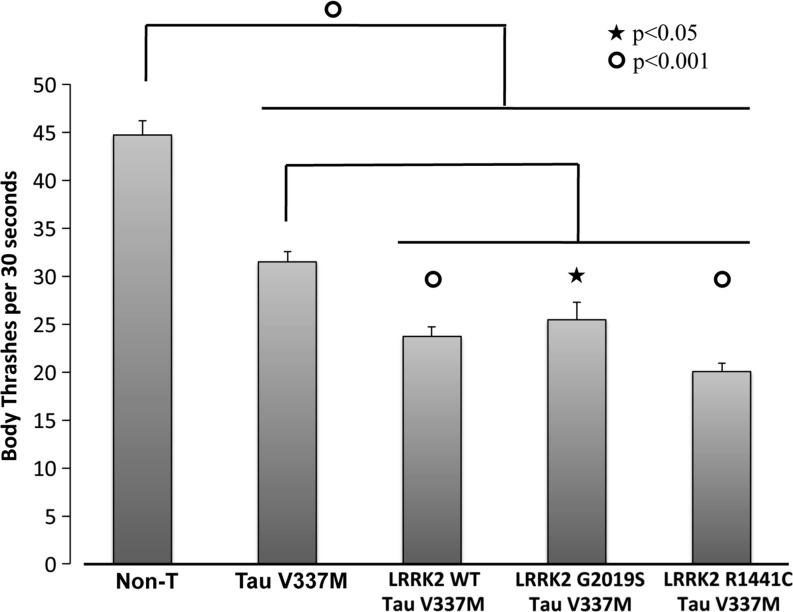
The crossing of nematodes expressing LRRK2 and Tau V337M produced viable offspring with no effects on lifespan or fertility but severe movement impairments. The motor deficits were quantified with the thrashing assay, as this has been shown to be a reliable indicator of Tau pathology in C. elegans (23, 26, 27). The combination of LRRK2 and Tau V337M produced a significant reduction in thrashing performance compared to both nontransgenic and Tau V337M alone (Fig. 1).
FIG. 1.
Bar graph of body trashes count in 30 sec in C. elegans transgenic and nontransgenic strains. Raw data are reported.
Total protein oxidation levels
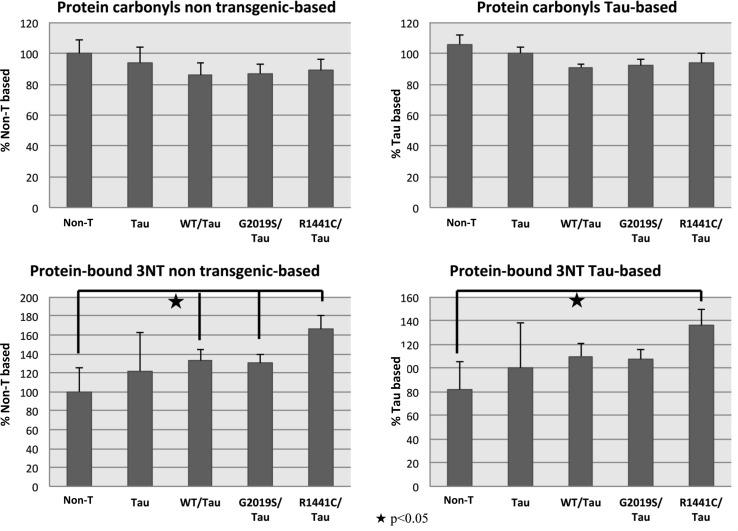
As a starting point for the study, we measured by slot blot analysis protein oxidative modification indexed as protein carbonylation, protein-resident 3-nitrotyrosine (3NT), and protein-bound HNE. In the analysis of total protein carbonyl levels shown in Figure 2, we saw no significant difference between nontransgenic (Non-T), Tau V337M transgenic (Tau), and LRRK2::Tau transgenic samples whether we compared them using as the control group Non-T or Tau. Interestingly, the carbonyl levels of LRRK2::Tau are lower than Non-T or TAU but did not reach statistical significance.
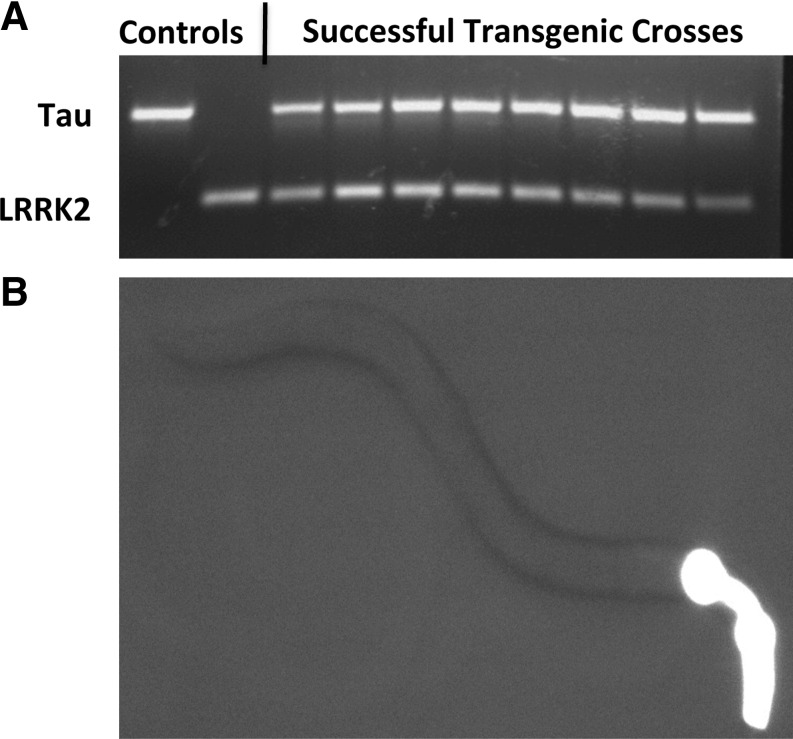
FIG. 2.
(A) PCR gel of tau and LRRK2 gene expression; (B) Expression of GFP in the pharyngeal muscles of a young adult transgenic C. elegans expressing Tau V337M. Fluorescence was used as a preliminary indicator of transgene expression. Image taken with a 20× objective.
Comparing protein-bound 3NT levels of Non-T with LRRK2::Tau samples (Fig. 2), we showed a significant increase of 133%, 130%, and 166% in WT/TAU, G2019S::Tau, and R1441C::Tau, respectively. The comparison of LRRK2::Tau transgenic samples with Tau transgenic samples in contrast showed a significant increase in protein-bound 3NT levels only for the R1441C::Tau sample.
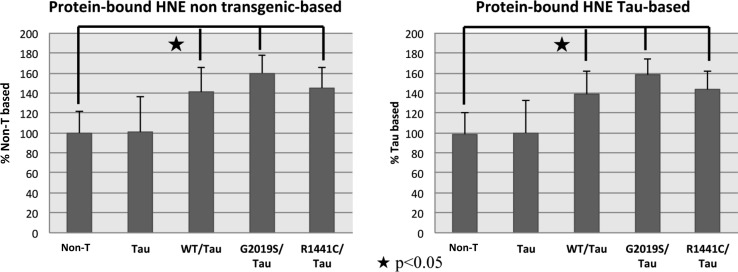
The last parameters of oxidative modification employed was protein-bound HNE. Figure 3 shows increased protein-bound HNE for LRRK2::Tau transgenic samples compared both with Non-T and with TAU that present nearly the same value: For the WT::Tau sample, the mean increase was 141% when compared to Non-T and 139% when compared to Tau samples; for G2019S/TAU the mean increase was 160% and 158%, and for R1441C::Tau the mean increase was 145% and 143%, respectively.
FIG. 3.
Bar graph of total protein carbonyls and total protein-bound 3NT in C. elegans transgenic and nontransgenic strains. Each value was compared before to the nontransgenic strain and after to the Tau strain, set as 100%. The symbols represent significant differences with p value<0.05.
Proteomics and redox proteomics analysis
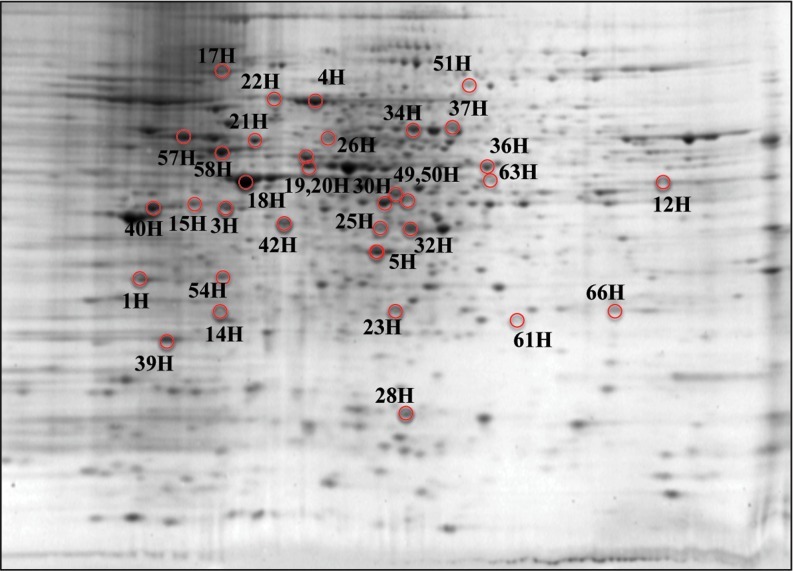
In the proteomics study, we analyzed the difference in protein expression levels, and for the redox proteomics analysis we identified proteins with altered protein-bound HNE levels. We considered for the analysis the seven groups previously described, showing first the comparison with nontransgenic samples and then with Tau transgenic samples as controls. The HNE blot values were normalized to the expression values on gels to yield specific protein-bound HNE levels per protein. The proteins with two or more percentage values listed were identified in two or more different spots. The proteomics and redox proteomics results are reported in Table 1 and shown in Figures 4, 5, and 6.
Table 1.
Proteins Identified by Proteomics and Redox Proteomics
|
Table 1A | |||||||
|---|---|---|---|---|---|---|---|
| Tau vs. Non transgenic | Protein name | Swiss prot accession number | Pb* | Peptides matched/searched | P value† | Fold increase/decrease | Function |
| 10E | Heat shock protein 25, isoform a | Q17849 | 1e-011 | 4/5 | 0.004 | 2.41 ↑ | Chaperone |
| 7E | 60S ribosomal protein L22 | P52819 | 5e-005 | 2/2 | 0.005 | 2.11 ↑ | Protein biosynthesis |
| 4E | Tubulin alpha-2 chain | P34690 | 1e-014 | 15/20 | 0.012 | 1.57 ↑ | Cell structure |
| 42H | 40S ribosomal protein SA | P46769 | 6e-014 | 8/14 | 0.040 | 3.47 ↑ | Protein biosynthesis |
| 49H 50H |
Fructose-bisphosphate aldolase 1 | P54216 | 1e-030 5e-015 |
9/19 6/9 |
0.014 0.042 |
2.76↑ 7.19 ↑ |
Energy metabolism |
| 51H | Propionyl-CoA carboxylase alpha chain, mitochondria | Q612F5 | 1e-007 | 2/3 | 0.045 | 0.42 ↓ | Energy metabolism |
| 37H | Aminopeptidase | Q27245 | 6e-006 | 5/13 | 0.040 | 7.70 ↑ | Protein degradation |
| 40H | Aspartic protease | O76830 | 2e-009 | 3/7 | 0.026 | 0.38 ↓ | Protein degradation |
|
Table 1B | |||||||
|---|---|---|---|---|---|---|---|
| LRRK2 wt::Tau vs. nontransgenic | Protein name | Swiss prot accession number | Pb | Peptides matched/searched | P value | Fold increase/decrease | Function |
| 19E | Glyceraldehyde-3-phosphate dehydrogenase | P17330 | 2e-009 | 6/48 | 0.004 | 0.60 ↓ | Energy metabolism |
| 16E | Dihydrolipoyl dehydrogenase | O17953 | 9e-012 | 4/7 | 0.005 | 0.45 ↓ | Pyruvate dehydrogenase complex |
| 25E | 60S ribosomal protein L7 | O01802 | 1e-013 | 2/3 | 0.005 | 0.48 ↓ | Protein biosynthesis |
| 38E | Bis(5'-nucleosyl)-tetraphosphatase | Q9U2M7 | 2e-009 | 2/2 | 0.008 | 2.01 ↑ | Induction of apoptosis |
| 2E | Histone H4 | P62784 | 3e-006 | 3/3 | 0.011 | 0.46 ↓ | Nucleosome structure |
| 22E | Protein F53F1.2, | P91997 | 3e-013 | 5/8 | 0.024 | 1.84 ↑ | Oxidoreductase activity |
| 23E | NADH ubiquinone oxidoreductase protein 2, | O01602 | 8e-012 | 4/8 | 0.027 | 3.27 ↑ | Energy metabolism |
| 12E | Nematode polyprotein allergen related protein | Q86S16 | 2e-010 | 2/4 | 0.028 | 0.37 ↓ | Development regulation |
| 41E | Vitellogenin-6 | P18948 | 2e-014 | 18/36 | 0.035 | 2.51 ↑ | Nutrient reservoir activity |
| 29E | Actin-depolymerizing factor 2 | Q07749 | 8e-013 | 6/15 | 0.037 | 0.56 ↓ | Cell structure |
| 5E | Myosin regulatory light chain 1 | P19625 | 4e-005 | 2/3 | 0.040 | 1.89 ↑ | Muscle structure |
| 3H | Disorganized muscle protein 1 | Q18066 | 4e-015 | 16/35 | 0.015 | 8.96 ↑ | Myofilaments structure |
| 42H | 40S ribosomal protein SA | P46769 | 6e-014 | 8/14 | 0.022 | 0.04 ↓ | Protein biosynthesis |
| 54H | Proteasome subunit alpha type-5 | Q95008 | 2e-016 | 10/21 | 0.042 | 0.18 ↓ | Protein degradation |
| 39H | Translationally-controlled tumor protein | Q93573 | 7e-013 | 4/15 | 0.014 | 36.98 ↑ | Microtubule stabilization |
|
Table 1C | |||||||
|---|---|---|---|---|---|---|---|
| LRRK2 wt::Tau vs. Tau | Protein name | Swiss prot accession number | Pb | Peptides matched/searched | P value | Fold increase/decrease | Function |
| 13E | Elongation factor 2 | P29691 | 2e-009 | 7/10 | 0.002 | 0.32 ↓ | Protein biosynthesis |
| 2E | Histone H4 | P62784 | 3e-006 | 3/3 | 0.003 | 0.46 ↓ | Nucleosome structure |
| 12E | Nematode polyprotein allergen related protein | Q86S16 | 2e-010 | 2/4 | 0.003 | 0.40 ↓ | Development regulation |
| 3E | V-type proton ATPase subunit A | Q9XW92 | 1e-012 | 24/45 | 0.011 | 3.66 ↑ | Lysosome structure/autophagy |
| 19E | Glyceraldehyde-3-phosphate dehydrogenase | P17330 | 2e-009 | 6/48 | 0.012 | 0.60 ↓ | Energy metabolism |
| 37E | Actin-depolymerizing factor 2, | Q07749 | 8e-010 | 5/7 | 0.019 | 6.64 ↑ | Cell structure |
| 34E 28E 36E |
Triosephosphate isomerase | Q10657 | 1e-010 7e-011 1e-010 |
7/17 5/7 3/4 |
0.022 0.004 0.022 |
2.59 ↑ 2.56 ↑ 6.60 ↑ |
Energy metabolism |
| 25E | 60S ribosomal protein L7 | O01802 | 1e-013 | 2/3 | 0.027 | 0.51 ↓ | Protein biosynthesis |
| 16E | Dihydrolipoyl dehydrogenase | O17953 | 9e-012 | 4/7 | 0.032 | 0.53 ↓ | |
| 9E | Fructose-1,6-biphosphatase protein 1 | Q9N2M2 | 6e-011 | 5/7 | 0.035 | 0.81 ↓ | Energy metabolism |
| 42E | Probable aconitate hydratase, mitochondrial | P34455 | 5e-014 | 13/24 | 0.044 | 0.54 ↓ | Energy metabolism |
| 33E | Proteasome subunit alpha type-1 | O44156 | 8e-013 | 2/2 | 0.047 | 1.90 ↑ | Protein degradation |
| 38E | Bis (5'-nucleosyl)-tetraphosphatase | Q9U2M7 | 2e-009 | 2/2 | 0.040 | 1.67 ↑ | Apoptosis induction |
| 1H | 60S ribosomal protein L7 | O01802 | 1e-030 | 2/4 | 0.035 | 5.69 ↑ | Protein biosynthesis |
| 3H | Disorganized muscle protein 1 | Q18066 | 4e-015 | 16/35 | 0.035 | 2.38 ↑ | Myofilament structure |
| 4H | Heat shock 70 kDa protein F, mitochondrial | P11141 | 5e-014 | 7/12 | 0.031 | 2.96 ↑ | Chaperone |
| 5H | beta-galactoside-binding lectin | P36573 | 1e-014 | 5/13 | 0.032 | 3.50 ↑ | Sugar binding |
|
Table 1D | |||||||
|---|---|---|---|---|---|---|---|
| LRRK2 G2019S::Tau vs. nontransgenic | Protein name | Swiss prot accession number | Pb | Peptides matched/searched | P value | Fold increase/decrease | Function |
| 4E | Tubulin alpha-2 chain | P34690 | 1e-014 | 15/20 | 0.0003 | 0.38 ↓ | Cell structure |
| 7E | 60S ribosomal protein L22 | P52819 | 5e-005 | 2/2 | 0.0003 | 7.31 ↑ | Protein biosynthesis |
| 32E | Calponin protein 4, | O44727 | 3e-010 | 2/3 | 0.005 | 1.71 ↑ | Muscle structure |
| 28E 34E |
Triosephosphate isomerase | Q10657 | 7e-011 1e-010 |
5/7 7/17 |
0.009 0.028 |
2.14 ↑ 1.87 ↑ |
Energy metabolism |
| 13E | Elongation factor 2 | P29691 | 2e-009 | 7/10 | 0.009 | 1.74 ↑ | Protein biosynthesis |
| 6E | Putative uncharacterized protein | A8WFJ0 | 5e-009 | 3/3 | 0.013 | 0.60 ↓ | ///// |
| 40E | Vitellogenin-6 | P18948 | 2e-014 | 19/36 | 0.026 | 3.17 ↑ | Nutrient reservoir activity |
| 8E | Fructose-1,6-biphosphatase protein 1 | Q9N2M2 | 6e-011 | 4/7 | 0.027 | 4.41 ↑ | Energy metabolism |
| 14E | Aconitate hydratase, mitochondrial | P34455 | 3e-013 | 6/8 | 0.039 | 2.06 ↑ | Energy metabolism |
| 38E | Bis (5'-nucleosyl)-tetraphosphatase | Q9U2M7 | 2e-009 | 2/2 | 0.040 | 1.91 ↑ | Apoptosis induction |
| 10E | Heat shock protein 25, isoform a | Q17849 | 1e-011 | 4/5 | 0.049 | 3.04 ↑ | Chaperone |
| 3H | Disorganized muscle protein 1 | Q18066 | 4e-015 | 16/35 | 0.035 | 3.05 ↑ | Myofilaments structure |
| 12H | Probable ornithine aminotransferase, mitochondrial | Q18040 | 2e-015 | 8/15 | 0.015 | 3.09 ↑ | Aminoacid biosynthesis |
| 23H | Proteasome subunit beta type-2 | P91477 | 2e-013 | 5/8 | 0.017 | 2.68 ↑ | Protein degradation |
| 58H | ATP synthase subunit beta, mitochondrial | P46561 | 1e-030 | 31/96 | 0.033 | 3.88 ↑ | Energy metabolism |
| 61H | Adenylate kinase | Q20140 | 4e-014 | 3/4 | 0.003 | 72.73 ↑ | Energy metabolism |
| 63H | Probable glutaryl-CoA dehydrogenase, mitochondrial | Q20772 | 8e-007 | 2/2 | 0.020 | 12.24 ↑ | Aminoacid metabolism |
| 28H | Fatty acid binding protein 2 | Q20224 | 3e-007 | 2/10 | 0.018 | 35.74 ↑ | Fatty acid transport |
| 32H | Guanine nucleotide binding protein beta 2 | Q21215 | 2e-010 | 8/29 | 0.036 | 5.87 ↑ | Signaling |
| 34H | ZK829.4 protein | Q23621 | 2e-006 | 13/33 | 0.023 | 2.60 ↑ | Oxidoreductase |
| 40H | Aspartic protease | O76830 | 2e-009 | 3/7 | 0.033 | 2.56 ↑ | Protein degradation |
|
Table 1E | |||||||
|---|---|---|---|---|---|---|---|
| LRRK2 G2019S::Tau vs. Tau | Protein name | Swiss prot accession number | Pb | Peptides matched/searched | P value | Fold increase/decrease | Function |
| 9E 8E |
Fructose-1,6-biphosphatase protein 1 | Q9N2M2 | 6e-011 6e-011 |
5/7 4/7 |
0.002 0.030 |
2.58↑ 1.82 ↑ |
Energy metabolism |
| 34E 28E 36E |
Triosephosphate isomerase | Q10657 | 1e-010 7e-011 1e-010 |
7/17 5/7 3/4 |
0.017 0.040 0.023 |
1.94 ↑ 1.83↑ 7.87 ↑ |
Energy metabolism |
| 27E 35E |
GTP-binding nuclear protein ran-1 | O17915 | 5e-010 7e-007 |
4/9 2/6 |
0.022 0.048 |
0.68 ↓ 0.43 ↓ |
Nuclear envelope assembly/cell division |
| 15E | Protein F01G10.1, | O17759 | 2e-011 | 11/32 | 0.024 | 0.67 ↓ | Nematode growth |
| 3E | V-type proton ATPase subunit A | Q9XW92 | 1e-012 | 24/45 | 0.026 | 2.27 ↑ | Lysosome structure, autophagy |
| 7E | 60S ribosomal protein L22 | P52819 | 5e-005 | 2/2 | 0.027 | 1.51 ↑ | Protein biosynthesis |
| 18E | Arginine kinase F46H5.3 | Q10454 | 1e-010 | 9/28 | 0.050 | 0.63 ↓ | Energy metabolism |
| 4H | Heat shock 70 kDa protein F, mitochondrial | P11141 | 5e-014 | 7/12 | 0.029 | 3.08 ↑ | Chaperone |
| 14H | 60S Ribosomal protein L6 | P47991 | 2e-009 | 3/4 | 0.003 | 6.56 ↑ | Protein biosynthesis |
| 15H | Probable inorganic pyrophosphatase 1 | Q18680 | 6e-013 | 9/14 | 0.047 | 2.49 ↑ | Cell growth, development |
| 17H | Vitellogenin-6 | P18948 | 2e-015 | 19/36 | 0.005 | 9.29 ↑ | Nutrients reservoir |
| 18H | Actin-1/3 | P10983 | 8e-012 | 11/21 | 0.046 | 9.62 ↑ | Cell structure |
| 19H 20H |
Enolase | Q27527 | 6e-016 7e-016 |
13/28 10/15 |
0.013 0.016 |
1.73↑ 1.88 ↑ |
Energy metabolism |
| 21H | V-type proton ATPase subunit B | Q19626 | 2e-015 | 15/31 | 0.035 | 3.57 ↑ | Lysosome structure, autophagy |
| 22H | Heat shock 70 kDa protein A | P09446 | 8e-015 | 14/26 | 0.049 | 19.11 ↑ | Chaperone |
| 23H | Proteasome subunit beta type-2 | P91477 | 2e-013 | 5/8 | 0.025 | 2.61 ↑ | Protein degradation |
| 25H | Eukaryotic translation initiation factor | A8WLV5 | 9e-011 | 2/2 | 0.002 | 5.85 ↑ | Translational process |
| 26H | Methylmalonate-semialdehyde dehydrogenase | P52713 | 3e-007 | 2/3 | 0.046 | 6.18 ↑ | Valine and pyrimidine metabolism |
| 28H | Fatty acid binding protein 2 | Q20224 | 3e-007 | 2/10 | 0.047 | 4.14 ↑ | Fatty acid transport |
| 30H | Probable arginine kinase | Q10454 | 1e-010 | 9/28 | 0.038 | 4.53 ↑ | Energy metabolism |
| 32H | Guanine nucleotide binding protein beta 2 | Q21215 | 2e-010 | 8/29 | 0.002 | 6.29 ↑ | Signaling |
| 34H | ZK829.4 protein | Q23621 | 2e-006 | 13/33 | 0.011 | 3.96 ↑ | Oxidoreductase |
| 36H | Citrate synthase mitochondrial | P34575 | 3e-009 | 10/32 | 0.027 | 2.85 ↑ | Energy metabolism |
| 37H | Aminopeptidase | Q27245 | 6e-006 | 5/13 | 0.018 | 16.31 ↑ | Protein degradation |
|
Table 1F | |||||||
|---|---|---|---|---|---|---|---|
| LRRK2 R1441C::Tau vs. non-transgenic | Protein name | Swiss prot accession number | Pb | Peptides matched/searched | P value | Fold increase/decrease | Function |
| 10E | Heat shock protein 25, isoform a | Q17849 | 1e-011 | 4/5 | 0.006 | 3.52 ↑ | Chaperone antioxidant |
| 26E 28E |
Triosephosphate isomerase | Q10657 | 2e-012 7e-011 |
9/34 5/7 |
0.008 0.028 |
1.61↑ 1.83 ↑ |
Energy metabolism |
| 1E 20E 21E |
Malate dehydrogenase | Q9UAV5 | 1e-011 2e-015 2e-014 |
3/5 8/31 4/6 |
0.036 0.014 0.031 |
3.31↑ 1.61↑ 1.45 ↑ |
Energy metabolism |
| 8E | Fructose-1,6-biphosphatase protein 1 | Q9N2M2 | 6e-011 | 4/7 | 0.019 | 2.42 ↑ | Energy metabolism |
| 2E | Histone H4 | P62784 | 3e-006 | 3/3 | 0.021 | 0.62 ↑ | Nucleosome structure |
| 41E | Vitellogenin-6 | P18948 | 2e-014 | 18/36 | 0.028 | 2.70 ↑ | Nutrient reservoir activity |
| 31E | Fatty-acid and retinol-binding protein 1 | P34382 | 0.0001 | 2/3 | 0.028 | 2.10 ↑ | Fatty acid transport |
| 7E | 60S ribosomal protein L22 | P52819 | 5e-005 | 2/2 | 0.032 | 3.50 ↑ | Protein biosynthesis |
| 32E | Calponin protein 4, | O44727 | 3e-010 | 2/3 | 0.043 | 1.56 ↑ | Muscle structure |
| 22E | Protein F53F1.2, | P91997 | 3e-013 | 5/8 | 0.045 | 1.63 ↑ | Oxidoreductase activity |
| 13E | Elongation factor 2 | P29691 | 2e-009 | 7/10 | 0.048 | 2.15 ↑ | Protein biosynthesis |
| 5H | Beta-galactoside-binding lectin | P36573 | 1e-014 | 5/13 | 1.7 E-06 | 0.03 ↓ | Sugar binding |
| 44H | Actin-1/3 | P10983 | 1e-030 | 20/47 | 0.020 | 3.20 ↑ | Cell structure |
| 49H 50H |
Fructose-bisphosphate aldolase 1 | P54216 | 1e-030 5e-015 |
9/19 6/9 |
0.003 | 9.84 ↑ | Energy metabolism |
| 57H | Tubulin beta-2 chain | P52275 | 9e-015 | 19/40 | 0.005 | 8.57 ↑ | Cell structure |
| 66H | Eukaryotic initiation factor 4A | P27639 | 3e-015 | 17/36 | 0.043 | 8.62 ↑ | Translational process |
|
Table 1G | |||||||
|---|---|---|---|---|---|---|---|
| LRRK2 R1441C::Tau vs. Tau | Protein name | Swiss prot accession number | Pb | Peptides matched/searched | P value | Fold increase/decrease | Function |
| 26E | Triosephosphate isomerase | Q10657 | 2e-012 | 9/34 | 0.003 | 1.28 ↑ | Energy metabolism |
| 3E | V-type proton ATPase catalytic subunit A | Q9XW92 | 1e-012 | 24/45 | 0.007 | 2.19 ↑ | Lysosome structure, autophagy |
| 17E | Fructose-bisphosphate aldolase 1 | P54216 | 5e-014 | 4/7 | 0.009 | 1.69 ↑ | Energy metabolism |
| 35E | GTP-binding nuclear protein ran-1 | O17915 | 7e-007 | 2/6 | 0.016 | 1.92 ↑ | Nuclear envelope assembly/cell division |
| 2E | Histone H4 | P62784 | 3e-006 | 3/3 | 0.022 | 0.63 ↓ | Nucleosome structure |
| 24E | 60S ribosomal protein L7 | O01802 | 6e-013 | 4/7 | 0.030 | 1.89 ↑ | Protein biosynthesis |
| 30E | Nucleoside diphosphate kinase | Q93576 | 2e-009 | 2/4 | 0.035 | 0.33 ↓ | Synthesis of nucleoside triphosphates |
| 33E | Proteasome subunit alpha type-1 | O44156 | 8e-013 | 2/2 | 0.039 | 1.87 ↑ | Protein degradation |
| 31E | Fatty-acid and retinol-binding protein 1 | P34382 | 0.0001 | 2/3 | 0.048 | 1.69 ↑ | Fatty acid transport |
| 5E | Myosin regulatory light chain 1 | P19625 | 4e-005 | 2/3 | 0.041 | 1.78 ↑ | Muscle structure |
| 1H | 60S Ribosomal protein L7 | O01802 | 1e-030 | 2/4 | 0.031 | 12.12 ↑ | Protein biosynthesis |
| 14H | 60S Ribosomal protein L6 | P47991 | 2e-009 | 3/4 | 0.021 | 15.30 ↑ | Protein biosynthesis |
| 17H | Vitellogenin-6 | P18948 | 2e-015 | 19/36 | 0.0002 | 6.04 ↑ | Nutrient reservoir |
| 19H 20H |
Enolase | Q27527 | 6e-016 7e-016 |
13/28 10/15 |
0.022 0.041 |
2.05 ↑ 2.00 ↑ |
Energy metabolism |
| 25H | Eukaryotic translation initiation factor | A8WLV5 | 9e-011 | 2/2 | 0.046 | 2.37 ↑ | Translational process |
| 42H | 40S ribosomal protein SA | P46769 | 6e-014 | 8/14 | 0.040 | 2.62 ↑ | Protein biosynthesis |
| 18H | Actin-1/3 | P10983 | 8e-012 | 11/21 | 0.004 | 4.02 ↑ | Cell structure |
| 39H | Translationally-controlled tumor protein | Q93573 | 7e-013 | 4/15 | 0.034 | 30.72 ↑ | Microtubule stabilization |
| 40H | Aspartic protease | O76830 | 2e-009 | 3/7 | 0.022 | 26.12 ↑ | Protein degradation |
Proteins altered in expression are marked with “E” after the identification number, whereas proteins altered in protein-bound HNE levels are marked with “H” after the identification number. The seven groups of analysis are divided in seven parts of Table 1 marked with alphabetical letters A to G.
The p value listed is the significance of the altered expression or protein-bound HNE levels relative to control samples with p<0.05. *Pb represents the probability of a false identity associated with the identification of each protein using the SEQUEST search algorithm.
FIG. 4.
Bar graph of total protein-bound HNE in C. elegans transgenic and nontransgenic strains. Each value was compared before to nontransgenic strain and after to Tau strain, set as 100%. The symbols represent significant differences with p value<0.05.
FIG. 5.
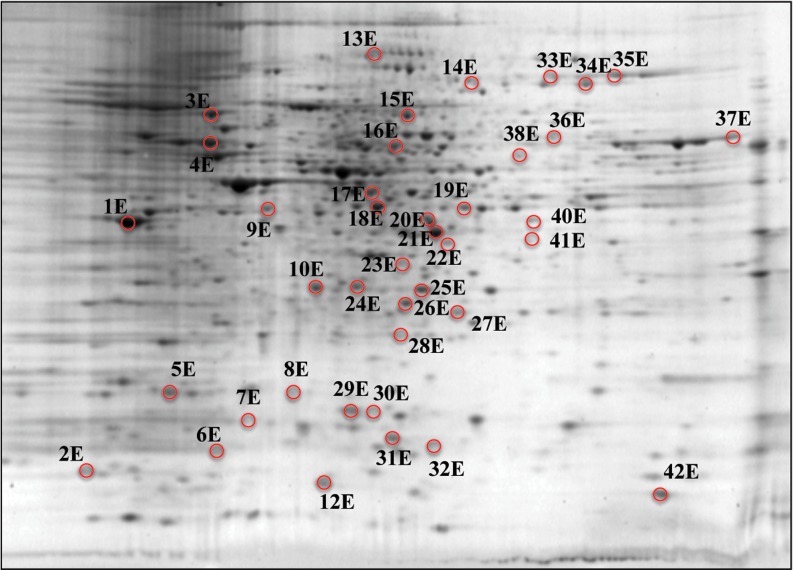
Representative 2D-gel with all the proteins identified in the expression proteomics study. The proteins are represented with identification numbers followed by the letter “e.” Consult Table 1 A–G for results of comparisons among the seven groups. (To see this illustration in color the reader is referred to the web version of this article at www.liebertonline.com/ars).
FIG. 6.
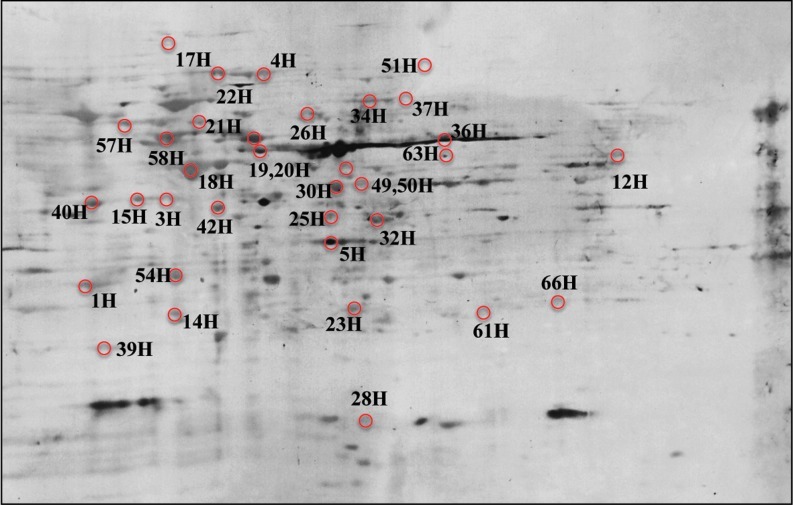
Representative 2D-gel with all the proteins identified in the redox proteomics study. The proteins are represented with identification numbers followed by the letter “h.” Consult Table 1 A–G for results of comparisons among the seven groups. (To see this illustration in color the reader is referred to the web version of this article at www.liebertonline.com/ars).
Tau vs. nontransgenic
In the comparison of Tau samples with Non-T we identified a significant increase in expression levels of three proteins in tau transgenic (Table 1A; Fig. 4). These proteins (with % increased values) are heat shock protein 25, isoform a (240%), 60S ribosomal protein L22 (211%), and tubulin alpha-2 chain (157%).
In the analysis of protein-bound HNE levels (Table 1A; Figs. 5 and 6), we identified for Tau samples compared with Non-T increased levels of three proteins: 40S ribosomal protein SA (347%), fructose-bisphosphate aldolase 1 (276% and 719%), and aminopeptidase (770%). We show also decreased levels of two proteins: propionyl-CoA carboxylase alpha chain mitochondrial (42%) and aspartic protease (38%).
LRRK2 WT::Tau vs. nontransgenic
The proteomics comparison between LRRK2 WT::Tau with Non-T samples identified increased expression levels of five proteins (Table 1B; Fig. 4): Bis(5'-nucleosyl)-tetraphosphatase (201%), protein F53F1.2 (184%), vitellogenin-6 (251%), NADH ubiquinone oxidoreductase protein 2 (327%), and myosin regulatory light chain 1 (189%); decreased expression levels of six proteins were identified: glyceraldehyde-3-phosphate dehydrogenase (60%), dihydrolipoyl dehydrogenase (45%), 60S ribosomal protein L7 (48%), histone H4 (46%), nematode polyprotein allergen related protein (37%), and actin-depolymerizing factor 2, isoform c (56%).
The analysis of protein-bound HNE levels identified increased levels in two proteins (Table 1B; Figs. 5 and 6): disorganized muscle protein 1 (846%) and transitionally controlled tumor protein (3968%); and decreased protein-bound HNE levels of two proteins: 40S ribosomal protein SA (4%) and proteasome subunit alpha type-5 (18%).
LRRK2 G2019S::Tau vs. nontransgenic
In the expression levels analysis between LRRK2 G2019S::Tau with Non-T samples, we identified nine proteins with increased levels (Table 1C; Fig. 4): 60S ribosomal protein L22 (731%), calponin protein 4 (171%), triosephosphate isomerase (214%, 187%), elongation factor 2 (214%), vitellogenin-6 (317%), fructose-1,6-biphosphatase protein 1 (441%), probable aconitate hydratase mitochondrial (206%), bis(5'-nucleosyl)-tetraphosphatase (191%), and heat shock protein 25, isoform a (304%); and decreased levels only for tubulin alpha-2 chain (38%).
The redox proteomics analysis identified 10 proteins with increased protein-bound HNE levels (Table 1C; Figs. 5 and 6): disorganized muscle protein 1 (305%), probable ornithine aminotransferase mitochondrial (309%), proteasome subunit beta type-2 (268%), ATP synthase subunit beta mitochondrial (388%), probable adenylate kinase isoenzyme F38B2.4 (727%), probable glutaryl-CoA dehydrogenase, mitochondrial (1222%), fatty acid binding protein 2 homologue (357%), guanine nucleotide binding protein beta 2 (587%), ZK829.4 protein (260%), and aspartic protease (256%).
LRRK2 R1441C::Tau vs. nontransgenic
In the expression levels analysis between LRRK2 R1441C::Tau with Non-T samples, we identified increased levels of 10 proteins (Table 1D; Fig. 4): heat shock protein 25 isoform a (352%), triosephosphate isomerase (161%, 183%), malate dehydrogenase (161%, 145%, 331%), fructose-1,6-biphosphatase protein 1 (242%), vitellogenin-6 (270%), fatty acid and retinol-binding protein 1 (210%), 60S ribosomal protein L22 (350%), calponin protein 4 (156%), protein F53F1.2 (163%), and elongation factor 2 (215%); and decreased expression levels only for histone H4 (62%).
The redox proteomics analysis of protein-bound HNE levels identified increased levels in four proteins (Table 1D; Figs. 5 and 6): actin-1/3 (320%), fructose-bisphosphate aldolase 1 (984%), tubulin beta-2 chain (857%), and eukaryotic initiation factor 4A (862%); and decreased protein-bound HNE levels only for beta-galactoside-binding lectin (3%).
LRRK2 WT::Tau vs. Tau
The proteomics comparison between LRRK2 WT::Tau transgenic with Tau samples identified decreased expression levels for eight proteins (Table 1E, Fig. 4): elongation factor 2 (32%), histone H4 (46%), nematode polyprotein allergen related protein (40%), glyceraldehyde-3-phosphate dehydrogenase (60%), 60S ribosomal protein L7 (51%), dihydrolipoyl dehydrogenase (53%), fructose-1,6-biphosphatase protein 1 (81%), and probable aconitate hydratase mitochondrial (54%); and increased levels for five proteins: V-type proton ATPase (366%), actin-depolymerizing factor 2, isoform c (664%), triosephosphate isomerase (259%, 660%), proteasome subunit alpha type-1 (190%), and bis(5'-nucleosyl)-tetraphosphatase (167%).
The redox proteomics analysis of protein-bound HNE levels identified increased levels of four proteins (Table 1E; Figs. 5 and 6): ribosomal protein L7 (569%), disorganized muscle protein 1 (238%), heat shock 70 kDa protein F mitochondrial (296%), and beta-galactoside-binding lectin (350%).
LRRK2 G2019S::Tau vs. Tau
In the proteomics expression levels analysis between LRRK2 G2019S::Tau with Tau samples, we identified increased levels of four proteins (Table 1F; Fig. 4): triosephosphate isomerase (194%, 797%, 183%), V-type proton ATPase (227%), 60S ribosomal protein L22 (151%), and fructose-1,6-biphosphatase protein 1 (182%); and decreased expression levels of three proteins: protein F01G10.1 (67%), GTP-binding nuclear protein ran-1 (68%), and arginine kinase (63%).
The redox proteomics analysis of protein-bound HNE identified increased levels of 17 proteins (Table 1F; Figs. 5 and 6): heat shock 70 kDa protein F mitochondrial (308%), ribosomal protein L6 (656%), probable inorganic pyrophosphatase 1 (249%), vitellogenin-6 (929%), actin-1/3 (923%), enolase (173%, 188%), V-type proton ATPase (357%), heat shock 70 kDa protein A (191%), proteasome subunit beta type-2 (261%), eukaryotic translation initiation factor (585%), methylmalonate-semialdehyde dehydrogenase (618%), fatty acid binding protein 2 homologue (414%), probable arginine kinase (453%), guanine nucleotide binding protein beta 2 (629%), ZK829.4 protein (396%), citrate synthase mitochondrial (285%), and aminopeptidase (163%).
LRRK2 R1441C::Tau vs. Tau
The proteomics comparison between LRRK2 R1441C::Tau transgenic samples with Tau samples identified increased expression levels of eight proteins (Table 1G; Fig. 4): triosephosphate isomerase (128%), V-type proton ATPase (219%), fructose-bisphosphate aldolase 1 (169%), GTP-binding nuclear protein ran-1 (192), 60S ribosomal protein L7 (189%), proteasome subunit alpha type-1 (187%), fatty-acid and retinol-binding protein 1 (169%), and myosin regulatory light chain 1 (178%); and decreased expression levels of two proteins: histone H4 (63%) and nucleoside diphosphate kinase (33%).
The redox proteomics analysis of protein-bound HNE identified increased levels of nine proteins (Table 1G; Figs. 5 and 6): ribosomal protein L7 (1221%), ribosomal protein L6 (1530%), vitellogenin-6 (604%), enolase (210%, 200%), eukaryotic translation initiation factor (237%), 40S ribosomal protein SA (262%), actin-1/3 (402%), transitionally-controlled tumor protein (307%), and aspartic protease (261%).
Redox proteomics data validation
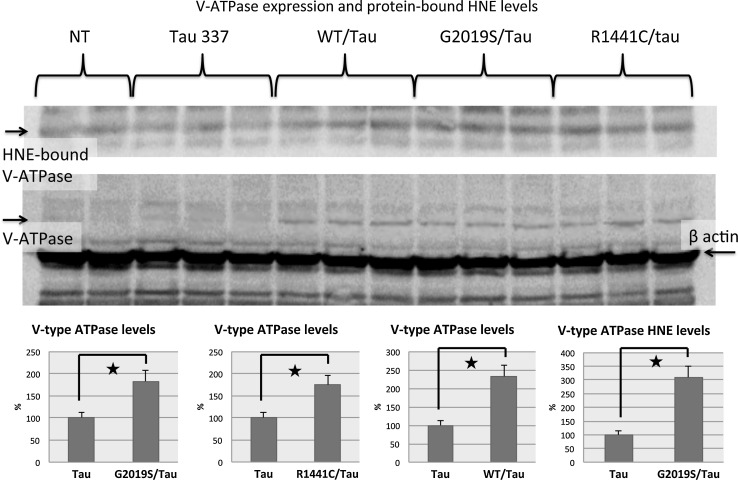
To verify the proteomics and redox proteomics data, a validation study on expression levels and protein-bound HNE levels of V-type proton ATPase, identified by mass spectrometry analysis in three different groups of analysis, was performed and is shown in Figure 7. V-type proton ATPase was identified with increased expression levels during proteomics analysis in the comparison of LRRK2 G2019S::Tau vs. Tau and LRRK2 R1441C::Tau vs. Tau, and with decreased expression levels in the comparison of LRRK2 WT::Tau vs. Tau. In the redox proteomics analysis, V-type proton ATPase was identified as having increased protein-bound HNE in the comparison of LRRK2 G2019S::Tau vs. Tau. We measured by Western blot the levels of V-type proton ATPase in these comparisons and the results are shown in Figure 7. They demonstrate the same trend seen in proteomics analysis for all the comparison groups considered. In detail we show an increase of 183% for LRRK2 G2019S::Tau vs. Tau and of 174% for LRRK2 R1441C::Tau vs. Tau, and an increase of 233% for LRRK2 WT::Tau vs. Tau, confirming the proteomics data.
FIG. 7.
Representative 2D Western blot with all the proteins identified with altered HNE levels using redox proteomics analysis. (To see this illustration in color the reader is referred to the web version of this article at www.liebertonline.com/ars.)
To validate redox proteomics analysis we immunoprecipitated V-type proton ATPase protein and we performed Western blot analysis probing with anti-HNE antibody to measure specific protein-bound HNE levels. As shown in Figure 7, we found increased protein-bound HNE levels of V-type proton ATPase (310%) in the comparison of LRRK2 G2019S::Tau vs. Tau as we obtained by redox proteomics analysis. The validation studies give confidence in the proteomics identifications presented here.
Increasing autophagic flux reduces deficits in C. elegans expressing Tau or LRRK2::Tau
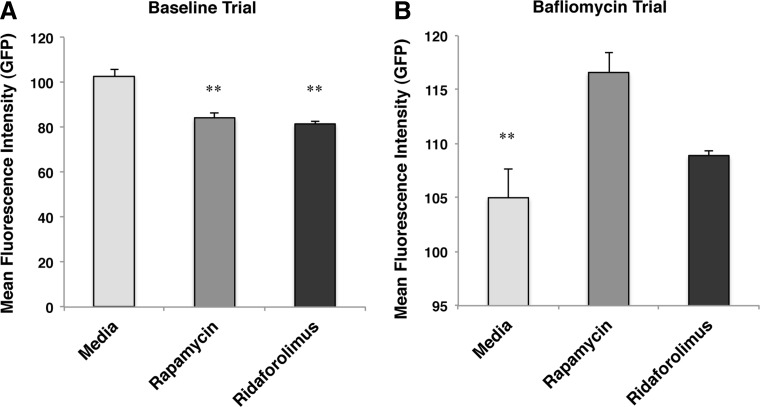
Ridaforolimus is a rapamycin analog able to inhibit mTOR signaling and induce autophagy. To test whether the motor deficits in the C. elegans lines were sensitive to the state of autophagy, we examined the effects of ridaforolimus. First, we measured the effect of ridadorolimus on autophagic flux, comparing the results to that observed with rapamycin, a well documented enhancer of autophagic flux. We measured autophagic flux in HEK293 cells expressing mCherry-GFP-LC3 at baseline and in the presence of bafliomycin, a lysosomal inhibitor. In Figures 8A and 8B, we show that ridaforolimus (200 nM) increases autophagic flux to the same degree as rapamycin, as evidenced by a loss of GFP-LC3 signal at baseline and recovery of GFP-LC3 signal in the presence of bafliomycin.
FIG. 8.
Proteomics data validation experiments. Top: Western blot of expression levels and protein-bound HNE levels of V-type proton ATPase in transgenic and nontransgenic C. elegans strains. Bottom: Bar graphs of significant differences between the groups analyzed are shown.
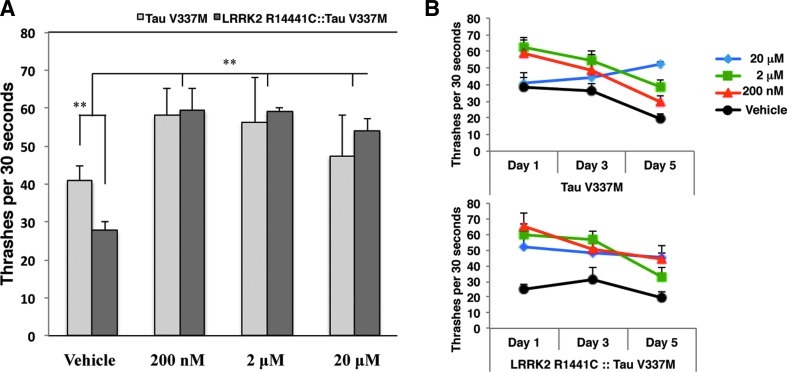
In order to test the role of autophagic pathway impairment in our C. elegans transgenic models, we administered ridaforolimus to all the strains and measured the thrashes for 30 seconds at day one of adulthood. In Figure 9A we show that ridaforolimus at different concentrations consistently improved thrash number in Tau and LRRK2 R1441C::Tau transgenic strains (as well as in the other transgenic models; data not shown) protecting motor neuron functionality through the restoration of autophagic pathway lost by mutant LRRK2 and tau insertion. In Figure 9B, we show that ridaforolimus maintains its protective activity, at various concentrations, even at 5 days of nematode adulthood, supporting a stable re-establishment of macroautophagy.
FIG. 9.
Comparision of GFP intensity between vehicle, rapamycin (200 nM) and ridaforolimus (200 nM) without (A) and with (B) bafliomycin; *p<0.05, **p<0.01, ***p<0.001.
Discussion
Increased expression of oxidative markers indexed by increased malondialdehyde-lysine (MDAL) and HNE, increased oxidative damage to selected proteins, with a particular involvement of energy metabolic pathways, have been observed in the cerebral cortex in sporadic PD even at relatively early stages of the disease (9, 10, 12, 21). Past studies, conducted on the involvement of oxidative damage linked to LRRK2 mutations in PD onset and progression, were associated with the G2019S LRRK2 mutation (22). G2019S, and to a lesser degree R1441G, seem crucial for neurite outgrowth, and their expression in primary cortical neurons leads to dramatic reductions in neuritic length and branching of axons and dendrites, loss of dopaminergic neurons, accumulation of abnormal proteins, such as alpha-synuclein and hyper-phosphorylated Tau, together with ubiquitin (17, 43, 48, 61).
A common feature in LRRK2 transgenic models is the presence of Tau alterations that are also present in a subset of LRRK2 patients. An attractive hypothesis is that Tau is one of the downstream targets of LRRK2 kinase activity (31,55). It was proposed that LRRK2 dysfunction impacts on Tau post-translational processing and compartmentalization and thus on its functional roles (14).
In our study we analyzed transgenic C. elegans strains expressing human LRRK2 WT, G2019S, and R1441C, and TauV337M genes to study the impact of human WT and mutated LRRK2 together with Tau on protein expression and oxidation. The results of total protein oxidative modifications show a general increase of lipid peroxidation indexed by protein-bound HNE in LRRK2 and TauV337M transgenic (TAU) strains compared to Tau or nontransgenic (Non-T) strains (Fig. 3). These data suggest increased oxidative stress levels involving lipid bilayer components of membranes with the formation of HNE end products that reacts directly with proteins by Michael addition (7). Moreover the presence of WT or mutated LRRK2 with Tau increase the total levels of protein-bound HNE, suggesting a direct or indirect interaction between these proteins as was previously postulated (31, 38, 43, 55).
We also founded a general increase in the levels of protein-bound 3NT (i.e., increased nitration levels) in LRRK2 and Tau strains compared to Non-T, but not to Tau. Protein carbonyls did not show significant differences between LRRK2 and Tau transgenic and nontransgenic strains. Considering the total results on protein oxidation, we chose to focus the redox proteomics analysis on protein-bound HNE in parallel with expression proteomics, mainly because the elevation of this oxidative stress marker was larger than that for 3NT.
Expression and redox proteomics are powerful tools that have been applied to neurodegenerative disorders such as Alzheimer's disease (4, 53) and has been used to identify oxidatively modified proteins in a C. elegans model of AD (6). The comparison of the Non-T with Tau strains shows few changes in protein expression, demonstrating slight alteration due to Tau insertion in the genome. The redox proteomics analysis show alteration of proteins mostly involved in energy metabolism or proteolytic pathways, but the undefined trend of oxidation towards one or the other group and the relative small number of proteins identified establish again a minor impact of the presence of Tau on cell redox status as seen alone for total oxidation levels. Accounting for these results, we propose that the overexpression of Tau by itself does not determine significant alterations on C. elegans protein expression and oxidation levels; however, this condition might change in the presence of others factors that could interact with Tau.
To test the hypothesis of association between WT and mutated LRRK2 and Tau in PD onset and development, we analyzed by proteomics and redox proteomics transgenic C. elegans strains containing human LRRK2 (WT, G2019S, and R1441C) and Tau genes compared to either Tau or Non-T. The LRRK2 WT::Tau strain demonstrated changes in the expressions levels of 11 and 12 proteins in the comparison with Non-T and Tau strains, respectively. Indeed, both the comparison groups displayed alterations of similar molecular pathways. Interestingly, many of the altered proteins, such as glyceraldehyde-3-phosphate dehydrogenase, are involved in the glucose metabolism pathway, suggesting glycolysis as a consistent target of LRRK2::Tau interaction in the C. elegans model. However, proteomics data show also the common alteration of proteins involved in apoptotic processes as well as nucleosome and ribosome structure, suggesting these specific pathways are also involved in the alterations. The altered protein-bound HNE levels were identified in, among the other proteins, ribosomal protein L7 and heat shock 70 kDa protein in the comparison of WT::TAU with TAU and in proteasome subunit alpha type-5 and 40S ribosomal protein SA in the comparison with Non-T. Some of the oxidized proteins identified by redox proteomics belong to molecular pathways also altered in the expression proteomics analysis. However, in both the groups of matching there is no clear trend of oxidation.
The overall proteomics and redox proteomics data suggest that the expression of LRRK2 WT with Tau does not have a strong impact on expression levels and protein oxidation in C. elegans. LRRK2 WT by itself, and in correlation with Tau is not able to cause large changes in the proteome, and the supposed downstream control that LRRK2 has on Tau PTM seems not to be altered when LRRK2 is not mutated. However, alteration of proteins involved in protein folding, biosynthesis, and degradation might depend by LRRK2 expression only and correlates with previous studies. A number of reports suggest a role for LRRK2 in regulating neuronal responses to stress, and in C. elegans it was reported that WT LRRK2 protects against mitochondrial dysfunction, but disease-related mutants of LRRK2 appear to produce responses that range from less protection to overt enhancement of toxicity (47). The co-expression of mutated LRRK2 (G2019S or R1441C) with Tau showed a higher effect on protein expression alterations and protein oxidation than WT::Tau co-expression for the number of proteins and molecular pathways involved and mainly for the trend of oxidatively modified proteins that clearly shifted towards the mutated forms. The expression profile of the C. elegans strain expressing LRRK2 G2019S::Tau compared both to Tau and Non-T strains demonstrate the alteration of 8 and 11 proteins, respectively, with several common identifications. In fact, changes in expression of proteins involved with energy metabolism are highly shared in both matching groups. This result was obtained also in the comparison with WT::Tau, confirming once again that LRRK2::Tau interaction consistently affects glucose metabolic pathways. Indeed, other proteins in the comparison of mutated LRRK2::Tau with Tau or Non-T display expression levels alterations such as ribosomal proteins, and interestingly cell structure proteins. These two groups of proteins are both shared between the two mutations considered and in common with the previous comparison group. The expression proteomics results show that the alterations in expression are mainly induced by the co-expression of LRRK2 (mutated or WT) with Tau that largely affect similar molecular pathways such as metabolic pathways, cell structure, or protein biosynthesis and degradation. However, a substantial difference between the co-expression of WT::Tau and mutated::Tau compared to control groups is that the expression levels mostly shift towards protein overexpression in presence of the mutations.
The redox proteomics results regarding protein-bound HNE show increased protein oxidative modification in strains with co-expression of mutated/tau compared to Tau, or Non-T. For G2019S::Tau we found increased HNE levels in 17 proteins compared to Tau, and 10 proteins in the comparison with Non-T. The number of proteins oxidatively modified identified by redox proteomics is consistent with the results of the total HNE levels, demonstrating increased lipid peroxidation in the presence of G2019S::Tau. In DA neurons, it was reported that the LRRK2 G2019S dominant mutant causes several dendritic defects, including Tau mislocalization, Tau hyperphosphorylation, and dendrite degeneration with the same behavior of Tau overexpression in DA neurons. Moreover, the co-expression of G2019S and Tau leads to neuronal loss and lack of microtubules in dendrites more intensely than either one alone (31). Hence, as Lin et al. suggested previously (31), our data support the notion that the interaction of mutated LRRK2 with Tau induces and enhances the detrimental activities of Tau in neurodegeneration.
Based on the identified proteins in expression proteomics analysis, one of the main pathways affected by G2019S::Tau expression is energy metabolism. These data correlate with previous studies on PD models about the oxidation of energy metabolism enzymes such as enolase (21), strengthening our C. elegans model as an appropriate PD model.
The R1441C/TAU co-expression present similar results to those to G2019S::Tau co-expression, suggesting common mechanisms that lead to increased oxidative stress.
The R1441C::Tau co-expression identified HNE modification of 4 and 9 proteins in the comparison with Tau and Non-T strains, respectively. The oxidative modifications affect metabolic proteins, but also like other LRRK2 transgenic strains, structural proteins and protein involved in protein biosynthesis and degradation are affected.
The redox proteomics data are consistent with the notion that the trend of oxidation is a result of the interaction of mutated LRRK2 and Tau. Mutated LRRK2 might increase its activity in a mechanism of gain of function (58), triggering the activation of several downstream signals that lead to Tau hyperphosphorylation and increased oxidative stress.
Analyzing specific protein alterations, this study reveals the different expression and the increased protein-bound HNE levels of several proteins involved in the autophagy process during the co-expression of LRRK2 WT or mutated and Tau. Recently, autophagy is gaining attention for its potential contribution to the pathogenesis of several neurodegenerative diseases (36, 41, 46). The increase in autophagic vacuoles in the substantia nigra of PD brains (5) supports the idea that autophagy is involved in PD. Several factors are implicated in autophagy alterations in PD, and recently Plowey et al. (42) suggested a strong link between LRRK2 and autophagy, consistent with an active role for autophagy induced by the LRRK2 pathological mutant. Expression of LRRK2 G2019S or R1441C leaded to the accumulation of increased autophagic vacuoles (2, 42). Here we found alteration of V-type proton ATPase, a vacuolar proton pump involved in lysosomal autophagy, in the presence of LRRK2::Tau co-expression. Supporting our results, it was previously reported that increased oxidative stress could affect the permeability of the lysosomal membrane or damage the lysosomal membrane by oxidizing proteins (60). Moreover, the findings that lysosomal malfunction (39) and mutations in ATP13A2, a lysosomal ATPase, led to a failure of autophagy execution in PD (15, 44) strengthen the correlation between LRRK2, autophagy and PD.
The further use of ridaforolimus supported our proteomics and redox proteomics results and is consistent with the notion of autophagic impairment in LRRK2/Tau transgenic strains. Ridaforolimus is a rapamycin analog and a macroautophagy activator through its inhibition of mTOR signaling (45, 52). We report in the present study a restoration of motor neuron functionality in the presence of ridaforolimus, through improved autophagy, in young and aged strains. Thus, by testing ridaforolimus in our nematode transgenic models, we confirmed both the involvement of autophagy impairment in mutated LRRK2 and Tau-modulated neurodegeneration and the potent protective effect of ridaforolimus against disease-associated mutant protein toxicity.
We also found in our study the altered expression and oxidative modification of several proteasome subunits during co-expression of LRRK2 WT or mutated and Tau. This result suggests that not only the autophagic way of protein degradation might be impaired by LRRK2 expression, but also the ubiquitin/proteasome system leading to the formation of protein aggregates as seen for the mutations of parkin (18) and UCH- L1 (35) components of the system. Increased oxidative stress, associated with depletion of ATP, is thought to contribute to the reduction of proteasome activity and aggregation of abnormal proteins (1, 32). The energy-related proteins identified in this study are consistent with this notion. It is generally believed that autophagy and the ubiquitin/proteasome system operate in concert playing a vital role in homeostasis and the cellular responses to stresses, such as endoplasmic reticulum stress, and mitochondrial damage.
Further, we identified various mitochondrial proteins, some energy-related and some not, such as NADH ubiquinone oxidoreductase protein 2 (LRRK2 specific) or mitochondrial HSP 70, that might suggest alterations and injury of the mitochondria. Dysfunction of mitochondria is implicated in the pathophysiology of PD (49, 56), and recent studies raised the possibility that LRRK2 modulates mitochondrial function and impairment (47). The increased expression of NADH ubiquinone oxidoreductase protein 2 is particularly interesting because it provides a mechanism through which LRRK2 might protect against rotenone, which we demonstrated previously (47). Upregulation of NADH ubiquinone oxidoreductase protein 2 would increase activity of complex I of the electron transport chain, which could protect against the deleterious effects of rotenone. Damaged mitochondria are a well-known source of pro-oxidant species, and mitophagy is thought to be a crucial process to protect cells from mitochondrial leakage of ROS/RNS. We suppose that the alterations of mitochondrial proteins could probably be related to changes in mitochondrial turnover (mitophagy) that are implicated for parkin and PINK1, and might be relevant to LRRK2.
We also reported cell structure impairment linked to LRRK2 expression for the alterations on actin depolymerizing factor 2 in LRRK2 WT::Tau, actin 1/3 and tubulin beta-2 chain in LRRK2 G2019S:: or R1441C::Tau. These findings nicely correlate with a recent study by Chen et al. showing regulation of LRRK2 by rac1 and linkage to actin remodeling (8). Moreover, three members of the ezrin/radixin/moesin (ERM) family, which regulate actin cytoskeleton reorganization and membrane curvature dynamics, are among the few recognized substrates of LRRK2 (3).
Another very interesting result concerns the involvement of protein biosynthetic machinery in the detrimental influence of LRRK2 and Tau co-expression in C. elegans transgenic strains. We identified here alterations of the eukaryotic translation initiation factor LRRK2 G2019S:: and R1441C::Tau and of eukaryotic initiation factor 4A LRRK2 R1441C::Tau that are part of the translational machinery. Moreover, we show altered expression and increased oxidation of several ribosomal subunits in most of the transgenic strains analyzed. A previous study by Imai et al. (25) reported 4E-BP, an interactor of the eukaryotic protein translation initiation factor eIF4E that in turn binds to capped mRNA species promoting their translation, as a potential substrate of LRRK2. LRRK2 phosphorylation of 4E-BP modulates the binding to eIF4E, controlling the repression of translation.
In conclusion, the co-expression of human WT LRRK2 and Tau lead to protein expression changes without altering the redox status, while mutated LRRK2 leads to increased protein oxidation. This result suggests that LRRK2 might interact with Tau through the activation of several downstream signals and the mutation could enhance LRRK2 effects on tau PTM and compartmentalization. Interestingly, we show that LRRK2 expression affects protein biosynthesis and degradation, and the mutations appear to enhance this unfavorable outcome. Involvement of protein degradation pathways, with a special attention on autophagy, in LRRK2 mediated degenerative process is gaining wide interest as a PD causative agent and represents a fascinating result of our study.
Materials and Methods
Chemicals
Ridaforolimus was supplied by ARIAD Pharmaceuticals (Cambridge, MA). All other chemicals were purchased from Sigma (Sigma, St Louis, MO), unless otherwise indicated.
Nematode culture and transgenic crossing
All LRRK2 lines were generated in the Wolozin Laboratory as previously described (47). The Tau V337M strain was generated and generously shared by Dr. Brian Kraemer of the University of Washington. The Bristol strain N2 was the nontransgenic line. Nematodes strains were grown at 20°C on standard NGM plates coated with a thin layer of OP50 bacteria. Crosses were produced by mating male LRRK2 lines with hermaphrodites from the Tau V337M lines. Progeny were examined by PCR for the presence of LRRK2 and propagated until homozygotes could be identified (Fig. 10). Liquid thrashing assays were performed in 20 (l of water on the lid of a plastic Petri dish. Age-synchronized worms were allowed to settle, and thrashes were counted for three 30 second intervals (n=10 per group).
FIG. 10.
Age synchronized nematodes cultured with varied doses of ridaforolimus. (A) Movement (thrashing) assessed on day 1 of adulthood. **p<0.01; (B) movement (thrashing) assessed on day 1, 3, and 5 of adulthood. (To see this illustration in color the reader is referred to the web version of this article at www.liebertonline.com/ars).
Primer sets
LRRK2 - forward - 5’-ATG GCT AGT GGC AGC TGT CAG-3’
LRRK2 - reverse - 5’-GAG TCC AAG ACG ATC AAC AGA-3’
Tau - forward - 5’-CAA GCT CGC ATG GTC AGT AA-3’
Tau - reverse - 5’-TTC TCA GTG GAG CCG ATC TT-3’
Sample preparation
The lyophilized samples were manually homogenized in ice-cold buffer (10 mM Tris pH 8, 0.32 M Sucrose, 0.1 mM MgCl2, 0.1 mM ethylenediaminetetraacetic acid, 10 μg/ml leupeptin, 10 μg/ml pepstatin, 10 μg/ml aprotinin) and sonicated for 10 sec on ice. Protein concentration was determined by the BCA method (Pierce, Rockford, IL). Proteins (150 (g) were precipitated in 15% final concentration of trichloroacetic acid for 10 min in ice. Samples were then spun down at 14000 rpm for 5 min, and precipitates were washed in ice-cold ethanol-ethyl acetate 1:1 solution four times.
The final pellets were dissolved in 200 μl of 8 M urea, 2% CHAPS, 2 M thiourea, 20 mM dithiothreitol, 0.2% of ampholytes (Bio-Rad, Hercules, CA) and bromophenol blue, incubated at room temperature for 90 min and sonicated for 5 sec.
Protein oxidation analysis
Total protein carbonyls, protein-bound HNE, and protein-bound 3NT levels were analyzed by slot blot method. Briefly, for protein carbonyls, the samples (5 (g of protein) were derivatized with 10 mM 2,4-dinitrophenylhydrazine in the presence of 5 (l of 12% SDS for 20 min at room temperature. The samples were neutralized with 7.5 (l of the neutralization solution (2 M Tris in 30% glycerol). The resulting solution was loaded into each well on nitrocellulose membrane under vacuum using a slot-blot apparatus (250 ng/lane). The membrane then was washed with wash buffer [10 mM Tris–HCl (pH 7.5), 150 mM NaCl, 0.05% Tween 20], blocked by incubation in the presence of 3% BSA, followed by incubation with rabbit polyclonal anti-DNPH antibody as primary antibody for 1 h. The membrane was washed with wash buffer and further incubated with alkaline phosphatase (ALP)-conjugated goat anti-rabbit antibody as secondary antibody for 1 h. The blot was developed using 5-bromo-4-chloro-3-indolyl-phosphate/nitroblue tetrazolium (BCIP/NBT) color developing reagent, scanned in TIF format using Adobe Photoshop on a Canoscan 8800F (Canon, USA) and quantified with ImageQuant TL 1D version 7.0 software (GE Healthcare, USA).
2D electrophoresis and blotting
Samples (200 μl, 200 μg) were loaded on 110-mm pH 3–10 immobilized pH gradients (IPG) strips in a Bio-Rad IEF Cell system (Bio-Rad). Following 18 h of active rehydration (50 V) isoelectric focusing was performed. For the second dimension, thawed strips were sequentially equilibrated for 15 min in the dark in 375 mM Tris pH 8.8, 6 M urea, 2% sodium dodecyl sulfate (SDS), 20% glycerol containing first 2% dithiothreitol and then 2.5% of iodoacetamide. SDS PAGE was performed in Criterion Tris-HCl Gels 8-16% (Bio-Rad) at 200 V for 1 h. Gels were fixed for 45 min in 10% methanol, 7% acetic acid, and stained overnight on the rocker with SYPRO Ruby gel stain (Bio-Rad). After destaining in deionized water, gels were scanned with a STORM UV transilluminator (λex=470 nm, λem=618 nm, Molecular Dynamics, Sunnyvale, CA).
The same amount of protein sample (200 μg) was used for 2D- immunoblotting analysis, and the electrophoresis was carried out as described above. The proteins from the 2D-electrophoresis gels were transferred to nitrocellulose membranes (Bio-Rad) using a Transblot-Blot SD Semi-Dry Transfer Cell (Bio-Rad) at 15 V for 2 h. HNE-protein adducts were detected on the nitrocellulose paper using a primary anti-HNE rabbit antibody (Chemicon, Temecula, CA) specific for HNE-bound protein (1:100) for 2 h at room temperature while rocking, followed by a secondary goat anti-rabbit IgG (Sigma) antibody (1:1300) diluted in wash blot buffer for 2 h RT. The resultant membrane was developed as previously described.
PD-Quest analysis
Gel imaging was software-aided using PD-Quest (Bio-Rad) imaging software. Briefly, a master gel was selected followed by normalization of all gels (nontransgenic and transgenic strain samples), according to the total spot density. Gel-to-gel analysis was then initiated in two parts: manual matching and automated matching. This process generates a large pool of data, approximately 350 spots. Only proteins showing computer-determined significant differential levels between the two groups being analyzed were considered for identification. A quantitative analysis set was created that recognized matched spots with differences in the number of pixels that occur in each spot and a statistical analysis set was created that used a Student's t-test at 95% confidence to identify spots with p values of <0.05. Spots with p<0.05 were considered significant. A Boolean analysis set was created that identified overlapping spots from the aforementioned quantitative and statistical sets. These spots were selected for subsequent mass spectrometric analysis.
Gel cutting and in-gel trypsin digestion
Protein spots statistically different than controls were digested in-gel by trypsin. Briefly, spots of interest were excised and then washed with 0.1 M ammonium bicarbonate (NH4HCO3) at room temperature for 15 min. Acetonitrile was added and incubated at room temperature for 15 min. This solvent mixture was then removed and gel pieces dried. The protein spots were then incubated with 20 μl of 20 mM DTT in 0.1 M NH4HCO3 at 56°C for 45 min. The DTT solution was removed and replaced with 20 μl of 55 mM iodoacetamide in 0.1 M NH4HCO3. The solution was then incubated at room temperature for 30 min. The iodoacetamide was removed and replaced with 0.2 ml of 50 mM NH4HCO3 at room temperature for 15 min. Acetonitrile (200 μl) was added. After 15 min incubation, the solvent was removed, and the gel spots were dried for 30 min. The gel pieces were rehydrated with 20 ng/μl-modified trypsin (Promega, Madison, WI) in 50 mM NH4HCO3 with the minimal volume enough to cover the gel pieces. The gel pieces were incubated overnight at 37°C in a shaking incubator.
Mass spectrometry
Protein spots of interest were excised, subjected to in-gel trypsin digestion, and resulting tryptic peptides were analyzed with an automated nanospray Nanomate Orbitrap XL MS/MS platform (6). The Orbitrap MS was operated in a data-dependent mode whereby the 8 most intense parent ions measured in the FT at 60,000 resolution were selected for ion trap fragmentation with the following conditions: injection time 50 ms, 35% collision energy. MS/MS spectra were measured in the FT at 7500 resolutions, and dynamic exclusion was set for 120 seconds. Each sample was acquired for a total of ∼2.5 min. MS/MS spectra were searched against the ipi Worms Database using SEQUEST with the following criteria: Xcorr>1.5, 2.0, 2.5, 3.0 for +1, +2, +3, and +4 charge states, respectively, and P-value (protein and peptide)<0.01. IPI accession numbers were cross-correlated with SwissProt accession numbers for final protein identification.
Western blot analysis
Proteins (50 μg) were added to sample buffer, denaturated for 5 min at 100°C, loaded on 8%–16 % precast Criterion gels (Bio-Rad) and separated by electrophoresis at 100 mA for 2 h. The gels were then transferred to nitrocellulose paper using the Transblot-BlotSD Semi-DryTransfer Cell at 20 mA for 2 h. The membranes were incubated with V-type proton ATPase mouse monoclonal primary antibody (AbCam) for expression analysis in PBST for 2 h at room temperature. The membranes were then washed three times for 5 min with PBST, followed by incubation with anti-mouse alkaline phosphatase secondary antibody (Sigma) in PBST for 2 h at room temperature. Membranes were then washed, developed, and scanned as previously described.
Immunoprecipitation
For the immunoprecipitation procedure, 150 μg of protein extracts were dissolved in 500 μl of RIPA buffer (10 mM Tris, pH 7.6; 140 mM NaCl; 0.5% NP40 including protease inhibitors) and then incubated with 1 μg of V-type proton ATPase antibody at 4°C overnight. Immunocomplexes were collected by using protein A/G suspension for 2 h at 4°C and washed five times with immunoprecipitation buffer. Immunoprecipitated V-type proton ATPase was recovered by resuspending the pellets in reducing SDS buffers and subjected to electrophoresis on 12% gels, followed by Western blot analysis.
Statistical analysis
Statistical analysis of protein levels matched with spots on 2D-gels from mutated LRRK2::Tau transgenic strains compared Non-T and Tau transgenic strains were carried out using Student's t-tests. A value of p<0.05 was considered statistically significant. Only proteins that were considered significantly different by Student's t-test were subjected to in-gel trypsin digestion and subsequent proteomic analysis.
Abbreviations Used
- HNE
4-hydroxy-2-nonenal
- LB/LN
Lewy bodies/Lewy neuritis
- LRRK2
leucine-rich repeat kinase 2
- Non-T
nontransgenic
- 3NT
3-nitrotyrosine
- PD
Parkinson disease
- PTM
post-translational modification
- ROS/RNS
reactive oxygen species/reactive nitrogen species
- Tau
tau V337M transgenic
- WT
wild type
Acknowledgments
This work was supported in part by grants to DAB (NIH AG-05119) and to BW (Alzheimer Association, NIEHS ES15567, NINDS NS060872). FDD was supported by a Fellowship from Istituto Pasteur–Fondazione Cenci Bolognetti.
Author Disclosure Statement
The authors state no financial conflict of interests in this study.
References
- 1.Abou-Sleiman PM. Muqit MM. Wood NW. Expanding insights of mitochondrial dysfunction in Parkinson's disease. Nat Rev Neurosci. 2006;7:207–219. doi: 10.1038/nrn1868. [DOI] [PubMed] [Google Scholar]
- 2.Alegre-Abarrategui J. Christian H. Lufino MM. Mutihac R. Venda LL. Ansorge O. Wade-Martins R. LRRK2 regulates autophagic activity and localizes to specific membrane microdomains in a novel human genomic reporter cellular model. Hum Mol Genet. 2009;18:4022–4034. doi: 10.1093/hmg/ddp346. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Alegre-Abarrategui J. Wade-Martins R. Parkinson disease, LRRK2 and the endocytic-autophagic pathway. Autophagy. 2009;5:1208–1210. doi: 10.4161/auto.5.8.9894. [DOI] [PubMed] [Google Scholar]
- 4.Aluise CD. Robinson RA. Cai J. Pierce WM. Markesbery WR. Butterfield DA. Redox proteomics analysis of brains from subjects with amnestic mild cognitive impairment compared to brains from subjects with preclinical Alzheimer's disease: Insights into memory loss in MCI. J Alzheimers Dis. 2011;23:257–269. doi: 10.3233/JAD-2010-101083. [DOI] [PubMed] [Google Scholar]
- 5.Anglade P. Vyas S. Javoy-Agid F. Herrero MT. Michel PP. Marquez J. Mouatt-Prigent A. Ruberg M. Hirsch EC. Agid Y. Apoptosis and autophagy in nigral neurons of patients with Parkinson's disease. Histol Histopathol. 1997;12:25–31. [PubMed] [Google Scholar]
- 6.Boyd-Kimball D. Poon HF. Lynn BC. Cai J. Pierce WM., Jr. Klein JB. Ferguson J. Link CD. Butterfield DA. Proteomic identification of proteins specifically oxidized in Caenorhabditis elegans expressing human Abeta(1-42): Implications for Alzheimer's disease. Neurobiol Aging. 2006;27:1239–1249. doi: 10.1016/j.neurobiolaging.2005.07.001. [DOI] [PubMed] [Google Scholar]
- 7.Butterfield DA. Bader Lange ML. Sultana R. Involvements of the lipid peroxidation product, HNE, in the pathogenesis and progression of Alzheimer's disease. Biochim Biophys Acta. 2010;1801:924–929. doi: 10.1016/j.bbalip.2010.02.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Chan D. Citro A. Cordy JM. Shen GC. Wolozin B. Rac1 Protein rescues neurite retraction caused by G2019S leucine-rich repeat kinase 2 (LRRK2) J Biol Chem. 2011;286:16140–16149. doi: 10.1074/jbc.M111.234005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Choi J. Levey AI. Weintraub ST. Rees HD. Gearing M. Chin LS. Li L. Oxidative modifications and down-regulation of ubiquitin carboxyl-terminal hydrolase L1 associated with idiopathic Parkinson's and Alzheimer's diseases. J Biol Chem. 2004;279:13256–13264. doi: 10.1074/jbc.M314124200. [DOI] [PubMed] [Google Scholar]
- 10.Choi J. Rees HD. Weintraub ST. Levey AI. Chin LS. Li L. Oxidative modifications and aggregation of Cu,Zn-superoxide dismutase associated with Alzheimer and Parkinson diseases. J Biol Chem. 2005;280:11648–11655. doi: 10.1074/jbc.M414327200. [DOI] [PubMed] [Google Scholar]
- 11.Dachsel JC. Farrer MJ. LRRK2 and Parkinson disease. Arch Neurol. 2010;67:542–547. doi: 10.1001/archneurol.2010.79. [DOI] [PubMed] [Google Scholar]
- 12.Dalfo E. Portero-Otin M. Ayala V. Martinez A. Pamplona R. Ferrer I. Evidence of oxidative stress in the neocortex in incidental Lewy body disease. J Neuropathol Exp Neurol. 2005;64:816–830. doi: 10.1097/01.jnen.0000179050.54522.5a. [DOI] [PubMed] [Google Scholar]
- 13.Dalle-Donne I. Scaloni A. Butterfield DA. Redox Proteomics: From Protein Modifications to Cellular Dysfunction and Diseases. New York: John Wiley and Sons; 2006. [DOI] [PubMed] [Google Scholar]
- 14.Devine MJ. Lewis PA. Emerging pathways in genetic Parkinson's disease: Tangles, Lewy bodies and LRRK2. FEBS J. 2008;275:5748–5757. doi: 10.1111/j.1742-4658.2008.06707.x. [DOI] [PubMed] [Google Scholar]
- 15.Di Fonzo A. Chien HF. Socal M. Giraudo S. Tassorelli C. Iliceto G. Fabbrini G. Marconi R. Fincati E. Abbruzzese G. Marini P. Squitieri F. Horstink MW. Montagna P. Libera AD. Stocchi F. Goldwurm S. Ferreira JJ. Meco G. Martignoni E. Lopiano L. Jardim LB. Oostra BA. Barbosa ER. Italian Parkinson Genetics N. Bonifati V. ATP13A2 missense mutations in juvenile parkinsonism and young onset Parkinson disease. Neurology. 2007;68:1557–1562. doi: 10.1212/01.wnl.0000260963.08711.08. [DOI] [PubMed] [Google Scholar]
- 16.Farrer MJ. Genetics of Parkinson disease: Paradigm shifts and future prospects. Nat Rev Genet. 2006;7:306–318. doi: 10.1038/nrg1831. [DOI] [PubMed] [Google Scholar]
- 17.Forno LS. Neuropathology of Parkinson's disease. J Neuropathol Exp Neurol. 1996;55:259–272. doi: 10.1097/00005072-199603000-00001. [DOI] [PubMed] [Google Scholar]
- 18.Foroud T. Uniacke SK. Liu L. Pankratz N. Rudolph A. Halter C. Shults C. Marder K. Conneally PM. Nichols WC. Parkinson Study G. Heterozygosity for a mutation in the parkin gene leads to later onset Parkinson disease. Neurology. 2003;60:796–801. doi: 10.1212/01.wnl.0000049470.00180.07. [DOI] [PubMed] [Google Scholar]
- 19.Galpern WR. Lang AE. Interface between tauopathies and synucleinopathies: A tale of two proteins. Ann Neurol. 2006;59:449–458. doi: 10.1002/ana.20819. [DOI] [PubMed] [Google Scholar]
- 20.Gloeckner CJ. Schumacher A. Boldt K. Ueffing M. The Parkinson disease-associated protein kinase LRRK2 exhibits MAPKKK activity and phosphorylates MKK3/6 and MKK4/7, in vitro. J Neurochem. 2009;109:959–968. doi: 10.1111/j.1471-4159.2009.06024.x. [DOI] [PubMed] [Google Scholar]
- 21.Gomez A. Ferrer I. Increased oxidation of certain glycolysis and energy metabolism enzymes in the frontal cortex in Lewy body diseases. J Neurosci Res. 2009;87:1002–1013. doi: 10.1002/jnr.21904. [DOI] [PubMed] [Google Scholar]
- 22.Gomez A. Ferrer I. Involvement of the cerebral cortex in Parkinson disease linked with G2019S LRRK2 mutation without cognitive impairment. Acta Neuropathol. 2010;120:155–167. doi: 10.1007/s00401-010-0669-y. [DOI] [PubMed] [Google Scholar]
- 23.Guthrie CR. Schellenberg GD. Kraemer BC. SUT-2 potentiates tau-induced neurotoxicity in Caenorhabditis elegans. Hum Mol Genet. 2009;18:1825–1838. doi: 10.1093/hmg/ddp099. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Hatano T. Kubo S. Imai S. Maeda M. Ishikawa K. Mizuno Y. Hattori N. Leucine-rich repeat kinase 2 associates with lipid rafts. Hum Mol Genet. 2007;16:678–690. doi: 10.1093/hmg/ddm013. [DOI] [PubMed] [Google Scholar]
- 25.Imai Y. Gehrke S. Wang HQ. Takahashi R. Hasegawa K. Oota E. Lu B. Phosphorylation of 4E-BP by LRRK2 affects the maintenance of dopaminergic neurons in Drosophila. EMBO J. 2008;27:2432–2443. doi: 10.1038/emboj.2008.163. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Kraemer BC. Burgess JK. Chen JH. Thomas JH. Schellenberg GD. Molecular pathways that influence human tau-induced pathology in Caenorhabditis elegans. Hum Mol Genet. 2006;15:1483–1496. doi: 10.1093/hmg/ddl067. [DOI] [PubMed] [Google Scholar]
- 27.Kraemer BC. Zhang B. Leverenz JB. Thomas JH. Trojanowski JQ. Schellenberg GD. Neurodegeneration and defective neurotransmission in a Caenorhabditis elegans model of tauopathy. Proc Natl Acad Sci USA. 2003;100:9980–9985. doi: 10.1073/pnas.1533448100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Lewis PA. Greggio E. Beilina A. Jain S. Baker A. Cookson MR. The R1441C mutation of LRRK2 disrupts GTP hydrolysis. Biochem Biophys Res Commun. 2007;357:668–671. doi: 10.1016/j.bbrc.2007.04.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Li X. Tan YC. Poulose S. Olanow CW. Huang XY. Yue Z. Leucine-rich repeat kinase 2 (LRRK2)/PARK8 possesses GTPase activity that is altered in familial Parkinson's disease R1441C/G mutants. J Neurochem. 2007;103:238–247. doi: 10.1111/j.1471-4159.2007.04743.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Li Y. Dunn L. Greggio E. Krumm B. Jackson GS. Cookson MR. Lewis PA. Deng J. The R1441C mutation alters the folding properties of the ROC domain of LRRK2. Biochim Biophys Acta. 2009;1792:1194–1197. doi: 10.1016/j.bbadis.2009.09.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Lin CH. Tsai PI. Wu RM. Chien CT. LRRK2 G2019S mutation induces dendrite degeneration through mislocalization and phosphorylation of tau by recruiting autoactivated GSK3ss. J Neurosci. 2010;30:13138–13149. doi: 10.1523/JNEUROSCI.1737-10.2010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Lin MT. Beal MF. Mitochondrial dysfunction and oxidative stress in neurodegenerative diseases. Nature. 2006;443:787–795. doi: 10.1038/nature05292. [DOI] [PubMed] [Google Scholar]
- 33.Ludolph AC. Kassubek J. Landwehrmeyer BG. Mandelkow E. Mandelkow EM. Burn DJ. Caparros-Lefebvre D. Frey KA. de Yebenes JG. Gasser T. Heutink P. Hoglinger G. Jamrozik Z. Jellinger KA. Kazantsev A. Kretzschmar H. Lang AE. Litvan I. Lucas JJ. McGeer PL. Melquist S. Oertel W. Otto M. Paviour D. Reum T. Saint-Raymond A. Steele JC. Tolnay M. Tumani H. van Swieten JC. Vanier MT. Vonsattel JP. Wagner S. Wszolek ZK Reisensburg Working Group for Tauopathies With P. Tauopathies with parkinsonism: Clinical spectrum, neuropathologic basis, biological markers, and treatment options. Eur J Neurol. 2009;16:297–309. doi: 10.1111/j.1468-1331.2008.02513.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.MacLeod D. Dowman J. Hammond R. Leete T. Inoue K. Abeliovich A. The familial Parkinsonism gene LRRK2 regulates neurite process morphology. Neuron. 2006;52:587–593. doi: 10.1016/j.neuron.2006.10.008. [DOI] [PubMed] [Google Scholar]
- 35.Maraganore DM. Lesnick TG. Elbaz A. Chartier-Harlin MC. Gasser T. Kruger R. Hattori N. Mellick GD. Quattrone A. Satoh J. Toda T. Wang J. Ioannidis JP. de Andrade M. Rocca WA. Consortium UGG. UCHL1 is a Parkinson's disease susceptibility gene. Ann Neurol. 2004;55:512–521. doi: 10.1002/ana.20017. [DOI] [PubMed] [Google Scholar]
- 36.Martinez-Vicente M. Cuervo AM. Autophagy and neurodegeneration: When the cleaning crew goes on strike. Lancet Neurol. 2007;6:352–361. doi: 10.1016/S1474-4422(07)70076-5. [DOI] [PubMed] [Google Scholar]
- 37.Mata IF. Wedemeyer WJ. Farrer MJ. Taylor JP. Gallo KA. LRRK2 in Parkinson's disease: Protein domains and functional insights. Trends Neurosci. 2006;29:286–293. doi: 10.1016/j.tins.2006.03.006. [DOI] [PubMed] [Google Scholar]
- 38.Melrose HL. Dachsel JC. Behrouz B. Lincoln SJ. Yue M. Hinkle KM. Kent CB. Korvatska E. Taylor JP. Witten L. Liang YQ. Beevers JE. Boules M. Dugger BN. Serna VA. Gaukhman A. Yu X. Castanedes-Casey M. Braithwaite AT. Ogholikhan S. Yu N. Bass D. Tyndall G. Schellenberg GD. Dickson DW. Janus C. Farrer MJ. Impaired dopaminergic neurotransmission and microtubule-associated protein tau alterations in human LRRK2 transgenic mice. Neurobiol Dis. 2010;40:503–517. doi: 10.1016/j.nbd.2010.07.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Meredith GE. Totterdell S. Petroske E. Santa Cruz K. Callison RC., Jr. Lau YS. Lysosomal malfunction accompanies alpha-synuclein aggregation in a progressive mouse model of Parkinson's disease. Brain Res. 2002;956:156–165. doi: 10.1016/s0006-8993(02)03514-x. [DOI] [PubMed] [Google Scholar]
- 40.Nuytemans K. Rademakers R. Theuns J. Pals P. Engelborghs S. Pickut B. de Pooter T. Peeters K. Mattheijssens M. Van den Broeck M. Cras P. De Deyn PP. van Broeckhoven C. Founder mutation p.R1441C in the leucine-rich repeat kinase 2 gene in Belgian Parkinson's disease patients. Eur J Hum Genet. 2008;16:471–479. doi: 10.1038/sj.ejhg.5201986. [DOI] [PubMed] [Google Scholar]
- 41.Pan T. Kondo S. Le W. Jankovic J. The role of autophagy-lysosome pathway in neurodegeneration associated with Parkinson's disease. Brain. 2008;131:1969–1978. doi: 10.1093/brain/awm318. [DOI] [PubMed] [Google Scholar]
- 42.Plowey ED. Cherra SJ., 3rd Liu YJ. Chu CT. Role of autophagy in G2019S-LRRK2-associated neurite shortening in differentiated SH-SY5Y cells. J Neurochem. 2008;105:1048–1056. doi: 10.1111/j.1471-4159.2008.05217.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Rajput A. Dickson DW. Robinson CA. Ross OA. Dachsel JC. Lincoln SJ. Cobb SA. Rajput ML. Farrer MJ. Parkinsonism, Lrrk2 G2019S, and tau neuropathology. Neurology. 2006;67:1506–1508. doi: 10.1212/01.wnl.0000240220.33950.0c. [DOI] [PubMed] [Google Scholar]
- 44.Ramirez A. Heimbach A. Grundemann J. Stiller B. Hampshire D. Cid LP. Goebel I. Mubaidin AF. Wriekat AL. Roeper J. Al-Din A. Hillmer AM. Karsak M. Liss B. Woods CG. Behrens MI. Kubisch C. Hereditary parkinsonism with dementia is caused by mutations in ATP13A2, encoding a lysosomal type 5 P-type ATPase. Nat Genet. 2006;38:1184–1191. doi: 10.1038/ng1884. [DOI] [PubMed] [Google Scholar]
- 45.Rivera VM. Squillace RM. Miller D. Berk L. Wardwell SD. Ning Y. Pollock R. Narasimhan NI. Iuliucci JD. Wang F. Clackson T. Ridaforolimus (AP23573; MK-8669), a potent mTOR inhibitor, has broad antitumor activity and can be optimally administered using intermittent dosing regimens. Mol Cancer Ther. 2011;10:1059–1071. doi: 10.1158/1535-7163.MCT-10-0792. [DOI] [PubMed] [Google Scholar]
- 46.Rubinsztein DC. The roles of intracellular protein-degradation pathways in neurodegeneration. Nature. 2006;443:780–786. doi: 10.1038/nature05291. [DOI] [PubMed] [Google Scholar]
- 47.Saha S. Guillily MD. Ferree A. Lanceta J. Chan D. Ghosh J. Hsu CH. Segal L. Raghavan K. Matsumoto K. Hisamoto N. Kuwahara T. Iwatsubo T. Moore L. Goldstein L. Cookson M. Wolozin B. LRRK2 modulates vulnerability to mitochondrial dysfunction in Caenorhabditis elegans. J Neurosci. 2009;29:9210–9218. doi: 10.1523/JNEUROSCI.2281-09.2009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Santpere G. Ferrer I. LRRK2 and neurodegeneration. Acta Neuropathol. 2009;117:227–46. doi: 10.1007/s00401-008-0478-8. [DOI] [PubMed] [Google Scholar]
- 49.Schapira AH. Mitochondrial dysfunction in neurodegenerative diseases. Neurochem Res. 2008;33:2502–2509. doi: 10.1007/s11064-008-9855-x. [DOI] [PubMed] [Google Scholar]
- 50.Shin N. Jeong H. Kwon J. Heo HY. Kwon JJ. Yun HJ. Kim CH. Han BS. Tong Y. Shen J. Hatano T. Hattori N. Kim KS. Chang S. Seol W. LRRK2 regulates synaptic vesicle endocytosis. Exp Cell Res. 2008;314:2055–2065. doi: 10.1016/j.yexcr.2008.02.015. [DOI] [PubMed] [Google Scholar]
- 51.Smith WW. Pei Z. Jiang H. Dawson VL. Dawson TM. Ross CA. Kinase activity of mutant LRRK2 mediates neuronal toxicity. Nat Neurosci. 2006;9:1231–1233. doi: 10.1038/nn1776. [DOI] [PubMed] [Google Scholar]
- 52.Squillace RM. Miller D. Cookson M. Wardwell SD. Moran L. Clapham D. Wang F. Clackson T. Rivera VM. Antitumor activity of ridaforolimus and potential cell-cycle determinants of sensitivity in sarcoma and endometrial cancer models. Mol Cancer Ther. 2011;10:1959–1968. doi: 10.1158/1535-7163.MCT-11-0273. [DOI] [PubMed] [Google Scholar]
- 53.Sultana R. Butterfield DA. Identification of the oxidative stress proteome in the brain. Free Radic Biol Med. 2011;50:487–494. doi: 10.1016/j.freeradbiomed.2010.11.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Taylor JP. Mata IF. Farrer MJ. LRRK2: A common pathway for parkinsonism, pathogenesis and prevention? Trends Mol Med. 2006;12:76–82. doi: 10.1016/j.molmed.2005.12.004. [DOI] [PubMed] [Google Scholar]
- 55.Taymans JM. Cookson MR. Mechanisms in dominant parkinsonism: The toxic triangle of LRRK2, alpha-synuclein, and tau. Bioessays. 2010;32:227–235. doi: 10.1002/bies.200900163. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Ved R. Saha S. Westlund B. Perier C. Burnam L. Sluder A. Hoener M. Rodrigues CM. Alfonso A. Steer C. Liu L. Przedborski S. Wolozin B. Similar patterns of mitochondrial vulnerability and rescue induced by genetic modification of alpha-synuclein, parkin, and DJ-1 in Caenorhabditis elegans. J Biol Chem. 2005;280:42655–42668. doi: 10.1074/jbc.M505910200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.West AB. Moore DJ. Biskup S. Bugayenko A. Smith WW. Ross CA. Dawson VL. Dawson TM. Parkinson's disease-associated mutations in leucine-rich repeat kinase 2 augment kinase activity. Proc Natl Acad Sci USA. 2005;102:16842–16847. doi: 10.1073/pnas.0507360102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.West AB. Moore DJ. Choi C. Andrabi SA. Li X. Dikeman D. Biskup S. Zhang Z. Lim KL. Dawson VL. Dawson TM. Parkinson's disease-associated mutations in LRRK2 link enhanced GTP-binding and kinase activities to neuronal toxicity. Hum Mol Genet. 2007;16:223–232. doi: 10.1093/hmg/ddl471. [DOI] [PubMed] [Google Scholar]
- 59.Wszolek ZK. Tsuboi Y. Ghetti B. Pickering-Brown S. Baba Y. Cheshire WP. Frontotemporal dementia and parkinsonism linked to chromosome 17 (FTDP-17) Orphanet J Rare Dis. 2006;1:30. doi: 10.1186/1750-1172-1-30. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Yang Q. Mao Z. Parkinson disease: A role for autophagy? Neuroscientist. 2010;16:335–341. doi: 10.1177/1073858409357118. [DOI] [PubMed] [Google Scholar]
- 61.Zimprich A. Biskup S. Leitner P. Lichtner P. Farrer M. Lincoln S. Kachergus J. Hulihan M. Uitti RJ. Calne DB. Stoessl AJ. Pfeiffer RF. Patenge N. Carbajal IC. Vieregge P. Asmus F. Muller-Myhsok B. Dickson DW. Meitinger T. Strom TM. Wszolek ZK. Gasser T. Mutations in LRRK2 cause autosomal-dominant parkinsonism with pleomorphic pathology. Neuron. 2004;44:601–607. doi: 10.1016/j.neuron.2004.11.005. [DOI] [PubMed] [Google Scholar]