Abstract
Salivary gland cells in the larvae of the dipteran Chironomus tentans offer unique possibilities to visualize the assembly and nucleocytoplasmic transport of a specific transcription product. Each nucleus harbors four giant polytene chromosomes, whose transcription sites are expanded, or puffed. On chromosome IV, there are two puffs of exceptional size, Balbiani ring (BR) 1 and BR 2. A BR gene is 35–40 kb, contains four short introns, and encodes a 1-MDa salivary polypeptide. The BR transcript is packed with proteins into a ribonucleoprotein (RNP) fibril that is folded into a compact ring-like structure. The completed RNP particle is released into the nucleoplasm and transported to the nuclear pore, where the RNP fibril is gradually unfolded and passes through the pore. On the cytoplasmic side, the exiting extended RNP fibril becomes engaged in protein synthesis and the ensuing polysome is anchored to the endoplasmic reticulum. Several of the BR particle proteins have been characterized, and their fate during the assembly and transport of the BR particle has been elucidated. The proteins studied are all added cotranscriptionally to the pre-mRNA molecule. The various proteins behave differently during RNA transport, and the flow pattern of each protein is related to the particular function of the protein. Because the cotranscriptional assembly of the pre-mRNP particle involves proteins functioning in the nucleus as well as proteins functioning in the cytoplasm, it is concluded that the fate of the mRNA molecule is determined to a considerable extent already at the gene level.
The organization of chromatin in a diploid cell nucleus is complex and dynamic. The chromosomes form chromosomal territories, each consisting of several more-or-less condensed and variable domains (1, 2). The individual territories are separated by a delicate network of thin channels, the interchromosomal space (2–4). The active genes are usually situated in the periphery of the domains and deliver the transcription products into the channel system (5, 6). The products move toward the periphery of the nucleus and leave the nucleus through the nuclear pores in the nuclear envelope (7, 8).
In the ordinary diploid nucleus, it has proven difficult to follow the flow of specific transcription products from the gene to the nuclear pores. At the light microscopy level, specific genes and their growing transcripts can be located by in situ hybridization (e.g., ref. 9), but the completed and released transcripts are usually too scarce in the nucleoplasm to be detected and traced in the channel system. In the electron microscope, it is difficult to identify specific active genes as well as the corresponding transcription products in transit from the gene to the periphery of the nucleus. However, in the polytene nuclei of dipteran insects, it is feasible, in exceptional cases, to visualize both the transcription process and the transport of the transcription product from the gene to the nuclear pores. The most extensively studied system in this respect is the Balbiani rings (BRs) on the polytene chromosomes in the larval salivary glands of the midge Chironomus tentans (8).
Polytene chromosomes consist of thousands of identical chromatids perfectly arranged side by side into well-defined cable-like structures (for review, see ref. 10). The transversely banded chromosomes allow specific chromosomal regions to be identified and synthetic events along the chromosomes to be studied. The transcriptionally active regions are blown-up, or puffed. In the salivary glands of C. tentans, there are three exceptionally large puffs, designated BR1, BR2, and BR3, which are all located on the short chromosome IV (Fig. 1). In the two largest BRs, BR1 and BR2, the transcriptionally active genes are 35–40 kb in size and contain four introns, three close to the 5′ end of the gene and one close to the 3′ end (11, 12). The introns are very short, and the BR1 and BR2 transcripts are, therefore, only minimally reduced in size during processing. The transcripts encode giant salivary polypeptides (about 1 MDa) that are secreted and form a proteinaceous tube in which the larva lives (13). As the BR transcripts are made large, remain large, and are abundant both on the gene and in the nucleoplasm, the BR transcription products are optimal for visualization of the assembly and transport of these transcription products; in fact, it has been possible to follow the formation of the product during transcription as well as the transport to and through the nuclear pores and finally the exit of the transcript and the formation of polysomes on the cytoplasmic side of the pore (8).
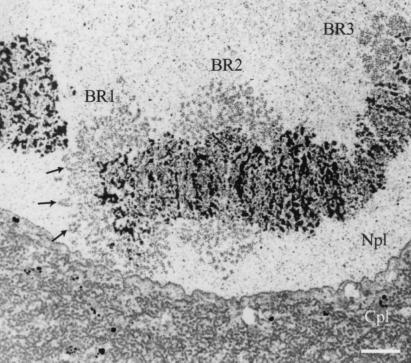
Figure 1.
Electron micrograph showing chromosome IV with its three giant puffs (BRs) in a salivary gland cell from C. tentans. The three BRs (BR1, BR2, and BR3) are indicated as well as the nucleoplasm (Npl) and cytoplasm (Cpl). The arrows mark a few prominent transcription loops (cf. Fig. 2D). (Bar equals 2 μm.)
Visualization of Assembly and Transport of BR Particles
The active BR genes have been studied both when spread on the surface of an electron microscopic grid (14) and when present within the cell (8, 15). The genes are heavily loaded with RNA polymerases and resemble in the electron microscope the well-known “Christmas-tree”-like ribosomal genes (16). The transcripts increase in size along the gene, and proteins associate with the growing RNAs to form thin ribonucleoprotein (RNP) fibrils. In spread preparations, the RNP fibrils are more or less extended because of the low salt conditions used. In situ, however, the packing of the RNP fibril into higher-order structure can be followed. At low resolution, an RNP fiber is first recognized, which is later on packed into a globular structure (Fig. 2D). At higher resolution, it can be seen how the thin RNP fibril is initially loosely coiled (forming the RNP fiber) and is subsequently tightly folded into a short ribbon, which is bent into a partial ring (the globule) (Fig. 3). When the particle is released from the gene, the RNP fiber is retracted into the globular portion, and the particle attains an almost ring-like conformation. The particle moves randomly in the interchromosomal space (17), although it can transiently bind to a fibrous network (18). When the particle gets to the nuclear pore complex and passes through the pore, the bent ribbon becomes straightened out, the RNP fibril unfolds and emerges extended on the cytoplasmic side, and protein synthesis is initiated (Fig. 3) (19). The translocation process has been studied in detail, and several discrete steps have been elucidated: binding of the BR RNP particle to the nucleoplasmic fibers of the nuclear pore complex, docking of the particle in front of the central channel of the pore complex, unrolling of the ribbon and translocation of the RNP complex with its 5′ end in the lead through the channel, exit of the unfolded RNP fibril into the cytoplasm, and formation of a polysome just outside the pore (8). Thus, the translocation of the BR RNP particle appears to be an ordered process with several well-defined stages. Furthermore, the spectacular conformational changes of the BR particle indicate that the process is quite dynamic, which is further supported by the observation that during translocation the BR particle loses proteins while others are presumably added (see below).
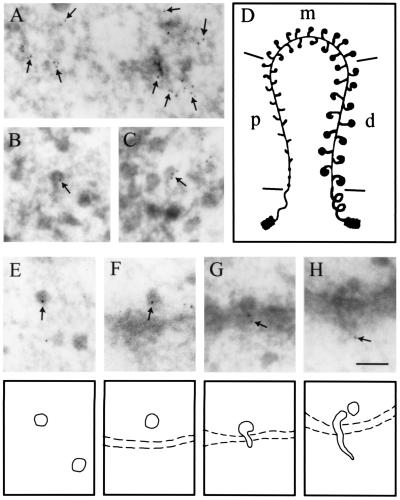
Figure 2.
Intracellular distribution of the cap-binding protein CBP20 in C. tentans salivary gland cells studied by immunoelectron microscopy. The assembly of the BR RNP particle is shown in A–D: proximal portions of the BR gene are displayed in A, distal portions in B and C, and a schematic drawing of the BR gene in D (p, proximal; m, middle; d, distal portions of the gene). The fate of the released BR particles is shown in E–H: BR particles are present in the nucleoplasm (E), at the pore (F), and in an unfolded conformation when passing through the pore (G and H). Gold particles are marked by arrows and indicate the position of CBP20. It should be noted that gold particles are at the leading 5′ end of the BR particle when it passes through the nuclear pore. (Bar equals 100 nm.) Modified from ref. 27; produced by permission of The Rockefeller University Press.
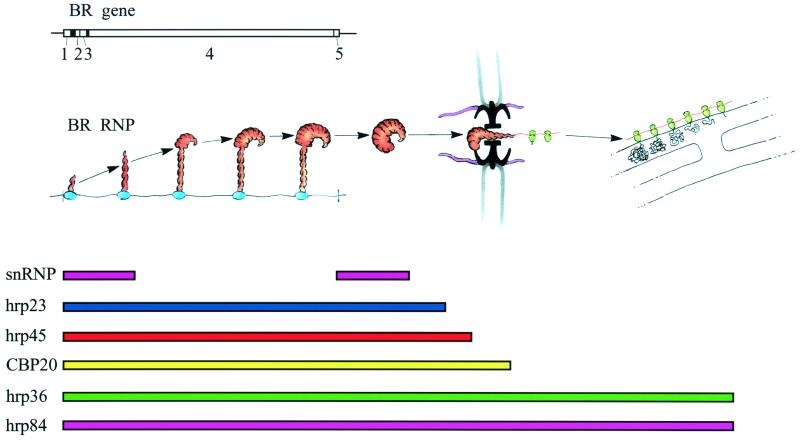
Figure 3.
Assembly and transport of the BR RNP particle and its relation to a number of BR RNA-associated proteins. The BR particle is assembled on the gene (left), passes through the nucleoplasm, unfolds, and translocates through the nuclear pore (middle). On the cytoplasmic side, the BR RNP fibril becomes engaged in protein synthesis and the polysomes anchor at the endoplasmic reticulum (right). The tripartite nuclear pore complex with its central channel is seen in black and its nuclear and cytoplasmic fibers are presented in pink. The BR gene with its five exons is displayed above the BR particle scheme, and the flow patterns of the BR RNA-associated proteins are outlined below. snRNP, small nuclear RNP. Modified from ref. 8; printed with permission from Elsevier Science.
Approach to Study BR RNA-Binding Proteins
Evidently proteins become associated with the RNA concomitant with transcription. In fact, the proteins seem to bind to the growing RNA molecule in the immediate vicinity of the RNA polymerase. Several questions are close at hand: What proteins are associated with the RNP particle? Are the proteins simply packaging proteins, or do they also play other functional roles? It has been estimated that there are 400–500 average-sized protein molecules in a BR particle (20).
It is well established that pre-mRNA is associated with many different proteins, usually designated hnRNP proteins (heterogeneous nuclear RNP proteins) (21). For example, in humans there are 30 major hnRNP proteins and a large number of minor ones (22). As a rule, the proteins can bind to a broad range of different sequences, some with higher affinity, others with lower affinity (21). Thus, as the hnRNP proteins show sequence preference in their interaction with RNA, they are likely to be nonrandomly bound to pre-mRNA. It has been directly shown in reconstitution experiments that each different RNA species is associated with a unique combination of hnRNP proteins (23). These studies were performed under conditions for binding sites and, therefore, resemble the in vivo situation in the cell nucleus. Furthermore, the hnRNP protein compositions at various puffs on polytene chromosomes in Drosophila (24) and Chironomus (25) differ quantitatively but also qualitatively, suggesting that each type of transcript binds a specific subset of hnRNP proteins. It is, therefore, an interesting possibility that the hnRNP proteins are not only unspecific RNA packaging proteins but also capable of exerting specific, transcript-related functions. To test such a hypothesis, it is attractive to study the protein set-up of individual specific transcripts and relate the individual proteins to the fate of the transcript.
It would have been most satisfactory if the proteins in the BR particles could have been studied by a direct approach. It is true that the BR particles can be isolated as a 300S fraction (20), but the quantities are not sufficient to allow a direct biochemical characterization. Instead, we adopted an indirect approach devised by Dreyfuss and coworkers (26). Nuclear RNA-binding proteins were isolated from C. tentans cultured cells by single-stranded DNA-Sepharose affinity chromatography and were used to raise monoclonal antibodies in mice. A collection of such antibodies was obtained (25). Antibodies that showed high specificity in Western blot experiments and bound to the BRs in immunocytochemical experiments were selected for further experiments. The antibodies were used to characterize the corresponding proteins by cDNA cloning and to study the fate of the proteins during the assembly and transport of the BR particle by using immunocytochemical and immunoelectron microscopy experiments.
As an example of a protein flow analysis, I have chosen the immunoelectron microscopic analysis of a cap-binding protein, CBP20 (27). CBP20 is known to bind to the 5′ end of the transcript in a cap-binding complex (CBC) together with another protein, CBP80 (28). An antibody raised against the human CBP20 was applied in the study of the BR particle. Cryosections through salivary gland cells were prepared and challenged with the anti-CBP20 antibody and subsequently with a secondary antibody coupled to gold. As shown in Fig. 2, the gold particles are present in the proximal portions of the active BR gene (Fig. 2A) as well as in the distal portions (Fig. 2 B and C). In the almost finished BR particles it can be seen that the gold is at the 5′ end of the particle (Fig. 2B; cf. schematic drawing in Fig. 3)—i.e., the position of the cap structure. Furthermore, it was noted that there is no increase in binding during the course of transcription, suggesting that the protein is added to the cap structure almost immediately upon initiation of transcription. BR particles released into the nucleoplasm are also labeled with gold (Fig. 2 E and F). Finally, during translocation through the nuclear pore, the leading 5′ end of the BR particle is labeled and the gold can also be seen on the cytoplasmic side of the nuclear pore complex (Fig. 2 G and H). Further out in the cytoplasm, there are no gold particles. We conclude that CBP20 is added cotranscriptionally and remains associated with the particle to and through the nuclear pore. On the cytoplasmic side, it is released from the particle and probably returns to the nucleus. These data are in good agreement with the observation that CBPs are shuttling proteins (28).
During the last couple of years a number of various RNA-binding proteins have been studied, and our results are summed up in Fig. 3. The flow patterns of the proteins are presented below the morphological description of the assembly and transport of the BR particle; the exon–intron organization of the BR gene is shown above. It is evident that the various proteins show quite different behavior during gene expression. Thus, not only the particle's morphology but also its protein composition during the transport from the gene to the cytoplasm is drastically changed.
Spliceosome Assembly and Disassembly
As a marker for spliceosome components we chose the snRNP proteins and used a monoclonal anti-snRNP antibody (Y12) to perform immunoelectron microscopy experiments (29). When the growing BR RNP products were studied in situ, it was noted that the snRNP proteins were present mainly in the proximal portion and only to a minor extent in the middle and distal portions of the active gene. Furthermore, nucleoplasmic BR particles, isolated, unfolded, and spread on a grid surface, showed labeling only at one end of the transcript, presumably the 3′ end. Thus, the snRNPs do not associate along the whole pre-mRNP fibril but rather bind to the 5′ and 3′ ends—i.e., the regions containing introns. These results nicely agree with an earlier analysis carried out at the RNA level, showing that the three 5′ end introns are spliced concomitantly with transcription in the promoter-proximal third of the gene, whereas the 3′ intron is spliced mainly posttranscriptionally (30). We conclude that the observed discontinuous distribution of snRNP proteins along the pre-mRNP fibril implies that spliceosomes both assemble and disassemble rapidly on the RNP fibril.
Proteins Confined to the Nucleus
Two of the studied proteins, hrp45 (31) and hrp23 (32), proved to be confined to the cell nucleus. The hrp45 protein contains two amino-terminal RNP-consensus RNA-binding domains (RBDs) and a carboxyl-terminal region rich in arginine-serine dipeptide repeats (RS domain), an organization characteristic of the SR family of RNA splicing factors (for review, see ref. 33). The hrp45 protein shows a high sequence homology to the human ASF-SF2 protein (34, 35) and the Drosophila SRp55 protein (36, 37), which are both known to be essential splicing factors (33). The hrp23 protein contains a single amino-terminal RBD and a carboxyl-terminal auxiliary region rich in glycine, arginine, and serine. It resembles the RBD-Gly type of hnRNP proteins (e.g., hnRNP A1), which contain one or two RBDs and a glycine-rich auxiliary domain. However, hrp23 share features with the SR proteins (e.g., several SR/RS dipeptides in the auxiliary domain), suggesting that hrp23 represents a group of proteins intermediate in structure between these two major groups of pre-mRNA-binding proteins. The hrp23 protein has a homologue in Drosophila, ROX21 (38), which has recently been shown to be a splicing repressor and, therefore, renamed RSF1 (repressor splicing factor 1) (39). Thus, the two BR particle proteins hrp45 and hrp23 are likely to be splicing factors.
Both hrp45 and hrp23 are added to the growing BR transcript along the large exon, and most likely along the entire transcript. Furthermore, they are both present in the nucleoplasmic BR particles, most of which contain fully spliced RNA (30). It should be stressed that neither of these putative splicing factors seems to behave as a genuine spliceosome component—i.e., a component that appears transiently on the pre-mRNP fibril and only at intron regions (compare the asymmetric distribution of snRNP proteins described above). Instead, they appear evenly along the transcript and remain with the fully spliced transcript in the nucleoplasm. It seems likely that the two proteins play important roles in the structural organization of the pre-mRNP particle, setting the stage for splicing rather than directly participating in the splicing process.
The two proteins are not released at the same time in conjunction with the translocation of the BR particle through the nuclear pore: whereas hrp23 is shed just before or at the binding of the particle to the pore, hrp45 is released when the particle enters the central channel. Thus, it seems likely that there is not a single protein-removal step at nucleocytoplasmic transport but rather a series of preparatory steps before the actual translocation of the RNP particle through the pore. It could be speculated that the shedding of hrp23 is required for binding of the particle to the nuclear pore complex, whereas the removal of hrp45 is closely connected to the translocation of the particle through the central channel. It is interesting to note that some mammalian hnRNP proteins—e.g., hnRNP C—contain a nuclear retention signal in the auxiliary domain (40). This signal can override nuclear export signals in the shuttling hnRNP proteins and, therefore, the nonshuttling proteins have to be actively displaced from the hnRNP complex before the nucleocytoplasmic translocation. We conclude that the hrp23 and hrp45 proteins are removed in a consecutive fashion beginning before or at the binding of the RNP particle to the nuclear pore complex. The fact that the proteins behave differently during nucleocytoplasmic translocation could imply that each of them plays a specific role during export of mRNA from the nucleus to the cytoplasm.
Proteins Accompanying the mRNA into the Cytoplasm
As discussed above, the CBP20 protein is bound to the cap structure early during transcription and accompanies the particle to and through the pore but is immediately dismissed just outside the pore. The rapid association of CBP20 with nascent RNA transcripts is consistent with the proposed role of the cap-binding complex (CBC) in splicing and 3′ end formation (28). Furthermore, the retention of the CBC on the RNP during translocation through the nuclear pore suggests that the CBC could also have a function at the recognition of the particle at the pore complex and/or in the translocation process itself when the 5′ end of the RNA is in the lead. Such a view is supported by the observation that the transport of snRNP particles is dependent on the cap structure and CBPs (41). However, although the cap structure facilitates transport of mRNA, it does not seem to be necessary (42, 43). Because the exiting 5′ end of the transcript is immediately engaged in protein synthesis, it is evident that the proteins bound to the cap structure are rapidly exchanged, CBPs being shed from the cap and translation initiation factors being recruited to the cap (28).
Three proteins, hrp36 (44), actin (45), and hrp84 (J. Zhao, D. Nashchekin, N. Visa, and B.D., unpublished data), have been found accompanying the BR RNA all the way from the gene via the nuclear pore into polysomes in the cytoplasm. The hrp36 protein is a 2xRBD-Gly protein and resembles the human hnRNP A1 protein and the Drosophila hrp40 protein (21).The hnRNP A1 protein is known to be a shuttling protein (46) and contain a nuclear export signal (NES) (47). It was early proposed that hnRNP A1 functions as a transport mediator for mRNA (46). The observation that hrp36 is associated with BR RNA during its translocation through the nuclear pore is in good agreement with such a concept (44). However, it is also remarkable that hrp36 stays with the mRNA also during protein synthesis and remains distributed along the messenger molecule. The role of hrp36 in polysomes is still only a matter of speculation, but the appearance of hrp36 along the entire message suggests a global role. One possibility could be that it keeps the RNA extended, thereby facilitating protein–RNA interactions and the translation process. Another possibility would be that it is available to package the RNA when not translated (compare DNA and nucleosomes). A third possibility would be that hrp36, like other hnRNP proteins, favors cap-dependent initiation of translation by preventing aberrant initiations along the message (48).
In our search for an export receptor binding to hrp36, we observed that actin forms a complex with hrp36 (45). It was shown first by immunoelectron microscopy that actin appears in the BR particle cotranscriptionally and remains attached to the particle in the nucleoplasm. Using DNase I affinity chromatography, we could demonstrate that actin is bound to hrp36 in nuclear as well as cytoplasmic extracts from C. tentans culture cells. The interaction is direct, as purified actin binds to recombinant hrp36 in an in vitro reconstitution experiment. Furthermore, the interaction between hrp36 and actin takes place in vivo as demonstrated by cross-linking. Thus, there is an hrp36–actin complex in the BR particle in the cell nucleus. This complex is also detected in the cytoplasm. As hrp36 enters polysomes, it seems likely that the complex is also present in the polysomes.
A central issue is whether the actin is monomeric or polymeric. In the fixed cells studied we found no evidence for actin filaments in the salivary gland cell nucleus. Most remarkably, many of the actin-containing BR particles do not seem to be associated with any fibers. Furthermore, no phalloidin staining was detected in the nucleus, although the brush border of the salivary gland cells, known to contain F-actin, was heavily stained. Finally, the anti-actin antibody used is known to have a strong preference for monomeric or short oligomeric actin. We conclude that in the fixed cells actin bound to hrp36 in the cell nucleus is likely to be in a monomeric or short oligomeric form. However, it has to be recalled that microfilaments can be extremely sensitive to fixation and could have disassembled during fixation. In fact, early microdissection experiments with C. tentans salivary gland cells showed that the polytene chromosomes are embedded in a labile gel (49), which has properties like the actin gel in amphibian oocyte nuclei (50). Thus, presently, the issue of the state of actin in the nuclear actin–hrp36 complex has to be left open. The state of actin in the cytoplasmic actin–hrp36 complex is also unclear, as it has not been possible to decide to what extent the immunolabeled actin in the cytoplasm reflects the distribution of the actin–hrp36 complex.
It can be speculated that the hrp36–actin complex is important for packing the RNA into a BR RNP fibril and further into well-defined higher-order structures (51). Other possibilities would be that actin promotes interaction of the BR particle with a fibrous network in the nucleoplasm, allows binding to export receptors (cf. ref. 52), or is involved in the dramatic conformational change of the particle upon translocation through the nuclear pore. Because actin remains bound to hrp36 in the cytoplasm, it is important to recall that hnRNP proteins have been found to affect translation efficiency, mRNA stability, and RNA location within cytoplasm (22). Evidently, a wide range of functional options have to be considered for the actin–hrp36 complex.
The third protein that is added cotranscriptionally to the BR transcript and accompanies the RNA through the nuclear pores and enters polysomes is hrp84, which is a putative RNA helicase. It belongs to the PL10 family of DEAD box proteins, which comprises, e.g., the human DBX (53), the mouse PL10 (54), the Xenopus An3 (55), and the yeast Ded1 proteins (56). The Ded1 protein is known to be important for initiation of translation (57). It is interesting to note that the mouse PL10 protein (57) and the human DBX (53) are probably also involved in the initiation of translation, as they can complement a deletion of the yeast gene DED1. Thus, it seems likely that hrp84 exerts its function in polysomes and presumably during the initiation of translation. We conclude that hrp84 represents a protein that functions in the polysomes in cytoplasm but is added to the transcript already in the nucleus.
The Cotranscriptional Loading Stage
The general picture that emerges from our studies of the proteins in the BR particle is that during the assembly of the particle the pre-mRNA molecule is loaded with proteins functioning early in the cell nucleus and with proteins functioning late in the cytoplasm. Thus, both the nuclear fate and the cytoplasmic fate of the mRNA are influenced by the proteins that are carried along with the RNA. This conclusion is supported by studies of gene expression in other species. The human hnRNP proteins are located predominantly in the cell nucleus, but many of them, including hnRNP A1, A2, D, E, and K, are shuttling between nucleus and cytoplasm (21, 46). In addition, more and more information accumulates showing that hnRNP proteins affect the fate of the mRNA in the cytoplasm—e.g., the transport of mRNA within the cytoplasm, the translational efficiency, and the mRNA turnover (for review, see ref. 22). It seems reasonable to assume that also these proteins travel with the mRNA from the nucleus to the cytoplasm, like hrp36 and hrp84 with BR RNA in C. tentans. Recently, it was shown in Drosophila by microinjection experiments that the proper cytoplasmic localization of fushi tarazu transcripts requires that the transcript enters the cytoplasm associated with the hnRNP protein hrp40 (58). We conclude that proteins loaded cotranscriptionally on pre-mRNA determine to a large extent the fate of the mRNA in both the nucleus and the cytoplasm.
The assembly of proteins along the pre-mRNA molecule is likely to be a complex process (Fig. 4). Some of the proteins, such as the cap-binding protein CBP20, bind to a specific sequence with high affinity, whereas most of the major RNA-binding proteins, such as the 2xRBD-Gly protein hrp36, bind with lower affinity at many sites along the pre-mRNA molecule. The presence of many RNA-binding proteins with limited sequence specificity will result in competition for available binding sites. The outcome of the assembly will, therefore, depend on the particular proteins present for binding and their relative abundance in the vicinity of the gene. It should be emphasized that the composition of nuclear hnRNP proteins is known to vary considerably between tissues and developmental stages (59). Thus, the set of proteins associated with a given transcript is not likely to be fixed but rather dependent on the cell type studied, physiological conditions, etc.
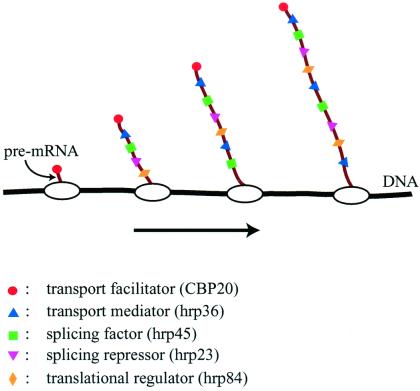
Figure 4.
Cotranscriptional loading of proteins onto growing BR pre-mRNA molecules. Some proteins bind to the pre-mRNA with high sequence specificity (e.g., CBP20), whereas others bind with lower specificity along the entire RNA molecule (e.g., the SR protein hrp45 and the 2xRBD-Gly protein hrp36). The RNP fibril formed serves as the substrate for trans-acting factors, and its structure affects a number of mRNA-related processes, including splicing, transport, and translation.
As most proteins bound to the pre-mRNA not only are packaging proteins but also exert more specific functions during the gene expression process, a modulated loading of the transcript with proteins will have functional implications. For example, it has been shown that the relative amounts of the two antagonistically acting RNA-binding proteins hnRNP A1 and ASF/SF2 decide the outcome of alternative splicing (60). The primary transcription product, the pre-mRNP fibril, should therefore be looked upon as a variable substrate for trans-acting factors, and the molecular organization of the fibril will influence not only splicing but also processes such as transport, translation, and mRNA degradation. Unfortunately, today we have only limited information on how the RNP fibril is organized at the molecular level, and we know even less about the rules that govern the assembly of the RNP fibril.
Conclusions
A specific transcription product, the BR RNP particle, has been studied during assembly on the gene and transport through the nucleoplasm to and through the nuclear pores. On the cytoplasmic side, the BR RNP particle appears as an extended RNP fibril that immediately engages in protein synthesis. A number of BR RNA-associated proteins have been identified, and their flow patterns have been studied in relation to the assembly and transport of the BR particle. The following major conclusions have been drawn:
(i) The BR RNP particle carries a specific subset of hnRNP proteins.
(ii) The proteins are added to the pre-mRNA cotranscriptionally.
(iii) The various proteins behave differently during RNA transport: some leave the transcript in the nucleoplasm or at the nuclear pore, others are shed subsequent to the translocation of the particle through the nuclear pore, whereas still others accompany the mRNA into polysomes.
(iv) The flow pattern of a protein seems related to the function of the protein.
(v) The particle proteins exert specific mRNA-related functions rather than merely serving as RNA-packaging devices.
(vi) The cotranscriptional assembly process sets the stage for both the nuclear and the cytoplasmic fate of the mRNA sequence.
Acknowledgments
I thank Sergej Masich, Birgitta Björkroth, and Birgitta Ivarsson for preparing the figures. The research was supported by the Swedish Natural Science Research Council, the Human Frontier Science Program Organization, the Knut and Alice Wallenberg Foundation, the Marianne and Marcus Wallenberg Foundation, and the Gunvor and Josef Anér Foundation.
Abbreviations
- BR
Balbiani ring
- RNP
ribonucleoprotein
- snRNP
small nuclear RNP
- hnRNP
heterogeneous nuclear RNP
- CBP
cap-binding protein
- RBD
RNA-binding domain
Footnotes
This paper was presented at the National Academy of Sciences colloquium, “Molecular Kinesis in Cellular Function and Plasticity,” held December 7–9, 2000, at the Arnold and Mabel Beckman Center in Irvine, CA.
References
- 1.Cremer T, Kurz A, Zirbel R, Dietzel S, Rinke B, Schröck E, Speicher M R, Mathieu U, Jauch A, Emmerich P, et al. Cold Spring Harbor Symp Quant Biol. 1973;58:777–792. doi: 10.1101/sqb.1993.058.01.085. [DOI] [PubMed] [Google Scholar]
- 2.van Driel R, Wansink D G, van Steensel B, Grande M A, Schul W, de Jong L. Int Rev Cytol. 1995;162A:151–188. doi: 10.1016/s0074-7696(08)61231-0. [DOI] [PubMed] [Google Scholar]
- 3.Spector D L. Proc Natl Acad Sci USA. 1990;87:147–151. doi: 10.1073/pnas.87.1.147. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Zachar Z, Kramer J, Mims I P, Bingham P M. J Cell Biol. 1993;121:729–742. doi: 10.1083/jcb.121.4.729. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Fakan S, Puvion E, Spohr G. Exp Cell Res. 1976;99:155–164. doi: 10.1016/0014-4827(76)90690-x. [DOI] [PubMed] [Google Scholar]
- 6.Kurz A, Lampel S, Nickolenko J E, Bradl J, Benner A, Zirbel R M, Cremer T, Lichter P. J Cell Biol. 1996;135:1195–1205. doi: 10.1083/jcb.135.5.1195. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Dworetzky S I, Feldherr C M. J Cell Biol. 1988;106:575–584. doi: 10.1083/jcb.106.3.575. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Daneholt B. Cell. 1997;88:585–588. doi: 10.1016/s0092-8674(00)81900-5. [DOI] [PubMed] [Google Scholar]
- 9.Xing Y, Johnson C V, Dobner P R, Lawrence J B. Science. 1993;259:1326–1330. doi: 10.1126/science.8446901. [DOI] [PubMed] [Google Scholar]
- 10.Case S T, Daneholt B. In: Biochemistry of Cell Differentiation II. Paul J, editor. Baltimore: University Park Press; 1977. pp. 45–77. [Google Scholar]
- 11.Wieslander L, Paulsson G. Proc Natl Acad Sci USA. 1992;89:4578–4582. doi: 10.1073/pnas.89.10.4578. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Wieslander L. Prog Nucleic Acids Res Mol Biol. 1994;48:275–313. doi: 10.1016/s0079-6603(08)60858-2. [DOI] [PubMed] [Google Scholar]
- 13.Case S T, Wieslander L. In: Biopolymers, Results and Problems in Cell Differentiation. Case S T, editor. Vol. 19. Berlin: Springer; 1992. pp. 187–226. [DOI] [PubMed] [Google Scholar]
- 14.Lamb M M, Daneholt B. Cell. 1979;17:835–848. doi: 10.1016/0092-8674(79)90324-6. [DOI] [PubMed] [Google Scholar]
- 15.Skoglund U, Andersson K, Björkroth B, Lamb M M, Daneholt B. Cell. 1983;34:847–855. doi: 10.1016/0092-8674(83)90542-1. [DOI] [PubMed] [Google Scholar]
- 16.Miller O L, Jr, Beatty B R. Science. 1969;164:955–957. doi: 10.1126/science.164.3882.955. [DOI] [PubMed] [Google Scholar]
- 17.Singh O P, Björkroth B, Masich S, Wieslander L, Daneholt B. Exp Cell Res. 1999;251:135–146. doi: 10.1006/excr.1999.4490. [DOI] [PubMed] [Google Scholar]
- 18.Miralles F, Öfverstedt L-G, Sabri N, Aissouni Y, Hellman U, Skoglund U, Visa N. J Cell Biol. 2000;148:271–282. doi: 10.1083/jcb.148.2.271. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Mehlin H, Daneholt B, Skoglund U. Cell. 1992;69:605–613. doi: 10.1016/0092-8674(92)90224-z. [DOI] [PubMed] [Google Scholar]
- 20.Wurtz T, Lönnroth A, Ovchinnikov L, Skoglund U, Daneholt B. Proc Natl Acad Sci USA. 1990;87:831–835. doi: 10.1073/pnas.87.2.831. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Dreyfuss G, Matunis M J, Pinol-Roma S, Burd C G. Annu Rev Biochem. 1993;62:289–321. doi: 10.1146/annurev.bi.62.070193.001445. [DOI] [PubMed] [Google Scholar]
- 22.Krecic A M, Swanson M S. Curr Opin Cell Biol. 1999;11:363–371. doi: 10.1016/S0955-0674(99)80051-9. [DOI] [PubMed] [Google Scholar]
- 23.Bennett M, Pinol-Roma S, Staknis D, Dreyfuss G, Reed R. Mol Cell Biol. 1992;12:3165–3175. doi: 10.1128/mcb.12.7.3165. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Matunis E L, Matunis M J, Dreyfuss G. J Cell Biol. 1993;121:219–228. doi: 10.1083/jcb.121.2.219. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Wurtz T, Kiseleva E, Nacheva G, Alzhanova-Ericsson A, Rosén A, Daneholt B. Mol Cell Biol. 1996;16:1425–1435. doi: 10.1128/mcb.16.4.1425. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Pinol-Roma S, Choi Y D, Matunis M M, Dreyfuss G. Genes Dev. 1988;2:215–227. doi: 10.1101/gad.2.2.215. [DOI] [PubMed] [Google Scholar]
- 27.Visa N, Izaurralde E, Ferreira J, Daneholt B, Mattaj I W. J Cell Biol. 1996;133:5–14. doi: 10.1083/jcb.133.1.5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Lewis J D, Izaurralde E. Eur J Biochem. 1997;247:461–469. doi: 10.1111/j.1432-1033.1997.00461.x. [DOI] [PubMed] [Google Scholar]
- 29.Kiseleva E, Wurtz T, Visa N, Daneholt B. EMBO J. 1994;13:6052–6061. doi: 10.1002/j.1460-2075.1994.tb06952.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Baurén G, Wieslander L. Cell. 1994;76:183–192. doi: 10.1016/0092-8674(94)90182-1. [DOI] [PubMed] [Google Scholar]
- 31.Alzhanova-Ericsson A, Sun X, Visa N, Kiseleva E, Wurtz T, Daneholt B. Genes Dev. 1996;10:2881–2893. doi: 10.1101/gad.10.22.2881. [DOI] [PubMed] [Google Scholar]
- 32.Sun X, Alzhanova-Ericsson A, Visa N, Aissouni Y, Zhao J, Daneholt B. J Cell Biol. 1998;142:1167–1180. doi: 10.1083/jcb.142.5.1181. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Fu X-D. RNA. 1995;1:663–680. [PMC free article] [PubMed] [Google Scholar]
- 34.Ge H, Zuo P, Manley J L. Cell. 1991;66:373–382. doi: 10.1016/0092-8674(91)90626-a. [DOI] [PubMed] [Google Scholar]
- 35.Krainer A R, Mayeda A, Kozak D, Binns G. Cell. 1991;66:383–394. doi: 10.1016/0092-8674(91)90627-b. [DOI] [PubMed] [Google Scholar]
- 36.Champlin D T, Frasch M, Saumweber H, Lis J T. Genes Dev. 1991;5:1611–1621. doi: 10.1101/gad.5.9.1611. [DOI] [PubMed] [Google Scholar]
- 37.Roth M B, Zahler A M, Stolk J A. J Cell Biol. 1991;115:587–596. doi: 10.1083/jcb.115.3.587. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Brand S F, Pichoff S, Noselli S, Bourbon H-M. Genes. 1995;154:187–192. doi: 10.1016/0378-1119(94)00840-o. [DOI] [PubMed] [Google Scholar]
- 39.Labourier E, Bourbon H-M, Gallouzi I, Fostier M, Allemand E, Tazi J. Genes Dev. 1999;13:740–753. doi: 10.1101/gad.13.6.740. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Nakielny S, Dreyfuss G. J Cell Biol. 1996;134:1365–1373. doi: 10.1083/jcb.134.6.1365. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Izaurralde E, Lewis J, Gamberi C, Jarmolowski A, McGuigan C, Mattaj I W. Nature (London) 1995;376:709–712. doi: 10.1038/376709a0. [DOI] [PubMed] [Google Scholar]
- 42.Hamm J, Mattaj I W. Cell. 1990;63:109–118. doi: 10.1016/0092-8674(90)90292-m. [DOI] [PubMed] [Google Scholar]
- 43.Jarmolowski A, Boelens W C, Izaurralde E, Mattaj I W. J Cell Biol. 1994;124:627–635. doi: 10.1083/jcb.124.5.627. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Visa N, Alzhanova-Ericsson, Sun S, Kiseleva E, Björkroth B, Wurtz T, Daneholt B. Cell. 1996;84:253–264. doi: 10.1016/s0092-8674(00)80980-0. [DOI] [PubMed] [Google Scholar]
- 45.Percipalle P, Zhao J, Pope B, Weeds A, Lindberg U, Daneholt B. J Cell Biol. 2001;153:229–236. doi: 10.1083/jcb.153.1.229. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Pinol-Roma S, Dreyfuss G. Nature. 1992;355:730–732. doi: 10.1038/355730a0. [DOI] [PubMed] [Google Scholar]
- 47.Michael W M, Choi M, Dreyfuss G. Cell. 1995;83:415–422. doi: 10.1016/0092-8674(95)90119-1. [DOI] [PubMed] [Google Scholar]
- 48.Svitkin Y V, Ovchinnikov L P, Dreyfuss G, Sonenburg N. EMBO J. 1996;15:7147–7155. [PMC free article] [PubMed] [Google Scholar]
- 49.D'Angelo E G. Biol Bull. 1946;90:71–87. [PubMed] [Google Scholar]
- 50.Clark T G, Merriam R W. Cell. 1977;12:883–891. doi: 10.1016/0092-8674(77)90152-0. [DOI] [PubMed] [Google Scholar]
- 51.Skoglund U, Andersson K, Strandberg B, Daneholt B. Nature (London) 1986;319:560–564. doi: 10.1038/319560a0. [DOI] [PubMed] [Google Scholar]
- 52.Wada A, Fukuda M, Mishima M, Nishida E. EMBO J. 1998;17:1635–1641. doi: 10.1093/emboj/17.6.1635. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Mamiya N, Worman H J. J Biol Chem. 1999;274:15751–15756. doi: 10.1074/jbc.274.22.15751. [DOI] [PubMed] [Google Scholar]
- 54.Leroy P, Alzari P, Sassoon D, Wolgemuth D, Fellous M. Cell. 1989;57:549–559. doi: 10.1016/0092-8674(89)90125-6. [DOI] [PubMed] [Google Scholar]
- 55.Gururajan R, Perry-O'Keefe H, Melton D A, Weeks D L. Nature (London) 1991;349:717–719. doi: 10.1038/349717a0. [DOI] [PubMed] [Google Scholar]
- 56.Jamieson D J, Rahe B, Pringle J, Beggs J D. Nature (London) 1991;349:715–717. doi: 10.1038/349715a0. [DOI] [PubMed] [Google Scholar]
- 57.Chuang R Y, Weaver P L, Liu Z, Chang T H. Science. 1997;275:1468–1471. doi: 10.1126/science.275.5305.1468. [DOI] [PubMed] [Google Scholar]
- 58.Lall S, Francis-Lang H, Flament A, Norvell A, Schüpbach T, Ish-Horowicz D. Cell. 1999;98:171–180. doi: 10.1016/s0092-8674(00)81012-0. [DOI] [PubMed] [Google Scholar]
- 59.Kamma H, Portman D S, Dreyfuss G. Exp Cell Res. 1995;221:187–196. doi: 10.1006/excr.1995.1366. [DOI] [PubMed] [Google Scholar]
- 60.Mayeda A, Krainer A R. Cell. 1992;68:365–375. doi: 10.1016/0092-8674(92)90477-t. [DOI] [PubMed] [Google Scholar]