Abstract
Experiments presented here show that each of the 1.5-kilobase inverted repeats of the kanamycin-resistance transposon Tn5 is transposable; we designate them IS50-L (left) and IS50-R (right). By DNA sequence analyses, IS50 is 1533 base pairs (bp) long and generates 9-bp direct repeats of target sequences. The ends of IS50 comprise a hyphenated 8-of-9-bp inverted repeat and are not used with equal efficiency; the outside ends are more active than the inside ends, suggesting that a strong transposase recognition site at the outside ends, suggesting that a strong transposase recognition site at the outside end extends beyond the 8 bp common to both ends.
Full text
PDF
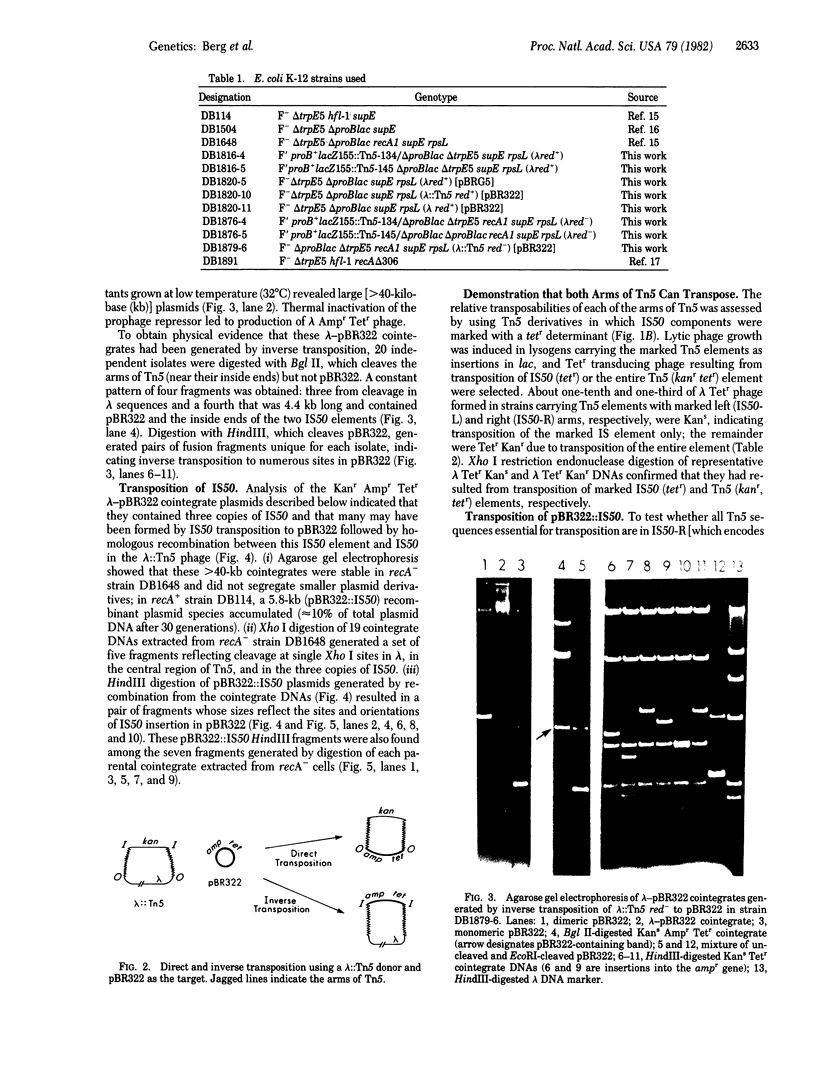
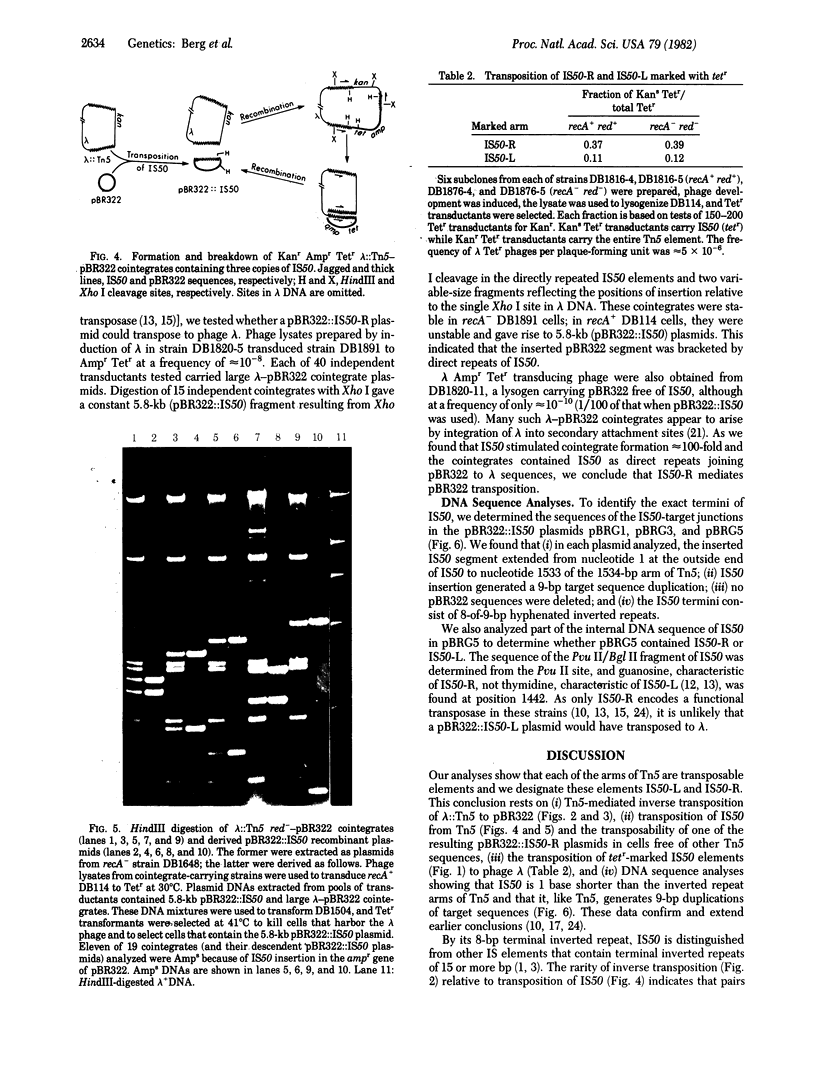



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Auerswald E. A., Ludwig G., Schaller H. Structural analysis of Tn5. Cold Spring Harb Symp Quant Biol. 1981;45(Pt 1):107–113. doi: 10.1101/sqb.1981.045.01.019. [DOI] [PubMed] [Google Scholar]
- Berg D. E., Drummond M. Absence of DNA sequences homologous to transposable element Tn5 (Kan) in the chromosome of Escherichia coli K-12. J Bacteriol. 1978 Oct;136(1):419–422. doi: 10.1128/jb.136.1.419-422.1978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Berg D. E., Egner C., Hirschel B. J., Howard J., Johnsrud L., Jorgensen R. A., Tlsty T. D. Insertion, excision, and inversion of Tn5. Cold Spring Harb Symp Quant Biol. 1981;45(Pt 1):115–123. doi: 10.1101/sqb.1981.045.01.020. [DOI] [PubMed] [Google Scholar]
- Berg D. E., Weiss A., Crossland L. Polarity of Tn5 insertion mutations in Escherichia coli. J Bacteriol. 1980 May;142(2):439–446. doi: 10.1128/jb.142.2.439-446.1980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Calos M. P., Miller J. H. Transposable elements. Cell. 1980 Jul;20(3):579–595. doi: 10.1016/0092-8674(80)90305-0. [DOI] [PubMed] [Google Scholar]
- Chandler M., Roulet E., Silver L., Boy de la Tour E., Caro L. Tn10 mediated integration of the plasmid R100.1 into the bacterial chromosome: inverse transposition. Mol Gen Genet. 1979 May 23;173(1):23–30. doi: 10.1007/BF00267687. [DOI] [PubMed] [Google Scholar]
- Egner C., Berg D. E. Excision of transposon Tn5 is dependent on the inverted repeats but not on the transposase function of Tn5. Proc Natl Acad Sci U S A. 1981 Jan;78(1):459–463. doi: 10.1073/pnas.78.1.459. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grindley N. D., Joyce C. M. Analysis of the structure and function of the kanamycin-resistance transposon Tn903. Cold Spring Harb Symp Quant Biol. 1981;45(Pt 1):125–133. doi: 10.1101/sqb.1981.045.01.021. [DOI] [PubMed] [Google Scholar]
- Guyer M. S., Reed R. R., Steitz J. A., Low K. B. Identification of a sex-factor-affinity site in E. coli as gamma delta. Cold Spring Harb Symp Quant Biol. 1981;45(Pt 1):135–140. doi: 10.1101/sqb.1981.045.01.022. [DOI] [PubMed] [Google Scholar]
- Heffron F., McCarthy B. J., Ohtsubo H., Ohtsubo E. DNA sequence analysis of the transposon Tn3: three genes and three sites involved in transposition of Tn3. Cell. 1979 Dec;18(4):1153–1163. doi: 10.1016/0092-8674(79)90228-9. [DOI] [PubMed] [Google Scholar]
- Isberg R. R., Syvanen M. Replicon fusions promoted by the inverted repeats of Tn5. The right repeat is an insertion sequence. J Mol Biol. 1981 Jul 25;150(1):15–32. doi: 10.1016/0022-2836(81)90322-3. [DOI] [PubMed] [Google Scholar]
- Jorgensen R. A., Rothstein S. J., Reznikoff W. S. A restriction enzyme cleavage map of Tn5 and location of a region encoding neomycin resistance. Mol Gen Genet. 1979;177(1):65–72. doi: 10.1007/BF00267254. [DOI] [PubMed] [Google Scholar]
- Kleckner N. Transposable elements in prokaryotes. Annu Rev Genet. 1981;15:341–404. doi: 10.1146/annurev.ge.15.120181.002013. [DOI] [PubMed] [Google Scholar]
- Maxam A. M., Gilbert W. Sequencing end-labeled DNA with base-specific chemical cleavages. Methods Enzymol. 1980;65(1):499–560. doi: 10.1016/s0076-6879(80)65059-9. [DOI] [PubMed] [Google Scholar]
- Pogue-Geile K. L., DasSarma S., King S. R., Jaskunas S. R. Recombination between bacteriophage lambda and plasmid pBR322 in Escherichia coli. J Bacteriol. 1980 Jun;142(3):992–1003. doi: 10.1128/jb.142.3.992-1003.1980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ross D. G., Grisafi P., Kleckner N., Botstein D. The ends of Tn10 are not IS3. J Bacteriol. 1979 Sep;139(3):1097–1101. doi: 10.1128/jb.139.3.1097-1101.1979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rothstein S. J., Jorgensen R. A., Yin J. C., Yong-di Z., Johnson R. C., Reznikoff W. S. Genetic organization of Tn5. Cold Spring Harb Symp Quant Biol. 1981;45(Pt 1):99–105. doi: 10.1101/sqb.1981.045.01.018. [DOI] [PubMed] [Google Scholar]
- Schmitt R., Altenbuchner J., Wiebauer K., Arnold W., Pühler A., Schöffl F. Basis of transposition and gene amplification by Tn1721 and related tetracycline-resistance transposons. Cold Spring Harb Symp Quant Biol. 1981;45(Pt 1):59–65. doi: 10.1101/sqb.1981.045.01.011. [DOI] [PubMed] [Google Scholar]
- So M., Heffron F., McCarthy B. J. The E. coli gene encoding heat stable toxin is a bacterial transposon flanked by inverted repeats of IS1. Nature. 1979 Feb 8;277(5696):453–456. doi: 10.1038/277453a0. [DOI] [PubMed] [Google Scholar]
- Sutcliffe J. G. pBR322 restriction map derived from the DNA sequence: accurate DNA size markers up to 4361 nucleotide pairs long. Nucleic Acids Res. 1978 Aug;5(8):2721–2728. doi: 10.1093/nar/5.8.2721. [DOI] [PMC free article] [PubMed] [Google Scholar]