Abstract
The integration of sample preparation methods into microfluidic devices provides automation necessary for achieving complete micro total analysis systems. We have developed a technique that combines on-chip sample enrichment with fluorescence labeling and purification. Polymer monoliths made from butyl methacrylate were fabricated in cyclic olefin copolymer microdevices and used for solid phase extraction. We studied the retention of fluorophores, amino acids and proteins on these columns. The retained samples were subsequently labeled with both Alexa Fluor 488 and Chromeo P503, and unreacted dye was rinsed off the column before sample elution. Additional purification was obtained from the differential retention of proteins and fluorescent labels. A linear relation between the eluted peak areas and concentrations of on-chip labeled heat shock protein 90 samples demonstrated the utility of this method for on-chip quantitation. Our fast and simple method of simultaneously concentrating and labeling samples on-chip is compatible with miniaturization and desirable for automated analysis.
1 Introduction
The integration of multiple functions in a single device can result in faster, cheaper and improved analysis compared to traditional laboratory methods [1]. Many such processes have been integrated in microfluidic devices, including extraction/purification [2-6], labeling [7,8], preconcentration [9,10], microdialysis [11,12], and detection [13,14]. One of the greatest difficulties in achieving completely miniaturized and integrated analysis has been the step of sample preparation [15], although important progress is being made in selected areas as noted below. Importantly, solid phase extraction (SPE) has been used in integrated sample processing, including extraction, purification and preconcentration [2,5,6].
SPE is a common sample preparation method wherein analytes are retained on a solid support and are subsequently eluted in a concentrated form [16]. The most common SPE modes in microfluidics are affinity [4,17,18] and reversed-phase [6,19,20]. Affinity SPE in microchips has been used to extract and quantify four cancer biomarkers in blood [4], to preconcentrate and purify PCR products [17], and to extract thiazole orange-conjugated adenosine monophosphate [18]. Reversed-phase columns are useful in the extraction of non-polar to moderately polar compounds. Silica-based materials are common reversed-phase SPE supports, having been used for the extraction of parabens and fluorescent dyes [6], the preconcentration of peptides and cytochrome c [19], and the concentration and separation of Rhodamine 123 and fluorescein isothiocyanate (FITC)-labeled ephedrine [20]. Monolithic columns are seeing increased usage because they can be easily prepared on-chip without the need for retaining structures like frits [5,21], and the porosity and surface area can be tuned by varying the monomer/porogen composition [22]. Neutral methacrylates are generally hydrophobic enough for reversed-phase SPE [23,24]. Cyclic olefin copolymer (COC) is a preferred polymer material for SPE microchips because of its stability in organic solvents such as acetonitrile that are used for elution [24,25]. Though photografting is generally used to modify the microchannel for enhanced monolith/wall adhesion [24], it has also been shown that monoliths fabricated in COC devices can be stable without surface pretreatment [26].
The integration of SPE with capillary electrophoresis or microchip electrophoresis (μCE) offers the advantages of improved sensitivity and sample cleanup, along with shorter analysis times, reduced sample loss and automation [20,27]. Typically, when SPE is coupled to μCE, an interface is used to control the transfer of analytes from the SPE column to the separation channel. In the analysis of dopamine by SPE-μCE, polydimethylsiloxane (PDMS) microvalves were used to segregate the processes of extraction, rinsing, sample elution and separation [23]. In a different setup, a nanoporous membrane sandwiched between two PDMS layers was used as an electrokinetic valve to separate the processes of SPE and electrophoretic separation [20]. In these PDMS SPE-μCE systems, separation of small molecules was done, which is less complicated since they typically do not bind as much as proteins to the device walls [28].
Many samples do not fluoresce naturally and have to be derivatized to take advantage of the superior sensitivity of laser-induced fluorescence detection. Labeling is often performed off-chip, but on-chip labeling has been achieved both in pre-column [7,8,29] and post-column [30-32] formats. Conventional dyes with high quantum yield, such as Alexa Fluor and fluorescein, are often used to label analytes. Additionally, fluorogenic reagents [7,8,33], which are weakly fluorescent until they react with a primary amine, have been used for on-chip derivatization because they produce lower background fluorescence, their reaction kinetics are fast, and they do not change the electrical charge of the sample [34]. The fluorogenic reagents, CE dye 503 [7], ThioGlo-1 [8], and naphthalene-2,3-dicarboxaldehyde [35], have been used for on-chip derivatization. While acceptable results were obtained from these integrated systems, lower limits of detection, avoiding on-line mixing of high concentrations of fluorescent dyes, and addressing system peaks and background fluorescence are all areas where improvement is desirable.
In this work we demonstrate a novel approach combining SPE with on-chip labeling and purification to improve over previous methods. We show that samples retained on a solid support can be concentrated and labeled on-chip prior to elution. Reversed-phase butyl methacrylate (BMA) porous polymer monoliths were formed in COC microdevices and used to study the retention of fluorophores, amino acids and proteins. The retained and concentrated samples were then labeled on-chip with Alexa Fluor 488 TFP ester or Chromeo P503. Subsequent rinsing to remove unreacted dye and selective elution of labeled sample relative to unconjugated fluorophore helped to greatly reduce the background fluorescence typically observed in on-chip labeling. On-chip labeling of heat shock protein 90 (HSP90) resulted in a concentration-dependent area of the eluted peak, demonstrating the ability of this method to quantify on-chip labeled samples. This work thus offers improved capabilities in on-chip labeling for miniaturized analysis.
2. Experimental
2.1 Reagents and materials
Zeonor 1020R (COC) was purchased from Zeon Chemicals (Louisville, KY, USA). Methyl methacrylate (MMA), BMA, lauryl methacrylate (LMA), 2,2-dimethoxy-2-phenylacetophenone (DMPA), 1-dodecanol, cyclohexanol, Tween 20, ethylene dimethacrylate (EDMA), and isopropyl alcohol were obtained from Sigma-Aldrich (St. Louis, MO, USA). Bovine serum albumin (BSA) was purchased from New England Biolabs (Ipswich, MA, USA) and HSP90 was from Sigma-Aldrich. The amino acids glycine, aspartic acid, phenylalanine and arginine were obtained from Sigma-Aldrich. The amino acids were labeled with FITC, while the proteins were labeled with Alexa Fluor 488 TFP ester. Both fluorophores were from Invitrogen (Carlsbad, CA, USA). Chromeo P503 was obtained from Active Motif (Carlsbad, CA, USA). Fluorescein (sodium salt) and dimethyl sulfoxide (DMSO) were purchased from Sigma-Aldrich. Hydroxypropyl cellulose (HPC, 100 kDa average molecular weight) was from Aldrich (Milwaukee, WI, USA). Sodium dodecyl sulfate (SDS) was obtained from Fisher Scientific (Pittsburgh, PA, USA). Buffer solutions were made from anhydrous sodium carbonate, sodium bicarbonate, acetonitrile (ACN), and sodium azide, all from EMD Chemicals (Gibbstown, NJ, USA). Amicon Ultra-0.5 centrifugal filter devices were obtained from Millipore (Billerica, MA, USA). All solutions were prepared with deionized water (18.3 MΩ cm) purified by a Barnstead EASYpure UV/UF system (Dubuque, IA, USA).
2.2 Device fabrication
COC plates were obtained by cutting the sheets into 2″ × 1″ pieces with a bandsaw. Holes in the cover plate were then drilled to serve as reservoirs in the bonded devices. The microdevices were fabricated using a combination of photolithographic patterning, etching, hot embossing and thermal bonding as described previously [36]. Bonding of COC was done at 110 °C for 20 min. Two different microchip designs were used for these experiments. A simple, two-reservoir layout (Fig. 1A) was used for initial testing while the design in Figure 1B was used for integrated experiments where no exchange of liquids in reservoirs was required. The channels in both designs were ~15 μm deep and ~50 μm wide. Before polymerization of a monolith, the channels were rinsed with isopropyl alcohol.
Figure 1.
Schematic of microfluidic devices used for on-chip labeling. a) Simple, two reservoir design used for initial testing. b) Layout used for integrated experiments. The reservoirs are: 1 – sample, 3 – fluorescent dye, 4 – rinse buffer, 5 – eluent, and 2 and 6 – buffer. The lengths from reservoirs 1, 2, 4 and 5 to the injection intersection are all 0.5 cm.
2.3 Preparation of SPE monoliths
Monoliths were made from a solution consisting of 25% BMA (or MMA or LMA), 15% EDMA, 25% (w/w) dodecanol, 10% cyclohexanol, and 25% Tween 20. 1% DMPA was added to the mixture as photoinitiator. The solution was sonicated for 10 min and degassed for 5 min. It was then filled into the device, and a mask was used to expose only the desired portion of the chip to UV radiation. Exposure was carried out with the use of a SunRay 600 UV floodlight from Uvitron International (West Springfield, MA, USA) at 50 mW/cm2 for 10 min. A 2 mm long monolith was formed in each microdevice in the location indicated in Figure 1. After polymerization, devices were rinsed with isopropyl alcohol followed by buffer (10 mM carbonate, pH 9.3). The morphology of the monoliths was characterized using a Philips XL30 FEG environmental scanning electron microscope (SEM) from FEI (Hillsboro, OR, USA).
To determine the loading capacity of the monoliths, increasing concentrations of BSA were loaded on column using a syringe pump (Harvard Apparatus, Holliston, MA, USA) operating at 20 μL/min. First, the column was preconditioned with carbonate buffer containing 30% ACN, and then each BSA solution was loaded for 10 min (200 μL total volume), followed by a 5 min rinse with aqueous carbonate buffer. BSA retention was monitored via the background-subtracted fluorescent intensity at the CCD detector.
2.4 Off-chip labeling
Amino acids were separately mixed with FITC at a 4:1 molar ratio and incubated at room temperature for 24 hr. This ratio ensures that almost no unreacted dye is left at the end of the labeling process. BSA and HSP90 were labeled with Alexa Fluor 488 TFP ester as described previously [9]. The labeled samples were analyzed by μCE as described previously [9], to confirm the lack of free dye, before loading into the monolithic column. A stock solution of Chromeo P503 was made by dissolving the dye in DMSO to a concentration of 0.8 mg/mL. For off-chip labeling of BSA with Chromeo P503, the protein and dye were mixed to final concentrations of 5 μg/mL and 70 μg/mL, respectively, and incubated for 30 min at room temperature.
2.5 Microdevice operation
Before sample loading, monolithic columns were preconditioned with different solutions to find the most favorable conditions for sample adsorption. The different pretreatments included rinsing the monolith with aqueous carbonate buffer (10 mM, pH 9.3), with carbonate buffer containing 30% ACN just prior to sample loading, or with carbonate buffer containing 30% ACN followed by thorough rinsing with aqueous carbonate buffer. We used pH 9.3 for all buffers except the eluent, because at pH values above 9 the ε-amino groups on proteins are mainly deprotonated [37], facilitating labeling.
After column preconditioning, device operation for the simple design (Fig. 1A) was as follows. For retention and preconcentration studies, labeled sample in carbonate buffer was loaded on the monolithic column by applying +400 V to reservoir 2 and grounding reservoir 1 for 5 min. Rinsing was done by replacing the sample in reservoir 1 with carbonate buffer and applying the same voltages as before for 2 min. For elution, the rinse buffer in reservoir 1 was replaced with eluent consisting of 85% ACN, 15% carbonate buffer (7.5 mM, pH 9.6), 0.05% HPC, and 0.05% SDS. Elution was accomplished by grounding reservoir 1 and applying +1000 V to reservoir 2. On-chip labeling with Alexa Fluor 488 (TFP ester) was done by transferring protein solution into reservoir 1 and applying the same column loading voltages as above for 10 min. Next, 50 μg/mL of labeling solution was placed in reservoir 1, which was grounded, and +400 V were applied to reservoir 2 for 15 min. This was followed by rinsing and elution as above. For on-chip labeling with Chromeo P503 the voltage polarity was reversed during loading of the dye and the rinse because of the positive charge on the label. The polarity was restored to positive for elution.
Experiments with the design in Figure 1B were done as follows (reservoirs not having a potential applied were allowed to float). Labeled sample was loaded by applying +400 V to reservoir 2 and grounding reservoir 1 for 5 min. Rinsing was achieved by applying the same potential to reservoir 2 and grounding reservoir 4. Elution was accomplished by applying +1000 V between reservoirs 5 and 2. For on-chip labeling with this design, unlabeled sample was loaded as before, then the dye was driven through the column by applying +400 V between reservoirs 3 and 2. This was followed by rinse and elution as above.
2.6 Instrumentation
The laser induced fluorescence system has been described previously [34,36]. A Nikon Eclipse TE300 inverted microscope equipped with a CCD camera (Coolsnap HQ, Roper Scientific, Sarasota, FL, USA) was used. A 10x expander was used to increase the laser beam diameter, which was directed to a 20x, 0.45 NA objective on the microscope. For fluorescence monitoring, the detection point was positioned either just to the left of reservoir 2 (Fig. 1A or B), or directly on the monolith. The collected CCD images were analyzed using V++ Precision Digital Imaging software (Auckland, New Zealand).
2.7 Data analysis
To evaluate the extent to which different samples adsorbed on the monolith, amino acids, fluorescent dyes and two proteins (BSA and HSP90) were loaded. Their retention was monitored via CCD detection by measuring the background-subtracted fluorescent intensity on the monolith after rinsing.
For protein preconcentration, the background-subtracted steady state fluorescent intensity before the sample reached the monolith, which is the signal without preconcentration, was compared to the elution intensity, obtained by subtracting the baseline from the peak maximum. The calculation of HSP90 concentration for the calibration curve was based on the eluted peak area.
3 Results and discussion
3.1 Preparation of monoliths
We fabricated thermally bonded COC microdevices [38] with reversed-phase monolithic columns for SPE. COC was chosen because of its resistance to common organic solvents like acetonitrile [25] that are normally used for elution in SPE. Fabrication of a 2 mm long monolith in these microchips was done with a ternary porogenic solvent system consisting of dodecanol, cyclohexanol, and Tween 20 [39]. Optimization of these porogens was carried out by Pagaduan et al. [39] who found that the monoliths formed were porous enough for water and buffer to flow through by capillary action. The monolith was made from a mixture consisting of a 60:40 porogen-to-monomer ratio, which provided more surface area and hence sample binding sites than monoliths with a higher ratio [40]. Importantly, experiments carried out in our monoliths showed that the application of voltage did not cause movement of the monolith, in agreement with Ladner et al [26]. Thus, pretreatment methods like photografting [24,26] that complicate fabrication, adsorb sample and cause channel clogging were avoided. The monolithic structure seen in Figure 2 showed clusters of globules with irregular through-pores, typical of high surface area monoliths.
Figure 2.

SEM image of a BMA monolith showing detailed morphology.
BMA was chosen over other monomers such as MMA and LMA, which formed monoliths with morphologies shown in Supplemental Figure S1. While MMA showed even packing and large pores, LMA produced a monolith with small through-pores. Even when the LMA monomer concentration was reduced to 25%, the very small through-pore dimensions were not favorable for flow. MMA was the least hydrophobic while LMA was the most hydrophobic of the monoliths fabricated. When LMA columns were used for protein extraction, elution was difficult because of the strong hydrophobic interactions with proteins. BMA, therefore, with intermediate properties between MMA and LMA, was best suited for selective retention and elution in these experiments. Monoliths prepared with BMA and EDMA have been shown to have comparable hydrophobicity to C18 beads [41]; indeed, BMA monoliths have been used for SPE of proteins [5,42].
3.2 Retention of samples on BMA monoliths
An initial study of column preparation and sample loading conditions for BMA monoliths is summarized in Supplemental Figure S2. Retention was greatest when the monolith was rinsed with carbonate buffer containing 30% ACN just before sample loading. It was also observed that samples dissolved in aqueous carbonate buffer or buffer having ≤0.5% ACN showed the best retention, while samples dissolved in carbonate buffer containing 2% ACN were retained less on the column. Rinsing reversed-phase monoliths with an ACN-aqueous buffer mixture has been shown to be necessary for preconditioning [43], removing impurities that may interfere with adsorption. Preconditioning with ACN also helps to activate and/or hydrate the monolith surface to provide adequate contact with the liquid sample [44].
Figure 3 shows the normalized retention of fluorescent dyes, amino acids and proteins on BMA monoliths. Retention of the fluorescent dyes (Fig. 3A) on the column was relatively low. Unreacted (free) Chromeo P503 has a low fluorescence signal [45], which explains its limited retention. The ionic sodium salt of fluorescein used does not partition significantly into the hydrophobic BMA column, accounting for its low retention. The slightly higher fluorescent intensities observed for Alexa Fluor 488 TFP ester and FITC are likely due to small differences in experimental conditions.
Figure 3.
Normalized retention of a) fluorescent dyes, and b) amino acids and proteins on an on-chip BMA column. Loading was the same for all samples, except the polarity was reversed for Chromeo P503. Values were normalized to 100 nM dye concentration in (a) and 100 ng/mL amino acid or protein concentration in (b).
Retention of amino acids (Fig. 3B) on our BMA monolith depended on their hydrophobicity. Hydropathicity index, a measure of hydrophobicity of amino acids and proteins [46], can predict their expected interaction with the monolith. More hydrophobic compounds have a more positive index value. Low retention was observed for aspartic acid and glycine, which have hydrophilic values of -3.5 and -0.4, respectively. As expected, phenylalanine, with an index of 2.8, was retained more. The two proteins in Figure 3B were more highly retained on BMA monoliths than any amino acids. Protein binding comes from the average surface hydrophobicity, which combines the contributions of all the amino acids present on the surface [47]. Some of these amino acids, such as phenylalanine, alanine, and methionine, are hydrophobic and form hydrophobic regions or “patches” on protein surfaces [48,49]. The non-specific adsorption of proteins like BSA to hydrophobic surfaces has been shown to be the result of interactions between these hydrophobic patches and surface [50]. These more extensive hydrophobic surface interactions for the two proteins studied (relative to the free amino acids) result in their greater retention on these monoliths. We determined the protein loading capacity of our 2 mm long monoliths to be 2 μg (30 pmol) of BSA.
3.3 Elution of samples
Figure 4 shows elution profiles of fluorescent dyes on a BMA column. The areas of the eluted peaks were correlated with the amount retained (see Fig. 3A). The elution profiles of some amino acids and the proteins, BSA and HSP90, from a BMA monolith can be seen in Figure 5. The areas of the eluted amino acid peaks (Figure 5A) followed the order of the amount retained (Fig. 3B) with aspartic acid having the lowest elution intensity and phenylalanine the highest. The earlier elution of HSP90 compared to BSA (Fig. 5B) is likely due to the fact that HSP90 has a greater negative charge density [51], and is not only more hydrophilic but also migrates faster in an electric field, both factors in electrochromatography [52]. A comparison of the intensities of the peaks at elution (Fig. 5B) to the signal before the sample reached the monolithic column gave the degree of preconcentration of each protein. The calculated enrichment factors were 11-fold for HSP90 and 6-fold for BSA, demonstrating that these proteins were concentrated on the reversed-phase column. The degree of preconcentration of HSP90 was higher than for BSA because it eluted in a ~2-fold narrower band, since it was less retained than BSA. It is expected that with longer columns and loading times the degree of preconcentration could be even greater.
Figure 4.
Elution profiles of fluorescent dyes from an on-chip BMA column. All signals were normalized to 100 nM dye concentration, as in Figure 3. Dyes (from bottom to top) are: Chromeo P503, fluorescein, FITC, and Alexa Fluor 488 TFP ester. Chromatograms are offset vertically for clarity. Positively charged Chromeo P503 dye was eluted with application of a negative voltage, while all others (negatively charged) required a positive voltage. The first 30 s of Chromeo P503 dye elution is expanded in the inset.
Figure 5.
Elution profiles of amino acids and two proteins (BSA and HSP90) from an on-chip BMA monolith. a) Chromatograms of amino acids (100 ng/mL) from bottom to top are: aspartic acid, glycine, and phenylalanine. Chromatograms are offset vertically for clarity. b) Elution of 1 μg/mL proteins (BSA – maximum around 55 s, and HSP90 – maximum around 30 s).
3.4 Off- and on-chip labeling with Chromeo P503
Figure 6 shows elution profiles for BSA labeled off- and on-chip with Chromeo P503 dye. Although Chromeo P503 dye is positively charged, after reaction with BSA the labeled protein is negatively charged at pH 9.3. Elution of Chromeo P503-labeled BSA from the column (Fig. 6A) is slower than Alexa Fluor 488-labeled BSA (Fig. 5B) because Chromeo P503-labeled protein has less negative charge [53,54]. Figure 6B shows that the elution of BSA labeled on-chip with Chromeo P503 is similar to that for protein labeled off-chip. For on-chip labeling, the oppositely charged Chromeo P503 and labeled protein migrate in opposite directions under the elution voltage, simplifying purification. This use of charge to selectively elute proteins relative to dye makes on-chip labeling of protein samples with Chromeo P503 appealing.
Figure 6.
Elution profiles of 1 μg/mL BSA labeled a) off-chip and b) on-chip with Chromeo P503 dye.
3.5 On-chip labeling with Alexa Fluor 488 (TFP ester)
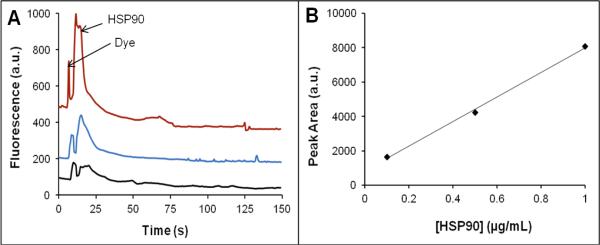
Alexa Fluor 488, which produces very bright and photostable conjugates [55,56], was used with HSP90 to demonstrate proof of principle of on-chip preconcentration, labeling and purification of protein from label within an integrated microfluidic platform. Figure 7 shows the elution of different concentrations of on-chip labeled HSP90 from a BMA column. A 15 min on-chip labeling of 1 μg/mL HSP90 with Alexa Fluor 488 TFP ester in Fig. 7A showed two peaks, the dye followed by HSP90. On-chip labeling of different concentrations of HSP90 retained on a BMA monolith showed increasing peak areas with concentration. A plot of peak area against HSP90 concentration (Fig. 7B) was linear, showing that concentration and signal can easily be correlated for an unknown sample after on-chip labeling. We note that these systems could be used to concentrate and label other proteins in a similar manner. Retention of more sample could be attained by increasing the surface area of the monolith through adjustment of polymerization conditions or usage of different branched crosslinkers, as demonstrated by Lee's group [57,58].
Figure 7.
On-chip labeling and elution of HSP90 using a BMA column. a) Elution traces. Chromatograms are offset vertically for clarity (bottom: 0.1 μg/mL, middle: 0.5 μg/mL and top: 1 μg/mL). b) Plot of peak area as a function of HSP90 concentration. The slope is 7200 ± 140, and the intercept is 830 ± 170.
Unlike typical on-chip labeling where the on-line mixing of high concentrations of fluorescent dyes with sample can interfere with the separation process, in our work protein samples are enriched in addition to being labeled and purified from the unreacted dye. These results demonstrate that unlabeled protein samples can be readily assayed by a simple and sensitive procedure that is favorable for automation. Integration of our reversed-phase columns with an up-stream affinity extraction module would further increase the selectivity of the method, especially for complex biological samples. Multiple analytes could also be extracted, labeled and separated in our integrated system.
4 Conclusion
The need for off-chip sample preparation has been a significant obstacle to miniaturized analysis, such that effective integration of sample labeling moves the field closer to the automation level necessary for application in point-of care diagnostics, for example. We have presented a miniaturized system that combines sample enrichment with on-chip labeling and purification. Samples are enriched through SPE on a BMA monolith in a microfluidic device. Analytes are fluorescently labeled while they are on the column, and much of the unreacted dye is removed in a rinsing step. The labeled protein is more strongly retained than the unattached label, enabling further purification during elution. We have characterized the retention and elution of fluorophores and fluorescently labeled amino acids and proteins. We have also developed conditions for automated on-chip fluorescent labeling and purification of proteins. Importantly, the eluted peak area for on-chip labeled HSP90 scaled linearly with concentration, demonstrating the ability of our method to quantify on-chip labeled samples. The combination of the proven enrichment capacity of SPE with fast labeling and separation yields a simple technique that is well suited for miniaturization. The integration of this approach with affinity extraction should provide an additional dimension of specificity, providing a powerful and automated method for bioanalysis.
Supplementary Material
Highlights (for review).
Sample enrichment, fluorescent labeling and purification are combined in a microchip.
Retention and elution of fluorophores, amino acids and proteins are characterized.
Samples are both enriched and fluorescently labeled on an on-chip monolithic column.
Eluted peak areas for on-chip labeled proteins scaled linearly with concentration.
Acknowledgement
This work was supported by the National Institutes of Health (R01 EB006124).
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- 1.Angelescu DE. Highly Integrated Microfluidics Design. Artech House. 2011 [Google Scholar]
- 2.Yang H, Mudrik JM, Jebrail MJ, Wheeler AR. Anal. Chem. 2011;83:3824. doi: 10.1021/ac2002388. [DOI] [PubMed] [Google Scholar]
- 3.Hagan KA, Reedy CR, Bienvenue JM, Dewald AH, Landers JP. Analyst. 2011;136:1928. doi: 10.1039/c0an00922a. [DOI] [PubMed] [Google Scholar]
- 4.Yang W, Yu M, Sun X, Woolley AT. Lab Chip. 2010;10:2527. doi: 10.1039/c005288d. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Hua Y, Jemere AB, Harrison DJ. J. Chromatogr. A. 2011;1218:4039. doi: 10.1016/j.chroma.2011.04.027. [DOI] [PubMed] [Google Scholar]
- 6.Tennico YH, Remcho VT. Electrophoresis. 2010;31:2548. doi: 10.1002/elps.201000256. [DOI] [PubMed] [Google Scholar]
- 7.Yu M, Wang Q, Patterson JE, Woolley AT. Anal. Chem. 2011;83:3541. doi: 10.1021/ac200254c. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Wu J, Ferrance JP, Landers JP, Weber SG. Anal. Chem. 2010;82:7267. doi: 10.1021/ac101182r. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Nge PN, Yang W, Pagaduan JV, Woolley AT. Electrophoresis. 2011;32:1133. doi: 10.1002/elps.201000698. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Beyor N, Yi L, Seo TS, Mathies RA. Anal. Chem. 2009;81:3523. doi: 10.1021/ac900060r. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Nandi P, Desai DP, Lunte SM. Electrophoresis. 2010;31:1414. doi: 10.1002/elps.200900612. [DOI] [PubMed] [Google Scholar]
- 12.Mecker LC, Martin RS. Anal. Chem. 2008;80:9257. doi: 10.1021/ac801614r. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Wu A, Wang L, Jensen E, Mathies R, Boser B. Lab Chip. 2010;10:519. doi: 10.1039/b922830f. [DOI] [PubMed] [Google Scholar]
- 14.Bhatta D, Michel AA, Marti Villalba M, Emmerson GD, Sparrow IJG, Perkins EA, McDonnell MB, Ely RW, Cartwright GA. Biosens. Bioelectron. 2011;30:78. doi: 10.1016/j.bios.2011.08.031. [DOI] [PubMed] [Google Scholar]
- 15.Verbarg J, Kamgar-Parsi K, Shields AR, Howell PB, Jr., Ligler FS. Lab Chip. 2012;12:1793. doi: 10.1039/c2lc21189k. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Svec F. J. Chromatogr. B. 2006;841:52. doi: 10.1016/j.jchromb.2006.03.055. [DOI] [PubMed] [Google Scholar]
- 17.Njoroge SK, Witek MA, Battle KN, Immethun VE, Hupert ML, Soper SA. Electrophoresis. 2011;32:3221. doi: 10.1002/elps.201100274. [DOI] [PubMed] [Google Scholar]
- 18.Nguyen T, Pei R, Stojanovic M, Lin Q. Microfluid. Nanofluid. 2009;6:479. [Google Scholar]
- 19.Wang C, Jemere AB, Harrison DJ. Electrophoresis. 2010;31:3703. doi: 10.1002/elps.201000317. [DOI] [PubMed] [Google Scholar]
- 20.Long Z, Shen Z, Wu D, Qin J, Lin B. Lab Chip. 2007;7:1819. doi: 10.1039/b711741h. [DOI] [PubMed] [Google Scholar]
- 21.Woodward SD, Urbanova I, Nurok D, Svec F. Anal. Chem. 2010;82:3445. doi: 10.1021/ac100392b. [DOI] [PubMed] [Google Scholar]
- 22.Yang Y, Li C, Lee KH, Craighead HG. Electrophoresis. 2005;26:3622. doi: 10.1002/elps.200500121. [DOI] [PubMed] [Google Scholar]
- 23.Kang Q-S, Li Y, Xu J-Q, Su L-J, Li Y-T, Huang W-H. Electrophoresis. 2010;31:3028. doi: 10.1002/elps.201000210. [DOI] [PubMed] [Google Scholar]
- 24.Liu J, Chen C-F, Tsao C-W, Chang C-C, Chu C-C, DeVoe DL. Anal. Chem. 2009;81:2545. doi: 10.1021/ac802359e. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Faure K, Albert M, Dugas V, Crétier G, Ferrigno R, Morin P, Rocca J-L. Electrophoresis. 2008;29:4948. doi: 10.1002/elps.200800235. [DOI] [PubMed] [Google Scholar]
- 26.Ladner Y, Crétier G, Faure K. J. Chromatogr. A. 2010;1217:8001. doi: 10.1016/j.chroma.2010.07.076. [DOI] [PubMed] [Google Scholar]
- 27.Ramautar R, de Jong GJ, Somsen GW. Electrophoresis. 2012;33:243. doi: 10.1002/elps.201100453. [DOI] [PubMed] [Google Scholar]
- 28.Huang B, Wu H, Kim S, Zare RN. Lab Chip. 2005;5:1005. doi: 10.1039/b509251e. [DOI] [PubMed] [Google Scholar]
- 29.Jacobson SC, Hergenroder R, Moore AW, Jr., Ramsey JM. Anal. Chem. 1994;66:4127. [Google Scholar]
- 30.Jacobson SC, Koutny LB, Hergenroeder R, Moore AW, Jr., Ramsey JM. Anal. Chem. 1994;66:3472. [Google Scholar]
- 31.Antes B, Oberkleiner P, Nechansky A, Szolar OHJ. J. Pharmaceut. Biomed. Anal. 2010;51:743. doi: 10.1016/j.jpba.2009.09.022. [DOI] [PubMed] [Google Scholar]
- 32.Ye M, Hu S, Schoenherr RM, Dovichi NJ. Electrophoresis. 2004;25:1319. doi: 10.1002/elps.200305841. [DOI] [PubMed] [Google Scholar]
- 33.Dickerson JA, Ramsay LM, Dada OO, Cermak N, Dovichi NJ. Electrophoresis. 2010;31:2650. doi: 10.1002/elps.201000151. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Yu M, Wang H-Y, Woolley AT. Electrophoresis. 2009;30:4230. doi: 10.1002/elps.200900349. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Yassine O, Morin P, Dispagne O, Renaud L, Denoroy L, Kleimann P, Faure K, Rocca J-L, Ouaini N, Ferrigno R. Anal. Chim. Acta. 2008;609:215. doi: 10.1016/j.aca.2007.12.045. [DOI] [PubMed] [Google Scholar]
- 36.Kelly RT, Woolley AT. Anal. Chem. 2003;75:1941. doi: 10.1021/ac0262964. [DOI] [PubMed] [Google Scholar]
- 37.Michielsen ECHJ, Diris JHC, Hackeng CM, Wodzig WKWH, Van Dieijen-Visser MP. Clin. Chem. 2005;51:222. doi: 10.1373/clinchem.2004.036251. [DOI] [PubMed] [Google Scholar]
- 38.Illa X, Ordeig O, Snakenborg D, Romano-Rodriguez A, Compton RG, Kutter JP. Lab Chip. 2010;10:1254. doi: 10.1039/b926737a. [DOI] [PubMed] [Google Scholar]
- 39.Pagaduan JV, Yang W, Woolley AT. Proc. SPIE. 2011;8031:80311V/1. [Google Scholar]
- 40.Urban J, Jandera P. J. Sep. Sci. 2008;31:2521. doi: 10.1002/jssc.200800182. [DOI] [PubMed] [Google Scholar]
- 41.Yu C, Davey MH, Svec F, Fréchet JMJ. Anal. Chem. 2001;73:5088. doi: 10.1021/ac0106288. [DOI] [PubMed] [Google Scholar]
- 42.Armenta JM, Gu B, Thulin CD, Lee ML. J. Chromatogr. A. 2007;1148:115. doi: 10.1016/j.chroma.2007.02.089. [DOI] [PubMed] [Google Scholar]
- 43.Augustin V, Jardy A, Gareil P, Hennion M-C. J. Chromatogr. A. 2006;1119:80. doi: 10.1016/j.chroma.2006.02.057. [DOI] [PubMed] [Google Scholar]
- 44.Marchiarullo DJ. Dissertation. Doctor of Philosophy, University of Virginia; 2009. Development of microfluidic technologies for on-site clinical and forensic analysis: Extraction, amplification, separation, and detection. [Google Scholar]
- 45.Wojcik R, Swearingen KE, Dickerson JA, Turner EH, Ramsay LM, Dovichi NJ. J. Chromatogr. A. 2008;1194:243. doi: 10.1016/j.chroma.2008.04.042. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Kyte J, Doolittle RF. J. Mol. Biol. 1982;157:105. doi: 10.1016/0022-2836(82)90515-0. [DOI] [PubMed] [Google Scholar]
- 47.Salgado JC, Rapaport I, Asenjo JA. J. Chromatogr. A. 2005;1098:44. doi: 10.1016/j.chroma.2005.08.039. [DOI] [PubMed] [Google Scholar]
- 48.Cummins PM, O'Connor BF. In: Methods in Molecular Biology. Walls D, Loughran ST, editors. Humana Press; 2011. p. 431. [Google Scholar]
- 49.Zhang Y, Martinez T, Woodruff B, Goetze A, Bailey R, Pettit D, Balland A. Anal. Chem. 2008;80:7022. doi: 10.1021/ac800928z. [DOI] [PubMed] [Google Scholar]
- 50.Mücksch C, Urbassek HM. Langmuir. 2011;27:12938. doi: 10.1021/la201972f. [DOI] [PubMed] [Google Scholar]
- 51. [May 2, 2012];The charge density differences between BSA and HSP90 were inferred from their ‘grand mean of hydropathicity’ (GRAVY) scores of -0.433 and -0.750 respectively, calculated using the PROTPARAM tool. http://us.expasy.org/tools/protparam.html.
- 52.Karenga S, El Rassi Z. J. Sep. Sci. 2008;31:2677. doi: 10.1002/jssc.200800310. [DOI] [PubMed] [Google Scholar]
- 53.Wetzl BK, Yarmoluk SM, Craig DB, Wolfbeis OS. Angew. Chem., Int. Ed. 2004;43:5400. doi: 10.1002/anie.200460508. [DOI] [PubMed] [Google Scholar]
- 54.Ramsay LM, Dickerson JA, Dovichi NJ. Electrophoresis. 2009;30:297. doi: 10.1002/elps.200800498. [DOI] [PubMed] [Google Scholar]
- 55.Jungbauer LM, Yu C, Laxton KJ, LaDu MJ. J. Mol. Recognit. 2009;22:403. doi: 10.1002/jmr.948. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Nichkova M, Dosev D, Gee SJ, Hammock BD, Kennedy IM. Anal. Biochem. 2007;369:34. doi: 10.1016/j.ab.2007.06.035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Liu K, Tolley HD, Lee ML. J. Chromatogr. A. 2012;1227:96. doi: 10.1016/j.chroma.2011.12.081. [DOI] [PubMed] [Google Scholar]
- 58.Li Y, Tolley HD, Lee ML. J. Chromatogr. A. 2011;1218:1399. doi: 10.1016/j.chroma.2011.01.028. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.