Abstract
Summary: We describe a tool for quantifying the uniformity of mapped reads in high-throughput sequencing experiments. Our statistic directly measures the uniformity of both read position and fragment length, and we explain how to compute a P-value that can be used to quantify biases arising from experimental protocols and mapping procedures. Our method is useful for comparing different protocols in experiments such as RNA-Seq.
Availability and implementation: We provide a freely available and open source python script that can be used to analyze raw read data or reads mapped to transcripts in BAM format at http://www.math.miami.edu/~vhower/ReadSpy.html
Contact: lpachter@math.berkeley.edu
Supplementary Information: Supplementary data are available at Bioinformatics online.
1 INTRODUCTION
In biological experiments, controlling the quality of data is fundamental to the reliability and reproducibility of the results. Quality control is especially important in modern sequencing experiments, where small biases in the preparation of DNA libraries can be amplified in the subsequent sequencing steps which typically yield millions of reads. Sequenced reads ideally represent fragments sampled uniformly at random from a library, and expected coverage statistics can be obtained from the classic Lander–Waterman model Lander and Waterman (1988). However, a key aspect of current experiments is variable fragment length, requiring the extension of the Lander–Waterman model to account for random fragment position as well as length. Such a generalization was described in Evans et al. (2010) where it was shown that the data from a sequencing experiment can be modeled by a two-dimensional spatial Poisson process. We present a statistical test for uniformity in the reads of a sequencing experiment based on this idea and show how it can be used to compare multiple sets of reads in order to assess different protocols. Our test requires the alignment of paired-end reads to a reference transcriptome so that fragment positions and lengths can be determined. Fortunately, such alignments are routinely produced after sequencing experiments, so that our test can easily be incorporated into sequencing analysis pipelines.
2 METHODS
An aligned read can be represented as an integer point in  as follows: The ‘t-coordinate’ corresponding to the read is its left-end point while the ‘l-coordinate’ is the length of the fragment. In Evans et al. (2010), it is shown that for any choice of fragment length distribution, the collection of points {(t, l)} from a sequencing experiment forms a two-dimensional Poisson process. This principle guides our further analysis of these points {(t, l)}, as we test for uniformity in both the t and l coordinates. The output of ReadSpy is a list of test statistics and P-values for each transcript. A statistically significant (low) P-value means we reject the fact that the dataset is uniform on that transcript. Thus, a higher P-value corresponds to a set of reads sampled uniformly, which is desired. In the next two sections, we describe the statistical test applied a each transcript. The test is formulated in terms of the genomic segment [a, b].
as follows: The ‘t-coordinate’ corresponding to the read is its left-end point while the ‘l-coordinate’ is the length of the fragment. In Evans et al. (2010), it is shown that for any choice of fragment length distribution, the collection of points {(t, l)} from a sequencing experiment forms a two-dimensional Poisson process. This principle guides our further analysis of these points {(t, l)}, as we test for uniformity in both the t and l coordinates. The output of ReadSpy is a list of test statistics and P-values for each transcript. A statistically significant (low) P-value means we reject the fact that the dataset is uniform on that transcript. Thus, a higher P-value corresponds to a set of reads sampled uniformly, which is desired. In the next two sections, we describe the statistical test applied a each transcript. The test is formulated in terms of the genomic segment [a, b].
We handle ambiguous reads in ReadSpy by first reporting all possible alignments, using eXpress Roberts and Pachter (Submitted for publication) (http://bio.math.berkeley.edu/eXpress/) to assign a probability to each alignment and then selecting one t-value per fragment according to the observed probabilities.
2.1 Hypothesis test for dataset
First, we describe the statistical test for a set of aligned reads from one sequencing experiment. The (t, l) coordinates corresponding to reads aligned to [a, b] satisfy
We define shifted coordinates in the (x, y)-plane as follows:
The new point set S of transformed datapoints satisfies  and is still homogeneous along the x-axis.
and is still homogeneous along the x-axis.
We use a simple χ2 statistic to test this partial homogeneity assumption using test constants C and D. First, we partition the points into horizontal strips
where  and the boundaries
and the boundaries  are chosen such that each
are chosen such that each  . Next, we ignore the y-coordinates within each strip and test the uniformity of the x coordinates. We divide the interval [0, 1] into D subintervals and define
. Next, we ignore the y-coordinates within each strip and test the uniformity of the x coordinates. We divide the interval [0, 1] into D subintervals and define
If the original point set forms a spatial Poisson process, then
 |
where  . Since all
. Since all  s come from mutually independent strips, we have
s come from mutually independent strips, we have
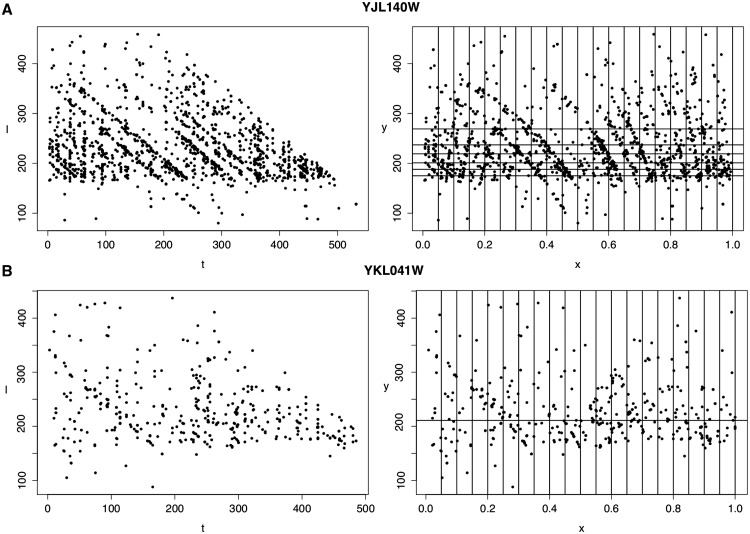
This allows us to calculate a P-value for the collection of aligned reads. This value may vary depending on the choice of constants C and D—a reasonable choice seems to be C = 200 and D = 20. Figure 1 shows the aligned reads for two yeast genes both in the (t, l)-plane and the (x, y)-plane. We also depict the vertical and horizontal grids for C = 200 and D = 20. The raw reads come from an RNA-Seq experiment using the dUTP protocol Levin et al. (2010) and were aligned to the yeast genome using Bowtie Langmead et al. (2009) using the ‘-X 600 -aS -n 3 -e 100’ option.
Fig. 1.
An illustration of the method. Fragments aligning to the Yeast genes YJL140W (A) and YKL041W (B) using dUTP protocol for RNA-Seq are depicted before (t, l-coordinates) and after (x, y-coordinates) our transformation. The horizontal boundaries in the (x, y)-plane are selected by requiring at least C = 200 points fall in each subdivision while the D = 20 vertical subdivisions are equally spaced. When comparing the these dUTP data sets to those from other RNA-Seq protocols, the point set in A is highly biased with a p-value of 1.2e−19 while B appears to be a random collection of points (p = 0.27)
2.2 Comparing multiple datasets
Assume we want to compare multiple sets of reads aligned to the same genomic interval [a, b]. Since the P-value of the above test depends on how the plane is divided into strips, the boundaries  are chosen to be the same across all the transformed point sets
are chosen to be the same across all the transformed point sets  . We impose the requirement that for each point set
. We impose the requirement that for each point set  , the horizontal strip
, the horizontal strip  contains at least C points. The resulting
contains at least C points. The resulting  s are then subsampled to produce
s are then subsampled to produce  such that for all i, we have
such that for all i, we have
We then apply our statistical test to each point set  using the horizontal strips
using the horizontal strips  with vertical subdivisions given by the constant D as above.
with vertical subdivisions given by the constant D as above.
3 RESULTS
We applied our statistical test to examine a variety of popular RNA-Seq protocols. Using data analyzed in Levin et al. (2010), we compare seven paired-end methods on 1464 transcripts in the yeast genome. Supplementary Table S1 gives transcript-by-transcript summary statistics for the log probabilities. There are multiple criteria for selecting an RNA-Seq protocal, but using ReadSpy we find that the dUTP protocol is statistically most random—meaning least biased—which agrees with Levin et al. (2010). Additionally, we make pairwise comparisons between the methods, which can be found in Supplementary Table S2. For each pair, the binomial test addresses statistically whether or not one method ‘wins’ on more often than the other. The P-values for this test are also listed in Supplementary Table S2. Our findings mostly agree with the results of Levin et al. (2010); however, our method illustrates the variability in sequence uniformity among genes. For instance, Figure 1 gives the point sets for dUTP protocol and two yeast genes: YDR007W (in A) and YJL078C (in B). When comparing all seven methods, dUTP achieved  for YJL140W and p = .27 for YKL041W. A low P-value indicates that the reads aligning to the transcript are not uniform, which is undesirable. One can visually see the difference in Figure 1 between non-random (in A) and random (in B).
for YJL140W and p = .27 for YKL041W. A low P-value indicates that the reads aligning to the transcript are not uniform, which is undesirable. One can visually see the difference in Figure 1 between non-random (in A) and random (in B).
Funding: V.H. was funded in part by NSF fellowship DMS-0902723. A.R and L.P. were funded in part by NIH R01 HG006129. A.R. was also funded in part by an NSF graduate research fellowship.
Conflict of Interest: none declared.
Supplementary Material
REFERENCES
- Evans S, et al. Coverage statistics for sequence census methods. BMC Bioinformatics. 2010;11:430. doi: 10.1186/1471-2105-11-430. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lander E, Waterman M. Genomic mapping by fingerprinting random clones: a mathematical analysis. Genomics. 1988;2:231–239. doi: 10.1016/0888-7543(88)90007-9. [DOI] [PubMed] [Google Scholar]
- Langmead B, et al. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 2009;10:R25. doi: 10.1186/gb-2009-10-3-r25. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Levin JZ, et al. Comprehensive comparative analysis of strand-specific RNA sequencing methods. Nat. Meth. 2010;7:709–715. doi: 10.1038/nmeth.1491. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.