Abstract
Changes in adipose tissue metabolism are central to adaptation of whole body energy homeostasis to pregnancy. To gain insight into the molecular mechanisms supporting tissue remodeling, we have characterized the longitudinal changes of the adipose transcriptome in human pregnancy. Healthy nonobese women recruited pregravid were followed in early (8–12 wk) and in late (36–38 wk) pregnancy. Adipose tissue biopsies were obtained in the fasting state from the gluteal depot. The adipose transcriptome was examined via whole genome DNA microarray. Expression of immune-related genes and extracellular matrix components was measured using real-time RT-PCR. Adipose mass, adipocyte size, and cell number increased in late pregnancy compared with pregravid measurements (P < 0.001) but remained unchanged in early pregnancy. The adipose transcriptome evolved during pregnancy with 10–15% of genes being differently expressed compared with pregravid. Functional gene cluster analysis revealed that the early molecular changes affected immune responses, angiogenesis, matrix remodeling, and lipid biosynthesis. Increased expression of macrophage markers (CD68, CD14, and the mannose-6 phosphate receptor) emphasized the recruitment of the immune network in both early and late pregnancy. The TLR4/NF-κB signaling pathway was enhanced specifically in relation to inflammatory adipokines and chemokines genes. We conclude that early recruitment of metabolic and immune molecular networks precedes the appearance of pregnancy-related physiological changes in adipose tissue. This biphasic pattern suggests that physiological inflammation is an early step preceding the development of insulin resistance, which peaks in late pregnancy.
Maternal body composition reflects homeostatic changes.
extensive modifications of maternal body composition take place during pregnancy. Total adipose mass and maternal blood volume increase significantly, whereas there is little change in maternal lean tissue mass (28, 32). The dynamic role of adipose tissue in the metabolic adaptations to pregnancy is highlighted by longitudinal changes in adipose tissue mass. There is a wide individual variation in fat mass accumulation that can range from −2 to >10 kg/woman depending on race, ethnicity, and nutritional and metabolic factors (30, 40). In lean women, the total gain in fat mass represents about 30% of gestational weight gain (22), but women with pre-ravid obesity usually gain less total weight than lean women (18). In uncomplicated pregnancies with appropriate weight gain, most fat deposition occurs during the second trimester and remains constant until term (22, 54). The mechanisms underlying the early adjustments for increased fat deposition have yet to be defined in either normal or metabolically compromised pregnancies.
Storage and endocrine function.
The changes in body composition during pregnancy are driven primarily by adaptations of maternal metabolic homeostasis. The ultimate goal of pregnancy-induced metabolic changes is to meet the high-energy demands of fetal development. Glucose, the primary energy fuel used by fetal tissues, needs to be readily available for transplacental transfer, whereas maternal tissues can rely on other energy substrates, such as lipids (3). The adaptations of lipid metabolism follow a well-described biphasic pattern. The first half of pregnancy is centered on storing maternal energy as adipose tissue triglycerides, whereas in late pregnancy the stored lipids are mobilized to be used by peripheral tissues and in preparation for lactation (25). These sequential adaptations are facilitated by modifications of insulin secretion and action. In early pregnancy, the higher insulin sensitivity facilitates cellular anabolism through activation of lipogenesis. Then, insulin resistance develops progressively to culminate in the third trimester (7, 43). Insulin resistance allows adipose tissue to mobilize the lipids stored earlier and skeletal muscle to utilize less glucose. These changes in maternal metabolic homeostasis result in increased circulating levels of insulin and triglycerides in late pregnancy (23). In addition to alterations in its capacity for energy storage, the endocrine function of adipose tissue also evolves during pregnancy. The synthesis and plasma concentration of leptin and adiponectin, two major adipokines, exhibit longitudinal changes parallel to those of insulin sensitivity. Whereas leptin concentrations increase early, there is a significant decrease in plasma adiponectin during the third trimester (9, 26). The longitudinal modifications of adipokines in healthy pregnancy are further enhanced in the context of pregnancy with diabetes and obesity (10, 24).
Remodeling.
White adipose tissue displays a remarkable flexibility, with an important and reversible capacity for expansion throughout adult life. Remodeling of the adipose tissue mass requires the integration of cellular mechanisms to support an increase in size and/or number of adipocytes. The turnover of fat cells is highly sensitive to variations in the metabolic and hormonal milieu, i.e., development, aging, diet, or metabolic disorders occurring throughout the lifespan (16, 33, 52). In obese individuals, the enlargement of adipose tissue requires active turnover of adipocytes with chronic inflammation elicited by accumulation of macrophages in the stromal vascular compartment (21, 60). Previously, we have shown that macrophage infiltration and increased adipocyte death contribute to inflammation in pregnant women with pregravid obesity (2, 12).
The aim of this study was to characterize the mechanisms responsible for the increase in adipose tissue mass during normal human pregnancy. We have conducted a longitudinal analysis beginning before and continuing throughout pregnancy to characterize the morphological and molecular traits of the adipose tissue in nonobese women. We show that molecular changes supporting remodeling of gluteal adipose tissue precede phenotypic changes in fat accretion and lipid metabolism that culminate in late pregnancy.
METHODS
Subjects.
Eleven women with singleton pregnancies were recruited prepregnancy (P) and followed up longitudinally in both early (E; 8–12 wk) and late pregnancy (L; 36–38 wk). The protocol was approved by the Institutional Review Board and Clinical Research Unit Scientific Review Committee at Case Western Reserve University. Volunteers gave their written informed consent, in accordance with the MetroHealth Medical Center (Cleveland, OH) Guidelines for the Protection of Human Subjects.
Metabolic and anthropometric measurements.
Anthropometric measurements and body composition were estimated using hydrodensitometry, as described previously (8, 27). Insulin sensitivity was measured with the euglycemic hyperinsulinemic clamp technique (7). Plasma was separated by centrifugation and kept frozen at −20°C until being assayed. Glucose was assessed by the glucose oxidase method (YSI, Yellow Springs, OH). Plasma insulin was measured using the sensitive human insulin radioimmunoassay kit (Linco Research, St. Charles, MO). The limit of sensitivity for the insulin assay is 0.2 μU/ml, with intra-assay coefficients of variation (CV) of 4.4–6.8%. Leptin and adiponectin were measured using ELISA kits (R & D Systems, Minneapolis, MN) with CVs of 3.0–6.2 and 6.2–8.4%, respectively. Interleukin-6 and interleukin-8 were assayed by ELISA (QuantiGlo; R & D Systems) with CVs of 5.3–7.8 and 2.6–3.4%, respectively. All plasma samples were run in duplicate.
Adipose tissue analysis: measurements of cell number and size.
Biopsies of adipose tissue (800–1,200 mg) from 11 women were obtained via liposuction of the subcutaneous gluteal depot at each time point (P, E, and L). One fragment of tissue was fixed immediately for immunohistochemistry, one fragment was snap-frozen in liquid nitrogen for RNA processing, and the remaining tissue was processed for collagenase digestion. After digestion with 1 mg/ml collagenase (Worthington Biochemical, Lakewood, NJ) for 45 min at 37°C, the mixture was filtered through a 100-μm gauze mesh. The infranatant was removed, and the floating layer of adipocytes was brought up to 10 ml for counting and size measurements. Adipocyte diameter was determined using a ×10 W. F. objective lens containing a ruler (Olympus BH-2 microscope; Olympus, Hiroshima, Japan). Numbers from 10 fields were averaged, and cellularity was defined as the number of adipocytes per gram of digested adipose tissue (44).
Immunohistochemistry.
Adipose tissue sections were fixed in 10% formalin and included in paraffin. Five-micrometer sections were counterstained using eosin and hematoxylin gill II (Sigma-Aldrich). Digitized images were obtained using a Nikon E600 microscope equipped with a DXM200 camera.
Protein expression.
Lysates of whole adipose tissue were prepared using 10 mM Tris-NaCl, pH 8, and lysis buffer with 1% Triton and protease inhibitors (Sigma-Aldrich). Protein lysates (2–200 μg/ml) were separated by SDS-PAGE (7% polyacrylamide gels) and immunoblotted or directly blotted to nitrocellulose for microfiltration (Hoefer). HL-60 whole cell lysate (Santa Cruz, CA) was used as positive control. Primary antibodies against GAPDH (1:3,000), glucose transporter 5 (1:500), lipoprotein lipase (LPL; 1:500), and adiponectin (1:500) were from Abcam. Actin (1:2,000), CCAAT/enhancer-binding protein (C/EBP) (1:500), and Toll-like receptor 4 (TLR4) (1:300) were from Santa Cruz Biotechnology; lipolysaccharide-binding protein (1:500) was from Origene Technologies. Detection was performed using anti-mouse or anti-rabbit HRP-coupled secondary antibodies (1:5,000) and ECL Plus Western blot chemiluminescence reagents (Amersham, GE Heathcare Sciences). Signal intensity was quantitated by gel doc imager (Bio-Rad).
Gene expression analysis.
Total RNA was isolated from intact adipose tissue using the Trizol (Invitrogen, Carlsbad, CA) extraction method. Gene expression was analyzed via whole genome microarray profiling, using U133 Affymetrix arrays and platforms as described (41). Selection of the significantly modified transcripts was performed by a multistep filter strategy. Among the genes that showed an absolute call of present according to MAS 5.0 algorithm, we selected the genes with a difference in signal detection of ≥4.5 times the average background minus the scaled noise. The hybridization intensities of significantly modified genes were examined by hierarchic cluster analysis with Gene-Spring GX (Agilent Technologies, Santa Clara, CA) and Treeview software (free). The genes that satisfied these criteria were then selected on the basis of a fold change >1.52 or <1.52 that was consistent in at least two comparisons. Genes that related to energy metabolism were identified according to the function of their putative encoded proteins from public databases. (IPA; Ingenuity Systems, Redwood City, CA). Changes in gene expression were validated by real-time PCR (Roche Thermocycler; Roche Applied Science, Indianapolis, IN) with LightCycler FastStart DNA SYBR Green 1 master mix and primers from Integrated DNA Technologies (Coralville, IA). Primers for specific target sequences were designed within the 3′ coding region of the genes.
Statistics.
All values are presented as means ± SE. Differences between dependent variables were examined with one-way or two-way repeated-measures analysis of variance. Significant mean differences among the three time points were identified with Fisher's protected least significant difference post hoc test. The data were analyzed using the StatView II statistical package (Abacus Concepts, Berkeley, CA). Statistical significance was set at P < 0.05.
RESULTS
Longitudinal phenotypic and metabolic changes.
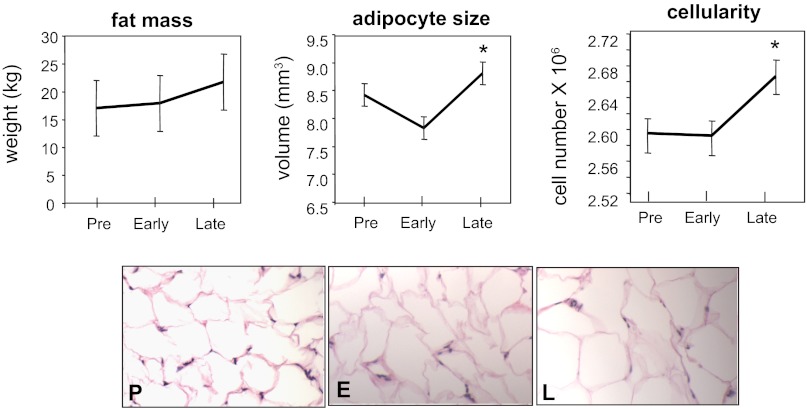
Longitudinal measurements were performed prepregnancy (P) and in both early (8–12 wk) (E) and late pregnancy (36–38 wk) (L). Metabolic and anthropometric parameters are listed in Table 1. Both plasma cholesterol and triglyceride concentrations were increased in L, whereas plasma free fatty acids were decreased compared with pregravid measurements. Insulin sensitivity decreased significantly in L compared with both E and P. Leptin concentrations increased significantly in E and remained elevated until L. Measurements of body composition showed that total body fat mass was increased in late gestation (P < 0.001; Fig. 1). An increase in the size and number of adipocytes was observed in L compared with pregravid but was not detected at early stages (P < 0.001; Fig. 1).
Table 1.
Anthropometric and metabolic characteristics of women before and during pregnancy
| Prepregnancy | Early Pregnancy | Late Pregnancy | |
|---|---|---|---|
| Weight kg | 70.2 ± 5.6 | 72.1 ± 5.2 | 84.3 ± 4.8* |
| BMI, kg/m2 | 24.0 ± 2.0 | 24.7 ± 1.8 | 28.9 ± 1.7* |
| Total body fat mass, kg | 19.9 ± 4.5 | 20.4 ± 4.1 | 25.9 ± 3.8* |
| Lean body mass, kg | 50.3 ± 1.8 | 51.6 ± 1.5 | 58.3 ± 2.0* |
| Triglycerides, mg/dl | 58 ± 10 | 63 ± 5 | 195 ± 55** |
| Cholesterol, mg/dl | 156 ± 11 | 154 ± 18 | 240 ± 18* |
| FFA basal, μmol/ml | 740 ± 21 | 610 ± 13 | 560 ± 12* |
| FFA clamp, μmol/ml | 140 ± 19 | 120 ± 40 | 202 ± 39* |
| Glucose, mg/dl | 90 ± 2 | 82 ± 3 | 79 ± 3* |
| Insulin, μU/ml | 12 ± 1.2 | 10 ± 1.8 | 16.2 ± 2.8 |
| ISI, mg·zmd·min−1·zmd·kg−1 | 8.6 ± 0.9 | 8.0 ± 0.8 | 5.2 ± 0.7* |
| Leptin, ng/ml | 12.6 ± 4.8 | 22.0 ± 8.1 | 26.5 ± 8.3* |
Results are means ± SE of n = 6 subjects whose adipose tissue was analyzed for gene expression.
BMI, body mass index; FFA, free fatty acids; ISI, insulin sensitivity index.
P < 0.001 and
P < 0.01, statistical significance compared with prepregnancy.
Fig. 1.
Morphological changes in adipose tissue in human pregnancy. Top: total adipose tissue mass, adipocyte size, and number of adipocytes prior to and during healthy uncomplicated pregnancy. Early: 8–11 wk of gestation; late: 34–36 wk of gestation. Statistical significance: *P < 0.05 vs. pregravid. Results are means ± SD for 11 women. Bottom: immunohistochemistry of subcutaneous gluteal adipose tissue sections. Initial magnification, ×20. P, pregravid; E, early pregnancy (8–12 wk); L, late pregnancy (34–36 wk).
Molecular characteristics of adipose tissue.
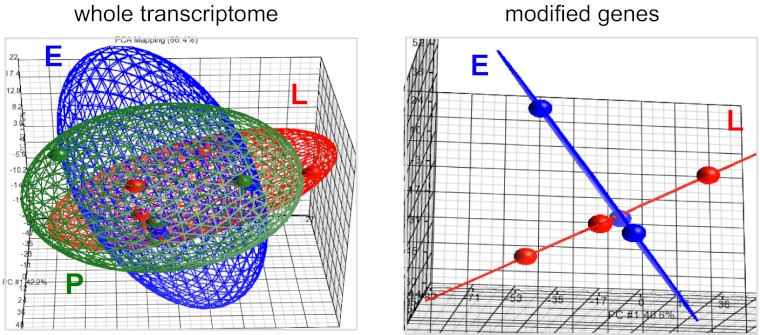
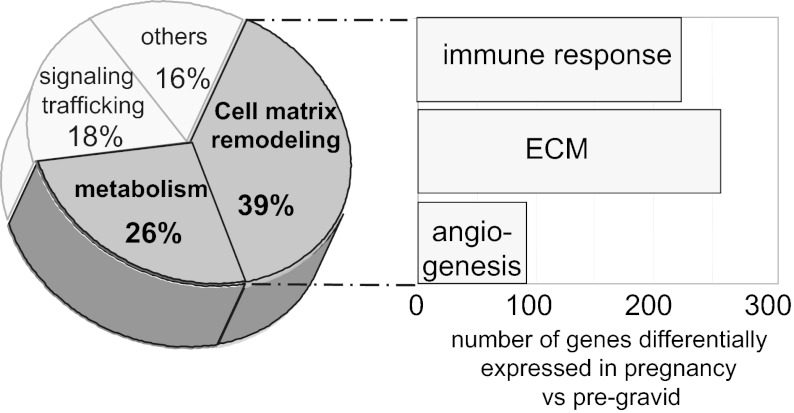
Using whole genome microarray analysis, the adipose tissue transcriptome, encompassing 12,897 ± 417 genes, remained quantitatively stable during pregnancy, with 13,510 ± 197 genes expressed during E and 13,097 ± 298 genes expressed during L. Principal component analysis revealed a large overlap in expression pattern between the pregravid, E, and L stages (Fig. 2). The genes differentially expressed in E (n = 1,286) and L (n = 1,111) represented 10–15% of the adipose tissue transcriptome. Comparison of global gene expression also revealed a distinct expression pattern in E and L (Fig. 2). Analysis of differentially modified genes based on their biological function indicated that 65% of pregnancy-associated changes were related to tissue remodeling (39%) and lipid metabolism (26%) (Tables 2 and 3 and Fig. 3). Genes involved in tissue remodeling were further separated into three main functional categories: mediators of the immune response, extracellular matrix components, and angiogenic factors (Fig. 3). Marked changes in genes regulating pathways for the inflammatory response and metabolism were noted in E compared with P. Innate immune responses elicited through the TLR4/NF-κB signaling pathway led to early increases in gene and protein expression of TLR4 as well as TNFα, IL-6, and small chemokines of the C-X-CL family (Fig. 4, A and B). Metabolic genes facilitating lipid accumulation through adipocyte differentiation and lipogenesis were most activated in E (Fig. 5, A and B, and Table 4). High expression of transcription factors of the forkhead box O (FOXO), peroxisome proliferator-activated receptor-γ (PPARγ), cAMP response element-binding protein, and sterol regulatory element-binding protein families were indicative of adipocyte differentiation. Increased VLDL receptor and LDL-related protein 1, fatty acid synthase, acetyl-CoA carboxylase, and stearoyl-CoA desaturase catalyzing fatty acid uptake, synthesis, and elongation were evidence of active lipogenesis. Extracellular matrix components with structural (elastin, collagen) and adhesive properties (laminin, fibronectin) were activated predominantly in E and included upregulation of metalloproteinases to enhance tissue plasticity through the release of insulin-like growth factor I (IGF-I). The concomitant activation of several angiogenic factors [VEGF, TGFβ, angiotensin, fibroblast growth factor receptor 2 (FGFR2)] suggested that neovascularization was associated with matrix remodeling (Table 2).
Fig. 2.
Three-dimensional scatter plot principal component analysis (PCA) of the adipose tissue transcriptome. Gluteal adipose tissue obtained longitudinally in P, E, and L from 6 of the 11 women. Full cohort was processed for microarray analysis. A: analysis of full data set (n = 22, 277 genes) showed overlap in transcriptome profiles before and during pregnancy. B: analysis of the genes modified significantly in E and L (n = 1,807 genes) compared with pregravid showed a distinctive pattern at each stage of pregnancy. Each knot represents 1 microarray data set.
Table 2.
Genes differentially modified in adipose tissue in early pregnancy
| NCBI Accession No. | Gene Name | Fold Change | Description |
|---|---|---|---|
| Lipid metabolism | |||
| NM_006415 | SPTLC1 | 1.5 | Serine palmitoyltransferase long chain |
| NM_004863 | SPTLC2 | 1.6 | Serine palmitoyltransferase long chain |
| NM_030791 | SGPP1 | 17.5 | Sphingosine-1 phosphate phosphatase |
| D85181 | SC5DL | 1.7 | Sterol-C5-desaturase |
| AF116616 | SCD | 1.5 | Stearoyl-CoA desaturase |
| S69189 | ACOX1 | −1.6 | Acyl coenzyme A oxidase |
| NM_013402 | FADS1 | 1.7 | Fatty acid desaturase |
| NM_001995 | ACSL1 | 1.5 | Acyl-CoA synthetase long-chain |
| NM_004104 | FASN | 1.7 | Fatty acid synthase |
| NM_198834 | ACACA | 1.5 | Acetyl-CoA carboxylase-α |
| NM_023928 | AACS | 1.5 | Acetoacetyl CoA synthetase |
| NM_024090 | ELOVL6 | 6.2 | Elongation of long chain fatty acids 6 |
| NM_022821 | ELOVL1 | 1.6 | Elongation of very long chain fatty acids 1 |
| U18197 | ACLY | 1.6 | ATP cytrate lyase |
| NM_002317 | LOX | 2.0 | Lysyl oxidase |
| NM_002979 | SCP2 | 1.5 | Sterol carrier protein |
| NM_006227 | PLTP | 1.5 | Phospholipid transfer protein |
| NM_000237 | LPL | 1.7 | Lipoprotein lipase |
| NM_015869 | PPARG | 1.6 | Peroxisome proliferator-activated receptor |
| NM_001455 | FOXO3B | 2.1 | forkhead box O3 |
| NM_002015 | FOXO1 | 1.5 | forkhead box O1 |
| NM_003355 | UCP2 | 2.5 | Uncoupling protein 2 |
| NM_000098 | CPT2 | −1.5 | Carnitine palmitoyltransferase 2 |
| NM_000017 | ACADS | −1.5 | Acyl-coenzyme A dehydrogenase short chain |
| NM_001608 | ACADL | −1.7 | Acyl-CoA dehydrogenase long chain |
| NM_005327 | HADHA | −1.5 | Hydroxyacyl-CoA dehydrogenase |
| NM_000041 | APOE | −1.7 | Apolipoprotein E |
| NM_207630 | CREM2 | 1.5 | cAMP-responsive element modulator |
| NM_204203 | C/EBP | 1.8 | CCAAT/enhancer binding protein-γ |
| NM_201988 | CREB | 1.7 | cAMP responsive element binding protein |
| NM_221147 | WWOX | 12.1 | WW domain containing oxidoreductase |
| Growth factors and matrix remodeling | |||
| NM_003376 | VEGFA | 1.7 | Vascular endothelial growth factor A |
| NM_001078 | VCAM1 | 1.8 | Vascular cell adhesion molecule |
| NM_000612 | INS-IGF2 | 4.6 | Insulin-like growth factor 2 |
| M37484 | IGF1 | 2.0 | Insulin like growth factor 1 |
| NM_022970 | FGFR2 | 1.5 | Fibroblast growth factor receptor 2 |
| NM_012098 | ANGPTL2 | 1.6 | Angiopoietin like 2 |
| AB000889 | PPAP2B | 1.9 | Phosphatidic acid phosphatase 2B |
| NM_004995 | MMP-14 | 2.0 | Matrix metalloproteinase 14 |
| NM_001846 | COL4A2 | 3.2 | Collagen type IV α2 |
| NM_003873 | NRP1 | 1.6 | Neuropilin 1 |
| NM_001069 | TUBB2B/2A | 2.8 | Tubulin-β 2A and 2B |
| NM_000224 | KER18 | 3.5 | Keratin 18 |
| NM_002964 | S100A8 | 1.5 | S100 calcium binding protein A8 |
| AF019888 | ARPC4 | 3.2 | Arp complex 20 kDa subunit |
| Immune responses | |||
| NM_002184 | IL6ST | 3.4 | Interleukin-6 signal transducer |
| NM_000878 | IL2RB | 1.9 | Interleukin-2 receptor-β |
| NM_004515 | ILF2 | 1.5 | Interleukin enhancer-binding factor |
| M97935 | STAT1 | 2.5 | Signal transducer and activator of transcription |
| NM_002982 | CCL2 | 2.2 | Chemokine (C-C motif) ligand |
| NM_002089 | CXCL2 | 2.0 | Chemokine (C-X-C motif) ligand |
| NM_005409 | CXCL11 | 2.4 | Chemokine (C-X-C motif) ligand |
| NM_002416 | CXCL9 | 1.6 | Chemokine(C-X-C motif) ligand |
| NM_005408 | CCL13 | 1.5 | Chemokine (C-C motif) ligand |
| NM_001565 | CXCL10 | 1.5 | Chemokine C-X-C motif ligand |
| NM_014452 | TNFRSF21 | 2.0 | Tumor necrosis factor receptor |
| NM_002546 | TNFRSF11B | 1.7 | Tumor necrosis factor receptor |
| NM_000416 | IFNGR1 | 1.5 | Interferon-γ receptor |
| NR_024168 | TLR4 | 1.5 | Toll-like receptor 4 |
| NM_000591 | CD14 | 1.5 | CD14 molecule |
| NM_004139 | LBP | 5.2 | Lipopolysaccharide-binding protein |
| NM_001556 | IKBKB | 1.8 | Inhibitor k light polypeptide enhancer |
| NM_003998 | NFKB1 | 1.5 | Nuclear factor k light polypeptide enhancer |
| NM_014294 | TRAM1 | 1.7 | Translocation associated membrane protein 1 |
| BC003388 | TANK | 1.6 | TRAF family member associated NFKB activator |
| NM_020056 | HLA-DQA1 | 1.6 | Major histocompatibility complex |
| NM_006512 | SAA4 | 1.5 | Serum amyloid A4 |
| NM_030754 | SAA1 | 1.6 | Serum amyloid A1 |
| NM_208703 | SAAP | 3.7 | Amyloid β (A4) precursor |
| NM_001212 | C1QBP | 1.5 | Complement component 1q binding protein |
NCBI, National Center for Biotechnology Information. Changes are expressed in fold change compared with pregravid. GSE37215 approval (NCBI tracking system no. 16555206).
Table 3.
Genes differentially modified in adipose tissue in late pregnancy
| NCBI Accession No, | Gene Name | Fold Change | Description |
|---|---|---|---|
| Immune-related genes | |||
| NM_004139 | 2 | LBP | Lipopolysaccharide-binding protein |
| NM_138554 | 1.6 | TLR4 | Toll like receptor 4 |
| NM_003265 | 2.5 | TLR3 | Toll-like receptor 3 |
| NM_003326 | 2.8 | TNFSF4 | Tumor necrosis factor ligand |
| NM_014452 | 1.9 | TNFRSF21 | Tumor necrosis factor receptor 21 |
| NM_014452 | 1.7 | TNFRSF20 | Tumor necrosis factor receptor 20 |
| NM_002389 | 1.6 | CD46 | CD46 molecule complement regulatory protein |
| NM_000331 | 1.6 | SAA1 | Serum amyloid A 1 |
| NM_006512 | 3.7 | SAA4 | Serum amyloid A4 |
| BC020795 | 2.3 | SAA2 | Serum amyloid A2 |
| NR_026576 | 9.1 | SAA3P | Serum amyloid A3 pseudogene |
| NM_004887 | 1.6 | CXCL14 | Chemokine (C-X-C motif) ligand 14 |
| NM_003150 | 1.5 | STAT3 | Signal transducer and activator of transcription |
| NM_002163 | 2.3 | IRF8 | Interferon regulatory factor 8 |
| NM_001161529 | 2.3 | CSF2RA | Colony-stimulating factor 2 receptor-α |
| NM_002852 | 2 | PTX3 | Pentraxin-related gene |
| NM_018643 | 2 | TREM1 | Triggering receptor expressed on myeolid cells |
| NM_001040059 | 1.9 | CD68 | CD68 molecule |
| NM_000757 | 1.9 | CSF1 | Colony-stimulating factor 1 |
| NM_001161529 | 1.9 | CSF2RA | Colony-stimulating factor 2, receptor A |
| NM_000395 | 1.5 | CSF2RB | Colony-stimulating factor 2, receptor B |
| AY312956 | 2.5 | TRADV3 | T cell receptor-δ chain |
| NM_005384 | 1.7 | NFIL3 | Nuclear factor interleukin-3 |
| NM_000585 | 1.5 | IL15 | Interleukin-15 |
| NM_001190981 | 1.9 | IL6ST | Interleukin-6 signal transducer |
| NM_000877 | 2.5 | IL1R1 | Interleukin-1 receptor type 1 |
| NM_001560 | 3.1 | IL13RA1 | Interleukin 13 receptor α1 |
| NM_030968 | 1.6 | C1QTNF1 | C1q and tumor necrosis factor related protein 1 |
| NM_016546 | 1.9 | C1RL | Complement component 1 receptor like |
| NM_001115131 | 3.2 | C6 | Complement component 6 |
| NM_002982 | 2.5 | CCL2 | Chemokine (C-C motif) ligand 2 |
| NM_001168298 | 1.7 | CXCR2 | Chemokine (C-X-C motif) receptor 2 |
| NM_002089 | 2.3 | CXCL2 | Chemokine (C-X-C motif) ligand 2 |
| NM_006271 | 2.8 | S100A1 | S100 calcium-binding protein A1 |
| NM_002964 | 3.5 | S100A8 | S100 calcium-binding protein A8 |
| NM_002965 | 3.5 | S100A9 | S100 calcium-binding protein A9 |
| Glucose and lipid metabolism | |||
| NM_001135585 | 24.2 | SLC2A5 | Facilitated glucose/fructose transporter |
| NM_014668 | 6.9 | GREB1 | GREB1 protein |
| NM_013402 | 5.7 | FADS1 | Fatty acid desaturase 1 |
| NM_004265 | 4.9 | FADS2 | Fatty acid desaturase 2 |
| NM_004104 | 4.3 | FASN | Fatty acid Synthase |
| NM_000384 | 3.5 | APOB | Apolipoprotein B |
| NM_005063 | 3.05 | SCD | Stearoyl-CoA desaturase-δ9 |
| NM_025225 | 2.5 | PNPLA3 | Patatin-like phospholipase domain 3 |
| NM_020376 | 1.7 | PNPLA2 | Patatin like phospholipase domain 2 |
| NM_001161587 | 2.5 | GYS1 | Glycogen synthase 1 |
| NM_021957 | 4.6 | GYS2 | Glycogen synthase 2 |
| NM_001018056 | 2.5 | VLDLR | Very-low-density lipoprotein receptor |
| NM_001161504 | 2.3 | ALDH4A1 | Aldehyde dehydrogenase 4 family A1 |
| NM_005589 | 1.7 | ALDH6A1 | Aldehyde dehydrogenase 6 family A1 |
| NM_001608 | 2 | ACADL | Acyl-coenzyme A dehydrogenase long chain |
| NM_001012727 | 1.9 | AGPAT2 | Acylglycerol-3 phosphate acyltransferase 2 |
| NM_018441 | 1.9 | PECR | Peroxisomal trans 2 aneoyl CoA reductase |
| NM_000284 | 1.7 | PDHA1 | Pyruvate dehydrogenase α1 |
| NM_005164 | 1.6 | ABCD2 | ATP-binding cassette subfamily D 2 |
| NM_001184705 | 1.6 | HADH | Hydroxylacyl CoA dehydrogenase |
| NM_000925 | 1.5 | PDHB | Pyruvate dehydrogenase |
| NM_024420 | 1.5 | PLA2GA4 | Phospholipase A 2 group IV |
| Growth factors | |||
| NM_000141 | 3.0 | FGFR2 | Fibroblast growth factor receptor 2 |
| NM_000618 | 2 | IGF-I | Insulin-like growth factor I |
| NM_000599 | 1.5 | IGFBP5 | Insulin-like growth factor-binding protein 5 |
| NM_001552 | 1.6 | IGFBP4 | Insulin-like growth factor-binding protein 4 |
| NM_005544 | 2.5 | IRS1 | Insulin receptor substrate-1 |
| NM_005542 | 1.9 | INSIG1 | Insulin-induced gene 1 |
| NM_016133 | 2.3 | INSIG2 | Insulin-induced gene 2 |
Levels of expression are expressed in fold change compared with pregravid. GSE37215 approval (NCBI tracking system no. 16555206).
Fig. 3.
Transcriptional changes in adipose tissue during pregnancy. Functional analysis of the genes whose expression was modified during pregnancy compared with pregravid identified 4 main gene clusters based on biological annotation of their DNA sequences. ECM, extracellular matrix.
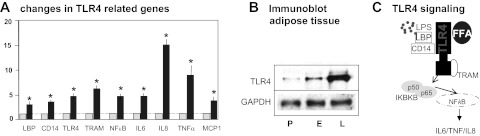
Fig. 4.
Pregnancy-related changes in adipose tissue Toll-like receptor 4 (TLR4) network. Genes implicated in TLR4 signaling pathways were selected from the lists in Table 2. A: mRNA levels were measured by quantitative RT-PCR analysis of adipose tissue from early pregnancy. Real-time threshold cycle (CT) values were normalized to actin and expressed in fold change vs. pregravid. B: immnublot longitudinal analysis of TLR4 protein content in adipose tissue at P, E, and L. C: diagram of the molecular pathways related to TLR4 activation identified with gene ontology analysis. Statistical significance: *P < 0.05. Gray bars, pregravid; black bars, early pregnancy; n = 6 independent determinations. LPS, lipopolysaccharide; LBP, lipopolysaccharide-binding protein; FFA, free fatty acid; TRAM, TRIF-related adaptor protein; IKBKB, inhibitor of κ-light polypeptide gene enhancer.
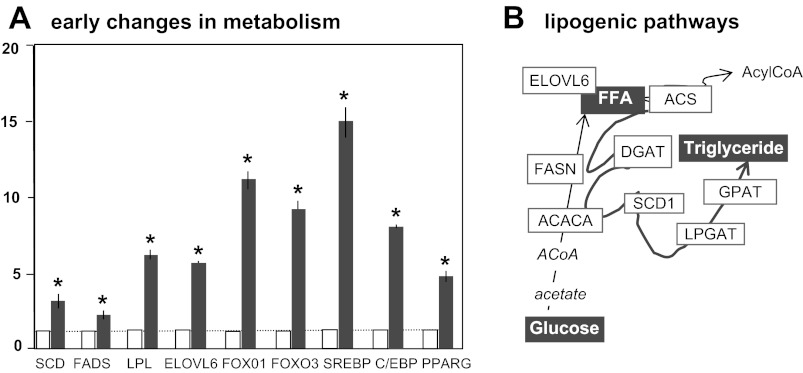
Fig. 5.
Pregnancy-related changes in adipose tissue metabolic pathways. mRNA levels of adipose-specific enzymes and transcription factors selected from Table 2 were measured by quantitative RT-PCR analysis. Real-time CT values were normalized to actin and expressed in fold change vs. pregravid. B: diagram of the molecular pathways identified with gene ontology analysis of differentially regulated genes in early pregnancy. Statistical significance: *P < 0.05. Open bars, pregravid; filled bars, early pregnancy for n = 6 independent determinations. ELOVL6, elongation of very long chain fatty acids protein 6; ACS, acyl-CoA synthetase; FASN, fatty acid synthase; DGAT, diacylglycerol acyltransferase; ACACA, acetyl-CoA carboxylase-1α; SCD, stearoyl-CoA desaturase; GPAT, glycerol phosphate acyltransferase; LPGAT, lisophosphatidylglycerol acyltransferase; ACoA, acetyl-CoA; FADS, fatty acid desaturase; LPL, lipoprotein lipase; FOXO, forkhead box O; SREBP, sterol regulatory element-binding protein; C/EBP, CCAAT/enhancer-binding protein; PPARγ, peroxisome proliferator-activated receptor-γ.
Table 4.
Adipose tissue protein expression quantitated by immunoblot
| Protein name | Prepregnancy | Early Pregnancy | Late Pregnancy |
|---|---|---|---|
| LBP | 1 | 1.9 ± 0.6* | 2.2 ± 0.6* |
| SLC2A5 (GLUT5) | 1 | 3.6 ± 1.1* | 5.4 ± 1.2* |
| Adiponectin | 1 | 1.8 ± 0.2* | 2.6 ± 0.6* |
| C/EBP | 1 | 2.2 ± 0.4* | 3.2 ± 0.7* |
| TLR4 | 1 | 2.9 ± 0.4* | 3.8 ± 1.1* |
| LPL | 1 | 1.5 ± 0.4ns | 3.3 ± 0.1* |
| GAPDH | 1 | 0.8 ± 0.1ns | 0.8 ± 0.1ns |
Results are means ± SD of n = 3 subjects; data are expressed as fold change over prepregnancy levels.
LBP, lipopolysaccharide-binding protein; SLC2A5, solute carrier family 2 (facilitated glucose transporter), member 5; GLUT5, glucose transporter 5; C/EBP, CCAAT/enhancer binding protein; LPL, lipoprotein lipase.
P = 0.05; nsnonsignificant statistical significance compared with prepregnancy.
DISCUSSION
Our study demonstrates an enlargement of adipose tissue during pregnancy, combining a higher cell number (hyperplasia) and a larger cell size (hypertrophy). This is in agreement with data for lower body depots in women (17, 50) but differs from the mechanisms described in obesity with hypertrophy occurring prior to hyperplasia. Hyperplasic growth appears to be genetically regulated, whereas hypertrophy is more nutritionally regulated (29). Growth is also dependent on each depot-unique gene signature, which defines the proliferation of adipocyte precursors (57). The longitudinal changes were characterized in the gluteal subcutaneous adipose depot. This depot is preferentially expanding in pregnancy (18), accessible to biopsy, and more prone to inflammatory changes than mesenteric or omental visceral depots (19). Further studies performed at visceral sites will help to achieve a global picture of adipose tissue changes.
Expansion and remodeling.
Our study reveals that adaptations of adipose tissue to healthy pregnancy develop in a temporal manner, with an array of molecular changes preceding the anthropometric expansion of adipose mass. Adipose tissue enlargement relies on molecular cross-talks between distinct cell types of the stromal-vascular fraction surrounding the adipocytes (45, 48). In agreement with this concept, our study suggests that pregnancy-induced adipose tissue expansion involves a combination of signaling pathways shared by preadipocytes and immune and endothelial cells (Fig. 6).
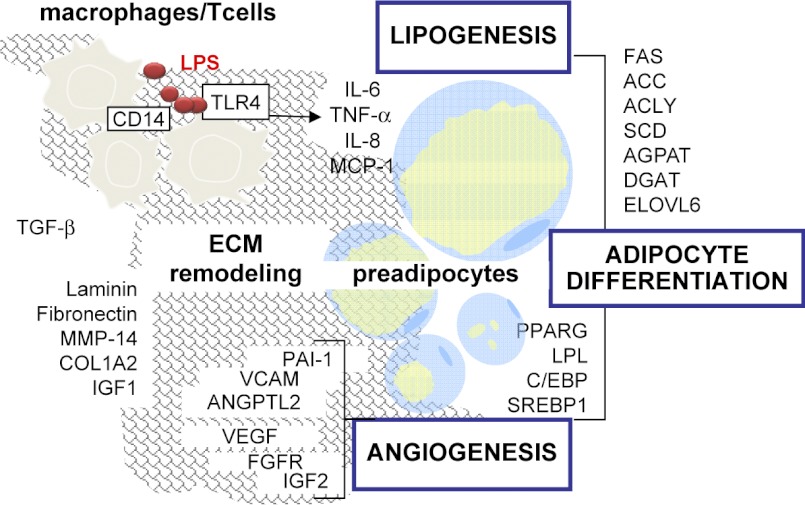
Fig. 6.
Model of functional networks contributing to adipose tissue remodeling in human pregnancy. Transcriptional activation patterns suggest that molecular networks from several adjacent cell types cooperate with the adipose tissue remodeling during pregnancy. ECM components and angiogenic factors are needed for vascular and adipocyte growth. Lipogenic genes and transcription factors are required for cell differentiation and lipid storage. Macrophages infiltrated between the stromal cells produce proinflammatory cytokines IL-6, IL-8, and TNF, which may 1) enhance neovascularization and 2) facilitate the development of insulin resistance. ACC, acetyl-CoA carboxylase; MCP-1, monocyte chemoattractant protein-1; ACLY, ATP citrate lyase; AGPAT, acylglycerol-3-phosphate acyltransferase; MMP-14, metalloproteinase-14; COL1A2, collagen type 1 α2; PAI-1, plasminogen activator inhibitor-1; VCAM, vascular cell adhesion molecule-1, ANGPTL2, angiopoietin-like 2; FGFR2, fibroblast growth factor receptor 2.
The flexibility of the cellular matrix surrounding the adipocytes plays a central role in the regulation of tissue expandability. Remodeling of extracellular matrix components is accelerated in a number of metabolic diseases that are associated with changes in the hormonal milieu (12, 48). Increases in metalloproteinases, fibronectin, and laminin accelerate preadipocyte differentiation in hypertrophic adipose tissue of obese individuals (20, 47). On the other hand, angiogenesis, which is also required for adipose tissue remodeling, may be facilitated by metalloproteinases and collagenases (11). VEGF-A and its most important functional partner, angiopoietin, are essential for initiation of the angiogenic program (59, 61). Hence, the combined increase in growth factors (IGF-II, FGFR2) and angiogenic factors (VEGF, TGFβ) is in line with the contribution of angiogenic mechanisms to the overall adipose tissue remodeling of early pregnancy (49).
The microenvironment surrounding the adipocyte may also be shaped by the production of adipokines that exert both paracrine and endocrine effects. TNFα and IL-6 exhibit increased expression in both the early and late stages of pregnancy, and therefore, they may regulate the expression of other genes that contribute to the negative action of insulin (21). The increase in leptin receptor expression is in agreement with a stimulatory role of leptin on angiogenesis, whereas the lack of leptin gene regulation is consistent with the rise in plasma leptin levels in pregnancy being contributed mainly by the placenta (34).
Adipocyte biology.
The 29% increase in fat mass measured in late pregnancy was associated with active cellular remodeling of adipose tissue. The activation of endothelial LPL suggested accelerated hydrolysis of circulating triglycerides into free fatty acids available for cellular uptake. Our findings bring support to a crucial role of LPL for increasing lipid storage in adipose tissue of pregnant rats (42). The upregulation of many other genes directly affecting lipogenesis and adipocyte differentiation indicated that those pathways are recruited early in pregnancy to enhance lipid storage in both mature and newly synthesized adipocytes. The forkhead transcription factor FOXO1 isoform, a master regulator of energy homeostasis in white adipose tissue, was one of the early induced genes. Activation of FOXO1 may contribute to the regulation of adipocyte size in response to excessive calorie intake (36, 37). Because FOXO1 is a direct PPARγ ligand, it may also contribute to the regulation of adipose size and adipogenic pathways (15, 53). The nutrient-sensitive genes C/EBPα, sterol regulatory element-binding protein-1, and PPARγ, which are mandatory regulators of both early and late programs of adipose differentiation (6, 14, 56), were also activated early in pregnancy. As a whole, the changes in the metabolic machinery indicated that enhanced adipocyte storage and increased differentiation of new adipose cells were initiated at early pregnancy stages. These findings emphasize the notion that the molecular mechanisms of the adipogenic process come into place well before the appearance of classical phenotypic markers. Our data also suggest a similarity between the mechanisms of adipose tissue physiological expansion in pregnancy and pathological expansion in obesity. One example is the early increase in secreted frizzled-related protein 1, an endogenous modulator of Wnt/β-catenin signaling that peaks in patients with mild obesity and gradually falls in morbidly obese subjects (Table 2 and Ref. 31).
Inflammation.
Signs of molecular inflammation were detected in adipose tissue from early stages throughout the end of pregnancy. The functional cluster of genes related to immune regulation represented 30% of all modified genes in pregnancy compared with the pregravid state (Fig. 2). Increased expression of the macrophage markers CD14, CD68, HLA-DR, and HLA-DQ and the mannose receptor was in line with a recent report of inflammatory changes in adipose tissue in mice in late pregnancy (60). mRNA of TLR4 and downstream modulators were increased in early pregnancy. TLR4 belongs to a family of membrane receptors first recruited in innate sensing through binding of the lipid A moiety of lipopolysaccharide released from gram-negative bacteria (46).
The environmental factors that may trigger the recruitment of innate immune pathways in early pregnancy are not known. The activation of TLR4 signal transduction has been proposed as a molecular link between diet-induced obesity and increased insulin resistance (5, 54). Nutritional changes through either maternal hyperphagia or dysphagia are potential candidates to impact adipose tissue receptors and function (4). Along this line, changes in microbiota have been documented in relation to gestational weight gain and may impact TLR4 signaling in pregnancy (13). Bacterial recognition by the serum amyloid A (SAA) receptor CD36 may also initiate inflammation in response to fat supply (5, 59). The acute-phase proteins of the SAA family are markers of low-grade inflammation and obesity and promote endothelial remodeling partly through TLR4-mediated pathways (39, 58, 1).
Conclusion.
Whereas substantial literature has established inflammation as an obligatory component of adipose tissue remodeling in obesity and other diseases, this report is the first indication of recruitment of inflammatory pathways in healthy human pregnancy. Enhancement of adipose immune response preceded the appearance of maternal phenotypic changes in body composition and insulin action. This biphasic pattern reveals physiological inflammation as an early step toward the development of physiological insulin resistance, which peaks later in pregnancy.
GRANTS
This study was supported by National Institute of Child Health and Human Development Grant RO1-HD-022965 to P. Catalano and S. Hauguel-de Mouzon.
DISCLOSURES
No conflicts of interest, financial or otherwise, are declared by the authors.
AUTHOR CONTRIBUTIONS
V.R. analyzed the data; V.R. and S.H.-dM. interpreted the results of the experiments; V.R. and M.H. prepared the figures; V.R. and S.H.-dM. drafted the manuscript; V.R., S. Basu, M.H., L.P., S. Bernard, P.M.C., and S.H.-dM. approved the final version of the manuscript; S. Basu, M.H., L.P., J.M., B.K., and S. Bernard performed the experiments; P.M.C. and S.H.-dM. did the conception and design of the research; P.M.C. and S.H.-dM. edited and revised the manuscript.
REFERENCES
- 1.Averill MM, Kerkhoff C, Bornfeldt KE. S100A8 and S100A9 in cardiovascular biology and disease. Arterioscler Thromb Vasc Biol 32: 223–229, 2012 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Basu S, Haghiac M, Surace P, Challier JC, Guerre-Millo M, Singh K, Waters T, Minium J, Presley L, Catalano PM, Hauguel-de Mouzon S. Pregravid obesity associates with increased maternal endotoxemia and metabolic inflammation. Obesity (Silver Spring) 19: 476–482, 2011 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Battaglia FC, Meschia G. Principal substrates of fetal metabolism. Physiol Rev 58: 499–527, 1978 [DOI] [PubMed] [Google Scholar]
- 4.Björntorp P. Growth hormone, insulin-like growth factor-I and lipid metabolism: interactions with sex steroids. Horm Res 46: 188–191, 1996 [DOI] [PubMed] [Google Scholar]
- 5.Baranova IN, Bocharov AV, Vishnyakova TG, Kurlander R, Chen Z, Fu D, Arias IM, Csako G, Patterson AP, Eggerman TL. CD36 is a novel serum amyloid A (SAA) receptor mediating SAA binding and SAA-induced signaling in human and rodent cells. J Biol Chem 285: 8492–8506, 2010 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Boizard M, Le Liepvre X, Lemarchand P, Foufelle F, Ferré P, Dugail I. Obesity-related overexpression of fatty-acid synthase gene in adipose tissue involves sterol regulatory element-binding protein transcription factors. J Biol Chem 273: 29164–29171, 1998 [DOI] [PubMed] [Google Scholar]
- 7.Catalano PM, Tyzbir ED, Roman NM, Amini SB, Sims EA. Longitudinal changes in insulin release and insulin resistance in nonobese pregnant women. Am J Obstet Gynecol 165: 1667–1672, 1991 [DOI] [PubMed] [Google Scholar]
- 8.Catalano PM, Wong WW, Drago NM, Amini SB. Estimating body composition in late gestation: a new hydration constant for body density and total body water. Am J Physiol Endocrinol Metab 268: E153–E158, 1995 [DOI] [PubMed] [Google Scholar]
- 9.Catalano PM, Hoegh M, Minium J, Huston-Presley L, Bernard S, Kalhan S, Hauguel-De Mouzon S. Adiponectin in human pregnancy: implications for regulation of glucose and lipid metabolism. Diabetologia 49: 1677–1685, 2006 [DOI] [PubMed] [Google Scholar]
- 10.Catalano PM, Presley L, Minium J, Hauguel-de Mouzon S. Fetuses of obese mothers develop insulin resistance in utero. Diabetes Care 32: 1076–1080, 2009 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Chavey C, Mari B, Monthouel MN, Bonnafous S, Anglard P, Van Obberghen E, Tartare-Deckert S. Matrix metalloproteinases are differentially expressed in adipose tissue during obesity and modulate adipocyte differentiation. J Biol Chem 278: 11888–11896, 2003 [DOI] [PubMed] [Google Scholar]
- 12.Cinti S, Mitchell G, Barbatelli G, Murano I, Ceresi E, Faloia E, Wang S, Fortier M, Greenberg AS, Obin MS. Adipocyte death defines macrophage localization and function in adipose tissue of obese mice and humans. J Lipid Res 46: 2347–2355, 2005 [DOI] [PubMed] [Google Scholar]
- 13.Collado MC, Isolauri E, Laitinen K, Salminen S. Distinct composition of gut microbiota during pregnancy in overweight and normal-weight women. Am J Clin Nutr 88: 894–899, 2008 [DOI] [PubMed] [Google Scholar]
- 14.Girard J, Ferré P, Foufelle F. Mechanisms by which carbohydrates regulate expression of genes for glycolytic and lipogenic enzymes. Annu Rev Nutr 17: 325–352, 1997 [DOI] [PubMed] [Google Scholar]
- 15.Dowell P, Otto TC, Adi S, Lane MD. Convergence of peroxisome proliferator-activated receptor gamma and Foxo1 signaling pathways. J Biol Chem 278: 45485–45491, 2003 [DOI] [PubMed] [Google Scholar]
- 16.Doyle SL, Donohoe CL, Lysaght J, Reynolds JV. Visceral obesity, metabolic syndrome, insulin resistance and cancer. Proc Nutr Soc 3: 1–9, 2011 [DOI] [PubMed] [Google Scholar]
- 17.Drolet R, Richard C, Sniderman AD, Mailloux J, Fortier M, Huot C, Rhéaume C, Tchernof A. Hypertrophy and hyperplasia of abdominal adipose tissues in women. Int J Obes 32: 283–291, 2008 [DOI] [PubMed] [Google Scholar]
- 18.Ehrenberg HM, Huston-Presley L, Catalano PM. The influence of obesity and gestational diabetes mellitus on accretion and the distribution of adipose tissue in pregnancy. Am J Obstet Gynecol 189: 944–948, 2003 [DOI] [PubMed] [Google Scholar]
- 19.Evans J, Goedecke JH, Söderström I, Burén J, Alvehus M, Blomquist C, Jonsson F, Hayes PM, Adams K, Dave JA, Levitt NS, Lambert EV, Olsson T. Depot- and ethnic-specific differences in the relationship between adipose tissue inflammation and insulin sensitivity. Clin Endocrinol (Oxf) 74: 51–59, 2011 [DOI] [PubMed] [Google Scholar]
- 20.Keophiphath M, Achard V, Henegar C, Rouault C, Clément K, Lacasa D. Macrophage-secreted factors promote a profibrotic phenotype in human preadipocytes. Mol Endocrinol 23: 11–24, 2009 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Kirwan JP, Hauguel-De Mouzon S, Lepercq J, Challier JC, Huston-Presley L, Friedman JE, Kalhan SC, Catalano PM. TNF-alpha is a predictor of insulin resistance in human pregnancy. Diabetes 51: 2207–2213, 2002 [DOI] [PubMed] [Google Scholar]
- 22.Kopp-Hoolihan LE, van Loan MD. Fat mass deposition during pregnancy using a four-component model. J Appl Physiol 87: 196–202, 1992 [DOI] [PubMed] [Google Scholar]
- 23.Knopp RH, Warth MR, Carrol CJ. Lipid metabolism in pregnancy. I. Changes in lipoprotein triglyceride and cholesterol in normal pregnancy and the effects of diabetes mellitus. J Reprod Med 10: 95–101, 1973 [PubMed] [Google Scholar]
- 24.Hauguel-de Mouzon S, Lepercq J, Catalano P. The known and unknown of leptin in pregnancy. Am J Obstet Gynecol 194: 1537–1545, 2006 [DOI] [PubMed] [Google Scholar]
- 25.Herrera E. Metabolic adaptations in pregnancy and their implications for the availability of substrates to the fetus. Eur J Clin Nutr 54, Suppl 1: S47–S51, 2000 [DOI] [PubMed] [Google Scholar]
- 26.Highman TJ, Friedman JE, Huston LP, Wong WW, Catalano PM. Longitudinal changes in maternal serum leptin concentrations, body composition, and resting metabolic rate in pregnancy. Am J Obstet Gynecol 178: 1010–1015, 1998 [DOI] [PubMed] [Google Scholar]
- 27.Huston Presley L, Wong WW, Roman NM, Amini SB, Catalano PM. Anthropometric estimation of maternal body composition in late gestation. Obstet Gynecol 96: 33–37, 2000 [DOI] [PubMed] [Google Scholar]
- 28.Hytten FE. Water storage in pregnancy. Midwife Health Visit 3: 60–62, 1967 [PubMed] [Google Scholar]
- 29.Jo J, Gavrilova O, Pack S, Jou W, Mullen S, Sumner AE, Cushman SW, Periwal V. Hypertrophy and/or Hyperplasia: Dynamics of Adipose Tissue Growth. PLoS Comput Biol 5: e1000324, 2009 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.King JC, Butte NF, Bronstein MN, Kopp LE, Lindquist SA. Energy metabolism during pregnancy: influence of maternal energy status. Am J Clin Nutr 59: 439S–445S, 1994 [DOI] [PubMed] [Google Scholar]
- 31.Lagathu C, Christodoulides C, Tan CY, Virtue S, Laudes M, Campbell M, Ishikawa K, Ortega F, Tinahones FJ, Fernández-Real JM, Orešič M, Sethi JK, Vidal-Puig A. Secreted frizzled-related protein 1 regulates adipose tissue expansion and is dysregulated in severe obesity. Int J Obes (Lond) 34: 1695–1705, 2010 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Larciprete G, Valensise H, Vasapollo B, Altomare F, Sorge R, Casalino B, De Lorenzo A, Arduini D. Body composition during normal pregnancy: reference ranges. Acta Diabetol 40: S225–S232, 2003 [DOI] [PubMed] [Google Scholar]
- 33.Lee MJ, Wu Y, Fried SK. Adipose tissue remodeling in pathophysiology of obesity. Curr Opin Clin Nutr Metab Care 13: 371–376, 2010 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Lepercq J, Cauzac M, Lahlou N, Timsit J, Girard J, Auwerx J, Hauguel-de Mouzon S. Overexpression of placental leptin in diabetic pregnancy: a critical role for insulin. Diabetes 47: 847–850, 1998 [DOI] [PubMed] [Google Scholar]
- 35.Mortensen OH, Nielsen AR, Erikstrup C, Plomgaard P, Fischer CP, Krogh-Madsen R, Lindegaard B, Petersen AM, Taudorf S, Pedersen BK. Calprotectin—a novel marker of obesity. PLoS One 4: e7419, 2009 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Nakae J, Kitamura T, Kitamura Y, Biggs WH, 3rd, Arden KC, Accili D. The forkhead transcription factor Foxo1 regulates adipocyte differentiation. Dev Cell 4: 119–129, 2003 [DOI] [PubMed] [Google Scholar]
- 37.Nakae J, Cao Y, Oki M, Orba Y, Sawa H, Kiyonari H, Iskandar K, Suga K, Lombes M, Hayashi Y. Forkhead transcription factor FoxO1 in adipose tissue regulates energy storage and expenditure. Diabetes 57: 563–576, 2008 [DOI] [PubMed] [Google Scholar]
- 39.Poitou C, Coussieu C, Rouault C, Coupaye M, Cancello R, Bedel JF, Gouillon M, Bouillot JL, Oppert JM, Basdevant A, Clément K. Serum amyloid A: a marker of adiposity-induced low-grade inflammation but not of metabolic status. Obesity (Silver Spring) 14: 309–318, 2006 [DOI] [PubMed] [Google Scholar]
- 40.Poppitt SD, Prentice AM, Goldberg GR, Whitehead RG. Energy-sparing strategies to protect human fetal growth. Am J Obstet Gynecol 171: 118–125, 1994 [DOI] [PubMed] [Google Scholar]
- 41.Radaelli T, Lepercq J, Varastehpour A, Basu S, Catalano PM, Hauguel-De Mouzon S. Differential regulation of genes for fetoplacental lipid pathways in pregnancy with gestational and type 1 diabetes mellitus. Am J Obstet Gynecol 201: 209.e1–209.e10, 2009 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Ramos MP, Crespo-Solans MD, del Campo S, Cacho J, Herrera E. Fat accumulation in the rat during early pregnancy is modulated by enhanced insulin responsiveness. Am J Physiol Endocrinol Metab 285: E318–E328, 2003 [DOI] [PubMed] [Google Scholar]
- 43.Ryan EA, O'Sullivan MJ, Skyler JS. Insulin action during pregnancy. Studies with the euglycemic clamp technique. Diabetes 34: 380–389, 1985 [DOI] [PubMed] [Google Scholar]
- 44.Salans LB, Horton ES, Sims EA. Experimental obesity in man: cellular character of the adipose tissue. J Clin Invest 50: 1005–1011, 1971 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Sethi JK, Vidal-Puig AJ. Thematic review series: adipocyte biology. Adipose tissue function and plasticity orchestrate nutritional adaptation. J Lipid Res 48: 1253–1262, 2007 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Shi H, Kokoeva MV, Inouye K, Tzameli I, Yin H, Flier J. TLR4 links innate immunity and fatty acid-induced insulin resistance. J Clin Invest 116: 3015–3025, 2006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Spencer M, Unal R, Zhu B, Rasouli N, McGehee RE, Jr, Peterson CA, Kern PA. Adipose tissue extracellular matrix and vascular abnormalities in obesity and insulin resistance. J Clin Endocrinol Metab 96: E1990–E1998, 2011 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Sun K, Kusminski CM, Scherer PE. Adipose tissue remodeling and obesity. J Clin Invest 121: 2094–2101, 2011 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Tam J, Duda DG, Perentes JY, Quadri RS, Fukumura D, Jain RK. Blockade of VEGFR2 and not VEGFR1 can limit diet-induced fat tissue expansion: role of local versus bone marrow-derived endothelial cells. PLoS One 4: e4974, 2009 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Tchoukalova YD, Koutsari C, Karpyak MV, Votruba SB, Wendland E, Jensen MD. Subcutaneous adipocyte size and body fat distribution. Am J Clin Nutr 87: 56–63, 2008 [DOI] [PubMed] [Google Scholar]
- 51.Tsukumo DM, Carvalho-Filho MA, Carvalheira JB, Prada PO, Hirabara SM, Schenka AA, Araújo EP, Vassallo J, Curi R, Velloso LA, Saad MJ. Loss-of-function mutation in Toll-like receptor 4 prevents diet-induced obesity and insulin resistance. Diabetes 56: 1986–1998, 2007 [DOI] [PubMed] [Google Scholar]
- 52.Van Harmelen V, Skurk T, Röhrig K, Lee YM, Halbleib M, Aprath-Husmann I, Hauner H. Effect of BMI and age on adipose tissue cellularity and differentiation capacity in women. Int J Obes Relat Metab Disord 27: 889–895, 2003 [DOI] [PubMed] [Google Scholar]
- 53.Vidal-Puig A, Jimenez-Liñan M, Lowell BB, Hamann A, Hu E, Spiegelman B, Flier JS, Moller DE. Regulation of PPAR gamma gene expression by nutrition and obesity in rodents. J Clin Invest 97: 2553–2561, 1996 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Villar J, Cogswell M, Kestler E, Castillo P, Menendez R, Repke JT. Effect of fat and fat-free mass deposition during pregnancy on birth weight. Am J Obstet Gynecol 167: 1344–1352, 1992 [DOI] [PubMed] [Google Scholar]
- 56.Wu Z, Rosen ED, Brun R, Hauser S, Adelmant G, Troy AE, McKeon C, Darlington GJ, Spiegelman BM. Cross-regulation of C/EBP alpha and PPAR gamma controls the transcriptional pathway of adipogenesis and insulin sensitivity. Mol Cell 3: 151–158, 1999 [DOI] [PubMed] [Google Scholar]
- 57.Xu H, Barnes GT, Yang Q, Tan G, Yang D, Chou CJ, Sole J, Nichols A, Ross JS, Tartaglia LA, Chen H. Chronic inflammation in fat plays a crucial role in the development of obesity-related insulin resistance. J Clin Invest 112: 1821–1830, 2003 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Yamamoto Y, Gesta S, Lee KY, Tran TT, Saadatirad P, Kahn CR. Adipose depots possess unique developmental gene signatures. Obesity (Silver Spring) 18: 872–878, 2010 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Yang RZ, Lee MJ, Hu H, Pollin TI, Ryan AS, Nicklas BJ, Snitker S, Horenstein RB, Hull K, Goldberg NH, Goldberg AP, Shuldiner AR, Fried SK, Gong DW. Acute-phase serum amyloid A: an inflammatory adipokine and potential link between obesity and its metabolic complications. PLoS Med 3: e287, 2006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Zhang L, Sugiyama T, Murabayashi N, Umekawa T, Ma N, Kamimoto Y, Ogawa Y, Sagawa N. The inflammatory changes of adipose tissue in late pregnant mice. J Mol Endocrinol 47: 157–165, 2011 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Zhang QX, Magovern CJ, Mack CA, Budenbender KT, Ko W, Rosengart TK. Vascular endothelial growth factor is the major angiogenic factor in omentum: mechanism of the omentum-mediated angiogenesis. J Surg Res 67: 147–154, 1997 [DOI] [PubMed] [Google Scholar]