Abstract
In the absence of HER2 overexpression, triple-negative breast cancers (TNBCs) rely on signaling by epidermal growth factor receptor (EGFR/ErbB1/HER1) to convey growth signals and stimulate cell proliferation. Soluble EGF-like ligands are derived from their transmembrane precursors by ADAM proteases, but the identity of the ADAM that is primarily responsible for ligand release and activation of EGFR in TNBCs is not clear. Using publicly available gene expression data for patients with lymph node-negative breast tumors who did not receive systemic treatment, we show that ADAM12L is the only ADAM whose expression level is significantly associated with decreased distant metastasis-free survival times. Similar effect was not observed for patients with ER-negative non-TNBCs. There was a positive correlation between ADAM12L and HB-EGF and EGFR in TNBCs, but not in ER-negative non-TNBCs. We further demonstrate that ectopic expression of ADAM12L increased EGFR phosphorylation in a mouse intraductal xenograft model of early breast cancer. Finally, we detect strong correlation between the level of anti-ADAM12L and anti-phospho-EGFR immunostaining in human breast tumors using tissue microarrays. These studies suggest that ADAM12L is the primary protease responsible for the activation of EGFR in early stage, lymph node-negative TNBCs. Thus, our results may provide novel insight into the biology of TNBC.
Keywords: Metalloprotease, epidermal growth factor, estrogen receptor, xenograft, tissue microarrays
Introduction
Triple-negative breast cancer (TNBC) is an aggressive form of the disease, with poor prognosis, and limited treatment options [1–3]. TNBC, which accounts for ~20% of all breast cancers, is characterized by the absence of estrogen receptor (ER) and progesterone receptor (PR) expression, and the lack of human epidermal growth factor receptor 2 (HER2) overexpression. Based on their gene expression profiles, the majority of TNBCs fall into the basal-like or claudin-low molecular subtypes [4,5]. Due to the absence of ER and HER2, established targeted therapies, such as endocrine therapy or anti-HER2 therapy, are not applicable.
HER2 is a member of the ErbB family of receptors that include epidermal growth factor receptor (EGFR/ErbB1/HER1), ErbB2/HER2/Neu, ErbB3/HER3, and ErbB4/HER4 [6]. EGFR requires ligand binding for dimerization, autophosphorylation, and stimulation of its tyrosine kinase activity. HER2 does not bind ligands but exists in an open conformation that is constitutively available for dimerization with other ErbBs. HER3 lacks intrinsic tyrosine kinase activity. Signaling by EGFR, HER2, and HER3 plays fundamental roles in breast tumor cell proliferation, survival, and metastasis [7]. HER4 also binds ligands and possesses tyrosine kinase activity but, unlike EGFR or HER2, it elicits anti-proliferative responses in mammary epithelium [8]. In the absence of HER2 overexpression, TNBCs depend on EGFR to convey growth signals, and ligand-mediated activation of EGFR becomes important for tumor progression. Ligands for EGFR include epidermal growth factor (EGF), tumor growth factor α (TGFα), heparin-binding EGF-like growth factor (HB-EGF), amphiregulin (AREG), epigen (EPGN), epiregulin (EREG), and betacellulin (BTC) [9]. They are all synthesized as transmembrane precursors that need to be shed from the membrane to generate soluble, biologically active agonists. The shedding is mediated by cell surface proteases (sheddases) that belong predominantly to the ADAM (A Disintegrin And Metalloprotease) family. Studies in cells cultured in vitro showed that several different ADAMs are capable of generating soluble EGFR ligands [10–13]. ADAM17 has emerged as the primary sheddase of EGF-like ligands in breast cancer [14,15], although its prognostic value is not limited to TNBC [15]. Whether the expression levels of other ADAMs are linked to breast cancer outcome and whether the potential risk is particularly increased in TNBC has not been investigated.
The ADAM family comprises 20 members, 9 out of which are catalytically active and expressed beyond the reproductive system: ADAM8, -9, -10, -12, -15, -17, -19, -28, and -33 [16]. ADAM12 is strongly upregulated in breast tumors and it has been previously implicated in breast cancer [17–20]. ADAM12 mRNA is alternatively spliced, giving rise to two different protein isoforms: ADAM12L (a transmembrane form) and ADAM12S (a secreted form) [17]. Recently, ADAM12S was reported to enhance tumor cell migration and invasion through Matrigel in vitro and to increase metastasis in mice in vivo [20]. It is currently unknown whether these pro-tumor roles of ADAM12 observed in laboratory settings translate into increased patient risk in clinical settings. Furthermore, despite the fact that ADAM12 was shown to be a potent sheddase of at least two different EGF-like ligands, EGF and BTC, in cultured cells [10], it is not clear whether it plays any role in the activation of EGFR in breast tumors in vivo.
The goal of this study was to examine whether ADAM12L plays a role in activation of EGFR in TNBC. Our results suggest that ADAM12L is the primary protease responsible for the activation of EGFR in an early stage, lymph node-negative TNBC.
Methods
Patient populations
We used four datasets that are publicly available at Gene Expression Omnibus (GEO, http://www.ncbi.nlm.nih.gov/geo/): EMC286 (GSE2034) [21,22], Erasmus (GSE5327) [22], TRANSBIG (GSE7390) [23], and Mainz (GSE11121) [24]. We applied the following criteria for dataset selection: (a) patients with lymph-node negative tumors, (b) patients did not receive systemic adjuvant treatment, (c) distant metastasis free survival times (DMFS) were available, (d) gene expression profiling was performed using Affymetrix HG-U133A platform, and (e) the number of TNBCs identified by gene expression profiling (ref. [25], see below) was greater than 15. Patient clinical characteristics are reported in Table 1.
Table 1.
Patient characteristics
| EMC286 | Erasmus | TRANSBIG | Mainz | |
|---|---|---|---|---|
| GEO Accession Number | GSE 2034 | GSE5327 | GSE7390 | GSE11121 |
| No. of patients | 286 | 58 | 198 | 200 |
| Age (y) | ||||
| Median | Unknown | Unknown | 46 | 60 |
| Range | Unknown | Unknown | 24–60 | 34–89 |
| T stage | ||||
| T1 | 0 | 0 | 102 | 111 |
| T2 | 0 | 0 | 96 | 81 |
| T3/4 | 0 | 0 | 0 | 8 |
| Unknown | 286 | 58 | 0 | 0 |
| Tumor grade | ||||
| Grade 1 | 0 | 0 | 30 | 29 |
| Grade 2 | 0 | 0 | 83 | 136 |
| Grade 3 | 0 | 0 | 83 | 35 |
| Unknown | 286 | 58 | 2 | 0 |
| Lymph node status | ||||
| Positive | 0 | 0 | 0 | 0 |
| Negative | 286 | 58 | 198 | 200 |
| Unknown | 0 | 0 | 0 | 0 |
| ER status | ||||
| Positive | 209 | 0 | 134 | 156 |
| Negative | 77 | 58 | 64 | 44 |
| Unknown | 0 | 0 | 0 | 0 |
| PR status | ||||
| Positive | 0 | 0 | 0 | 130 |
| Negative | 0 | 0 | 0 | 70 |
| Unknown | 286 | 58 | 198 | 0 |
| HER2 status | ||||
| Positive | 0 | 0 | 0 | 0 |
| Negative | 0 | 0 | 0 | 0 |
| Unknown | 286 | 58 | 198 | 200 |
| Endocrine therapy | 0 | 0 | 0 | 0 |
| Chemotherapy | 0 | 0 | 0 | 0 |
| Median follow-up (y) | 7.2 | 7.2 | 13.6 | 7.7 |
| Distant metastasis | 107 | 11 | 62 | 46 |
| Reference | 21 | 22 | 23 | 24 |
Data analysis
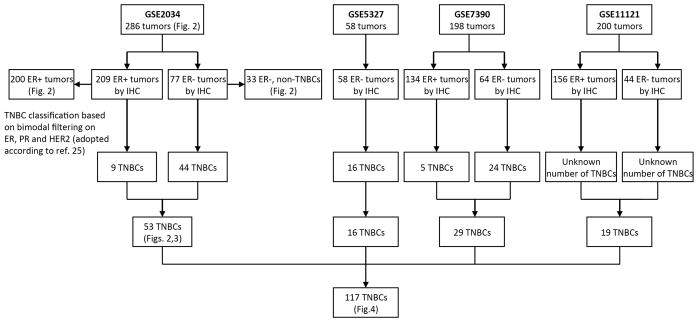
The EMC286, Erasmus, and TRANSBIG datasets contained information on the ER, but not PR or HER2 status. The Mainz dataset lacked information on ER, PR, and HER2. To identify TNBCs, we have adopted tumor classification based on bimodal filtering on ER, PR, and HER2 expression determined by Lehman et al. [25]. Previous studies have shown that microarray-based readout of ER, PR, and HER2 correlates well with immunohistochemical (IHC) determination of the ER, PR, and HER2 status [26,27]. Overall, 53 tumors in the EMC286 dataset were classified as TNBCs. Nine tumors that were originally categorized as ER-positive based on IHC, but were classified as TNBC based on ER/PR/HER2 mRNA expression, were considered TNBC in our analyses. Consistently, the set of ER-negative tumors comprised 77 tumors that were called ER-negative by IHC, plus 9 ER-positive tumors that were classified as TNBC based on gene expression profiling. The remaining 200 tumors were considered ER-positive. Erasmus, TRANSBIG and Mainz datasets contained 16, 29, and 19 TNBCs, respectively. To evaluate the prognostic value of ADAM12L in the combined set TNBCs from the fours cohorts, ADAM12L expression data were merged. Data were first log2-transformed, median-centered, and Z-score transformed [28] to normalize ADAM12L expression across all samples in each cohort. The flowchart of data analysis is shown in Fig. 1. The list of Affymetrix probes used to retrieve gene expression profiles is provided in Online Resource Table S1.
Fig. 1.
Flowchart of analysis according to REMARK criteria [59]
Statistical analyses
All data analyses were performed using GraphPad Prism version 5.0d. Survival curves were calculated using the Kaplan-Meier method and compared by the log-rank (Mantel-Cox) test. All reported p values are two-sided.
Additional methods are described in Online Resource Methods.
Results
High expression level of ADAM12L is associated with poor prognosis in lymph node-negative TNBC without systemic therapy
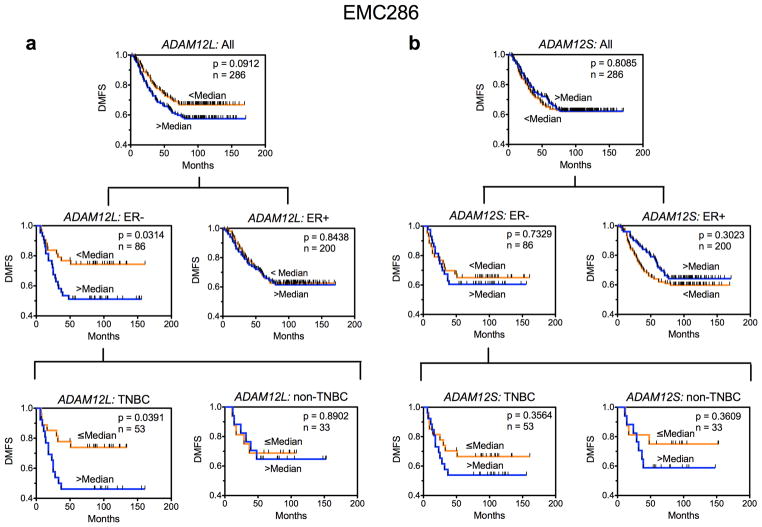
We assessed the prognostic value of ADAM12L and ADAM12S in 286 early stage breast cancer patients treated at the Erasmus Medical Center (the EMC286 dataset) [21]. All tumors were lymph node-negative, and patients did not receive neoadjuvant or adjuvant systemic therapy. Patients were divided into two groups with low and high expression of ADAM12L (Fig. 2a) or ADAM12S (Fig. 2b), with cut-offs set as median expression values. When all 286 tumors were analyzed together, the group with high expression of ADAM12L had worse prognosis than the group with low expression of ADAM12L, but the difference was not statistically significant. When ER-negative and ER-positive tumors were considered separately, a striking difference in the prognostic value of ADAM12L became evident. In ER-negative tumors, high expression of ADAM12L was associated with significantly lower DMFS rates (hazard ratio (HR) 2.154, 95% confidence interval (CI) 1.071 to 4.332, p = 0.0314). In ER-positive tumors, there was no difference in survival rates for patients with high or low levels of ADAM12L. Most importantly, within ER-negative tumors, poor prognosis associated with high ADAM12L was restricted to TNBCs (HR 2.492, 95% CI 1.047 to 5.933, p = 0.0391). Remarkably, in ER-negative non-TNBC, high expression of ADAM12L was not linked to poor outcome. Assuming that the difference between TNBC and ER-negative non-TNBC was mainly due to HER2 overexpression (the mean expression level of HER2 in TNBC was indeed ~12-fold lower than in ER-negative non-TNBC, p<0.0001, data not shown), this result indicates that the poor prognostic value of ADAM12L was manifested only in the absence of HER2 overexpression. Interestingly, although TNBCs overlap to a large extent with basal cancers (a majority of basal cancer are triple-negative, and 60–80% of TNBCs are basal tumors [1–4]), the prognostic value of ADAM12L in basal cancers was lower than in TNBCs (Online Resource Fig. S1). Thus, decreased DMFS in TNBC expressing high ADAM12L were not a simple consequence of poor prognostic value of ADAM12L in basal cancers. Given the fact that the mean values of ADAM12L were not statistically different between TNBCs and other tumor groups (Online Resource Fig. S2), these results suggest that ADAM12L may activate EGFR via the cleavage of EGF-like ligands, and this function is masked or irrelevant in the presence of HER2 overexpression. In a marked contrast to ADAM12L, ADAM12S did not have a significant prognostic value in any of the tumor groups examined (Fig. 2b).
Fig. 2.
High expression of ADAM12L is associated with poor prognosis in TNBC patients from the EMC286 cohort. Kaplan-Meier analysis of distant metastasis-free survival fractions for all patients (top), patients with ER-negative or ER-positive tumors (middle), and patients with TNBC or ER-negative non-TNBC (bottom). Patients were rank-ordered according to the expression of ADAM12L (a) or ADAM12S (b) and divided into two groups with mRNA expression levels below or above the median values.
Efficient activation of the EGFR pathway requires three components: the receptor, a ligand, and a sheddase that generates the soluble ligand. While the expression of these components may be regulated by entirely separate mechanisms, it is clear that increased levels of all three components create a particular advantage for tumor cells. Therefore, we examined the correlation between ADAM12L, EGF-like ligand mRNAs and EGFR. We observed no positive correlation in all 286 tumors or in ER-positive cancers (Table 2). In contrast, in ER-negative tumors, there was a positive correlation between ADAM12L and HB-EGF (Pearson r = 0.3038, p = 0.0045; Table 2). Most importantly, this positive correlation between ADAM12L and HB-EGF (Pearson r = 0.3632, p = 0.0075) and EGFR (Pearson r = 0.2728, p = 0.0481) was retained in the TNBC subset of ER-negative tumors. In the remaining ER-negative non-TNBCs, ADAM12L was not significantly correlated with any of the mRNAs tested. Interestingly, despite their correlation with ADAM12L, high expression of HB-EGF or EGFR mRNA alone was not significantly associated with decreased DMFS in TNBC (Online Resource Fig. S1). This is not surprising, since high HB-EGF or EGFR protein levels do not inevitably lead to the activation of EGFR unless a sheddase, i.e., ADAM12L, is also expressed at high level.
Table 2.
Correlation analysis of ADAM12L, EGF-like ligands, and EGFR in breast tumors from the EMC286 cohort. Values represent Pearson r.
| All
| |||||||
|---|---|---|---|---|---|---|---|
| EGF | HB-EGF | TGFα | AREG | BTC | EREG | EGFR | |
| ADAM12L | −0.0525 (p = 0.3766) | 0.0603 (p = 0.3095) | −0.0781 (p = 0.1877) | 0.0432 (p = 0.4663) | −0.1097 (p = 0.0640) | 0.0278 (p = 0.6392) | 0.0067 (p = 0.9104) |
| ER+
| |||||||
|---|---|---|---|---|---|---|---|
| EGF | HB-EGF | TGFα | AREG | BTC | EREG | EGFR | |
| ADAM12L | −0.0520 (p = 0.4646) | −0.0068 (p = 0.9243) | −0.1431 (p = 0.0432) | 0.0019 (p = 0.9784) | −0.1141 (p = 0.1077) | 0.0258 (p = 0.7174) | 0.0203 (p = 0.7750) |
| ER−
| |||||||
|---|---|---|---|---|---|---|---|
| EGF | HB-EGF | TGFα | AREG | BTC | EREG | EGFR | |
| ADAM12L | −0.08296 (p = 0.4476) | 0.3038 (p = 0.0045) | −0.0214 (p = 0.8446) | 0.1193 (p = 0.2738) | −0.0497 (p = 0.6493) | 0.0006 (p = 0.9954) | 0.1703 (p = 0.1169) |
| TNBC
| |||||||
|---|---|---|---|---|---|---|---|
| EGF | HB-EGF | TGFα | AREG | BTC | EREG | EGFR | |
| ADAM12L | −0.1004 (p = 0.4744) | 0.3632 (p = 0.0075) | 0.0504 (p = 0.7202) | 0.2246 (p = 0.1060) | −0.1060 (p = 0.4500) | −0.0276 (p-0.8445) | 0.2728 (p = 0.0481) |
| non-TNBC
| |||||||
|---|---|---|---|---|---|---|---|
| EGF | HB-EGF | TGFα | AREG | BTC | EREG | EGFR | |
| ADAM12L | −0.1030 (p = 0.5685) | 0.2593 (p = 0.1450) | −0.2398 (p = 0.1789) | 0.0735 (p = 0.6843) | 0.1051 (p = 0.5605) | 0.1093 (p = 0.5448) | 0.1751 (p = 0.3297) |
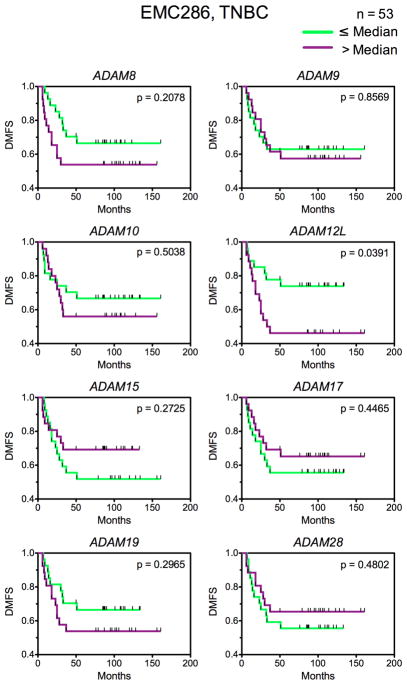
Next, we asked whether other transmembrane, catalytically active ADAMs also contribute to poor prognosis in TNBCs. Out of eight other catalytically active ADAMs whose expression is not limited to the reproductive system, the following seven were represented on the Affymetrix U133A array: ADAM8, -9, -10, -15, -17, -19, and -28. Among TNBC patients from the EMC286 cohort, ADAM12L was the only ADAM that was significantly associated with poor prognosis (Fig. 3).
Fig. 3.
The prognostic value of ADAM12L in TNBCs from the EMC286 dataset is the highest among other ADAMs encoding membrane-anchored, catalytically-active ADAM proteases. Kaplan-Meier plots of distant metastasis-free survival fractions for TNBC patients from the EMC286 cohort are shown.
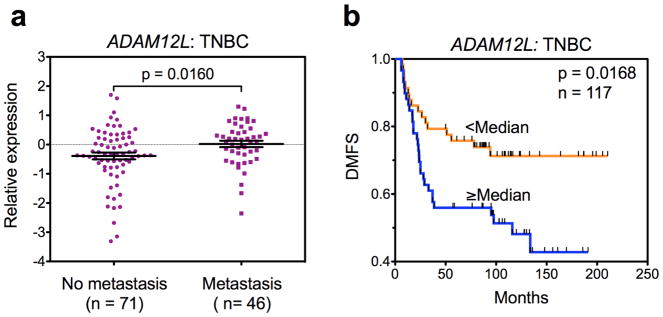
To extend our investigations beyond the EMC286 cohort, we combined ADAM12L expression profiles in TNBCs from the EMC286 dataset and from three other cohorts, Erasmus [22], TRANSBIG [23], and Mainz [24]. All tumors were node-negative and patients did not receive adjuvant chemotherapy. In this combined dataset of 117 TNBCs, ADAM12L levels were significantly higher in patients with distant metastases than in patients without metastasis (Fig. 4a). Patients with high expression of ADAM12L had significantly lower survival rates than patients with low expression of ADAM12L (HR 2.030, 95% CI 1.136 to 3.628, p = 0.0168; Fig. 4b).
Fig. 4.
High expression of ADAM12L is associated with poor prognosis in a combined set of node-negative TNBCs without chemotherapy. Gene expression profiles obtained in four independent studies (GEO accession numbers GSE2034, GSE5327, GSE7390, and GSE11121) were merged, as described in Methods. a Log2-transformed, median-centered ADAM12L expression values in TNBC patients without vs with distant metastasis. Mean expression values ± SEM are indicated. b Kaplan-Meier plots of distant metastasis-free survival fractions are shown.
ADAM12L augments EGFR phosphorylation in vivo
Next, we examined whether there is a cause-and-effect relationship between the level of expression of ADAM12L protein and the extent of EGFR phosphorylation in vivo in a mouse model of an early ER-negative breast cancer. We used the MIND (mouse intraductal) transplantation model, in which ER-negative MCF10DCIS.com breast tumor cells are transplanted directly into the mammary ducts [29].
MCF10DCIS.com cells express high levels of EGFR [29]. We confirmed that this receptor is fully functional, as it became readily phosphorylated at Tyr1173, one of the major autophosphorylation sites [30], in response to added EGF (Fig. 5a). The endogenous expression level of ADAM12L protein in MCF10DCIS.com cells was low and poorly detected by immunoblotting (Fig. 5b). Using a retroviral delivery system and selection with puromycin, we established cells with stable overexpression of ADAM12L. The ectopically expressed ADAM12L produced two bands in Western blots (Fig. 5b). The ~120-kDa band represented the inactive intracellular ADAM12L, and the ~90-kDa band represented the processed, active form that resides at the cell surface [31]. Flow cytometry using live cells further demonstrated surface expression of ADAM12L in stably transduced cells (Fig. 5b).
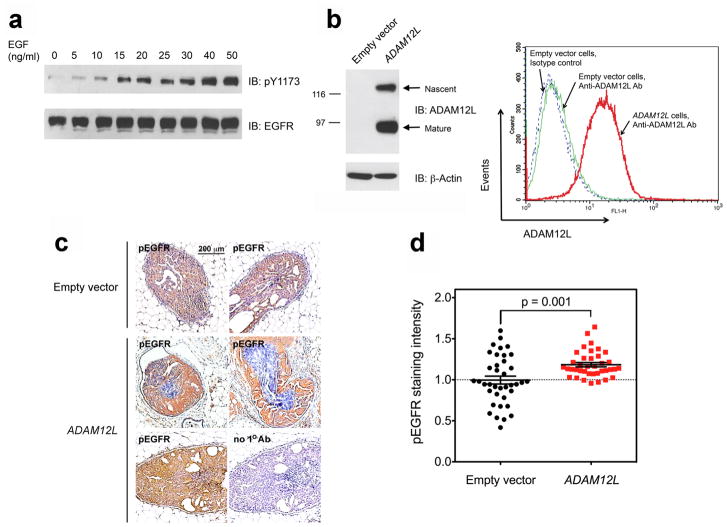
Fig. 5.
Increased expression of ADAM12L augments EGFR phosphorylation in a mouse-intraductal (MIND) xenograft model of basal breast cancer. a Detection of a functional EGFR in MCF10DCIS.com cells. Cells were serum-starved for 16 h and then incubated for 30 min with indicated concentrations of EGF. The extent of EGFR phosphorylation (pY1173) was examined by Western blotting. b Stable expression of ADAM12L in MCF10DCIS.com cells. Cells stably transduced with ADAM12L or with empty vector were analyzed by Western blotting using an antibody against the cytoplasmic tail of ADAM12L (left). The nascent form represents the full-length, catalytically inactive, intracellular form of ADAM12L. The mature form represents ADAM12L lacking its N-terminal pro-domain, which is catalytically active and resides predominantly at the cell surface. Cell surface localization of the mature form of ADAM12L was confirmed by flow cytometry after staining of live cells with an antibody recognizing the extracellular domain of ADAM12L. c Immunohistochemical detection of phosphorylated EGFR in mouse intraductal xenografts. MCF10DCIS.com cells stably transduced with ADAM12L or with empty vector were injected into mammary ducts of NSG mice. Six weeks after injection, paraffin-embedded mammary tissue sections were stained with anti-phospho-EGFR (Y1173) antibody, visualized with DAB (brown), and counterstained with hematoxylin (blue). Four representative images are shown in the top and middle rows, respectively. The specificity of staining was confirmed by incubation of duplicate sections with or without primary antibody (bottom row). d Comparison of anti-phospho-EGFR staining intensities in intraductal xenografts of MCF10DCIS.com cells stably transduced with empty vector or ADAM12L. For each cell type, 38 tissue sections from three different mice were analyzed. Quantification was performed as described in Online Resource Methods. All staining intensity values (in arbitrary units) were normalized to the mean value obtained for cells transduced with empty vector.
ADAM12L overexpressing cells and control cells transduced with vector only were transplanted into mammary ducts. Six weeks later, the extent of EGFR phosphorylation in intraductal lesions was examined by immunohistochemistry. As reported earlier, most of the lesions at this time point were DCIS-like, with some lesions already progressing to invasion [29]. Fig. 5c shows examples of anti-phospho-EGFR (Y1173) staining in the intraductal lesions. To assess the effect of ADAM12L on the extent of EGFR phosphorylation, the intensity of anti-phospho-EGFR staining was quantified, as described in Online Resource Methods. Analysis of 76 different intraductal lesions (38 lesions per each cell type, 6 mice total) showed significantly higher anti-phospho-EGFR staining in lesions generated by ADAM12L-overexpressing cells than control cells (Fig. 5d). This result suggested that ADAM12L was capable of inducing EGFR phosphorylation in the MIND model in vivo. As MCF10DCIS.com cells express oncogenic H-RasG12V [32], it was not feasible to study downstream effects of EGFR activation.
EGFR phosphorylation in human breast tumors strongly correlates with ADAM12L expression level
To assess an association between EGFR phosphorylation and the level of ADAM12L protein in human breast tumors, we performed IHC analysis of tissue microarrays. We analyzed 142 samples of 72 different breast tumors, using anti-ADAM12L and anti-phospho-EGFR (Y1173) antibodies (Fig. 6a,b). The results show a strong correlation between staining intensities for ADAM12L and for phospho-EGFR (Pearson r = 0.8353, p < 0.0001) (Fig. 6c). Interestingly, this strong correlation was not limited just to TNBC, as a similar degree of correlation was observed when only TNBC samples were included in the analysis (Pearson r = 0.7663, p < 0.0001) (Fig. 6c). Thus, ADAM12L may be a universal modulator of EGFR activation in all breast tumors, but this activity of ADAM12L becomes biologically relevant only in the context of TNBC, where the EGFR pathway controls cell proliferation.
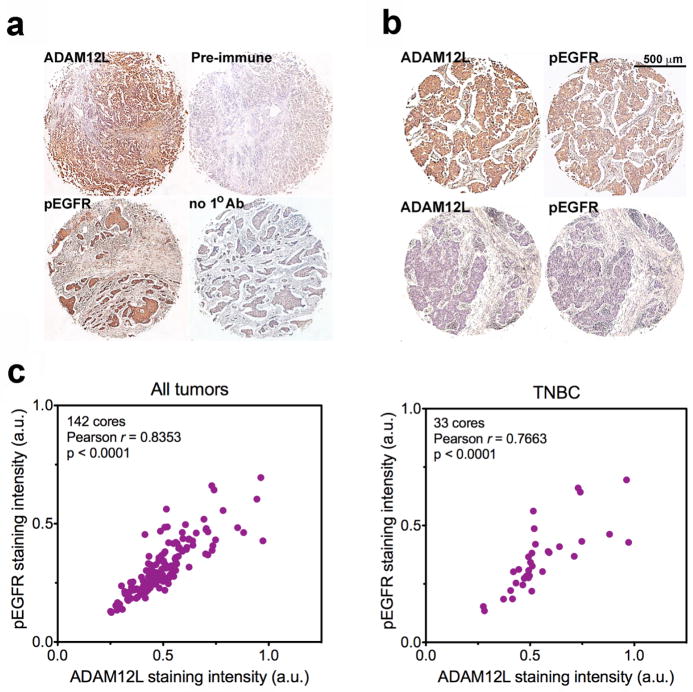
Fig. 6.
Positive correlation between the level of ADAM12L expression and the extent of EGFR phosphorylation in human breast cancer tissue microarrays. a Validation of the specificities of the antibodies used for IHC staining of tissue microarrays. Two cores of the same tumor were stained with anti-ADAM12L antibody or with pre-immune serum (top). Similarly, two cores of the same tumor were stained with anti-pEGFR (Y1173) or with the secondary antibody only (bottom). Positive staining was observed mostly in epithelial cells, with much weaker staining in the stromal compartment. b Two examples of duplicate sections in which epithelial cells stained strongly (top) or weakly (bottom) for both ADAM12L and pEGFR. c Correlation analysis of ADAM12L staining versus pEGFR staining intensities in 142 cores, including 33 cores of TNBC. Staining intensities (in arbitrary units) were quantified as described in Online Resource Methods.
Discussion
Due to the absence of ER and HER2 in TNBC, anti-estrogen or anti-HER2 therapies are not possible. The only remaining option for systemic treatment is conventional chemotherapy. However, as chemotherapy typically aims at basic cellular processes such as cell division, it is non-specific and harmful to normal dividing cells. Furthermore, many TNBCs acquire chemoresistance after prolonged exposure to cytotoxic drugs, compromising the efficacy of the treatment. Thus, finding new molecular targets in TNBC and identifying novel predictive markers are of the utmost urgency.
The ongoing clinical trials for patients with TNBC include novel cytotoxic drugs, poly(ADP-ribose) polymerase (PARP) inhibitors, antiangiogenic agents, Src and Abl kinase inhibitors, and EGFR-targeting agents [33]. EGFR seems to be a logical target in TNBC for several reasons. First, in the absence of HER2-overexpression, TNBCs depend on signaling by EGFR to stimulate tumor cell proliferation and survival. Second, 60–70% of all TNBCs show increased expression of EGFR [34–37]. Third, KRAS mutations are infrequent in TNBC [38], making the tumors amenable to EGFR inhibition. Current EGFR-targeting agents being evaluated in clinical trials include small molecule inhibitors of the EGFR tyrosine kinase, as well as monoclonal antibodies (Cetuximab) [3,39]. However, so far these strategies have produced limited success [1,39]. One possible reason could be that the clinical studies involving EGFR inhibitors were conducted in unselected TNBC patients, and were not directed by any mechanistic hypotheses. Since ligand binding is an absolute requirement for EGFR activation, and since activating mutations in EGFR that would nullify this requirement are rare [40,41], limiting ligand availability represents another logical approach to block the EGFR pathway. Furthermore, one can postulate that TNBC patients with the highest sheddase activity, in which soluble EGF-like ligands are produced with the highest efficiency, are the best candidates for the existing anti-EGFR therapies. This approach requires, however, precise knowledge of the sheddase responsible for the release of EGF-like ligands and activation of EGFR.
In this study, we demonstrate that high levels of ADAM12L are associated with decreased DMFS in TNBC and we link this effect to the activation of EGFR. Importantly, ADAM12L was the only ADAM encoding a transmembrane, catalytically active protease whose expression level was significantly linked to poor prognosis in TNBC patients. We further used an in vivo model to confirm a causal role of ADAM12L protein in tyrosine phosphorylation of EGFR, which often serves as a read-out of the activation status of the receptor. Importantly, we uncovered a strong correlation between ADAM12L protein level and the extent of EGFR phosphorylation using tissue arrays.
Although the significance of ADAM-mediated shedding of EGF-like ligands in breast cancer biology has been well recognized [42–44], the identity of the “perpetrator” ADAM that is actually responsible for ligand cleavage and activation of EGFR has not been conclusively determined. To our best knowledge, the current study represents the first evaluation of the prognostic values of all catalytically active ADAMs in systemically untreated TNBCs. This is also the first report that specifically links ADAM12L to the activation of EGFR in TNBC. This function of ADAM12L has not been previously revealed by cell biology approaches or by mouse models. Interestingly, the two breast cancer-associated loss-of-function mutations in ADAM12, i.e., D301H and G479E, were not found in TNBC [45,46]. The third mutation, L792F, was found in TNBC, but this mutation does not alter the proteolytic activity of ADAM12L (ref. [47] and our unpublished observations). Thus, it appears that TNBCs express the active form of ADAM12L, which is consistent with its role in EGFR activation in TNBC described here.
One unexpected result of our survival analyses is the discrepancy between the prognostic values of ADAM12L and ADAM12S. The distinction between ADAM12L and ADAM12S is important, because these two mRNAs encode transmembrane and secreted proteases, respectively, with potentially diverse biological functions during tumor progression and metastasis. Recently, ADAM12S has been recently shown to be more potent than ADAM12L in inducing metastasis in an orthotopic mouse xenograft model [20]. This result does not agree with the results of our survival analyses presented here, but the discrepancy may be explained by the fact that the model used in the referenced study did not mimic human TNBC.
Finally, an important question remains: what is the functional mechanism by which ADAM12L promotes metastasis in node-negative TNBC, despite complete removal of the primary tumor? One possibility is that the ADAM12L/EGFR axis increases the chance of an early dissemination of a small number of cancer cells prior to surgery, and these cells go undetected in the node analysis. EGFR activation has been shown to promote epithelial-to-mesenchymal-like transition (EMT), down-regulation of E-cadherin, tumor cell migration, and invasion [48–53], and these functions fit the proposed model of ADAM12L/EGFR enhancing tumor cell dissemination. Another possibility is that ADAM12L-mediated activation of the EGFR pathway might enhance tumor cell colonization at a metastatic site. Recent studies have shown that shedding of EGF-like ligands from tumor cells modulates in a paracrine manner the bone microenvironment and promotes osteolytic metastasis [54]. The sheddases implicated in this process are ADAMTS1 (a disintegrin and metalloproteinase with thrombospondin motifs1) and MMP1 (matrix metalloprotease 1). These two proteases are not linked to poor prognosis in TNBCs from the EMC286 cohort (results not shown). However, as the pattern of metastatic spread in TNBC is different from other types of breast cancer, with brain and liver metastases being more frequent than metastasis to bone or lung [55–57], the identity of the sheddases and the spectrum of EGF-like ligands involved may also be different. Recently, HB-EGF has been shown to be a mediator of cancer cell passage through the blood-brain barrier [58]. Our preliminary examination of the gene expression data suggested that the expression level of ADAM12L might be indeed higher in TNBCs that metastasized to the brain than in other TNBCs, but because of the limited number of cases available, the difference did not reach statistical significance (results not shown).
The discovery of ADAM12L as a prognostic factor in TNBC and linking its function to the EGFR pathway may have at least two potential implications. First, it should promote further studies to establish whether ADAM12L may serve as a novel therapeutic target in TNBC. It should also encourage a quest for highly specific inhibitors that would effectively discriminate between ADAM12L and other catalytically active ADAMs. Second, stratification of TNBC patients based on the expression level of ADAM12L may identify a group that could benefit the most from inhibition of EGFR and thus it may increase the efficacy of anti-EGFR treatment.
Supplementary Material
Acknowledgments
This work was supported by NIH grant 1R15CA151065 and Innovative Research Award from Terry C. Johnson Center for Basic Cancer Research at KSU to AZ, and by NIH grant 5R00CA127462 to FB. This is contribution 12-470-J from Kansas Agricultural Experiment Station.
Footnotes
Conflict of interest
The authors declare that they have no conflict of interest.
References
- 1.Carey L, Winer E, Viale G, Cameron D, Gianni L. Triple-negative breast cancer: disease entity or title of convenience? Nat Rev Clin Oncol. 2010;7(12):683–692. doi: 10.1038/nrclinonc.2010.154. [DOI] [PubMed] [Google Scholar]
- 2.Foulkes WD, Smith IE, Reis-Filho JS. Triple-negative breast cancer. New Engl J Med. 2010;363(20):1938–1948. doi: 10.1056/NEJMra1001389. [DOI] [PubMed] [Google Scholar]
- 3.Pal SK, Childs BH, Pegram M. Triple negative breast cancer: unmet medical needs. Breast Cancer Res Treat. 2011;125(3):627–636. doi: 10.1007/s10549-010-1293-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Sotiriou C, Pusztai L. Gene-expression signatures in breast cancer. New Engl J Med. 2009;360(8):790–800. doi: 10.1056/NEJMra0801289. [DOI] [PubMed] [Google Scholar]
- 5.Prat A, Parker JS, Karginova O, Fan C, Livasy C, Herschkowitz JI, He X, Perou CM. Phenotypic and molecular characterization of the claudin-low intrinsic subtype of breast cancer. Breast Cancer Res. 2010;12(5):R68. doi: 10.1186/bcr2635. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Hynes NE, MacDonald G. ErbB receptors and signaling pathways in cancer. Curr Opin Cell Biol. 2009;21(2):177–184. doi: 10.1016/j.ceb.2008.12.010. [DOI] [PubMed] [Google Scholar]
- 7.Foley J, Nickerson NK, Nam S, Allen KT, Gilmore JL, Nephew KP, Riese DJ., 2nd EGFR signaling in breast cancer: bad to the bone. Semin Cell Dev Biol. 2010;21(9):951–960. doi: 10.1016/j.semcdb.2010.08.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Muraoka-Cook RS, Feng SM, Strunk KE, Earp HS., 3rd ErbB4/HER4: role in mammary gland development, differentiation and growth inhibition. J Mammary Gland Biol Neoplasia. 2008;13(2):235–246. doi: 10.1007/s10911-008-9080-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Wilson KJ, Gilmore JL, Foley J, Lemmon MA, Riese DJ., 2nd Functional selectivity of EGF family peptide growth factors: implications for cancer. Pharmacol Ther. 2009;122(1):1–8. doi: 10.1016/j.pharmthera.2008.11.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Horiuchi K, Le Gall S, Schulte M, Yamaguchi T, Reiss K, Murphy G, Toyama Y, Hartmann D, Saftig P, Blobel CP. Substrate selectivity of epidermal growth factor-receptor ligand sheddases and their regulation by phorbol esters and calcium influx. Mol Biol Cell. 2007;18(1):176–188. doi: 10.1091/mbc.E06-01-0014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Sunnarborg SW, Hinkle CL, Stevenson M, Russell WE, Raska CS, Peschon JJ, Castner BJ, Gerhart MJ, Paxton RJ, Black RA, Lee DC. Tumor necrosis factor-α converting enzyme (TACE) regulates epidermal growth factor receptor ligand availability. J Biol Chem. 2002;277(15):12838–12845. doi: 10.1074/jbc.M112050200. [DOI] [PubMed] [Google Scholar]
- 12.Sternlicht MD, Sunnarborg SW, Kouros-Mehr H, Yu Y, Lee DC, Werb Z. Mammary ductal morphogenesis requires paracrine activation of stromal EGFR via ADAM17-dependent shedding of epithelial amphiregulin. Development. 2005;132(17):3923–3933. doi: 10.1242/dev.01966. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Sahin U, Weskamp G, Kelly K, Zhou HM, Higashiyama S, Peschon J, Hartmann D, Saftig P, Blobel CP. Distinct roles for ADAM10 and ADAM17 in ectodomain shedding of six EGFR ligands. J Cell Biol. 2004;164(5):769–779. doi: 10.1083/jcb.200307137. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Kenny PA, Bissell MJ. Targeting TACE-dependent EGFR ligand shedding in breast cancer. J Clin Invest. 2007;117(2):337–345. doi: 10.1172/JCI29518. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.McGowan PM, McKiernan E, Bolster F, Ryan BM, Hill AD, McDermott EW, Evoy D, O’Higgins N, Crown J, Duffy MJ. ADAM-17 predicts adverse outcome in patients with breast cancer. Annals of oncology. 2008;19(6):1075–1081. doi: 10.1093/annonc/mdm609. [DOI] [PubMed] [Google Scholar]
- 16.Edwards DR, Handsley MM, Pennington CJ. The ADAM metalloproteinases. Mol Aspects Med. 2008;29(5):258–289. doi: 10.1016/j.mam.2008.08.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Kveiborg M, Albrechtsen R, Couchman JR, Wewer UM. Cellular roles of ADAM12 in health and disease. Int J Biochem Cell Biol. 2008;40(9):1685–1702. doi: 10.1016/j.biocel.2008.01.025. [DOI] [PubMed] [Google Scholar]
- 18.Kveiborg M, Frohlich C, Albrechtsen R, Tischler V, Dietrich N, Holck P, Kronqvist P, Rank F, Mercurio AM, Wewer UM. A role for ADAM12 in breast tumor progression and stromal cell apoptosis. Cancer Res. 2005;65(11):4754–4761. doi: 10.1158/0008-5472.CAN-05-0262. [DOI] [PubMed] [Google Scholar]
- 19.Frohlich C, Nehammer C, Albrechtsen R, Kronqvist P, Kveiborg M, Sehara-Fujisawa A, Mercurio AM, Wewer UM. ADAM12 produced by tumor cells rather than stromal cells accelerates breast tumor progression. Mol Cancer Res. 2011;9(11):1449–1461. doi: 10.1158/1541-7786.MCR-11-0100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Roy R, Rodig S, Bielenberg D, Zurakowski D, Moses MA. ADAM12 transmembrane and secreted isoforms promote breast tumor growth: a distinct role for ADAM12-S protein in tumor metastasis. J Biol Chem. 2011;286(23):20758–20768. doi: 10.1074/jbc.M110.216036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Wang Y, Klijn JG, Zhang Y, Sieuwerts AM, Look MP, Yang F, Talantov D, Timmermans M, Meijer-van Gelder ME, Yu J, Jatkoe T, Berns EM, Atkins D, Foekens JA. Gene-expression profiles to predict distant metastasis of lymph-node-negative primary breast cancer. Lancet. 2005;365(9460):671–679. doi: 10.1016/S0140-6736(05)17947-1. [DOI] [PubMed] [Google Scholar]
- 22.Minn AJ, Gupta GP, Padua D, Bos P, Nguyen DX, Nuyten D, Kreike B, Zhang Y, Wang Y, Ishwaran H, Foekens JA, van de Vijver M, Massague J. Lung metastasis genes couple breast tumor size and metastatic spread. Proc Nat Acad Sci USA. 2007;104(16):6740–6745. doi: 10.1073/pnas.0701138104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Desmedt C, Piette F, Loi S, Wang Y, Lallemand F, Haibe-Kains B, Viale G, Delorenzi M, Zhang Y, d’Assignies MS, Bergh J, Lidereau R, Ellis P, Harris AL, Klijn JG, Foekens JA, Cardoso F, Piccart MJ, Buyse M, Sotiriou C. Strong time dependence of the 76-gene prognostic signature for node-negative breast cancer patients in the TRANSBIG multicenter independent validation series. Clin Cancer Res. 2007;13(11):3207–3214. doi: 10.1158/1078-0432.CCR-06-2765. [DOI] [PubMed] [Google Scholar]
- 24.Schmidt M, Bohm D, von Torne C, Steiner E, Puhl A, Pilch H, Lehr HA, Hengstler JG, Kolbl H, Gehrmann M. The humoral immune system has a key prognostic impact in node-negative breast cancer. Cancer Res. 2008;68(13):5405–5413. doi: 10.1158/0008-5472.CAN-07-5206. [DOI] [PubMed] [Google Scholar]
- 25.Lehmann BD, Bauer JA, Chen X, Sanders ME, Chakravarthy AB, Shyr Y, Pietenpol JA. Identification of human triple-negative breast cancer subtypes and preclinical models for selection of targeted therapies. J Clin Invest. 2011;121(7):2750–2767. doi: 10.1172/JCI45014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Press MF, Finn RS, Cameron D, Di Leo A, Geyer CE, Villalobos IE, Santiago A, Guzman R, Gasparyan A, Ma Y, Danenberg K, Martin AM, Williams L, Oliva C, Stein S, Gagnon R, Arbushites M, Koehler MT. HER-2 gene amplification, HER-2 and epidermal growth factor receptor mRNA and protein expression, and lapatinib efficacy in women with metastatic breast cancer. Clin Cancer Res. 2008;14(23):7861–7870. doi: 10.1158/1078-0432.CCR-08-1056. [DOI] [PubMed] [Google Scholar]
- 27.Roepman P, Horlings HM, Krijgsman O, Kok M, Bueno-de-Mesquita JM, Bender R, Linn SC, Glas AM, van de Vijver MJ. Microarray-based determination of estrogen receptor, progesterone receptor, and HER2 receptor status in breast cancer. Clin Cancer Res. 2009;15(22):7003–7011. doi: 10.1158/1078-0432.CCR-09-0449. [DOI] [PubMed] [Google Scholar]
- 28.Cheadle C, Vawter MP, Freed WJ, Becker KG. Analysis of microarray data using Z score transformation. J Mol Diagn. 2003;5(2):73–81. doi: 10.1016/S1525-1578(10)60455-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Behbod F, Kittrell FS, LaMarca H, Edwards D, Kerbawy S, Heestand JC, Young E, Mukhopadhyay P, Yeh HW, Allred DC, Hu M, Polyak K, Rosen JM, Medina D. An intraductal human-in-mouse transplantation model mimics the subtypes of ductal carcinoma in situ. Breast Cancer Res. 2009;11(5):R66. doi: 10.1186/bcr2358. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Olayioye MA, Neve RM, Lane HA, Hynes NE. The ErbB signaling network: receptor heterodimerization in development and cancer. EMBO J. 2000;19(13):3159–3167. doi: 10.1093/emboj/19.13.3159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Cao Y, Kang Q, Zhao Z, Zolkiewska A. Intracellular processing of metalloprotease disintegrin ADAM12. J Biol Chem. 2002;277(29):26403–26411. doi: 10.1074/jbc.M110814200. [DOI] [PubMed] [Google Scholar]
- 32.Miller FR, Santner SJ, Tait L, Dawson PJ. MCF10DCIS.com xenograft model of human comedo ductal carcinoma in situ. J Natl Cancer Inst. 2000;92 (14):1185–1186. doi: 10.1093/jnci/92.14.1185a. [DOI] [PubMed] [Google Scholar]
- 33.Carey LA. Directed therapy of subtypes of triple-negative breast cancer. Oncologist. 2011;16(Suppl 1):71–78. doi: 10.1634/theoncologist.2011-S1-71. [DOI] [PubMed] [Google Scholar]
- 34.Shien T, Tashiro T, Omatsu M, Masuda T, Furuta K, Sato N, Akashi-Tanaka S, Uehara M, Iwamoto E, Kinoshita T, Fukutomi T, Tsuda H, Hasegawa T. Frequent overexpression of epidermal growth factor receptor (EGFR) in mammary high grade ductal carcinomas with myoepithelial differentiation. J Clin Pathol. 2005;58(12):1299–1304. doi: 10.1136/jcp.2005.026096. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Arnes JB, Begin LR, Stefansson I, Brunet JS, Nielsen TO, Foulkes WD, Akslen LA. Expression of epidermal growth factor receptor in relation to BRCA1 status, basal-like markers and prognosis in breast cancer. J Clin Pathol. 2009;62(2):139–146. doi: 10.1136/jcp.2008.056291. [DOI] [PubMed] [Google Scholar]
- 36.Viale G, Rotmensz N, Maisonneuve P, Bottiglieri L, Montagna E, Luini A, Veronesi P, Intra M, Torrisi R, Cardillo A, Campagnoli E, Goldhirsch A, Colleoni M. Invasive ductal carcinoma of the breast with the “triple-negative” phenotype: prognostic implications of EGFR immunoreactivity. Breast Cancer Res Treat. 2009;116(2):317–328. doi: 10.1007/s10549-008-0206-z. [DOI] [PubMed] [Google Scholar]
- 37.Corkery B, Crown J, Clynes M, O’Donovan N. Epidermal growth factor receptor as a potential therapeutic target in triple-negative breast cancer. Annals Oncol. 2009;20(5):862–867. doi: 10.1093/annonc/mdn710. [DOI] [PubMed] [Google Scholar]
- 38.Sanchez-Munoz A, Gallego E, de Luque V, Perez-Rivas LG, Vicioso L, Ribelles N, Lozano J, Alba E. Lack of evidence for KRAS oncogenic mutations in triple-negative breast cancer. BMC Cancer. 2010;10:136. doi: 10.1186/1471-2407-10-136. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Peddi PF, Ellis MJ, Ma C. Molecular basis of triple negative breast cancer and implications for therapy. Int J Breast Cancer. 2012;2012:217185. doi: 10.1155/2012/217185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Bhargava R, Gerald WL, Li AR, Pan Q, Lal P, Ladanyi M, Chen B. EGFR gene amplification in breast cancer: correlation with epidermal growth factor receptor mRNA and protein expression and HER-2 status and absence of EGFR-activating mutations. Modern Pathol. 2005;18(8):1027–1033. doi: 10.1038/modpathol.3800438. [DOI] [PubMed] [Google Scholar]
- 41.Reis-Filho JS, Pinheiro C, Lambros MB, Milanezi F, Carvalho S, Savage K, Simpson PT, Jones C, Swift S, Mackay A, Reis RM, Hornick JL, Pereira EM, Baltazar F, Fletcher CD, Ashworth A, Lakhani SR, Schmitt FC. EGFR amplification and lack of activating mutations in metaplastic breast carcinomas. J Pathol. 2006;209(4):445–453. doi: 10.1002/path.2004. [DOI] [PubMed] [Google Scholar]
- 42.Blobel CP, Carpenter G, Freeman M. The role of protease activity in ErbB biology. Exp Cell Res. 2009;315(4):671–682. doi: 10.1016/j.yexcr.2008.10.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Duffy MJ, McKiernan E, O’Donovan N, McGowan PM. Role of ADAMs in cancer formation and progression. Clin Cancer Res. 2009;15(4):1140–1144. doi: 10.1158/1078-0432.CCR-08-1585. [DOI] [PubMed] [Google Scholar]
- 44.Duffy MJ, Mullooly M, O’Donovan N, Sukor S, Crown J, Pierce A, McGowan PM. The ADAMs family of proteases: new biomarkers and therapeutic targets for cancer? Clin Proteomics. 2011;8(1):9. doi: 10.1186/1559-0275-8-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Sjoblom T, Jones S, Wood LD, Parsons DW, Lin J, Barber TD, Mandelker D, Leary RJ, Ptak J, Silliman N, Szabo S, Buckhaults P, Farrell C, Meeh P, Markowitz SD, Willis J, Dawson D, Willson JK, Gazdar AF, Hartigan J, Wu L, Liu C, Parmigiani G, Park BH, Bachman KE, Papadopoulos N, Vogelstein B, Kinzler KW, Velculescu VE. The consensus coding sequences of human breast and colorectal cancers. Science. 2006;314(5797):268–274. doi: 10.1126/science.1133427. [DOI] [PubMed] [Google Scholar]
- 46.Dyczynska E, Syta E, Sun D, Zolkiewska A. Breast cancer-associated mutations in metalloprotease disintegrin ADAM12 interfere with the intracellular trafficking and processing of the protein. Int J Cancer. 2008;122(11):2634–2640. doi: 10.1002/ijc.23405. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Stautz D, Wewer UM, Kveiborg M. Functional Analysis of a Breast Cancer-Associated Mutation in the Intracellular Domain of the Metalloprotease ADAM12. PLoS One. 2012;7(5):e37628. doi: 10.1371/journal.pone.0037628. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Lu Z, Jiang G, Blume-Jensen P, Hunter T. Epidermal growth factor-induced tumor cell invasion and metastasis initiated by dephosphorylation and downregulation of focal adhesion kinase. Mol Cell Biol. 2001;21(12):4016–4031. doi: 10.1128/MCB.21.12.4016-4031.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Lu Z, Ghosh S, Wang Z, Hunter T. Downregulation of caveolin-1 function by EGF leads to the loss of E-cadherin, increased transcriptional activity of beta-catenin, and enhanced tumor cell invasion. Cancer Cell. 2003;4 (6):499–515. doi: 10.1016/s1535-6108(03)00304-0. [DOI] [PubMed] [Google Scholar]
- 50.Hager MH, Morley S, Bielenberg DR, Gao S, Morello M, Holcomb IN, Liu W, Mouneimne G, Demichelis F, Kim J, Solomon KR, Adam RM, Isaacs WB, Higgs HN, Vessella RL, Di Vizio D, Freeman MR. DIAPH3 governs the cellular transition to the amoeboid tumour phenotype. EMBO Mol Med. 2012 doi: 10.1002/emmm.201200242. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Nie F, Yang J, Wen S, An YL, Ding J, Ju SH, Zhao Z, Chen HJ, Peng XG, Wong ST, Zhao H, Teng GJ. Involvement of epidermal growth factor receptor overexpression in the promotion of breast cancer brain metastasis. Cancer. 2012 doi: 10.1002/cncr.27553. [DOI] [PubMed] [Google Scholar]
- 52.Samanta S, Sharma VM, Khan A, Mercurio AM. Regulation of IMP3 by EGFR signaling and repression by ERbeta: implications for triple-negative breast cancer. Oncogene. 2012 doi: 10.1038/onc.2011.620. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Jin W, Chen BB, Li JY, Zhu H, Huang M, Gu SM, Wang QQ, Chen JY, Yu S, Wu J, Shao ZM. TIEG1 inhibits breast cancer invasion and metastasis by inhibition of epidermal growth factor receptor (EGFR) transcription and the EGFR signaling pathway. Mol Cell Biol. 2012;32(1):50–63. doi: 10.1128/MCB.06152-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Lu X, Wang Q, Hu G, Van Poznak C, Fleisher M, Reiss M, Massague J, Kang Y. ADAMTS1 and MMP1 proteolytically engage EGF-like ligands in an osteolytic signaling cascade for bone metastasis. Genes & development. 2009;23(16):1882–1894. doi: 10.1101/gad.1824809. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Fulford LG, Reis-Filho JS, Ryder K, Jones C, Gillett CE, Hanby A, Easton D, Lakhani SR. Basal-like grade III invasive ductal carcinoma of the breast: patterns of metastasis and long-term survival. Breast Cancer Res. 2007;9(1):R4. doi: 10.1186/bcr1636. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Hicks DG, Short SM, Prescott NL, Tarr SM, Coleman KA, Yoder BJ, Crowe JP, Choueiri TK, Dawson AE, Budd GT, Tubbs RR, Casey G, Weil RJ. Breast cancers with brain metastases are more likely to be estrogen receptor negative, express the basal cytokeratin CK5/6, and overexpress HER2 or EGFR. Am J Surg Pathol. 2006;30(9):1097–1104. doi: 10.1097/01.pas.0000213306.05811.b9. [DOI] [PubMed] [Google Scholar]
- 57.Dent R, Hanna WM, Trudeau M, Rawlinson E, Sun P, Narod SA. Pattern of metastatic spread in triple-negative breast cancer. Breast Cancer Res Treat. 2009;115(2):423–428. doi: 10.1007/s10549-008-0086-2. [DOI] [PubMed] [Google Scholar]
- 58.Bos PD, Zhang XH, Nadal C, Shu W, Gomis RR, Nguyen DX, Minn AJ, van de Vijver MJ, Gerald WL, Foekens JA, Massague J. Genes that mediate breast cancer metastasis to the brain. Nature. 2009;459(7249):1005–1009. doi: 10.1038/nature08021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.McShane LM, Altman DG, Sauerbrei W, Taube SE, Gion M, Clark GM. Reporting recommendations for tumor marker prognostic studies (REMARK) J Nat Cancer Inst. 2005;97(16):1180–1184. doi: 10.1093/jnci/dji237. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.