Abstract
Lysosomal enzymes catalyze the breakdown of macromolecules in the cell. In humans, loss of activity of a lysosomal enzyme leads to an inherited metabolic defect known as a lysosomal storage disorder. The human lysosomal enzyme galactosamine-6-sulfatase (GALNS, also known as N-acetylgalactosamine-6-sulfatase and GalN6S; E.C. 3.1.6.4) is deficient in patients with the lysosomal storage disease mucopolysaccharidosis IV A (also known as MPS IV A and Morquio A). Here we report the three-dimensional structure of human GALNS, determined by x-ray crystallography at 2.2 Å resolution. The structure reveals a catalytic gem diol nucleophile derived from modification of a cysteine side chain. The active site of GALNS is a large, positively charged trench suitable for binding polyanionic substrates such as keratan sulfate and chondroitin-6-sulfate. Enzymatic assays on the insect cell-expressed human GALNS indicate activity against synthetic substrates and inhibition by both substrate and product. Mapping 120 MPS IV A missense mutations onto the structure reveals that a majority of mutations affect the hydrophobic core of the structure, indicating that most MPS IV A cases result from misfolding of GALNS. Comparison of the structure of GALNS to paralogous sulfatases shows a wide variety of active site geometries in the family, but strict conservation of the catalytic machinery. Overall, the structure and the known mutations establish the molecular basis for MPS IV A and for the larger MPS family of diseases.
Keywords: sulfatase structure, lysosomal storage disease, enzyme replacement therapy, x-ray crystallography, cysteine modification
Introduction
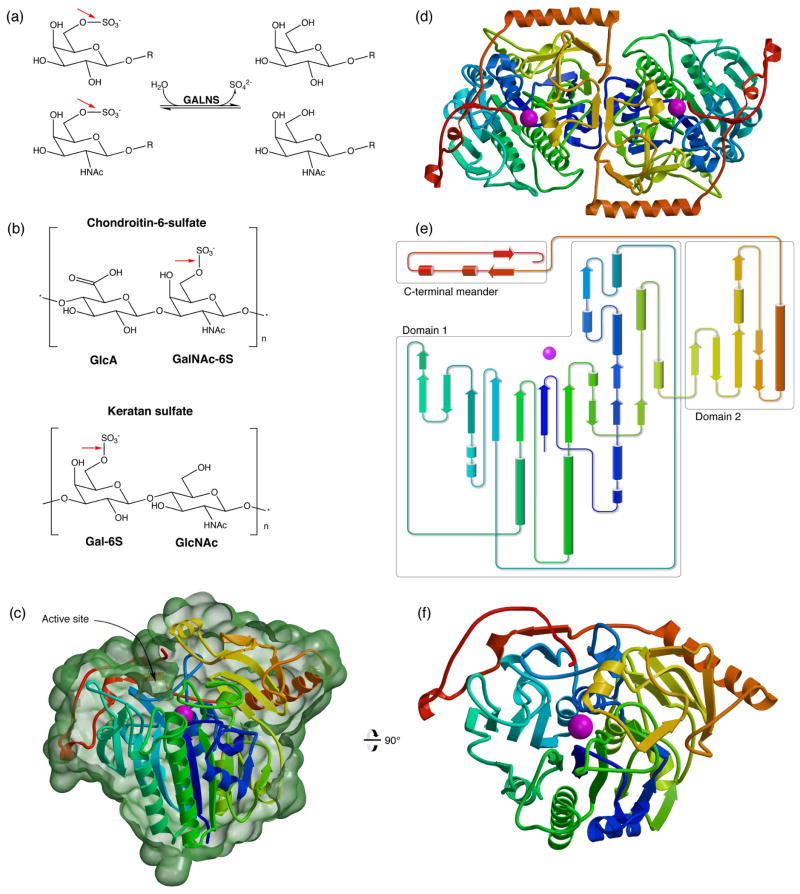
Lysosomal enzymes are responsible for the catabolism of biomolecules. Deficiencies in lysosomal enzymes result in accumulation of undegraded substrates in tissues, leading eventually to lysosomal storage diseases. The human lysosomal galactosamine-6-sulfatase (GALNS, also known as N-acetylgalactosamine-6-sulfatase and GalN6S; E.C. 3.1.6.4) removes sulfate groups from a terminal N-acetylgalactosamine-6-sulfate (or galactose-6-sulfate) in mucopolysaccharides such as keratan sulfate and chondroitin-6-sulfate (Fig. 1a and b). Defects in GALNS lead to accumulation of substrates, resulting in the development of the lysosomal storage disease mucopolysaccharidosis IV A (also known as MPS IV A and Morquio A disease).1
Fig. 1.
Human GALNS reaction and overall structure
(a): GALNS cleaves 6-sulfate attached to Gal and GalNAc saccharides. (b): GALNS substrates include chondroitin-6-sulfate and keratan sulfate. Red arrows show the bond cleaved by GALNS. (c): The GALNS monomer structure colored from blue to red from N to C terminus and Ca2+ in magenta. (This coloring scheme is matched in subsequent panels.) The molecular surface in green (calculated in Povscript43) is cut away to indicate the trench defining the active site. (d): An overview of the GALNS dimer structure viewed down the molecular dyad. (e): A topology diagram of the monomer shows domains boxed in grey. (f): An orthogonal view of the GALNS monomer viewed down the large β sheet in domain 1.
The autosomal recessive MPS IV A affects approximately 1 in 200,000 live births and is characterized by a wide variety of symptoms, including severe skeletal abnormalities, hearing loss, corneal clouding, heart valve disease, and other impairments.2 The disease presents two phenotypes depending on its severity: a mild form, which generally allows the patient a full lifespan, and a severe form, which often results in death before the second decade of life.2 However, the correlation between genotype and phenotype is not fully understood. At present, 157 different mutations have been identified in the GALNS gene in patients with MPS IV A, 120 of which are missense mutations leading to a change of a single side chain residue in the protein.3 While there is no currently approved treatment for MPS IV A, enzyme replacement therapy, where patients are injected weekly with recombinant enzyme, is in Phase III clinical trials.4
Despite the intense clinical interest in GALNS and the large amounts of purified protein available for decades, to date there has been no structure of the protein reported. GALNS was first purified from human placenta in 1979,5 and later was isolated from liver cells6 and fibroblasts. The recombinant enzyme used in clinical trials is purified from Chinese hamster ovary cells.9 Because of the importance of GALNS and its relationship to disease, those interested in the structure have resorted to homology modeling10 using the best available paralogs, the 36% and 28% identical human arylsulfatase A11 (ASA) and arylsulfatase B12 (ASB) structures.
In general, sulfatases catalyze the hydrolysis of sulfate ester bonds from a diverse range of substrates. In vitro, most sulfatases can hydrolyze synthetic substrates such as 4-methylumbelliferyl sulfate (4-MU-S), which shares only a sulfate group with native substrates. Therefore, determining native substrate specificity for sulfatases is non-trivial.
Sulfatases require a modified amino acid nucleophile for catalytic function. The polypeptide encodes a cysteine (or occasionally serine), which is then enzymatically converted to a formylglycine aldehyde by formylglycine-generating enzyme (FGE).13–16 The ubiquitous FGE recognizes a specific sequence motif (typically CXPXR), and oxidative desulfurization of the side chain leads to an aldehyde. Hydration of the aldehyde leads to a dihydroxyalanine (DHA) nucleophile with geminal hydroxy groups.21–23
To elucidate the molecular basis for MPS IV A, we determined the structure of human GALNS to 2.2 Å resolution. Using stably transfected insect cells, we expressed human GALNS glycoprotein and demonstrated enzyme activity comparable to endogenous enzyme. Using x-ray crystallography and mass spectrometry, we identified FGE modification of the GALNS nucleophile in our insect cell expression system. To extend our structural knowledge about GALNS to the effects seen in patients with MPS IV A, we mapped the disease-causing mutations onto the structure. The mutations fall into three categories: disruption of the active site, perturbation of overall fold, and surface exposure, suggesting potential treatments depending on a patient’s genotype. Overall, these results will improve the understanding of the molecular defects in MPS IV A and will provide insight into lysosomal storage diseases and other protein folding diseases.
Results
Protein expression
We purified GALNS from Trichoplusia ni (Tn5) insect cells, using both baculovirus-infected and stable-cell line approaches. The protein sequence contained the native signal sequence for secretion and a C-terminal hexahistidine tag for purification. Baculovirus-infected cells yielded about 0.2 mg purified GALNS per liter of culture while stably transfected cell lines yielded approximately 0.5 mg per liter. Monoclonal selection of the stable cell lines increased expression to 0.7 mg per liter of culture.
Overall description of the structure
The structure of the recombinant human GALNS was determined by X-ray crystallography to a resolution of 2.2 Å (Table 1). The X-ray structure reveals GALNS as a homodimeric glycoprotein with each monomer having an N-terminal domain containing the active site (domain 1), a second domain with antiparallel β-strands (domain 2), and a C-terminal meander (Fig. 1c–f). Domain 1 (residues 28-379) has an α/β topology formed from a core ten-stranded β-sheet at the center of six α-helices. Domain 2 (residues 380-481) comprises a four anti-parallel stranded β-sheet perpendicular to a long α-helix (Fig. 1e). Domain 2 is followed by a C-terminal meander (residues 482-510), which threads back into domain 1 and defines a portion of the active site. The GALNS active site contains a calcium liganded to the catalytic nucleophile and the side chains of four other residues. Each monomer contains two N-linked glycosylation sites at Asn 204 and Asn 423, the latter of which falls close to the molecular dyad, and the attached carbohydrates pack against one another. Each monomer contains three disulfide bonds (308-419, 489-518, and 501-507), and an unpaired cysteine (164). The interface between the two monomers in the dimer buries 3094 Å2 of surface area, and the large interface is consistent with the dimeric GALNS used in clinical trials as enzyme replacement therapy for MPS IV A.24
Table 1.
X-ray crystallographic data
| PDB: | 4FDI | 4FDJ |
|---|---|---|
| Ligand Soak | - | GalNAc |
| Beamline | APS 24-ID-C | BNL X6A |
| Wavelength, Å | 0.9795 | 1.100 |
| Space Group | P21 | P21 |
| Cell Lengths, Å | 61.4, 155.5, 62.6 | 60.7, 155.8, 61.8 |
| Cell β angle, ° | 113.8 | 113.4 |
| Resolution, Å (last shell) | 50-2.2 (2.26-2.2) | 50-2.8 (2.85-2.8) |
| Observations overall | 202,615 (9,634) | 71,414 (2,536) |
| Unique observations (last shell) | 54,154 (2,676) | 24,690 (1,055) |
| Completeness, % (last shell) | 99.7 (99.3) | 96.4 (83.7) |
| Multiplicity (last shell) | 3.7 (3.6) | 2.9 (2.4) |
| Rsym, % (last shell) | 11.0 (27.7) | 14.3 (27.8) |
| <I/σI> (last shell) | 14.3 (4.2) | 8.6 (2.5) |
|
| ||
| Refinement | ||
| Rwork/Rfree (%) | 16.4/21.1 | 20.7/25.1 |
| No. of atoms: total | 8,682 | 8,080 |
| Protein | 7,811 | 7,787 |
| Carbohydrate | 56 | 86 |
| Water | 775 | 205 |
| Other | 40 | 2 |
| Average B-factors, Å2 | 22.8 | 40.0 |
| Protein | 21.8 | 39.9 |
| Carbohydrate | 56.0 | 64.5 |
| Water | 29.9 | 33.2 |
| Ramachandran Plot, % | ||
| Favored | 96.7 | 95.9 |
| Allowed | 3.3 | 4.1 |
| Outlier | 0.0 | 0.0 |
| RMS deviations | ||
| Bonds, Å | 0.007 | 0.008 |
| Angles, ° | 1.10 | 1.16 |
The formylglycine modification and enzyme mechanism
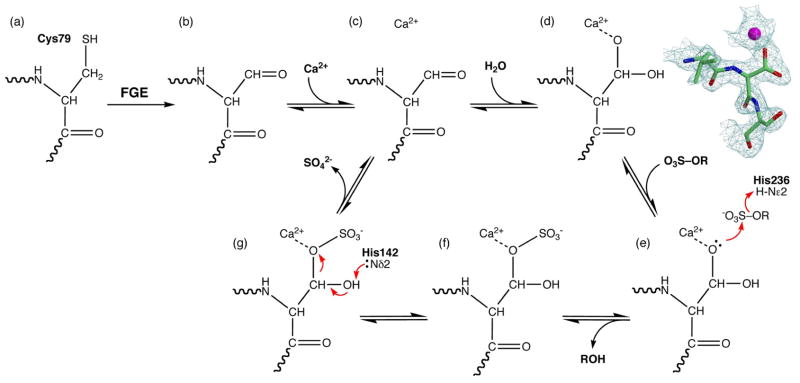
Sulfatases require maturation of a side chain residue into a catalytic nucleophile. In GALNS, the polypeptide chain encodes a cysteine at residue 79 at the start of the motif CXPXR (Fig. 2a). FGE recognizes this motif and converts Cys 79 into a formylglycine aldehyde (Fig. 2b).25 Addition of Ca2+ in the active site (Fig. 2c) and hydration of the aldehyde by a water molecule generates the gem diol nucleophile DHA (Fig. 2d). The GALNS crystal structure contains branched electron density at residue 79, corresponding to the hydrated form of the aldehyde (Fig. 2d). In the denatured and metal-chelated GALNS protein examined by mass spectrometry, residue 79 contains an overwhelming abundance of formylglycine (MW 85) with little presence of cysteine (MW 103) and undetectable amounts of DHA (MW 102) (Figs. 2 and SF1). The equilibrium between the aldehyde and the hydrated gem diol is shifted toward the aldehyde in the denatured GALNS and toward the gem diol in the crystal structure. The insect-cell expressed GALNS demonstrates nearly 100% post-translational modification of Cys 79, as seen by both mass spectrometry and crystallography.
Fig. 2.
GALNS maturation and mechanism
The GALNS gene encodes a cysteine at position 79 (a), which formylglycine generating enzyme (FGE) enzymatically converts into a formylglycine aldehyde (b). In the crystal, the aldehyde binds the calcium cofactor (c) and hydrates into the gem diol DHA (d). Substrate binding leads to nucleophilic attack on the sulfate (e), resulting of transfer of sulfate to the enzyme (f). Finally, intramolecular hydrolysis releases the sulfate (g) and regenerates the aldehyde. The protein in the crystal appears in state (d), while denaturing the enzyme for mass spectrometry disrupts the metal binding site and shifts the equilibrium to the aldehyde seen in state (b). The electron density in (d) is a σA-weighted 2Fo-Fc map contoured at 1.8s of the tripeptide centered at residue 79.
Both Ca2+ and Mg2+ have been found in the active sites of sulfatases. Mg2+ can appear in the active site when present at high concentrations during the purification and/or crystallization process, but Ca2+ is the preferred ion for biochemical activity. With GALNS, crystallographic refinement of Mg2+ in the active site resulted in positive peaks of 4–5 σ in difference electron density maps while Ca2+ led to a more consistent fit to the electron density. Additionally, we checked the bond valence of the metal using the software Valence,27 which showed a value of 1.92 for Ca2+ and a value of 0.918 for Mg2+, strongly suggesting that the ion in GALNS is Ca2+.
After the discovery of FGE and its creation of an active site aldehyde in sulfatases, it was unclear whether the sulfatase mechanism involved the sulfated substrate attacking the aldehyde on the enzyme or the DHA nucleophile on the enzyme attacking the sulfated substrate.22 Our structure lends support to the latter mechanism, that upon substrate binding, nucleophilic attack of the sulfate by DHA cleaves the substrate (Fig. 2e and f). Subsequent intramolecular hydrolysis of the sulfated enzyme intermediate uses the unique gem diol to regenerate the apoenzyme (Fig. 2g).
The active site and ligand binding
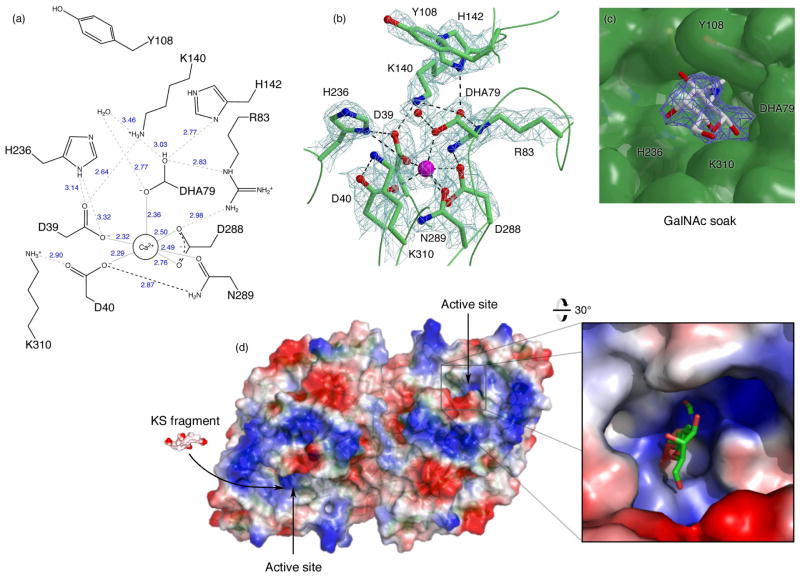
The active site of GALNS is found in a pocket at the center of the N-terminal α/β domain. The primary active site residues include Asp 39, Asp 40, Arg 83, Tyr 108, Lys 140, His 142, His 236, Asp 288, Asn 289, Lys 310, and DHA 79 (Fig. 3a). The active site contains a Ca2+ hexavalently coordinated by oxygens from the nucleophile at 79, Asp 39, Asp 40, Asp 288 (2 atoms), and Asn 289 (Fig. 3b). A citrate molecule from the buffer binds in the active site. The active site contains a high concentration of basic residues, leading to a considerable positive electrostatic surface potential, as expected for an enzyme that binds highly sulfated substrates (Fig. 3d). GALNS has a predicted isoelectric point of 6.3, and the electrostatic surface shows a molecule of pronounced negative potential with patches of positive potential near the active site.
Fig. 3.
GALNS active site and ligand binding
(a): Schematic of interactions in the GALNS active site. NCS-averaged interatomic distances in Å are shown in blue. (b) σA-weighted 2Fo-Fc map of the active site residues contoured at 1.8σ. (c): Ligand-bound structure. The electron density shows a σA-weighted 2Fo-Fc map contoured at 1.3σ around the GalNAc ligand. The molecular surface shows the large substrate-binding cavity with key residues indicated. (d): The GALNS electrostatic surface potential (calculated in Pymol42) plotted from −57kT (red) to +57kT (blue) shows the large areas of positive charge suitable for interacting with polyanionic substrates. The active sites are indicated with arrows, and the inset shows the active site with GalNAc bound. A tetrasaccharide fragment of the keratan sulfate substrate (labeled “KS fragment”) is shown for scale.
To better understand the molecular basis for ligand binding in GALNS, we determined the crystal structure of the protein soaked with a catalytic product, the N-acetylgalactosamine (GalNAc) monosaccharide, at 2.8 Å resolution (Fig. 3c and Table 1). The sugar binds in a non-productive orientation, with the 1-OH 2.6 Å from the catalytic nucleophile and the 6-OH (which is sulfated in the substrate) pointing away from the nucleophile. GalNAc binds GALNS using a common carbohydrate-recognition motif, where the plane of the sugar makes van der Waals contacts with the aromatic side chain Tyr 108.
The GALNS active site lies at the base of a deep conical crevice surrounded by positively charged groups (Fig. 3d). Unlike lysosomal glycosidases, which have exquisite specificity for the saccharide they recognize, GALNS and other lysosomal sulfatases have substantial promiscuity in substrate recognition. For example, GALNS will efficiently hydrolyze the synthetic substrate 4-MU-S, which shares only a sulfate group with the natural substrate. This promiscuity in substrate recognition is reflected in the substrate-binding pocket, a large open depression on the surface of the enzyme, allowing a wide variety of molecules access to the catalytic site. With the natural substrate keratan sulfate, the multiple negative charges on the polysulfated substrate presumably help guide the substrate into the active site (Fig. 3d).
Enzyme kinetics and inhibition
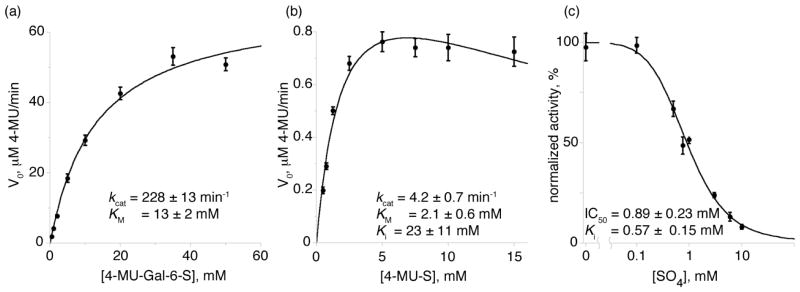
In the lysosome, GALNS removes sulfate groups from 6-sulfated galactosides and 6-sulfated N-acetylgalactosaminides in keratan sulfate and chondroitin-6-sulfate (Fig. 1a and b). To examine the enzymatic activity of purified GALNS, we measured the kinetic parameters KM and kcat of the enzyme in vitro using two synthetic substrates: 4-methylumbelliferyl-β-D-galactose-6-sulfate (4-MU-Gal-6-S)30 and 4-MU-S (Fig. 4). The 4-MU-Gal-6-S substrate mimics the terminal sulfated sugar found in the natural substrates, and GALNS has reported KM values of 0.05 mM for keratan sulfate6 and 0.01 mM for chondroitin-6-sulfate.31 With the 4-MU-Gal-6-S substrate, recombinant GALNS has a kcat of 228 ± 13 min−1 and a KM of 13 ± 2 mM (Fig. 4a), comparable to the values measured for ASA towards a synthetic substrate containing a 4-MU fluorophore.22 With the 4-MU-S substrate, GALNS has a kcat of 4.2 ± 0.7 min−1 and a KM of 2.1 ± 0.6 mM and showed substrate inhibition (with a substrate Ki of 23 mM), suggesting a non-productive binding mode (Fig. 4c). Because the natural substrates show considerably lower KM values than their smaller synthetic homologs,6,31 sulfatases such as GALNS interact with more of the natural substrates beyond the terminal sulfated monosaccharide. The kinetic parameters of the insect cell-expressed human GALNS are comparable to those from GALNS purified from endogenous sources.6
Fig. 4.
GALNS reaction kinetics
(a) and (b): Michaelis-Menten velocity vs. substrate concentration plots of GALNS with the synthetic substrates 4-MU-Gal-6-S (a) and 4-MU-S (b). 4-MU-S showed evidence of substrate inhibition (see text). (c): GALNS cleavage of 4-MU-S is inhibited by the presence of sulfate, which shows an IC50 of 0.89 mM in the assay.
To enable inhibitor testing of human GALNS, we established an assay for measuring inhibition of cleavage of the 4-MU-S substrate by GALNS. We hypothesized that sulfate (the catalytic product of the GALNS reaction) would act as a competitive inhibitor, and the inhibition assay showed an IC50 of 0.89 ± 0.23 mM and a Ki of 0.57 ± 0.15 mM for sulfate inhibition of GALNS (Fig. 4c). This assay will allow us to test small molecules for inhibition and potential pharmacological chaperone activity.
Comparison to other sulfatases
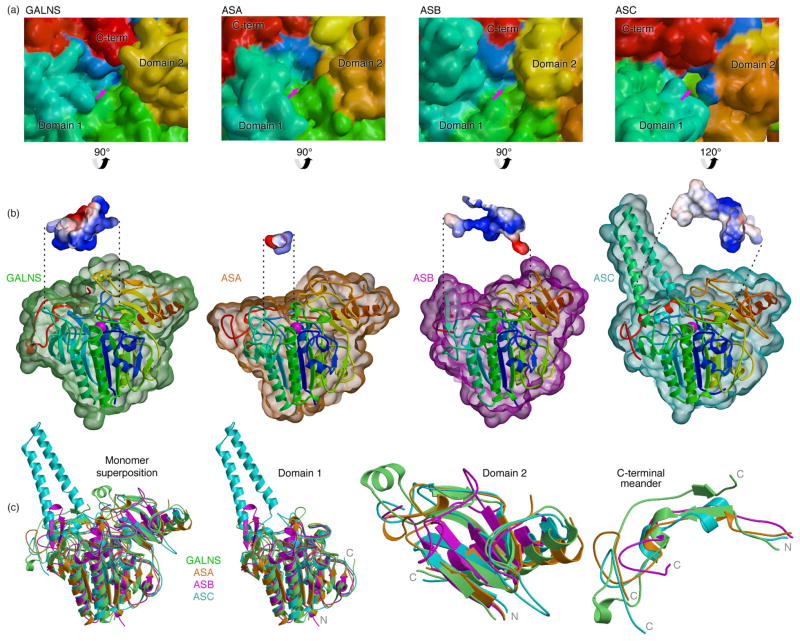
We compared the structure of GALNS to the most closely related paralogs with known structures, human ASA (PDB ID: 1AUK), ASB (PDB ID: 1FSU), and arylsulfatase C (ASC; PDB ID: 1P49; Figs. 5 and SF2). Overall GALNS has with a root-mean-square (RMS) deviation of 1.31 Å with ASA (for 408 Cαs, or 83% of the total), of 1.37 Å with ASB (339 Cαs, 72%), and of 1.24 Å with ASC (381 Cαs, 70%). Domains 1 and 2 superpose better than the divergent C-terminal meander (Fig. 5c). GALNS domain 1 superposes with an RMS deviation of 1.17 Å with ASA (318 Cαs, 90%), of 1.24 Å with ASB (291 Cαs, 84%), and of 1.06 Å with ASC (312 Cαs, 75%), and domain 2 superposes with an RMS deviation of 1.17 Å with ASA (73 Cαs, 75%), of 1.53 Å with ASB (60 Cαs, 58%), and of 1.15 Å with ASC (59 Cαs, 60%). Less than 40% of the residues in the small C-terminal meander superpose across the structures. The pairwise sequence identity32 across the structures is highest in domain 1, where it approaches 40%, decreases in domain 2 to 13–25%, and drops to undetectable in the C-terminal meander (Fig. SF2).
Fig. 5.
Comparison of GALNS to other human sulfatases
(a) Surface representations of GALNS (PDB ID 4FDI), ASA (PDB ID 1AUK), ASB (PDB ID 1FSU), and ASC (PDB ID 1P49), colored as in Fig. 1 and viewed into the active sites (indicated by arrows). (b) Surfaces of GALNS (green), ASA (orange), ASB (purple), and ASC (cyan) are shown around ribbon representation of the monomers (colored as in Fig. 1). The front surfaces are cropped to show the active site in the vicinity of the metal-binding site (magenta), and insets show the surface cavities (calculated in Pymol42) colored by electrostatic potentials as in Fig. 3. (c) Superposition of GALNS (green), ASA (orange), ASB (purple), and ASC (cyan) are separated into panels corresponding to domain 1, domain 2, and the C-terminal meander, with the termini of each chain indicated.
One remarkable feature of the GALNS structure is the participation of the extreme C-terminal segment of the C-terminal meander in defining the active-site pocket (Figs. 1 and 5). GALNS contains a novel conformation of its C-terminal meander, where the polypeptide reverses direction and forms a wall of the active site contributing to substrate selectivity. We compared the substrate selectivity in the sulfatase family by examining the active-site pockets of GALNS and the paralog structures. The active-site machinery in domain 1, including the requirement for a metal, the metal-binding residues, and the post-translationally modified nucleophile, is strictly conserved across the family.33 All of the family members have positively charged active-site pockets suitable for attracting sulfated substrates, but the overall sizes, shapes, and electrostatics of the pockets vary extensively (Fig. 5a and b). For example, GALNS and ASB (which cleave sulfates from polyanionic substrates like chondroitin sulfate) contain open, flat, positively charged active sites.12 In contrast, ASA (which cleaves sulfates off of sphingolipid sulfate esters solublized by saposin presentation molecules) contains a small active site complementary to the sulfate head group of the substrate.11 ASC (a membrane bound enzyme which cleaves sulfates from 3-β-hydroxysteroid sulfates) contains an active site with both charged and hydrophobic regions.26 In sulfatases, highly conserved catalytic machinery couples with diverse active site geometry, leading to cleavage of a wide variety of substrates.
MPS IV A disease mutations
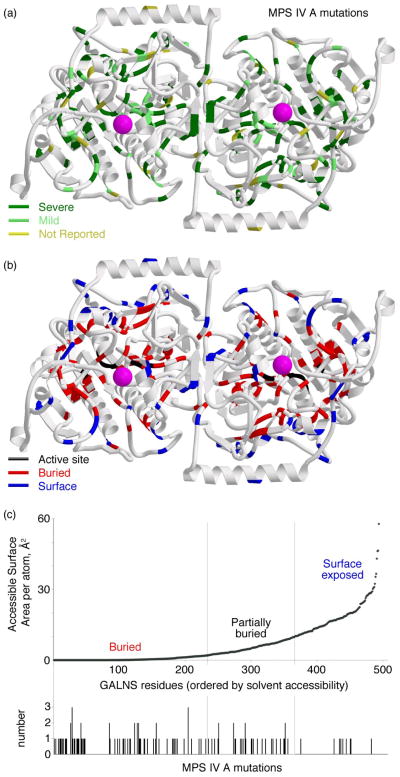
To gain further insight into the defects in GALNS that lead to disease, we mapped the locations of 120 disease-causing missense mutations found in MPS IV A patients (Fig. 6 and Table 2). Surprisingly, only 6 mutations (5%) affect active site residues, while 78 (65%) map to buried residues in the core of the protein and 32 mutations (27%) that map to surface residues. Domain 1 contains a larger number of mutations that map to buried residues (68) vs. surface residues (19). In contrast, mutations that map to domain 2 are found nearly equally in buried residues (9) and surface residues (10).
Fig. 6.
GALNS mutations in MPS IV A
(a): Disease-causing mutations leading to substitutions in the protein were mapped onto the GALNS structure. In the top panel, reported mutations are colored by their disease phenotype, as severe (less than 125cm in stature, dark green), mild (over 125cm, light green), or not reported (yellow). In the bottom panel, the mutations are mapped to their location on the protein, as near the active site (black), buried in the hydrophobic core (red), or on the surface (blue). (b): GALNS residues were ranked by solvent accessibility and vertical bars indicate a substitution leading to MPS IV A disease. Most of the mutations lead to changes in buried residues, meaning MPS IV A is most often a protein-folding disease.
Table 2.
Effect of Morquio A Disease-Causing Mutations
| Position | Mutation | Atomic acc. surf. | Effect on GALNS | Disease Phenotype | Mutation Category | Ref. |
|---|---|---|---|---|---|---|
| M1V | 1A>G | In signal sequence | Severe | Processing | 48 | |
| W10X | 29G>A | In signal sequence | Severe | Processing | 49 | |
| L15M | 43C>A | In signal sequence | Mild | Processing | 50 | |
| G23R | 67G>C | In signal sequence | Mild | Processing | 51 | |
| L36P | 107 T>C | 0.0 | Buried middle of in β strand; Pro would kink strand | Mild | Buried | 48 |
| M41L | 121 A>T | 0.2 | Little room for branched side chain | Mild | Buried | 3 |
| G42E | 125 G>A | 0.0 | No room for side chain | Severe | Buried | 50 |
| G47R | 139 G>A | 0.0 | No room for large side chain | Severe | Buried | 51 |
| S53F | 158 C>T | 3.8 | Polar region near dimer interface; little room for large side chain | NR | Surface | 50 |
| D60N | 178G>A | 7.5 | Unknown | Mild | Surface | 52 |
| R61W | 181C>T | 17.0 | Solvent exposed hydrophobic residue | Mild | Surface | 53 |
| G66R | 196G>C | 0.0 | No room for large side chain | Severe | Buried | 50 |
| F69V | 205T>G | 0.0 | Loss of hydrophobic packing | Severe | Buried | 48 |
| P77R | 230C>G | 0.0 | No room for large side chain | Severe | Buried | 54 |
| C79Y | 236G>A | 1.3 | Loss of nucleophile | Severe | Active Site | 55 |
| S80L | 239C>T | 0.0 | Buried in polar environment, possible disruption of formylglycine modification | Severe | Active Site | 49 |
| T88I | 263C>T | 0.0 | Loss of buried hydrogen bonding | NR | Buried | 56 |
| R90W | 268C>T | 3.4 | Partially buried; no room for Trp | Severe | Buried | 57 |
| R94G | 280C>G | 3.2 | Loss of ion pair to Glu121 | Mild | Buried | 50 |
| R94C | 280C>T | 3.2 | Loss of ion pair to Glu121 | Mild | Buried | 3 |
| R94L | 281G>T | 3.2 | Loss of ion pair to Glu121 | Severe | Buried | 10 |
| G96C | 286G>T | 0.1 | Gly-specific phi/psi angles | Severe | Buried | 50 |
| G96V | 287G>T | 0.1 | Gly-specific phi/psi angles | Severe | Buried | 3 |
| F97V | 289T>G | 0.0 | Loss of hydrophobic packing | Mild | Buried | 52 |
| A107T | 319G>A | 0.0 | No room for larger side chain | NR | Buried | 53 |
| Q111X | 331C>T | 0.7 | Stop | Severe | - | 49 |
| Q111R | 332A>G | 0.7 | Loss of buried hydrogen bonding | Mild | Buried | 58 |
| I113F | 337A>T | 0.0 | Buried; no room for larger side chain | Severe | Buried | 3 |
| G116S | 346G>A | 0.0 | Buried; no room for larger side chain | Severe | Buried | 57 |
| P125L | 374C>T | 0.0 | Buried; no room for larger side chain | Severe | Buried | 3 |
| K129X | 385A>T | 17.6 | Stop | Severe | - | 51 |
| S135R | 405C>G | 0.2 | Buried; no room for larger side chain | Severe | Buried | 59 |
| V138A | 413T>C | 0.0 | Loss of hydrophobic packing | Mild | Buried | 50 |
| G139S | 415G>A | 0.1 | Gly-specific phi/psi angles | Severe | Buried | 48 |
| W141R | 421T>A | 0.0 | Buried in hydrophobic core; near active site | Severe | Buried | 60 |
| W141C | 423G>T | 0.0 | Buried in hydrophobic core; near active site | Severe | Buried | 61 |
| H142R | 425A>G | 0.7 | Active site residue | Severe | Active Site | 3 |
| Q148X | 442C>T | 18.5 | Stop | Severe | - | 3 |
| H150Y | 448C>T | 1.9 | Buried; little room for larger side chain | Mild | Buried | 52 |
| P151S | 451C>T | 0.2 | Buried in hydrophobic core | Severe | Buried | 50 |
| P151L | 452C>T | 0.2 | Buried in hydrophobic core | Severe | Buried | 49 |
| G155R | 463G>A | 2.5 | Gly-specific phi/psi angles | Severe | Surface | 3 |
| G155E | 464G>A | 2.5 | Gly-specific phi/psi angles | Severe | Surface | 50 |
| F156S | 467T>C | 0.0 | Buried in hydrophobic core | Mild | Buried | 53 |
| F156C | 467T>G | 0.0 | Buried in hydrophobic core | Severe | Buried | 10 |
| W159C | 477G>T | 0.2 | Buried in hydrophobic core | Severe | Buried | 50 |
| S162F | 485C>T | 0.1 | Buried; no room for larger side chain | Severe | Buried | 3 |
| P163H | 488C>A | 0.6 | Buried; little room for larger side chain | Mild | Buried | 3 |
| N164T | 491A>C | 0.8 | Disruption of buried hydrogen bonds | NR | Buried | 62 |
| H166Q | 498C>G | 0.0 | Disruption of buried ion pair with Asp233 | Severe | Buried | 53 |
| F167V | 499T>G | 15.2 | Near active site; mutation may affect substrate recognition or protein binding | Mild | Surface | 55 |
| G168R | 502G>A | 1.6 | Gly-specific phi/psi angles | Severe | Buried | 50 |
| D171A | 512A>C | 16.7 | Unknown | Mild | Surface | 49 |
| P179S | 535C>T | 0.0 | Completely buried | Severe | Buried | 56 |
| P179H | 536C>A | 0.0 | Buried; little room for larger side chain | Severe | Buried | 60 |
| P179L | 536C>T | 0.0 | Buried; little room for larger side chain | Severe | Buried | 57 |
| E185G | 554A>G | 17.9 | Unknown | Severe | Surface | 50 |
| A203V | 608C>T | 0.0 | Buried; little room for branched side chain | NR | Buried | 57 |
| N204K | 612C>G | 6.0 | Glycosylation site | Mild | Surface | 49 |
| Q211X | 631C>T | 17.8 | Stop | Severe | - | 49 |
| W230G | 688T>G | 0.0 | Loss of hydrophobic packing | Severe | Buried | 60 |
| W230X | 689G>A | 0.0 | Stop | Severe | - | 3 |
| D233N | 697G>A | 0.3 | Disruption of buried ion pair with His166 | Severe | Buried | 50 |
| H236D | 706C>G | 4.7 | Active site | Mild | Active Site | 50 |
| V239F | 715G>T | 0.0 | Buried; no room for larger side chain | Severe | Buried | 60 |
| G247D | 740G>A | 18.0 | Unknown | Severe | Surface | 56 |
| R253W | 757C>T | 6.6 | Partially buried; little room for Trp | Mild | Surface | 57 |
| A257T | 769G>A | 0.0 | Buried; little room for branched side chain | Severe | Buried | 3 |
| R259Q | 776G>A | 9.8 | Unknown | Mild | Surface | 49 |
| E260D | 780G>C | 0.0 | Loss of buried hydrogen bonding | NR | Buried | 51 |
| F284V | 850T>G | 0.2 | Loss of hydrophobic packing | Mild | Buried | 51 |
| S287L | 860C>T | 0.0 | Loss of buried hydrogen bonding | Severe | Buried | 53 |
| N289S | 866A>G | 0.0 | Active site, metal binding residue | Mild | Active Site | 3 |
| G290S | 868G>A | 0.0 | Little room for larger side chain; near active site | Severe | Buried | 49 |
| A291T | 871G>A | 0.4 | Buried; little room for larger side chain | Severe | Buried | 49 |
| A291D | 872C>A | 0.4 | Buried; little room for larger side chain | Mild | Buried | 51 |
| S295F | 884C>T | 3.5 | Partly buried; little room for larger side chain | Mild | Surface | 3 |
| G301C | 901G>T | 3.7 | Buried; little room for larger side chain | Severe | Buried | 49 |
| L307P | 920T>C | 0.4 | Buried in hydrophobic pocket | Severe | Buried | 50 |
| G309R | 925G>A | 0.2 | Buried; no room for larger side chain | Severe | Buried | 52 |
| K310N | 930G>C | 4.5 | Active site | Mild | Active Site | 56 |
| T312A | 934A>G | 0.0 | Loss of hydrophobic packing | NR | Buried | 51 |
| T312S | 935C>G | 0.0 | Loss of hydrophobic packing | Mild | Buried | 50 |
| G316V | 947G>T | 0.1 | Buried; no room for larger side chain | NR | Buried | 60 |
| M318R | 953T>G | 0.0 | Buried; little room for larger side chain | Severe | Buried | 51 |
| A324E | 971C>A | 0.0 | Buried in hydrophobic core | Mild | Buried | 50 |
| W325X | 974G>A | 0.0 | Stop | Severe | - | 63 |
| W325C | 975G>T | 0.0 | Buried in hydrophobic pocket | NR | Buried | 52 |
| Q338X | 1012 C>T | 0.2 | Stop | Severe | - | 3 |
| G340D | 1019 G>A | 0.4 | Buried in hydrophobic pocket | Severe | Buried | 51 |
| S341R | 1023 C>A | 0.8 | Little room for larger side chain | Severe | Buried | 57 |
| M343L | 1027 A>T | 1.6 | Buried; little room for branched side chain | Severe | Buried | 64 |
| M343R | 1028 T>G | 1.6 | Buried; little room for larger side chain | Severe | Buried | 51 |
| D344N | 1030 G>A | 0.1 | Loss of buried ion pair with Arg380 | Severe | Buried | 52 |
| D344E | 1032 C>G | 0.1 | Disruption of buried ion pair with Arg380 | Severe | Buried | 48 |
| L345P | 1034 T>C | 0.7 | Little room for Pro side chain within helix | Severe | Buried | 48 |
| F346L | 1038 C>A | 0.0 | Loss of hydrophobic packing | Severe | Buried | 50 |
| A351V | 1052 C>T | 5.6 | Little room for branched side chain | Severe | Surface | 50 |
| L352P | 1055 T>C | 2.0 | Partly buried in hydrophobic pocket | Mild | Buried | 49 |
| P357L | 1070 C>T | 8.1 | Unknown | Severe | Surface | 3 |
| R361G | 1081 A>G | 6.6 | Ion pair with Asp360 | Severe | Surface | 48 |
| L366F | 1096 C>T | 7.1 | Unknown | NR | Surface | 53 |
| L369P | 1106 T>C | 3.6 | Little room for Pro side chain within helix | Severe | Surface | 3 |
| Q374X | 1120 C>T | 17.0 | Stop | Severe | - | 63 |
| R376Q | 1127 G>A | 16.6 | Unknown | Severe | Surface | 3 |
| R380T | 1139 G>C | 1.2 | Loss of buried ion pair with Asp344 | Severe | Buried | 56 |
| R380S | 1140 G>T | 1.2 | Loss of buried ion pair with Asp344 | Mild | Buried | 51 |
| R386C | 1156 C>T | 1.1 | Disruption of hydrophobic core | Severe | Buried | 48 |
| R386H | 1157 G>A | 1.1 | Disruption of hydrophobic core | Severe | Buried | 57 |
| D388N | 1162 G>A | 5.1 | Loss of ion pair with His103 | Mild | Surface | 57 |
| M391V | 1171 A>G | 0.0 | Disruption of hydrophobic core | Mild | Buried | 65 |
| A392V | 1175 C>T | 0.0 | Little room for larger side chain | NR | Buried | 3 |
| L395V | 1183 C>G | 4.1 | Unknown | Severe | Surface | 57 |
| L395P | 1184 T>C | 4.1 | Phi-psi angles not favored for Pro | NR | Surface | 48 |
| H398D | 1192 C>G | 2.5 | Loss of ion pair with Glu460 | Mild | Surface | 57 |
| H401Y | 1201 C>T | 0.1 | Loss of buried ion pair with Glu450 | NR | Buried | 53 |
| N407H | 1219 A>C | 0.4 | Disruption of buried hydrogen bonds | Severe | Buried | 52 |
| W409S | 1226 G>C | 7.5 | Unknown | Mild | Surface | 3 |
| G421E | 1262 G>A | 2.3 | Little room for larger side chain | Severe | Surface | 50 |
| Q422K | 1264 C>A | 4.6 | Disruption of hydrogen bonds | Severe | Surface | 50 |
| Q422X | 1264 C>T | 4.6 | Stop | Severe | - | 51 |
| E450V | 1349 A>T | 0.0 | Loss of buried ion pair with His401 | Severe | Buried | 62 |
| F452I | 1354 T>A | 7.7 | Loss of hydrophobic packing | Severe | Surface | 66 |
| F452L | 1354 T>C | 7.7 | Loss of hydrophobic packing | Severe | Surface | 53 |
| S470P | 1408 T>C | 24.3 | Little room for Pro side chain in helix | NR | Surface | 51 |
| Q473X | 1417 C>T | 20.2 | Stop | Severe | - | 50 |
| P484S | 1450 C>T | 4.1 | Unknown | Mild | Surface | 57 |
| N487S | 1460 A>G | 6.9 | Disruption of hydrogen bonds | Severe | Surface | 50 |
| V488M | 1462 G>A | 17.5 | Unknown | NR | Surface | 56 |
| M494V | 1480 A>G | 1.8 | Little room for branched side chain | NR | Buried | 49 |
Discussion
Many nearly identical carbohydrate-containing substrates appear in the lysosome. For example, glucosides and galactosides differ by a single chiral center (at the 4 position of the sugar ring), α- and β- sugars differ by a single chiral configuration at the anomeric center, etc. In response, lysosomal glycosidases have evolved exquisite specificity for their substrates, as they are able to distinguish among the many similar substrates differing by a single chiral center. In contrast, lysosomal sulfatases have greater promiscuity in their substrate recognition. The differences between the lysosomal glycosidases and sulfatases are manifested in different active site geometries: exoglycosidases have small active sites and make direct interactions with every functional group on the saccharide, while sulfatases have larger and deeper active sites that are capable of binding a wide variety of substrates. The GALNS structure suggests that the specificity of lysosomal sulfatases derives from a large number of interactions with the substrate, beyond the sulfated monosaccharide proximal to the catalytic nucleophile.
The active sites of sulfatases including GALNS are often broad trenches that accommodate a variety of potential substrates (including synthetic substrates such as 4-MU-S, which share only a sulfate group with endogenous substrates). However, the promiscuity of the sulfatases is limited in vivo, since the multiple sulfatases in the lysosome are non-redundant and loss of one sulfatase cannot be compensated by the activity of a related enzyme. For example, MPS IV A symptoms develop due to the loss of one sulfatase (GALNS), but the missing activity in patients is not compensated by the presence of other fully functional sulfatases like ASA and ASB, although all three sulfatases have overlapping specificity in vitro.
MPS IV A is primarily a disease of protein folding, because the majority of disease-causing mutations result in disruption of the hydrophobic core of the protein. Domain 1 appears to be a critical scaffold for the overall structure of GALNS, as 78% of the mutations in domain 1 map to the hydrophobic core of the domain. In contrast, 53% of the mutations in domain 2 map to surface residues, which are less likely to perturb the folding of GALNS. The prevalence of disease-causing mutations mapping to surface residues in domain 2 suggests that this domain may participate in required intermolecular contacts for correct protein function. Along this line, GALNS has been reported to participate in a lysosomal complex with the β-galactosidase, sialidase-1, and protective protein/cathepsin A (PPCA) proteins.
Human GALNS tolerates a variety of active site ligands. In addition to the cellular substrates keratan sulfate and chondroitin-6-sulfate, the active site can cleave synthetic substrates like 4-MU-Gal-6-S and 4-MU-S and can bind inhibitors including GalNAc and sulfate. Multiple sugar-binding possibilities occur within the GALNS active site, as revealed by catalytically productive binding of 4-MU-S, by non-productive binding of 4-MU-S (i.e. substrate inhibition), and by non-productive GalNAc binding (in the cocrystal structure). Other sulfatases including ASB show similar non-productive binding of monosaccharides.12 In GALNS, the key active-site interaction appears to be sulfate recognition, because sulfate acts as an effective inhibitor and 4-MU-S (lacking a sugar) is a competent substrate. Critical ionic interactions between GALNS and a sulfated ligand explain previous observations of GALNS inhibition by sulfate, phosphate, and chloride,31 and citrate binding in the unliganded crystal structure.
Pharmacological chaperone therapy, or the use of a small molecule to stabilize a lysosomal enzyme as it transits from the ER to the lysosome, has been proposed and tested for many lysosomal storage diseases. Pharmacological chaperones have been developed for lysosomal glycosidases, but not yet for lysosomal sulfatases. The highly specific substrate recognition performed by lysosomal glycosidases can be exploited to make tight-binding pharmacological chaperones from substrate and product analogs. However, the larger and less specific substrate-binding pockets found in lysosomal sulfatases make pharmacological chaperone design more complex. Additionally, the preference of sulfatases for anionic substrates could present pharmacokinetic problems. The structure of GALNS reported here reveals a large and convoluted substrate-binding pocket well suited to the rational design of ligands that might serve as pharmacological chaperones or mechanism-based inhibitors.
In enzyme replacement therapy for MPS IV A, recombinant enzyme is injected into patients to replace the missing GALNS enzymatic activity. In an effort to improve delivery of recombinant GALNS to the bones of MPS IV A patients, one proposed variant of GALNS contains six additional glutamate residues at the N terminus.36 The GALNS structure indicates that modification of the N terminus is likely to be well tolerated by the protein, but additions to the C terminus could perturb the substrate-binding properties of the variant.
Purification of GALNS from endogenous sources led to proteins that were processed into smaller fragments, thought to be a requirement for full enzymatic activity. The structure lends no support to the processing requirement for activity, because the insect-cell expressed GALNS is unprocessed and shows comparable enzymatic activity to GALNS purified from endogenous sources. It appears that endogenous GALNS may be occasionally nicked by a cellular protease, presumably at a site or sites between the 308-419 disulfide, because the fragments remain disulfide-linked.
In summary, the crystal structure reported here provides for the first time a molecular basis for accurately predicting the effect of a MPS IV A mutation on the GALNS protein. Additionally, the high-resolution structure reveals a large, convoluted active site highly suited to rational drug-design experiments. The human GALNS structure furthers our understanding of the molecular defects that underlie the family of inherited metabolic defects known as lysosomal storage disorders.
Materials and Methods
Molecular biology
Human GALNS cDNA (NCBI Sequence ID: NM_000512.4) was purchased from Open Biosystems. The open reading frame, including the 26-residue native signal sequence (for secretion into the media) and a C-terminal hexahistidine affinity tag, was amplified by PCR using Hot Start Phusion polymerase (NEB). For transfection into insect cells, the PCR product was gel purified, incubated with Taq polymerase to add 3′ A-overhangs, and cloned into the pIB/V5-His-TOPO TA vector per the manufacturer’s protocol (Invitrogen). Top 10 E. coli were transformed with the GALNS plasmid DNA; single clones were isolated, analyzed by restriction enzyme digestion, and sequenced.
Insect cell protein expression and purification
GALNS plasmid DNA was used to transfect Trichoplusia ni (High Five) insect cells. 72 h post transfection, 100 mg/mL blasticidin was added for 2 weeks to select for GALNS integration. Subsequently, cells at 50–60% confluence were diluted to 1 cell/well into 96-well plates containing 10 μg/mL blasticidin and then grown to confluence. Supernatants were tested by immunoblotting to assess GALNS expression, and the best-expressing monoclonal cell was amplified for large-scale suspension culture. Typically 5–10 L cultures were grown in multiple 3 L baffled Fernbach flasks (Corning). Cultures were inoculated at 5×105 cells/mL in fresh serum-free media and grown for 3 days until viability dropped below 95%. Cells were removed by centrifugation, and the supernatant clarified by centrifuging for 45 min at 4700g prior to concentration and exchange into Buffer A (50 mM sodium phosphate, pH 7.0, and 250 mM sodium chloride) by tangential flow filtration.
GALNS was purified by nickel affinity chromatography and cation exchange chromatography. Clarified supernatant was loaded into a 5 mL Ni-NTA column and eluted with a gradient of Buffer B (50 mM sodium phosphate, pH 7.0, 250 mM sodium chloride, 400 mM imidazole) over 70 column volumes on an FPLC at 4°C. Fractions with GALNS were pooled and exchanged into sodium acetate, pH 5.1 prior to loading onto a 5 mL SOURCE-S cation-exchange column (GE Healthcare) in Buffer C (20 mM sodium acetate, pH 5.1). Proteins were eluted with a linear gradient of Buffer D (20 mM sodium acetate, pH 5.1, and 1 M sodium chloride) over 100 column volumes at 4°C. Fractions containing GALNS were pooled, exchanged into 20 mM Bis–Tris, pH 6.0, and concentrated to 10 mg/ml for crystallography.
Crystallography
We crystallized fully glycosylated GALNS by vapor diffusion against 10–15% PEG 6000, 0.1M citric acid, pH 5.0. Despite extensive optimization efforts, all crystals grew as stacked plates, which necessitated mechanical separation. Harvested crystals were transferred to cryoprotectant buffer (20% PEG 6000, 0.1M citric acid, pH 5.0, and 25% glycerol) and cooled in liquid nitrogen. GalNAc ligand data were collected from crystals soaked in cryoprotectant supplemented with 50 mM GalNAc. Diffraction data were collected at beamline 24-ID-C at the Advanced Photon Source and at beamline X6A at Brookhaven National Lab. X-ray data were integrated and scaled using the HKL2000 software,38 and high mosaicity, poor spot shape, and anisotropy in the diffraction led to poor merging statistics. Several resolution limits were tested, and the reported limits were based upon the quality of the electron density maps. Molecular replacement in the CCP4 program AMoRe39 using protein atoms from the 34% identical human arylsulfatase A (PDB ID: 1N2K) led to an initial Rfactor of 51.0% with the next best solution at 53.3%. The structure was completed from repeated cycles of model building in Coot41 and refinement in REFMAC5.39 Alternate interpretations of DHA (as multiple cysteine conformations) and Ca2+ (as other ions) led to poorer fits to the electron density. Molecular figures were made in Pymol42 and Povscript.43 Sequence alignments were calculated using ClustalW44 and Blastp,45 and Ramachandran statistics using Rampage.46 Structural superpositions were calculated in LSQMAN32 using a distance cutoff of 3.5 Å for Cα positions.
Mass spectrometry
Liquid chromatography/mass spectrometry (LC/MS) was used to confirm the post-translational modification of GALNS residue 79. GALNS was denatured by addition of guanidinium HCl (GuHCl) and ethylenediaminetetraacetic acid (EDTA) to a final concentration of 50 μM GALNS, 6 M GuHCl, and 1 mM EDTA. The protein was then reduced in 20 mM dithiothreitol for 30 minutes and cysteines carboxymethylated in 20 mM iodoacetic acid for 30 minutes in the dark. The protein was exchanged into 100 mM Tris HCl, pH 8.0 on a NAP-5 desalting column prior to trypsin incubation for five hours with immobilized trypsin (Promega). After trypsin removal, GALNS peptides were injected into a NanoLC system (Dionex) for separation on a C18 reverse phase column and analysis by a QStar XL hybrid quadrupole time-of-flight mass spectrometer (ABI/Sciex). In parallel with LC/MS, the more abundant peptide ions were isolated and subjected to tandem mass spectrometry (MS/MS) for fragmentation by collision-activated dissociation (CAD). This method allowed peptide identification and measurement of the mass of the amino acid at position 79.
Kinetic assays
Kinetic assays measured the hydrolysis of the fluorescent reporter 4-MU from two sulfated substrates: 4-MU-S (Sigma) and 4-MU-Gal-6-S (Glycosynth and Toronto Research Chemicals). For the 4-MU-S assays, 4-MU release was measured directly by fluorescence excitation at 362 nm and emission at 448 nm. The starting GALNS stock (calculated using an extinction coefficient of 1.6 mg/ml protein/OD280) contained 2.4 μM GALNS in 10 mM Bis-Tris, 10 mM sodium chloride, pH 6.0. The enzyme stock was diluted 8-fold into 100 mM sodium acetate buffer pH 4.5 at eight 4-MU-S substrate concentrations from 0.25 to 15 mM. Aliquots were taken every 5 minutes for 30 minutes and diluted into 250 mM glycine, pH 10.2 prior to reading 4-MU fluorescence in a microplate reader.
For the 4-MU-Gal-6-S reactions, cleavage of substrate was monitored with a coupled reaction where GALNS first removed the sulfate and then β-galactosidase hydrolyzed the glycosidic linkage, releasing the fluorescent product 4-MU. GALNS was diluted to 0.3 μM and incubated in 100 mM sodium acetate, pH 4.5 at eight substrate concentrations from 0.50 to 50 mM. Aliquots were diluted 11-fold into 0.1 mg/ml E. coli β-galactosidase (Sigma) in 100 mM sodium potassium phosphate, pH 6.2 for 30 minutes. Aliquots were then diluted 30-fold into 250 mM glycine, pH 10.2, prior to reading in a microplate reader. Control experiments lacking either of the two enzymes showed no fluorescence above background. Fluorescence of free 4-MU at 448 nm was converted to 4-MU concentration using a standard curve. Error bars represent the standard deviations of triplicate measurements. Velocities were calculated from linear fits of [4-MU] vs. time plots, and Michaelis-Menten hyperbolae were fit in KaleidaGraph to obtain KM and kcat. The 4-MU-S data showed substrate inhibition, so KM, Ki, and kcat fit better to a modified Michaelis-Menten function47:
where V0 is the initial velocity, [E]T and [S] are concentrations of enzyme and substrate, respectively, and Ki is the equilibrium constant for non-productive binding of substrate.
The inhibition assay used sub-inhibitory concentrations of 4-MU-S substrate. 0.3 μM GALNS, 1.2 mM 4-MU-S, and 100 mM sodium acetate, pH 4.5 were combined with 0 to 10 mM ammonium sulfate. Samples were taken every 5 minutes for 25 minutes and diluted 30-fold into 250 mM glycine, pH 10.2 prior to fluorimetry. The IC50 was fit in KaleidaGraph from the sigmoidal dose-response curve according to
where b is the Hill slope. The IC50 was converted into the Ki for a competitive inhibitor47 according to
Mutation analysis
Accessible surfaces areas were calculated in the CCP4 program AREAIMOL,39 and the average accessible surface area per side chain atom was calculated in Excel (Microsoft). Non-glycine side chains were considered buried if the average accessible surface area per side chain atom was less than 2 Å2.
Supplementary Material
Highlights.
We have determined the crystal structure of human GALNS
The structure reveals the molecular basis of MPS IV A (Morquio A) disease
The structure shows post-translational modification of an active site cysteine
The structure reveals that MPS IV A (Morquio A) is a protein-folding disease
Human GALNS is a promising target for rational drug design
Acknowledgments
This work was funded by NIH grant R01 DK76877 to S.C.G., by NIH T32 GM008515 to Y.R.-C., and by an HHMI summer research internship to E.K.S. We gratefully acknowledge Igor Kaltashov for assistance with mass spectrometry, and Matthew C. Metcalf and Elih M. Velázquez-Delgado for assistance with X-ray experiments. We thank Jean Jankonic, Marc Allaire, and Vivian Stojanoff at the National Synchrotron Light Source X6A beam line, funded by the National Institute of General Medical Sciences, National Institute of Health under agreement GM-0080. We thank Igor Kourinov and the staff of the Advanced Photon Source Northeastern Collaborative Access Team beamlines, which are supported by grants from the National Center for Research Resources (5P41RR015301-10) and the National Institute of General Medical Sciences (8 P41 GM103403-10) from the National Institutes of Health.
Footnotes
Accession Numbers
Coordinates and structure factors have been deposited in the Protein Data Bank with accession numbers 4FDI and 4FDJ.
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- 1.Neufeld EF, Muenzer J. The Mucopolysaccharidoses. In: Scriver CR, Beaudet AL, Sly WS, Valle D, editors. The Metabolic and Molecular Bases of Inherited Disease. 8. McGraw-Hill; New York: 2001. pp. 3421–3452. [Google Scholar]
- 2.Northover H, Cowie RA, Wraith JE. Mucopolysaccharidosis type IVA (Morquio syndrome): a clinical review. J Inherit Metab Dis. 1996;19:357–65. doi: 10.1007/BF01799267. [DOI] [PubMed] [Google Scholar]
- 3.Tomatsu S, Montano AM, Nishioka T, Gutierrez MA, Pena OM, Tranda Firescu GG, Lopez P, Yamaguchi S, Noguchi A, Orii T. Mutation and polymorphism spectrum of the GALNS gene in mucopolysaccharidosis IVA (Morquio A) Hum Mutat. 2005;26:500–12. doi: 10.1002/humu.20257. [DOI] [PubMed] [Google Scholar]
- 4.Martell L, Lau K, Mei M, Burnett V, Decker C, Foehr ED. Biomarker analysis of Morquio syndrome: identification of disease state and drug responsive markers. Orphanet J Rare Dis. 2011;6:84. doi: 10.1186/1750-1172-6-84. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Glössl J, Truppe W, Kresse H. Purification and properties of N-acetylgalactosamine 6-sulphate sulphatase from human placenta. Biochem J. 1979;181:37–46. doi: 10.1042/bj1810037. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Bielicki J, Hopwood JJ. Human liver N-acetylgalactosamine 6-sulphatase. Purification and characterization. Biochem J. 1991;279 (Pt 2):515–20. doi: 10.1042/bj2790515. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Glössl J, Maroteaux P, Di Natale P, Kresse H. Different properties of residual N-acetylgalactosamine-6-sulfate sulfatase in fibroblasts from patients with mild and severe forms of Morquio disease type A. Pediatr Res. 1981;15:976–978. [Google Scholar]
- 8.Glössl J, Kresse H. Impaired degradation of keratan sulphate by Morquio A fibroblasts. Biochem J. 1982;203:335–8. doi: 10.1042/bj2030335. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Dvorak-Ewell M, Wendt D, Hague C, Christianson T, Koppaka V, Crippen D, Kakkis E, Vellard M. Enzyme replacement in a human model of mucopolysaccharidosis IVA in vitro and its biodistribution in the cartilage of wild type mice. PLoS One. 2010;5:e12194. doi: 10.1371/journal.pone.0012194. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Sukegawa K, Nakamura H, Kato Z, Tomatsu S, Montano AM, Fukao T, Toietta G, Tortora P, Orii T, Kondo N. Biochemical and structural analysis of missense mutations in N-acetylgalactosamine-6-sulfate sulfatase causing mucopolysaccharidosis IVA phenotypes. Hum Mol Genet. 2000;9:1283–90. doi: 10.1093/hmg/9.9.1283. [DOI] [PubMed] [Google Scholar]
- 11.Lukatela G, Krauss N, Theis K, Selmer T, Gieselmann V, von Figura K, Saenger W. Crystal structure of human arylsulfatase A: the aldehyde function and the metal ion at the active site suggest a novel mechanism for sulfate ester hydrolysis. Biochemistry. 1998;37:3654–64. doi: 10.1021/bi9714924. [DOI] [PubMed] [Google Scholar]
- 12.Bond CS, Clements PR, Ashby SJ, Collyer CA, Harrop SJ, Hopwood JJ, Guss JM. Structure of a human lysosomal sulfatase. Structure. 1997;5:277–89. doi: 10.1016/s0969-2126(97)00185-8. [DOI] [PubMed] [Google Scholar]
- 13.Cosma MP, Pepe S, Annunziata I, Newbold RF, Grompe M, Parenti G, Ballabio A. The multiple sulfatase deficiency gene encodes an essential and limiting factor for the activity of sulfatases. Cell. 2003;113:445–56. doi: 10.1016/s0092-8674(03)00348-9. [DOI] [PubMed] [Google Scholar]
- 14.Dierks T, Schmidt B, Borissenko LV, Peng J, Preusser A, Mariappan M, von Figura K. Multiple sulfatase deficiency is caused by mutations in the gene encoding the human Ca-formylglycine generating enzyme. Cell. 2003;113:435–44. doi: 10.1016/s0092-8674(03)00347-7. [DOI] [PubMed] [Google Scholar]
- 15.Dierks T, Schlotawa L, Frese MA, Radhakrishnan K, von Figura K, Schmidt B. Molecular basis of multiple sulfatase deficiency, mucolipidosis II/III and Niemann-Pick C1 disease - Lysosomal storage disorders caused by defects of non-lysosomal proteins. Biochim Biophys Acta. 2009;1793:710–25. doi: 10.1016/j.bbamcr.2008.11.015. [DOI] [PubMed] [Google Scholar]
- 16.Bojarova P, Williams SJ. Sulfotransferases, sulfatases and formylglycine-generating enzymes: a sulfation fascination. Curr Opin Chem Biol. 2008;12:573–81. doi: 10.1016/j.cbpa.2008.06.018. [DOI] [PubMed] [Google Scholar]
- 17.Rush JS, Bertozzi CR. New aldehyde tag sequences identified by screening formylglycine generating enzymes in vitro and in vivo. J Am Chem Soc. 2008;130:12240–1. doi: 10.1021/ja804530w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Rabuka D, Rush JS, Dehart GW, Wu P, Bertozzi CR. Site-specific chemical protein conjugation using genetically encoded aldehyde tags. Nat Protoc. 2012;7:1052–67. doi: 10.1038/nprot.2012.045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Roeser D, Preusser-Kunze A, Schmidt B, Gasow K, Wittmann JG, Dierks T, von Figura K, Rudolph MG. A general binding mechanism for all human sulfatases by the formylglycine-generating enzyme. Proc Natl Acad Sci U S A. 2006;103:81–6. doi: 10.1073/pnas.0507592102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.von Figura K, Schmidt B, Selmer T, Dierks T. A novel protein modification generating an aldehyde group in sulfatases: its role in catalysis and disease. Bioessays. 1998;20:505–10. doi: 10.1002/(SICI)1521-1878(199806)20:6<505::AID-BIES9>3.0.CO;2-K. [DOI] [PubMed] [Google Scholar]
- 21.Ghosh D. Human sulfatases: a structural perspective to catalysis. Cell Mol Life Sci. 2007;64:2013–22. doi: 10.1007/s00018-007-7175-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Hanson SR, Best MD, Wong CH. Sulfatases: structure, mechanism, biological activity, inhibition, and synthetic utility. Angew Chem Int Ed Engl. 2004;43:5736–63. doi: 10.1002/anie.200300632. [DOI] [PubMed] [Google Scholar]
- 23.Boltes I, Czapinska H, Kahnert A, von Bulow R, Dierks T, Schmidt B, von Figura K, Kertesz MA, Uson I. 1.3 A structure of arylsulfatase from Pseudomonas aeruginosa establishes the catalytic mechanism of sulfate ester cleavage in the sulfatase family. Structure. 2001;9:483–91. doi: 10.1016/s0969-2126(01)00609-8. [DOI] [PubMed] [Google Scholar]
- 24.Tomatsu S, Montano AM, Gutierrez M, Grubb JH, Oikawa H, Dung VC, Ohashi A, Nishioka T, Yamada M, Tosaka Y, Trandafirescu GG, Orii T. Characterization and pharmacokinetic study of recombinant human N-acetylgalactosamine-6-sulfate sulfatase. Mol Genet Metab. 2007;91:69–78. doi: 10.1016/j.ymgme.2007.01.004. [DOI] [PubMed] [Google Scholar]
- 25.von Figura K, Gieselmann V, Jaeken J. Metachromatic leukodystrophy. In: Scriver CR, Beaudet AL, Sly WS, Valle D, editors. The Metabolic and Molecular Bases of Inherited Disease. 8. McGraw-Hill; New York: 2001. pp. 3695–3724. [Google Scholar]
- 26.Hernandez-Guzman FG, Higashiyama T, Pangborn W, Osawa Y, Ghosh D. Structure of human estrone sulfatase suggests functional roles of membrane association. J Biol Chem. 2003;278:22989–97. doi: 10.1074/jbc.M211497200. [DOI] [PubMed] [Google Scholar]
- 27.Brown ID. Recent developments in the methods and applications of the bond valence model. Chem Rev. 2009;109:6858–6919. doi: 10.1021/cr900053k. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Weis WI, Drickamer K. Structural basis of lectin-carbohydrate recognition. Annu Rev Biochem. 1996;65:441–73. doi: 10.1146/annurev.bi.65.070196.002301. [DOI] [PubMed] [Google Scholar]
- 29.Elgavish S, Shaanan B. Lectin-carbohydrate interactions: different folds, common recognition principles. Trends Biochem Sci. 1997;22:462–7. doi: 10.1016/s0968-0004(97)01146-8. [DOI] [PubMed] [Google Scholar]
- 30.van Diggelen OP, Zhao H, Kleijer WJ, Janse HC, Poorthuis BJ, van Pelt J, Kamerling JP, Galjaard H. A fluorimetric enzyme assay for the diagnoisis of Morquio disease type A (MPS IV A) Clinica Chimica Acta. 1990;187:131–139. doi: 10.1016/0009-8981(90)90339-t. [DOI] [PubMed] [Google Scholar]
- 31.Bielicki J, Fuller M, Guo XH, Morris CP, Hopewood JJ, Anson DS. Expression, purification and characterization of recombinant human N-acetylgalactosamine-6-sulphatase. Biochem J. 1995;311 (Pt 1):333–9. doi: 10.1042/bj3110333. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Kleywegt GJ, Jones TA. Halloween...masks and bones. In: Bailey S, Hubbard R, Waller DA, editors. Proceedings of the CCP4 Study Weekend. From First Map to Final Model. SERC Daresbury Laboratory; Warrington: 1994. pp. 59–66. [Google Scholar]
- 33.Ghosh D. Three-dimensional structures of sulfatases. Methods Enzymol. 2005;400:273–93. doi: 10.1016/S0076-6879(05)00016-9. [DOI] [PubMed] [Google Scholar]
- 34.Pshezhetsky AV, Ashmarina M. Lysosomal multienzyme complex: biochemistry, genetics, and molecular pathophysiology. Prog Nucleic Acid Res Mol Biol. 2001;69:81–114. doi: 10.1016/s0079-6603(01)69045-7. [DOI] [PubMed] [Google Scholar]
- 35.Pshezhetsky AV, Potier M. Association of N-acetylgalactosamine-6-sulfate sulfatase with the multienzyme lysosomal complex of β-galactosidase, cathepsin A, and neuraminidase. Possible implication for intralysosomal catabolism of keratan sulfate. J Biol Chem. 1996;271:28359–65. doi: 10.1074/jbc.271.45.28359. [DOI] [PubMed] [Google Scholar]
- 36.Tomatsu S, Montano AM, Dung VC, Ohashi A, Oikawa H, Oguma T, Orii T, Barrera L, Sly WS. Enhancement of drug delivery: enzyme-replacement therapy for murine Morquio A syndrome. Mol Ther. 2010;18:1094–102. doi: 10.1038/mt.2010.32. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Masue M, Sukegawa K, Orii T, Hashimoto T. N-acetylgalactosamine-6-sulfate sulfatase in human placenta: purification and characteristics. J Biochem. 1991;110:965–70. doi: 10.1093/oxfordjournals.jbchem.a123697. [DOI] [PubMed] [Google Scholar]
- 38.Otwinowski Z, Minor W. Processing of X-ray diffraction data collected in oscillation mode. In: Carter CW, Sweet RM, editors. Methods in Enzymology: Macromolecular Crystallography, part A. Vol. 276. Academic Press; 1997. pp. 307–326. [DOI] [PubMed] [Google Scholar]
- 39.Collaborative Computational Project, N. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. 1994;D50:760–763. doi: 10.1107/S0907444994003112. [DOI] [PubMed] [Google Scholar]
- 40.Chruszcz M, Laidler P, Monkiewicz M, Ortlund E, Lebioda L, Lewinski K. Crystal structure of a covalent intermediate of endogenous human arylsulfatase A. J Inorg Biochem. 2003;96:386–92. doi: 10.1016/s0162-0134(03)00176-4. [DOI] [PubMed] [Google Scholar]
- 41.Emsley P, Cowtan K. Coot: model-building tools for molecular graphics. Acta Crystallogr D Biol Crystallogr. 2004;60:2126–32. doi: 10.1107/S0907444904019158. [DOI] [PubMed] [Google Scholar]
- 42.DeLano WL. The PyMOL Molecular Graphics System. DeLano Scientific LLC; Palo Alto, CA, USA: 2008. [Google Scholar]
- 43.Fenn TD, Ringe D, Petsko GA. POVScript+: a program for model and data visualization using persistence of vision ray-tracing. J Appl Cryst. 2003;36:944–947. [Google Scholar]
- 44.Thompson JD, Higgins DG, Gibson TJ. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994;22:4673–80. doi: 10.1093/nar/22.22.4673. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 1997;25:3389–402. doi: 10.1093/nar/25.17.3389. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Lovell SC, Davis IW, Arendall WB, 3rd, de Bakker PI, Word JM, Prisant MG, Richardson JS, Richardson DC. Structure validation by Cα geometry: ϕ,ψ and Cβ deviation. Proteins. 2003;50:437–50. doi: 10.1002/prot.10286. [DOI] [PubMed] [Google Scholar]
- 47.Copeland RA. Enzymes: A Practical Introduction to Structure, Mechanism, and Data Analysis. 2. Wiley-VCH; New York: 2000. [Google Scholar]
- 48.Tomatsu S, Filocamo M, Orii KO, Sly WS, Gutierrez MA, Nishioka T, Serrato OP, Di Natale P, Montano AM, Yamaguchi S, Kondo N, Orii T, Noguchi A. Mucopolysaccharidosis IVA (Morquio A): identification of novel common mutations in the N-acetylgalactosamine-6-sulfate sulfatase (GALNS) gene in Italian patients. Hum Mutat. 2004;24:187–8. doi: 10.1002/humu.9265. [DOI] [PubMed] [Google Scholar]
- 49.Wang Z, Zhang W, Wang Y, Meng Y, Su L, Shi H, Huang S. Mucopolysaccharidosis IVA mutations in Chinese patients: 16 novel mutations. J Hum Genet. 2010;55:534–40. doi: 10.1038/jhg.2010.65. [DOI] [PubMed] [Google Scholar]
- 50.Bunge S, Kleijer WJ, Tylki-Szymanska A, Steglich C, Beck M, Tomatsu S, Fukuda S, Poorthuis BJ, Czartoryska B, Orii T, Gal A. Identification of 31 novel mutations in the N-acetylgalactosamine-6-sulfatase gene reveals excessive allelic heterogeneity among patients with Morquio A syndrome. Hum Mutat. 1997;10:223–32. doi: 10.1002/(SICI)1098-1004(1997)10:3<223::AID-HUMU8>3.0.CO;2-J. [DOI] [PubMed] [Google Scholar]
- 51.Ogawa T, Tomatsu S, Fukuda S, Yamagishi A, Rezvi GM, Sukegawa K, Kondo N, Suzuki Y, Shimozawa N, Oru T. Mucopolysaccharidosis IVA: screening and identification of mutations of the N-acetylgalactosamine-6-sulfate sulfatase gene. Hum Mol Genet. 1995;4:341–9. doi: 10.1093/hmg/4.3.341. [DOI] [PubMed] [Google Scholar]
- 52.Tomatsu S, Nishioka T, Montano AM, Gutierrez MA, Pena OS, Orii KO, Sly WS, Yamaguchi S, Orii T, Paschke E, Kircher SG, Noguchi A. Mucopolysaccharidosis IVA: identification of mutations and methylation study in GALNS gene. J Med Genet. 2004;41:e98. doi: 10.1136/jmg.2003.018010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Yamada N, Fukuda S, Tomatsu S, Muller V, Hopwood JJ, Nelson J, Kato Z, Yamagishi A, Sukegawa K, Kondo N, Orii T. Molecular heterogeneity in mucopolysaccharidosis IVA in Australia and Northern Ireland: nine novel mutations including T312S, a common allele that confers a mild phenotype. Hum Mutat. 1998;11:202–8. doi: 10.1002/(SICI)1098-1004(1998)11:3<202::AID-HUMU4>3.0.CO;2-J. [DOI] [PubMed] [Google Scholar]
- 54.Qubbaj W, Al-Aqeel AI, Al-Hassnan Z, Al-Duraihim A, Awartani K, Al-Rejjal R, Coskun S. Preimplantation genetic diagnosis of Morquio disease. Prenat Diagn. 2008;28:900–3. doi: 10.1002/pd.2081. [DOI] [PubMed] [Google Scholar]
- 55.Kato Z, Fukuda S, Tomatsu S, Vega H, Yasunaga T, Yamagishi A, Yamada N, Valencia A, Barrera LA, Sukegawa K, Orii T, Kondo N. A novel common missense mutation G301C in the N-acetylgalactosamine-6-sulfate sulfatase gene in mucopolysaccharidosis IVA. Hum Genet. 1997;101:97–101. doi: 10.1007/s004390050594. [DOI] [PubMed] [Google Scholar]
- 56.Tomatsu S, Dieter T, Schwartz IV, Sarmient P, Giugliani R, Barrera LA, Guelbert N, Kremer R, Repetto GM, Gutierrez MA, Nishioka T, Serrato OP, Montano AM, Yamaguchi S, Noguchi A. Identification of a common mutation in mucopolysaccharidosis IVA: correlation among genotype, phenotype, and keratan sulfate. J Hum Genet. 2004;49:490–4. doi: 10.1007/s10038-004-0178-8. [DOI] [PubMed] [Google Scholar]
- 57.Tomatsu S, Fukuda S, Cooper A, Wraith JE, Ferreira P, Di Natale P, Tortora P, Fujimoto A, Kato Z, Yamada N, Isogai K, Yamagishi A, Sukegawa K, Suzuki Y, Shimozawa N, Kondo N, Sly WS, Orii T. Fourteen novel mucopolysaccharidosis IVA producing mutations in GALNS gene. Hum Mutat. 1997;10:368–75. doi: 10.1002/(SICI)1098-1004(1997)10:5<368::AID-HUMU6>3.0.CO;2-B. [DOI] [PubMed] [Google Scholar]
- 58.Terzioglu M, Tokatli A, Coskun T, Emre S. Molecular analysis of Turkish mucopolysaccharidosis IVA (Morquio A) patients: identification of novel mutations in the N-acetylgalactosamine-6-sulfate sulfatase (GALNS) gene. Hum Mutat. 2002;20:477–8. doi: 10.1002/humu.9088. [DOI] [PubMed] [Google Scholar]
- 59.Fukuda S, Tomatsu S, Masue M, Sukegawa K, Iwata H, Ogawa T, Nakashima Y, Hori T, Yamagishi A, Hanyu Y, et al. Mucopolysaccharidosis type IVA. N-acetylgalactosamine-6-sulfate sulfatase exonic point mutations in classical Morquio and mild cases. J Clin Invest. 1992;90:1049–53. doi: 10.1172/JCI115919. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Tomatsu S, Fukuda S, Cooper A, Wraith JE, Rezvi GM, Yamagishi A, Yamada N, Kato Z, Isogai K, Sukegawa K, et al. Mucopolysaccharidosis type IVA: identification of six novel mutations among non-Japanese patients. Hum Mol Genet. 1995;4:741–3. doi: 10.1093/hmg/4.4.741. [DOI] [PubMed] [Google Scholar]
- 61.Montano AM, Kaitila I, Sukegawa K, Tomatsu S, Kato Z, Nakamura H, Fukuda S, Orii T, Kondo N. Mucopolysaccharidosis IVA: characterization of a common mutation found in Finnish patients with attenuated phenotype. Hum Genet. 2003;113:162–9. doi: 10.1007/s00439-003-0959-8. [DOI] [PubMed] [Google Scholar]
- 62.Laradi S, Tukel T, Khediri S, Shabbeer J, Erazo M, Chkioua L, Chaabouni M, Ferchichi S, Miled A, Desnick RJ. Mucopolysaccharidosis type IV: N-acetylgalactosamine-6-sulfatase mutations in Tunisian patients. Mol Genet Metab. 2006;87:213–8. doi: 10.1016/j.ymgme.2005.11.001. [DOI] [PubMed] [Google Scholar]
- 63.Tomatsu S, Fukuda S, Cooper A, Wraith JE, Yamada N, Isogai K, Kato Z, Sukegawa K, Kondo N, Suzuki Y, et al. Two new mutations, Q473X and N487S, in a Caucasian patient with mucopolysaccharidosis IVA (Morquio disease) Hum Mutat. 1995;6:195–6. doi: 10.1002/humu.1380060218. [DOI] [PubMed] [Google Scholar]
- 64.Cole DE, Fukuda S, Gordon BA, Rip JW, LeCouteur AN, Rupar CA, Tomatsu S, Ogawa T, Sukegawa K, Orii T. Heteroallelic missense mutations of the galactosamine-6-sulfate sulfatase (GALNS) gene in a mild form of Morquio disease (MPS IVA) Am J Med Genet. 1996;63:558–65. doi: 10.1002/(SICI)1096-8628(19960628)63:4<558::AID-AJMG9>3.0.CO;2-K. [DOI] [PubMed] [Google Scholar]
- 65.Tomatsu S, Fukuda S, Cooper A, Wraith JE, Rezvi GM, Yamagishi A, Yamada N, Kato Z, Isogai K, Sukegawa K, et al. Mucopolysaccharidosis IVA: identification of a common missense mutation I113F in the N-Acetylgalactosamine-6-sulfate sulfatase gene. Am J Hum Genet. 1995;57:556–63. [PMC free article] [PubMed] [Google Scholar]
- 66.Carraresi L, Parini R, Filoni C, Caciotti A, Sersale G, Tomatsu S, Orlando C, Zammarchi E, Guerrini R, Donati MA, Morrone A. GALNS gene expression profiling in Morquio A patients’ fibroblasts. Clin Chim Acta. 2008;397:72–6. doi: 10.1016/j.cca.2008.07.021. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.