Abstract
Malignant pleural mesothelioma (MPM) is a highly aggressive neoplasm arising from the mesothelial cells lining the parietal pleura and it exhibits poor prognosis. Although there has been significant progress in MPM treatment, development of more efficient therapeutic approaches is needed. BMAL1 is a core component of the circadian clock machinery and its constitutive overexpression in MPM has been reported. Here, we demonstrate that BMAL1 may serve as a molecular target for MPM. The majority of MPM cell lines and a subset of MPM clinical specimens expressed higher levels of BMAL1 compared to a nontumorigenic mesothelial cell line (MeT-5A) and normal parietal pleural specimens, respectively. A serum shock induced a rhythmical BMAL1 expression change in MeT-5A but not in ACC-MESO-1, suggesting that the circadian rhythm pathway is deregulated in MPM cells. BMAL1 knockdown suppressed proliferation and anchorage-dependent and independent clonal growth in two MPM cell lines (ACC-MESO-1 and H290) but not in MeT-5A. Notably, BMAL1 depletion resulted in cell cycle disruption with a substantial increase in apoptotic and polyploidy cell population in association with downregulation of Wee1, cyclin B and p21WAF1/CIP1 and upregulation of cyclin E expression. BMAL1 knockdown induced mitotic catastrophe as denoted by disruption of cell cycle regulators and induction of drastic morphological changes including micronucleation and multiple nuclei in ACC-MESO-1 cells that expressed the highest level of BMAL1. Taken together, these findings indicate that BMAL1 has a critical role in MPM and could serve as an attractive therapeutic target for MPM.
Keywords: apoptosis, BMAL1, mesothelioma, targeted therapy, mitotic catastrophe
Malignant pleural mesothelioma (MPM) is a highly aggressive neoplasm arising from the mesothelial or submesothelial cells lining the parietal pleura. MPM has a particularly poor prognosis with a median survival of approximately 12 months from the onset of diagnosis.1 Asbestos exposure is considered the most important etiologic factor that has been mentioned in relation to MPM.2 Although a significant progress in MPM treatment has been achieved, there is an urgent need for developing new therapeutic approaches to improve the clinical outcome of patients with MPM.3
Many physiological, biological and metabolic processes are controlled by a homeostatic system called the circadian clock. This system is regulated by a circadian pacemaker located in the suprachiasmatic nucleus of the anterior hypothalamus that controls peripheral clocks over the 24 hr.4 Recent studies have shown that peripheral tissues also have advanced molecular mechanisms that regulate circadian events, many of which have also been observed in established cell lines.5–7
Several circadian clock genes have been reported to control circadian rhythms in peripheral tissues, including three period proteins (PER1, PER2 and PER3), two cryptochromes (CRY1 and CRY2), CLOCK, NPAS2 and BMAL proteins. BMAL1 is an indispensible core component in the circadian clock machinery. It can form heterodimer complexes with CLOCK or NPAS2 genes; this complex drives transcription from E-box elements found in the promoters of circadian-responsive genes.8 Period and cryptochrome proteins negatively regulate CLOCK/BMAL1 dimer-mediated transcription, thereby forming the feedback loop that regulates the timing of clock gene transcription.9
Disruption of the circadian clock has been associated with a wide variety of human disorders including cancer.10 Previous studies have shown that clock genes are involved in the pathogenesis of human cancers. These genes seem to function primarily as tumor suppressors.11 Several studies have reported the involvement of BMAL1 in human cancers. High BMAL1 expression was associated with poor patients’ prognosis and distant metastasis in colorectal and breast cancer.12,13 In addition, vascular endothelial growth factor is transcriptionally upregulated by BMAL1.14 These reports suggest an oncogenic role for BMAL1. By contrast, Taniguchi et al.4 showed that BMAL1 expression is inactivated by promoter methylation in hematologic malignancies but not in solid cancers and that exogenously overexpressed BMAL1 suppresses in vitro and in vivo growth of a lymphoma cell line, indicating a tumor suppressive role of BMAL1 that may be specific for hematologic malignancies.
Recently, expression microarray analysis of MPM showed overexpression of several circadian rhythm genes compared to normal parietal pleural. Specifically, the BMAL1 transcript was found to be overexpressed in MPM whereas negative regulators of BMAL1 were expressed at lower levels. These findings raise the possibility that BMAL1 could contribute to the aggressive malignant phenotypes of MPM.15 To the best of our knowledge, no prior studies have analyzed the functional roles of BMAL1 in MPM and thus we sought to investigate the role of BMAL1 in the pathogenesis of MPM and its potential utility as a therapeutic target for MPM.
Material and Methods
Cell lines and tissue culture
Thirteen MPM cell lines and a nontumorgenic mesothelial cell line (MeT-5A) were used in this study. We purchased H2452, H2052, MSTO-211H, H28 and MeT-5A cell lines from the American Type Culture Collection and confirmed their authenticity by short tandem repeat (STR) analysis. H290 and H2373 were gifts from Dr Adi F. Gazdar (University of Texas Southwestern Medical Center, Dallas, TX). ACC-MESO-1, Y-MESO-12, Y-MESO-9, ACC-MESO-4, Y-MESO-22 (epithelioid) Y-MESO-14 (biphasic), and Y-MESO-8D (sarcomatoid) cell lines are established by ourselves.16 Cells were grown in monolayer cultures in RPMI 1640 (Sigma-Aldrich Corp., St. Louis, MO, USA) containing 10% fetal bovine serum, 2 mmol/L glutamine and 1 mmol/L sodium pyruvate at 37°C in a humidified atmosphere of 95% air and 5% CO2. MeT-5A cells were cultured in Medium 199 with Earle’s balanced salt solution, 0.75 mM l-glutamine and 1.25 g/L sodium bicarbonate supplemented with 3.3 nM epidermal growth factor, 400 nM hydrocortisone, 870 nM insulin, 20 mM 4–2-hydroxyethyl-1-piperazineetha-nesulfonic acid and 10% fetal bovine serum.
RNA isolation and quantitative real-time reverse transcriptase-PCR analysis
For mRNA analysis, 5 µg of total RNA isolated using Trizol (Invitrogen, Carlsbad, CA, USA) were reverse transcribed with Super script III First-Strand Synthesis System using Random primer system (Invitrogen, Carlsbad, CA, USA). Quantitative real-time reverse transcriptase-PCR (qRT-PCR) analysis of BMAL1, NPAS2 and CLOCK was performed as described previously. 17 GAPDH (Assays-on-Demand; Applied Biosystems, Foster City, CA, USA) was used as an internal control.
Transfection of short interfering RNA
Cells (4.5 × 105) were plated in 10 cm2 dish plate. Next day, cells were transiently transfected with either 10 nM predesigned short interfering RNA (siRNA) [Stealth Select RNA interference (RNAi)] targeting BMAL1 or control siRNA purchased from Invitrogen using Lipofectamine RNAiMAX (Invitrogen Corp., Carlsbad, CA, USA) according to the manufacturer’s protocol. After 48 hr, the transfected cells were harvested for further analysis or plated for growth assays.
Western blot analysis
Cells were collected and washed twice in 1× phosphate-buffered saline (PBS), then lysed in ice-cold lysis buffer (0.5 M Tris-HCl with pH 7.4, 1.5 M NaCl, 2.5% deoxycholic acid, 10 mM EDTA, 10% NP-40, 0.5 mM DTT, 1 mM phenylmethylsulfonyl fluoride, 5 mg/mL leupeptin and 10 mg/mL aprotinin) for 5 min. The lysate was centrifuged at 13,000 rpm for 20 min at 4°C, and protein content of the supernatant was measured. Total cell lysates (30 µg/well) were separated by SDS-PAGE and the gels transferred into nitrocellulose membranes (Whatman, Piscataway, NJ). Membranes were blocked with 5% nonfat dry milk in PBS containing Tween-20 (PBST) (1× PBS, 0.1% Tween-20) for 1 hr at room temperature and incubated with primary antibody at 4°C overnight. Membranes were then washed three times with PBST and probed with appropriate horseradish peroxidase-conjugated secondary antibody for 1 hr at room temperature. The membranes were washed three times in PBST and bands were visualized using Western blot chemiluminescence reagent (BioRad, Hercules, CA 94547, USA). Antibodies were obtained from Santa Cruz (CA. 95060, USA) and Cell Signaling (Danvers, MA 01923, USA) Biotechnologies and used at the following dilutions (BMAL1, 1:1,000; Cyclin B, 1:1,000; Cyclin E, 1:1,000; Wee1, 1:1,000; p21WAF1/CIP1, 1:1,000; Cleaved caspase 3, 1:1,000; beta-actin, 1:5,000).
Immunofluorescence staining
Forty-eight-hour post-transfection with BMAL1-siRNA or control-siRNA oligos, in vitro growing cells were seeded at 2 × 104 cells/chamber in two-well Lab-Tek™ Chamber Slide System, incubated overnight at 37°C and 5% CO2. Next day, cells were washed with PBS and fixed with 4% paraformaldehyde solution, permeabilized with 0.01% Triton X-100 and blocked in 1% BSA in PBS for 20 min at room temperature. Subsequent antibody incubations were in PBS containing 1% BSA–0.01% Triton X-100. Antibody reagents were mouse anti-α-tubulin (1:100, Sigma-Aldrich St. Louis, MO, USA) and rabbit anti-BMAL1 H-170 (1:200, Santa Cruz Biotechnology, CA. 95060, USA). Incubation with primary antibody was kept overnight at 4°C, followed by three washes with PBS solution. Secondary reagent with Alexa Flour® 488-cojugated goat antimouse IgG, goat antirabbit IgG (1:1,000, Molecular probes, Invitrogen, Eugene, OR 97402, USA) and Alexa Flour® 594-cojugated goat antimouse IgG (1:700, Molecular probes, Invitrogen Eugene, OR 97402, USA). Secondary antibodies were incubated for 1 hr at room temperature. After PBS washing, independent mounting was done with Prolong Gold antifade reagent supplemented with 4,6-diamidino-2-phenylindole (DAPI) (Invitrogen, Carlsbad, CA, USA). All stained cells were visualized by confocal Eclipse TE 2000-E microscope (Nikon, Tokyo, Japan) with 20×, 40× and 60× objectives.
Tumor specimens of patients with MPM
Sixteen resected MPM tissue specimens were obtained from patients diagnosed with MPM who underwent surgery at the Department of Thoracic Surgery, Nagoya University Hospital between 2005 and 2011. Eleven patients received neoadjuvant chemotherapy with pemetrexed and cisplatin, while one patient received adjuvant chemotherapy and radiotherapy. Fifteen specimens were obtained through extra-pleural pneumonectomy and one was obtained by pleurectomy. Diagnosis of mesothelioma was made based on clinical evaluation, histopathologic examination, and the clinical stage was determined according to the International Mesothelioma Interest Group.18 Patients’ overall survival was defined as the length of time from the date of surgery to that of death. Four noncancerous (normal parietal pleura) tissue specimens were obtained from patients who underwent thoracic surgery for different causes (i.e., not including mesothelioma) and were used as normal controls. The study protocol was approved by the Institutional Review Boards of Nagoya University Graduate School of Medicine. Informed consent was obtained from the patients following institutional guidelines. The expression level of the circadian clock protein, BMAL1, was examined in the aforementioned surgically resected specimens. Sections from formalin-fixed paraffin-embedded were treated for immunostaining with commercially available BMAL1 antibody (Sanata Cruz Biotechnology, CA.95060, USA) according to the procedures described elsewhere.13
Cell growth assays
Colorimetric proliferation assay was performed using WST-1 assay kit (Roche, Basel, Switzerland) according to manufacturer’s instruction. Liquid and soft agar colony formation assays were done as described previously.19
Cell cycle analysis
Post-transfection with BMAL1-siRNA or control oligos, cells were synchronized by serum starvation for 12 hr or by double-thymidine treatment (to block mitosis and induce late G1/early S phase arrest; subconfluent cell cultures were incubated in complete medium containing 2 mM thymidine for 18 hr, and then, the thymidine medium was removed and replaced with complete medium lacking thymidine for 12 hr followed by another 18-hr incubation in the presence of thymidine). Synchronized populations were harvested and washed in ice-cold PBS. Following centrifugation at 900g for 5 min, cells were suspended in 150 µL of cold PBS while vortex gently, and cells were fixed by dropwise addition of 350 µL ice-cold ethanol. Fixed cells were stored at −20°C for at least 30 min. For staining, pelleted cells were washed twice with cold PBS and resuspended in 0.5 mL PBS containing 200 µg/mL RNase, and stained with propidium iodide 20 µg. Cells were incubated at 37°C for 30 min and maintained at 4°C before analysis, cells were filtered through 40 µM nylon mesh and analyzed by flow cytometry for cell cycle status [FACS Calibur instrument (Becton Dickinson), with BD Cell Quest™ Pro Ver. 5.2.1 (BD) Bioscience, Franklin Lakes, NJ, USA].
Apoptosis analysis
Apoptosis was quantified by detecting surface exposure of phosphatidylserine in apoptotic cells using a phycoerythrin (PE)—Annexin V Apoptosis Detection Kit I (BD Biosciences). Cells were harvested 5 days after transfection of siRNA oligos, treated according to the manufacturer’s instructions and measured with PE 7-amino-actinomycin D (7-AAD) staining using flow cytometry [FACS Calibur instrument (Becton Dickinson), with BD Cell Quest™ Pro Ver. 5.2.1 (BD) Bioscience, Franklin Lakes, NJ, USA].
Hematoxylin and eosin staining
Post-transfection with BMAL1-siRNA or control oligos, ACC-MESO-1 cells were harvested after 48 hr and 10,000 cells were plated/chambered in eight-well Lab-Tek™ Chamber Slide System, and then incubated overnight at 37°C and 5% CO2. Next day, cells washed with PBS and fixed with 1% glutaraldehyde for 5 min and subsequently stained with Mayer’s Hematoxylin solution (Muto Puro Chemical LTD, Tokyo, Japan) for 2–3 min, rinsed with distilled water and then submerged in Scott’s tap water substitute (0.2% NaHCO3 and 2% MgSO4) for bluing, finally cells were washed with distilled water once and examined under the microscope.
Statistics
SPSS Ver. 18 software was used for all statistics analysis in this study. Mann–Whitney U-test was used for analyzing difference between two groups.
Results
MPM cell lines express higher levels of BMAL1 than normal pleural mesothelial cells
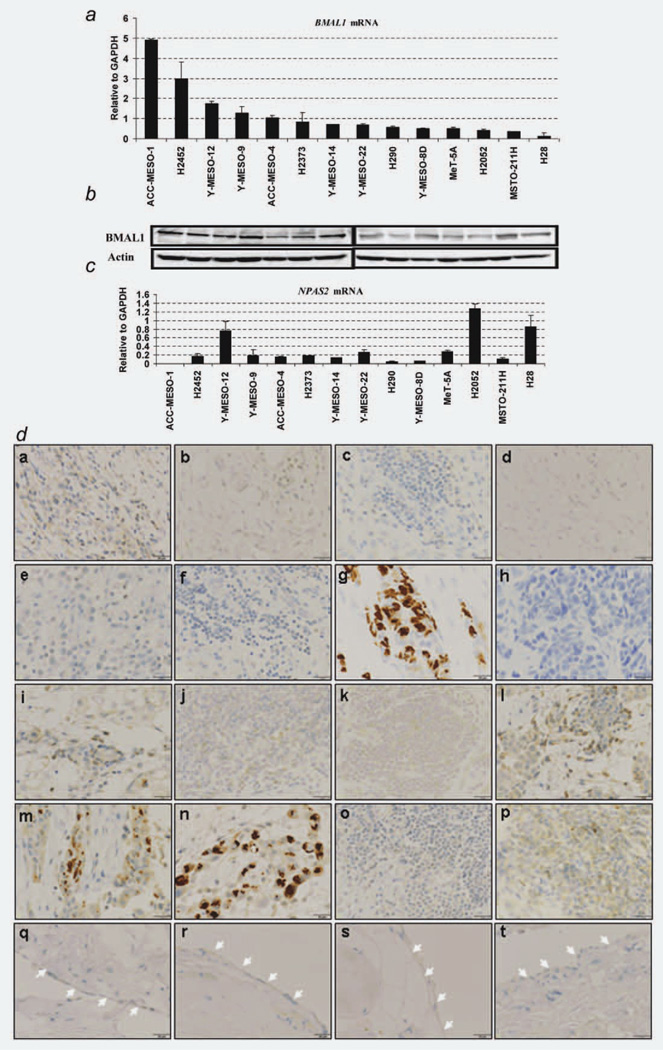
Quantitative detection of BMAL1 and NPAS2 mRNA in a panel of MPM and MeT-5A cell lines was performed using qRT-PCR. Ten out of 13 (77%) MPM cell lines expressed higher levels of BMAL1 mRNA than MeT-5A, while NPAS2 expression profile seems different from BMAL1 profile in MPM cell lines (Figs. 1a and 1c). Western blot analysis for the same set of cell lines clearly demonstrated variable degrees of BMAL1 protein expression (Fig. 1b). Furthermore, we investigated BMAL1 expression levels in 16 surgically annotated MPM and four normal parietal pleural specimens (Table 1). Through immunohistochemical analysis, BMAL1 was detected in a subset of MPM specimens with nuclear and/or cytoplasmic localization, while none of the four normal parietal pleural samples showed detectable levels of BMAL1 protein (Fig. 1d), suggesting that BMAL1 may be important in the development of MPM.
Figure 1.
BMAL1 expression levels in MPM and normal parietal pleura. (a) qRT-PCR analysis of BMAL1 in 13 MPM cell lines and an immortalized pleural mesothelial cell line MeT-5A (control). The cell lines are aligned by expression levels of BMAL1 mRNA from high (left) to low (right). The result is a representative of two independent qRT-PCR experiments done in duplicated reactions. (b) Western blots of BMAL1 in MPM cell lines. Actin was used as a loading control. (c) qRT-PCR analysis of NPAS2 in 13 MPM cell lines and MeT-5A. (d) Immunohistochemical staining of BMAL1 in surgically annotated MPM and normal parietal pleural specimens. Immunohistochemical analysis showed an overexpression of BMAL1 in a subset of MPM specimen (a–p), whereas normal pleural mesothelial cells showed negative BMAL1 immunoreactivity (white arrows, q–t). Details of MPM and normal parietal pleural specimens were mentioned in “Material and Methods” Section. ×400 magnification and 25 µm scale bars were used for all images. [Color figure can be viewed in the online issue, which is available at wileyonlinelibrary.com.]
Table 1.
Clinical features of 16 patients with MPM
| ID | Age/Sex | Histology | BMAL1 Status (IHC score) |
Asbestos exposure |
OS (M) | Staging |
Surgery | Therapy | |
|---|---|---|---|---|---|---|---|---|---|
| Clinical | Pathological | ||||||||
| 1 | 50/M | Biphasic | Negative (1+) | + | 2.5 | T2N0M0/II | T3N2M0/III | Right EPP | No |
| 2 | 54/M | Biphasic | Negative (1+) | + | 24.0 | T2N0M0/II | T2N0M0/II | Left EPP | No |
| 3 | 56/M | Epithelioid | Negative (1+) | + | 25.3 | T3N2M0/III | T4N0M0/IV | Left EPP | No |
| 4 | 65/M | Epithelioid | Negative (0) | + | 8.8 | T3N1M0/III | T3N2M0/III | Right EPP | Neoadjuvant CT |
| 5 | 46/F | Epithelioid | Negative (1+) | − | 36.0 | T1bN0M0/Ib | T3N2M0/III | Left EPP | Adjuvant CT + LRT |
| 6 | 70/F | Epithelioid | Negative (0) | − | 27.6 | T1bN1M0/III | T2N1M0/III | Left EPP | Neoadjuvant CT |
| 7 | 65/M | Epithelioid | Positive (3+) | + | 8.4 | T3N0M0/III | T4N0M0/IV | Right EPP | Neoadjuvant CT |
| 8 | 60/M | Biphasic | Negative (0) | + | 31.3 | T3N2M0/III | T3N0M0/III | Right EPP | Neoadjuvant CT + PHR |
| 9 | 62/M | Epithelioid | Positive (2+) | + | 13.0 | T3N0M0/III | T4NxM0/IV | Left pleurectomy | Neoadjuvant CT |
| 10 | 67/M | Biphasic | Negative (1+) | − | 11.6 | T2N1M0/III | T3N0M0/III | Right EPP | Neoadjuvant CT + PHR |
| 11 | 66/M | Biphasic | Negative (0) | + | 8.3 | T2N0M0/II | T3N0M0/III | Right EPP | Neoadjuvant CT |
| 12 | 67/M | Epithelioid | Positive (2+) | + | 3.6 | T2N0M0/II | T3N2M0/III | Right EPP | No |
| 13 | 68/M | Epithelioid | Positive (3+) | − | 7.7 | T3N0M0/III | T3N0M0/III | Left EPP | Neoadjuvant CT + PHR |
| 14 | 63/M | Epithelioid | Positive (3+) | + | 1.9 | T2N0M0/II | T3N0M0/III | Lt EPP | Neoadjuvant CT |
| 15 | 68/M | Sarcomatoid | Negative (0) | + | 1.0 | T3N2M0/III | T3N0M0/III | Right EPP | Neoadjuvant CT |
| 16 | 64/M | Epithelioid | Negative (1+) | + | 0.8 | T2N0M0/II | T3N2M0/III | Left EPP | Neoadjuvant CT |
Abbreviations: Id, patient’s number; IHC, immunohistochemical score for BMAL1; OS, overall survival; M, month; EPP, extra-pleural pneumonectomy; neoadjuvant CT, neoadjuvant chemotherapy (cisplatin + pemetrexed); adjuvant chemotherapy (cisplatin + pemetrexed); LRT, local radiation therapy; PHR, post-operative hemithoracic radiation.
Expression profile of BMAL1 and CLOCK over 24 hr in ACC-MESO-1 and MeT-5A cells
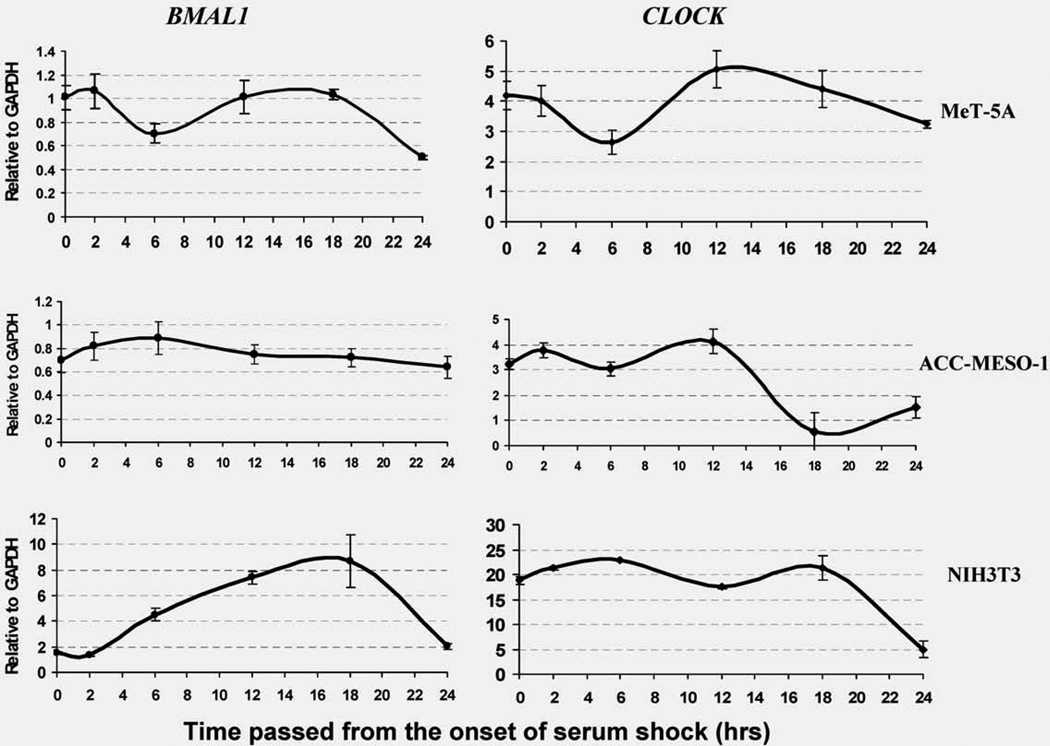
Based on previous data showing that expression of BMAL1 followed a rhythmical pattern in animal models,20 we decided to examine the expression of BMAL1 and CLOCK by qRT-PCR analysis to evaluate their oscillation over 24 hr in ACC-MESO-1 and MeT-5A cells. We used the mouse fibroblast cell line (NIH3T3) as a positive control as it shows a rhythmical expression of BMAL1.20 Serum shock was done as described previously.5 BMAL1 rhythmical expression was found in serum-shocked normal nontumorigenic mesothelial cells but not in ACC-MESO-1 cells, whereas CLOCK showed rhythmical expression in both MPM and normal mesothelial cells (Fig. 2). These findings indicate that BMAL1 rhythmical expression was intact in normal mesothelial cells but not in MPM cells.
Figure 2.
Expression profiles of BMAL1 and CLOCK genes over 24 hr in ACC-MESO-1 and MeT-5A cells. qRT-PCR analysis of BMAL1 and CLOCK mRNA in ACC-MESO-1, MeT-5A at time indicated above. NIH3T3 cells were used as a positive control. Serum shock was performed as mentioned in the “Materials and Methods” Section. We found that serum shock induces rhythmical expression changes of BMAL1 and CLOCK mRNA in NIH3T3 as well as MeT-5A cells. Serum shocked ACC-MESO-1 cells showed rhythmical changes of CLOCK mRNA, but not BMAL1 (GAPDH was used as an internal control). The results are averages of two independent experiments done in duplicate.
BMAL1 knockdown suppresses proliferation, anchorage-dependent and independent clonal growth of MPM cells
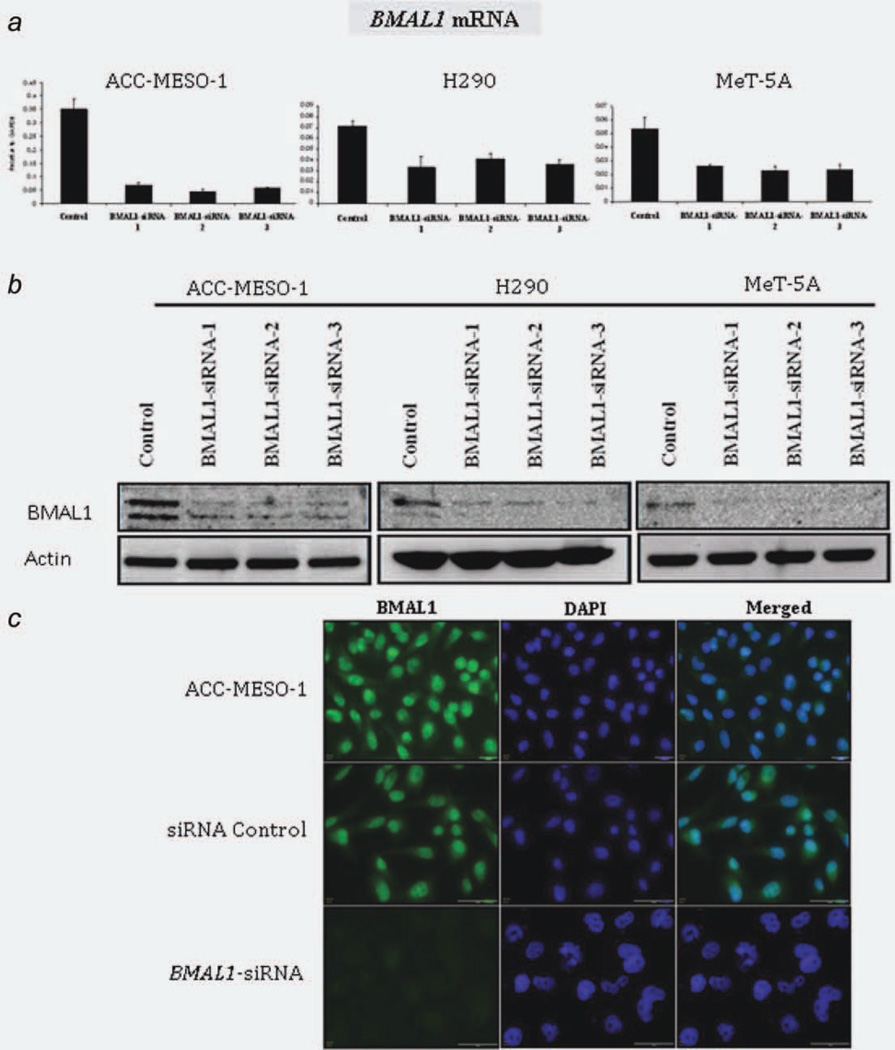
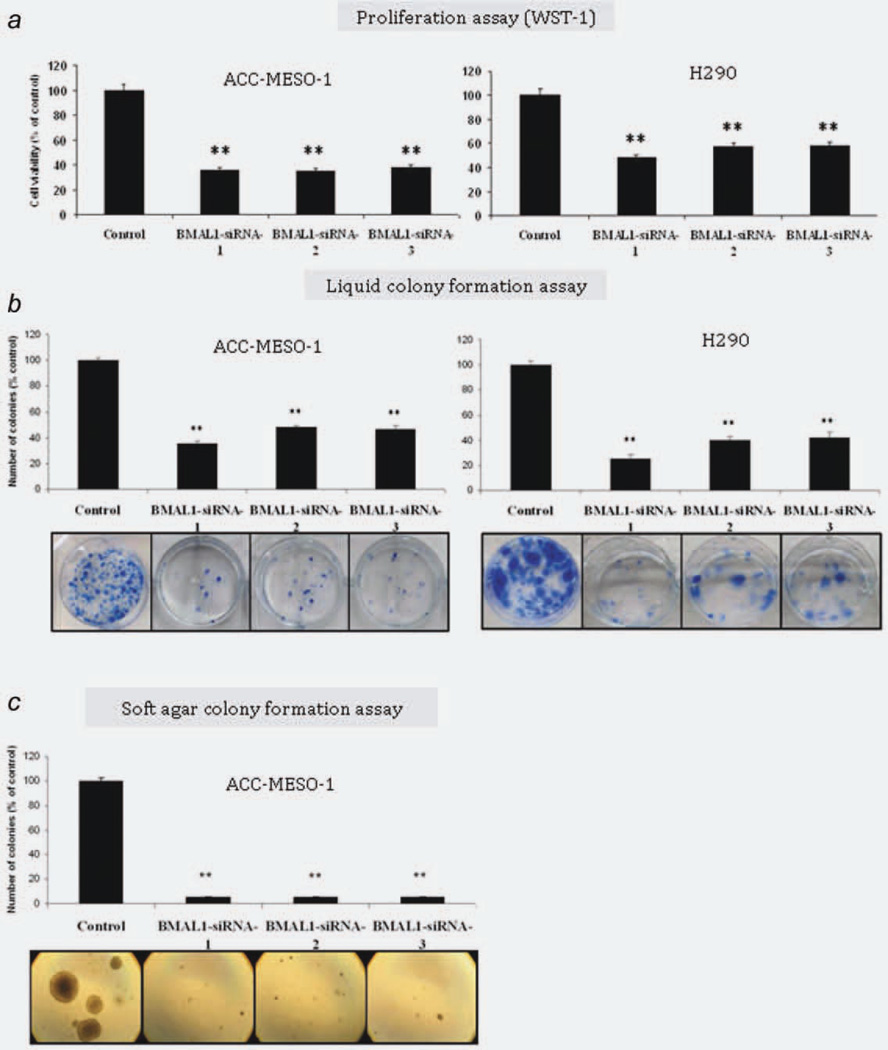
To investigate the role of BMAL1 in MPM cell growth, we performed RNAi-mediated gene silencing against BMAL1. ACC-MESO-1 and H290 cell lines were selected as the MPM cell models to be used for further investigations (Fig. 1a). MeT-5A was used as a normal control. To reduce the off target effects, we used low dose (10 nM) stealth selected RNAi (Invitrogen Carlsbad, CA, USA) which includes three siRNA oligos with nonoverlapping sequences targeting BMAL1. Efficient BMAL1 knockdown was confirmed by qRT-PCR, western blotting and immunostaining (Fig. 3). Next, to evaluate the effect of BMAL1 knockdown on cell proliferation in mass culture and clonogenic growth in anchorage-dependent and independent conditions, we performed colorimetric growth, liquid and soft agar colony formation assays. We found that in ACC-MESO-1 and H290 cells, BMAL1 knockdown significantly suppressed proliferation and dramatically suppressed colony formation in anchorage-dependent (liquid colony formation assay) and anchorage-independent conditions (soft agar assay) (Fig. 4). By contrast, we did not see significant suppression of proliferation in MeT-5A (Supporting Information, Fig. S1a).
Figure 3.
BMAL1 knockdown in MPM and MeT-5A cells. Confirmation of BMAL1 knockdown by (a) qRT-PCR, (b) Western blot analysis and (c) immunofluorescence (IF) assay. IF was used to examine BMAL1 levels in the parental, siRNA control or siRNA-BMAL1-treated ACC-MESO-1 cells. All data are averages of three independent experiments done in duplicates. [Color figure can be viewed in the online issue, which is available at wileyonlinelibrary.com.]
Figure 4.
BMAL1 knockdown inhibits proliferation and suppresses clonal growth of MPM cells in anchorage-dependent and -independent conditions. (a) WST-1 proliferation. (b) Liquid colony formation assays for ACC-MESO-1 and H290 cells transfected with BMAL1-siRNA or control oligos. (c) Soft agar colony formation assay for ACC-MESO-1 cells transfected with BMAL1-siRNA or control oligos. Results are from three independent experiments and shown as mean ± SD. In liquid colony and soft agar assay colony, numbers of cells transfected with control oligos are set as 100%. ** indicate p < 0.01 (Mann–Whitney U test). All data are averages of three independent experiments done in duplicates. [Color figure can be viewed in the online issue, which is available at wileyonlinelibrary.com.]
BMAL1 depletion induces massive apoptosis in MPM cells, with limited consequences in the normal pleural mesothelial cells
Next, we investigated whether the antiproliferative effect of BMAL1 depletion is due to cell death. BMAL1 knockdown resulted in massive apoptosis and necrosis in MPM cells (Fig. 5a), as evidenced by increases in cleaved caspase-3 activation in ACC-MESO-1 cells (Fig. 5b). By contrast, subtle apoptosis could be found in MeT-5A cells following BMAL1 knockdown (Fig. 5a). These results indicate that BMAL1 knockdown-induced growth inhibition occurs in part through apoptotic mechanisms and MPM cells are more dependent on BMAL1 expression for their survival than normal mesothelial cells.
Figure 5.
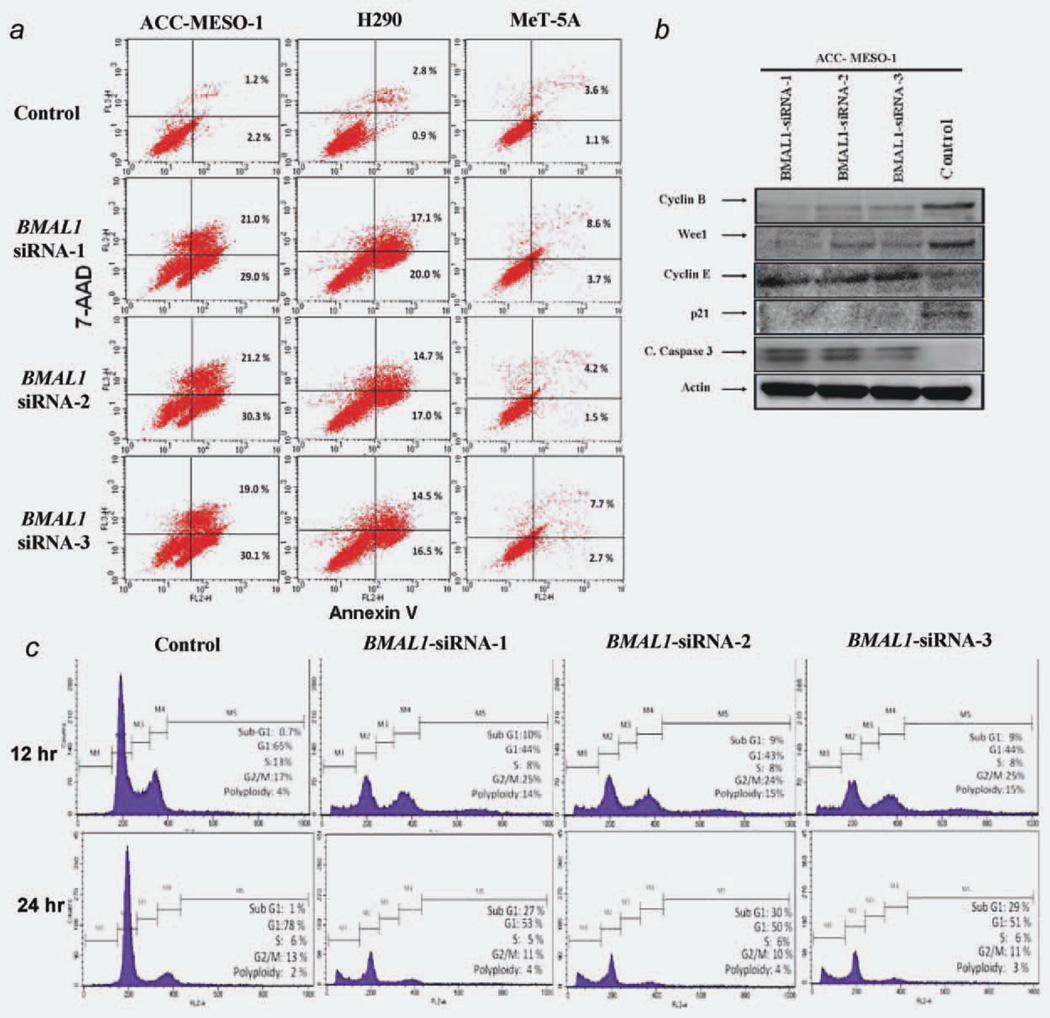
BMAL1 knockdown induces apoptosis and cell cycle disruption (a) FACS analysis of cells costained with anti-annexin V and 7-AAD. High Annexin V and low 7-AAD cells are undergoing apoptosis while cells with low Annexin and high 7-AAD are undergoing necrosis. (b) Immunoblot showing effects of BMAL1 knockdown on its targets in ACC-MESO-1 cells. (c) Cell cycle analysis of ACC-MESO-1 cells transfected with BMAL1-siRNA or control oligos. Forty-eight-hour post-transfection cells underwent serum starvation for 12 hr (upper panel) and for 24 hr (lower panel) then were harvested with both adherent and floating cells were combined and prepared for cell cycle analysis by flow cytometry, as described in “Material and Methods” Section. Results are the average of two independent experiments. [Color figure can be viewed in the online issue, which is available at wileyonlinelibrary.com.]
BMAL1 knockdown leads to cell cycle disruption in ACC-MESO-1 cells with a clear rise of polyploidy
We investigated whether BMAL1 knockdown-induced growth inhibition also was caused by cell cycle arrest. Consistent with induction of apoptosis as measured by cleaved caspase-3 expression, there was an increase in subG1 DNA content in ACC-MESO-1 transfected with BMAL1-siRNA oligos compared to cells transfected with control oligos (Fig. 5c). The proportion of cells in G1 phase decreased whereas a modest increase in the proportion of cells in G2/M phase. Notably, the profiling also showed an increase in the fraction of cells whose DNA contents exceed 4 N that correspond to polyploidy cell population. With much longer times in culture, high percentages of those cells (in G2/M phase and polyploidy region) decreased while subG1 population significantly increased (Fig. 5c), implying that polyploidy cells underwent apoptosis. By contrast, we did not see significant changes in cell cycle profiling of MeT-5A following BMAL1 knockdown (Supporting Information, Fig. S1b).
Depletion of BMAL1 induces drastic morphological alterations indicative of mitotic catastrophe in ACC-MESO-1 cells
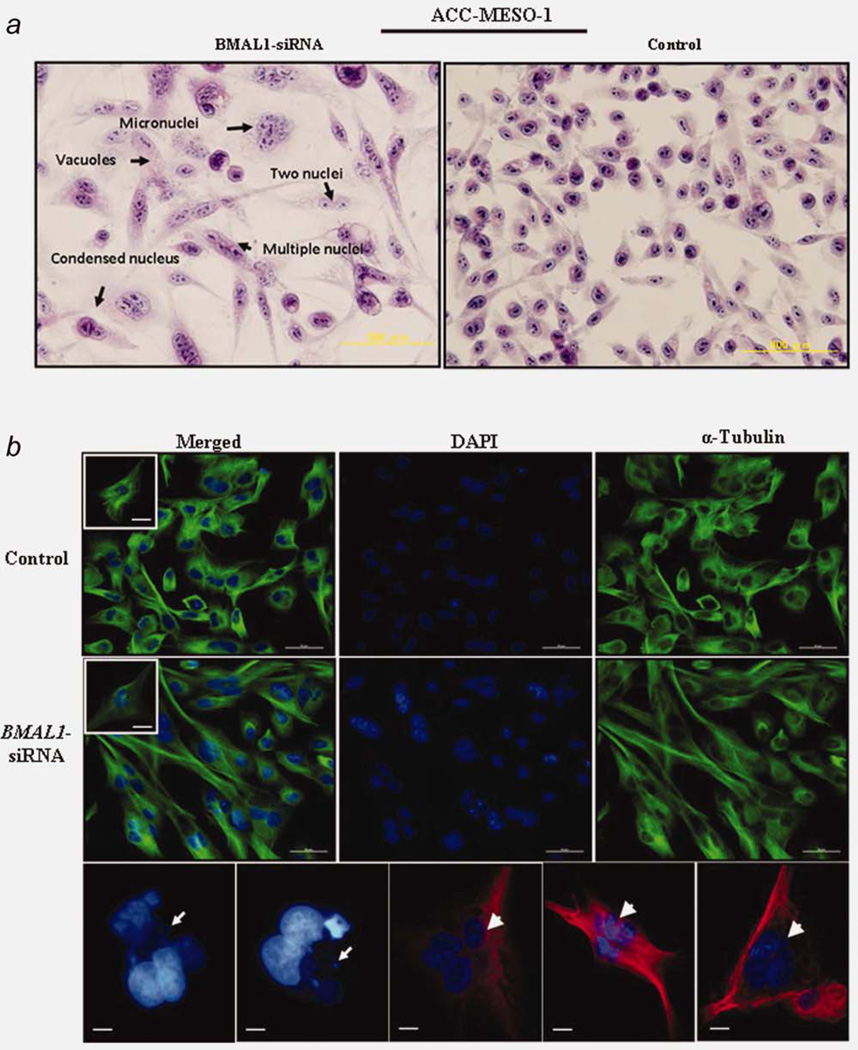
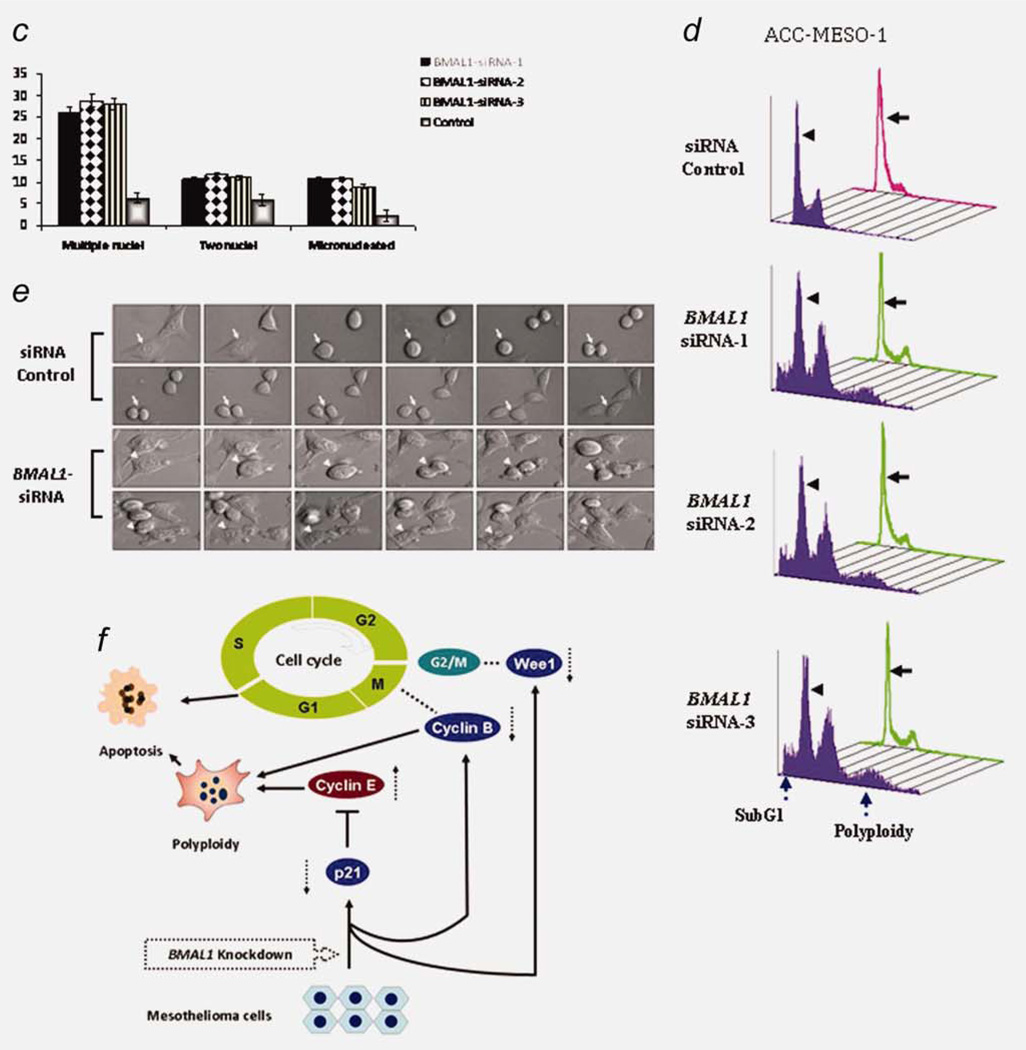
We noted that following BMAL1 knockdown, ACC-MESO-1 cells underwent drastic morphological changes; the cells enlarged and elongated by visual examination under phase contrast microscopy. Next, we performed hematoxlin and eosin (H–E) staining for these cells. As shown in Figure 6a, dramatic morphological alterations were seen in ACC-MESO-1 cells after BMAL1 knockdown; cells exhibit much enlarged flattened shape, micronucleation, multiple nuclei and vacuolization occasionally were found. Next, we examined BMAL1 siRNA-treated ACC-MESO-1 cells by detailed morphological analysis of DAPI-stained cell nuclei and α-tubulin immunostaining to visualize the cytoskeletal alterations associated with changes in nuclear morphology. Multiple morphological defects were identified which are consistent with the findings observed by H–E staining (Figs. 6b and 6c). Importantly, the existence of micronucleation is a highly indicative sign of catastrophic mitosis. To examine the possibility of mitotic catastrophe as a sequel of BMAL1 knockdown and to explain its role in BMAL1-induced cell death during the cell cycle, ACC-MESO-1 cells transfected with BMAL1-siRNA or control oligos synchronized using a double thymidine block (to induce pharmacological block of mitosis in these cells). Interestingly, inhibition of mitosis in ACC-MESO-1 transfected with BMAL1 siRNA resulted in a marked decrease of polyploidy and subG1 population (Fig. 6d), suggesting aberrant mitosis as a cause of cell death after BMAL1 knockdown. To confirm the occurrence of mitotic catastrophe following BMAL1 knockdown, we performed time lapse microscopic examination for ACC-MESO-1 cells transfected with BMAL1-siRNA or control oligos. Examination showed that cells rounding up as they entered mitosis, then attempting to undergo cytokinesis. ACC-MESO-1 cells transfected with control siRNA successfully completed mitosis, but BMAL1 siRNA-treated cells failed to divide properly and exhibited large cell volume and micronucleation (Fig. 6e), indicating mitotic catastrophe as a cell fate following BMAL1 knockdown in those cells.
Figure 6.
BMAL1 knockdown induces dramatic morphological alterations in ACC-MESO-1 cells. (a) H–E stain showing the nuclear morphological changes identified in ACC-MESO-1 cells after ablation of BMAL1. (b) IF of α-tubulin and DAPI stains. The upper panels represent ACC-MESO-1 cells treated with siRNA control. The lower panels represent the most frequent morphological changes (arrow indicates micronucleation and arrow head indicates multiple nuclei) in single ACC-MESO-1 cell after BMAL1 depletion. The middle panels represent ACC-MESO-1 cells treated with BMAL1 siRNA. (c) Quantification of the binuclear, multinuclear and micronuclear phenotypes in ACC-MESO-1 cells after BMAL1 knockdown. (d) Cell cycle profiling of ACC-MESO-1 cells showing marked decrease of BMAL1-induced cell death and polyploidy formation following mitosis block. ACC-MESO-1 cells transfected with BMAL1 siRNA or control oligos with double-thymidine (black arrows) and without thymidine treatment (arrow heads) were harvested for analyses of DNA content by flow cytometry. Synchronized cells at late G1/early S by double-thymidine escaped from BMAL1 knockdown-induced cell death with marked reduction in subG1 and polyploidy formation. (e) Time lapse microscopic examination showing the aberrant mitosis in ACC-MESO-1 cells transfected with BMAL1 siRNA (white arrow head) and intact mitosis in cells transfected with control oligos (white arrow). (f) Proposed molecular mechanism of BMAL1 knockdown-induced mitotic catastrophe in ACC-MESO-1 cells. Decreasing cyclin B below a critical level results in mitosis skipping and enhances polyploidy formation. Downregulation of p21WAF1/CIP1 results in accumulation of cyclin E which could lead to increased number of polyploidy cells. Wee1 downregulation could contribute to eventual escape from G2/M arrest, which in turns participates in impairment of mitotic integrity. [Color figure can be viewed in the online issue, which is available at wileyonlinelibrary.com.]
RNAi-mediated knockdown of BMAL1 results in expression alterations of the cell cycle regulators in ACC-MESO-1 cells
Recent data suggest that there is a strong relationship between the circadian clock system and regulation of the cell cycle.21,22 In particular, BMAL1 is considered a key regulator of cancer cell proliferation through coordinating the activity of cell cycle proteins including p21WAF1/CIP1 and cyclin B. Specifically, some data suggest that the circadian clock controls mitotic process through Wee1, known to be clock target gating the G2/M transition.10,21,23 Therefore, we investigated the link between BMAL1 knockdown and the status of the cell cycle proteins. Notably, ACC-MESO-1 cells after BMAL1 knockdown showed profound alterations in cell cycle regulators, including significant decrease of Wee1, p21WAF1/CIP1, cyclin B proteins and accumulation of cyclin E protein (Fig. 5b).
Discussion
In this report, we show that most MPM cell lines and a subset of surgically annotated MPM specimens expressed higher levels of BMAL1 than MeT-5A and normal parietal pleural specimens, respectively. Silencing of BMAL1 resulted in suppression of MPM cell growth and induction of apoptosis in these cells but limited effect was observed in MeT-5A. BMAL1 depletion from ACC-MESO-1 cells, which expressed the highest level of BMAL1, led to cell cycle disruption with a substantial increase in apoptotic and polyploidy cell population as well as decreased levels of Wee1, cyclin B and p21WAF1/CIP1 expression and upregulation of cyclin E. BMAL1 knockdown in ACC-MESO-1 cells induced mitotic catastrophe denoted by marked disruption of cell cycle regulator proteins and drastic morphological changes including micronucleation, multiple nuclei and increased cellular volume of ACC-MESO-1 cells.
By immunohistochemical analysis, we demonstrated that BMAL1 is constitutively expressed in a subset of clinical MPM samples. Two out of three Stage IV-MPM patients were positive for BMAL1, whereas three out of 12 Stage III-MPM patients were positive (Table 1). This suggests that BMAL1 expression may be associated with advanced stage-MPM, but the small patients’ number makes it difficult to draw a firm conclusion on this possible association (Supporting Information, Fig. S2). It would be of importance to analyze the association between BMAL1 expression and clinicopathological features of MPM using a large cohort. In line with our provisional results, high BMAL1 expression was observed in metastatic breast, colorectal and liver cancers,12,13 these findings suggest that BMAL1 has an oncogenic activities in a variety of human cancers.
A prior study reported that clock genes oscillation was found in different tissues including muscle, liver and adipose tissues.24 BMAL1 rhythmic expression was observed also in NIH3T3 cell following synchronization by serum shock.25 Here, we found that the rhythmic expression of BMAL1 over the 24 hr period was intact and clearly observed in NIH3T3 cells and MeT-5A as well. Intriguingly, ACC-MESO-1 cells showed constant levels of BMAL1 mRNA. This result is consistent with a recent report20 showing that BMAL1 is rhythmically expressed in mouse prostate while serum-shocked synchronized prostate cancer cells showed disrupted circadian rhythmicity of BMAL1 gene. In line with Roe et al.’s15 results who reported that BMAL1 was found to be overexpressed concomitantly with downregulation of its negative counterpart in MPM compared to the normal parietal pleural tissue, our findings support the notion that the circadian rhythm pathway could be deregulated in MPM.
We demonstrated that inhibition of BMAL1 expression using RNAi technique significantly suppressed proliferation, anchorage-dependent and independent clonal growth in MPM cells. To explore how BMAL1 knockdown induced growth inhibition, we performed apoptosis assays and found that depletion of BMAL1 expression by siRNA resulted in a significant increase in the fraction of apoptotic and necrotic cells in the MPM cell lines (ACC-MESO-1 and H290) with limited effect in MeT-5A. We also examined the regulatory role of BMAL1 in MPM cell cycle and its importance in sustained cell proliferation. Interestingly, we found that RNAi-mediated knockdown of BMAL1 resulted in cell cycle disruption of ACC-MESO-1 cells but not in MeT-5A cells.
Following BMAL1 transient knockdown, we observed multiple morphological abnormalities consistent with aberrant mitotic process. There was an increase of binucleated cells, which may correspond to the modest increase in the G2/M phase cell population. Furthermore, some other cells (about 10%) showed micronucleation. Bergman et al.26 reported that the occurrence of cells with double nucleus suggests that these cells undergo cell division without segregating their DNA and could be explained by the increase in cells with 4 N DNA content seen with flow cytometry. Importantly, micronucleation is highly indicative of mitotic catastrophe and could be resulted from chromosomal mis-segregation caused by DNA breaks. It is quite possible that ACC-MESO-1 cells underwent mitotic catastrophe due to impairment of cell cycle regulator proteins, such as, cyclin B, p21WAF1/CIP1 and Wee1 (Fig. 6f). It is demonstrated that decreasing cyclin B below a critical level results in mitosis skipping and enhances polyploidy formation, which is considered as one of the characteristics of mitotic catastrophe.27 Polyploidy cells pass an extra round of cell cycle and finally undergo apoptosis.28 In addition, Wee1 downregulation could contribute to eventual escape from G2/M arrest, which in turns participates in impairment of mitotic integrity. In this study, we observed downregulation of p21WAF1/CIP1 and cyclin E upregulation following BMAL1 knockdown, and the latter could also lead to increased number of polyploidy cells. Other investigators noted that BMAL1 modulate the transcriptional activity of p53 toward its target p21 and clearly showed that BMAL1 knockdown caused a decrease in p21 level.22 Moreover, a previous study reported that overexpression of cyclin E led to impairment of mitosis and polyploidy formation.29 We believe that accumulation of polyploidy, large multinucleated cells and micronucleation in ACC-MESO-1 after BMAL1 knockdown were consistent with cell death mechanism involving mitotic catastrophe. It has been shown that mitotic catastrophe could be considered as an important safeguard to prevent the proliferation of polyploidy cells30 and apoptosis frequently follows mitotic catastrophe.31 Taken together, these findings show that BMAL1 plays a critical role in the mitotic process of cancer cells. To our knowledge, this study is the first to shed light on the involvement of BMAL1 in polyploidy formation, impairment of mitotic events, and also to highlight the role of circadian clock genes in MPM. In conclusion, we provide evidence that BMAL1 plays an important role and it may be a promising therapeutic target for MPM, but careful consideration is needed to avoid the counter effect on tissues dependant on BMAL1 such as muscles and liver.
Supplementary Material
Abbreviations
- BMAL1
brain and muscle ARNT-like protein-1
- STR
short tandem repeat
- GAPDH
glyceraldehyde 3-phosphate dehydrogenase
- BSS
balanced salt solution
- H–E
Hematoxlin and Eosin
- HEPES
4-2-hydroxyethyl-1-piperazineetha-nesulfonic acid
- MPM
malignant pleural mesothelioma
- NPAS2
neuronal PAS domain protein 2
- qRT-PCR
quantitative real-time reverse transcriptase-PCR
- siRNA
short interfering RNA
- DAPI
4,6-diamidino-2-phenylindole
Footnotes
Additional Supporting Information may be found in the online version of this article.
References
- 1.Robinson BW, Musk AW, Lake RA. Malignant mesothelioma. Lancet. 2005;366:397–408. doi: 10.1016/S0140-6736(05)67025-0. [DOI] [PubMed] [Google Scholar]
- 2.Sekido Y. Molecular biology of malignant mesothelioma. Environ Health Prev Med. 2008;13:65–70. doi: 10.1007/s12199-007-0015-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Tsao AS, Wistuba I, Roth JA, et al. Malignant pleural mesothelioma. J Clin Oncol. 2009;27:2081–2090. doi: 10.1200/JCO.2008.19.8523. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Taniguchi H, Fernandez AF, Setien F, et al. Epigenetic inactivation of the circadian clock gene BMAL1 in hematologic malignancies. Cancer Res. 2009;69:8447–8454. doi: 10.1158/0008-5472.CAN-09-0551. [DOI] [PubMed] [Google Scholar]
- 5.Balsalobre A, Damiola F, Schibler U. A serum shock induces circadian gene expression in mammalian tissue culture cells. Cell. 1998;93:929–937. doi: 10.1016/s0092-8674(00)81199-x. [DOI] [PubMed] [Google Scholar]
- 6.Straif K, Baan R, Grosse Y, et al. Carcinogenicity of shift-work, painting, and fire-fighting. Lancet Oncol. 2007;8:1065–1066. doi: 10.1016/S1470-2045(07)70373-X. [DOI] [PubMed] [Google Scholar]
- 7.Canaple L, Kakizawa T, Laudet V. The days and nights of cancer cells. Cancer Res. 2003;63:7545–7552. [PubMed] [Google Scholar]
- 8.Bunger MK, Wilsbacher LD, Moran SM, et al. Mop3 is an essential component of the master circadian pacemaker in mammals. Cell. 2000;103:1009–1017. doi: 10.1016/s0092-8674(00)00205-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Gauger MA, Sancar A. Cryptochrome, circadian cycle, cell cycle checkpoints, and cancer. Cancer Res. 2005;65:6828–6834. doi: 10.1158/0008-5472.CAN-05-1119. [DOI] [PubMed] [Google Scholar]
- 10.Matsuo T, Yamaguchi S, Mitsui S, et al. Control mechanism of the circadian clock for timing of cell division in vivo. Science. 2003;302:255–259. doi: 10.1126/science.1086271. [DOI] [PubMed] [Google Scholar]
- 11.Fu L, Lee CC. The circadian clock: pacemaker and tumour suppressor. Nat Rev Cancer. 2003;3:350–361. doi: 10.1038/nrc1072. [DOI] [PubMed] [Google Scholar]
- 12.Oshima T, Takenoshita S, Akaike M, et al. Expression of circadian genes correlates with liver metastasis and outcomes in colorectal cancer. Oncol Rep. 2011;25:1439–1446. doi: 10.3892/or.2011.1207. [DOI] [PubMed] [Google Scholar]
- 13.Kuo SJ, Chen ST, Yeh KT, et al. Disturbance of circadian gene expression in breast cancer. Virchows Arch. 2009;454:467–474. doi: 10.1007/s00428-009-0761-7. [DOI] [PubMed] [Google Scholar]
- 14.Koyanagi S, Kuramoto Y, Nakagawa H, et al. A molecular mechanism regulating circadian expression of vascular endothelial growth factor in tumor cells. Cancer Res. 2003;63:7277–7283. [PubMed] [Google Scholar]
- 15.Roe OD, Anderssen E, Helge E, et al. Genome-wide profile of pleural mesothelioma versus parietal and visceral pleura: the emerging gene portrait of the mesothelioma phenotype. PLoS One. 2009;4:e6554. doi: 10.1371/journal.pone.0006554. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Usami N, Fukui T, Kondo M, et al. Establishment and characterization of four malignant pleural mesothelioma cell lines from Japanese patients. Cancer Sci. 2006;97:387–394. doi: 10.1111/j.1349-7006.2006.00184.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Takeyama Y, Sato M, Horio M, et al. Knockdown of ZEB1, a master epithelial-to-mesenchymal transition (EMT) gene, suppresses anchorage-independent cell growth of lung cancer cells. Cancer Lett. 2010;296:216–224. doi: 10.1016/j.canlet.2010.04.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Husain AN, Colby TV, Ordonez NG, et al. Guidelines for pathologic diagnosis of malignant mesothelioma: a consensus statement from the International Mesothelioma Interest Group. Arch Pathol Lab Med. 2009;133:1317–1331. doi: 10.5858/133.8.1317. [DOI] [PubMed] [Google Scholar]
- 19.Sato M, Vaughan MB, Girard L, et al. Multiple oncogenic changes (K-RAS(V12), p53 knockdown, mutant EGFRs, p16 bypass, telomerase) are not sufficient to confer a full malignant phenotype on human bronchial epithelial cells. Cancer Res. 2006;66:2116–2128. doi: 10.1158/0008-5472.CAN-05-2521. [DOI] [PubMed] [Google Scholar]
- 20.Cao Q, Gery S, Dashti A, et al. A role for the clock gene per1 in prostate cancer. Cancer Res. 2009;69:7619–7625. doi: 10.1158/0008-5472.CAN-08-4199. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Grechez-Cassiau A, Rayet B, Guillaumond F, et al. The circadian clock component BMAL1 is a critical regulator of p21WAF1/CIP1 expression and hepatocyte proliferation. J Biol Chem. 2008;283:4535–4542. doi: 10.1074/jbc.M705576200. [DOI] [PubMed] [Google Scholar]
- 22.Mullenders J, Fabius AW, Madiredjo M, et al. A large scale shRNA barcode screen identifies the circadian clock component ARNTL as putative regulator of the p53 tumor suppressor pathway. PLoS One. 2009;4:e4798. doi: 10.1371/journal.pone.0004798. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Wood PA, Du-Quiton J, You S, et al. Circadian clock coordinates cancer cell cycle progression, thymidylate synthase, and 5-fluorouracil therapeutic index. Mol Cancer Ther. 2006;5:2023–2033. doi: 10.1158/1535-7163.MCT-06-0177. [DOI] [PubMed] [Google Scholar]
- 24.Yang X, Downes M, Yu RT, et al. Nuclear receptor expression links the circadian clock to metabolism. Cell. 2006;126:801–810. doi: 10.1016/j.cell.2006.06.050. [DOI] [PubMed] [Google Scholar]
- 25.Yagita K, Tamanini F, van Der Horst GT, et al. Molecular mechanisms of the biological clock in cultured fibroblasts. Science. 2001;292:278–281. doi: 10.1126/science.1059542. [DOI] [PubMed] [Google Scholar]
- 26.Bergman LM, Birts CN, Darley M, et al. CtBPs promote cell survival through the maintenance of mitotic fidelity. Mol Cell Biol. 2009;29:4539–4551. doi: 10.1128/MCB.00439-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Galimberti F, Thompson SL, Ravi S, et al. Anaphase catastrophe is a target for cancer therapy. Clin Cancer Res. 2011;17:1218–1222. doi: 10.1158/1078-0432.CCR-10-1178. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Li S, Szymborski A, Miron MJ, et al. The adenovirus E4orf4 protein induces growth arrest and mitotic catastrophe in H1299 human lung carcinoma cells. Oncogene. 2009;28:390–400. doi: 10.1038/onc.2008.393. [DOI] [PubMed] [Google Scholar]
- 29.Keck JM, Summers MK, Tedesco D, et al. Cyclin E overexpression impairs progression through mitosis by inhibiting APC(Cdh1) J Cell Biol. 2007;178:371–385. doi: 10.1083/jcb.200703202. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Marrazzo E, Marchini S, Tavecchio M, et al. The expression of the DeltaNp73beta isoform of p73 leads to tetraploidy. Eur J Cancer. 2009;45:443–453. doi: 10.1016/j.ejca.2008.09.024. [DOI] [PubMed] [Google Scholar]
- 31.Rivera A, Mavila A, Bayless KJ, et al. Cyclin A1 is a p53-induced gene that mediates apoptosis, G2/M arrest, and mitotic catastrophe in renal, ovarian, and lung carcinoma cells. Cell Mol Life Sci. 2006;63:1425–1439. doi: 10.1007/s00018-006-5521-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.