Abstract
One of the key aspects of neuronal differentiation is the array of neurotransmitters and neurotransmitter receptors that each neuron possesses. One important goal of developmental neuroscience is to understand how these differentiated properties are established during development. In this paper, we use fluorescence activated cell sorting and RNA-seq to determine the transcriptome of the Drosophila CNS midline cells, which consist of a small number of well-characterized neurons and glia. These data revealed that midline cells express 9 neuropeptide precursor genes, 13 neuropeptide receptor genes, and 31 small-molecule neurotransmitter receptor genes. In situ hybridization and high-resolution confocal analyses were carried-out to determine the midline cell identity for these neuropeptides and the neuropeptide receptors. The results revealed a surprising level of diversity. Neuropeptide genes are expressed in a variety of midline cell types, including motoneurons, GABAergic interneurons, and midline glia. These data revealed previously unknown functional differences among the highly-related iVUM neurons. There also exist segmental differences in expression for the same neuronal sub-type. Similar experiments on midline-expressed neuropeptide receptor genes reveal considerable diversity in synaptic inputs. Multiple receptor types were expressed in midline interneurons and motoneurons, and, in one case, link feeding behavior to gut peristalsis and locomotion. There were also segmental differences, variations between the 3 iVUMs, and three hormone receptor genes were broadly expressed in most midline cells. The Drosophila Castor transcription factor is present at high levels in iVUM5, which is both GABAergic and expresses the short neuropeptide F precursor gene. Genetic and misexpression experiments indicated that castor specifically controls expression of the short neuropeptide F precursor gene, but does not affect iVUM cell fate or expression of Gad1. This indicates a novel function for castor in regulating neuropeptide gene expression.
Keywords: castor, CNS midline, Drosophila, FACS, neuropeptide, RNA-seq
Introduction
One of the key goals of developmental neuroscience is to understand the origins of neuronal and glial diversity. One of the important characteristics of differentiated neurons is their complement of neurotransmitters and neurotransmitter receptors. By coupling the isolation of specific CNS cell types and RNA-seq transcriptomic analysis with in situ hybridization/immunostaining of expressed genes, it is possible to characterize the differentiated properties of individual CNS cells. In this manner the following questions can be addressed: (1) do individual neurons express multiple neurotransmitters and neurotransmitter receptors, (2) how similar are different types of neurons (e.g. dopaminergic vs. GABAergic neurons), (3) how similar are related neurons (e.g. dopaminergic neurons in distinct regions of the CNS), (4) do homologous neurons differ in different segments or regions, (5) do glia express neurotransmitter or neurotransmitter receptors, and (6) what are the regulatory mechanisms that control differentiation? In this paper, we isolate a small set of neurons and glia that reside at the midline of the Drosophila embryonic CNS, perform RNA-seq to define the midline transcriptome, validate the information with high-resolution in situ hybridization to determine the cell-type specificity of each gene, and identify a transcriptional regulator of a peptide neurotransmitter.
The Drosophila CNS midline cells have been well-characterized and provide an excellent system to study the genetic origins of neuronal and glial diversity. In the late stage embryonic CNS, there are ~22 midline cells per segment (Fig. 2B). These include 2 peptidergic MP1 neurons, the dopaminergic H-cell interneuron, the glutamatergic H-cell sib interneuron, 3 GABAergic iVUM (Ventral Unpaired Median interneurons) interneurons, and 3 octopaminergic/glutamatergic mVUM motoneurons that innervate body wall muscles and the female reproductive system (Wheeler et al., 2006). These neurons are derived from a set of Midline Precursors (MPs) that each divide once to generate 2 neurons. MP1 generates the MP1 neurons, MP3 divides into H-cell and H-cell sib, and MP4-6 each divides into an iVUM/mVUM pair (iVUM4-6 and mVUM4-6) (Wheeler et al., 2008). A median neuroblast (MNB) generates ~8 embryonic neurons (MNB progeny; MNBp) and continues to divide post-embryonically (Truman and Bate, 1988; Wheeler et al., 2006). There are two distinct populations of midline glia (MG), the anterior midline glia (AMG) and posterior midline glia (PMG) (Dong and Jacobs, 1997; Wheeler et al., 2012; Wheeler et al., 2006). The PMG undergo apoptosis during late embryonic development and their function is unknown. Only a subset of the AMG survive, and they ensheath the axon commissures.
Fig. 2.
RNA-seq data accurately predicts CNS midline-enriched gene expression. (A) List showing 14–16 hour midline and non-midline FPKM values for 6 genes with enriched midline expression. (B) Diagram of a typical stage 16/17 midline segment showing midline cells depicted with the cell-type specific markers used in this manuscript. Anterior left, dorsal top. (C–G) Alkaline phosphatase in situ hybridization experiments of stage 16 whole-mount embryos showing strong CNS midline enrichment of (C) cry, (D) dmrt99B, (E) HLH3B, (F) per, and (G) vg. Horizontal views are shown, with anterior left. (H–K) Stage 16/17 3.7sim>mCD8.GFP embryos were stained with anti-GFP to visualize midline cells (blue), and hybridized with RNA probes (lower-case italics) or immunostained with antibodies (upper-case, non-italicized). Each image is a 1 μm confocal slice showing a sagittal view of the relevant portion of a single segment with anterior left and dorsal top. (H) cry colocalizes with odd in MP1 neurons. (I) dmrt99B and (J) vg colocalize with Zfh1 in mVUMs. Only 2 mVUMs can be seen in the confocal plane shown in (I). (K) HLH3B colocalizes with En in iVUMs. Non-midline HLH3B expression is also seen in this image. (L–N) Venn diagrams illustrating the extent of overlap of genes expressed with FPKM values ≥5 between: (L) the 6–8 hour (green) and 14–16 hour (red) midline samples; (M) midline (green) and non-midline (red) cells at 6–8 hours AEL; and (N) midline (green) and non-midline (red) cells at 14–16 hours AEL.
In addition to the cellular characterization of midline development, the molecular composition of the Drosophila CNS midline cells is well-defined. Previously, we analyzed the developmental distribution of 286 genes expressed in midline cells by in situ hybridization and immunostaining (Kearney et al., 2004; Wheeler et al., 2006). In addition, we analyzed the midline expression of 77 genes by confocal analysis, identifying the midline cell-type specificity of each during development (Wheeler et al., 2008; Wheeler et al., 2006). This data is accessible on the Drosophila CNS Midline Gene Expression Database (MidExDB; http://midline.bio.unc.edu;) (Wheeler et al., 2009b). Thus, each midline cell type can be identified at all stages of development, aiding in genetic analyses of midline cell development. Nevertheless, there has been relatively little characterization of the unique differentiated properties of each midline neuronal cell type, such as the presence and distribution of neuropeptides and neurotransmitter receptors.
In this paper, we use fluorescence activated cell sorting (FACS) to isolate midline cells at two major developmental periods and carry-out RNA-seq on each cell sample. The results were validated by comparison to the extensive knowledge of midline gene expression and by using in situ hybridization to verify midline expression of genes previously not known to be expressed in midline cells. In this paper, we focus on the midline expression of neurotransmitters and their receptors. The data indicate that 9 neuropeptide precursor genes are expressed in subsets of CNS midline cells, including MP1 neurons, iVUMs, MNB progeny, and surprisingly, MG. In addition, 31 small-molecule neurotransmitter receptors and 13 neuropeptide receptor genes are expressed in a variety of midline cell types. In terms of the expression patterns of these genes, there exist both segmental differences and differences in specific iVUMs and MNB progeny. Among these results, the short neuropeptide F precursor (sNPF) gene is expressed in iVUM5, which has high levels of the Castor (Cas) transcription factor compared to iVUM4 and iVUM6. Genetic analyses demonstrate that sNPF expression in iVUM5 is absent in cas mutant embryos, while other differentiated properties of iVUM5 are unaffected. Thus, we propose a novel role of Cas in regulating sNPF neuropeptide gene expression. The combined use of FACS cell purification, RNA-seq analysis, in situ hybridization, and genetics allows the detailed analysis of how unique neurons acquire their differentiated state.
Materials and methods
Drosophila strains and genetics
3.7sim-Gal4 (Xiao et al., 1996) and y1 w*; PinYt/CyO; P{UAS-mCD8-GFP.L}LL6 (Lee and Luo, 1999) were used to generate the w1118; 3.7sim-Gal4; UAS-mCD8.GFP strain (3.7sim>mCD8.GFP) used for cell sorting and in situ hybridization and immunostaining experiments. The w1118; 3.7sim-Gal4 UAS-tau.GFP strain (3.7sim>tau.GFP) (Brand, 1995; Wheeler et al., 2006) was also used for in situ hybridization and immunostaining experiments. UAS-cas strains (Kambadur et al., 1998) were provided by W. Odenwald (NINDS, Bethesda, MD), and the cas24 and cas39 mutant strains (Cui and Doe, 1992) were provided by C. Doe (University of Oregon, HHMI).
Collection and dissociation of embryos for cell sorting
3.7sim>mCD8.GFP flies were propagated at low density in 6 oz bottles with cornmeal-based food at 25°C. For embryo collections, recently eclosed flies were transferred to 900 cm3 cages and acclimated for several days. Typically, 3 bottles were used to populate each cage, and 6–8 cages were used for each sample. On collection days, 100 mm grape agar plates with yeast paste were changed 3X for 2 hours/each to clear older embryos. Embryos were then collected for 2 hours on fresh yeasted grape agar plates and aged either 6–8 hours or 14–16 hours after egg laying (AEL) at 25°C. After staging, embryos were dechorionated with 100% bleach for 45 sec for 6–8 hour embryos or 1 min for 14–16 hour embryos, washed with 1X phosphate buffered saline (PBS) + 0.1% Tween-20, and placed on ice in 1X PBS. An aliquot of embryos was removed for immediate fixation and subsequent staging analysis. The remaining embryos were split into 1.5 ml microfuge tubes with ~25 μl packed embryos/tube. Generally, a total of 75–150 μl of packed embryos were used for each sample. PBS was removed and 100 μl of chilled dissociation buffer (9:1 10X Trypsin-EDTA:10X PBS) was added. Embryos were gently ground with a plastic pestle, and 400 μl of additional dissociation buffer was added. The cell solution was rocked on a nutator at room temperature for 10 min for 6–8 hour embryos and 20 min for 14–16 hour embryos. Cells were occasionally disrupted by pipetting. To terminate the trypsin digestion, tubes were placed on ice and 25 μl of heat-inactivated fetal bovine serum was added. Cells were filtered through a 41 μm nylon net into new microfuge tubes and 500 μl of chilled FACS buffer (1X PBS; 2% BSA; 2.5 mM EDTA) was added (FACS buffer was filtered through an 0.22 μm filter prior to use to remove particles that are detected by the flow cytometer). Cells were pelleted at 4°C for 4 min at 700 × g, washed with 500 μl chilled FACS buffer, pelleted, and resuspended in 200 μl of chilled FACS buffer. At this point, all tubes corresponding to a single embryo collection were combined, cells were filtered through a 41 μm nylon net, counted on a hemacytometer, and diluted with chilled FACS buffer to a concentration of 106 cells/ml. Cells were kept on ice in the dark until sorted.
Cell sorting and cDNA library preparation
Cell sorting was performed on the same day as the embryo dissociation. Prior to sorting, propidium iodide was added to a final concentration of 1 μg/ml. Cells were sorted into populations of GFP+ midline cells and GFP− non-midline cells on a Sony/iCyt Reflection cell sorter at the Flow Cytometry Core Facility (http://flowcytometry.med.unc.edu) at UNC Chapel Hill School of Medicine. Cell sorting was based on forward scatter and back scatter properties, the presence or absence of propidium iodide fluorescence to exclude dead or ruptured cells, and GFP fluorescence intensity. Cells were sorted directly into RNeasy Mini Kit Lysis Buffer (Qiagen) and frozen in a dry ice/ethanol bath. Before and after the main sort, an aliquot of cells were sorted into PBS and analyzed in the flow cytometer to estimate the accuracy of sorting.
Total RNA was prepared using the RNeasy Mini Kit, treated with RNase-free DNase (Qiagen), and re-purified with the RNeasy Mini Kit. mRNA was isolated using the Poly(A)Purist MAG Kit (Ambion), fragmented at 70°C for exactly 2 min with 10X Fragmentation Reagent (Ambion), and precipitated. First-strand cDNA was generated with SuperScript III Reverse Transcriptase (Invitrogen) using random hexamers. Second-strand cDNA was generated and purified according to the “mRNA Sequencing, Sample Preparation Guide” (Illumina; Rev. D, Sept. 2009) using 5X Second Strand Buffer (Invitrogen), DNA polymerase I (New England BioLabs; NEB), and the QIAquick PCR Purification Kit (Qiagen). End repair of cDNA fragments, adenylation of 3′ ends, and ligation of adapters were performed as described in “Preparing Samples for Sequencing Genomic DNA/Chip-Seq (v2.5)” provided by the UNC High-Throughput Sequencing Facility (HTSF). Adapters were obtained from the HTSF, and all other reagents were obtained from NEB. Purification of samples between steps was done using Qiagen kits as described in the Illumina protocol mentioned above. Adapter-ligated cDNA fragments (200–300 bp in length) were purified from a 2% agarose gel using the QIAquick Gel Extraction Kit. The adapter-ligated fragments were enriched with 18 cycles of PCR using Phusion High-Fidelity DNA polymerase (NEB) with primers obtained from the HTSF. The generated cDNA library was purified with the QIAquick PCR Purification Kit and sequenced on an Illumina GAIIx at the HTSF to generate single-ended 76 nucleotide reads.
Alignment of high-throughput sequencing reads
From the Illumina-generated fastq files, sequence reads were aligned to the D. melanogaster genome (build 5) using Bowtie (version 0.12.7) and TopHat (version 1.3.1.OSX_x86_64) with the following non-default parameters: -i 20 -I 142000 -g 1 --solexa1.3-quals (Langmead et al., 2009; Trapnell et al., 2009). The chosen minimum (-i) and maximum (-I) allowable intron lengths were based on known, annotated intron lengths in the D. melanogaster genome as reported on FlyBase (McQuilton et al., 2012). Only reads mapping uniquely to the genome were considered (-g 1). Cufflinks (version 1.0.3.OSX_x86_64) was run with the D. melanogaster reference annotation r5.39 to associate mapped reads with genes and calculate expression values (Trapnell et al., 2010). A mask file was used to eliminate the mapping of reads to rRNA, tRNA, snRNA, snoRNA, snmRNA, and ribosomal protein genes during this process.
All samples were processed individually; however, the two CNS midline samples from each time-point were also processed together in TopHat as replicates (Table 1). In the analyses that follow, a gene was considered to be expressed in the CNS midline cells above a certain threshold value if the following two conditions were met: (1) The average of the individual FPKM values (Fragments Per Kilobase of exon per Million fragments mapped) for that gene from both CNS midline samples was above the threshold value. (2) The FPKM value for that gene when the two CNS midline samples were processed as replicates was above the threshold value. All CNS midline expression values reported here are the FPKM values obtained when the two CNS midline samples were processed as replicates.
Table 1.
Summary of FACS and RNA-seq. For each sample, the numbers of cells isolated via FACS, 76 nucleotide sequencing reads generated from the cDNA library, and reads mapping uniquely to the genome are shown.
| Sample Name | Cell Population | Time-point | # of Sorted Events (Cells) | # of Reads Generated | # of Uniquely Mapped Reads |
|---|---|---|---|---|---|
| 6Mid1 | CNS Midline | 6–8 hours AEL | 20,000 | 7,162,776 | 4,471,612 |
| 6Mid2 | CNS Midline | 6–8 hours AEL | 40,000 | 8,179,000 | 6,213,434 |
| 6Mid1+2 | CNS Midline | 6–8 hours AEL | N/A | N/A | 10,701,922 |
| 6nonMid | Non-Midline | 6–8 hours AEL | 50,000 | 8,923,570 | 6,062,148 |
| 14Mid1 | CNS Midline | 14–16 hours AEL | 42,000 | 23,995,806 | 10,363,609 |
| 14Mid2 | CNS Midline | 14–16 hours AEL | 62,000 | 29,547,037 | 23,829,053 |
| 14Mid1+2 | CNS Midline | 14–16 hours AEL | N/A | N/A | 34,209,443 |
| 14nonMid | Non-Midline | 14–16 hours AEL | 50,000 | 23,758,803 | 15,216,077 |
In situ hybridization and immunostaining
Standard protocols for alkaline phosphatase and fluorescent in situ hybridization, and immunostaining were previously described (Kearney et al., 2004). These protocols are available on MidExDB (http://midline.bio.unc.edu/MDB_Display_protocols.aspx). For in situ hybridization experiments, digoxigenin or biotin-labeled RNA probes were generated from cDNA clones in the Drosophila Gene Collection Releases 1 and 2 (Berkeley Drosophila Genome Project) or from cDNA amplicons generated using a 3′ primer containing a T3 RNA polymerase binding site (Supplemental Tables S1, S2). RNA detection was carried-out using anti-digoxigenin-AP Fab fragments (Roche), anti-Digoxigenin-POD Fab fragments (Roche), streptavidin-HRP (PerkinElmer), and the Tyramide Signal Amplification (TSA) system (PerkinElmer). For immunostaining, the primary antibodies used were: rabbit anti-GFP (Abcam), mouse anti-Tau (Sigma-Aldrich), rabbit anti-Cas (W. Odenwald, NINDS) (Kambadur et al., 1998), guinea pig anti-Zfh1 (Fortini et al., 1991), and mouse anti-En (4D9; Developmental Studies Hybridoma Bank). Secondary antibodies were conjugated with Alexa Fluor dyes (Invitrogen).
Data distribution
Raw sequencing read files and Cufflinks processed genes.fpkm_tracking files have been deposited in the Gene Expression Omnibus (GEO) at the National Center for Biotechnology Information (NCBI). FPKM values of genes are also accessible on MidExDB (http://midline.bio.unc.edu).
Results
Isolation of fluorescently-labeled CNS midline cells
To further understand the development and differentiated properties of Drosophila embryonic midline neurons and glia, we used RNA-seq to determine the midline cell transcriptome. The protocol involved using FACS isolation of GFP-labeled embryonic midline cells, followed by mRNA isolation and RNA-seq analysis. High-quality, biologically-relevant results require careful attention to multiple criteria. (1) Midline cell expression of the fluorescent marker (GFP) must be sufficiently strong for the cell sorter to distinguish labeled cells from unlabeled cells. (2) There should be minimal expression of the fluorescent marker outside of the CNS midline. (3) Fluorescent marker expression must be consistently strong in all midline cells throughout the developmental periods analyzed. (4) Expression of the transgenic marker should have no adverse effects on embryonic development. Multiple Gal4 and UAS lines were analyzed, and a 3.7sim-Gal4; UAS-mCD8.GFP (3.7sim>mCD8.GFP) strain best matched the criteria described above. This strain uses the early midline enhancer from the single-minded (sim) gene (Nambu et al., 1991; Xiao et al., 1996), which encodes a bHLH-PAS protein and is a master regulator of CNS midline gene expression (Nambu et al., 1991).
The two embryonic developmental time-points selected for this study were: (1) 6–8 hours after egg laying (AEL) (stages 11–12), which represents a time when the CNS midline neurons have recently arisen from their precursors and are acquiring their respective cell identities and beginning to differentiate, and (2) 14–16 hours AEL (stages 16–17), a time in which CNS midline neurons and glia are well-differentiated (Wheeler et al., 2008; Wheeler et al., 2006). Staining 3.7sim>mCD8.GFP embryos with anti-GFP revealed that embryos had strong and specific GFP expression in all CNS midline cells at both 6–8 hours and 14–16 hours AEL (Fig. 1A–D). Furthermore, >99% of 3.7sim>mCD8.GFP embryos aged 6–8 hours and 14–16 hours AEL were at the proper stages of development.
Fig. 1.
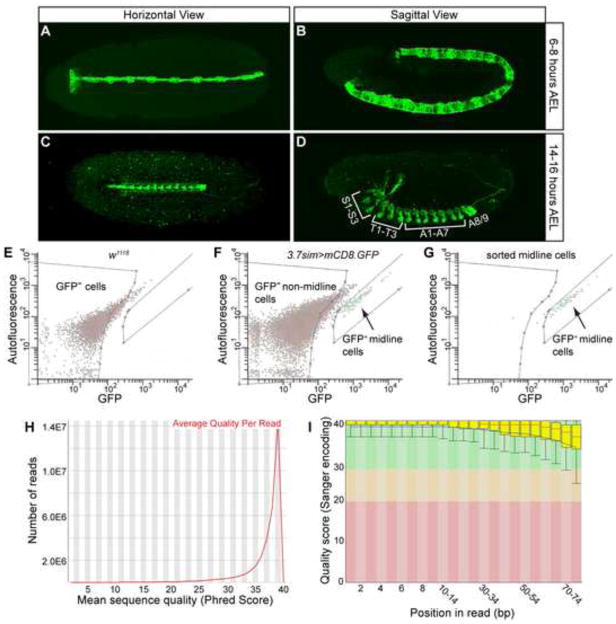
High-quality RNA-seq reads were obtained from purified CNS midline cells. (A–D) 3.7sim>mCD8.GFP embryos were stained with anti-GFP (green) and imaged by confocal microscopy. Images represent (A,C) horizontal views and (B,D) sagittal views of (A,B) 6–8 hour AEL embryos and (C,D) 14–16 hour AEL embryos. (C) The speckled GFP signal seen outside of the midline is from mVUM neurites. (D) Midline expression of GFP is present in all midline cells from segments S1 to A8/9. (E–G) Representative histograms plotting fluorescence profiles of cells from (E) dissociated w1118 embryos, (F) dissociated 3.7sim>mCD8.GFP embryos, and (G) sorted GFP-positive cells from 3.7sim>mCD8.GFP embryos. Purities of >95% were achieved in sorting GFP-positive cells. Boxes indicate gates used for sorting GFP− and GFP+ cells. (H,I) Representative charts generated using FastQC depicting: (H) the average quality of individual sequencing reads, and (I) the quality of base calls at each position in a sequencing read across all reads.
Dissociated embryonic cells were FACS-sorted into populations of GFP+ CNS midline cells and GFP− non-midline cells (Fig. 1E,F). GFP+ midline cells were successfully sorted with >95% purity, based on pre- and post-sorting checks of accuracy on the flow cytometer (Fig. 1G). Two independently-collected CNS midline samples and one non-midline sample were analyzed for each time-point (Table 1).
mRNA sequencing and data validation
For each of the sorted cell samples, mRNA was isolated and sequenced on an Illumina GAIIx. Single-ended 76 nucleotide reads were generated, and analyses with FastQC software indicated high quality reads (Fig. 1H,I). Uniquely mapping reads were aligned to the annotated D. melanogaster genome using the Bowtie, TopHat, and Cufflinks programs (Langmead et al., 2009; Trapnell et al., 2010; Trapnell et al., 2009).
Surveying well-characterized genes with no known midline expression indicated that these genes had FPKM (Fragments Per Kilobase of exon per Million fragments mapped) values <5 in the CNS midline RNA-seq data (Supplemental Table S3). These genes include those expressed in the lateral CNS, sensory cells, midline accessory cells (channel glia, dorsal median cells), muscle, and trachea. Previous studies have characterized the CNS midline cell-type specific expression of 77 genes by confocal microscopy (Wheeler et al., 2009a; Wheeler et al., 2006). To initially validate the fidelity of the midline RNA-seq data, the FPKM values for these midline-expressed genes were assessed. Of the 77 genes, 75 are expressed between stages 11–17, the time-points used in our experiments. All 75 of these genes had an FPKM value >5 in either the 6–8 hour or 14–16 hour samples (or both) (Supplemental Table S4). Thus, we designate genes with a midline FPKM value ≥5 as “expressed” in midline cells, and an FPKM value <5 as “not expressed, or expressed at a low level.” This MidExDB data set included genes that are expressed in a single midline cell type. Since these genes often showed high midline FPKM values in the RNA-seq data, these results further confirm the sensitivity of the RNA-seq data and that all cell types were present in the sorted CNS midline samples.
RNA-seq data predicts CNS midline gene expression
To test the predictive abilities of the RNA-seq data, we addressed whether these data could identify factors that are enriched in the midline cells. To concentrate on transcription factors and transcriptional regulators, we first generated a list of 859 genes that contain the term “transcription” in their FlyBase gene ontology entries. We find that 508 of these genes are expressed in the 14–16 hour midline sample with FPKM values ≥5 and 292 with FPKM values ≥20. We then searched this list for genes with FPKM values ≥20 in the 14–16 hour midline sample and <5 in the 14–16 hour non-midline sample, reasoning that a 4-fold difference of FPKM values will identify genes with strong midline expression and relatively low expression outside the midline when analyzed by in situ hybridization. This search identified 6 genes: cryptochrome (cry), doublesex-Mab related 99B (dmrt99B), HLH3B, period (per), sim, and vestigial (vg) (Fig. 2A) (note that Drosophila cry encodes a blue light photoreceptor, and is not a transcription factor in Drosophila). Analysis in vivo by in situ hybridization confirmed that the expression of all 5 genes, in addition to sim, is highly enriched in the CNS midline (Fig. 2C–G). These genes were further analyzed for midline cell expression using fluorescent in situ hybridization in combination with midline cell type-specific markers (Fig. 2B). The two circadian rhythm-associated genes, cry and per, show expression in non-overlapping midline neurons. Expression of the cry gene colocalizes with the MP1 neuron marker odd skipped (odd) (Fig. 2H), whereas per expression is specific to H-cell sib, iVUM4-6, and the MNBp, as previously shown for a per-Gal4 transgenic strain (Wheeler et al., 2006). Transcripts for dmrt99B and vg both colocalize with Zinc finger homeodomain 1 (Zfh1) protein, indicating expression in the 3 mVUM motoneurons (Fig. 2I,J) (Guss et al., 2008). Expression of HLH3B was observed in Engrailed (En)+ iVUMs (Fig. 2K) and H-cell sib, which has high levels of Vesicular glutamate transporter (VGlut) expression (data not shown). Thus, the ability of the RNA-seq and in situ hybridization data to identify transcriptional regulatory genes with high midline specificity expands the list of known transcription factors that may control midline neuron-specific gene expression.
Quantitative analysis of the CNS midline transcriptome
The examination of the midline RNA-seq data indicated that both 6–8 hour and 14–16 hour midline samples are a valid representation of the midline transcriptome, allowing a quantitative assessment of midline transcription during those two time periods. At the time of these experiments, there were 13909 protein coding genes annotated for D. melanogaster (McQuilton et al., 2012). When examining genes for which reliable RNA-seq data were obtained for both time-points, we see 6094 genes expressed in the CNS midline at 6–8 hours AEL with FPKM values ≥5, and 6529 genes at 14–16 hours AEL. Of these genes, 5041 (66.5%) were expressed at both time-points with 2541 genes (33.5%) expressed in only one time interval (Fig. 2L). The large difference in gene expression between the 2 stages reflects the rapid changes in development that occur as the cells transition from a group of cells at stages 11–12 that are undergoing dramatic changes in cell fate, migration, and apoptosis to differentiated neurons and glia (stages 16–17). Comparing CNS midline to non-midline gene expression, we find that at either time-point, most genes (79–84%) expressed with FPKM values ≥5 are shared between the 2 samples, while only 4–6% of genes are specific to the CNS midline cells (Fig. 2M,N). Not surprisingly, more genes (10–17%) are expressed only in non-midline cells compared to those only expressed in midline cells.
CNS midline neurons utilize a diverse array of neuropeptides
The CNS midline neurons have been well-characterized with respect to the neurotransmitters they utilize: MP1 neurons (Pigment-dispersing factor (Pdf) and Proctolin (Proct)), H-cell (dopamine), H-cell sib (glutamate), iVUMs (GABA), and mVUMs (glutamate, octopamine) (Wheeler et al., 2006). Consistent with these assignments, the RNA-seq data generally showed high levels of expression for genes involved in the generation, metabolism, and transport of these neurotransmitters (data not shown). The RNA-seq data was further screened for the utilization of additional neurotransmitters. None of the midline neurons are likely to utilize acetylcholine, histamine, or serotonin, based on a lack of expression of Choline acetyltransferase (FPKM=3.76), Histidine decarboxylase (FPKM=0.79), or Serotonin transporter (FPKM=2.62). However, analysis of 33 Drosophila neuropeptide precursor genes in the 14–16 hour RNA-seq data revealed that 12 of these genes have FPKM values ≥5, including Pdf and Proct (Fig. 3A,B). Here, we characterize the expression of the remaining 10 neuropeptide precursor genes in midline segments S1-A7, and arrange their expression by cell type (Fig. 3L).
Fig. 3.
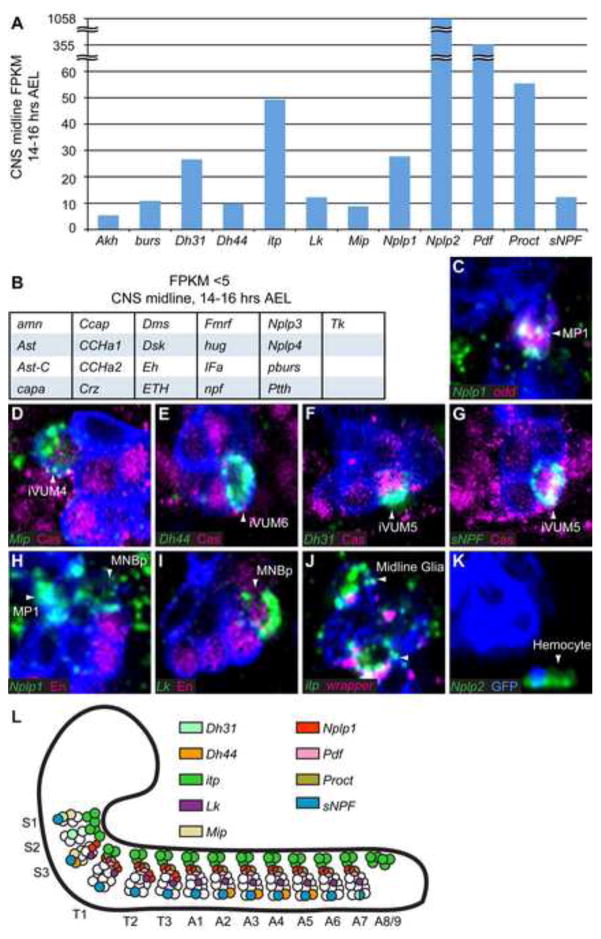
Neuropeptide precursor gene expression in the CNS midline cells. (A) 12 neuropeptide precursor genes have CNS midline FPKM values ≥5 at 14–16 hours AEL, while (B) 21 genes have FPKM values <5. (C–K) In situ hybridization of 8 neuropeptide precursor genes (green) are shown in sagittal views of stage 16/17 3.7sim>GFP embryos. (C,H–K) mCD8.GFP was detected with anti-GFP (blue), and (D–G) tau.GFP was detected with anti-Tau (blue). Various midline cell markers are shown in magenta, and arrowheads indicate the neuropeptide gene-expressing cells. Anterior left, dorsal top. (C) Nplp1 is expressed in odd+ MP1 neurons. (D) Mip is expressed in Caslo iVUM4, (E) Dh44 in Cas− iVUM6, and (F) Dh31 and (G) sNPF in Cashi iVUM5 neurons. (H) Nplp1 is also expressed in an En− MNBp neuron, while (I) Lk is expressed in an En+ MNBp neuron. (J) Expression of itp is in wrapper+ AMG. (K) Nplp2 expression is seen in hemocytes that have engulfed apoptotic GFP-expressing midline cells. (L) Diagram of segmental neuropeptide expression in the CNS midline at 14–16 hours AEL.
MP1 neurons
The Neuropeptide-like precursor 1 (Nplp1) gene is expressed in odd+ MP1 neurons in all segments (Fig. 3C). Along with Pdf and Proct, this increases the number of neuropeptide genes expressed in MP1 neurons to three. The MP1 neurons undergo apoptosis in segments S3-A4 at late embryogenesis (Miguel-Aliaga and Thor, 2004). In larval segments A8/9, the MP1s likely innervate the hindgut, suggesting that Pdf, Proct, and Nplp1 peptides may regulate hindgut activity.
iVUM neurons
Four neuropeptide precursor genes are expressed in subsets of the GABAergic iVUM interneurons. iVUMs are local interneurons of unknown function. Myoinhibiting peptide precursor (Mip), which has been implicated in ecdysis in Drosophila and inhibiting ecdysteroid synthesis in other insects (Kim et al., 2006; Veelaert et al., 1996), is expressed in only 1 En− cell of segment S1 and 1 cell of segment S3. While the lineages of the suboesophageal midline cells have not been characterized in detail, the Mip+ cell in segment S3 is En+ Caslo, indicative of iVUM4 neurons (Fig. 3D). The diuretic hormone precursor genes Dh31 and Dh44, are also expressed in iVUM neurons. Diuretic hormone peptides are implicated in peristalsis and in controlling water and ion balance by acting on the Malpighian tubules (Cabrero et al., 2002; Coast et al., 2001; LaJeunesse et al., 2010). Dh44 is expressed in iVUM6 (En+ Cas−) of segments A2-A5 and an En+ Cas− cell of segment S3 (Fig. 3E), while Dh31 is expressed in iVUM5 (En+ Cashi) of segment A7 and an En− Cas+ cell of segments S1 and S2 (Fig. 3F). The short neuropeptide F precursor (sNPF) gene, which encodes peptides involved in feeding behavior (Lee et al., 2004), is expressed in iVUM5 (En+ Cashi) of segments S3-A7, and an En+ cell of segment S1 (Fig. 3G). These results revealed several novel findings. (1) The GABAergic iVUM interneurons likely utilize a variety of neuropeptides in neurotransmission. (2) Each neuropeptide is expressed only in a specific iVUM – either iVUM4, iVUM5, or iVUM6. This indicates that each iVUM is distinct physiologically, despite their general similarities in gene expression and axonogenesis. (3) There exist segmental differences in the embryonic expression of each neuropeptide with the most extreme case being Mip, which is only expressed in segments S1 and S3, indicating there are functional differences between iVUM neurons of different segments. It remains possible that expression of these neuropeptide genes may emerge in additional segments later in development, but even so, there would still exist segmental differences in neuropeptide gene appearance.
MNB progeny
In segments S3-T3, we observed Nplp1 expression in 2 En− MNBp (Fig. 3H). Expression of Leucokinin (Lk), which has been implicated in regulating diuresis and meal size (Al-Anzi et al., 2010; Terhzaz et al., 1999), is present in a single En+ MNBp in segments S3-A7 (Fig. 3I).
MG
In Drosophila, the neuropeptide ion transport peptide (itp) is expressed in dorsal and ventral lateral clock neurons (Dircksen et al., 2008; Johard et al., 2009). While its function in these neurons is unknown; in locusts, Itp has been shown to play an antidiuretic role, involved in the reabsorption of ions and fluids in the hindgut (Audsley et al., 1992; King et al., 1999). Interestingly, in the CNS midline, itp is not expressed in neurons, but in MG (Fig. 3J). Expression was present in AMG, but not PMG, in all segments.
Non-midline RNA-seq+ genes
Three neuropeptide precursor genes, bursicon (burs), Neuropeptide-like precursor 2 (Nplp2), and Adipokinetic hormone (Akh) were not detected by in situ hybridization in CNS midline cells, yet were present in the 14–16 hour midline RNA-seq data with FPKM values ≥5. The 3′ end of the coding sequence of the burs gene is 139 bp downstream of the annotated 3′-UTR of the GABA-B-R2 gene, which has an FPKM value of 87.69 in the 14–16 hour midline sample (Table 2). One possibility is that the 3′-UTR of GABA-B-R2 may overlap the burs gene leading to burs being scored as midline-expressed. Expression of Nplp2 is observed in hemocytes that are also GFP+ (Fig. 3K). During embryogenesis, ~70% of MG undergo apoptosis and are engulfed by hemocytes (Sonnenfeld and Jacobs, 1995; Wheeler et al., 2012). Consequently, hemocytes that engulf GFP+ MG will be FACS-sorted into the GFP+ sample, explaining the occurrence of high Nplp2 FPKM values. For Akh, the obtained FPKM value is 5.34, just above the threshold for considering a gene as “expressed” in the midline, and it likely represents a false-positive.
Table 2.
Midline FPKM values for small-molecule neurotransmitter receptors at 14–16 hours AEL. Genes with FPKM values ≥5 are shown on left, and those with FPKM values <5 are shown on right.
| 14–16 hr midline FPKM ≥5 | 14–16 hr midline FPKM <5 | ||||||
|---|---|---|---|---|---|---|---|
| Symbol | FPKM | Gene Symbol | FPKM | Gene Symbol | FPKM | Gene Symbol | FPKM |
| 5-HT1A | 74.82 | GABA-B-R3 | 10.03 | nAcRα-96Ab | 100.85 | clumsy | 1.12 |
| 5-HT1B | 8.52 | gfA | 36.58 | nAcRβ-21C | 7.41 | GluRIIA | 1.08 |
| 5-HT2 | 5.44 | Glu-RI | 50.34 | nAcRβ-64B | 264.02 | GluRIIB | 0.13 |
| 5-HT7 | 15.43 | Lcch3 | 83.81 | nAcRβ-96A | 97.23 | GluRIIC | 0.89 |
| CG3822 | 88.53 | mAcR-60C | 73.04 | Nmdar1 | 69.71 | HisCl1 | 1.01 |
| CG42796 | 5.21 | mGluRA | 7.67 | Nmdar2 | 77.55 | ort | 0.16 |
| D2R | 50.92 | nAcRα-30D | 87.68 | Oamb | 20.64 | TyrR | 2.54 |
| DopR | 10.49 | nAcRα-34E | 109.21 | Octβ2R | 62.22 | ||
| DopR2 | 6.58 | nAcRα-7E | 10.03 | Rdl | 166.58 | ||
| GABA-B-R1 | 39.71 | nAcRα-80B | 9.04 | ||||
| GABA-B-R2 | 87.69 | nAcRα-96Aa | 174.64 | ||||
Neuropeptide receptors are expressed in diverse CNS midline neuronal cell types
Previous work has identified a variety of neurotransmitter receptor genes expressed in midline neurons, including Glu-RI, 5-HT1A, and NPFR1 (Wheeler et al., 2006). We have utilized the midline 14–16 hour RNA-seq data to further characterize neurotransmitter receptor expression in midline cells. Of 38 neurotransmitter receptors genes analyzed, 31 have midline FPKM values ≥5 at 14–16 hours AEL (Table 2), including Glu-RI and 5-HT1A. This suggests that the midline cells are able to respond to a wide variety of neural inputs. Complementing this analysis, we also analyzed the expression of 19 Drosophila neuropeptide receptors and found that 13 have 14–16 hour midline FPKM values ≥5, including NPFR1 (Fig. 4A,B). Using in situ hybridization, we further investigated their expression in segments T1-A7.
Fig. 4.
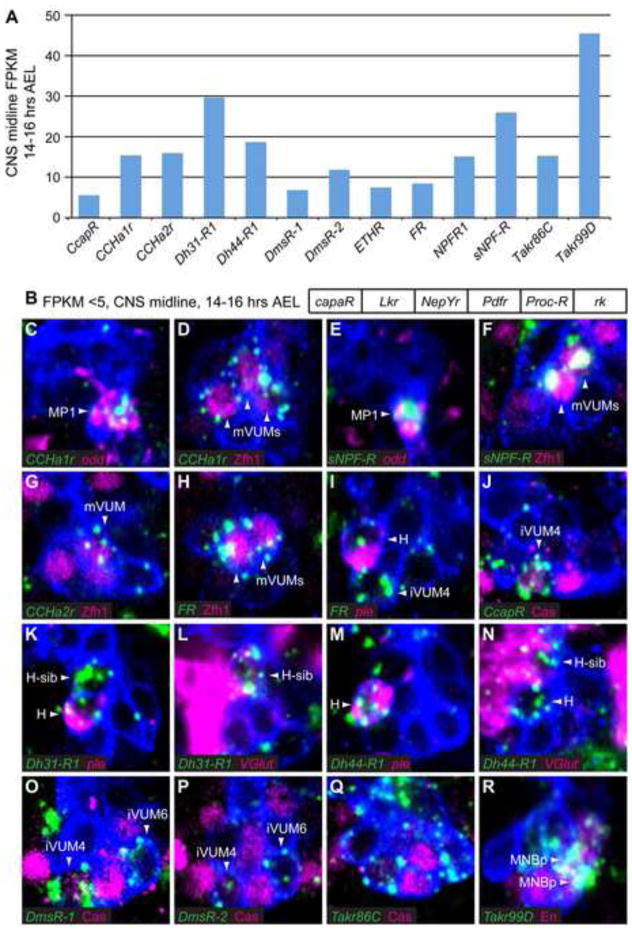
Neuropeptide receptor expression in the CNS midline cells. (A) 13 neuropeptide receptor genes have CNS midline FPKM values ≥5 at 14–16 hours, while (B) 6 neuropeptide receptor genes have FPKM values <5. (C–R) In situ hybridization of 11 neuropeptide precursor genes (green) are shown in sagittal views of stage 16/17 3.7sim>GFP embryos. (C–I,K–N,R) mCD8.GFP was detected with anti-GFP (blue), and (J,O–Q) tau.GFP was detected with anti-Tau (blue). Midline cell markers are shown in magenta. Arrowheads indicate the neuropeptide receptor-expressing cells. Anterior left, dorsal top. (C,E) CCHa1r and sNPF-R expression is present in MP1 neurons. (D,F–H). CCHa1r, sNPF-R, CCHa2r, and FR expression is present in mVUMs. (I) FR expression is also present in H-cell and iVUM4 of several segments. (J) CcapR is expressed in Caslo iVUM4. (K,M) Dh31-R1 and Dh44-R1 colocalize with ple in H-cell neurons and (L,N) with high VGlut in H-cell sib neurons. (O,P) Expression of DmsR-1 and DmsR-2 is observed in Caslo iVUM4 and Cas− iVUM6. (Q,R) Takr86C and Takr99D expression is present in most midline neurons with Takr99D expression highest in the MNBp (arrowheads).
Motoneurons
The CCHamide-1 receptor (CCHa1r) and short neuropeptide F receptor (sNPF-R) genes are both expressed in MP1 and mVUM motoneurons (Fig. 4C–F). Drosophila sNPF is implicated in appetite control and food uptake (Lee et al., 2004), and presence of the sNPF receptor on MP1s and all 3 mVUMs links feeding behavior to midline-controlled gut peristalsis (MP1s) and locomotion (mVUMs). The CCHamide-2 receptor (CCHa2r) is also expressed in mVUMs, although its expression is restricted to abdominal segments and is highest in the posterior-most mVUM6 neurons (Fig. 4G). The Fmrf Receptor (FR) is expressed in all 3 mVUM neurons of segments T3-A7 (Fig. 4H).
Interneurons
Midline interneurons also express a variety of neuropeptide receptors. Expression of FR is segmentally restricted in several interneurons, with expression seen in iVUM4 neurons of segments T1–T2 and H-cell neurons of segments T1-A1 (Fig. 4I; data not shown). Cardioacceleratory peptide receptor (CcapR) showed the most restricted expression domain in the CNS midline of all neuropeptide receptors examined, with expression present in iVUM4 neurons of segments A5–A7 (Fig. 4J). The diuretic hormone receptor genes Dh31-R1 and Dh44-R1, have overlapping midline expression patterns, with both present in H-cell and H-cell sib (Fig. 4K–N). The Dromyosuppressin receptor genes DmsR-1 and DmsR-2 show overlapping patterns of expression in iVUM4 and iVUM6, but not iVUM5 (Fig. 4O,P).
Broad expression
Three hormone receptor genes are broadly expressed in the CNS midline neurons. The Tachykinin-like receptor genes Takr86C and Takr99D are expressed in most midline neurons (Fig. 4Q,R), with Takr99D expressed at highest levels in an En+ MNBp. Ecdysis-triggering hormone receptor (ETHR) is the most broadly expressed of the neuropeptide receptors, with expression in all midline neurons and MG (data not shown). ETHR has two splice variants, ETHR-RA and RB, (Iversen et al., 2002). Our 14–16 hour midline RNA-seq data show both variants expressed at roughly equal levels (FPKM values of 4.3 and 3.2 for RA and RB, respectively), similar to the whole animal RNA-Seq data (Graveley et al., 2011).
Castor is a positive regulator of sNPF expression in iVUM5
The ability to assign expression profiles for each neuropeptide and neurotransmitter receptor to specific midline cells facilitates the genetic analysis of their regulation. The most interesting aspect of sNPF expression is that it is present in only iVUM5 and not iVUM4 or iVUM6. While most characterized transcription factors are expressed in all 3 iVUMs, levels of the Castor (Cas) zinc finger transcription factor protein vary for each iVUM: being low in iVUM4, high in iVUM5, and absent in iVUM6 (Fig. 5A) (Wheeler et al., 2008). The high levels of Cas in iVUM5 suggest that it might control sNPF expression. Examining two castor protein-null alleles, cas24 and cas39, there was a complete absence of iVUM5-specific sNPF expression in both homozygous and trans-heterozygous mutant embryos (Fig. 5B,C). This effect is specific to the midline, since non-midline CNS expression of sNPF is present in cas mutant embryos (Fig. 5B′,C′). In general, midline neurons and glia are present in cas mutants, indicating no gross disorganization. More specifically, expression of en and Glutamic acid decarboxylase 1 (Gad1) is unaffected in cas mutant embryos in all 3 iVUMs (Fig. 5F–I). This is consistent with iVUM neurons being properly specified and undergoing differentiation to become GABAergic in the cas mutants. Therefore, the defect in iVUM5 neuronal differentiation in cas mutants is limited to the loss of sNPF expression for the genes analyzed.
Fig. 5.
Expression of sNPF in iVUM5 is regulated by cas. Sagittal views of stage 16/17 3.7sim-Gal4 UAS-tau.GFP embryos showing a sagittal view of a single segment of the (A–G) CNS midline cells or (B′-E′) corresponding slices ~8 μm lateral to the midline. Embryos are stained with anti-Tau (blue) and anti-Cas (magenta) and hybridized to (B–E) an sNPF probe (green) or (F,G) a Gad1 probe. In wild-type embryos, sNPF expression is (B) present in iVUM5 (arrowhead), and (B′) present in a lateral CNS Cas− cell (dotted circle). Arrow shows non-midline Cas protein in wild-type embryos. (C) In cas24/cas39 mutant embryos, sNPF expression is absent in iVUM5, although (C′) sNPF is present in lateral neurons (dotted circle). (D,D′) In 3.7sim-Gal4 UAS-tau.GFP/UAS-cas, misexpression of cas occurs in all midline cells, yet has no effect on sNPF expression. (E,E′) In 3.7sim-Gal4 UAS-tau.GFP/UAS-cas; cas24/cas39, misexpression of cas throughout the CNS midline in a cas mutant background rescues CNS midline expression of sNPF in iVUM5 (arrowhead). (F) In a wild-type embryo, Gad1 is expressed at high levels in iVUM4-6 (arrowheads) and MNBp. (G) In cas24 mutant embryos, Gad1 expression is unaffected and is present in all iVUMs, including iVUM5. (H,I) In both (H) wild-type and (I) cas39 mutant embryos, En is present in iVUM neurons (arrowheads). Note the absence of Cas immunostaining in panels G and I indicating that these embryos were homozygous mutant for cas.
Misexpression of high levels of cas in all midline neurons and glia in 3.7sim-Gal4 UAS-cas embryos did not result in ectopic sNPF expression (Fig. 5D), indicating that other factors, in addition to high levels of Cas, are necessary for sNPF expression. Supporting this hypothesis, midline misexpression of cas (3.7sim-Gal4 UAS-cas) in cas mutants completely rescues expression of sNPF in only a single cell per midline segment (Fig. 5E). One model that would explain these observations includes a transcriptional activator that is present in iVUM5, but absent from iVUM4 and iVUM6. In this model, both Cas and a second coactivator are necessary for sNPF expression. A second model proposes the presence of a transcriptional repressor in iVUM4 and iVUM6. This repressor would keep sNPF expression off in these cells, even when cas is misexpressed at high levels. The cas gene also controls development of the Drosophila Lk, FMRFa, Nplp1, and Capa peptidergic neurons (Benito-Sipos et al., 2010; Gabilondo et al., 2011; Losada-Pérez et al., 2010). However, in those cases, it is proposed that cas is acting as a temporal identity gene required for neuronal cell fate, whereas in iVUM5, we propose that cas does not direct neuronal cell fate, but influences differentiation – specifically neuropeptide expression.
Discussion
In this paper, we have combined the use of FACS purification of midline cells, RNA-seq, and in situ hybridization to study the differentiation of midline neurons. Based on this data, we have defined a midline-expressed gene as having an FPKM ≥ 5 in the midline GPF+ samples. Examining the RNA-seq data for expression values of 75 genes known to be expressed in midline cells, we found that all 75 had an FPKM > 5 (5.30 – 1351.15). Similarly, we examined 24 genes that are not thought to be expressed in midline cells, and they all had an FPKM < 5 (0.00 – 4.98). As a test of our RNA-seq data, we used fluorescent in situ hybridization to examine the expression of 22 neuropeptide precursor and neuropeptide receptor genes with FPKM values > 5 (5.34 – 1058). Of the 22 genes tested, 19 had midline expression. Of the three genes that had undetectable midline expression, one was expressed in hemocytes and was likely detected because they had engulfed a GFP+ midline cell, one may represent an annotation error due to proximity to a midline–expressed gene, and the other had an FPKM just barely above our threshold of 5. Thus, the false negative rate from our analyses is 0% (0/75) and the false positive rate is <14% (3/22). Consequently, the RNA-seq data and our threshold designations are strong indicators of midline expression.
The expression of neuropeptide precursor genes in a variety of midline neurons indicate that these neurons commonly co-utilize small molecule and peptide neurotransmitters. The 3 iVUMs are GABAergic interneurons with similar axonal trajectories. We showed that iVUM4 expresses Mip, iVUM5 expresses sNPF (plus Dh31 in segment A7), and iVUM6 expresses Dh44. Thus, despite near identity in their previously studied gene expression, iVUM4, iVUM5, and iVUM6 possess differences in gene expression and in their likely modes of neurotransmission. These results also indicate that iVUMs are peptidergic in addition to utilizing GABA as a neurotransmitter. Co-transmission of neuropeptides with small-molecule neurotransmitters has been documented in both vertebrates and invertebrates (Hökfelt et al., 2000; Burnstock, 2004; Nässel and Homberg, 2006). Recent work on the Drosophila brain central complex has described FMRFamide-related (Fmrf) neuropeptide gene expression in GABAergic neurons, Mip and Tachykinin (Tk) expression in cholinergic neurons, and sNPF expression in both cholinergic and glutamatergic neurons (Kahsai and Winther, 2011). In the case of iVUMs, it needs to be demonstrated that these cells generate neuropeptides that play functionally relevant roles in neurotransmission.
One unexpected observation is that midline expression of itp is present in MG, but absent in neurons. Neuropeptide expression in glia is observed in vertebrate systems and has been implicated in processes like axon guidance, neuronal survival, neuronal proliferation, and immunity (Ubink et al., 2003). To our knowledge, this is the first reported case in Drosophila of glial neuropeptide expression. Since MG are in close apposition to commissural axons (e.g. Wheeler et al., 2009a), as well as extend laterally through gliopodia (Vasenkova et al., 2006), it is possible that MG may be modulating neurotransmission, or playing some other developmental, physiological, or homeostatic role.
It is important to note that our expression analysis is relegated to late stage embryos, and that the patterns of expression may differ in larvae and adults. In fact, comparing our embryonic midline expression data to the comprehensive analysis of neuropeptide distribution in 3rd instar larvae (Park et al., 2008) reveals significant differences for most of the genes analyzed. In the larvae, there are no detectable levels of DH44, ITP, LK, NPLP1, or sNPF in midline cells, and even though DH31, PDF, and PROCT may be present in midline cells, their distribution is more restricted than embryonic precursor gene expression. It will be informative to examine neuropeptide appearance in 1st instar larvae to see whether the neuropeptide distribution fits with embryonic expression of the precursor genes. If so, this would indicate significant changes in neuropeptide gene expression during larval development. Alternatively, the 3rd instar levels of neuropeptide may be present in midline cells reflecting embryonic precursor gene expression, but too low to clearly detect. In many large neurosecretory cells of the nervous system, the bHLH transcription factor Dimmed (Dimm) acts as a scaling factor that up-regulates genes involved in neurosecretion (Mills and Taghert, 2012). Our RNA-seq data suggest that dimm is not highly expressed in the peptidergic neurons of the CNS midline cells at stages 16–17 (FPKM <5), consistent with the lack of anti-Dimm immunostaining in embryonic midline cells (Park et al., 2008), except in Pdf+ neurons in segment A8/9. This suggests that neuropeptide production in midline neurons may be relatively low, possibly reflecting differences in function from high-producing neurons.
Small-molecule neurotransmitter receptor expression is diverse in midline neurons, with 31 receptor genes expressed with FPKM values >5. These include inputs for acetylcholine, dopamine, GABA, glutamate, octopamine, and serotonin. Additionally, 13 neuropeptide receptors are expressed in subsets of midline cells. Interestingly, there are several examples where midline neurons secrete specific neuropeptides, and different midline neurons possess their receptors. For example, Dh31-R1 and Dh44-R1 are expressed in H-cell and H-cell sib, and iVUM5 and iVUM6 neuron express Dh31 and Dh44, respectively. In addition, iVUM5 expresses sNPF while the MP1 neurons and mVUMs express sNPF-R. However, at this time, it is unknown whether the midline-directed neurosecretion directly influences midline neurotransmission.
Using data obtained from our RNA-seq experiments with information already known about CNS midline cell gene expression, we were also able to link the activity of a transcription factor with neuropeptide expression. The role of Cas as a temporal identity factor important during the specification of lateral CNS cells has been well established (Grosskortenhaus et al., 2006; Isshiki et al., 2001). In the CNS midline cells, we found that cas does not specify the neurons it is expressed in, but is required in iVUM5 neurons for the expression of the sNPF neuropeptide gene. Our observation that cas is not sufficient to drive sNPF expression in other CNS midline cells, including iVUM4 and iVUM6, suggests that other iVUM5-specific factors are required in addition to cas. Since cas affects iVUM5 sNPF expression, but not that of Gad1, this indicates that multiple regulatory proteins control different aspects of iVUM5 differentiation. This contrasts with the general view of neuronal differentiation in C. elegans in which neuronal differentiation is largely co-regulated (e.g. Kratsios et al., 2012; Hobert, 2011). This manuscript has focused on only a small fraction of the midline RNA-seq data, and future analyses of the data will be likely reveal additional insight into the regulation of neural and glial differentiation. One important extension of this approach will be to purify subsets of CNS midline neurons and glia followed by RNA-seq analysis to assess the similarities and differences of individual midline cell types.
Supplementary Material
Drosophila embryonic CNS midline cells were FACS purified followed by RNA-seq
The midline transcriptome was determined at two major developmental time points
In situ hybridization revealed diversity of midline neural function gene expression
9 neuropeptides and 44 neurotransmitter receptors are present in midline cells
castor regulates neuropeptide gene expression in a midline interneuron subtype
Acknowledgments
The authors would like to thank Trevor Parton for help with in situ hybridizations, Feng Liu for protocols and advice on embryo dissociation and cell sorting, Dan McKay and Erin Osborne for discussions about analyzing high throughput sequencing data, Ward Odenwald for providing UAS-cas strains and anti-Cas antibody, Chris Doe for providing cas mutant strains, the Developmental Studies Hybridoma Bank for antibodies, and the Bloomington Drosophila Stock Center for fly strains. We thank Lisa Bixby at the UNC Flow Cytometry Core Facility for her expertise with cell sorting, Piotr Mieczkowski at the UNC-HTSF for protocols and sequencing, and Simon Andrews at the Babraham Institute for the FastQC software. We also thank Joe Pearson, Stephanie Stagg, Joseph Watson, and Scott Wheeler for helpful discussions. This work was supported by NIH grants R01 NS64264 (NINDS) and R37 RD25251 (NICHD) to S.T.C.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- Al-Anzi B, Armand E, Nagamei P, Olszewski M, Sapin V, Waters C, Zinn K, Wyman RJ, Benzer S. The leucokinin pathway and its neurons regulate meal size in Drosophila. Curr Biol. 2010;20:969–978. doi: 10.1016/j.cub.2010.04.039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Audsley N, McIntosh C, Phillips JE. Isolation of a neuropeptide from locust corpus cardiacum which influences ileal transport. J Exp Biol. 1992;173:261–274. doi: 10.1242/jeb.173.1.261. [DOI] [PubMed] [Google Scholar]
- Benito-Sipos J, Estacio-Gómez A, Moris-Sanz M, Baumgardt M, Thor S, Díaz-Benjumea FJ. A genetic cascade involving klumpfuss, nab and castor specifies the abdominal leucokinergic neurons in the Drosophila CNS. Development. 2010;137:3327–3336. doi: 10.1242/dev.052233. [DOI] [PubMed] [Google Scholar]
- Brand A. GFP in Drosophila. Trends Genet. 1995;11:324–325. doi: 10.1016/s0168-9525(00)89091-5. [DOI] [PubMed] [Google Scholar]
- Burnstock G. Cotransmission. Curr Opin Pharmacol. 2004;4:47–52. doi: 10.1016/j.coph.2003.08.001. [DOI] [PubMed] [Google Scholar]
- Cabrero P, Radford JC, Broderick KE, Costes L, Veenstra JA, Spana EP, Davies SA, Dow JA. The Dh gene of Drosophila melanogaster encodes a diuretic peptide that acts through cyclic AMP. J Exp Biol. 2002;205:3799–3807. doi: 10.1242/jeb.205.24.3799. [DOI] [PubMed] [Google Scholar]
- Coast GM, Webster SG, Schegg KM, Tobe SS, Schooley DA. The Drosophila melanogaster homologue of an insect calcitonin-like diuretic peptide stimulates V-ATPase activity in fruit fly Malpighian tubules. J Exp Biol. 2001;204:1795–1804. doi: 10.1242/jeb.204.10.1795. [DOI] [PubMed] [Google Scholar]
- Cui X, Doe CQ. ming is expressed in neuroblast sublineages and regulates gene expression in the Drosophila central nervous system. Development. 1992;116:943–952. doi: 10.1242/dev.116.4.943. [DOI] [PubMed] [Google Scholar]
- Dircksen H, Tesfai LK, Albus C, Nässel DR. Ion transport peptide splice forms in central and peripheral neurons throughout postembryogenesis of Drosophila melanogaster. J Comp Neurol. 2008;509:23–41. doi: 10.1002/cne.21715. [DOI] [PubMed] [Google Scholar]
- Dong R, Jacobs JR. Origin and differentiation of supernumerary midline glia in Drosophila embryos deficient for apoptosis. Dev Biol. 1997;190:165–177. doi: 10.1006/dbio.1997.8688. [DOI] [PubMed] [Google Scholar]
- Fortini ME, Lai ZC, Rubin GM. The Drosophila zfh-1 and zfh-2 genes encode novel proteins containing both zinc-finger and homeodomain motifs. Mech Dev. 1991;34:113–122. doi: 10.1016/0925-4773(91)90048-b. [DOI] [PubMed] [Google Scholar]
- Gabilondo H, Losada-Pérez M, Del Saz D, Molina I, León Y, Canal I, Torroja L, Benito-Sipos J. A targeted genetic screen identifies crucial players in the specification of the Drosophila abdominal Capaergic neurons. Mech Dev. 2011;128:208–221. doi: 10.1016/j.mod.2011.01.002. [DOI] [PubMed] [Google Scholar]
- Graveley BR, Brooks AN, Carlson JW, Duff MO, Landolin JM, Yang L, Artieri CG, van Baren MJ, Boley N, Booth BW, Brown JB, Cherbas L, Davis CA, Dobin A, Li R, Lin W, Malone JH, Mattiuzzo NR, Miller D, Sturgill D, Tuch BB, Zaleski C, Zhang D, Blanchette M, Dudoit S, Eads B, Green RE, Hammonds A, Jiang L, Kapranov P, Langton L, Perrimon N, Sandler JE, Wan KH, Willingham A, Zhang Y, Zou Y, Andrews J, Bickel PJ, Brenner SE, Brent MR, Cherbas P, Gingeras TR, Hoskins RA, Kaufman TC, Oliver B, Celniker SE. The developmental transcriptome of Drosophila melanogaster. Nature. 2011;471:473–479. doi: 10.1038/nature09715. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grosskortenhaus R, Robinson KJ, Doe CQ. Pdm and Castor specify late-born motor neuron identity in the NB7-1 lineage. Genes Dev. 2006;20:2618–2627. doi: 10.1101/gad.1445306. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guss KA, Mistry H, Skeath JB. Vestigial expression in the Drosophila embryonic central nervous system. Dev Dyn. 2008;237:2483–2489. doi: 10.1002/dvdy.21664. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hobert O. Regulation of terminal differentiation programs in the nervous system. Annu Rev Cell Dev Biol. 2011;27:681–696. doi: 10.1146/annurev-cellbio-092910-154226. [DOI] [PubMed] [Google Scholar]
- Hökfelt T, Broberger C, Xu ZQ, Sergeyev V, Ubink R, Diez M. Neuropeptides--an overview. Neuropharmacology. 2000;39:1337–1356. doi: 10.1016/s0028-3908(00)00010-1. [DOI] [PubMed] [Google Scholar]
- Isshiki T, Pearson B, Holbrook S, Doe CQ. Drosophila neuroblasts sequentially express transcription factors which specify the temporal identity of their neuronal progeny. Cell. 2001;106:511–521. doi: 10.1016/s0092-8674(01)00465-2. [DOI] [PubMed] [Google Scholar]
- Iversen A, Cazzamali G, Williamson M, Hauser F, Grimmelikhuijzen CJ. Molecular identification of the first insect ecdysis triggering hormone receptors. Biochem Biophys Res Commun. 2002;299:924–931. doi: 10.1016/s0006-291x(02)02798-5. [DOI] [PubMed] [Google Scholar]
- Johard HA, Yoishii T, Dircksen H, Cusumano P, Rouyer F, Helfrich-Förster C, Nässel DR. Peptidergic clock neurons in Drosophila: ion transport peptide and short neuropeptide F in subsets of dorsal and ventral lateral neurons. J Comp Neurol. 2009;516:59–73. doi: 10.1002/cne.22099. [DOI] [PubMed] [Google Scholar]
- Kahsai L, Winther ÅM. Chemical neuroanatomy of the Drosophila central complex: distribution of multiple neuropeptides in relation to neurotransmitters. J Comp Neurol. 2011;519:290–315. doi: 10.1002/cne.22520. [DOI] [PubMed] [Google Scholar]
- Kambadur R, Koizumi K, Stivers C, Nagle J, Poole SJ, Odenwald WF. Regulation of POU genes by castor and hunchback establishes layered compartments in the Drosophila CNS. Genes Dev. 1998;12:246–260. doi: 10.1101/gad.12.2.246. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kearney JB, Wheeler SR, Estes P, Parente B, Crews ST. Gene expression profiling of the developing Drosophila CNS midline cells. Dev Biol. 2004;275:473–92. doi: 10.1016/j.ydbio.2004.08.047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim YJ, Zitnan D, Galizia CG, Cho KH, Adams ME. A command chemical triggers an innate behavior by sequential activation of multiple peptidergic ensembles. Curr Biol. 2006;16:1395–1407. doi: 10.1016/j.cub.2006.06.027. [DOI] [PubMed] [Google Scholar]
- King DS, Meredith J, Wang YJ, Phillips JE. Biological actions of synthetic locust ion transport peptide (ITP) Insect Biochem Mol Biol. 1999;29:11–18. doi: 10.1016/s0965-1748(98)00098-8. [DOI] [PubMed] [Google Scholar]
- Kratsios P, Stolfi A, Levine M, Hobert O. Coordinated regulation of cholinergic motor neuron traits through a conserved terminal selector gene. Nat Neurosci. 2011;15:205–214. doi: 10.1038/nn.2989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- LaJeunesse DR, Johnson B, Presnell JS, Catignas KK, Zapotoczny G. Peristalsis in the junction region of the Drosophila larval midgut is modulated by DH31 expressing enteroendocrine cells. BMC Physiol. 2010;10:14. doi: 10.1186/1472-6793-10-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Langmead B, Trapnell C, Pop M, Salzberg SL. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 2009;10:R25. doi: 10.1186/gb-2009-10-3-r25. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee KS, You KH, Choo JK, Han YM, Yu K. Drosophila short neuropeptide F regulates food intake and body size. J Biol Chem. 2004;279:50781–50789. doi: 10.1074/jbc.M407842200. [DOI] [PubMed] [Google Scholar]
- Lee T, Luo L. Mosaic analysis with a repressible cell marker for studies of gene function in neuronal morphogenesis. Neuron. 1999;22:451–461. doi: 10.1016/s0896-6273(00)80701-1. [DOI] [PubMed] [Google Scholar]
- Losada-Pérez M, Gabilondo H, del Saz D, Baumgardt M, Molina I, León Y, Monedero I, Díaz-Benjumea F, Torroja L, Benito-Sipos J. Lineage-unrelated neurons generated in different temporal windows and expressing different combinatorial codes can converge in the activation of the same terminal differentiation gene. Mech Dev. 2010;127:458–471. doi: 10.1016/j.mod.2010.08.003. [DOI] [PubMed] [Google Scholar]
- McQuilton P, St Pierre SE, Thurmond J, FlyBase Consortium. FlyBase 101--the basics of navigating FlyBase. Nucleic Acids Res. 2012;40:D706–14. doi: 10.1093/nar/gkr1030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miguel-Aliaga I, Thor S. Segment-specific prevention of pioneer neuron apoptosis by cell-autonomous, postmitotic Hox gene activity. Development. 2004;131:6093–6105. doi: 10.1242/dev.01521. [DOI] [PubMed] [Google Scholar]
- Mills JC, Taghert PH. Scaling factors: transcription factors regulating subcellular domains. Bioessays. 2012;34:10–16. doi: 10.1002/bies.201100089. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nambu JR, Lewis JL, Wharton KA, Crews ST. The Drosophila single-minded gene encodes a helix-loop-helix protein which acts as a master regulator of CNS midline development. Cell. 1991;67:1157–1167. doi: 10.1016/0092-8674(91)90292-7. [DOI] [PubMed] [Google Scholar]
- Nässel DR, Homberg U. Neuropeptides in interneurons of the insect brain. Cell Tissue Res. 2006;326:1–24. doi: 10.1007/s00441-006-0210-8. [DOI] [PubMed] [Google Scholar]
- Park D, Veenstra JA, Park JH, Taghert PH. Mapping peptidergic cells in Drosophila: where DIMM fits in. PLoS One. 2008;3:e1896. doi: 10.1371/journal.pone.0001896. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sonnenfeld MJ, Jacobs JR. Apoptosis of the midline glia during Drosophila embryogenesis: a correlation with axon contact. Development. 1995;121:569–578. doi: 10.1242/dev.121.2.569. [DOI] [PubMed] [Google Scholar]
- Terhzaz S, O’Connell FC, Pollock VP, Kean L, Davies SA, Veenstra JA, Dow JA. Isolation and characterization of a leucokinin-like peptide of Drosophila melanogaster. J Exp Biol. 1999;202:3667–3676. doi: 10.1242/jeb.202.24.3667. [DOI] [PubMed] [Google Scholar]
- Trapnell C, Pachter L, Salzberg SL. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics. 2009;25:1105–1111. doi: 10.1093/bioinformatics/btp120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Trapnell C, Williams BA, Pertea G, Mortazavi A, Kwan G, van Baren MJ, Salzberg SL, Wold BJ, Pachter L. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol. 2010;28:511–515. doi: 10.1038/nbt.1621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Truman JW, Bate M. Spatial and temporal patterns of neurogenesis in the central nervous system of Drosophila melanogaster. Dev Biol. 1988;125:145–157. doi: 10.1016/0012-1606(88)90067-x. [DOI] [PubMed] [Google Scholar]
- Ubink R, Calza L, Hökfelt T. ‘Neuro’-peptides in glia: Focus on NPY and galanin. Trends Neurosci. 2003;26:604–609. doi: 10.1016/j.tins.2003.09.003. [DOI] [PubMed] [Google Scholar]
- Vasenkova I, Luginbuhl D, Chiba A. Gliopodia extend the range of direct glia-neuron communication during the CNS development in Drosophila. Mol Cell Neurosci. 2006;31:123–130. doi: 10.1016/j.mcn.2005.10.001. [DOI] [PubMed] [Google Scholar]
- Veelaert D, Devreese B, Schoofs L, Van Beeumen J, Vanden Broeck J, Tobe SS, De Loof A. Isolation and characterization of eight myoinhibiting peptides from the desert locust, Schistocerca gregaria: new members of the cockroach allatostatin family. Mol Cell Endocrinol. 1996;122:183–190. doi: 10.1016/0303-7207(96)03889-0. [DOI] [PubMed] [Google Scholar]
- Wheeler SR, Kearney JB, Guardiola AR, Crews ST. Single-cell mapping of neural and glial gene expression in the developing Drosophila CNS midline cells. Dev Biol. 2006;294:509–524. doi: 10.1016/j.ydbio.2006.03.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wheeler SR, Stagg SB, Crews ST. Multiple Notch signaling events control Drosophila CNS midline neurogenesis, gliogenesis and neuronal identity. Development. 2008;135:3071–3079. doi: 10.1242/dev.022343. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wheeler SR, Banerjee S, Blauth K, Rogers SL, Bhat MA, Crews ST. Neurexin IV and Wrapper interactions mediate Drosophila midline glial migration and axonal ensheathment. Development. 2009a;136:1147–1157. doi: 10.1242/dev.030254. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wheeler SR, Stagg SB, Crews ST. MidExDB: a database of Drosophila CNS midline cell gene expression. BMC Dev Biol. 2009b;9:56. doi: 10.1186/1471-213X-9-56. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wheeler SR, Pearson JC, Crews ST. Time-lapse imaging reveals stereotypical patterns of Drosophila midline glial migration. Dev Biol. 2012;361:232–244. doi: 10.1016/j.ydbio.2011.10.024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xiao H, Hrdlicka LA, Nambu JR. Alternate functions of the single-minded and rhomboid genes in development of the Drosophila ventral neuroectoderm. Mech Dev. 1996;58:65–74. doi: 10.1016/s0925-4773(96)00559-x. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.