Abstract
Background
The lavender phenotype in quail is a dilution of both eumelanin and phaeomelanin in feathers that produces a blue-grey colour on a wild-type feather pattern background. It has been previously demonstrated by intergeneric hybridization that the lavender mutation in quail is homologous to the same phenotype in chicken, which is caused by a single base-pair change in exon 1 of MLPH.
Results
In this study, we have shown that a mutation of MLPH is also associated with feather colour dilution in quail, but that the mutational event is extremely different. In this species, the lavender phenotype is associated with a non-lethal complex mutation involving three consecutive overlapping chromosomal changes (two inversions and one deletion) that have consequences on the genomic organization of four genes (MLPH and the neighbouring PRLH, RAB17 and LRRFIP1). The deletion of PRLH has no effect on the level of circulating prolactin. Lavender birds have lighter body weight, lower body temperature and increased feed consumption and residual feed intake than wild-type plumage quail, indicating that this complex mutation is affecting the metabolism and the regulation of homeothermy.
Conclusions
An extensive overlapping chromosome rearrangement was associated with a non-pathological Mendelian trait and minor, non deleterious effects in the lavender Japanese quail which is a natural knockout for PRLH.
Background
Coat and plumage colours result from the interplay of a number of genes, and the variety of visible colours associated with specific mutations or combination of mutations may be used to gain an insight into underlying gene action. The available knowledge of mouse coat colour mutants is quite extensive, but little is as yet known about the genes and the mutations that cause the highly variable plumage colour of birds. Recently, causal mutations for quail plumage colours yellow[1], recessive black[2], and roux[3] have been characterized and found to be homologous to known coat colour mutations in mice. Increasing our knowledge of colour mutations common to mammals and birds should make it possible to compare more systematically colour genes and their associated phenotypic effects in these two animal classes. Indeed, the comprehensive phenotyping of mutants, which is a common practice in mice [4] for example, has begun in quail [5,6], but more contributions are needed to enable more extensive comparisons between mammals and birds.
Until now, all the reported causal mutations in MLPH (melanophilin) of humans, mice and other species have been single-base substitutions or small deletions, the effects of which were limited to the dilution of hair [7,8] or feather [9] colour. It was suggested [10] that MLPH formed a complex with MyoVa and RAB27a for transporting melanosomes from melanocyte to keratinocyte. The MLPH-associated dilution of coat or plumage pigmentation should then result in the defective transport of melanosomes. This produces a diluted, leaden or lavender blue-grey colour and has been reported in several mammals: humans (Griscelli syndrome type 3 [7]), mice [10], cats [11], dogs [12] and minks [13]. More recently, this effect was also found in the chicken, with the identification in MLPH of the mutation responsible for the lavender plumage [9].
In the Japanese quail (Coturnix japonica), the lavender plumage colour (Figure 1) is determined by a recessive autosomal mutation [14,15], like in chicken [16], which leads to the dilution of both eumelanin (from black to light blue-grey) and phaeomelanin (from red to buff). Intergeneric hybridisation between lavender Gallus and Coturnix has produced hybrids with a lavender plumage [14]. This result indicated that, as in chicken, MLPH was associated with the lavender plumage colour in quail, but did not give any information on the nature of the causal mutation.
Figure 1.
Wild-type (left) and lavender (right) Japanese quail.
The primary purpose of the present study was to characterise the mutation within MLPH that could be responsible for the “lavender” plumage in Japanese quail, and to compare lavender and wild-type quail for possible phenotypic differences associated with the mutation.
Methods
Quail
All Japanese quail were produced and maintained at the INRA Experimental Unit PEAT in Nouzilly, France (Pôle d'Expérimentation Avicole de Tours, F-37380 Nouzilly, authorization B37-175-1, 2007) in accordance with European Union Guidelines for animal care, under authorization 37–002 delivered to D. Gourichon by the French Ministry of Agriculture. Animal procedures were approved by the Departmental Direction of Veterinary Services of Indre-et-Loire. A homozygous lav/lav quail line was developed in the 1980s from a single individual with lavender plumage given by a fancy breeder and that was crossed initially to a “wild-type” experimental line showing no visible plumage colour mutations in segregation. The line was then maintained by group mating of about 15 males and 30 females per generation. First, heterozygous lav/+ quail were produced specifically for the experiment by crosses with the “wild-type” line. For the study of co-transmission between MLPH genotypes and quail phenotypes, 6 lavender lav/lav quail and 42 lav/+ or +/+ birds with wild-type plumage were obtained in a single hatch from 3 single-pair matings between non-sib heterozygous lav/+ quail. Blood samples were collected from the six parents and the 48 progeny for DNA analysis. For the study of other possible phenotypic differences between lavender and wild-type animals, 130 quail (63 lavender and 67 wild-type) were obtained from 21 single-pair matings between lav/lav and lav/+ quail, in a single hatch following a two-week egg collection period.
DNA extraction and genomic analyses
High molecular-weight DNA was obtained from blood samples using classical DNA extraction procedure [17]. All genomic analyses were carried out using DNA samples from three +/+ and three lav/lav quail. The annotation of the region was based on information from the chicken genome sequence (assembly v2.1), from the UCSC (genome.ucsc.edu) and Ensembl ( http://www.ensembl.org) genome browsers, under the assumption that the organization of chicken and quail genomes is highly similar [18]. Primers were designed using Primer3 software (frodo.wi.mit.edu/primer3/) and quail genomic sequence as a template or chicken sequence when the quail sequence was not available. When the chicken sequence was used, new quail-specific primers were designed after successful amplification and sequencing of the region. PCRs were undertaken using the Hot Start Master Mix (Qiagen). All Sanger sequencing was conducted by the Eurofins-MWG company (Ebersberg, Germany) using PCR as templates and PCR primers as sequencing primers. The list of primers used is detailed in Additional file 1: Table S1.
Chromosome walking
To carry out this approach [19], a first PCR was completed using a biotinylated primer specific to the known region and a partly degenerate primer coupled to a unique tail sequence. After purification with stretptavidin-coated beads, a second PCR was conducted on this template using a nested primer from the known region and a primer complementary to the unique tail sequence, then the product was sequenced using these primers as sequencing primers. The primers used are detailed in Additional file 1: Table S1.
Tissue samples and RNA extraction
Tissues (liver, dorsal skin, lung, brain and pectoral muscle) were frozen in liquid nitrogen immediately after sampling from 3 +/+ and 3 lav/lav adult quail, and kept at −80 °C before use. Samples were extracted by grinding tissues in liquid nitrogen followed by re suspension in Qiazol (Qiagen). After addition of chloroform and centrifugation, the aqueous phase was recovered and supplemented with isopropanol, and total RNA was precipitated by centrifugation after ethanol washing. Quantification was performed using 1 μL of the solution in a Nanodrop (ThermoScientific), and quality (profile and RNA Integrity Number (RIN) value) controlled by analysis of 1 μL of the solution in a Bioanalyzer (Agilent).
Gene expression
Using total RNA, RT-PCRs were performed with SuperScript II (Invitrogen) for DNA first-strand synthesis, followed by PCR using Hot-Start Master Mix (Qiagen). For MLPH, 6 RT-PCRs were tested, corresponding to different parts of the gene, encompassing or not the deleted region (between exons 1 to 2, 5 to 7, 1 to 7, 8 to 10, 1 to 10 and 5 to 10). For PRLH, 2 RT-PCRs were tested, between exons 1 to 2 and 1 to 3. For RAB17, 2 RT-PCRs were tested, between exons 1 to 4 and 3 to 4. Amplification was confirmed by electrophoresis on 2% agarose gel, and the sizes of amplified fragments determined by comparison to a reference ladder. The list of primers used is detailed in Additional file 1: Table S1.
Genotyping
A PCR test was developed for genotyping the lavender locus in quail. A three-primer PCR test was used, with one primer in a conserved region (Gen_F) and a second primer specific to the genomic organisation of each genotype: inside the deleted region for wild-type (Gen_wt_R) and inside the inverted region for lavender (Gen_lav_R). Primer sequences are detailed in Additional file 1: Table S1. After PCR, amplification was confirmed by electrophoresis on 1.5% agarose gel, and the sizes of amplified fragments determined by comparison to a reference ladder. The expected sizes of PCR products were 423 bp for the band specific to wild-type genomic organization, and 630 bp for the band specific to lavender genomic organization.
Husbandry and traits
Chicks were reared in group cages and given a starter diet until they were 5 weeks of age. They were sexed at 3 weeks, and males and females were kept separately thereafter. Next, they were transferred to adult individual cages in a single battery under artificial lighting for 14 hours per day, and were given a standard commercial diet. Drinking water was available at all times. All husbandry procedures followed French regulations. Individual body weights were measured weekly between the ages of 1 and 9 weeks. A 3-week feed trial was started with 96 5-month quail of both genders (48 males and 48 females) and plumage colours (52 lavender and 44 wild-type quail) from the 21 full-sib families. Individual feed intake during the 3-week period was registered, and body weights were measured at the beginning and the end of the test, after 12 hours’ fasting, in order to estimate body weight gain and metabolic body weight on test. For each female, the number of eggs laid during the test was recorded, and three eggs laid consecutively were weighed to estimate the total egg mass produced during the feed test. Body temperature was measured at the end of the test. Four quail did not complete the feed trial. After the feed test, all quail were fed ad libitum until 6 months of age when they were weighed and slaughtered at the experimental unit. Sacrificed quail were weighed before dissection (carcass body weight). Abdominal adipose tissue, right pectoralis major and pectoralis minor muscles, and liver were collected and weighed for each quail.
Blood samples were obtained from 20 lavender and 6 wild-type quail and plasma concentration of prolactin were measured by radioimmunoassay at the CEBC [20]. Pooled plasma samples taken from quail resulted in dose–response curves that paralleled the chicken standard curves (AFP4444B). Thus, the cross-reactivity of the chicken prolactin antibody with prolactin was equivalent in both species, and this heterologous assay could be used to assess relative levels of quail prolactin. The detection limit of the assay was 2.8 ng/ml and the intra-assay coefficient of variation was 12.5%.
Statistical analyses
Individual growth was assessed using the nonlinear monomolecular model [21]: body weight = A – B-kt, where A is the asymptotic body weight, B is the range of body weights from hatching to asymptotic body weight, k is the relative rate of growth, and t is the age in days, and parameters of the curve were obtained by using the NLIN procedure [22]. Single curves were also adjusted for each genotype (lav/lav and lav/+) to estimate the fit (coefficient of determination) of the model. Residual feed intake (RFI), which is the part of the feed consumed that is not used for growth, maintenance or production [23] was estimated. In females, RFI was estimated for each bird as the residual (over the 21-day feed-test period) of the multiple regression of individual feed intake (FI) on body weight gain on test (BWG), metabolic body weight (MBW), and egg mass (EM). The multiple regression fitted to the data (R2 = 0.57) was FI = 23.3 + 1.50 BWG + 6.56 MBW +1.075 EM. To estimate individual RFI in males, the multiple regression (R2 = 0.72) was FI = 255.2 + 3.35 BWG + 2.91 MBW.
Parameters of individual growth curves, body weights, feed intake and carcass weight were analysed by an analysis of variance with family, sex and genotype (lav/lav or lav/+) as main effects (Full model). An analysis without the effect of genotype (Reduced model) was also run to evaluate the variation explained by plumage colour. The linear model used for the analysis of egg number and egg weight was similar but did not include the effect of sex. Body temperatures and dissection traits were analysed by an analysis of covariance, with family, sex and genotype as main effects, and with a covariable, which was the contemporaneous body weight for measures of rectal temperature and the carcass weight for dissection traits. The analyses of variance and covariance were carried out using the GLM procedure [22].
Results
Characterisation of the lavender mutation
The genomic region of MLPH was compared in wild-type and lavender Japanese quail to determine the causal mutation of the lavender phenotype. A preliminary screening with PCR amplification of several targets distributed over the region revealed that some consecutive PCRs failed to produce any amplification in DNA from lavender birds (PCR E to L, Figure 2), whereas all PCRs amplified the target region in DNA from wild-type animals. Moreover, the flanking PCRs produced the expected amplification in both wild-type and lavender quail (PCR A to D and M to P, Figure 2). All PCR fragments were directly sequenced from both ends and their sequences corresponded to the region expected from the chicken genomic sequence. This pattern was compatible with a deletion encompassing a large part of the MLPH gene, the entire PRLH gene and a large part of the RAB17 gene, according to the chicken genome annotation. Additional PCR tests confirmed that a deletion was likely to be associated with lavender in Japanese quail, and allowed us to narrow down the two regions containing each putative breakpoint to less than 1 kb (see respectively overlapping PCRs D–E and L–M). But all attempts to close the deletion by a PCR with primers designed in the flanking regions failed, and it was concluded that the mutation was more complex than a simple deletion.
Figure 2.
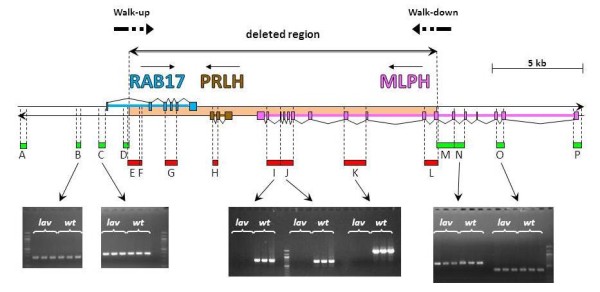
Results of the PCR screening of the genomic region comprising MLPH ,PRLH and RAB17. MLPH, PRLH and RAB17 genes are coloured in purple, brown, and light blue, respectively. Exons and introns are depicted as outlined boxes and broken lines, respectively. PCR working in both lavender and wild-type (A-D and M-P) are in green, PCR working only in wild-type (E-L) are in red, with band visualization after gel electrophoresis for PCR B, C, I, J, K, N and O on 3 lavender and 3 wild-type animals. Degenerate PCR Walk-up and Walk-down are indicated by arrows.
Chromosome walking from the flanking regions was used to determine the genomic structure of the region for the lavender allele: PCR assays were done with one primer anchored in each flanking region (corresponding respectively to regions D and M) and a degenerate one with a specific tail. After a second nested PCR, the products were sequenced directly from both ends, which revealed that a complex pattern of rearrangements had occurred to produce the lavender allele.
From the region upstream of the first putative breakpoint (region D), the sequences of PCR products after chromosome walking (named Walk-up on Figures 2 and 3) matched successively to two different locations in the genome. The first one (from region D, on the positive strand) was the region between the nested primer and the breaking point, followed, after the breaking point, by the second one that matched to a region 27.6 kb upstream of the breakpoint on the opposite strand.
Figure 3.
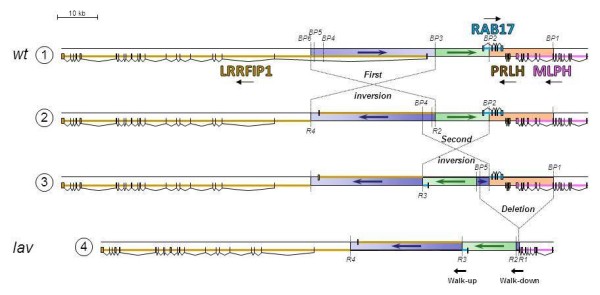
Reconstruction of chromosomal events from wild-type genomic (1, wt ) organization to lavender (4, lav ) genomic organization. MLPH, PRLH, RAB17 and LRRFIP1 genes are coloured in purple, brown, light blue and orange, respectively. Exons and introns are depicted as outlined boxes and broken lines, respectively. Breakpoints are numbered from BP1 to BP6 in the wild-type sequence, and recombined points are numbered from R1 to R4 in the lavender sequence. Degenerate Walk-up and Walk-down PCRs are indicated by arrows.
From the region downstream of the second putative breakpoint (region M), the sequenced PCR products after chromosome walking (named Walk-down on Figures 2 and 3) matched successively to three different locations of the genome. The first one (from region M, on the negative strand) was the region between the nested primer and the breaking point, the second one was a 246-bp fragment corresponding to a region 30 kb upstream on the same negative strand, followed by the third one, matching to a region 3 kb upstream, but on the positive strand.
Figure 3 shows the most parsimonious sequence of chromosomal events that could lead from the organization of the wild-type allele (state 1) to that observed in the lavender allele (state 4). The organization of the genomic region for lavender has been constructed using the information provided by the chromosome-walking approach and complementary PCR analyses. The complex mutation spans 60 kb, and the sequence of chromosomal events consists of three ordered steps. The first one is the inversion of a 30.9-kb region upstream of MLPH that contains the first exon and a large part of the first intron of LRRFIP1 (between breakpoints BP3 and BP6). The second step is another inversion of 16.4 kb overlapping the first one, which includes again a part of LRRFIP1 intron 1 on one side, exon 1 and a large part of intron 1 of RAB17 on the other side (between breakpoints BP4 and BP2). Finally, the last step is a 16-kb deletion overlapping the second inversion, which removes most of RAB17, the entire PRLH gene and a large part of MLPH (between breakpoints BP5 and BP1). BP1 is located between MLPH exons 9 and 10, and the only remaining genomic region is corresponding to exons 1 to 9. The breakpoints have been named in the wild-type allele according to their order along the sequence from MLPH (from BP1 to BP6), and named similarly in the lavender allele (from R1 to R4, where R stands for rearrangement). The breakpoints after the first inversion are, respectively, R4 and R2; the remaining breakpoint of the second inversion is R3 (the second one being lost during the deletion), and the scar of the deletion is R1. All the breakpoints (from BP1 to BP6 for the wild-type allele, and from R1 to R4 for the lavender allele) have been confirmed by specific PCR amplifications (Figure 2 and Additional file 2: Figure S1) followed by sequencing, except for BP3 and R4 for which nonspecific PCR amplification led to poor sequence quality (Additional file 1: Table S2). Larger local differences between chicken and quail genomic sequences upstream of the BP3 breakpoint are the most likely reason for this results, given that unsuccessful preliminary mapping of quail NGS reads on the chicken reference genome was already observed in this region (F. Pitel, personal communication). Sequences of wild-type and lavender breakpoint regions have been deposited into the nucleotide GenBank database, under accession numbers JX266433 to JX266439 as follows: JX266433 for BP1_wt, JX266434 for BP2_wt, JX266435 for BP4_wt, JX266436 for BP5_wt, JX266437 for BP6_wt, JX266438 for R1-R2_lav, and JX266439 for R3_lav (Additional file 1: Table S3).
Heterozygous parents lav/+ (n = 6) and offspring (n = 48) from the informative cross were genotyped using the three-primer PCR test. The genotypes of all parents were confirmed as heterozygous, whereas the genotypes of offspring were in perfect agreement with their observed phenotypes: homozygous lav/lav for lavender birds (n = 6), heterozygous lav/+ (n = 19) or homozygous +/+ (n = 23) for wild-type quail.
Gene expression
The RNA integrity number (RIN) for the extracted RNA ranged between 7.4 and 7.8. All RT-PCR amplifications were successful for MLPH and RAB17 pairs of primers in liver, dorsal skin, lung, brain and pectoral muscle samples from wild-type adults. In lavender adults, we obtained no amplification for RAB17 and for MLPH exons 1 to 10, 8 to 10 and 5 to 10, but the same products as obtained in the wild type were detected for MLPH exons 1 to 2, 1 to 7 and 5 to 7. No amplification was obtained for RT-PCR targeting PRLH in both lavender and wild-type quail, although the gene is likely to be present in wild-type birds: there are 2 chicken transcribed sequences for PRLH in GenBank (EST BX933358 / BU445468 (from ovary) and mRNA EF418015), and PRLH was reported to be present on chromosome 7 of the chicken [24].
Association between the complex mutation revealed by the plumage colour and various traits Parameters of the growth curves are listed and compared in Table 1. Asymptotic body weight and body weight range were influenced by the complex mutation (P < 0.05), and lavender quail had a consistently lighter body weight during the growth period. The relative rate of growth was similar for the two genotypes, however, indicating that the overall shape of the growth curve was not affected. Results of the feed intake trial are given in Table 2. Measures of body weight confirmed that there was only a marginal difference between genotypes (P < 0.05 or NS). Body weight gain on test was similar for the two quail groups. Daily feed intake and residual feed intake were higher (P < 0.001) for the lavender quail, which had a lower body temperature (P < 0.001) than wild-type quail. Egg number and egg weight were not affected. Measures taken at 6 months (Table 3) confirmed the smaller body weight (P < 0.05) and lower body temperature (P < 0.01) of lavender quail. Gross dissection showed that the pectoralis skeletal muscles were lighter (P < 0.001), and the liver was heavier (P < 0.001) in the lavender quail, whereas there was no difference in abdominal adipose tissue associated with the genotype. Plasma prolactin mean levels were not significantly different (P = 0.30) for lavender (82.4 ng/mL) and wild-type (54.0 ng/mL) quail.
Table 1.
Parameters (mean ± SD) of the growth curves1for 59 lavender (lav/lav) and 66 wild-type (lav/+) Japanese quail, and effects of the family, sex and plumage colour on the parameters of individual curves
|
Parameter of the curve |
Plumage colour (genotype) |
Percentage of the total variance explained by the model2(R2) |
Significance of the effect of plumage colour in the Full model |
||
|---|---|---|---|---|---|
| Lavender (lav/lav) | Wild-type (lav/+) | Full model | Reduced model | ||
|
A |
204.4 ± 32.1 |
215.9 ± 38.3 |
67 |
65 |
* |
|
B |
221.4 ± 34.3 |
233.0 ± 39.6 |
71 |
69 |
* |
| k | 0.0405 ± 0.00604 | 0.0401 ± 0.00759 | 43 | 43 | NS |
1 with the monomolecular growth model: body weight = A-B-kt, where A is the asymptotic body weight, B is the range of body weight from hatching to asymptotic weight, k is the relative rate of growth, and t is the age in days.
2 The Full model includes the effects of (full-sib) family, sex and plumage colour, and the Reduced model does not include plumage colour.
*: p ≤ 0.05; NS: not significant.
Table 2.
Effects of family, sex and plumage colour on body weights, feed consumption, egg production and body temperature (mean ± SD) in a 3-week feed trial of 9-week oldlavender(n = 51) and wild-type (n = 41) Japanese quail from 18 full-sib families
|
Trait |
Plumage colour (genotype) |
Percentage of the total variance explained by the model1(R2) |
Significance of the effect of plumage colour in the Full model |
||
|---|---|---|---|---|---|
| Lavender (lav/lav) | Wild-type (lav/+) | Full model | Reduced model | ||
| 64-d BW after 12 h fasting (g) |
173.3 ± 22.2 |
178.6 ± 23.9 |
75 |
73 |
* |
| 85-d BW after 12 h fasting (g) |
181.7 ± 20.1 |
186.5 ± 20.3 |
61 |
59 |
NS |
| Daily feed intake (g) |
24.5 ± 4.3 |
23.0 ± 4.0 |
72 |
65 |
*** |
| BW gain in 3-week feed test (g) |
8.7 ± 11.7 |
6.7 ± 10.0 |
45 |
44 |
NS |
| Egg number2 on 3-week test |
19.5 ± 2.0 |
18.4 ± 3.5 |
27 |
25 |
NS |
| Egg weight1 (g) |
9.8 ± 0.9 |
9.8 ± 1.3 |
62 |
61 |
NS |
| Residual feed intake (g) |
19.2 ± 33.5 |
−22.8 ± 24.4 |
44 |
17 |
*** |
| 85-d body temperature3 after 12 h fasting (°C) | 41.02 ± 0.28 | 41.16 ± 0.27 | 60 | 47 | *** |
1The Full model includes the effects of family, sex and plumage colour, and the Reduced model does not include plumage colour.
2 from 23 lavender and 23 wild-type females.
3 85-d BW was used as a covariable for the analysis of body temperature.
* p ≤ 0.05; *** p ≤ 0.001; NS not significant.
Table 3.
Effects of family, sex, and plumage colour on gross body composition (mean ± SD) of 6-month oldlavender(n = 51) and wild-type (n = 41) Japanese quail from 18 full-sib families
|
Trait |
Plumage colour (genotype) |
Percentage of the total variance explained by the model1(R2) |
Significance of the effect of plumage colour in the Full model |
||
|---|---|---|---|---|---|
| Lavender (lav/lav) | Wild-type (lav/+) | Full model | Reduced model | ||
| 6-month body weight (g) |
187.3 ± 21.1 |
194.3 ± 22.1 |
67 |
64 |
* |
| 6-month body temperature2 (°C) |
40.89 ± 0.35 |
41.03 ± 0.26 |
52 |
45 |
*** |
| Carcass weight (g) |
179.6 ± 19.8 |
187.3 ± 21.0 |
65 |
61 |
* |
| Abdominal adipose tissue3 (g) |
4.20 ± 2.14 |
4.06 ± 1.99 |
72 |
71 |
NS |
|
Pectoralis muscles3,4 (g) |
12.7 ± 1.4 |
14.0 ± 1.7 |
83 |
76 |
*** |
| Liver weight3 (g) | 4.29 ± 2.03 | 4.10 ± 1.62 | 91 | 89 | *** |
1 The Full model includes the effects of family, sex and plumage colour, and the Reduced model does not include plumage colour.
2 Live body weight was used as a covariable for the analysis of body temperature of 6-month fed quail.
3 Carcass weight was used as a covariable for the analyses of abdominal adipose tissue, breast muscle weight, liver weight, and heart weight.
4 Total weight of the right pectoralis major and pectoralis minor muscles.
*: p ≤ 0.05; ***: p ≤ 0.001; NS: not significant.
Discussion
We have shown that a complex mutation was associated with the lavender phenotype in Japanese quail, and genotyping this mutation in an informative cross confirmed that the mutation co-segregated with the phenotype. The region affected by the complex mutation is highly conserved between birds and mammals, and MLPH, PRLH, RAB17 and LRRFIP1 are found in the same order and relative orientation in Zebra Finch (chromosome 7), Chicken (chromosome 7), Mouse (chromosome 1) and Human (chromosome 2). The same region can be found in the genome of the turkey, but it is distributed on several contigs because an assembly is not yet available. This conserved synteny indicates that the genes involved in the present mutation and those already studied in mammals or chicken are orthologous, and that this region is not generally affected by genomic instability.
The characterization of the detailed genomic structure of the mutation leading to the lavender phenotype in quail showed that an homologous phenotype between quail and chicken, demonstrated by the production of lavender intergeneric hybrids [14], was apparently the consequence of very different mutational events: a single base-pair substitution in chicken [9] versus a complex and large mutation with three successive chromosomal rearrangements (two inversions and one deletion), affecting four consecutive genes in quail. The reconstruction of the most likely sequence of chromosome rearrangements leading to the lavender allele raises the question of the molecular mechanism involved. Intermediate steps (2 and 3, Figure 3) between wild-type and lavender alleles were not observed in the population (PCR screening of the breakpoints, data not shown). Then, it was not possible to determine whether the rearrangements occurred as three unrelated events, with intermediate genotypes now lost or not represented in the population, or in only one step. If the three mutations occurred independently, something may have increased the probability of recurrent chromosome rearrangements in the same region. Otherwise, a complex and dramatic change in genome organization took place during a single meiosis, leading to the lavender allele. Screening the genomic region with chicken or quail sequence data (acquired during the study) did not reveal any particular structure, such as repeats or transposable elements that might have facilitated chromosomal rearrangements.
Very few mutations similar to the one observed here, i.e. overlapping chromosomal rearrangements, have been reported. In Human, complex chromosome rearrangements have been observed in cancer cells, and some are even characteristic of specific cancers [25], and other genetic diseases. For example, the Wiskott-Aldrich syndrome results from a complex mutation due to two deletions and one inversion in a 4.5-kb region [26]. In that example, the three chromosomal events were contiguous, but it has not been confirmed that they were overlapping, and the family study revealed that the complex mutation appeared in a single step. Similarly, another complex human mutation [27] resulted from two deletions and one inversion in a 2-Mb region, and all were contiguous. The pattern of the mutation suggested two overlapping chromosomal changes: an inversion followed by a deletion, and the family study indicated that this complex mutation occurred as a single event. These observations of similar de novo complex chromosomal changes suggest that the chromosomal rearrangements leading to the lavender quail may have also taken place in a single step.
Mechanisms proposed to explain complex mutational events combining inversion and deletion are based on strand misalignment of sequences with complementary features during DNA replication or meiosis. We have carefully checked the breakpoint regions for a common DNA pattern. These regions were not located on repeated or transposable elements, but some sequences flanking breakpoints shared a common G(C)3-5A pattern (Additional file 1: Table S2). Interestingly, a 2.2-kb DNA sequence (between BP4 and BP5, Figure 3) appears to be involved in all the chromosomal changes. The first inversion changed its sequence orientation, the second one restored it, and the deletion removed this DNA portion together with the region containing RAB17, PRLH and MLPH, leaving only a short 246-bp region (between BP6 and BP5, Figure 3) translocated 30 kb downstream by the two inversions. The sequence of this DNA segment did not show any particular structure, however.
The complex mutation found in the Japanese quail is not lethal, although it has a major impact on the DNA sequence of MLPH, but also on PRLH, which is completely missing, RAB17, which is mostly deleted, and to a lesser extent on LRRFIP1. Consequently, only expression of a truncated gene is possible for MLPH, and RAB17 and PRLH are not expressed. This is confirmed by the RT-PCR results, with no expression of RAB17 in lavender, and no transcript for MLPH beyond exon 9. PRLH (Prolactin Releasing Hormone) is one of the hypothalamic peptide hormones that regulate the production of pituitary hormones, and it was first identified as a prolactin-releasing factor in mammals. PRLH also acts as a neuromodulator of pituitary products, and is involved in the control of metabolism, energy homoeostasis and food intake [24]. The absence of the whole DNA region for PRLH in lavender quail was not associated with any difference in plasma prolactin concentration. Indeed, secretion of prolactin in quail is probably under the control of the vasoactive intestinal peptide (VIP), as in turkey [28] and zebra finch [29], which would explain the unchanged prolactin levels of lavender quail. RAB17 is a member of the RAB family (Ras proteins involved in membrane trafficking, part of the RAS oncogene superfamily). It encodes the RAS-associated protein 17 found in epithelial cells [30] and may be involved in membrane trafficking. It is almost completely deleted in lavender quail, but it is possible that its loss is compensated by another gene of the same family through functional redundancy, since the RAB gene family has many members (for example, 67 protein-coding genes in humans, and 53 protein-coding genes in chicken are annotated in Homologene NCBI database: http://www.ncbi.nlm.nih.gov/homologene). RAB17 particularly, is phylogenetically very close to RAB5 genes (RAB5A, RAB5B and RAB5C) [31]. RAB17 expression [32] is regulated by MITF in mammalian cell lines (melanomas) and melanocytes with RAB17 knockdown have reduced filopodia formation, leading to impaired melanosome transfer. Then, RAB17 might make some contribution to the quail plumage colour but it is probably not responsible for lavender in quail because hybrids between lavender chicken and quail would have the RAB17 function rescued by the normal chicken RAB17, and should have shown a wild-type, undiluted, plumage colour, which they did not [14]. LRRFIP1 (Leucine-Rich Repeat (in Flightless I) Interacting Protein-1) encodes a protein involved in the regulation of the TLR signalling pathway [33] and the production of type-I interferon in response to pathogens [34]. This immune function has not been investigated in the present study, but it is likely that this gene was only affected marginally by the complex mutation because it lies mostly outside the deleted and inverted region of the lavender quail DNA sequence. Indeed, only its putative first exon was involved, whereas gene annotation in human and mouse indicates that only one alternatively spliced variant out of five described is starting from this first exon, with relatively few ESTs including this first exon.
Past results in chicken [9], mice [10], cats [11], and dogs [12] point to the large deletion in MLPH as the most likely cause of the “lavender” plumage colour in the Japanese quail. In the present study, the lavender genotype was also associated with slightly lower growth, as for albino[35], roux[5] and silver[36] mutations in quail. The percentage of the phenotypic variation of the body weight traits which was accounted for by the linear model (R2) did not increase by more than 3 points (Tables 1, 2 and 3) when plumage colour was included (Full model), despite statistical significance. Yet, as similar associations between lower body weight and diluted plumage colour due to various mutations were also reported in the chicken (eg: [35]), these convergent observations in quail and chicken across several independent genes for plumage colour suggest that the association may be due to a pleiotropic effect related to melanin expression. Indeed, it was found recently that the feathers of roux, lavender and albino quail, which have lower body weight than wild-type birds, each had lower total melanin content than wild-type and yellow quail [37] with similar adult body weight [6]. Further study will be needed to find the possible causes of this covariation because melanins have several suggested functions (eg: [38,39]). Most other associated phenotypic effects observed in lavender quail are probably not attributable to MLPH alone, since melanophilin defects in MLPH-mutated humans or mice have not been found to be associated with any noticeable phenotypic consequence other than the colour dilution [7].
The very significant differences in higher residual feed intake and lower body temperature in lavender quail during the feed trial (Table 2) were associated with a fair portion of the variation as R2 increased by 27 and 13 points, respectively, when the effect of plumage colour was included in the linear model. This phenotypic pattern might be related to central homeothermy-related deficiency induced by the effect of the complex mutation on PRLH as was reported in PRLH-deficient rodents [40]. Of course, other genes linked to lavender might also be involved in some of the physiological changes observed in this study.
Conclusion
As in Humans, complex mutations may occur in birds, and an extensive overlapping chromosome rearrangement was associated with a non-pathological Mendelian trait and minor, non deleterious effects in the lavender Japanese quail which is a natural knockout for PRLH.
Competing interests
The authors declare they have no competing interests.
Authors’ contributions
JLC carried out the genomic analyses, the gene expression and the genotyping, and MV and SF contributed to the early genomic work. DG supervised the production of experimental birds. OC carried out the study of PRLH activity. BB and FM designed the study, and wrote the paper. TB contributed to the study and to the redaction of the paper. All authors read and approved the final manuscript.
Supplementary Material
Table S1. List and sequence of primers used during the study. Table S2: Sequence at breakpoints for the three chromosomal changes. Table S3: Sequences containing breakpoints uploaded in GenBank database.
Figure S1. Results of the confirmation PCR for lavender breakpoints R1, R2, both R1 and R2, and R3 with band visualization after gel electrophoresis. PCR containing both R1 and R2 breakpoints was performed using primers R2_F and R1_R (see Additional file 1: Table S1). PCR are working on all lavender samples (lav) and not on wild-type samples (wt).
Contributor Information
Bertrand Bed’hom, Email: bertrand.bedhom@jouy.inra.fr.
Mohsen Vaez, Email: vaez_m@yahoo.com.
Jean-Luc Coville, Email: Jean-Luc.Coville@jouy.inra.fr.
David Gourichon, Email: David.Gourichon@tours.inra.fr.
Olivier Chastel, Email: chastel@cebc.cnrs.fr.
Sarah Follett, Email: safollett@hotmail.com.
Terry Burke, Email: t.a.burke@sheffield.ac.uk.
Francis Minvielle, Email: francis.minvielle@jouy.inra.fr.
Acknowledgments
Excellent animal care was provided by the staff at PEAT in Nouzilly. Carl-Johan Rubin is acknowledged for providing the chromosome walking protocol. At the Centre d’Etudes Biologiques de Chizé, we thank A. Lacroix and C. Trouvé for their excellent technical assistance with the prolactin assay. We are grateful to Dr. A.F. Parlow for kindly providing us with a chicken kit for prolactin assay. The Sheffield group was supported by a grant from the UK Natural Environment Research Council.
Author details
1UMR 1313 INRA/AgroParisTech, Génétique Animale et Biologie Intégrative GABI, 78352 Jouy-en-Josas, France. 2Department of Animal and Plant Sciences, University of Sheffield, Sheffield, S10 2TN, UK. 3UE 1295 INRA, Pôle d’Expérimentation Animale de Tours PEAT, 37380, Nouzilly, France. 4UPR 1934 CNRS, Centre d’Etudes Biologiques de Chizé CEBC, 79360, Beauvoir sur Niort, France. 5Present address: Division of Genetics, Department of Biology, University of Isfahan, Isfahan, Iran. 6INRA-GABI, bat 211, Centre de Recherches de Jouy, 78352 Jouy-en-Josas, France.
References
- Nadeau NJ, Minvielle F, Ito S, Inoue-Murayama M, Gourichon D, Follett SA, Burke T, Mundy NI. Characterization of Japanese quail yellow as a genomic deletion upstream of the avian homologue of the mammalian ASIP (agouti) gene. Genetics. 2008;178:777–786. doi: 10.1534/genetics.107.077073. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hiragaki T, Inoue-Murayama M, Miwa M, Fujiwara A, Mizutani M, Minvielle F, Ito S. Recessive black is allelic to the yellow plumage locus in Japanese quail and associated with a frameshift deletion in the ASIP gene. Genetics. 2008;178:771–775. doi: 10.1534/genetics.107.077040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nadeau NJ, Mundy NI, Gourichon D, Minvielle F. Association of a single nucleotide substitution in TYRP1 with roux in Japanese quail (Coturnix japonica) Anim Genet. 2007;38:609–613. doi: 10.1111/j.1365-2052.2007.01667.x. [DOI] [PubMed] [Google Scholar]
- Elvert R, Ehrhardt N, Taube M, Klingenspor M, Gailus-Durner V, Fuchs H, De Angelis MH, Blanquet V, Heldmaier G. Metabolic phenotyping of mouse mutants in the German Mouse Clinic. Integr Zool. 2006;1:122–125. doi: 10.1111/j.1749-4877.2006.00026.x. [DOI] [PubMed] [Google Scholar]
- Minvielle F, Hirigoyen E, Boulay M. Associated effects of the roux plumage colour mutation on growth, carcass traits, egg production and reproduction of Japanese quail. Poult Sci. 1999;78:1479–1484. doi: 10.1093/ps/78.11.1479. [DOI] [PubMed] [Google Scholar]
- Minvielle F, Gourichon D, Ito S, Inoue-Murayama M, Riviere S. Effects of the dominant lethal Yellow mutation on reproduction, growth, feed consumption, body temperature, and body composition in Japanese quail. Poult Sci. 2007;86:1646–1650. doi: 10.1093/ps/86.8.1646. [DOI] [PubMed] [Google Scholar]
- Ménasché G, Ho CH, Sanal O, Feldmann J, Tezcan I, Ersoy F, Houdusse A, Fischer A, de Saint Basile G. Griscelli syndrome restricted to hypopigmentation results from a melanophilin defect (GS3) or a MYO5A F-exon deletion (GS1) J Clin Invest. 2003;112:450–456. doi: 10.1172/JCI18264. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Al-Idrissi E, ElGhazali G, AlZahrani M, Ménasché G, Schmid JP, de Saint Basile G. Premature birth, respiratory distress, intracerebral hemorrhage, and silvery-gray hair: differential diagnosis of the 3 types of Griscelli syndrome. J Pediatr Hematol Oncol. 2010;23:365–374. doi: 10.1097/MPH.0b013e3181e34dc9. [DOI] [PubMed] [Google Scholar]
- Vaez M, Follett SA, Bed’hom B, Gourichon D, Tixier-Boichard M, Burke T. A single point mutation within the melanophilin gene causes the lavender plumage colour dilution phenotype in the chicken. BMC Genet. 2008;9:7. doi: 10.1186/1471-2156-9-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Matesic LE, Yip R, Reuss AE, Swing DA, O’Sullivan TN, Fletcher CF, Copeland NG, Jenkins NA. Mutations in Mlph, encoding a member of the Rab effector family, cause the melanosome transport defects observed in leaden mice. Proc Natl Acad Sci USA. 2001;98:10238–10243. doi: 10.1073/pnas.181336698. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ishida Y, David VA, Eizirik E, Schäffer AA, Neelam BA, Roelke ME, Hannah SS, O’Brien SJ, Menotti-Raymond M. A homozygous single-base deletion in MLPH causes the dilute coat colour phenotype in the domestic cat. Genomics. 2006;88:698–705. doi: 10.1016/j.ygeno.2006.06.006. [DOI] [PubMed] [Google Scholar]
- Welle M, Philipp U, Rüfenacht S, Roosje P, Scharfenstein M, Schütz E, Brenig B, Linek M, Mecklenburg L, Grest P, Drögemüller M, Haase B, Leeb T, Drögemüller C. MLPH genotype-melanin phenotype correlation in dilute dogs. J Hered. 2009;100:S75–S79. doi: 10.1093/jhered/esp010. [DOI] [Google Scholar]
- Anistoroaei R, Christensen K. Mapping of the silver gene in mink and its association with the dilution gene in dog. Cytogenet Genome Res. 2007;116:316–318. doi: 10.1159/000100417. [DOI] [PubMed] [Google Scholar]
- Minvielle F, Gourichon D, Monvoisin JL. Testing homology of loci for two plumage colours, « lavender » and « recessive white », with hybrids between chickens and Japanese quail. J Hered. 2002;93:73–76. doi: 10.1093/jhered/93.1.73. [DOI] [PubMed] [Google Scholar]
- Minvielle F, Gourichon D, Monvoisin JL. Effects of two-locus combinations, using the roux, lavender and beige mutations, on plumage colour of Japanese quail. J Hered. 2003;94:517–522. doi: 10.1093/jhered/esg091. [DOI] [PubMed] [Google Scholar]
- Brumbaugh JA, Chatterjee G, Hollander WF. Adentritic melanocytes: a mutation in linkage group II of the fowl. J Hered. 1972;63:19–25. doi: 10.1093/oxfordjournals.jhered.a108211. [DOI] [PubMed] [Google Scholar]
- Chang CM, Coville JL, Coquerelle G, Gourichon D, Oulmouden A, Tixier-Boichard M. Complete association between a retroviral insertion in the tyrosinase gene and the recessive white mutation in chickens. BMC Genomics. 2006;7:19. doi: 10.1186/1471-2164-7-19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kayang BB, Fillon V, Inoue-Murayama M, Miwa M, Leroux S, Fève K, Monvoisin JL, Pitel F, Vignoles M, Mouilhayrat C, Beaumont C, Ito S, Minvielle F, Vignal A. Integrated maps in quail (Coturnix japonica) confirm the high degree of synteny conservation with chicken (Gallus gallus) despite 35 million years of divergence. BMC Genomics. 2006;7:101. doi: 10.1186/1471-2164-7-101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sørensen AB, Duch M, Jorgensen P, Pedersen FS. Amplification and sequence-analysis of DNA flanking integrated proviruses by a simple 2-step polymerase chain-reaction method. J Virol. 1993;67:7118–7124. doi: 10.1128/jvi.67.12.7118-7124.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chastel O, Lacroix A, Weimerskirch H, Gabrielsen GW. Modulation of prolactin but not corticosterone responses to stress in relation to parental effort in a long-lived bird. Horm Behav. 2005;47:459–466. doi: 10.1016/j.yhbeh.2004.10.009. [DOI] [PubMed] [Google Scholar]
- France J, Dijkstra J, Dhanoa MS. Growth functions and their applications in animal science. Ann Zootech. 1996;45:165–174. doi: 10.1051/animres:19960637. [DOI] [Google Scholar]
- Institute SAS. SAS User’s Guide: Statistics. SAS Institute Inc, Cary; 1999. [Google Scholar]
- Herd RM, Arthur PF. 2009 Physiological basis for residual feed intake. J Anim Sci. 2009;87:E64–E71. doi: 10.2527/jas.2008-1345. [DOI] [PubMed] [Google Scholar]
- Lagerström MC, Frederiksson R, Bjarnadottir TK, Schiöth HB. The ancestry of the prolactin-releasing hormone precursor. Ann NY Acad Sci. 2005;1040:368–370. doi: 10.1196/annals.1327.064. [DOI] [PubMed] [Google Scholar]
- Heim S, Mitelman F. Cancer Cytogenetics. 3. John Wiley & Sons Inc, Hoboken; 2009. [Google Scholar]
- Andreu N, García-Rodríguez M, Volpini V, Frecha C, Molina IJ, Fontan G, Fillat C. A novel Wiskott-Aldrich syndrome protein (WASP) complex mutation identified in a WAS patient results in an aberrant product at the C-terminus from two transcripts with unusual polyA signals. J Hum Genet. 2006;51:92–97. doi: 10.1007/s10038-005-0328-7. [DOI] [PubMed] [Google Scholar]
- Wheway JM, Yau SC, Nihalani V, Ellis D, Irving M, Splitt M, Roberts RG. A complex deletion-inversion-deletion event results in a chimeric IL1RAPL1-dystrophin transcript and a contiguous gene deletion syndrome. J Med Genet. 2003;40:127–131. doi: 10.1136/jmg.40.2.127. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Halawani El ME, Silsby JL, Mauro LJ. Vasoactive intestinal peptide is a hypothalamic prolactin-releasing neuropeptide in the turkey (Meleagris gallopavo) Gen Comp Endocrinol. 1990;78:66–73. doi: 10.1016/0016-6480(90)90048-Q. [DOI] [PubMed] [Google Scholar]
- Christensen D, Vleck CM. Prolactin release and response to vasoactive intestinal peptide in an opportunistic breeder, the zebra finch (Taeniopygia guttata) Gen Comp Endocrinol. 2008;157:91–98. doi: 10.1016/j.ygcen.2008.04.013. [DOI] [PubMed] [Google Scholar]
- Lütcke A, Jansson S, Parton RG, Chavrier P, Valencia A, Huber LA, Lehtonen E, Zerial M. Rab 17, a novel small GTPase, is specific for epithelial cells and is induced during cell polarization. J Cell Biol. 1993;121:553–564. doi: 10.1083/jcb.121.3.553. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Diekmann Y, Seixas E, Gouw M, Tavares-Cadete F, Seabra MC, Pereira-Leal JB. Thousands of Rab GTPases for the cell biologist. PLoS Comput Biol. 2011;7:e1002217. doi: 10.1371/journal.pcbi.1002217. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Beaumont KA, Hamilton NA, Moores MT, Brown DL, Ohbayashi N, Cairncross O, Cook AL, Smith AG, Misaki R, Fukuda M, Taguchi T, Sturm RA, Stow JL. The recycling endosome protein Rab17 regulates melanocytic filopodia formation and melanosome trafficking. Traffic. 2011;12:627–643. doi: 10.1111/j.1600-0854.2011.01172.x. [DOI] [PubMed] [Google Scholar]
- Arakawa R, Bagashev A, Song L, Maurer K, Sullivan KE. Characterization of LRRFIP1. Biochem Cell Biol. 2010;88:899–906. doi: 10.1139/o10-014. [DOI] [PubMed] [Google Scholar]
- Yang P, An H, Liu X, Wen M, Zheng Y, Rui Y, Cao X. The cytolsolic nucleic acid sensor LRRFIP1 mediates the production of type I interferon by a beta-catenin-dependent pathway. Nat Immunol. 2010;11:487–494. doi: 10.1038/ni.1876. [DOI] [PubMed] [Google Scholar]
- Mérat P, Bordas A, Coquerelle G. The relationship of several genes suppressing plumage colour with body weight, food intake and feather loss of laying hens. Br Poult Sci. 1979;20:587–594. doi: 10.1080/00071667908416625. [DOI] [PubMed] [Google Scholar]
- Minvielle F, Bed’hom B, Coville JL, Ito S, Inoue-Murayama M, Gourichon D. The “silver” Japanese quail and the MITF gene: causal mutation, associated traits and homology with the “blue” chicken plumage. BMC Genet. 2010;11:15. doi: 10.1186/1471-2156-11-15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Minvielle F, Cecchi T, Passamonti P, Gourichon D, Renieri C. Plumage colour mutations and melanins in the feathers of the Japanese quail: a first comparison. Anim Genet. 2009;40:971–974. doi: 10.1111/j.1365-2052.2009.01929.x. [DOI] [PubMed] [Google Scholar]
- Kansaku N, Ohkubo T, Okabayashi H, Guéméné D, Kuhnlein U, Zadworny D, Shimada K. Cloning of duck PRL cDNA and genomic DNA. Gen Comp Endocrinol. 2005;141:39–47. doi: 10.1016/j.ygcen.2004.11.017. [DOI] [PubMed] [Google Scholar]
- Chakarov N, Boerner M, Krüger O. Fitness in common buzzards at the cross-point of opposite melanin-parasite interactions. Funct Ecol. 2008;22:1062–1069. doi: 10.1111/j.1365-2435.2008.01460.x. [DOI] [Google Scholar]
- Takayanagi Y, Matsumoto H, Nakata M, Mera T, Fukusumi S, Hinuma S, Ueta Y, Yada T, Leng G, Onaka T. Endogenous prolactin-releasing peptide regulates food intake in rodents. J Clin Invest. 2008;118:4014–4024. doi: 10.1172/JCI34682. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Table S1. List and sequence of primers used during the study. Table S2: Sequence at breakpoints for the three chromosomal changes. Table S3: Sequences containing breakpoints uploaded in GenBank database.
Figure S1. Results of the confirmation PCR for lavender breakpoints R1, R2, both R1 and R2, and R3 with band visualization after gel electrophoresis. PCR containing both R1 and R2 breakpoints was performed using primers R2_F and R1_R (see Additional file 1: Table S1). PCR are working on all lavender samples (lav) and not on wild-type samples (wt).



