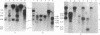Abstract
mRNA for the large subunit (LS) of ribulose-1,5-bisphosphate carboxylase [3-phospho-D-glycerate carboxylase (dimerizing), EC 4.1.1.39] of Zea mays is complementary to an uninterrupted 1600-base-pair-long chloroplast DNA sequence that has been mapped precisely within the 4350-base-pair-long chloroplast DNA fragment Bam 9 to which it had been traced earlier [Bedbrook, J. R., Coen, D. M., Beaton, A. R., Bogorad, L. & Rich, A. (1979) J. Biol. Chem. 254, 905-910]. An additional 1400-base-pair-long uninterrupted region that is colinear with a chloroplast RNA has been detected on Bam 9. The transcript from this region is part of a 2200-nucleotide-long RNA. The remainder of the DNA sequence for the 2200-base-pair RNA maps outside Bam 9. The 1600-base-pair LS gene and the gene for the 2200-nucleotide transcript are close to one another. They are separated by an untranscribed intercistronic “gap” about 330 base pairs long. These two closely packed genes are inverted on the chromosome—i.e., their 3′ termini are at opposite ends of the untranscribed gap and they map on opposite strands.
Keywords: ribulosebisphosphate carboxylase, 2.2-kilobase gene, plasmid
Full text
PDF


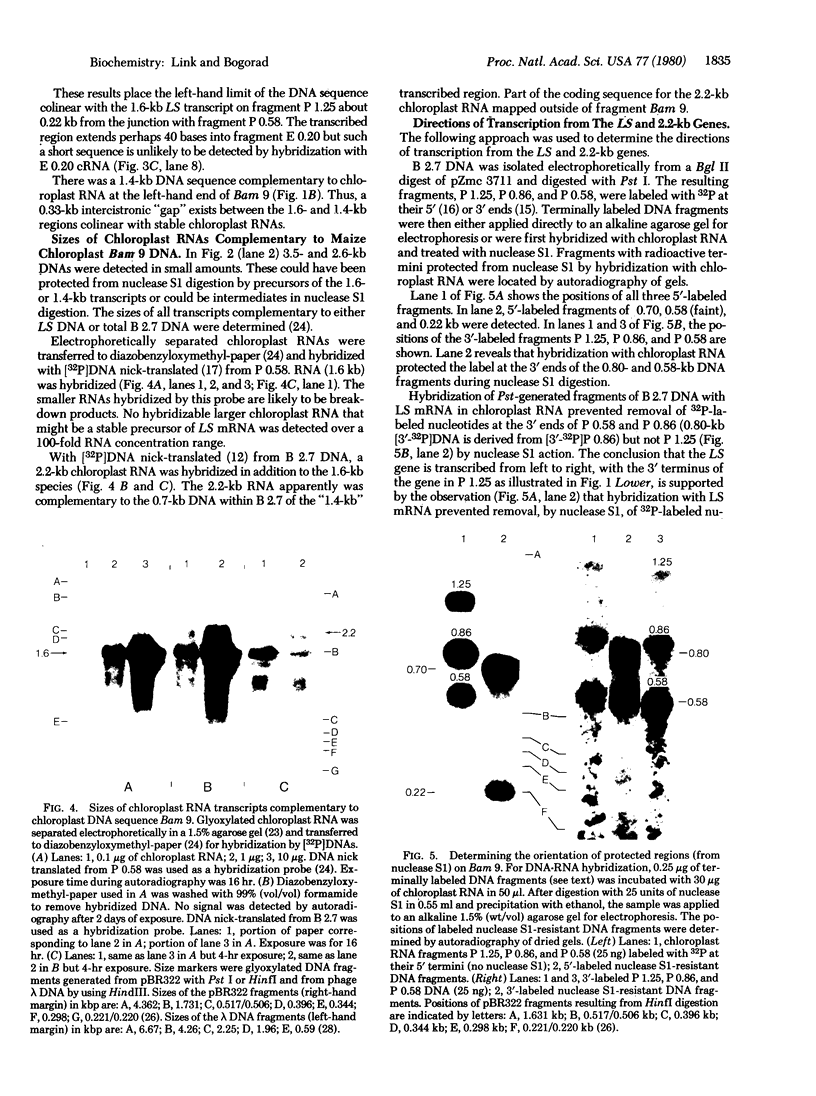

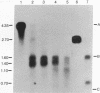
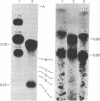
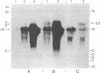
Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Alwine J. C., Kemp D. J., Stark G. R. Method for detection of specific RNAs in agarose gels by transfer to diazobenzyloxymethyl-paper and hybridization with DNA probes. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5350–5354. doi: 10.1073/pnas.74.12.5350. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Alwine J. C., Kemp D. J., Stark G. R. Method for detection of specific RNAs in agarose gels by transfer to diazobenzyloxymethyl-paper and hybridization with DNA probes. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5350–5354. doi: 10.1073/pnas.74.12.5350. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bedbrook J. R., Bogorad L. Endonuclease recognition sites mapped on Zea mays chloroplast DNA. Proc Natl Acad Sci U S A. 1976 Dec;73(12):4309–4313. doi: 10.1073/pnas.73.12.4309. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bedbrook J. R., Coen D. M., Beaton A. R., Bogorad L., Rich A. Location of the single gene for the large subunit of ribulosebisphosphate carboxylase on the maize chloroplast chromosome. J Biol Chem. 1979 Feb 10;254(3):905–910. [PubMed] [Google Scholar]
- Bedbrook J. R., Kolodner R., Bogorad L. Zea mays chloroplast ribosomal RNA genes are part of a 22,000 base pair inverted repeat. Cell. 1977 Aug;11(4):739–749. doi: 10.1016/0092-8674(77)90288-4. [DOI] [PubMed] [Google Scholar]
- Berk A. J., Sharp P. A. Sizing and mapping of early adenovirus mRNAs by gel electrophoresis of S1 endonuclease-digested hybrids. Cell. 1977 Nov;12(3):721–732. doi: 10.1016/0092-8674(77)90272-0. [DOI] [PubMed] [Google Scholar]
- Berk A. J., Sharp P. A. Spliced early mRNAs of simian virus 40. Proc Natl Acad Sci U S A. 1978 Mar;75(3):1274–1278. doi: 10.1073/pnas.75.3.1274. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bolivar F., Rodriguez R. L., Greene P. J., Betlach M. C., Heyneker H. L., Boyer H. W., Crosa J. H., Falkow S. Construction and characterization of new cloning vehicles. II. A multipurpose cloning system. Gene. 1977;2(2):95–113. [PubMed] [Google Scholar]
- Burgess R. R., Jendrisak J. J. A procedure for the rapid, large-scall purification of Escherichia coli DNA-dependent RNA polymerase involving Polymin P precipitation and DNA-cellulose chromatography. Biochemistry. 1975 Oct 21;14(21):4634–4638. doi: 10.1021/bi00692a011. [DOI] [PubMed] [Google Scholar]
- Casey J., Davidson N. Rates of formation and thermal stabilities of RNA:DNA and DNA:DNA duplexes at high concentrations of formamide. Nucleic Acids Res. 1977;4(5):1539–1552. doi: 10.1093/nar/4.5.1539. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clewell D. B., Helinski D. R. Supercoiled circular DNA-protein complex in Escherichia coli: purification and induced conversion to an opern circular DNA form. Proc Natl Acad Sci U S A. 1969 Apr;62(4):1159–1166. doi: 10.1073/pnas.62.4.1159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Coen D. M., Bedbrook J. R., Bogorad L., Rich A. Maize chloroplast DNA fragment encoding the large subunit of ribulosebisphosphate carboxylase. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5487–5491. doi: 10.1073/pnas.74.12.5487. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gray P. W., Hallick R. B. Physical mapping of the Euglena gracilis chloroplast DNA and ribosomal RNA gene region. Biochemistry. 1978 Jan 24;17(2):284–289. doi: 10.1021/bi00595a015. [DOI] [PubMed] [Google Scholar]
- Heffron F., Sublett R., Hedges R. W., Jacob A., Falkow S. Origin of the TEM-beta-lactamase gene found on plasmids. J Bacteriol. 1975 Apr;122(1):250–256. doi: 10.1128/jb.122.1.250-256.1975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Link G., Coen D. M., Bogorad L. Differential expression of the gene for the large subunit of ribulose bisphosphate carboxylase in maize leaf cell types. Cell. 1978 Nov;15(3):725–731. doi: 10.1016/0092-8674(78)90258-1. [DOI] [PubMed] [Google Scholar]
- Maxam A. M., Gilbert W. A new method for sequencing DNA. Proc Natl Acad Sci U S A. 1977 Feb;74(2):560–564. doi: 10.1073/pnas.74.2.560. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McDonell M. W., Simon M. N., Studier F. W. Analysis of restriction fragments of T7 DNA and determination of molecular weights by electrophoresis in neutral and alkaline gels. J Mol Biol. 1977 Feb 15;110(1):119–146. doi: 10.1016/s0022-2836(77)80102-2. [DOI] [PubMed] [Google Scholar]
- Murray K., Murray N. E. Phage lambda receptor chromosomes for DNA fragments made with restriction endonuclease III of Haemophilus influenzae and restriction endonuclease I of Escherichia coli. J Mol Biol. 1975 Nov 5;98(3):551–564. doi: 10.1016/s0022-2836(75)80086-6. [DOI] [PubMed] [Google Scholar]
- Rigby P. W., Dieckmann M., Rhodes C., Berg P. Labeling deoxyribonucleic acid to high specific activity in vitro by nick translation with DNA polymerase I. J Mol Biol. 1977 Jun 15;113(1):237–251. doi: 10.1016/0022-2836(77)90052-3. [DOI] [PubMed] [Google Scholar]
- Rochaix J. D., Malnoe P. Anatomy of the chloroplast ribosomal DNA of Chlamydomonas reinhardii. Cell. 1978 Oct;15(2):661–670. doi: 10.1016/0092-8674(78)90034-x. [DOI] [PubMed] [Google Scholar]
- Roychoudhury R., Jay E., Wu R. Terminal labeling and addition of homopolymer tracts to duplex DNA fragments by terminal deoxynucleotidyl transferase. Nucleic Acids Res. 1976 Apr;3(4):863–877. doi: 10.1093/nar/3.4.863. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Southern E. M. Detection of specific sequences among DNA fragments separated by gel electrophoresis. J Mol Biol. 1975 Nov 5;98(3):503–517. doi: 10.1016/s0022-2836(75)80083-0. [DOI] [PubMed] [Google Scholar]
- Sutcliffe J. G. pBR322 restriction map derived from the DNA sequence: accurate DNA size markers up to 4361 nucleotide pairs long. Nucleic Acids Res. 1978 Aug;5(8):2721–2728. doi: 10.1093/nar/5.8.2721. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thuring R. W., Sanders J. P., Borst P. A freeze-squeeze method for recovering long DNA from agarose gels. Anal Biochem. 1975 May 26;66(1):213–220. doi: 10.1016/0003-2697(75)90739-3. [DOI] [PubMed] [Google Scholar]
- Whitfeld P. R., Herrmann R. G., Bottomley W. Mapping of the ribosomal RNA genes on spinach chloroplast DNA. Nucleic Acids Res. 1978 Jun;5(6):1741–1751. doi: 10.1093/nar/5.6.1741. [DOI] [PMC free article] [PubMed] [Google Scholar]