Abstract
Aptamers are single-stranded oligonucleotides that bind to targets with high affinity and selectivity. Their use as molecular recognition elements has emerged as a viable approach for biosensing, diagnostics, and therapeutics. Despite this potential, relatively few aptamers exist that bind to small molecules. Small molecules are important targets for investigation due to their diverse biological functions as well as their clinical and commercial uses. Novel, effective molecular recognition probes for these compounds are therefore of great interest. This paper will highlight the technical challenges of aptamer development for small molecule targets, as well as the opportunities that exist for their application in biosensing and chemical biology.
1. Aptamers as Molecular Recognition Elements
Historically, nucleic acids were associated with the storage and genetic coding of information and have long been thought to be less complex than proteins [1]. However, like proteins, nucleic acids are able to fold into intricate tertiary structures that have the potential to perform a variety of functions including gene-regulation, catalytic activity and ligand-binding [2]. Interest in these so-called “functional” nucleic acids was prompted by the ever-increasing number of discoveries of non-coding ribonucleic acids (RNAs) displaying catalytic or binding properties [2].
Two decades ago, several researchers revolutionized molecular recognition by developing synthetic RNA motifs that bound specifically to molecular targets [3–5]. These RNA structures, called aptamers, were selected using an in vitro selection procedure called systematic evolution of ligands by exponential enrichment (SELEX) [3]. Like antibodies, these synthetically derived molecular recognition probes were found to be selective and able to bind to their targets with high affinity.
Currently, there is a growing need for rapid, robust, and inexpensive methods for sensing and diagnostic purposes [6]. As molecular recognition is the cornerstone of sensing, there has been increased focus on the development of new molecular recognition probes for sensing applications [7]. While antibodies have long been considered to be the standard in molecular recognition and the use of antibodies as recognition probes predates the 1950s, the relatively new technology of aptamers offers several advantages [8]. Firstly, the in vitro aptamer selection process allows a greater control over aptamer binding conditions. Nonphysiological salt concentrations, temperatures and pH can be used in successful selections [9]. Due to the robustness of the phosphodiester backbone, aptamers can exhibit an improved stability over their protein-based antibody counterparts. In particular, aptamers can be reversibly denatured by changing the surrounding conditions. For example, a change in pH, temperature, ionic strength, or use of denaturants irreversibly denatures antibodies, while aptamers simply unfold. The aptamer structure can then regain functionality upon return of the original binding conditions [6]. Due to the nucleic acid nature of aptamers, they bind to complementary nucleic acids as well as their targets, which can be exploited in sensing schemes or as “antidotes” in vivo. Aptamer generation has been achieved for a wide variety of targets including small molecules [10], proteins [11], viruses [12], and whole cells [13]. Unlike antibodies, aptamers can also be generated for targets that are toxic as well as for targets that do not elicit an immune response in vivo [8]. Once selected, aptamers are manufactured using well-established automated chemical solid-phase synthesis [14, 15]. The accuracy and reproducibility of this procedure allows for a relative ease in producing aptamers at large scales, with very little batch-to-batch variation in activity [16]. Additionally, aptamer sequences can be modified with reporter molecules throughout this solid-phase synthesis; this allows for labeling at judiciously chosen nucleotide positions to minimize any effect on the functionality of the aptamer [17, 18].
Aptamers also offer advantages over other synthetically created molecular recognition systems such as molecular imprinted polymers (MIPs). While MIP synthesis can be simple and cheap, and the resulting MIPs are unaffected by changes in heat and pH [19], MIPs typically display high cross-reactivity [20] and are not particularly amenable to chemical modifications.
Of course, aptamers are not without their disadvantages. Unlike antibodies or MIPS, their tertiary structure is highly dependent on solution conditions, and they are easily degraded in blood. Furthermore, antibodies have a significantly higher chemical diversity with 20 amino acids. However, some of these problems can be addressed, for example, through chemical modifications to increase nuclease resistance or increase the diversity of the nucleic acid pools.
1.1. Systematic Evolution of Ligands by Exponential Enrichment (SELEX)
The concept of in vitro evolution was first reported in the 1960s with the observation that, in a cell-free system, the RNA genome of the Qβ bacteriophage could be evolved during replication to form RNAs that were more efficiently copied by the viral replicase [21]. Later, they were able to evolve sequences for other traits such as resistance to ethidium bromide [22]. Despite the importance of these early discoveries, however, the true potential of in vitro evolution was not realized until several decades later, after the introduction of modern biotechnological advances such as the invention of polymerase chain reaction (PCR), the isolation of reverse transcriptase and the ability to generate long oligonucleotides containing random nucleotide regions using solid-phase synthesis. Equipped with these modern techniques, in 1990, three separate groups reported in vitro selection and evolution of functional nucleic acids [23]. Tuerk and Gold [3] used the term SELEX for their process of selecting RNA ligands against T4 DNA polymerase; Ellington and Szostak [4] performed in vitro selection to select RNA ligands (for which they coined the term “aptamers”) against various organic dyes; Robertson and Joyce [5] evolved the Tetrahymena self-splicing intron to carry out a DNA cleavage reaction.
Since its invention, several researchers have performed SELEX to isolate nucleic acids with a wide variety of functions. While several modifications of the procedure have been made by various groups, the general SELEX process remains the same (Figure 1 shows the process for DNA aptamers). Typically, SELEX begins with an initial library (often referred to as “a pool”) of random nucleic acid sequences (either RNA or DNA depending on the nature of the research). SELEX libraries ideally consist of 30–80 random nucleotide positions flanked by primer-binding sites necessary for PCR amplification [24]. The library is then incubated with the target of interest and several washing steps are employed to remove nonfunctional sequences. For small molecule targets, the target is usually immobilized onto a solid-support matrix to permit partitioning of binding and nonbinding sequences [25]. The next steps in SELEX include the elution of the binding sequences from the target and the polymerase chain reaction (PCR) amplification (reverse transcription PCR for the RNA aptamers) of those binding sequences to yield an enriched library for subsequent, more stringent, selection rounds [10]. As the interactions that lead to molecular recognition between the binding sequences and the target are noncovalent in nature, mild conditions can be used to separate the two species. Elution using heat, high concentrations of the target molecule, or chaotropic agents, such as urea, can be performed. The strength of the molecular interactions within the target-aptamer complex will dictate the conditions required for elution [7].
Figure 1.
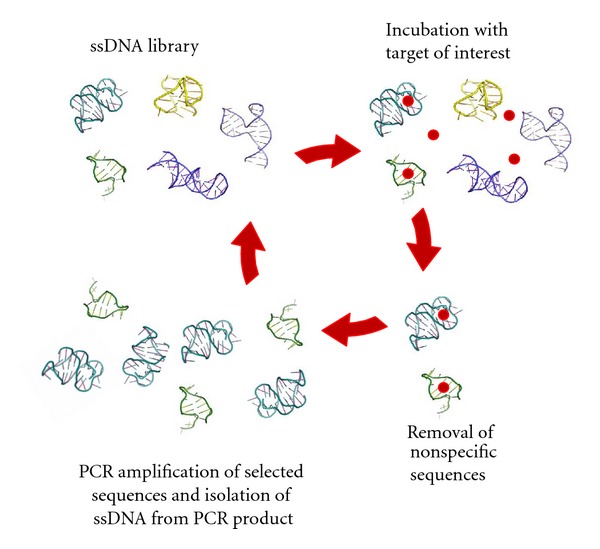
The systematic evolution of ligands by exponential enrichment process (SELEX). Beginning with a large library of DNA, iterative cycles of target incubation, library partitioning, and amplification are performed to select aptamers.
Once separated from the target, the few binding DNA sequences are amplified by PCR to yield a practical amount of sample to continue the process. As PCR generates double-stranded DNA and aptamers are single stranded, the DNA aptamer sequence is separated from its complement using one of a number of techniques, such as gel electrophoresis or using an agarose resin [26, 27]. Single-stranded RNA aptamers are generated from the double-stranded DNA PCR products by in vitro transcription [10], thus no further processing is required before reintroduction of an enriched RNA pool into the next SELEX round.
SELEX progress can be monitored by modifying the aptamer strand with a traceable label, to determine when more stringent conditions should be applied [28]. The enriched library generated from a round of selection is subjected to further selection rounds that serve to increase the affinity of the library for the target molecule (positive selections). After several rounds, the enriched library is cloned, sequenced and characterized to isolate aptamers with the desired properties. Once these sequences are elucidated, solid-phase chemical synthesis is used to reproducibly synthesize aptamers in large quantities.
1.2. Adaptations to the SELEX Process
An enormous advantage of SELEX is its flexibility with respect to methodology, binding conditions and library design. The first SELEX modifications introduced the inclusion of negative or counter selection steps to eliminate sequences displaying affinity for either the solid-support matrix or compounds sharing structural similarity to the target. The majority of more recent SELEX modifications involve changing the stringency, the platform on which selection is performed or the type of target [29, 30]. The library used in the selection can also be modified to include fixed regions of known functionality or increase the diversity of structures available for selection either through initial pool design [28, 31, 32], or by inclusion of mutagenic PCR to alter the pool from round to round [33]. The SELEX process has also been automated by several groups [34–36]. Table 1 lists several modifications to the original SELEX process. Regardless of whether the listed SELEX modifications involved changes to target immobilization, nucleic acid library, selection stringency, amplification, or monitoring of the enrichment, the goal of these changes was to either generate improved aptamers or to simplify the SELEX procedure.
Table 1.
A list of modifications to the SELEX process and their descriptions.
| Method | Description | Reference |
|---|---|---|
| Atomic force microscopy (AFM)-SELEX | AFM-SELEX uses a dynamic atomic force microscopy tip to pick up and visualize aptamer-target complexes. This SELEX requires only one round of selection. | [99] |
| Automated SELEX | This SELEX uses automated systems for the procedure to reduce the time and labour required. | [34] |
| Blended SELEX | In this technique, a lead chemical compound is attached covalently or non-covalently to a nucleic acid library. Each nucleic acid conjugate in the starting library is a variant of the chemical compound moiety and allows up to 1015 variants of the small molecule to be screened for the most active of these composite assemblies. | [100] |
| Cell-SELEX | Cell-SELEX generates aptamers that can bind specifically to a cell of interest. Commonly, a cancer cell line is used as the target to generate aptamers that can differentiate that cell from other cancers or normal cells. | [101] |
| Capillary electrophoresis (CE)-SELEX | The separation of bound and nonbound oligonucleotides is performed using capillary electrophoresis. | [102] |
| Chimeric SELEX | Chimeric SELEX uses two or more different oligonucleotide libraries for production of chimeric aptamers with more than one wanted feature or function. Each of the parent libraries will be selected first to a distinct feature; the resulting aptamers are then fused together. | [103] |
| Conditional SELEX | This SELEX uses regulator molecules during the selection, thus, allowing aptamer binding to the target to be regulated. | [104] |
| Counter selection/ subtractive SELEX | This technique employs additional rounds of SELEX to remove sequences that bind to similar target structures. | [44] |
| Covalent/ Crosslinking SELEX | This process is used to select aptamers that contain reactive groups which are capable of covalent linking to a target protein. | [105] |
| Deconvolution SELEX | Deconvolution SELEX is used to generate aptamers for complex targets. Typically selection is performed on mixtures (or a cell). Once aptamers have been generated, a second part of SELEX involves discriminating which aptamers bind to which parts of the complex mixture. | [106] |
| Electrophoretic mobility shift assay (EMSA)-SELEX | The partitioning step of SELEX occurs through the use of electrophoretic mobility shift assay (EMSA) at every round. | [107] |
| Expression cassette SELEX | This is a special form of blended SELEX that involves transcription factors and optimizes aptamer activity for gene therapy applications. | [108] |
| Fluorescence-activated cell sorting (FACS) SELEX | This SELEX makes use of fluorescence-activated cell sorting to differentiate and separate aptamer-bound cells. | [59] |
| FluMag SELEX | Here the library is modified with fluorescein instead of radiolabels for quantification purposes. Additionally, the target is immobilized to magnetic beads instead of agarose. | [109] |
| Genomic SELEX | The SELEX library is constructed from an organism's genome and target proteins and metabolites from the same organism are used to elucidate meaningful interactions. | [110] |
| In vivo SELEX | In vivo SELEX uses transient transfection in an iterative procedure in cultured vertebrate cells to select for RNA-processing signals. | [111] |
| Indirect SELEX | The target used in the selection is not the aptamer binder; however, it becomes required for aptamer binding to the new target. | [112] |
| Mod-SELEX | Mod-SELEX uses a library of oligonucleotides with chemical substitutions that result in nuclease-resistant aptamers. | [113] |
| Multivalent aptamer isolation (MAI) SELEX | This process is used to generate aptamer pairs for a given target. | [114] |
| Microfluidics SELEX | This SELEX uses microfluidic technologies, creating an automatic, and miniature SELEX platform for fast aptamer screening. | [115, 116] |
| Monolex | Monolex involves a single affinity chromatography step, followed by physical segmentation of the affinity material, to obtain the highest affinity aptamers. | [117] |
| Multiplexed massively parallel SELEX | This allows analysis of large numbers of transcription factors in parallel through the use of affinity-tagged proteins, bar-coded selection oligonucleotides, and multiplexed sequencing. | [118] |
| Multi-stage SELEX | Multistage SELEX is a modified version of chimeric selex. Here, the fused aptamer components then go through an additional selection with all the targets. | [119] |
| Negative selection | An additional step, performed typically at the beginning of selection, removes sequences that have an affinity for the selection matrix. | [48] |
| Next generation SELEX | This SELEX uses designed oligonucleotide libraries that tile through a pre-mRNA sequence. The pool is then partitioned into bound and unbound fractions, which are quantified by a two-color microarray. | [120] |
| Non-SELEX (NCEEM) | This process involves repetitive steps of partitioning with no amplification steps. | [121] |
| Photo SELEX | Aptamers bearing photo-reactive groups that can photo cross-link to a target and/or photo activate a target molecule are used. | [122] |
| Primer-free SELEX | This SELEX involves removal of the primer-annealing sequences from the library prior to selection, preventing unwanted primer-based secondary structures. | [123] |
| Serial analysis of gene expression (SAGE) or high- throughput SELEX | SAGE SELEX links oligomers from SELEX with longer DNA molecules that can be efficiently sequenced. | [124] |
| Spiegelmer technology | The aptamer selection is performed with the natural D-nucleic acids but on the opposite enantiomer of the chiral target molecule. After sequencing, the aptamers are synthesized as L-isomers for binding to the desired enantiomer of the target. | [125] |
| Slow off-rate modified aptamers (SOMAmer) | The selection is performed with oligonucleotide libraries that are uniformly functionalized at the 5′-position resulting in high-quality aptamers. | [28] |
| Tailored SELEX | This is an integrated method to identify aptamers with only 10 fixed nucleotides through ligation and removal of primer binding sites within the SELEX process. | [126] |
| Target expressed on cell surface (TECS) SELEX | Recombinant proteins on the cell surface are used directly as the selection target. | [127] |
| Tissue-SELEX | This method is for generating aptamers capable of binding to tissue targets. | [106] |
| Toggle-SELEX | The selection is performed on different targets in alternating rounds. | [128] |
| Yeast Genetic SELEX | This method optimizes in vitro selected aptamers by creating a library of degenerate aptamers and performing a secondary selection in vivo using a yeast three (one)-hybrid system. | [129] |
2. SELEX Targets
As can be noted from Figure 2, less than a quarter of existing aptamers have been generated for small molecule targets. With the success of the first in vitro selection experiments to small organic dyes [4], much of the original SELEX focus was on developing aptamers for small molecules. However, once it was found that aptamers could be easily selected for proteins and cells, new aptamers for small molecules became less prevalent. These larger targets, containing more functional groups and structural motifs, give a higher probability of finding sequences that can interact with the target via hydrogen bonds, electrostatic interactions, and hydrophobic interactions [6].
Figure 2.

Breakdown, by target type, of aptamers selected between 1990 and 2011. This list was generated using the Aptamer Base [41] http://aptamerbase.semanticscience.org/ (accessed July 9, 2012) (accessed July 9, 2012).
2.1. Small Molecule-Binding Aptamers
Despite this trend towards larger targets, there are many compelling reasons for pursuing the identification of new small molecule-binding aptamers. Small molecules play key roles in many biological processes due to their ability to diffuse across cell membranes [37]. These targets may be harmful, such as toxins and carcinogens, or beneficial, such as drugs or nutrients. In cells, small molecules serve as cell signaling molecules, pigments, or as part of defense mechanisms [38]. In molecular biology, they can be used as antibiotics or other important drugs [39]. In the food industry, small molecules are important for energy storage or can act as pesticides [40]. Aptamers for small molecules may be applied to a wide variety of applications in medicine, agriculture, and environmental analysis. Tables 2 and 3 list the small molecule targets for which DNA and RNA aptamers, respectively, have been characterized.
Table 2.
A listing of DNA aptamers reported in the open literature* (up until July 2012) that have been confirmed to bind to small molecule targets. The dissociation constant (K d), a measure of binding affinity, is included as well as the year of aptamer development.
| Target | Binding affinity (K d) | Year | Reference |
|---|---|---|---|
| Reactive green 19 | 33 μM | 1992 | [130] |
| Adenosine monophosphate and adenosine triphosphate | 6 μM | 1995 | [131] |
| L-arginine | 2.5 mM | 1995 | [132] |
| L-argininamide | 0.25 mM | 1995 | [132] |
| Anionic porphyrins | 0.4–4.9 μM | 1996 | [88] |
| Sulforhodamine B | 190 nM | 1998 | [61] |
| Cellobiose | 600 nM | 1998 | [133] |
| 7,8-dihydro-8-hydroxy-2′-deoxyguanosine | 270 nM | 1998 | [134] |
| Cholic acid | 5–67.5 μM | 2000 | [135] |
| Hematoporphyrin | 1.6 μM | 2000 | [136] |
| L-tyrosinamide | 4.5 μM | 2001 | [137] |
| Sialyllactose | 4.9 μM | 2004 | [138] |
| Ethanolamine | 6–19 nM | 2005 | [139] |
| (R)-thalidomide | 1 μM | 2007 | [49] |
| Hoechst derivative 7e | 878 nM | 2007 | [140] |
| 17β-estradiol | 0.13 μM | 2007 | [141] |
| Lys-Arg-Azobenzene-Arg | 0.33 μM | 2007 | [142] |
| Tetracycline | 64 nM | 2008 | [143] |
| L and D arginine | 580–810 μM | 2008 | [144] |
| Daunomycin | 10 nM | 2008 | [145] |
| Oxytetracycline | 10 nM | 2008 | [54] |
| Ochratoxin A | 200 nM | 2008 | [26] |
| Dopamine | 700 nM | 2009 | [146] |
| 8-hydroxy-2′-deoxyguanosine | 100 nM | 2009 | [147] |
| Diclofenac | 42.7–166.34 nM | 2009 | [148] |
| (S) and (R)-ibuprofen | 1.5–5.2 μM | 2010 | [50] |
| Adenosine triphosphate | 3.7 μM | 2010 | [31] |
| Fumonisin B1 (FB1) | 100 nM | 2010 | [7] |
| Acetamiprid | 4.98 μM | 2011 | [149] |
| Kanamycin | 78.8 nM | 2011 | [150] |
| L-tryptophan | 1.757 μM | 2011 | [151] |
| Bisphenol A | 8.3 nM | 2011 | [53] |
| Ochratoxin A | 96–293 nM | 2011 | [152] |
| Phenylphosphonic dichloride | >50 μM | 2011 | [153] |
| Organophosphorus pesticides (phorate, profenofos, isocarbophos and omethoate) | 0.8–2.5 μM | 2012 | [154] |
| Polychlorinated biphenyls (PCB77) | 4.02, 8.32 μM | 2012 | [155] |
| Polychlorinated biphenyls (PCB72 and PCB106) | 60–100 nM | 2012 | [156] |
| Ampicillin | 9.4–13.4 nM | 2012 | [157] |
*Only aptamer sequences that have experimentally determined K d values were included in this table.
Table 3.
A listing of RNA aptamers reported in the open literature* (up until July 2012) that have been confirmed to bind to small molecule targets. The dissociation constant (K d), a measure of binding affinity, is included as well as the year of aptamer development.
| Target | Binding affinity (K d) | Year | Reference |
|---|---|---|---|
| Organic dyes | 100–600 μM | 1990 | [158] |
| D-tryptophan | 18 μM | 1992 | [46] |
| L-valine | 2.9 mM | 1994 | [159] |
| Theophylline | 100 nM | 1994 | [44] |
| Cyanocobalamin | 88 nM | 1994 | [160] |
| L-citrulline | 62–68 μM | 1994 | [47] |
| Flavin mononucleotide | 0.5 μM | 1994 | [161] |
| Flavin adenine dinucleotide | 137–273 μM | 1994 | [161] |
| Kanamycin A | ≤300 nM | 1995 | [162] |
| Neomycin | 100 nM | 1995 | [163] |
| Tobramycin | 2–3 nM | 1995 | [164] |
| Lividomycin | ≤300 nM | 1995 | [162] |
| Nicotinamide adenine dinucleotide | 2.5 μM | 1995 | [165] |
| Riboflavin | 1–5 μM | 1995 | [165] |
| Biotin | 5 μM | 1995 | [166] |
| L-arginine | 330 nM | 1996 | [48] |
| Dopamine | 2.8 μM | 1997 | [167] |
| 7-methyl-guanosine | 5 μM | 1997 | [168] |
| CCdApPuro | 10 nM | 1997 | [169] |
| Chloramphenicol | 25–65 μM | 1997 | [170] |
| Viomycin | 11–21 μM | 1997 | [171] |
| Sulforhodamine B | 310 nM | 1998 | [172] |
| Streptomycin | 1–10 μM | 1998 | [173] |
| L-isoleucine | 200–500 μM | 1998 | [174] |
| 7,8-dihydro-8-hydroxy-2′-deoxyguanosine (8-oxodG) | 0.27–2.8 μM | 1998 | [134] |
| Xanthine | 3.3 μM | 1998 | [175] |
| Guanine | 1.3 μM | 1998 | [175] |
| Malachite green | ≤1 μM | 1999 | [176] |
| Phosphatidylcholine | ≥100 μM | 1999 | [177] |
| Cyclic adenosine monophosphate | 10 μM | 2000 | [178] |
| Adenosine triphosphate | 127–223 μM | 2000 | [179] |
| L-tyrosine | 35 μM | 2000 | [180] |
| S-adenosyl homocysteine | 0.2–0.8 μM | 2000 | [181] |
| Neomycin | 1.8 μM | 2000 | [182] |
| Moenomycin A | 300–400 nM | 2001 | [183] |
| Sialyl Lewis X | 0.085–10 nM | 2001 | [184] |
| Tetracycline | 1 μM | 2001 | [185] |
| Kanamycin B | 180 nM | 2001 | [186] |
| Adenine | 10 μM | 2002 | [187] |
| Flavin adenine dinucleotide | 50 μM | 2002 | [188] |
| L-isoleucine | 1–7 mM | 2003 | [189] |
| Adenosine triphosphate | 2 μM | 2003 | [190] |
| Morpholine-based GTP analog | 20, 33 μM | 2003 | [191] |
| 4,4′-methylenedianiline | 0.45–15 μM | 2004 | [192] |
| Tobramycin | 30–100 nM | 2004 | [34] |
| Kanamycin | 10–30 nM | 2004 | [34] |
| Adenosine triphosphate | 5 μM | 2004 | [193] |
| Isoleucine | 0.9 mM | 2005 | [194] |
| L-histidine | 8–54 μM | 2005 | [195] |
| Codeine | 2.5–4 μM | 2006 | [33] |
| Mesomesoprotoporphyrin IX | 188–445 nM | 2006 | [87] |
| Thyroxine | 50 μM | 2007 | [196] |
| Tobramycin | 16 μM | 2007 | [197] |
| 10-carboxy-2,7-di-t-butyl-trans-12c,12d-dimethyl-12c,12d-dihydrobenzo[e]pyrene | 2.7 μM | 2007 | [198] |
| Dimethylindole red | 87 nM | 2008 | [199] |
| Cyanine 3 dye | 60 μM | 2010 | [200] |
| Aniline-substituted sulforhodamine analogue | 3.5 μM | 2010 | [201] |
| Atrazine | 2 μM | 2010 | [202] |
| Sphingosylphosphorylcholine | 20–250 nM | 2010 | [203] |
| Black hole quencher | 4.7 μM | 2011 | [204] |
| 4-dimethylaminobenzylidene imidazolinone | 464 nM | 2011 | [205] |
| Glutathione | 41.8, 48.9 nM | 2011 | [206] |
| Heteroaryldihydropyrimidine | 50 nM | 2011 | [207] |
*Only aptamer sequences that have experimentally determined K d values were included in this table.
2.2. Conceptual Challenges for Small Molecule Aptamers
Although they represent a minor proportion of all aptamers, small molecule-binding aptamers are among the most successful and widely studied aptamers in the literature. For example, the ATP aptamer is second only to the thrombin aptamer in terms of the number of publications using the sequence in an aptamer-based assay, sensor, or biosensor in the last ten years. The cocaine and theophylline aptamers are the fifth and seventh most frequently used aptamers for biosensing, respectively [42]. It has already been described that aptamers are ideal molecular recognition probes for small molecules [8, 43], based on their ability to achieve a remarkably high degree of selectivity. The first example of this unparalleled selectivity was observed in 1994, when the selected RNA aptamer for theophylline displayed a 10,000 times weaker binding affinity to caffeine, a xanthine that differs by a single methyl group. This selectivity was found to be a 10-fold improvement on the selectivity for the antibodies for these targets [44]. Several groups have also exploited the ability of aptamers to distinguish between small molecule enantiomers [45]. Initially, several RNA aptamers displayed partial discrimination between various L and D amino acids [46, 47]. Then, in 1996, Geiger et al. [48] reported the selection of RNA aptamers that bound to L-arginine with high affinity and enantioselectivity. More recently, enantioselective DNA aptamers have been selected for the small molecule drug (R)-thalidomide [49] and separate aptamers have been identified for (S) and (R)-ibuprofen [50].
One possible explanation for the scarcity of new small molecule aptamers is the impression that aptamers cannot bind these smaller targets with the high affinity required for most sensing applications. Work by Carothers et al. attempted to determine the effect of target structure and size on binding affinity [51]. Using aptamers for 6 small molecule targets from the literature, as well as aptamers obtained from his own selections for two other small molecules, Carothers determined that the target molecular weight was proportional to the resulting aptamer affinity (larger targets resulted in lower K d values). This finding was consistent with the findings of other studies between affinity and target mass [52]. However, the target theophylline which has a very small mass (180 g/mol) did not follow this general trend. It was therefore concluded that targets with fewer rotatable bonds, and therefore fewer degrees of freedom, can result in improved aptamer affinity. Nevertheless, while many aptamers that bind to small molecule targets display affinities in the low to mid micromolar range, there are several aptamers that have recently been isolated with K d values in the low nanomolar range (e.g., BPA [53] and oxytetracycline [54]). Furthermore, riboswitches, which are widely considered as containing “natural aptamers,” bind exclusively to small molecules and ions, and several of these display remarkably strong binding. For example, the guanine riboswitch has a K d of 5 nM [55] and the thiamine pyrophosphate-sensing riboswitch has an affinity in the picomolar range [56]. The glycine riboswitch is particularly noteworthy for its ability to selectively bind to one of the smallest target of any natural or artificial aptamer [57]. Nature's effectiveness at developing small molecule aptamers should provide an indication that there is considerable untapped potential in this field.
In the early 1990s, in an effort to promote the power of SELEX, numerous papers and reviews boasted that in vitro selection is facile, inexpensive, and fail-safe, which may have contributed to little interest in publications for new selections. On the contrary, SELEX can be very laborious and it has been estimated that less than 30% of selections result in aptamers [28]. Additionally, patents for virtually every application of aptamers have placed a stranglehold on aptamer innovation [58]. As a result, very few research groups have chosen to invest the time and expense to develop aptamers for new small molecule targets, especially considering the unique technical challenges that arise when selecting for small molecule binding aptamers, as is discussed in the next section.
3. Technical Challenges for Small Molecule Aptamers
3.1. Target Immobilization
The separation of target-bound sequences from those with no affinity for the target is a critical step in the SELEX process. For protein targets, partitioning can be achieved using a matrix that selectively adsorbs the target and any interacting aptamer sequences. For example, nitrocellulose filters are a cheap and convenient matrix for this purpose due to their nucleic acid permeability and their ability to retain proteins by hydrophobic adsorption. With cell targets, partitioning can be accomplished by centrifugation, fluorescence activated cell sorting (FACS), [59] or by gentle washing of adherent cells [13]. In the case of both these target types, the selection can be accomplished without chemical modification of the target; this is ideal since it increases the likelihood of finding aptamers capable of binding the molecule in its unaltered form. This is typically not possible with small molecule aptamer selections. Thus, the primary complication arises from the need to immobilize the target to a solid support matrix, for example, magnetic beads, acrylic beads, agarose/sepharose, to facilitate the partitioning process. Early small molecule aptamers were selected for targets for which premade agarose material was commercially available [10]. In the absence of commercially available material, there is a wide array of conjugation chemistries that are available for preparing these materials for SELEX experiments. However, these are all dependent on the presence of certain functional groups that allow for coupling, which are not always present on the desired target. For cases where conjugation is possible, the proportionally large amount of column material, in comparison to the target, that is presented to the nucleic acid pool during each SELEX round can result in high nonspecific binding of the library. As chemical modification of the target is required to facilitate column immobilization, the library is exposed to chemically-modified target rather than the desired, unmodified target molecule, increasing the likelihood of selecting sequences that display binding properties towards the matrix and/or the linker arm. Despite negative selection steps, carry-over of such sequences is difficult to avoid [60]. Many aptamer applications, particularly those in vivo, require the selected sequences to bind the target free in solution. Therefore, any aptamer affinity derived from partial binding to the matrix or from chemical modifications will reduce the functionality of the aptamer in the intended applications. For example, the published rhodamine aptamer displays a weaker binding to the target rhodamine when in solution compared to when it is immobilized on the matrix used in the selection [61].
3.2. Measurement of Binding Affinity (K d)
While new methods for the determination of binding affinity are constantly being developed, this is often the limiting factor in the rapid development and testing of aptamers. This is particularly true for small molecule binding aptamers. To measure K d, a constant concentration of either the aptamer or target is titrated with an increasing concentration of the other component to yield a binding isotherm. A list of common methods for determining aptamer binding affinity can be found in the Table 4. A brief evaluation of their applicability to small molecule binding aptamers is also provided below.
Table 4.
Methods for determining aptamer binding affinity.
| Method | Description of method and applicability to small molecules | Sample reference |
|---|---|---|
| Spectroscopy-based methods | ||
|
| ||
| Fluorescence intensity | The fluorescence of the aptamer or target may be quenched or increased upon binding. This method requires a fluorescent small molecule target or requires labelling of the target. | [208] |
| Fluorescence polarization | A fluorophore is excited with polarized light and, due to rotational diffusion, the size of the fluorophore will dictate the proportion of polarized light that is emitted. This method requires a fluorescent small molecule target or target labelling. It can be used with a fluorescently tagged aptamer, however, the method is less sensitive as the overall change in mass upon binding a small molecule will be less dramatic. | [209] |
| UV-vis absorption | This method requires a change in intensity or wavelength of absorption in either the aptamer or target's UV-vis spectrum. In some cases, melting studies can be used to determine K d. | [210] |
| Circular dichroism (CD) | CD refers to the differential absorption of left and right circularly polarized light. Upon aptamer binding to the target, the CD spectra may change but a significant difference in conformation upon target binding is required for this method to have good sensitivity. | [211] |
| Nuclear magnetic spectroscopy (NMR) | By comparing the heteronuclear single quantum coherence spectroscopy (HSQC) of individual amide protons in the free and bound aptamer, it is possible to observe changes in the chemical shifts of the peaks. This method requires conformation changes in the aptamer for good sensitivity. | [212] |
|
| ||
| Mass-sensitive surface-based measurements | ||
|
| ||
| Surface plasmon resonance (SPR) | Either the target or aptamer can be coupled to a chip; by flowing various concentrations of the nontethered ligand, changes in refractive index can be measured as the aptamer-target complex forms. If the small molecule target is immobilized, its ability to bind to the aptamer may be compromised. Immobilization of the aptamer, however, leads to a less sensitive measurement as the smaller target will cause less of a change at the surface. | [33] |
| Quartz crystal microbalance (QCM) | This method uses piezoelectric crystals to correlate the mass accumulated (target binding) on the surface with a decrease of the resonance frequency of the quartz crystal. Once again, small molecule target immobilization could affect binding affinity. Immobilization of the aptamer leads to a less sensitive measurement because there will less of a mass change upon target binding. | [213] |
|
| ||
| Separation-based methods | ||
|
| ||
| High-performance liquid chromatography (HPLC) | Zone separations of the free aptamer, target, and aptamer-target complex can be used to assess the equilibrium distribution of these components. This method is particularly difficult with small molecule targets as they have less of an effect on the separation of aptamer-target complex from the free aptamer. | [214] |
| Capillary electrophoresis (CE), kinetic capillary electrophoresis (KCE), affinity probe capillary electrophoresis (APCE) | This method is similar to HPLC except that it using an electric field to separate the components of the mixture by size and charge. Small molecule targets can be a challenge, typically requiring labeling of the small molecule although label-free KCE UV has recently been described. Once again, separation of the aptamer-target complex from the free aptamer can be more difficult in the case of small molecule targets. | [64, 215, 216] |
| Microfree-flow electrophoresis (μ-FFE) | This technique separates aptamer and aptamer-target complex based on their electrophoretic mobilities. Sample is continuously streamed into a planar flow channel while an electric field is applied perpendicularly to the direction of flow, deflecting analyte streams as they travel through the flow channel according to their mobility. Once again, this method is less effective with small molecule targets. | [217] |
| Equilibrium dialysis | Equilibrium dialysis allows the aptamer, target and the complex to equilibrate in a two compartment cell separated by a semipermeable membrane that allows only the smallest component to pass through. This method can be hampered by nonspecific adsorption of small molecule targets to the membrane. | [26] |
| Ultrafiltration/ nitrocellulose filtration | This method is similar to dialysis. The aptamer and target are incubated to allow binding. The fraction of the smallest unbound component is forced through a filter and measured. Once again, nonspecific adsorption to the membrane can cause this method to be unreliable. | [218] |
| Affinity chromatography | Either the target or aptamer is covalently immobilized to a solid-phase support. The other component is incubated with the support and the amount of binding is calculated. As with other methods, chemical modification of the target or the aptamer to allow for immobilization can affect binding. | [7] |
| Electrophoretic mobility shift assay (EMSA) | The presence of the target will cause an increase in molecular weight of the aptamer-target complex, resulting in a change in electrophoretic mobility and a gel shift. This approach is not effective with small molecule targets unless a significant conformational change is observed upon binding. | [219] |
| Optical thermophoresis | Based on the directed movement of molecules along temperature gradients, the thermophoresis of an aptamer typically differs significantly from that of an aptamer-target complex because of changes in size, charge, or solvation energy. This method requires fluorescent labelling which could affect binding. Also, it could be less sensitive for small molecule aptamers due to the smaller change in mass upon target binding. | [220] |
|
| ||
| Other methods | ||
|
| ||
| Isothermal titration calorimetry (ITC) | This method allows simultaneous determination of K d, stoichiometry, and thermodynamic properties. It relies on the fact that formation of the aptamer-target complex is an exothermic process. Effective for small molecule aptamers, particularly if a large conformational change occurs upon target binding. | [221] |
| High-throughput affinity quantitative PCR binding assay | With this method, an aptamer duplex is incubated with the target. The concentration of aptamer released by this binding event is then measured using real time PCR. | [222] |
| In-line probing | Spontaneous cleavage of the RNA backbone is affected by local structural characteristics, which in turn are impacted by target binding. Can be effective for small molecule aptamers but requires conformational changes upon target binding and is only applicable to RNA aptamers. | [223] |
| Footprinting assays | This method determines the region of aptamer sequence where target binding occurs by exploiting that the target may protect the aptamer from enzymatic cleavage/chemical reactions. Footprinting assays are easier with larger targets or require conformational changes with target binding. | [224, 225] |
As can be seen in Table 4, relatively few of these common K d methods are effective for measuring aptamer binding to small molecules. Separation-based techniques are among the most common approaches for determining binding affinity, and many of these are more challenging for small molecule targets than for proteins. In particular, separation-based methods that rely on a dramatic change in the size of the aptamer-target complex upon target binding are of limited use when the target is much smaller than the aptamer. Other methods require that the target has some intrinsic fluorescence/absorbance, which is often not the case. For targets lacking these properties, an alternative is to label the target, which can affect the chemical properties of the small molecule and interfere with aptamer binding. Surface mass-sensitive detection methods such as QCM and SPR are typically limited to large targets such as proteins [62]. These approaches generally require one binding partner to be tethered to the surface. In cases where the aptamer is surface-bound, the sensitivity of the technique may be compromised by the small overall mass change caused by small molecule binding. As an alternative, the target could be attached to the surface, but once again this chemical modification of the target can negatively impact binding affinity. Other methods that detect a change in aptamer conformation upon binding to target can be applicable to small molecule aptamers, but measurable conformation change is not a universal property of all aptamers [63]. Some recent reports of approaches for determining aptamer binding affinity have recognized the unique challenges for small molecule aptamers and attempted to address them using more innovative approaches such as automated microchip electrophoresis and atomic force spectroscopy, although no technique can be considered generally applicable to small molecule aptamers at this stage [64, 65].
4. Opportunities for Small Molecule Aptamers
While aptamer technology has existed for over two decades, the challenges imposed on the development of aptamers for small molecules has resulted in very few novel aptamers that can bind to practical small molecular targets. Nevertheless, there are many opportunities for the innovative application of small molecule binding aptamers in biosensing and chemical biology. A diverse range of natural and synthetic compounds fall under the designation of small molecules, including organic compounds, amino acids, steroids, carbohydrates, and nucleotides. These molecules play a variety of beneficial roles; they are therapeutics, dyes, cofactors, metabolites, and neurotransmitters. Unfortunately, they may also be harmful substances, such as pollutants, food adulterants, carcinogens, and drugs of abuse. There are many reasons why effective tools for the detection of small molecules are needed now more than ever before. These include the growing recognition of the role of small molecules in biological systems, the extensive application of synthetic small molecules as drugs, and the increasing need to monitor contamination our environment and food supply. Many examples of aptamer-based sensing approaches have been designed for the detection of small molecules, however, most are proof-of-concept systems using the ATP, cocaine, or theophylline aptamer. Electrochemical, fluorescence, colorimetric, and other approaches that have been employed for aptamer-based sensing have been highlighted in several recent reviews [66–69]. However, the general applicability of many of these biosensors to small molecule targets, other than the three main targets mentioned, has yet to be confirmed. As shown in Tables 2 and 3, however, there are many small molecule aptamer sequences that are available for biosensor development. Several of these targets are relevant to applications in environmental monitoring, agriculture, food safety, and medicine [70]; this is a clear motivation to move aptamer-based biosensing past the few proof-of-concept systems and to validate aptamer-based sensing approaches for real-world bioanalytical applications.
One notable approach to small-molecule biosensing has been the “structure-switching” strategy, described by Li and colleagues, which relates to the unique ability of nucleic aptamers to bind to both their cognate target and to a complementary sequence [71]. This ability to switch from a nucleic acid duplex to an aptamer-target complex, and its concomitant structural changes, have been shown to be a generally applicable method for converting a recognition event into detectable signal. As such, it has been applied to several fluorescence, electrochemical, or colorimetric (nanoparticle-based) assays (see Figure 3). This strategy is particularly appealing given that a large conformational change upon target binding is virtually guaranteed, thus many of the typical limitations encountered in small molecule detection are avoided. Both DNA and RNA aptamers can be employed in this approach [72] and it has been used in solution and on surfaces [73]. Although many reports use this methodology for the sensing of adenosine or ATP [71], this method has been applied to detect several other small molecule targets including theophylline [72], cocaine [74], histidine [75], OTA [76], L-argininamide [77], tyrosinamide [78], GTP [79], and arginine [80].
Figure 3.
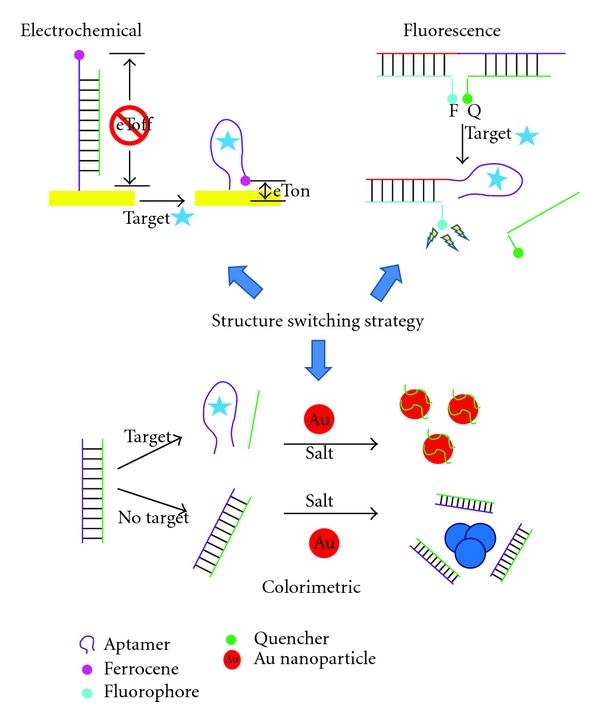
Illustration of sample electrochemical, fluorescence, and colorimetric assays using the structure switching strategy and small molecule-binding aptamers.
Recent reviews have examined how aptamers can be combined with other functional moieties without affecting their ability to recognize and bind to their cognate target [81, 82]. This property can allow aptamers to serve as regulatory elements for nucleic acid enzymes (either natural ribozymes or synthetic DNAzymes) or other actuator parts, allowing control of a variety of functions such as gene-expression regulation [83]. Small molecule-binding sequences are particularly convenient for the preparation of these chimeras. This is perhaps not surprising considering that natural riboswitches, which are known to be key regulators of several biosynthetic pathways, contain an aptamer domain that serves as a high affinity sensor for a specific small molecule. In one of the original examples of these chimeric systems, one of the stem-loop sections of the widely studied hammerhead ribozyme was replaced with the ATP-binding aptamer. As a result, ATP binding was required for activation [84]. A similar approach can be applied to the development of allosteric aptamers for sensing. Conjugation of the malachite green aptamer to the flavin mononucleotide (FMN) aptamer created a FMN sensor where binding of malachite green and the concomitant increase in fluorescence could only be achieved after FMN bound to its aptamer domain [85]. ATP and theophylline sensors were also made using the same approach. This strategy has been suggested for the fluorescence detection of cellular metabolites; a combination of the endogenously expressed aptamer conjugate and cellular dyes could enable intracellular detection. Thus, the continued development of biologically relevant small molecule aptamers for these chimeric systems is important in order to enable further advances in areas such as in vivo imaging and synthetic biology.
Small molecule-binding-aptamers can also serve as important tools for elucidating the role that biologically-important small molecules play in modulating critical cellular regulatory circuits. Intramers, aptamers specific for intracellular target proteins, have already been shown to be useful tools for probing important protein-based networks in vivo [86]. By perturbing the intracellular pools of physiologically important small molecules, small-molecule-binding aptamers also have the potential to improve our understanding of biological systems. This strategy has been explored by Marletta using intracellular expression of a heme-binding RNA aptamer to predictably modulate E. coli heme biosynthesis as a model for a product feedback inhibited system. This group demonstrated that in vitro selected, heme-binding RNA aptamers could specifically sequester intracellular heme when expressed in vivo and perturb the heme-mediated inhibition of the heme biosynthetic pathway in a measurable way [87]. Using heme-binding DNA aptamers [88], Marletta and colleagues also demonstrated that aptamers can successfully control an essential, P. falciparum-specific metabolic pathway [89]. In this study, the pathway of interest was the parasite's heme detoxification pathway, a key target for malaria controls. Hemoglobin ingested by the parasite is degraded in vacuolar structures at low pH (between 4.5 and 5.5), releasing free heme. This free heme is cytotoxic, thus, heme detoxification by polymerization into hemozoin is a critical process for plasmodia survival. The heme-binding DNA aptamers were shown to interfere with hemozoin formation in two in vitro assays, one using a model lipid-catalyzed system and the other using parasite-derived lysates containing the native hemozoin formation components. IC50 values for inhibition by the aptamers were comparable to that of chloroquine, a known inhibitor of hemozoin production. Additionally, when preloaded into red blood cells, nuclease resistant heme-binding aptamers induced parasite toxicity in a manner consistent with inhibiting the hemozoin production process in early stage parasites (see Figure 4). These examples show the potential utility of small molecule-binding aptamers as new chemical tools for probing biophysical processes as well as their potential use as leads in antimalarial drug development.
Figure 4.
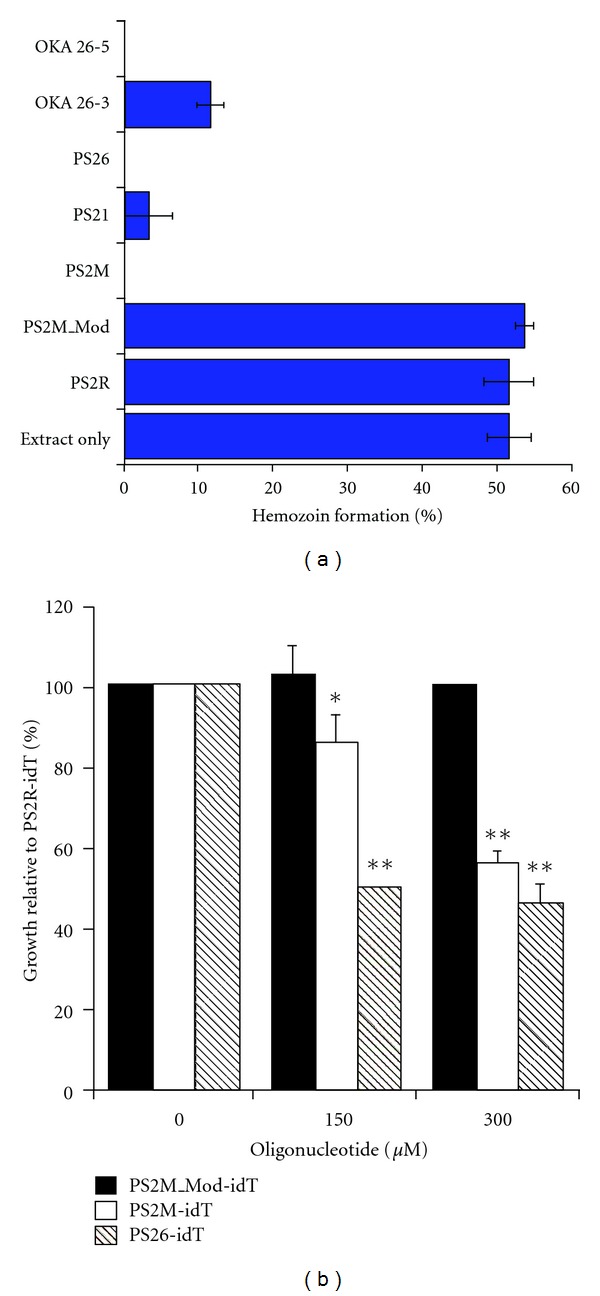
(a) Confirmation of aptamer-mediated inhibition of hemozoin formation within parasite lysates. Heme-binding DNA aptamers (OKA 26-5 and 26-3; PS26, PS21, and PS2M) inhibit hemozoin formation but control oligonucleotides (PS2M_Mod and PS2R) have no effect. (b) The growth of parasites incubated in red blood cells that had been preloaded with nuclease resistant DNA aptamers (PS2M_idT and PS26-idT) is significantly inhibited in comparison to those exposed to red blood cells loaded with control oligonucleotides (PS2M_Mod-idT). Used with permission from PNAS.
Recently, the in vivo utility of a dopamine-binding DNA aptamer as a tool to investigate neurobiological processes was demonstrated in a preclinical animal model of schizophrenia [90]. In a similar fashion as the heme example, the role of the dopamine aptamer was to sequester excess dopamine in a particular brain region (the nucleus accumbens) and to monitor whether an abnormal behavior could be reversed as a result. The drug MK-801, an N-Methyl-D-aspartate (NMDA) receptor antagonist, has been used to model the cognitive dysfunction observed in individuals with schizophrenia. In this schizophrenia model, rats are trained to press a bar for chocolate pellet rewards. After 5 days of training, the rats undergo an “extinction” test, where no chocolate is presented upon lever pressing. While normal rats quickly realize this and stop pressing the lever, rats under the influence of MK-801 show a cognitive defect known as perseveration, meaning that they continued to press the lever at a high rate and are unable to inhibit this behavioral tendency. This defect has been linked to high levels of dopamine in the nucleus accumbens region of the brain. The ability of a dopamine-binding DNA aptamer to reverse these MK-801-induced cognitive deficits when injected directly into the nucleus accumbens was determined. Figure 5(a) shows that injection of the dopamine aptamer (filled triangles) reversed the MK-801-induced elevation in lever pressing to levels as seen in rats not treated with MK-801 (empty squares, X with dashed line). Injection of buffer (empty diamonds) or of a random oligonucleotide (dashed line) had no effect on moderating the pressing behavior. Thus, it appears that the aptamer was successful in sequestering the excess dopamine within the nucleus accumbens, resulting in “normal” behavior. Interestingly, it was also shown that the aptamer treatment did not impair locomotor activity in the animals (Figure 5(b)). The near-selective effect of the aptamer on reversing cognitive deficits without drastic negative motor consequences lends support for the use of DNA aptamers in the further study of preclinical animal models of mental health disease and as possible drug leads. New neurotransmitter-binding aptamers, as well as strategies for the delivery of aptamers across the blood-brain-barrier, will be required to realize the full potential of aptamers in this regard.
Figure 5.
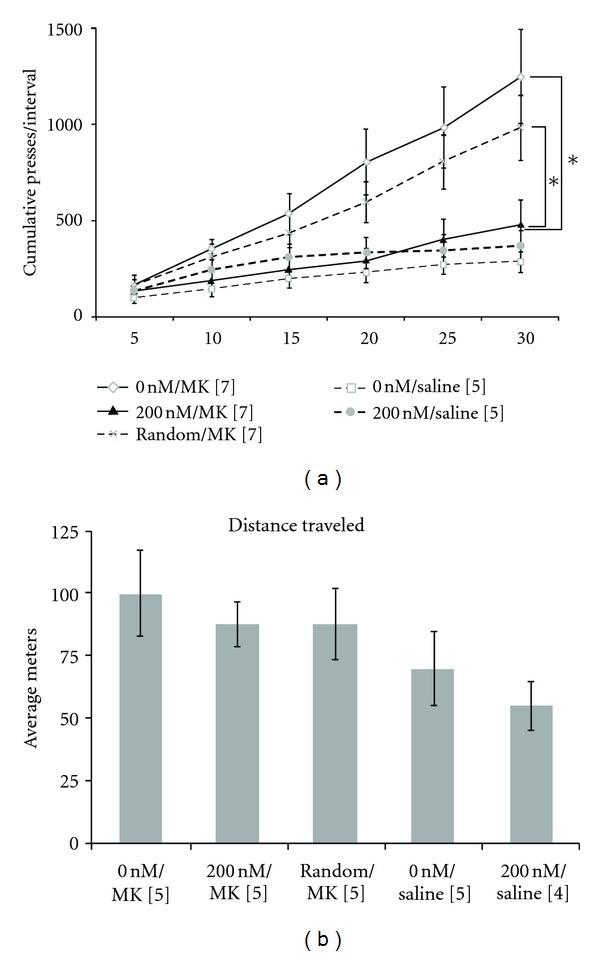
(a) Pretreatment of animals with the dopamine aptamer reversed the effects of MK-801 administration. Animals given MK-801 (empty diamonds; 0 nM/MK) show higher cumulative presses in this behavioural test in comparison to animals not given this drug (empty squares, dashed line; 0 nM/Saline). The group receiving aptamer pretreatment (filled triangles; 200 nM/MK), however, showed similar levels of cumulative presses as those that were not given any MK-801. A random oligonucleotide pretreatment, however, had no dampening effect on the number of presses (X with dashed line; Random/MK). (b) Aptamer pretreatment (200 nM/MK) did not significantly affect locomotor activity as measured by distance traveled in an elevated cross maze. Used with permission from PLoS One.
5. Conclusions and Outlook
The selection and application of small molecule-binding aptamers come with a unique set of challenges that have hampered their research and commercialization. Nevertheless, efforts to expand the suite of aptamers for pertinent small molecules need to continue. A major bottleneck in small molecule aptamer development and application occurs at the point of K d determination. Approaches for binding affinity measurement that are more generally applicable to small molecule targets are required. Additionally, there is a negative perception that there are published aptamers that have little affinity to their targets [91]. Because of these issues associated with K d measurements for small molecules, several complementary methods may be required in order to achieve a true sense of the aptamer binding affinity for its target. Ideally, these methods should allow aptamer-target binding to occur free in solution, to remove any contribution from matrix binding. A new aptamer database is available (http://aptamerbase.semanticscience.org/) containing valuable information about the experimental conditions under which the aptamers were selected and their binding affinity quantified [41]. Efforts to mine this data to better tailor SELEX and binding affinity experiments for small molecules is currently underway. Aptamer-based assays also need to begin to move away from the proof-of-concept targets and exploit the largely untapped resource of existing small molecule-binding aptamers. Signs of this shift to more relevant aptamer-based assays can be seen with the large number of biosensors developed using the ochratoxin A (OTA) aptamer. This small molecule mycotoxin contaminates a wide variety of food commodities such as cereals and wine. Several recent reports have used OTA-binding sequences [26] to develop affinity clean-up columns [92, 93] and biosensors [94–98] for evaluation under actual food testing conditions. Applications of small molecule-binding aptamers in other burgeoning areas, such as metabolomics, drug discovery, and synthetic biology, could also soon see dramatic growth. Continued effort in the development of aptamers for important small molecules is required in order for this field to realize its full potential.
Acknowledgments
The authors thank Alexander Wahba for assistance and the Natural Sciences and Engineering Council of Canada (NSERC), the Canadian Foundation for Innovation (CFI), the Ontario Research Fund (ORF), and an Ontario Early Researcher Award for funding.
References
- 1.Deigan KE, Ferré-D'Amaré AR. Riboswitches: discovery of drugs that target bacterial gene-regulatory RNAs. Accounts of Chemical Research. 2011;44(12):1329–1338. doi: 10.1021/ar200039b. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Weigand JE, Suess B. Aptamers and riboswitches: perspectives in biotechnology. Applied Microbiology and Biotechnology. 2009;85(2):229–236. doi: 10.1007/s00253-009-2194-2. [DOI] [PubMed] [Google Scholar]
- 3.Tuerk C, Gold L. Systemic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science. 1990;249(4968):505–510. doi: 10.1126/science.2200121. [DOI] [PubMed] [Google Scholar]
- 4.Ellington AD, Szostak JW. In vitro selection of RNA molecules that bind specific ligands. Nature. 1990;346(6287):818–822. doi: 10.1038/346818a0. [DOI] [PubMed] [Google Scholar]
- 5.Robertson DL, Joyce GF. Selection in vitro of an RNA enzyme that specifically cleaves single-stranded DNA. Nature. 1990;344(6265):467–468. doi: 10.1038/344467a0. [DOI] [PubMed] [Google Scholar]
- 6.Mascini M. Aptamers in Bioanalysis. Hoboken, NJ, USA: John Wiley & Sons; 2009. [Google Scholar]
- 7.McKeague M, Bradley CR, De Girolamo A, Visconti A, David Miller J, DeRosa MC. Screening and initial binding assessment of fumonisin B1 aptamers. International Journal of Molecular Sciences. 2010;11(12):4864–4881. doi: 10.3390/ijms11124864. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Jayasena SD. Aptamers: an emerging class of molecules that rival antibodies in diagnostics. Clinical Chemistry. 1999;45(9):1628–1650. [PubMed] [Google Scholar]
- 9.Nimjee SM, Rusconi CP, Sullenger BA. Aptamers: an emerging class of therapeutics. Annual Review of Medicine. 2005;56:555–583. doi: 10.1146/annurev.med.56.062904.144915. [DOI] [PubMed] [Google Scholar]
- 10.Jhaveri S, Ellington A. In vitro selection of RNA aptamers to a small molecule target. Current Protocols in Nucleic Acid Chemistry. 2002;(Chapter 9, Unit 9.5) doi: 10.1002/0471142700.nc0905s08. [DOI] [PubMed] [Google Scholar]
- 11.Bardoczy V, Meszaros T. Aptamer selection for macromolecular (Protein) and for small molecule targets. Proceedings of the Periodica Polytechnica Abstracts of PhD Conference; 2006. [Google Scholar]
- 12.Balogh Z, Lautner G, Bardóczy V, Komorowska B, Gyurcsányi RE, Mészáros T. Selection and versatile application of virus-specific aptamers. FASEB Journal. 2010;24(11):4187–4195. doi: 10.1096/fj.09-144246. [DOI] [PubMed] [Google Scholar]
- 13.Sefah K, Shangguan D, Xiong X, O’Donoghue MB, Tan W. Development of DNA aptamers using Cell-SELEX. Nature protocols. 2010;5(6):1169–1185. doi: 10.1038/nprot.2010.66. [DOI] [PubMed] [Google Scholar]
- 14.Beaucage SL, Caruthers MH. Deoxynucleoside phosphoramidites-A new class of key intermediates for deoxypolynucleotide synthesis. Tetrahedron Letters. 1981;22(20):1859–1862. [Google Scholar]
- 15.Beaucage SL, Iyer RP. Advances in the synthesis of oligonucleotides by the phosphoramidite approach. Tetrahedron. 1992;48(12):2223–2311. [Google Scholar]
- 16.Keefe AD, Pai S, Ellington A. Aptamers as therapeutics. Nature Reviews Drug Discovery. 2010;9(7):537–550. doi: 10.1038/nrd3141. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Beaucage SL, Iyer RP. The synthesis of modified oligonucleotides by the phosporamidite approach and their applications. Tetrahedron. 1993;49(28):6123–6194. [Google Scholar]
- 18.Beaucage SL, Iyer RP. The functionalization of oligonucleotides via phosphoramidite derivatives. Tetrahedron. 1993;49(10):1925–1963. [Google Scholar]
- 19.Bui BTS, Haupt K. Molecularly imprinted polymers: synthetic receptors in bioanalysis. Analytical and Bioanalytical Chemistry. 2010;398(6):2481–2492. doi: 10.1007/s00216-010-4158-x. [DOI] [PubMed] [Google Scholar]
- 20.Xu ZX, Gao HJ, Zhang LM, Chen XQ, Qiao XG. The biomimetic immunoassay based on molecularly imprinted polymer: a comprehensive review of recent progress and future prospects. Journal of Food Science. 2011;76(2):R69–R75. doi: 10.1111/j.1750-3841.2010.02020.x. [DOI] [PubMed] [Google Scholar]
- 21.Mills DR, Peterson RL, Spiegelman S. An extracellular Darwinian experiment with a self-duplicating nucleic acid molecule. Proceedings of the National Academy of Sciences of the United States of America. 1967;58(1):217–224. doi: 10.1073/pnas.58.1.217. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Saffhill R, Schneider-Bernloehr H, Orgel LE, Spiegelman S. In vitro selection of bacteriophage Qβ ribonucleic acid variants resistant to ethidium bromide. Journal of Molecular Biology. 1970;51(3):531–539. doi: 10.1016/0022-2836(70)90006-9. [DOI] [PubMed] [Google Scholar]
- 23.Wilson DS, Szostak JW. In vitro selection of functional nucleic acids. Annual Review of Biochemistry. 1999;68:611–647. doi: 10.1146/annurev.biochem.68.1.611. [DOI] [PubMed] [Google Scholar]
- 24.Silverman S, K S. Artificial functional nucleic acids: aptamers, ribozymes, and deoxyribozymes identified by in vitro selection. Functional Nucleic Acids For Analytical Applications. 2009;1:47–108. [Google Scholar]
- 25.Vant-Hull B, Gold L, Zichi DA. Theoretical principles of in vitro selection using combinatorial nucleic acid libraries. Current Protocols in Nucleic acid Chemistry. 2000;(Chapter 9, Unit 9.1) doi: 10.1002/0471142700.nc0901s00. [DOI] [PubMed] [Google Scholar]
- 26.Cruz-Aguado JA, Penner G. Determination of ochratoxin A with a DNA aptamer. Journal of Agricultural and Food Chemistry. 2008;56(22):10456–10461. doi: 10.1021/jf801957h. [DOI] [PubMed] [Google Scholar]
- 27.Svobodova M, Pinto A, Nadal P, OSullivan CK. Comparison of different methods for generation of single-stranded DNA for SELEX processes. Analytical and Bioanalytical Chemistry. 2012;404(3):835–842. doi: 10.1007/s00216-012-6183-4. [DOI] [PubMed] [Google Scholar]
- 28.Gold L, Ayers D, Bertino J, et al. Aptamer-based multiplexed proteomic technology for biomarker discovery. PLoS One. 2010;5(12) doi: 10.1371/journal.pone.0015004.15004 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Stoltenburg R, Reinemann C, Strehlitz B. SELEX-A (r)evolutionary method to generate high-affinity nucleic acid ligands. Biomolecular Engineering. 2007;24(4):381–403. doi: 10.1016/j.bioeng.2007.06.001. [DOI] [PubMed] [Google Scholar]
- 30.Davis JH, Szostak JW. Isolation of high-affinity GTP aptamers from partially structured RNA libraries. Proceedings of the National Academy of Sciences of the United States of America. 2002;99(18):11616–11621. doi: 10.1073/pnas.182095699. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Luo X, Mckeague M, Pitre S, et al. Computational approaches toward the design of pools for the in vitro selection of complex aptamers. RNA. 2010;16(11):2252–2262. doi: 10.1261/rna.2102210. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Ruff KM, Snyder TM, Liu DR. Enhanced functional potential of nucleic acid aptamer libraries patterned to increase secondary structure. Journal of the American Chemical Society. 2010;132(27):9453–9464. doi: 10.1021/ja103023m. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Win MN, Klein JS, Smolke CD. Codeine-binding RNA aptamers and rapid determination of their binding constants using a direct coupling surface plasmon resonance assay. Nucleic Acids Research. 2006;34(19):5670–5682. doi: 10.1093/nar/gkl718. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Cox JC, Rudolph P, Ellington AD. Automated RNA selection. Biotechnology Progress. 1998;14(6):845–850. doi: 10.1021/bp980097h. [DOI] [PubMed] [Google Scholar]
- 35.Goertz PW, Cox JC, Ellington AD. Automated selection of aminoglycoside aptamers. Journal of the Association for Laboratory Automation. 2004;9(3):150–154. [Google Scholar]
- 36.Wochner A, Cech B, Menger M, Erdmann VA, Glökler J. Semi-automated selection of DNA aptamers using magnetic particle handling. BioTechniques. 2007;43(3):344–353. doi: 10.2144/000112532. [DOI] [PubMed] [Google Scholar]
- 37.Cho MJ, Juliano R. Macromolecular versus small-molecule therapeutics: drug discovery, development and clinical considerations. Trends in Biotechnology. 1996;14(5):153–158. doi: 10.1016/0167-7799(96)10024-X. [DOI] [PubMed] [Google Scholar]
- 38.Ashour ML, Wink M. Genus Bupleurum: a review of its phytochemistry, pharmacology and modes of action. Journal of Pharmacy and Pharmacology. 2011;63(3):305–321. doi: 10.1111/j.2042-7158.2010.01170.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Roemer T, Davies J, Giaever G, Nislow C. Bugs, drugs and chemical genomics. Nature Chemical Biology. 2012;8(1):46–56. doi: 10.1038/nchembio.744. [DOI] [PubMed] [Google Scholar]
- 40.Walsh TA. The emerging field of chemical genetics: potential applications for pesticide discovery. Pest Management Science. 2007;63(12):1165–1171. doi: 10.1002/ps.1452. [DOI] [PubMed] [Google Scholar]
- 41.Cruz-Toledo J, McKeague M, Zhang X, et al. Aptamer base: a collaborative knowledge base to describe aptamers and SELEX experiments. Database. 2012;2012 doi: 10.1093/database/bas006.bas006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Mascini M, Palchetti I, Tombelli S. Nucleic acid and peptide aptamers: fundamentals and bioanalytical aspects. Angewandte Chemie. 2012;51:1316–1332. doi: 10.1002/anie.201006630. [DOI] [PubMed] [Google Scholar]
- 43.Michael F. Oligonucleotide aptamers that recognize small molecules . Current Opinion in Structural Biology. 1999;9:324–329. doi: 10.1016/S0959-440X(99)80043-8. [DOI] [PubMed] [Google Scholar]
- 44.Jenison RD, Gill SC, Pardi A, Polisky B. High-resolution molecular discrimination by RNA. Science. 1994;263(5152):1425–1429. doi: 10.1126/science.7510417. [DOI] [PubMed] [Google Scholar]
- 45.Michaud M, Jourdan E, Villet A, Ravel A, Grosset C, Peyrin E. A DNA aptamer as a new target-specific chiral selector for HPLC. Journal of the American Chemical Society. 2003;125(28):8672–8679. doi: 10.1021/ja034483t. [DOI] [PubMed] [Google Scholar]
- 46.Famulok M, Szostak JW. Stereospecific recognition of tryptophan agarose by in vitro selected RNA. Journal of the American Chemical Society. 1992;114:3990–3991. [Google Scholar]
- 47.Famulok M. Molecular recognition of amino acids by RNA-aptamers: an L-citrulline binding RNA motif and its evolution into an L-arginine binder. Journal of the American Chemical Society. 1994;116(5):1698–1706. [Google Scholar]
- 48.Geiger A, Burgstaller P, Von der Eltz H, Roeder A, Famulok M. RNA aptamers that bind L-arginine with sub-micromolar dissociation constants and high enantioselectivity. Nucleic Acids Research. 1996;24(6):1029–1036. doi: 10.1093/nar/24.6.1029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Shoji A, Kuwahara M, Ozaki H, Sawai H. Modified DNA aptamer that binds the (R)-isomer of a thalidomide derivative with high enantioselectivity. Journal of the American Chemical Society. 2007;129(5):1456–1464. doi: 10.1021/ja067098n. [DOI] [PubMed] [Google Scholar]
- 50.Kim YS, Hyun CJ, Kim IA, Gu MB. Isolation and characterization of enantioselective DNA aptamers for ibuprofen. Bioorganic and Medicinal Chemistry. 2010;18(10):3467–3473. doi: 10.1016/j.bmc.2010.03.074. [DOI] [PubMed] [Google Scholar]
- 51.Carothers JM, Goler JA, Kapoor Y, Lara L, Keasling JD. Selecting RNA aptamers for synthetic biology: investigating magnesium dependence and predicting binding affinity. Nucleic Acids Research. 2010;38(8):2736–2747. doi: 10.1093/nar/gkq082.gkq082 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Pfeffer P, Gohlke H. DrugScoreRNA—knowledge-based scoring function to predict RNA—Ligand interactions. Journal of Chemical Information and Modeling. 2007;47(5):1868–1876. doi: 10.1021/ci700134p. [DOI] [PubMed] [Google Scholar]
- 53.Jo M, Ahn JY, Lee J, et al. Development of single-stranded DNA aptamers for specific bisphenol a detection. Oligonucleotides. 2011;21(2):85–91. doi: 10.1089/oli.2010.0267. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Niazi JH, Lee SJ, Kim YS, Gu MB. Bioorg. Med. Chem. 2008;16:1254–1261. doi: 10.1016/j.bmc.2007.10.073. [DOI] [PubMed] [Google Scholar]
- 55.Mandal M, Boese B, Barrick JE, Winkler WC, Breaker RR. Riboswitches control fundamental biochemical pathways in Bacillus subtilis and other bacteria. Cell. 2003;113(5):577–586. doi: 10.1016/s0092-8674(03)00391-x. [DOI] [PubMed] [Google Scholar]
- 56.Welz R, Breaker RR. Ligand binding and gene control characteristics of tandem riboswitches in Bacillus anthracis. RNA. 2007;13(4):573–582. doi: 10.1261/rna.407707. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Kwon M, Strobel SA. Chemical basis of glycine riboswitch cooperativity. RNA. 2008;14(1):25–34. doi: 10.1261/rna.771608. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Missailidis S, Hardy A. Aptamers as inhibitors of target proteins. Expert Opinion on Therapeutic Patents. 2009;19(8):1073–1082. doi: 10.1517/13543770903042337. [DOI] [PubMed] [Google Scholar]
- 59.Mayer G, Ahmed MSL, Dolf A, Endl E, Knolle PA, Famulok M. Fluorescence-activated cell sorting for aptamer SELEX with cell mixtures. Nature Protocols. 2010;5(12):1993–2004. doi: 10.1038/nprot.2010.163. [DOI] [PubMed] [Google Scholar]
- 60.Schneider DJ, Vanderslice R, Gold L. Flow cell SELEX. US Patent 5,861,254, 1999.
- 61.Wilson C, Szostak JW. Isolation of a fluorophore-specific DNA aptamer with weak redox activity. Chemistry and Biology. 1998;5(11):609–617. doi: 10.1016/s1074-5521(98)90289-7. [DOI] [PubMed] [Google Scholar]
- 62.Yao C, Qi Y, Zhao Y, Xiang Y, Chen Q, Fu W. Aptamer-based piezoelectric quartz crystal microbalance biosensor array for the quantification of IgE. Biosensors and Bioelectronics. 2009;24(8):2499–2503. doi: 10.1016/j.bios.2008.12.036. [DOI] [PubMed] [Google Scholar]
- 63.White RJ, Rowe AA, Plaxco KW. Re-engineering aptamers to support reagentless, self-reporting electrochemical sensors. Analyst. 2010;135(3):589–594. doi: 10.1039/b921253a. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Hu J, Easley C. A simple and rapid approach for measurement of dissociation constants of DNA aptamers against proteins and small molecules via automated microchip electrophoresis. Analyst. 2011;136(17):3461–3468. doi: 10.1039/c0an00842g. [DOI] [PubMed] [Google Scholar]
- 65.Nguyen TH, Steinbock LJ, Butt HJ, Helm M, Berger R. Measuring single small molecule binding via rupture forces of a split aptamer. Journal of the American Chemical Society. 2011;133(7):2025–2027. doi: 10.1021/ja1092002. [DOI] [PubMed] [Google Scholar]
- 66.Lau PS, Li Y. Functional nucleic acids as molecular recognition elements for small organic and biological molecules. Current Organic Chemistr. 2011;15(4):557–575. [Google Scholar]
- 67.Cheng AKH, Sen D, Yu HZ. Design and testing of aptamer-based electrochemical biosensors for proteins and small molecules. Bioelectrochemistry. 2009;77(1):1–12. doi: 10.1016/j.bioelechem.2009.04.007. [DOI] [PubMed] [Google Scholar]
- 68.Wang RE, Zhang Y, Cai J, Cai W, Gao T. Aptamer-based fluorescent biosensors. Current Medicinal Chemistry. 2011;18:4175–4184. doi: 10.2174/092986711797189637. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Cho EJ, Lee JW, Ellington AD. Applications of aptamers as sensors. Annual Review of Analytical Chemistry . 2009:241–264. doi: 10.1146/annurev.anchem.1.031207.112851. [DOI] [PubMed] [Google Scholar]
- 70.McKeague M, Giamberardino A, DeRosa MC. Advances in aptamer-based biosensors for food safety. In: Somerset V, editor. Environmental Biosensors. InTech; 2011. pp. 17–42. [Google Scholar]
- 71.Nutiu R, Li Y. Structure-switching signaling aptamers. Journal of the American Chemical Society. 2003;125(16):4771–4778. doi: 10.1021/ja028962o. [DOI] [PubMed] [Google Scholar]
- 72.Lau PS, Coombes BK, Li Y. A General approach to the construction of structure-switching reporters from RNA aptamers. Angewandte Chemie International. 2010;49:7938–7942. doi: 10.1002/anie.201002621. [DOI] [PubMed] [Google Scholar]
- 73.Carrasquilla C, Lau PS, Li Y, Brennan JD. Stabilizing structure-switching signaling RNA aptamers by entrapment in sol-gel derived materials for solid-phase assay. Journal of the American Chemical Society. 2012;134:10998–11005. doi: 10.1021/ja304064a. [DOI] [PubMed] [Google Scholar]
- 74.Zheng D, Zou R, Lou X. free fluorescent detection of ions, proteins, and small molecules using structure-switching aptamers, SYBR gold, and exonuclease. Analytical Chemistry. 2012;84:3554–3560. doi: 10.1021/ac300690r. [DOI] [PubMed] [Google Scholar]
- 75.Liang J, Chen Z, Guo L, Li L. Electrochemical sensing of L-histidine based on structure-switching DNAzymes and gold nanoparticle-graphene nanosheet composites . Chemical Communications . 2011;47:5476–5478. doi: 10.1039/c1cc10965k. [DOI] [PubMed] [Google Scholar]
- 76.Chen J, Fang Z, Liu J, Zeng L. A simple and rapid biosensor for ochratoxin A based on a structure-switching signaling aptamer. Food Control. 2012;25:555–560. [Google Scholar]
- 77.Hun X, Wang Z. L-Argininamide biosensor based on S1 nuclease hydrolysis signal amplification. Microchimica Acta. 2012;176:209–216. [Google Scholar]
- 78.Zhu Z, Schmidt T, Mahrous M, et al. Optimization of the structure-switching aptamer-based fluorescence polarization assay for the sensitive tyrosinamide sensing. Analytica Chimica Acta. 2011;707:191–196. doi: 10.1016/j.aca.2011.09.022. [DOI] [PubMed] [Google Scholar]
- 79.Nutiu R, Li Y. In vitro selection of structure-switching signaling aptamers. Angewandte Chemie . 2005;44:1061–1065. doi: 10.1002/anie.200461848. [DOI] [PubMed] [Google Scholar]
- 80.Null EL, Lu Y. Rapid determination of enantiomeric ratio using fluorescent DNA or RNA aptamers. Analyst. 2010;135(2):419–422. doi: 10.1039/b921267a. [DOI] [PubMed] [Google Scholar]
- 81.Famulok M, Hartig JS, Mayer G. Functional aptamers and aptazymes in biotechnology, diagnostics, and therapy. Chemical Reviews. 2007;107(9):3715–3743. doi: 10.1021/cr0306743. [DOI] [PubMed] [Google Scholar]
- 82.Vinkenborg JL, Karnowski N, Famulok M. Aptamers for allosteric regulation. Nature Chemical Biology. 2011;7(8):519–527. doi: 10.1038/nchembio.609. [DOI] [PubMed] [Google Scholar]
- 83.Win MN, Liang JC, Smolke CD. Frameworks for Programming Biological Function through RNA Parts and Devices. Chemistry and Biology. 2009;16(3):298–310. doi: 10.1016/j.chembiol.2009.02.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Tang J, Breaker RR. Rational design of allosteric ribozymes. Chemistry and Biology. 1997;4(6):453–459. doi: 10.1016/s1074-5521(97)90197-6. [DOI] [PubMed] [Google Scholar]
- 85.Stojanovic MN, Kolpashchikov DM. Modular aptameric sensors. Journal of the American Chemical Society. 2004;126(30):9266–9270. doi: 10.1021/ja032013t. [DOI] [PubMed] [Google Scholar]
- 86.Famulok M, Blind M, Mayer G. Intramers as promising new tools in functional proteomics. Chemistry and Biology. 2001;8(10):931–939. doi: 10.1016/s1074-5521(01)00070-9. [DOI] [PubMed] [Google Scholar]
- 87.Niles JC, Marletta MA. Utilizing RNA aptamers to probe a physiologically important heme-regulated cellular network. ACS Chemical Biology. 2006;1(8):515–524. doi: 10.1021/cb6002527. [DOI] [PubMed] [Google Scholar]
- 88.Li Y, Geyer CR, Sen D. Recognition of anionic porphyrins by DNA aptamers. Biochemistry. 1996;35(21):6911–6922. doi: 10.1021/bi960038h. [DOI] [PubMed] [Google Scholar]
- 89.Niles JC, DeRisi JL, Marletta MA. Inhibiting Plasmodium falciparum growth and heme detoxification pathway using heme-binding DNA aptamers. Proceedings of the National Academy of Sciences of the United States of America. 2009;106(32):13266–13271. doi: 10.1073/pnas.0906370106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Holahan MR, Madularu D, McConnell EM, Walsh R, DeRosa MC. Intra-accumbens injection of a dopamine aptamer abates MK-801-induced cognitive dysfunction in a model of schizophrenia. PLoS One. 2011;6(7) doi: 10.1371/journal.pone.0022239.e22239 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Penner G. IVD Technology, 2012.
- 92.De Girolamo A, McKeague M, Miller JD, DeRosa MC, Visconti A. Determination of ochratoxin A in wheat after clean-up through a DNA aptamer-based solid phase extraction column. Food Chemistry. 2011;127(3):1378–1384. doi: 10.1016/j.foodchem.2011.01.107. [DOI] [PubMed] [Google Scholar]
- 93.De Girolamo A, Le L, Penner G, Schena R, Visconti A. Analytical performances of a DNA-ligand system using time-resolved fluorescence for the determination of ochratoxin A in wheat. Analytical and Bioanalytical Chemistry. 2012;403:2627–2634. doi: 10.1007/s00216-012-6076-6. [DOI] [PubMed] [Google Scholar]
- 94.Yang C, Lates V, Prieto-Simón B, Marty J, Yang X. Aptamer-DNAzyme hairpins for biosensing of Ochratoxin A . Biosensors and Bioelectronics. 2012;32:208–212. doi: 10.1016/j.bios.2011.12.011. [DOI] [PubMed] [Google Scholar]
- 95.Bonel L, Vidal JC, Duato P, Castillo JR. An electrochemical competitive biosensor for ochratoxin A based on a DNA biotinylated aptamer. Biosensors and Bioelectronics. 2011;26(7):3254–3259. doi: 10.1016/j.bios.2010.12.036. [DOI] [PubMed] [Google Scholar]
- 96.Guo Z, Ren J, Wang J, Wang E. Single-walled carbon nanotubes based quenching of free FAM-aptamer for selective determination of ochratoxin A. Talanta. 2011;85(5):2517–2521. doi: 10.1016/j.talanta.2011.08.015. [DOI] [PubMed] [Google Scholar]
- 97.Wang L, Ma W, Chen W, et al. An aptamer-based chromatographic strip assay for sensitive toxin semi-quantitative detection. Biosensors and Bioelectronics. 2011;26(6):3059–3062. doi: 10.1016/j.bios.2010.11.040. [DOI] [PubMed] [Google Scholar]
- 98.Kuang H, Chen W, Xu D, et al. Fabricated aptamer-based electrochemical ‘signal-off’ sensor of ochratoxin A. Biosensors and Bioelectronics. 2010;26(2):710–716. doi: 10.1016/j.bios.2010.06.058. [DOI] [PubMed] [Google Scholar]
- 99.Miyachi Y, Shimizu N, Ogino C, Kondo A. Selection of DNA aptamers using atomic force microscopy. Nucleic Acids Research. 2010;38(4, article e21) doi: 10.1093/nar/gkp1101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Smith D, Kirschenheuter GP, Charlton J, Guidot DM, Repine JE. In vitro selection of RNA-based irreversible inhibitors of human neutrophil elastase. Chemistry and Biology. 1995;2(11):741–750. doi: 10.1016/1074-5521(95)90102-7. [DOI] [PubMed] [Google Scholar]
- 101.Kim Y, Liu C, Tan W. Aptamers generated by Cell SELEX for biomarker discovery. Biomarkers in Medicine. 2009;3(2):193–202. doi: 10.2217/bmm.09.5. [DOI] [PubMed] [Google Scholar]
- 102.Mendonsa SD, Bowser MT. In vitro selection of high-affinity DNA ligands for human IgE using capillary electrophoresis. Analytical Chemistry. 2004;76(18):5387–5392. doi: 10.1021/ac049857v. [DOI] [PubMed] [Google Scholar]
- 103.Burke DH, Willis JH. Recombination, RNA evolution, and bifunctional RNA molecules isolated through chimeric SELEX. RNA. 1998;4(9):1165–1175. doi: 10.1017/s1355838298980542. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Smith JD, Gold L. Conditional-selex. US Patent 6706482, 2004.
- 105.Jensen KB, Atkinson BL, Willis MC, Koch TH, Gold L. Using in vitro selection to direct the covalent attachment of human immunodeficiency virus type 1 Rev protein to high-affinity RNA ligands. Proceedings of the National Academy of Sciences of the United States of America. 1995;92(26):12220–12224. doi: 10.1073/pnas.92.26.12220. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106.Morris KN, Jensen KB, Julin CM, Weil M, Gold L. High affinity ligands from in vitro selection: complex targets. Proceedings of the National Academy of Sciences of the United States of America. 1998;95(6):2902–2907. doi: 10.1073/pnas.95.6.2902. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Tsai RYL, Reed RR. Identification of DNA recognition sequences and protein interaction domains of the multiple-Zn-finger protein Roaz. Molecular and Cellular Biology. 1998;18(11):6447–6456. doi: 10.1128/mcb.18.11.6447. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108.Martell RE, Nevins JR, Sullenger BA. Optimizing aptamer activity for gene therapy applications using expression cassette SELEX. Molecular Therapy. 2002;6(1):30–34. doi: 10.1006/mthe.2002.0624. [DOI] [PubMed] [Google Scholar]
- 109.Stoltenburg R, Reinemann C, Strehlitz B. FluMag-SELEX as an advantageous method for DNA aptamer selection. Analytical and Bioanalytical Chemistry. 2005;383(1):83–91. doi: 10.1007/s00216-005-3388-9. [DOI] [PubMed] [Google Scholar]
- 110.Dobbelstein M, Shenk T. In vitro selection of RNA ligands for the ribosomal L22 protein associated with Epstein-Barr virus-expressed RNA by using randomized and cDNA-derived RNA libraries. Journal of Virology. 1995;69(12):8027–8034. doi: 10.1128/jvi.69.12.8027-8034.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Coulter LR, Landree MA, Cooper TA. Identification of a new class of exonic splicing enhancers by in vivo selection. Molecular and Cellular Biology. 1997;17(4):2143–2150. doi: 10.1128/mcb.17.4.2143. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112.Kawakami J, Imanaka H, Yokota Y, Sugimoto N. In vitro selection of aptamers that act with Zn2+ . Journal of Inorganic Biochemistry. 2000;82(1–4):197–206. doi: 10.1016/s0162-0134(00)00158-6. [DOI] [PubMed] [Google Scholar]
- 113.Keefe AD, Cload ST. SELEX with modified nucleotides. Current Opinion in Chemical Biology. 2008;12(4):448–456. doi: 10.1016/j.cbpa.2008.06.028. [DOI] [PubMed] [Google Scholar]
- 114.Gong Q, Wang J, Ahmad KM, et al. Selection strategy to generate aptamer pairs that bind to distinct sites on protein targets. Analytical Chemistry. 2012;84(12):5365–5371. doi: 10.1021/ac300873p. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115.Huang CJ, Lin HI, Shiesh SC, Lee GB. Integrated microfluidic system for rapid screening of CRP aptamers utilizing systematic evolution of ligands by exponential enrichment (SELEX) Biosensors and Bioelectronics. 2010;25(7):1761–1766. doi: 10.1016/j.bios.2009.12.029. [DOI] [PubMed] [Google Scholar]
- 116.Lou X, Qian J, Xiao Y, et al. Micromagnetic selection of aptamers in microfluidic channels. Proceedings of the National Academy of Sciences of the United States of America. 2009;106(9):2989–2994. doi: 10.1073/pnas.0813135106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 117.Nitsche A, Kurth A, Dunkhorst A, et al. One-step selection of Vaccinia virus-binding DNA aptamers by MonoLEX. BMC Biotechnology. 2007;7(article no. 48) doi: 10.1186/1472-6750-7-48. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 118.Jolma A, Kivioja T, Toivonen J, et al. Multiplexed massively parallel SELEX for characterization of human transcription factor binding specificities. Genome Research. 2010;20(6):861–873. doi: 10.1101/gr.100552.109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.Wu L, Curran JF. An allosteric synthetic DNA. Nucleic Acids Research. 1999;27(6):1512–1516. doi: 10.1093/nar/27.6.1512. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 120.Reid DC, Chang BL, Gunderson SI, Alpert L, Thompson WA, Fairbrother WG. Next-generation SELEX identifies sequence and structural determinants of splicing factor binding in human pre-mRNA sequence. RNA. 2009;15(12):2385–2397. doi: 10.1261/rna.1821809. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121.Berezovski M, Musheev M, Drabovich A, Krylov SN. Non-SELEX selection of aptamers. Journal of the American Chemical Society. 2006;128(5):1410–1411. doi: 10.1021/ja056943j. [DOI] [PubMed] [Google Scholar]
- 122.Brody EN, Willis MC, Smith JD, Jayasena S, Zichi D, Gold L. The use of aptamers in large arrays for molecular diagnostics. Molecular Diagnosis. 1999;4(4):381–388. doi: 10.1016/s1084-8592(99)80014-9. [DOI] [PubMed] [Google Scholar]
- 123.Wen JD, Gray DM. Selection of genomic sequences that bind tightly to Ff gene 5 protein: primer-free genomic SELEX. Nucleic acids research. 2004;32(22, article e182) doi: 10.1093/nar/gnh179. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124.Roulet E, Busso S, Camargo AA, Simpson AJG, Mermod N, Bucher P. High-throughput SELEX-SAGE method for quantitative modeling of transcription-factor binding sites. Nature Biotechnology. 2002;20(8):831–835. doi: 10.1038/nbt718. [DOI] [PubMed] [Google Scholar]
- 125.Klußmann S, Nolte A, Bald R, Erdmann VA, Fürste JP. Mirror-image RNA that binds D-adenosine. Nature Biotechnology. 1996;14(9):1112–1115. doi: 10.1038/nbt0996-1112. [DOI] [PubMed] [Google Scholar]
- 126.Vater A, Jarosch F, Buchner K, Klussmann S. Short bioactive Spiegelmers to migraine-associated calcitonin gene-related peptide rapidly identified by a novel approach: tailored-SELEX. Nucleic acids research. 2003;31(21, article 130) doi: 10.1093/nar/gng130. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 127.Ohuchi SP, Ohtsu T, Nakamura Y. Selection ofRNA aptamers againstrecombinant transforming growth factor-β type III receptor displayed oncell surface. Biochimie. 2006;88(7):897–904. doi: 10.1016/j.biochi.2006.02.004. [DOI] [PubMed] [Google Scholar]
- 128.White R, Rusconi C, Scardino E, et al. Generation of species cross-reactive aptamers using “toggle” SELEX. Molecular Therapy. 2001;4(6):567–573. doi: 10.1006/mthe.2001.0495. [DOI] [PubMed] [Google Scholar]
- 129.Cassiday LA, Maher III LJ. Yeast genetic selections to optimize RNA decoys for transcription factor NF-κB. Proceedings of the National Academy of Sciences of the United States of America. 2003;100(7):3930–3935. doi: 10.1073/pnas.0736013100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 130.Ellington AD, Szostak JW. Selection in vitro of single-stranded DNA molecules that fold into specific ligand-binding structures. Nature. 1992;355(6363):850–852. doi: 10.1038/355850a0. [DOI] [PubMed] [Google Scholar]
- 131.Huizenga DE, Szostak JW. A DNA aptamer that binds adenosine and ATP. Biochemistry. 1995;34(2):656–665. doi: 10.1021/bi00002a033. [DOI] [PubMed] [Google Scholar]
- 132.Harada K, Frankel AD. Identification of two novel arginine binding DNAs. EMBO Journal. 1995;14(23):5798–5811. doi: 10.1002/j.1460-2075.1995.tb00268.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 133.Yang Q, Goldstein IJ, Mei H-Y, Engelke DR. DNA ligands that bind tightly and selectively to cellobiose. Proceedings of the National Academy of Sciences of the United States of America. 1998;95(10):5462–5467. doi: 10.1073/pnas.95.10.5462. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 134.Rink SM, Shen JC, Loeb LA. Creation of RNA molecules that recognize the oxidative lesion 7,8-dihydro-8-hydroxy-2′-deoxyguanosine (8-oxodG) in DNA. Proceedings of the National Academy of Sciences of the United States of America. 1998;95(20):11619–11624. doi: 10.1073/pnas.95.20.11619. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 135.Kato T, Takemura T, Yano K, Ikebukuro K, Karube I. In vitro selection of DNA aptamers which bind to cholic acid. Biochimica et Biophysica Acta. Gene Structure and Expression. 2000;1493(1-2):12–18. doi: 10.1016/s0167-4781(00)00080-4. [DOI] [PubMed] [Google Scholar]
- 136.Okazawa A, Maeda H, Fukusaki E, Katakura Y, Kobayashi A. In vitro selection of hematoporphyrin binding DNA aptamers. Bioorganic and Medicinal Chemistry Letters. 2000;10(23):2653–2656. doi: 10.1016/s0960-894x(00)00540-0. [DOI] [PubMed] [Google Scholar]
- 137.Vianini E, Palumbo M, Gatto B. In vitro selection of DNA aptamers that bind L-tyrosinamide. Bioorganic and Medicinal Chemistry. 2001;9(10):2543–2548. doi: 10.1016/s0968-0896(01)00054-2. [DOI] [PubMed] [Google Scholar]
- 138.Masud MM, Kuwahara M, Ozaki H, Sawai H. Sialyllactose-binding modified DNA aptamer bearing additional functionality by SELEX. Bioorganic and Medicinal Chemistry. 2004;12(5):1111–1120. doi: 10.1016/j.bmc.2003.12.009. [DOI] [PubMed] [Google Scholar]
- 139.Mann D, Reinemann C, Stoltenburg R, Strehlitz B. In vitro selection of DNA aptamers binding ethanolamine. Biochemical and Biophysical Research Communications. 2005;338(4):1928–1934. doi: 10.1016/j.bbrc.2005.10.172. [DOI] [PubMed] [Google Scholar]
- 140.Sando S, Narita A, Aoyama Y. Light-up Hoechst-DNA aptamer pair: generation of an aptamer-selective fluorophore from a conventional DNA-staining dye. ChemBioChem. 2007;8(15):1795–1803. doi: 10.1002/cbic.200700325. [DOI] [PubMed] [Google Scholar]
- 141.Kim YS, Jung HS, Matsuura T, Lee HY, Kawai T, Gu MB. Electrochemical detection of 17β-estradiol using DNA aptamer immobilized gold electrode chip. Biosensors and Bioelectronics. 2007;22(11):2525–2531. doi: 10.1016/j.bios.2006.10.004. [DOI] [PubMed] [Google Scholar]
- 142.Hayashi G, Hagihara M, Dohno C, Nakatani K. Photoregulation of a peptide-RNA interaction on a gold surface. Journal of the American Chemical Society. 2007;129(28):8678–8679. doi: 10.1021/ja071298x. [DOI] [PubMed] [Google Scholar]
- 143.Niazi JH, Lee SJ, Gu MB. Single-stranded DNA aptamers specific for antibiotics tetracyclines. Bioorganic and Medicinal Chemistry. 2008;16(15):7245–7253. doi: 10.1016/j.bmc.2008.06.033. [DOI] [PubMed] [Google Scholar]
- 144.Ohsawa K, Kasamatsu T, Nagashima JI, et al. Arginine-modified DNA aptamers that show enantioselective recognition of the dicarboxylic acid moiety of glutamic acid. Analytical Sciences. 2008;24(1):167–172. doi: 10.2116/analsci.24.167. [DOI] [PubMed] [Google Scholar]
- 145.Wochner A, Menger M, Orgel D, et al. A DNA aptamer with high affinity and specificity for therapeutic anthracyclines. Analytical Biochemistry. 2008;373(1):34–42. doi: 10.1016/j.ab.2007.09.007. [DOI] [PubMed] [Google Scholar]
- 146.Walsh R, DeRosa MC. Retention of function in the DNA homolog of the RNA dopamine aptamer. Biochemical and Biophysical Research Communications. 2009;388(4):732–735. doi: 10.1016/j.bbrc.2009.08.084. [DOI] [PubMed] [Google Scholar]
- 147.Miyachi Y, Shimizu N, Ogino C, Fukuda H, Kondo A. Selection of a DNA aptamer that binds 8-OHdG using GMP-agarose. Bioorganic and Medicinal Chemistry Letters. 2009;19(13):3619–3622. doi: 10.1016/j.bmcl.2009.04.130. [DOI] [PubMed] [Google Scholar]
- 148.Joeng CB, Niazi JH, Lee SJ, Gu MB. ssDNA aptamers that recognize diclofenac and 2-anilinophenylacetic acid. Bioorganic and Medicinal Chemistry. 2009;17(15):5380–5387. doi: 10.1016/j.bmc.2009.06.044. [DOI] [PubMed] [Google Scholar]
- 149.He J, Liu Y, Fan M, Liu X. Isolation and identification of the DNA aptamer target to acetamiprid. Journal of Agricultural and Food Chemistry. 2011;59(5):1582–1586. doi: 10.1021/jf104189g. [DOI] [PubMed] [Google Scholar]
- 150.Song KM, Cho M, Jo H, et al. Gold nanoparticle-based colorimetric detection of kanamycin using a DNA aptamer. Analytical Biochemistry. 2011;415(2):175–181. doi: 10.1016/j.ab.2011.04.007. [DOI] [PubMed] [Google Scholar]
- 151.Yang X, Bing T, Mei H, Fang C, Cao Z, Shangguan D. Characterization and application of a DNA aptamer binding to l-tryptophan. Analyst. 2011;136(3):577–585. doi: 10.1039/c0an00550a. [DOI] [PubMed] [Google Scholar]
- 152.Barthelmebs L, Jonca J, Hayat A, Prieto-Simon B, Marty JL. Enzyme-Linked Aptamer Assays (ELAAs), based on a competition format for a rapid and sensitive detection of Ochratoxin A in wine. Food Control. 2011;22(5):737–743. [Google Scholar]
- 153.Renaud De La Faverie A, Hamon F, Di Primo C, et al. Nucleic acids targeted to drugs: SELEX against a quadruplex ligand. Biochimie. 2011;93(8):1357–1367. doi: 10.1016/j.biochi.2011.05.022. [DOI] [PubMed] [Google Scholar]
- 154.Wang L, Liu X, Zhang Q, et al. Selection of DNA aptamers that bind to four organophosphorus pesticides. Biotechnology Letters. 2012;34(5):869–874. doi: 10.1007/s10529-012-0850-6. [DOI] [PubMed] [Google Scholar]
- 155.Xu S, Yuan H, Chen S, Xu A, Wang J, Wu L. Selection of DNA aptamers against polychlorinated biphenyls as potential biorecognition elements for environmental analysis. Analytical Biochemistry. 2012;423(2):195–201. doi: 10.1016/j.ab.2012.01.026. [DOI] [PubMed] [Google Scholar]
- 156.Mehta J, Rouah-Martin E, Van Dorst B, et al. Selection and characterization of PCB-binding DNA aptamers. Analytical Chemistry. 2012;84(3):1669–1676. doi: 10.1021/ac202960b. [DOI] [PubMed] [Google Scholar]
- 157.Song K-M, Jeong E, Jeon W, Cho M, Ban C. Aptasensor for ampicillin using gold nanoparticle based dual fluorescence-colorimetric methods. Analytical and Bioanalytical Chemistry. 2012;402(6):2153–2161. doi: 10.1007/s00216-011-5662-3. [DOI] [PubMed] [Google Scholar]
- 158.Szostak JW. Enzymatic activity of the conserved core of a group I self- splicing intron. Nature. 1986;322(6074):83–86. doi: 10.1038/322083a0. [DOI] [PubMed] [Google Scholar]
- 159.Majerfeld I, Yarus M. An RNA pocket for an aliphatic hydrophobe. Nature Structural Biology. 1994;1(5):287–292. doi: 10.1038/nsb0594-287. [DOI] [PubMed] [Google Scholar]
- 160.Lorsch JR, Szostak JW. In vitro selection of RNA aptamers specific for cyanocobalamin. Biochemistry. 1994;33(4):973–982. doi: 10.1021/bi00170a016. [DOI] [PubMed] [Google Scholar]
- 161.Burgstaller P, Famulok M. Isolation of RNA aptamers for biological cofactors by in vitro selection. Angewandte Chemie. 1994;33(10):1084–1087. [Google Scholar]
- 162.Lato SM, Boles AR, Ellington AD. In vitro selection of RNA lectins: using combinatorial chemistry to interpret ribozyme evolution. Chemistry and Biology. 1995;2(5):291–303. doi: 10.1016/1074-5521(95)90048-9. [DOI] [PubMed] [Google Scholar]
- 163.Wallis MG, Von Ahsen U, Schroeder R, Famulok M. A novel RNA motif for neomycin recognition. Chemistry and Biology. 1995;2(8):543–552. doi: 10.1016/1074-5521(95)90188-4. [DOI] [PubMed] [Google Scholar]
- 164.Wang Y, Rando RR. Specific binding of aminoglycoside antibiotics to RNA. Chemistry and Biology. 1995;2:281–290. doi: 10.1016/1074-5521(95)90047-0. [DOI] [PubMed] [Google Scholar]
- 165.Lauhon CT, Szostak JW. RNA aptamers that bind flavin and nicotinamide redox cofactors. Journal of the American Chemical Society. 1995;117(4):1246–1257. doi: 10.1021/ja00109a008. [DOI] [PubMed] [Google Scholar]
- 166.Wilson C, Nix J, Szostak J. Functional requirements for specific ligand recognition by a biotin-binding rna pseudoknot. Biochemistry. 1998;37(41):14410–14419. doi: 10.1021/bi981371j. [DOI] [PubMed] [Google Scholar]
- 167.Mannironi C, Di Nardo A, Fruscoloni P, Tocchini-Valentini GP. In vitro selection of dopamine RNA ligands. Biochemistry. 1997;36(32):9726–9734. doi: 10.1021/bi9700633. [DOI] [PubMed] [Google Scholar]
- 168.Haller AA, Sarnow P. In vitro selection of a 7-methyl-guanosine binding RNA that inhibits translation of capped mRNA molecules. Proceedings of the National Academy of Sciences of the United States of America. 1997;94(16):8521–8526. doi: 10.1073/pnas.94.16.8521. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 169.Welch M, Majerfeld I, Yarus M. 23S rRNA similarity from selection for peptidyl transferase mimicry. Biochemistry. 1997;36(22):6614–6623. doi: 10.1021/bi963135j. [DOI] [PubMed] [Google Scholar]
- 170.Burke DH, Hoffman DC, Brown A, Hansen M, Pardi A, Gold L. RNA aptamers to the peptidyl transferase inhibitor chloramphenicol. Chemistry and Biology. 1997;4(11):833–843. doi: 10.1016/s1074-5521(97)90116-2. [DOI] [PubMed] [Google Scholar]
- 171.Wallis MG, Streicher B, Wank H, et al. In vitro selection of a viomycin-binding RNA pseudoknot. Chemistry and Biology. 1997;4(5):357–366. doi: 10.1016/s1074-5521(97)90126-5. [DOI] [PubMed] [Google Scholar]
- 172.Holeman LA, Robinson SL, Szostak JW, Wilson C. Isolation and characterization of fluorophore-binding RNA aptamers. Folding and Design. 1998;3(6):423–431. doi: 10.1016/S1359-0278(98)00059-5. [DOI] [PubMed] [Google Scholar]
- 173.Wallace ST, Schroede R. In vitro selection and characterization of streptomycin-binding RNAs: recognition discrimination between antibiotics. RNA. 1998;4(1):112–123. [PMC free article] [PubMed] [Google Scholar]
- 174.Majerfeld I, Yarus M. Isoleucine:RNA sites with associated coding sequences. RNA. 1998;4(4):471–478. [PMC free article] [PubMed] [Google Scholar]
- 175.Kiga D, Futamura Y, Sakamoto K, Yokoyama S. An RNA aptamer to the xanthine/guanine base with a distinctive mode of purine recognition. Nucleic Acids Research. 1998;26(7):1755–1760. doi: 10.1093/nar/26.7.1755. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 176.Grate D, Wilson C. Laser-mediated, site-specific inactivation of RNA transcripts. Proceedings of the National Academy of Sciences of the United States of America. 1999;96(11):6131–6136. doi: 10.1073/pnas.96.11.6131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 177.Khvorova A, Kwak YG, Tamkun M, Majerfeld I, Yarus M. RNAs that bind and change the permeability of phospholipid membranes. Proceedings of the National Academy of Sciences of the United States of America. 1999;96(19):10649–10654. doi: 10.1073/pnas.96.19.10649. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 178.Koizumi M, Breaker RR. Molecular recognition of cAMP by an RNA aptamer. Biochemistry. 2000;39(30):8983–8992. doi: 10.1021/bi000149n. [DOI] [PubMed] [Google Scholar]
- 179.Jhaveri S, Rajendran M, Ellington AD. In vitro selection of signaling aptamers. Nature Biotechnology. 2000;18(12):1293–1297. doi: 10.1038/82414. [DOI] [PubMed] [Google Scholar]
- 180.Mannironi C, Scerch C, Fruscoloni P, Tocchini-Valentini GP. Molecular recognition of amino acids by RNA aptamers: the evolution into an L-tyrosine binder of a dopamine-binding RNA motif. RNA. 2000;6(4):520–527. doi: 10.1017/s1355838200991763. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 181.Gebhardt K, Shokraei A, Babaie E, Lindqvist BH. RNA aptamers to S-adenosylhomocysteine: kinetic properties, divalent cation dependency, and comparison with anti-S-adenosylhomocysteine antibody. Biochemistry. 2000;39(24):7255–7265. doi: 10.1021/bi000295t. [DOI] [PubMed] [Google Scholar]
- 182.Cowan JA, Ohyama T, Wang D, Natarajan K. Recognition of a cognate RNA aptamer by neomycin B: quantitative evaluation of hydrogen bonding and electrostatic interactions. Nucleic Acids Research. 2000;28(15):2935–2942. doi: 10.1093/nar/28.15.2935. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 183.Schürer H, Stembera K, Knoll D, et al. Aptamers that bind to the antibiotic moenomycin A. Bioorganic and Medicinal Chemistry. 2001;9(10):2557–2563. doi: 10.1016/s0968-0896(01)00030-x. [DOI] [PubMed] [Google Scholar]
- 184.Jeong S, Eom T-Y, Kim S-J, Lee S-W, Yu J. In vitro selection of the RNA Aptamer against the Sialyl Lewis X and its inhibition of the cell adhesion. Biochemical and Biophysical Research Communications. 2001;281(1):237–243. doi: 10.1006/bbrc.2001.4327. [DOI] [PubMed] [Google Scholar]
- 185.Berens C, Thain A, Schroeder R. A tetracycline-binding RNA aptamer. Bioorganic and Medicinal Chemistry. 2001;9(10):2549–2556. doi: 10.1016/s0968-0896(01)00063-3. [DOI] [PubMed] [Google Scholar]
- 186.Kwon M, Chun SM, Jeong S, Yu J. In vitro selection of RNA against kanamycin B. Molecules and Cells. 2001;11(3):303–311. [PubMed] [Google Scholar]
- 187.Meli M, Vergne J, Décout J-L, Maurel M-C. Adenine-aptamer complexes. A bipartite RNA site that binds the adenine nucleic base. Journal of Biological Chemistry. 2002;277(3):2104–2111. doi: 10.1074/jbc.M107130200. [DOI] [PubMed] [Google Scholar]
- 188.Roychowdhury-Saha M, Lato SM, Shank ED, Burke DH. Flavin recognition by an RNA aptamer targeted toward FAD. Biochemistry. 2002;41(8):2492–2499. doi: 10.1021/bi015719d. [DOI] [PubMed] [Google Scholar]
- 189.Lozupone C, Changayil S, Majerfeld I, Yarus M. Selection of the simplest RNA that binds isoleucine. RNA. 2003;9(11):1315–1322. doi: 10.1261/rna.5114503. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 190.Vaish NK, Larralde R, Fraley AW, Szostak JW, McLaughlin LW. A novel, modification-dependent ATP-binding aptamer selected from an RNA library incorporating a cationic functionality. Biochemistry. 2003;42(29):8842–8851. doi: 10.1021/bi027354i. [DOI] [PubMed] [Google Scholar]
- 191.Huang Z, Szostak JW. Evolution of aptamers with a new specificity and new secondary structures from an ATP aptamer. RNA. 2003;9(12):1456–1463. doi: 10.1261/rna.5990203. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 192.Brockstedt U, Uzarowska A, Montpetit A, Pfau W, Labuda D. In vitro evolution of RNA aptamers recognizing carcinogenic aromatic amines. Biochemical and Biophysical Research Communications. 2004;313(4):1004–1008. doi: 10.1016/j.bbrc.2003.12.030. [DOI] [PubMed] [Google Scholar]
- 193.Sazani PL, Larralde R, Szostak JW. A small aptamer with strong and specific recognition of the triphosphate of ATP. Journal of the American Chemical Society. 2004;126(27):8370–8371. doi: 10.1021/ja049171k. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 194.Legiewicz M, Yarus M. A more complex isoleucine aptamer with a cognate triplet. Journal of Biological Chemistry. 2005;280(20):19815–19822. doi: 10.1074/jbc.M502329200. [DOI] [PubMed] [Google Scholar]
- 195.Majerfeld I, Puthenvedu D, Yarus M. RNA affinity for molecular L-histidine; genetic code origins. Journal of Molecular Evolution. 2005;61(2):226–235. doi: 10.1007/s00239-004-0360-9. [DOI] [PubMed] [Google Scholar]
- 196.Lévesque D, Beaudoin JD, Roy S, Perreault JP. In vitro selection and characterization of RNA aptamers binding thyroxine hormone. Biochemical Journal. 2007;403(1):129–138. doi: 10.1042/BJ20061216. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 197.Morse DP. Direct selection of RNA beacon aptamers. Biochemical and Biophysical Research Communications. 2007;359(1):94–101. doi: 10.1016/j.bbrc.2007.05.072. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 198.Lee HW, Robinson SG, Bandyopadhyay S, Mitchell RH, Sen D. Reversible photo-regulation of a hammerhead ribozyme using a diffusible effector. Journal of Molecular Biology. 2007;371(5):1163–1173. doi: 10.1016/j.jmb.2007.06.042. [DOI] [PubMed] [Google Scholar]
- 199.Constantin TP, Silva GL, Robertson KL, et al. Synthesis of new fluorogenic cyanine dyes and incorporation into RNA fluoromodules. Organic Letters. 2008;10(8):1561–1564. doi: 10.1021/ol702920e. [DOI] [PubMed] [Google Scholar]
- 200.Endo K, Nakamura Y. A binary Cy3 aptamer probe composed of folded modules. Analytical Biochemistry. 2010;400:103–109. doi: 10.1016/j.ab.2010.01.015. [DOI] [PubMed] [Google Scholar]
- 201.Lee J, Lee KH, Jeon J, Dragulescu-Andrasi A, Xiao F, Rao J. Combining SELEX screening and rational design to develop light-up fluorophore-RNA aptamer pairs for RNA tagging. ACS Chemical Biology. 2010;5(11):1065–1074. doi: 10.1021/cb1001894. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 202.Sinha J, Reyes SJ, Gallivan JP. Reprogramming bacteria to seek and destroy an herbicide. Nature Chemical Biology. 2010;6(6):464–470. doi: 10.1038/nchembio.369. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- 203.Horii K, Omi K, Yoshida Y, et al. Development of a sphingosylphosphorylcholine detection system using RNA aptamers. Molecules. 2010;15(8):5742–5755. doi: 10.3390/molecules15085742. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 204.Murata A, Sato SI, Kawazoe Y, Uesugi M. Small-molecule fluorescent probes for specific RNA targets. Chemical Communications. 2011;47(16):4712–4714. doi: 10.1039/c1cc10393h. [DOI] [PubMed] [Google Scholar]
- 205.Paige JS, Wu KY, Jaffrey SR. RNA mimics of green fluorescent protein. Science. 2011;333(6042):642–646. doi: 10.1126/science.1207339. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 206.Bala J, Bhaskar A, Varshney A, Singh AK, Dey S, Yadava P. In vitro selected RNA aptamer recognizing glutathione induces ROS-mediated apoptosis in the human breast cancer cell line MCF 7. RNA Biology. 2011;8(1):101–111. doi: 10.4161/rna.8.1.14116. [DOI] [PubMed] [Google Scholar]
- 207.Lau JL, Baksh MM, Fiedler JD, et al. Evolution and protein packaging of small-molecule RNA aptamers. ACS Nano. 2011;5:7722–7729. doi: 10.1021/nn2006927. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 208.Flinders J, DeFina SC, Brackett DM, Baugh C, Wilson C, Dieckmann T. Recognition of planar and nonplanar ligands in the malachite green—RNA aptamer complex. ChemBioChem. 2004;5(1):62–72. doi: 10.1002/cbic.200300701. [DOI] [PubMed] [Google Scholar]
- 209.Cruz-Aguado JA, Penner G. Fluorescence polarization based displacement assay for the determination of small molecules with aptamers. Analytical Chemistry. 2008;80(22):8853–8855. doi: 10.1021/ac8017058. [DOI] [PubMed] [Google Scholar]
- 210.Guedin A, Lacroix L, Mergny JL. Thermal melting studies of ligand DNA interactions. Methods in Molecular Biology. 2010;613:25–35. doi: 10.1007/978-1-60327-418-0_2. [DOI] [PubMed] [Google Scholar]
- 211.Lin P, Chen R, Lee C, Chang Y, Chen C, Chen W. Studies of the binding mechanism between aptamers and thrombin by circular dichroism, surface plasmon resonance and isothermal titration calorimetry. Colloids and Surfaces B. 2011;88:552–558. doi: 10.1016/j.colsurfb.2011.07.032. [DOI] [PubMed] [Google Scholar]
- 212.Lee JH, Canny MD, De Erkenez A, et al. A therapeutic aptamer inhibits angiogenesis by specifically targeting the heparin binding domain of VEGF165. Proceedings of the National Academy of Sciences of the United States of America. 2005;102(52):18902–18907. doi: 10.1073/pnas.0509069102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 213.Sultan Y, Walsh R, Monreal C, DeRosa MC. Preparation of functional aptamer films using layer-by-layer self-assembly. Biomacromolecules. 2009;10(5):1149–1154. doi: 10.1021/bm8014126. [DOI] [PubMed] [Google Scholar]
- 214.Deng Q, German I, Buchanan D, Kennedy RT. Retention and separation of adenosine and analogues by affinity chromatography with an aptamer stationary phase. Analytical Chemistry. 2001;73(22):5415–5421. doi: 10.1021/ac0105437. [DOI] [PubMed] [Google Scholar]
- 215.Drabovich AP, Berezovski M, Okhonin V, Krylov SN. Selection of smart aptamers by methods of kinetic capillary electrophoresis. Analytical Chemistry. 2006;78(9):3171–3178. doi: 10.1021/ac060144h. [DOI] [PubMed] [Google Scholar]
- 216.Bao J, Krylova SM, Reinstein O, Johnson PE, Krylov SN. Label-free solution-based kinetic study of aptamer-small molecule interactions by kinetic capillary electrophoresis with UV detection revealing how kinetics control equilibrium. Analytical Chemistry. 2011;83:8387–8390. doi: 10.1021/ac2026699. [DOI] [PubMed] [Google Scholar]
- 217.Turgeon RT, Fonslow BR, Jing M, Bowser MT. Measuring aptamer equilibria using gradient micro free flow electrophoresis. Analytical Chemistry. 2010;82(9):3636–3641. doi: 10.1021/ac902877v. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 218.Hall B, Arshad S, Seo K, et al. In vitro selection of RNA aptamers to a protein target by filter immobilization. Current Protocols in Molecular Biology. 2009;(88):24.3.1–24.3.27. doi: 10.1002/0471142727.mb2403s88. [DOI] [PubMed] [Google Scholar]
- 219.Gaillard C, Strauss F. DNA loops and semicatenated DNA junctions. BMC Biochemistry. 2000;1, article no. 1:1–7. doi: 10.1186/1471-2091-1-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 220.Baaske P, Wienken CJ, Reineck P, Duhr S, Braun D. Optical thermophoresis for quantifying the buffer dependence of aptamer binding. Angewandte Chemie. 2010;49(12):2238–2241. doi: 10.1002/anie.200903998. [DOI] [PubMed] [Google Scholar]
- 221.Potty ASR, Kourentzi K, Fang H, et al. Biophysical characterization of DNA aptamer interactions with vascular endothelial growth factor. Biopolymers. 2009;91(2):145–156. doi: 10.1002/bip.21097. [DOI] [PubMed] [Google Scholar]
- 222.Oh SS, Plakos K, Lou X, Xiao Y, Soh HT. In vitro selection of structure-switching, self-reporting aptamers. Proceedings of the National Academy of Sciences of the United States of America. 2010;107(32):14053–14058. doi: 10.1073/pnas.1009172107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 223.Regulski EE, Breaker RR. In-line probing analysis of riboswitches. Methods in Molecular Biology. 2008;419:53–67. doi: 10.1007/978-1-59745-033-1_4. [DOI] [PubMed] [Google Scholar]
- 224.Yoshida W, Sode K, Ikebukuro K. Homogeneous DNA sensing using enzyme-inhibiting DNA aptamers. Biochemical and Biophysical Research Communications. 2006;348(1):245–252. doi: 10.1016/j.bbrc.2006.07.069. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 225.McManus SA, Li Y. Multiple occurrences of an efficient self-phosphorylating deoxyribozyme motif. Biochemistry. 2007;46(8):2198–2204. doi: 10.1021/bi061613c. [DOI] [PubMed] [Google Scholar]


