Abstract
MicroRNA (miRNA), a small non-coding RNA that functions as a mediator in gene silencing, plays important roles in gene regulation in various vital functions and activities. Here we show that the miR-29 members are upregulated in klotho-deficient [klotho(−/−)] mice, a senescence-model animal, and also in normal elderly ICR mice relative to wild-type littermates and young ICR mice. In addition, levels of type IV collagen, a major component of basement membranes and a putative target of miR-29, were lower in klotho(−/−) and elderly ICR mice than in wild-type littermates and young ICR mice. RNA degradation mediated by miR-29 may participate in the suppression of type IV collagen, both in vivo and in vitro. Taken together, our current findings suggest that the miR-29 upregulated in aging may be involved in the downregulation of type IV collagen, leading to a possible weakening of the basal membrane in senescent tissues, and miR-29 may be a useful molecular marker of senescence.
Introduction
Senescence is a set of biological changes that occurs in an individual with increasing age. A common change is the decline of physiology such as a decrease in hearing, visual activity, and memory skills; and behind such a physiological decline, there are presumably biochemical changes involved. In order to understand such complex senescent phenomena at a molecular level, finding biological molecules that show quantitative and qualitative variation with increasing age is vital; and such molecules may be useful as senescence makers.
MicroRNA (miRNA) is a 21∼23-nucleotide-long small non-coding RNA and functions as a mediator in gene silencing. Hundreds of miRNA genes have been found in animals and plants [see the microRNA database (miRBase): http://www.mirbase.org/index.shtml]. miRNAs play essential roles in gene regulation by inhibiting translation of messenger RNAs (mRNAs) that are partially complementary to the miRNAs, and by digestion of mRNAs that are nearly complementary to the miRNAs, or by RNA interference (RNAi), during cell proliferation, differentiation and development [1]–[5]. Expression profiles of miRNAs are quite useful in understanding complex gene regulation involving miRNAs and in characterizing miRNAs themselves. Many expression analyses have been performed in various tissues and cells, and revealed tissue- and stage-specific expression of miRNAs. In mammals, it has been found that a major small RNA class transition from retrotransposon-derived small interfering RNAs (siRNAs) and Piwi-interacting RNAs (piRNAs) to zygotically expressed miRNAs occurs during early development of pre-implantation embryos [6], and tissue- or organ-specific expression patterns of miRNAs are generated thereafter [4], [7]–[9].
Age-associated change in gene expression involving miRNAs is an important research field. We investigated the expression profile of miRNAs in mouse brain tissue by means of DNA chip and reverse-transcription quantitative polymerase chain reaction (RT-qPCR), and found that the alteration of the miRNA expression occurred during the course of mouse brain growth [10].
In this study we investigated miRNAs in klotho-deficient [klotho(−/−)] mice, which are a senescence model showing a number of phenotypes resembling human premature aging [11], and also miRNAs in 1 month and 14–18 month old outbred ICR mice studied as normal young and elderly mice, respectively. The data indicated that the miR-29 members were markedly increased in klotho(−/−) and elderly ICR mice compared with wild-type littermates and young ICR mice.
Materials and Methods
Mice
Klotho-deficient male mice and ICR male mice were purchased from CLEA Japan, Inc (Tokyo, Japan). Mice were housed, fed and maintained in the laboratory animal facility according to the National Institute of Neuroscience animal care guidelines. This study was carried out in strict accordance with the recommendations in the Guide for the Care and Use of Laboratory Animals of the National Institutes of Neuroscience. The protocol was approved by the Committee on the Ethics of Animal Experiments of the National Institutes of Neuroscience (Permit Number: 2012003).
DNA and RNA oligonucleotides
DNA oligonucleotides and small RNA duplexes used in this study were synthesized by and purchased from Sigma-Aldrich (St Louis, MO, USA).
DNA chip analysis
Genopal®-MICM DNA chips (Mitsubishi Rayon, Tokyo, Japan) for detection of mouse miRNAs were used. DNA oligonucleotide probes for 200 miRNAs are installed on the DNA chip, and the list of the detectable miRNAs is available at the following address: http://www.mrc.co.jp/genome/pdf/micm07_list.pdf
Tissues were surgically removed from sacrificed mice and total RNAs were immediately isolated from the fresh tissues using TRIzol reagent (Invitrogen, Carlsbad, CA, USA) according to the manufacturer's instructions. Preparation of small-sized RNAs, fluorescent labeling, hybridization with DNA chips and detection of hybridization signals were carried out as described previously [10], [12], [13].
Construction of reporter genes and hydrodynamic delivery method
To examine the knockdown potencies of endogenous miR-29a and miR-29b in vivo, we constructed reporter genes with the psiCheck2 vector (Promega): the Renilla luciferase gene carrying the complementary (target) sequence of miR-29a or miR-29b in the 3′ untranslated region was constructed as described previously [14]. The target sequences synthesized in the construction were as follows:
For the target of miR-29a;
Sense-strand; 5′- TCGAGTAACCGATTTCAGATGGTGCTATTACTAGT-3′
Antisense-strand; 5′- ACTAGTAATAGCACCATCTGAAATCGGTTAC-3′
For the target of miR-29b;
Sense-strand; 5′- TCGAGAACACTGATTTCAAATGGTGCTATTACTAGT-3′
Antisense-strand; 5′- ACTAGTAATAGCACCATTTGAAATCAGTGTTC-3′
The reporter plasmid and the pSV-β-Galactosidase control vector (Promega) as a control were administered to young (5-week-old) and elderly (8∼18-month-old) ICR mice by a hydrodynamic delivery method. Briefly, 100 µg of the reporter plasmid and 40 µg of the pSV-β-Galactosidase control vector were mixed in 2 ml of sterile saline solution. ICR mice were anesthetized with somnopentyl (50 mg/kg b.w.), and 2 ml of the plasmid mixture was systemically administered via the lateral tail vein within 5 seconds. Three days after administration, the heart, lung, liver and kidney were excised from the mice, and the tissue extracts were prepared. The expression levels of the luciferase and β-galactosidase reporter genes were measured using a Dual-Luciferase Reporter Assay System (Promega) and a Beta-Glo Assay System (Promega), respectively, by the Fusion Universal Microplate Analyzer (PerkinElmer, Waltham, MA, USA).
Systemic administration of the synthetic miR-29b duplex
The synthetic miR-29b duplex and siControl (non-silencing siRNA; QIAGEN, Venlo, Netherlands) were each prepared with atelocollagen (AteloGene Systemic Use; KOKEN, Tokyo, Japan) according to the manufacturer's instructions. The resultant RNA/atelocollagen complexes (1.0 mg/kg b.w.) were systemically administered to normal male ICR mice (5-week-old; n = 5/group) via the lateral tail vein. Four systemic administrations were carried out (Day 1, 4, 8, and 12), and two days after the last administration (Day 14) the treated mice were examined. The sequences of the synthetic miR-29b duplexes were as follows:
Sense- strand; 5′- UAGCACCAUUUGAAAUCAGUGUU-3′
Antisense-strand; 5′- CACUGAUUUCAAAUGGUGUUAUU-3′
Cell culture, transfection of miR-29 mimics and cell viability assay
Neuro2a (N2a), a mouse neuroblastoma cell line originally established by RJ Kleber and FH Ruddle (1969) [15], was used in this study. Human embryonic kidney 293 (HEK293) cells were obtained from the Health Science Research Resources Bank. The cells were grown in Dulbecco's modified Eagle's medium (Wako, Osaka, Japan) supplemented with 10% fetal bovine serum (Invitrogen), 100 U/ml penicillin and 100 µg/ml streptomycin (Wako) at 37°C in 5% CO2-humidified chamber as described previously [13], [16], [17]. The day before transfection, cells were trypsinized, diluted with culture medium without antibiotics, and seeded into 96-well culture plates at a cell density 5×104 cells/cm2. To each well, 20 nM (final concentration) of each MISSION microRNA mimic of has-miR-29a, has-miR-29b, or negative control miRNA (Sigma-Aldrich) was transfected into the cells using Lipofectamine2000 transfection reagent (Invitrogen) according to the manufacturer's instructions. Four hours after transfection, the culture medium was replaced with fresh medium containing antibiotics. After the 3-day incubation, total RNA isolation as described above or cell viability assay was carried out. Cell viability was examined with the CellTiter 96 AQueous One Solution Cell Proliferation Assay (Promega, Fitchburg, WI, USA) according to the manufacturer's instructions.
For assessment of the knockdown potency of miR-29a or miR-29b, the constructed reporter plasmids (see above) together with or without synthetic miR-29 duplexes were transfected into N2a cells, and 24 hour after transfection the dual luciferase assay was carried out as described previously [14].
Reverse transcription (RT)-(real time)-polymerase chain reaction (PCR)
Total RNAs extracted by TRIzol reagent (Invitrogen) were treated with Turbo DNase I (Applied Biosystems, Carlsbad, CA, USA) according to the manufacturer's instructions and subjected to RT-(real time)-PCR. Real time PCR (quantitative PCR; qPCR) was performed using the AB 7300 Real Time PCR System (Applied Biosystems) with a TaqMan Universal PCR Master Mix together with TaqMan® MicroRNA Assays (Applied Biosystems) according to the manufacturer's instructions.
The TaqMan® MicroRNA Assays used were as follows (ABI Assay IDs are indicated in parentheses):
hsa-miR-29a (2112), hsa-miR-29b (413) and snoRNA202 (1232).
For examination of protein-coding genes, total RNAs were subjected to cDNA synthesis using Oligo(dT)15 primers (Promega) and a SuperScript III reverse transcriptase (Invitrogen) according to the manufacturer's instructions. The cDNAs were examined by qPCR using the AB 7300 Real Time PCR System (Applied Biosystems) with SYBR Green PCR Master Mix (Applied Biosystems) and TaqMan Universal PCR Master Mix together with Perfect Real Time Primers (TAKARA BIO, Otsu, Shiga, Japan) and TaqMan® Gene Expression Assays (Applied Biosystems), respectively, according to the manufacturer's instructions.
The Perfect Real Time Primers used were as follows (TAKARA BIO primer-set IDs are indicated in parentheses):
Col4a1 (MA118711), Col4a2 (MA112149), Col4a3 (MA124802), Col4a4 (MA077950), Col4a5 (MA113961), Col4a6 (MA087283), Cabin1 (MA069171), Dnmt1 (MA060016), Dicer1 (MA043537), Lamc1 (MA059108), Gapdh, (MA050371), COL4A1 (HA156982), COL4A2 (HA174126), CABIN1 (HA183487), DNMT1 (HA002770), DICER1 (HA174786), LAMC1 (HA083467) and GAPDH (HA067812).
TaqMan® Gene Expression Assays used were as follows (ABI Assay IDs are indicated in parentheses):
klotho (Mm00502002_m1) and Gapdh (Mm99999915_g1).
Western blotting
Equal amounts of protein was separated by SDS-PAGE and electrophoretically blotted onto PVDF membranes (Millipore, Billerica, MA, USA). Membranes were blocked in blocking solution (4% bovine serum albumin and 0.05% Tween-20 in TBS) and incubated with rabbit polyclonal anti-collagen IV (mouse) antibody (250485; ABBIOTEC, San Diego, CA, USA) or mouse monoclonal anti-α-tubulin antibody (Sigma-Aldrich), followed by washing in TBS containing 0.05% Tween-20, and then incubated with horseradish peroxidase-conjugated anti-rabbit or anti-mouse anti-goat IgG. Antigen-antibody complexes were visualized using a chemiluminescent reagent (Millipore).
Blood tests
Blood samples drawn from mice were subjected to biochemical tests. To examine renal function, plasma creatinine levels were measured with a creatinine assay kit (BioVision Inc., Milpitas, CA, USA). The plasma levels of hexanoyl-lysine adduct (HEL) and 8-hydroxy-2′-deoxyguanosine (8OH-dG) as stress markers were measured by HEL ELISA Kit [Japan Institute for the Control of Aging (JaICA), NIKKEN SEIL Co., Ltd., Shizuoka, Japan] and High Sensitive 8-OHdG Check ELISA Kit (JaICA), respectively.
Statistical analysis
Differences in the levels of expression of miRNAs were analyzed by Student's t-test (two-tailed).
Accession numbers
The Gene Expression Omnibus (GEO) accession numbers for the miRNA expression data used in this study are GSE34531 (miRNAs in Klotho(−/−) and wild-type mouse tissues) and GSE34532 (miRNAs in young and elderly ICR mouse tissues).
Results
miRNA expression profiles of klotho-deficient mice
The klotho gene encodes a single-pass transmembrane protein involved in the maintenance of homeostasis, and klotho-deficient [klotho(−/−)] mice are known to be an model of aging that shows various senescence-like phenotypes resembling human premature aging such as a short lifespan, decreased bone mineral density and atrophy of lymphopoietic and reproductive organs [11]. We investigated miRNAs in klotho(−/−) mice and wild-type littermates by means of DNA chip. Duplicated DNA chip analyses with different individual mice exhibited reproducible results except for the data of the klotho(−/−) spleen (Fig. S1); the discrepancy for the spleen miRNAs may be due to individual differences.
Fold changes (FC) in the expression levels of miRNAs between klotho(−/−) and wild-type mice were calculated, and the miRNAs that showed more than 1.5 FC values or less than 0.5 FC values are listed (Table 1 and Table S1): the former (>1.5 FC) represents the miRNA that were markedly increased in klotho(−/−) mice versus wild-type mice, and the latter (<0.5 FC) represents the miRNA markedly decreased in klotho(−/−) mice. From the FC analysis, the miR-29 members (miR-29a, -29b and -29c) appear to be commonly upregulated in klotho(−/−) mice (Table 1).
Table 1. Difference in the expression of miRNAs between klotho-deficient and wild-type mice.
| Exp.1 | Exp.2 | ||||||
| Tissue | miRNA | Kl | Wt | FC | Kl | Wt | FC |
| Lung | mmu-miR-29b | 791 | 332 | 2.38 | 567 | 150 | 3.79 |
| mmu-miR-29c | 502 | 256 | 1.96 | 449 | 133 | 3.37 | |
| mmu-miR-409 | 42 | 22 | 1.88 | 41 | 26 | 1.55 | |
| mmu-miR-296 | 124 | 67 | 1.86 | 122 | 45 | 2.71 | |
| mmu-miR-29a | 978 | 575 | 1.70 | 899 | 266 | 3.38 | |
| mmu-miR-101a | 228 | 147 | 1.55 | 214 | 88 | 2.43 | |
| Liver | mmu-miR-29b | 171 | 22 | 7.70 | 122 | 39 | 3.13 |
| mmu-miR-29a | 216 | 47 | 4.60 | 154 | 67 | 2.30 | |
| mmu-miR-29c | 106 | 25 | 4.30 | 90 | 36 | 2.48 | |
| mmu-miR-145 | 46 | 14 | 3.35 | 31 | 16 | 1.89 | |
| mmu-miR-101a | 102 | 32 | 3.19 | 77 | 51 | 1.52 | |
| mmu-miR-27a | 104 | 34 | 3.09 | 75 | 49 | 1.51 | |
| mmu-miR-24 | 120 | 42 | 2.86 | 89 | 57 | 1.57 | |
| mmu-miR-126-3p | 97 | 36 | 2.71 | 72 | 48 | 1.50 | |
| mmu-miR-26a | 219 | 84 | 2.62 | 198 | 121 | 1.63 | |
| mmu-miR-101b | 131 | 55 | 2.40 | 116 | 75 | 1.56 | |
| mmu-miR-30b | 44 | 19 | 2.33 | 37 | 24 | 1.56 | |
| Kidney | mmu-miR-29a | 437 | 158 | 2.76 | 361 | 192 | 1.88 |
| mmu-miR-29b | 358 | 143 | 2.50 | 293 | 171 | 1.72 | |
| mmu-miR-34a | 99 | 41 | 2.41 | 105 | 49 | 2.16 | |
| mmu-miR-31 | 17 | 7 | 2.34 | 15 | 10 | 1.53 | |
| mmu-let-7b | 1099 | 471 | 2.33 | 487 | 315 | 1.55 | |
| mmu-miR-199a | 65 | 29 | 2.23 | 51 | 33 | 1.54 | |
| mmu-miR-92 | 54 | 25 | 2.20 | 34 | 19 | 1.83 | |
| mmu-miR-125a | 161 | 74 | 2.19 | 105 | 61 | 1.73 | |
| mmu-miR-21 | 418 | 193 | 2.17 | 323 | 212 | 1.53 | |
| mmu-miR-296 | 120 | 56 | 2.16 | 55 | 36 | 1.51 | |
| mmu-miR-125b | 212 | 101 | 2.10 | 152 | 94 | 1.61 | |
| mmu-let-7c | 1563 | 755 | 2.07 | 806 | 515 | 1.57 | |
| Heart | mmu-miR-298 | 64 | 4 | 15.66 | 27 | 5 | 5.88 |
| mmu-miR-291a-3p | 35 | 2 | 15.50 | 18 | 2 | 7.49 | |
| mmu-miR-370 | 30 | 4 | 7.05 | 13 | 6 | 2.02 | |
| mmu-miR-328 | 39 | 10 | 4.03 | 13 | 8 | 1.63 | |
| mmu-miR-150 | 39 | 16 | 2.46 | 25 | 15 | 1.64 | |
| mmu-miR-194 | 18 | 8 | 2.28 | 14 | 7 | 1.86 | |
| mmu-miR-29b | 320 | 143 | 2.24 | 240 | 136 | 1.77 | |
| mmu-miR-92 | 47 | 31 | 1.52 | 28 | 17 | 1.64 | |
| mmu-miR-146 | 12 | 22 | 0.56 | 8 | 14 | 0.57 | |
Hybridization signal intensities obtained from DNA chip analysis [duplicated experiments (Exp.) 1 and 2] are indicated.
Fold changes (FC) in the expression of miRNAs between klotho (Kl) and wild-type (Wt) mice are calculated.
Upregulation of miR-29 in elderly ICR mice
To see if normal aged-mice, like klotho(−/−) mice, have a similar increase in the expression of miR-29, we measured miR-29a and miR-29b in outbred ICR mice at age 14 or 18 months (14 m, 18 m; elderly mice) and 1 month (1 m; young mice) by means of RT-qPCR; note that five different individual ICR mice in each group were examined to account for individual differences. The results of the qPCR analysis consistently indicated that the levels of either miR-29a or miR-29b were significantly higher in elderly mouse tissues than in young tissues (Fig. 1).
Figure 1. miR-29a and miR-29b expression in young and elderly ICR mice.
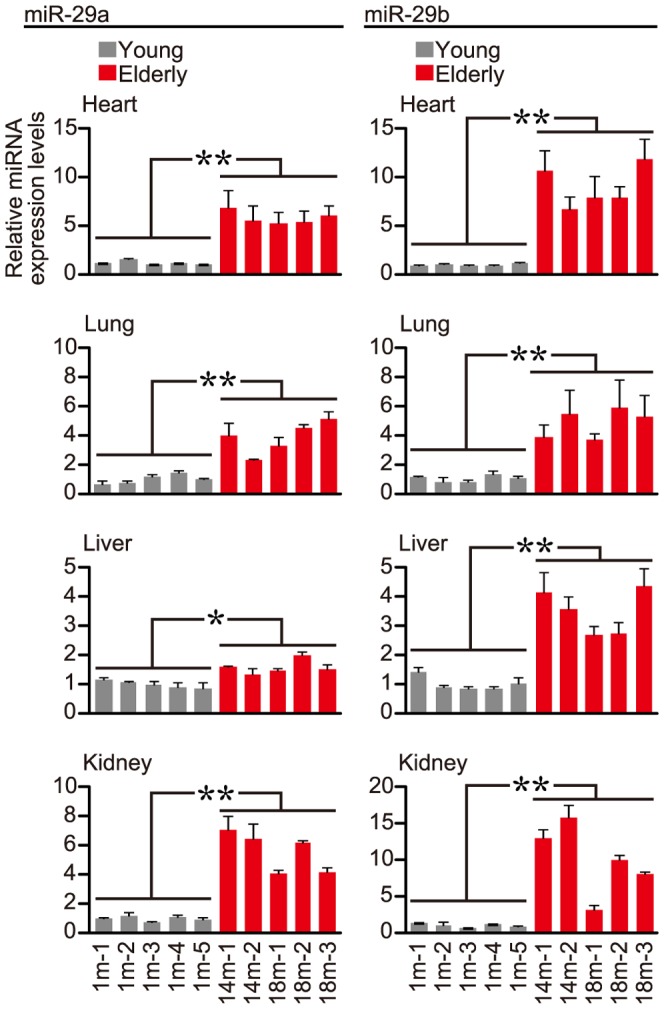
Total RNAs were extracted from indicated tissues of young (1-month-old: 1 m) and elderly (14- or 18-month-old: 14 m or 18 m) ICR mice, and subjected to RT-qPCR analysis. The miR-29a, miR-29b and snoRNA202 (as a control) expressions were examined and analyzed by the delta-delta Ct method. Five individual mice in each group were investigated to account for individual variability, and three measurements by qPCR per sample were carried out. Error bars represent standard deviations. The normalized miR-29a and miR-29b expression levels in the young (gray bars) and elderly (red bars) groups in each tissue were further examined by Student's t-test, and significant differences between the two groups in all the tissues examined were detected (* P<0.05, **P<0.01).
We further investigated whether the upregulated miR-29 members in elderly mice enhanced their mediated knockdown activities. To address this, the luciferase reporter genes carrying the complementary (target) sequences of miR-29a and miR-29b, respectively, in their 3′ untranslated regions (3′UTRs) were constructed (Fig. 2), and administered to elderly and young ICR mice by a hydrodynamic delivery method. Three days after administration, the expression of the reporter gene was examined. Neither heart nor lung had expression of the luciferase reporter genes, whereas liver and kidney exhibited expression; however, the expression data in liver, particularly the levels of the luciferase carrying no target sequence (control empty vector), were poorly-reproducible, so that they were not able to be further evaluated (Fig. S2). The expression data for kidney were further normalized to the data obtained with the control empty vector. The normalized expression levels of the reporter gene carrying either the miR-29a or miR-29b target sequence was markedly decreased in elderly kidney than in young kidney (Fig. 3). Therefore, miR-29 upregulation in elderly kidney may result in the enhancement of gene silencing mediated by it.
Figure 2. Assessment of knockdown potency using reporter genes.

(A) Schematic drawing of constructed reporter genes. The psiCheck2-backbone reporter plasmid encoding the Renilla luciferase gene carrying the target sequence of miR-29a (psiCheck2-miR29a) or miR-29b (psiCheck2-miR29b) was constructed. The SV40 promoter, Renilla luciferase coding region (CDR) and 3′ untranslated region (3′UTR) of the reporter genes are indicated. The target sequences of miR-29a and miR-29b are indicated in the 3′UTRs. (B, C) Suppression of reporter genes by synthetic miR-29 mimics. The reporter plasmids indicated were transfected together with synthetic miRNA mimics of miR-29a (mimic_29a), miR-29b (mimic_29b) or a negative control miRNA (negative_miR) into N2a cells. 24 hr after transfection, dual-luciferase assay was carried out. The target (Renilla) luciferase activity was normalized to control (Photinus) luciferase activity and further normalized to the data obtained from N2a cells transfected with only the reporter plasmid (non-treatment). Data are averages of four independent experiments. Error bars represent standard deviations. The normalized data were analyzed by one-way analysis of variance (ANOVA), followed by Dunnett's test. Significant differences are indicated by * (P<0.05).
Figure 3. In vivo knockdown potency mediated by endogenous miR-29a or miR-29b.
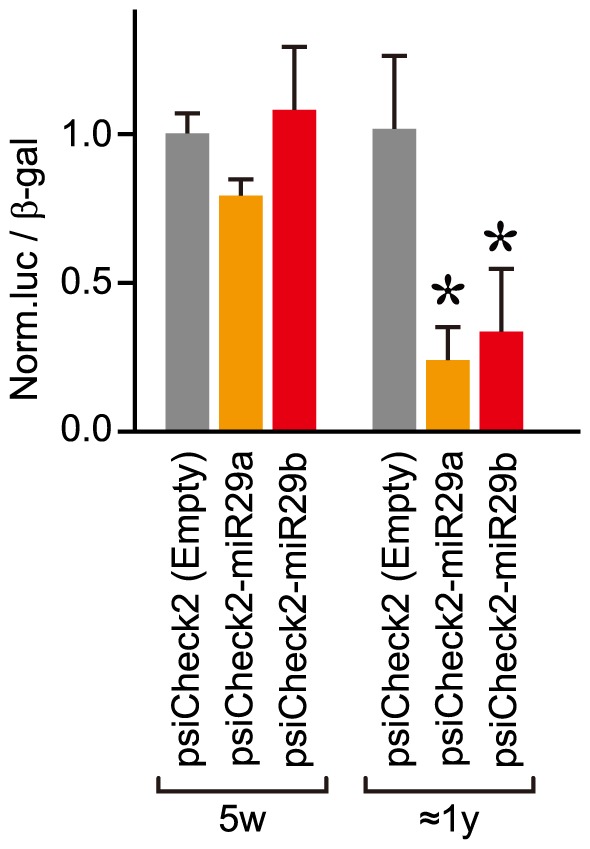
The constructed reporter plasmid carrying the miR-29a or miR-29b target sequence (Fig. 2A) and the β-galactosidase expression plasmid as a control were systemically administrated to young (5-week-old: 5w) and elderly (8∼18-month-old: ≈1 y) ICR mice by a hydrodynamic delivery method. Three days after administration, kidney cell extracts were prepared from the treated mice, and then the luciferase and β-galactosidase activities were examined. The expression levels of luciferase were normalized to those of β-galactosidase, and further normalized to the data obtained with a control empty vector (Empty). Data are averages of three different individual's data. Error bars represent standard deviations. The normalized data in the young (5w) and elderly (≈1 y) groups were analyzed by Student's t-test (one-tailed), and significant differences from the data of each empty vector are indicated by * (P<0.05).
Klotho expression in elderly ICR mice
The upregulation of miR-29 members was seen in elderly ICR mice as well as klotho(−/−) mice lacking klotho expression; this raised the question of whether klotho might influence the expression of the miR-29 members, although klotho is predominantly expressed in kidney, and hardly expressed in other tissues including heart, lung and liver. We investigated klotho expression levels in elderly and young ICR mice by RT-qPCR. The results indicate that there was no significant difference in the expression of klotho between elderly and young ICR mice (Fig. 4). Therefore, miR-29 expression is probably unaffected by klotho expression levels.
Figure 4. Klotho expression in young and elderly mice.
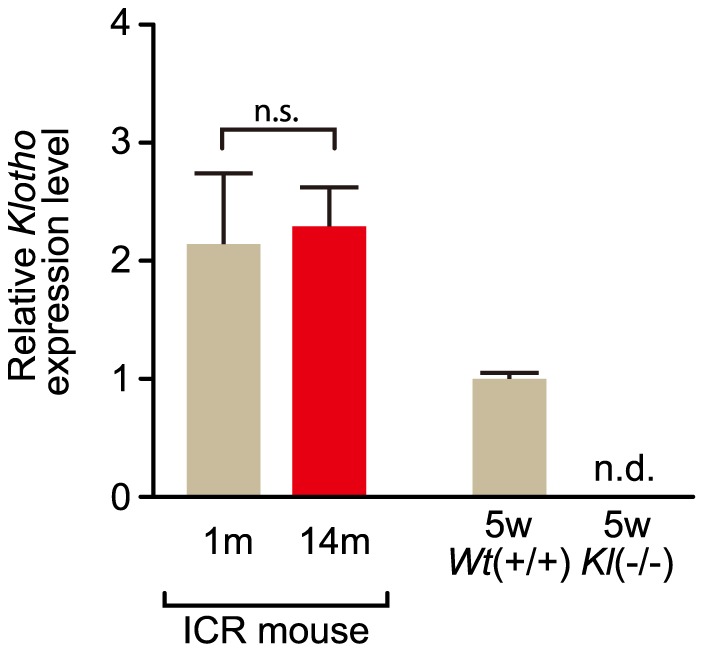
Because klotho is predominantly expressed in kidney [11], klotho expression was examined in this tissue. Total RNAs were extracted from kidney of young (1-month-old: 1 m) and elderly (14-month-old: 14 m) ICR mice and also of 5-week-old (5w) klotho-deficient [Kl(−/−)] and wild-type littermate [Wt(+/+)] mice, and examined by RT-qPCR for klotho and Gapdh. Three measurements by qPCR per each sample were carried out. The expression levels of klotho and Gapdh (as a control) were analyzed by the delta-delta Ct method. Error bars represent standard deviations. n.s., no statistical significance; n.d., no detection.
Target genes of miR-29
Target genes controlled by the miR-29 upregulated in elderly mice may be also involved in natural aging. We searched for candidate gene targets of miR-29 using the TargetScan program (http://www.targetscan.org/vert_40/), and selected the type IV alpha collagen gene family as a potential target. The type IV collagen gene family is composed of the Col4α1, Col4α2, Col4α3, Col4α4, Col4α5 and Col4α6 genes, encoding the α1, α2, α3, α4, α5 and α6 chains of type IV collagen, respectively; and the genes carry putative binding sequences of the miR-29 members in their 3′UTRs. Additionally, previous studies with cultured mammalian cells suggest a significant association between miR-29 members and the type I or type IV collagen genes [18]–[20].
Western blot analyses of type IV collagen indicated that the levels of type IV collagen in heart, lung, liver and kidney of elderly ICR mice was less than that of young mouse tissues (Fig. 5A; upper panel). Interestingly, similar results were also observed between klotho(−/−) and wild-type littermate mice (Fig. 5A; lower panel).
Figure 5. Expression profile of type IV collagen.
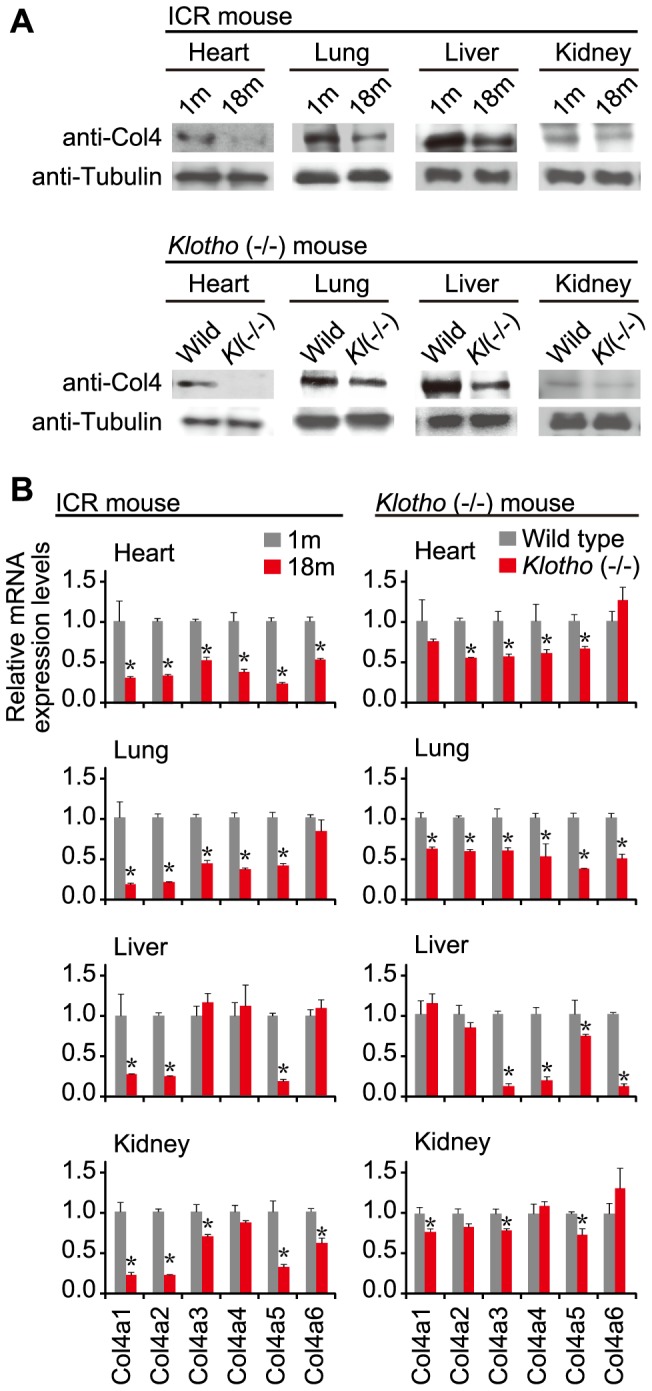
(A) Western blot analysis of type IV collagen. Protein samples prepared from indicated tissues of young (1-month-old: 1 m) and elderly (18-month-old: 18 m) ICR mice and also of klotho(−/−) [Kl(−/−)] and wild-type littermate mice were examined by Western blotting with anti-collagen IV antibody (anti-Col4) and anti-α-tubulin antibody (anti-Tubulin) as an internal loading control. (B) Expression of Col4α1–α6. The expression levels of Col4α1–α6 belonging to the type IV collagen gene family, and Gapdh as a control in indicated tissues of young (1 m) and elderly (18 m) ICR mice, and in klotho(−/−) and wild-type littermate mice were examined by RT-qPCR followed by analyses with the delta Ct method. The levels of Col4α1–α6 expression were normalized to those of Gapdh, and further normalized to the data of young ICR mice or wild-type littermates, each of which was set as 1. Error bars represent standard deviations. The normalized expression data were analyzed by Student's t-test (two-tailed). Significant differences between young (1 m) and elderly (18 m) ICR mice and between klotho(−/−) and wild-type littermate mice are indicated by * (P<0.05).
Next we examined the Col4α1–α6 transcripts in the tissues by means of RT-qPCR. The levels of most of the transcripts in elderly ICR and klotho(−/−) tissues were less than those in young and wild-type littermate tissues, although there were a few exceptions (Fig. 5B). Based on the data, levels of type IV collagen appear to correlate with levels of Col4α1–α6 transcripts.
In vitro gene silencing mediated by synthetic miR-29
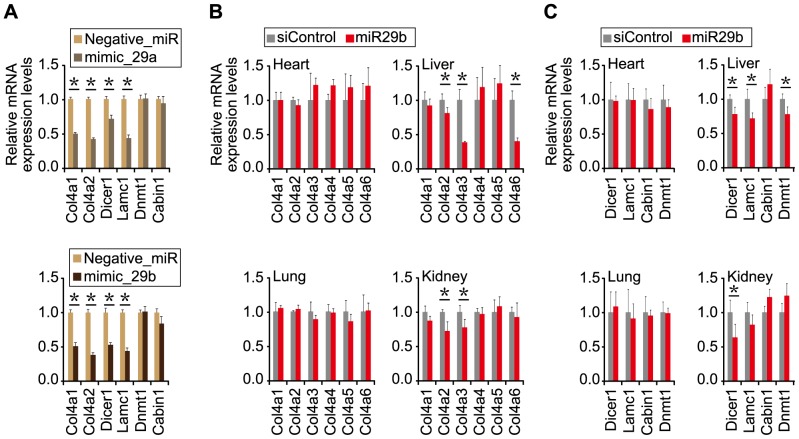
To verify whether the gene silencing mediated by miR-29 could influence the regulation of type IV collagen gene expression, we introduced synthetic miR-29a (miR-29a mimic) or miR-29b (miR-29b mimic) into mouse Neuro2a (N2a) cells and examined their effects on gene silencing against endogenous type IV collagen and other genes (Fig. 6A). RT-qPCR analyses indicated that: (i) levels of Col4α1 and Col4α2 were significantly reduced by introduction of either miR-29a mimic or miR-29b mimic relative to the negative control, (ii) levels of calcineurin binding protein 1 (Cabin1) and DNA methyltransferase 1 (Dnmt1) genes carrying no putative binding sequence of miR-29 remained almost unchanged, and (iii) levels of Dicer1 and laminin, gamma 1 (Lamc1) genes carrying a putative binding site for miR-29 decreased in the presence of the miR-29 mimic. Similar results were also obtained when human embryonic kidney 293 (HEK293) cells were used instead of N2a cells (Fig. S3). Therefore, these findings indicate that the gene silencing mediated by miR-29 suppressed type IV collagen, Dicer1 or Lamc1, each of which carries a putative binding site(s) of miR-29, and that RNA degradation may partially contribute to the gene silencing.
Figure 6. Gene expression profiles in miR-29-treated N2a cells and young ICR mice.
(A) Expression profiles in miR-29-treated N2a cells. MISSION microRNA Mimic (Sigma-Aldrich) has-miR-29a (mimic_29a), has-miR-29b (mimic_29b) or negative control miRNA (negative_miR) was transfected into N2a cells. Three days after transfection, total RNAs were extracted and examined by RT-qPCR followed by analysis using the delta-delta Ct method using the expression levels of Gapdh as a control. The data were further normalized to the data obtained with the negative_miR. Error bars represent standard deviations. The genes examined are indicated: Col4α1, Col4α2, Dicer1, Dnmt1, Lamc1 and Cabin1. The Col4α3–α6 genes were examined as well, but their expression was not detected in the cells. (B, C) Gene expression profiles in miR-29b-treated ICR mice. Normal ICR mice (5-week-old) were subjected to four systemic administrations (Day 1, 4, 8, and 12) of miR-29b duplex or siControl (as a negative control). Two days after the last administration, the treated mice were examined. RT-qPCR analyses were carried out as in A. The examined tissues and genes are indicated: (B) type IV collagen gene family and (C) other genes. Statistically significant decrease in the expression of genes in the miR-29b-treated mice is indicated by * (p<0.05).
In vivo gene silencing mediated by synthetic miR-29b
To further confirm the effects of miR-29 on the expression of its target genes in vivo, we administered a synthetic miR-29b duplex or siControl duplex to 5-week-old (young) ICR mice. After four systemic administrations of the individual small RNA duplexes, the treated ICR mice were examined. RT-qPCR analyses indicated a significant decrease in the expression of some of the Col4α genes in liver and kidney of the miR-29b-treated mice compared with the siControl-treated mice (Fig. 6B). In contrast, no significant difference in the expression was seen in heart and lung between the mice (Fig. 6B); this may be due to the failure of delivery of the small RNA duplexes into these tissues as in the case of using the reporter plasmids described above. A similar decreasing tendency of the expression of Dicer1 and Lamc1 in liver and kidney of the miR-29b-treated ICR mice was also detected (Fig. 6C). Together with the results of Figs. 1 and 3, the findings suggest that the introduced synthetic miR-29b can enhance in vivo gene silencing against its target genes such as Col4α, Dnmt1 and Cabin1 even in young (5-week-old) ICR mice in which the endogenous miR-29 is expressed at a low level. Accordingly, it is conceivable that the miR-29 members spontaneously upregulated in elderly mice may have gradually contributed to such gene silencing (regulation) of target genes.
Plasma biochemical analyses
To examine renal function and oxidative stress in elderly and young ICR mice and also in miR-29b- and siControl-administered ICR mice, we measured the levels of creatinine, hexanoyl-lysine adduct (HEL) and 8-hydroxy-2′-deoxyguanosine (8OH-dG) in plasma: creatinine is an index of renal function, and HEL and 8OH-dG are stress markers. An increasing tendency of either HEL or 8OH-dG in elderly mice compared with young mice was observed (Fig. 7); this may suggest an increase in oxidative stress with increasing age. As for the others, there was no significant difference observed.
Figure 7. Plasma creatinine, HEL, and 8OHdG levels in elderly and young mice and miRNA-treated mice.
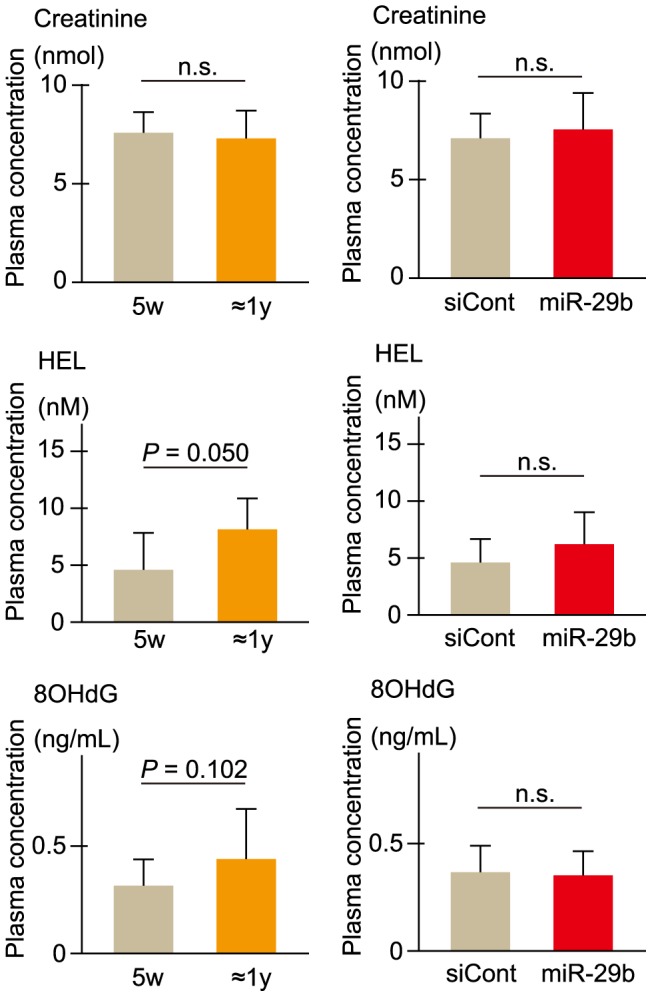
Plasma samples were prepared from young (5-week-old; 5w) and elderly (8∼18-month-old: ≈1 y) ICR mice and also from miR-29b- and siControl (siCont)-treated ICR mice (5-week-old), and then subjected to plasma biochemical tests. The level of creatinine was measured as an index of renal function, and HEL and 8OHdG were each examined as a stress marker. Data are averages of three different individual's measurements. Error bars represent standard deviations. Differences were statistically analyzed by Student's t-test. n.s., no statistical significance.
Discussion
Gene regulation involving miRNAs plays an essential role in various life functions and activities. Senescence is a general deterioration in functioning, in which many biological changes including the alteration of gene expression are presumed to be involved. In this study we examined miRNAs in normal elderly ICR mice as well as in klotho(−/−) mice, a senescence-model animal, and the findings revealed that the miR-29 members were upregulated in both elderly and senescence-model mice. Interestingly, the upregulation of miRNAs was seen in major tissues of elderly ICR as well as klotho(−/−) mice except for the klotho(−/−) brain [10]; thus, miR-29 upregulation may be a common event associated with aging. As for why the klotho(−/−) brain differed from other tissues, this issue remains to be resolved.
In this study, we specifically used outbred ICR mice as a normal mouse strain with natural aging. Since ICR mice possess heterogeneous genomes, ICR may be a suitable lab animal for studying natural aging like the aging of human beings, who also have heterogeneous genetic backgrounds. While there are differences in genetic background between mouse and human, basic genes or gene regulatory systems essential for the fundamental mechanism of natural aging, if any, may be common between them, and such genes or regulatory systems may be capable of becoming useful molecular indicators for studying natural aging. In the current study using outbred ICR mice, we found that the miR-29 members were upregulated in elderly ICR mice although there were more or less individual differences in the degree of their upregulation. The miR-29 upregulated in elderly individuals may be closely related to natural aging, and this raises the possibility that upregulation of miR-29 may be encoded as a genetic program associated with aging.
As with previous studies, our current study indicate that the type IV collagen (Col4α1–α6) genes are potential targets of miR-29. Type IV collagen is localized in the basement membrane and involved in the maintenance of the structure of extracellular matrix. Our current study further indicates that the type IV collagen transcript and protein levels in elderly mouse tissues are less than in young tissues; this may represent weakening of the basement membranes in elderly tissues, and may be compatible with the progression of aging. While, the transcription attenuation of Col4α1–α6 may be a major cause of the reduction of type IV collagen in elderly tissues, our findings suggest another contribution of miR-29 to the suppression of type IV collagen. Thus, miR-29 appears to be involved in the reduction of type IV collagen transcripts and other gene transcripts carrying putative binding sites for miR-29. Taken together, the miR-29 upregulated in elderly tissues may work as a mediator in RNA degradation of target transcripts as well as in translation inhibition, which is the primary function of miRNAs.
Previous in vitro studies using cultured mammalian cells indicated that miR-29 is associated with cellular senescence: the miR-29 members were upregulated during cellular senescence in cultured mammalian cells [21], and appeared to be directly and indirectly engaged in gene regulation of cellular senescence-related B-Myb and p53 genes, respectively [22]–[25]. In the current study, the miR-29-administered N2a and HEK293 cells appeared to be weakened (Fig. 8 and Fig. S4); this might reflect cellular senescence triggered by an increase in the amount of miR-29 in the cells. Taken together, multiple senescence-associated genes may be regulated by the miR-29 members, and a gene regulatory network may participate in the progression or signs of aging.
Figure 8. Cell viability of N2a cells treated with miR-29 mimics.
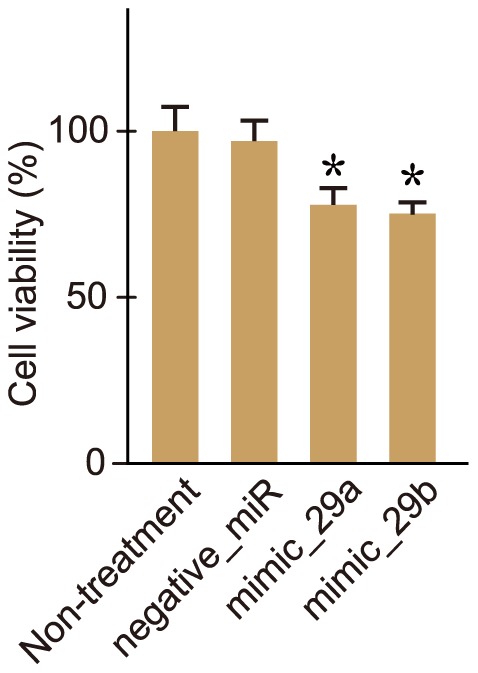
The miR-29 mimics (mimic_29a and mimic_29b) and a negative control miRNA (negative_miR) were transfected into N2a cells as in Fig. 6A. Three days after transfection, cell viability assay was performed. Data are averages of four measurements. Error bars represent standard deviations. Difference between non-treated cells and each of the treated cells was statistically analyzed by ANOVA, followed by the Dunnett's test (* P<0.05).
Finally, miR-29 may be a common miRNA coupled to the progression of aging in various tissues, and may have a key role in senescence. Further studies on the relationship between miR-29 and the aging process need to be carried out. Additionally, confirmatory experiments to see whether human beings deliver similar results need to be performed. If the results are positive, the miR-29 members and also their target Col4α1–α6 genes could become useful molecular markers of general senescence.
Supporting Information
Reproducibility of miRNA expression profiles. Expression profile analyses of miRNAs in indicated tissues of klotho-deficient [klotho(−/−)] and wild-type littermate mice were carried out using the Genopal®-MICM DNA chip, and duplicated with two different individual mice (No. 1 and No. 2). The expression profile data were compared to each other by scatter-plot graphs. The expression data of miRNAs were represented by hybridization signal intensities and indicated by arbitrary units.
(TIF)
Evaluation of in vivo knockdown potency mediated by endogenous miR-29 . The reporter plasmids (Fig. 2A) and β-galactosidase expression plasmid as a control were systemically administered to young (5 week-old; 5w) ICR mice and examined as described in Fig. 3. The normalized levels of luciferase activities in liver and kidney were plotted by scatter graphs. As a result, the data obtained with psiCheck2 (empty vector) as a control appeared to be poorly-reproducible in the liver; in contrast, the kidney data remained stable. Therefore, the evaluation of in vivo knockdown potency mediated by endogenous miR-29 in the liver could not be carried out because the control data varied widely. Averaged values of the data are indicated by bars (−).
(TIF)
Gene expression profiles in miR-29 -treated HEK293 cells. MISSION microRNA Mimics (mimic_29a and mimic_29b) and a negative control miRNA (negative_miR) were transfected into HEK293 cells as in Fig. 6A. Twenty four hours after transfection, total RNAs were extracted and subjected to RT-qPCR followed by analysis using the delta-delta Ct method with the expression level of GAPDH as a control. The data were further normalized to the data obtained with the negative_miR. Data are average of four independent measurements. Error bars represent standard deviations. The genes examined are indicated: COL4A1, COL4A2, DICER1, LAMC1, CABIN1 and DNMT1. Differences between the negative control and mimic_29a or _29b were statistically analyzed by ANOVA, followed by Dunnett's test (* P<0.05). n.s., no statistical significance.
(TIF)
Cell viability of HEK293 cells treated with miR-29 mimics. The miR-29 mimics (mimic_29a and mimic_29b) and a negative control miRNA (negative_miR) were transfected into HEK293 cells as in Figure S3. Three days after transfection, cell viability was examined as in Fig. 8. Data are averages of four measurements. Error bars represent standard deviations. Difference between non-treated cells and each treated cells was statistically analyzed by ANOVA, followed by Dunnett's test (* P<0.05).
(TIF)
Difference in the expression of miRNAs between klotho -deficient and wild-type mice. Hybridization signal intensities obtained from DNA chip analysis [duplicated experiments (Exp.) 1 and 2] are indicated. Fold changes (FC) in the expression of miRNAs between the klotho (Kl) and wild-type (Wt) mice are calculated.
(TIF)
Acknowledgments
We would like to thank H. Oshima for his kind assistance.
Funding Statement
This work was supported by research grants from the Ministry of Health, Labour and Welfare of Japan and by Grants-in-Aid for Scientific Research (B) from the Japan Society for the Promotion of Science. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
- 1. Cheng AM, Byrom MW, Shelton J, Ford LP (2005) Antisense inhibition of human miRNAs and indications for an involvement of miRNA in cell growth and apoptosis. Nucleic Acids Res 33: 1290–1297. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Doench JG, Petersen CP, Sharp PA (2003) siRNAs can function as miRNAs. Genes Dev 17: 438–442. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Krichevsky AM, King KS, Donahue CP, Khrapko K, Kosik KS (2003) A microRNA array reveals extensive regulation of microRNAs during brain development. RNA 9: 1274–1281. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Liu CG, Calin GA, Meloon B, Gamliel N, Sevignani C, et al. (2004) An oligonucleotide microchip for genome-wide microRNA profiling in human and mouse tissues. Proc Natl Acad Sci U S A 101: 9740–9744. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Zeng Y, Yi R, Cullen BR (2003) MicroRNAs and small interfering RNAs can inhibit mRNA expression by similar mechanisms. Proc Natl Acad Sci U S A 100: 9779–9784. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Ohnishi Y, Totoki Y, Toyoda A, Watanabe T, Yamamoto Y, et al. (2010) Small RNA class transition from siRNA/piRNA to miRNA during pre-implantation mouse development. Nucleic Acids Res 38: 5141–5151. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Babak T, Zhang W, Morris Q, Blencowe BJ, Hughes TR (2004) Probing microRNAs with microarrays: tissue specificity and functional inference. RNA 10: 1813–1819. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Lagos-Quintana M, Rauhut R, Yalcin A, Meyer J, Lendeckel W, et al. (2002) Identification of tissue-specific microRNAs from mouse. Curr Biol 12: 735–739. [DOI] [PubMed] [Google Scholar]
- 9. Lee Y, Kim M, Han J, Yeom KH, Lee S, et al. (2004) MicroRNA genes are transcribed by RNA polymerase II. EMBO J 23: 4051–4060. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Eda A, Takahashi M, Fukushima T, Hohjoh H (2011) Alteration of microRNA expression in the process of mouse brain growth. Gene 485: 46–52. [DOI] [PubMed] [Google Scholar]
- 11. Kuro-o M, Matsumura Y, Aizawa H, Kawaguchi H, Suga T, et al. (1997) Mutation of the mouse klotho gene leads to a syndrome resembling ageing. Nature 390: 45–51. [DOI] [PubMed] [Google Scholar]
- 12. Hohjoh H, Fukushima T (2007) Expression profile analysis of microRNA (miRNA) in mouse central nervous system using a new miRNA detection system that examines hybridization signals at every step of washing. Gene 391: 39–44. [DOI] [PubMed] [Google Scholar]
- 13. Hohjoh H, Fukushima T (2007) Marked change in microRNA expression during neuronal differentiation of human teratocarcinoma NTera2D1 and mouse embryonal carcinoma P19 cells. Biochem Biophys Res Commun 362: 360–367. [DOI] [PubMed] [Google Scholar]
- 14. Tamura Y, Yoshida M, Ohnishi Y, Hohjoh H (2009) Variation of gene silencing involving endogenous microRNA in mammalian cells. Mol Biol Rep 36: 1413–1420. [DOI] [PubMed] [Google Scholar]
- 15. Klebe R, Ruddle F (1969) Neuroblastoma: Cell culture analysis of a differentiating stem cell system. J Cell Boil 43: 69a. [Google Scholar]
- 16. Sakai T, Hohjoh H (2006) Gene silencing analyses against amyloid precursor protein (APP) gene family by RNA interference. Cell Biol Int 30: 952–956. [DOI] [PubMed] [Google Scholar]
- 17. Takahashi M, Watanabe S, Murata M, Furuya H, Kanazawa I, et al. (2010) Tailor-made RNAi knockdown against triplet repeat disease-causing alleles. Proc Natl Acad Sci U S A 107: 21731–21736. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Luna C, Li G, Qiu J, Epstein DL, Gonzalez P (2009) Role of miR-29b on the regulation of the extracellular matrix in human trabecular meshwork cells under chronic oxidative stress. Mol Vis 15: 2488–2497. [PMC free article] [PubMed] [Google Scholar]
- 19. Villarreal G Jr, Oh DJ, Kang MH, Rhee DJ (2011) Coordinated regulation of extracellular matrix synthesis by the microRNA-29 family in the trabecular meshwork. Invest Ophthalmol Vis Sci 52: 3391–3397. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Kwiecinski M, Noetel A, Elfimova N, Trebicka J, Schievenbusch S, et al. (2011) Hepatocyte growth factor (HGF) inhibits collagen I and IV synthesis in hepatic stellate cells by miRNA-29 induction. PLoS One 6: e24568. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Martinez I, Cazalla D, Almstead LL, Steitz JA, DiMaio D (2011) miR-29 and miR-30 regulate B-Myb expression during cellular senescence. Proc Natl Acad Sci U S A 108: 522–527. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Masselink H, Vastenhouw N, Bernards R (2001) B-myb rescues ras-induced premature senescence, which requires its transactivation domain. Cancer Lett 171: 87–101. [DOI] [PubMed] [Google Scholar]
- 23. Ben-Porath I, Weinberg RA (2005) The signals and pathways activating cellular senescence. Int J Biochem Cell Biol 37: 961–976. [DOI] [PubMed] [Google Scholar]
- 24. Hara E, Tsurui H, Shinozaki A, Nakada S, Oda K (1991) Cooperative effect of antisense-Rb and antisense-p53 oligomers on the extension of life span in human diploid fibroblasts, TIG-1. Biochem Biophys Res Commun 179: 528–534. [DOI] [PubMed] [Google Scholar]
- 25. Johung K, Goodwin EC, DiMaio D (2007) Human papillomavirus E7 repression in cervical carcinoma cells initiates a transcriptional cascade driven by the retinoblastoma family, resulting in senescence. J Virol 81: 2102–2116. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Reproducibility of miRNA expression profiles. Expression profile analyses of miRNAs in indicated tissues of klotho-deficient [klotho(−/−)] and wild-type littermate mice were carried out using the Genopal®-MICM DNA chip, and duplicated with two different individual mice (No. 1 and No. 2). The expression profile data were compared to each other by scatter-plot graphs. The expression data of miRNAs were represented by hybridization signal intensities and indicated by arbitrary units.
(TIF)
Evaluation of in vivo knockdown potency mediated by endogenous miR-29 . The reporter plasmids (Fig. 2A) and β-galactosidase expression plasmid as a control were systemically administered to young (5 week-old; 5w) ICR mice and examined as described in Fig. 3. The normalized levels of luciferase activities in liver and kidney were plotted by scatter graphs. As a result, the data obtained with psiCheck2 (empty vector) as a control appeared to be poorly-reproducible in the liver; in contrast, the kidney data remained stable. Therefore, the evaluation of in vivo knockdown potency mediated by endogenous miR-29 in the liver could not be carried out because the control data varied widely. Averaged values of the data are indicated by bars (−).
(TIF)
Gene expression profiles in miR-29 -treated HEK293 cells. MISSION microRNA Mimics (mimic_29a and mimic_29b) and a negative control miRNA (negative_miR) were transfected into HEK293 cells as in Fig. 6A. Twenty four hours after transfection, total RNAs were extracted and subjected to RT-qPCR followed by analysis using the delta-delta Ct method with the expression level of GAPDH as a control. The data were further normalized to the data obtained with the negative_miR. Data are average of four independent measurements. Error bars represent standard deviations. The genes examined are indicated: COL4A1, COL4A2, DICER1, LAMC1, CABIN1 and DNMT1. Differences between the negative control and mimic_29a or _29b were statistically analyzed by ANOVA, followed by Dunnett's test (* P<0.05). n.s., no statistical significance.
(TIF)
Cell viability of HEK293 cells treated with miR-29 mimics. The miR-29 mimics (mimic_29a and mimic_29b) and a negative control miRNA (negative_miR) were transfected into HEK293 cells as in Figure S3. Three days after transfection, cell viability was examined as in Fig. 8. Data are averages of four measurements. Error bars represent standard deviations. Difference between non-treated cells and each treated cells was statistically analyzed by ANOVA, followed by Dunnett's test (* P<0.05).
(TIF)
Difference in the expression of miRNAs between klotho -deficient and wild-type mice. Hybridization signal intensities obtained from DNA chip analysis [duplicated experiments (Exp.) 1 and 2] are indicated. Fold changes (FC) in the expression of miRNAs between the klotho (Kl) and wild-type (Wt) mice are calculated.
(TIF)