Abstract
Cooperative activation of actin-myosin interaction by tropomyosin (Tm) is central to regulation of contraction in muscle cells and cellular and intracellular movements in nonmuscle cells. The steric blocking model of muscle regulation proposed 40 y ago has been substantiated at both the kinetic and structural levels. Even with atomic resolution structures of the major players, how Tm binds and is designed for regulatory function has remained a mystery. Here we show that a set of periodically distributed evolutionarily conserved surface residues of Tm is required for cooperative regulation of actomyosin. Based on our results, we propose a model of Tm on a structure of actin-Tm-myosin in the “open” (on) state showing potential electrostatic interactions of the residues with both actin and myosin. The sites alternate with a second set of conserved surface residues that are important for actin binding in the inhibitory state in the absence of myosin. The transition from the closed to open states requires the sites identified here, even when troponin + Ca2+ is present. The evolutionarily conserved residues are important for actomyosin regulation, a universal function of Tm that has a common structural basis and mechanism.
Keywords: actin filament structure, cell motility, cytoskeleton, muscle contraction, motility assay
Actin and myosin are found in almost all eukaryotic cells and are required for diverse cellular functions, including muscle contraction, cell motility, cell adhesion, cytokinesis, and organelle transport (1). The actin–myosin interaction produces two types of movements: force generation between actin filaments leading to contractions, such as in muscle contraction, cell motility, and cytokinesis; and transport of subcellular organelles and macromolecular complexes by myosin motors along actin filaments. The actomyosin contractile apparatus is best characterized in striated muscle contraction, which is regulated in a Ca2+-dependent manner by the proteins, tropomyosin (Tm) and troponin (Tn).
Tm is a universal regulator of the actin cytoskeleton. It is a two-chained, α-helical coiled-coil actin binding protein that associates end-to-end to form a continuous strand along both sides of actin filaments and regulates the functions and stability of actin filaments (2, 3). Tm regulates the interaction of actin filaments with actin binding proteins, including myosin, Tn, ADF-cofilin, Arp2/3, formin, and tropomodulin. Tm can activate or inhibit actomyosin, depending on the myosin and Tm isoforms (4–7). The Tm-dependent regulation is widespread, having been reported in fission yeast and mammalian muscle and nonmuscle systems. The regulation of actomyosin by Tm may be viewed as universal, although there is little understanding of the mechanism or structural basis, the focus of the present work.
In striated muscle, regulation of myosin binding to the thin filament (actin-Tm-Tn) is described by a three-state model, where the three kinetic states are thought to correspond to three positions of Tm on the actin filament (8–10). The three states are defined as: (i) “blocked” or “off” state, in the presence of Tn and absence of Ca2+, where Tm blocks the interaction of myosin with actin; (ii) “closed” state, in the presence of Ca2+ and Tn, where Tm shifts azimuthally on the actin filament to partially expose the myosin binding site and allow weak myosin binding; and (iii) “open” or “on” state, in the presence of myosin heads (subfragment 1 or S1), where Tm azimuthally shifts further on the actin filament, allowing strong myosin binding and force production (11, 12). The population of thin filaments in each state is modulated by the presence of Ca2+ and myosin S1. The binding of myosin S1 to thin filaments shifts the equilibrium of the filaments from the closed to open state, in which both Tm and myosin bind to actin with higher affinity (13–17). This cooperative activation of the thin filament by myosin from the closed to open state can be produced by Tm alone, independent of Tn and Ca2+, and is likely to be a general mechanism of regulation of actomyosin by Tm in nonmuscle cells, where Tn is absent.
Previous studies showed that individual periodic repeats of Tm had varying degrees of effect on thin filament regulation, indicating that specific regions of Tm contribute differently to its regulatory function (18–21). EM and 3D reconstruction of actin-Tm filaments with different Tm isoforms have shown that in the absence of Tn and myosin, Tm occupies the blocked or closed position on the actin filament, depending on the Tm and actin isoform (22). This finding implies that amino acid sequence differences among isoforms can modulate the regulatory functions of Tm. However, the specific residues of Tm that interact with actin in the different regulatory states as well as their effect on myosin regulation are not known.
We previously reported an evolutionary analysis of Tms from 26 species, ranging from cnidarians to vertebrates, and determined the conserved residues of Tm (23). Mutations at conserved surface positions b, c, and f of the heptad repeat of the Tm coiled-coil revealed that residues in the first halves of Tm’s periodic repeats or periods (P) 2–6 of 7 repeats are important for actin affinity. Mutations in the second halves of P2–P6 had a relatively small effect on actin affinity. We hypothesize that these residues are important for another conserved function of Tm, myosin regulation.
In the present study, we determined the effect of Tm mutations at conserved surface positions in the second halves of P2– P6 on myosin regulation using in vitro motility assays. Tm with mutations in P3– P6 showed a large inhibition of filament velocities compared with WT Tm. The period 3 and period 6 mutants inhibited filament velocities, even in the presence of N-ethylmaleimide (NEM)-S1 or Tn + Ca2+. These results, supported by kinetic and fluorescent probe studies, indicate that the mutated residues in second halves of P3– P6 are required for shifting the equilibrium of actin-Tm filaments from closed to open state and may be binding sites for actin or myosin in the open or activated state. A molecular model for interacting sites of Tm with actin and myosin in the open regulatory state is presented to give additional insights into this universal regulatory mechanism.
Results
The striated muscle αTm constructs used in this study have an Ala-Ser extension at the N terminus and have three to four conserved residues mutated to Ala in the second halves of periods P2–P6 (blue in Fig. 1). A first-half mutant, P3-1-EE, was used as a control for the effect of mutations in the first half of the periods (red in Fig. 1). This mutant was selected because it had the smallest effect on actin affinity (23) compared with other first-half mutants, to separate the effects of Tm mutations on myosin regulation from the effect on regulation caused by weaker actin affinity of Tms in the absence of myosin. The mutations in the second half have nonmeasurable or small effects on stability and actin affinity (Table 1) (23), minimizing the concern that a major consequence of the surface mutations is a global effect on the structure and actin binding.
Fig. 1.
Tm mutations at evolutionarily conserved surface residues (23). (A) The rat striated αTm sequence showing conserved b, c, and f residues that were mutated to Ala in P2–P6. Each mutant has three to four mutations within the first-half (red) or second-half (blue) of each period. (B) Tm mutations shown in the 7 Å striated muscle αTm structure (1C1G) (43). Mutations in the first-half of periods 2–6 are in red, and mutations in the second-half are in blue. The periodic repeats do not correspond to the half-turns of the coiled-coil in the 7 Å structure because there are five and three-quarter half-turns of the supercoil per molecule, not seven (43, 44). That is, the half-turns do not correspond to the seven actins along the length of one Tm molecule.
Table 1.
Actin affinity, TM from ellipticity at 222 nm (CD), filament speeds and kinetic data of Tm mutants
| Mutant | Kapp† (×106 M−1) | TM† (°C) | Speed‡ (μm/s), (% moving) | kfast(max)§ (s−1) | kfast(max)/Kact,fast§ (μM−1⋅s−1) | kslow(max)§ (s−1) | kslow(max)/Kact,slow§ (μM-1⋅s−1) |
| Actin | — | — | 4.7 ± 0.2, n = 8, (95) | 178.2 ± 29 | 4.6 ± 0.6 | 11.4 ± 1.1 | 1.2 ± 0.3 |
| WT | 2.1 ± 0.1 | 49.0 | 3.1 ± 1.0*, n = 8, (90) | 139.0 ± 16 | 1.7 ± 0.1 | 8.4 ± 1.0 | 0.6 ± 1.1 |
| P2-2-EKD | 2.6 ± 0.2 | 49.0 | 2.4 ± 0.6, n = 4, (78) | — | — | ||
| P3-1-EE | 1.2 ± 0.1 | 49.0 | 3.5 ± 0.4, n = 2, (96) | — | — | ||
| P3-2-EKDE | 1.1 ± 0.1 | 49.2 | 1.0 ± 0.6**, n = 4 (40), | 129.5 ± 53 | 0.9 ± 0.1 | 4.8 ± 0.7 | 0.3 ± 0.1 |
| P4-2-EEEE | 0.9 ± 0.1 | 47.6 | 1.7 ± 0.7***, n = 3, (68) | — | — | ||
| P5-2-EKNK | 1.3 ± 0.1 | 49.2 | 1.3 ± 0.1**, n = 3, (54) | — | — | ||
| P6-2-KEE | 1.9 ± 0.1 | 48.3 | 0.6 ± 0.5**, n = 4 (29), | 202.7 ± 60 | 1.1 ± 0.1 | 6.8 ± 0.6 | 0.3 ± 0.1 |
†The values for the binding constant of Tm to actin, Kapp, shown with SE, and TM, the temperature at which the ellipticity at 222 nm, normalized to a scale of 0–1, is equal to 0.5, from Barua et al. (23).
‡Filament speeds ± SD of actin and actin-Tm from in vitro motility assays in the absence of NEM-S1 and Tn.
*P < 0.005 compared with actin, **P < 0.005 compared with WT Tm, ***P < 0.05 compared with WT Tm (unpaired Student t test) (Fig. 2A). The average values for the percent moving filaments are in parentheses. The total number of filaments ranged between 140 and 290 in all experiments.
§kfast(max) and kslow(max) ± SE are the fast and slow components of the maximal rates of product dissociation at saturating actin concentration, and kfast(max)/Kact,fast and kslow(max)/Kact,slow ± SE are the second-order rate constants of the fast and slow components of rates of product dissociation (Fig. 3).
In Vitro Motility of Actin-Tm.
In vitro motility assays were carried out in the absence of Tn and Ca2+ to determine the effects of Tm mutations on myosin-dependent regulation. A myosin surface density (40 µg/mL) was selected that allows maximal velocity of unregulated actin filaments (Fig. S1). A weighted probability of the filament velocity for hundreds of events was fitted to a Gaussian distribution to determine a mean velocity ± SD for each experimental condition (SI Materials and Methods and Fig. S2). The average of the velocities obtained from two or more independent experiments is reported (Fig. 2 and Table 1). The velocity of actin-Tm(wild-type) [A-Tm(WT)] filaments was lower by ∼40% relative to actin alone, indicating inhibition (P < 0.005) (black bars, Fig. 2A). The inhibition by A-Tm(WT) is consistent with previous reports (24, 25) and with kinetics and EM structural evidence that Tm is predominantly in the closed state in the absence of Tn (11, 22). The velocities of A-Tm(mutant) filaments, P2-2-EKD and P3-1-EE, were not significantly different from A-Tm(WT) (P > 0.05). The velocities of all other A-Tm(mutant) filaments were inhibited by ∼40–80% relative to A-Tm(WT) (P < 0.05) (Fig. 2A and Table 1). P3-2-EKDE and P6-2-KEE showed the largest inhibition. The fraction of filaments moving varied from 30–80% for the Tm second-half mutants compared with 90–95% for WT Tm and unregulated actin, consistent with inhibition of motility of A-Tm(mutant) filaments (Table 1).
Fig. 2.
Filament speeds of actin and actin-Tm in in vitro motility assays. (A) Filament speeds in the absence of NEM-S1 and Tn (black bars) (*P < 0.005 compared with actin, **P < 0.005 and ***P < 0.05 compared with WT Tm, unpaired Student t test); in the presence of NEM-S1 (white bars); and in the presence of Tn + Ca2+ (gray bars). (B and C) Filament speeds at increasing concentrations of NEM-S1. The values are mean ± SD from two to eight experiments (Table 1 and SI Materials and Methods). The data with no error bars are from one experiment. An antimyosin subfragment 2 monoclonal antibody was bound to nitrocellulose-coated glass cover-slips before incubation with chicken skeletal myosin (40 μg/mL) at 4 °C for 2 h. The myosin was subjected to “dead-head” removal before incubation (Fig. S1). The cover-slips were transferred to 15-μL drops of 2 nM rhodamine-phalloidin–labeled chicken skeletal actin or actin-Tm (1 µM Tm) or actin-Tm-Tn (1 µM Tm, 1.3 µM Tn) in motility buffer in a small parafilm ring fixed on an alumina slide. Movement of actin filaments from 1 to 2 min of continuous video were recorded from several fields for each experiment and analyzed with semiautomated filament tracking programs (SI Materials and Methods). NEM-S1 was added to actin-Tm at the indicated concentrations. The Tn + Ca2+ experiments were in the presence of 0.2 mM CaCl2. The Tn − Ca2+ experiments were in the presence of 0.2 mM EGTA and showed complete inhibition of motion (Table S1). Assay conditions: 25 mM imidazole, pH 7.6, 25 mM KCl, 4 mM MgCl2, 7.6 mM MgATP, 50 mM DTT, 0.5% methyl cellulose, and an oxygen scavenger system (0.1 mg/mL glucose oxidase, 0.02 mg/mL catalase, 2.5 mg/mL glucose); Temperature: 27 °C.
Activation of Motility by NEM-S1.
Because the velocity of A-Tm filaments was inhibited >40% for most Tm constructs compared with actin alone, we carried out motility assays in the presence of NEM-S1, a mimic of the rigor complex between actin and myosin S1, known to activate regulated thin filaments (26–28). NEM-S1 binds tightly and activates thin filaments in the presence of MgATP. The addition of NEM-S1 increased the velocity of WT and mutant A-Tm filaments, illustrating activation (Fig. 2 and Table S1). The dependence of filament velocities on NEM-S1 concentration varied according to the mutant. A-Tm(WT) was activated at NEM-S1 concentrations ≥200 nM, with a velocity equivalent to the velocity of unregulated actin in the absence of NEM-S1, indicating full activation (Fig. 2 A and B). The P2-2-EKD, P3-1-EE, and P4-2-EEEE mutants also showed maximal or near maximal activation at NEM-S1 concentrations ≥200 nM (Fig. 2 B and C). The P3-2EKDE, P5-2-EKNK, and P6-2-KEE mutants reached maximal or near maximal activation at higher NEM-S1 concentrations (≥400 nM), inferring that these mutants require more NEM-S1 for activation, compared with WT (Fig. 2 B and C). The P6-2-KEE mutant was not fully activated within the range of NEM-S1 concentrations used in our experiments (Fig. 2C). The velocities of unregulated actin filaments were competitively inhibited ∼40% by binding of the inactive NEM-S1 heads, as were P3-1-EE and P6-2-KEE at higher NEM-S1 concentrations (Fig. 2 B and C). We conclude that the conserved surface residues in the second halves of P3–P6 are required for activation of actin-Tm filaments by myosin, and regulation of myosin binding to the thin filament is not related to affinity of Tm to actin in the absence of myosin.
In Vitro Motility with Troponin + Ca2+.
In the presence of Tn and Ca2+, the velocity of A-Tm(WT), A-Tm(P2-2-EKD), and A-Tm(P3-1-EE) filaments increased by about 40% relative to the velocity with no Tn, indicating Ca2+-dependent activation (gray bars, Fig. 2A). The velocities of the P3-2-EKDE, P4-2-EEEE, P5-2-EKNK, and P6-2-KEE mutants were 40–80% inhibited compared with WT in the presence of Tn + Ca2+ (Table S1). Of these mutants, Tn + Ca2+ had no effect on the velocity of P3-2-EKDE or P4-2-EEEE, compared with the velocity in the absence of Tn (Fig. 2A). We conclude that the inhibition of velocity by the Tm mutants observed in the motility assays is independent of the presence of Tn + Ca2+, and the mutations do not affect Ca2+-dependent regulation. The results are consistent with the prevalent view that the cooperativity of the activation of the regulated thin filament (actin-Tm-Tn) primarily depends on Tm (28–30). In the presence of Tn without Ca2+, WT and mutant Tms showed complete inhibition of motion (Table S1), indicating normal relaxation of the thin filament and intact Tn-binding sites on Tm.
Kinetics of Phosphate Dissociation from Actomyosin-ADP-Pi.
A possible mechanism for inhibition of filament velocity by Tm mutants may be through inhibition of the actomyosin ATPase cycle, either by changing the rate of product dissociation or the affinity of myosin for the actin filament. To test this, we determined the effect of the P3-2-EKDE and P6-2-KEE mutants, which showed the largest inhibition of filament velocity in motility assays, on the rate of product dissociation from actomyosin. We measured the rate of dissociation of the fluorescent ADP analog, deoxymantADP (mdADP), from actomyosin-S1-mdADP-phosphate (AM-mdADP-Pi) (12). The rate of dissociation is biphasic, with an initial rapid decrease in fluorescence followed by a slower decrease. For skeletal actomyosin, the rate of mdADP dissociation is fast and the rapid fluorescence phase is limited by and provides a measure of the rate of dissociation of Pi.
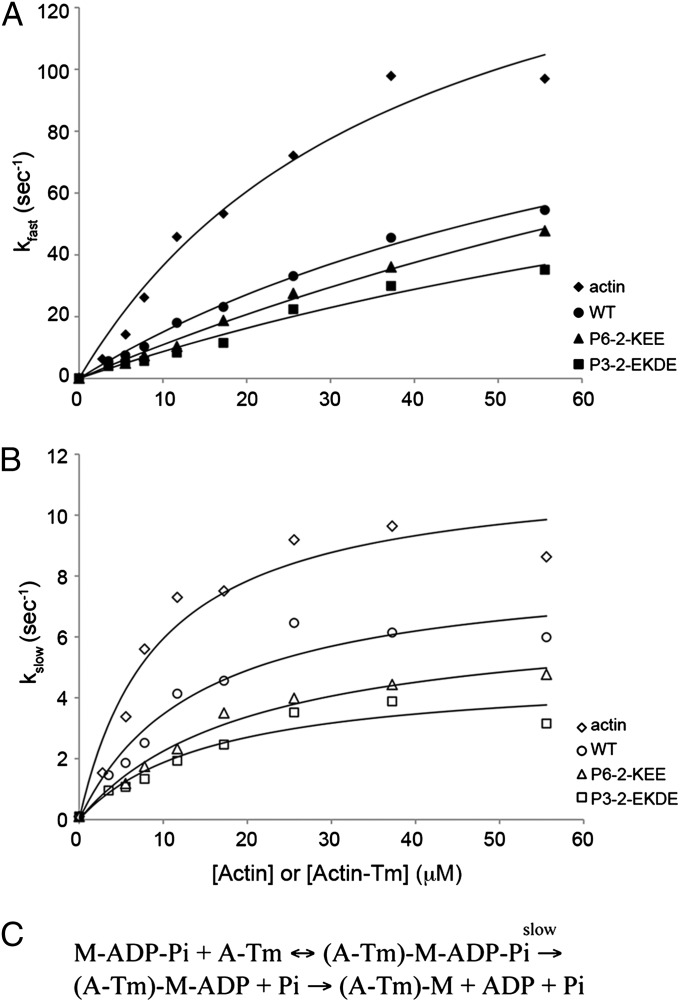
We measured the dependence of observed rates of the fast (kfast) and slow (kslow) components of mdADP dissociation upon actin or actin-Tm (WT, P3-2-EKDE, or P6-2-KEE) concentration (Fig. 3 A and B). The actin concentration dependence of the rates was fit to a hyperbola that gives maximum rates of product dissociation at saturating actin concentration [kfast(max), kslow(max)] and second-order rate constants [kfast(max)/Kact,fast and kslow(max)/Kact,slow] of M-ADP-Pi binding to A-Tm followed by product dissociation, as shown in Fig. 3C (Table 1). Kact,fast and Kact,slow are the actin concentrations required to obtain half the maximal observed rates. There is a 2-fold decrease in the second order rate constant of the fast phase for A-(P3-2-EKD) and a 1.5-fold decrease for A-(P6-2-KEE) compared with A-(WT) (Table 1). This finding implies that the inhibition of velocity shown by P3-2-EKDE and P6-2-KEE may be attributed to a slower rate of product dissociation compared with WT, which could either be because of a slower rate of Pi dissociation or weaker binding of M-ADP-Pi to A-Tm. The rate of these steps is threefold slower for A-(WT) compared with actin alone, consistent with the inhibition of velocity of A-(WT) compared with unregulated actin.
Fig. 3.
Rate of dissociation of mdADP from actin-myosin-S1-mdADP-Pi or actin-Tm-myosin-S1-mdADP-Pi. Double mixing stopped-flow experiments were performed as described in SI Materials and Methods. 6 μM myosin-S1 and 4 µM mdADP were mixed, incubated for 1 s, and then mixed with actin or actin-Tm and 1 mM ATP. The actin-Tm complexes were preformed at molar ratios (Tm:actin) of 1:5 for WT and P6-2-KEE, and 1:2.5 for P3-2-EKDE, to ensure that the actin was saturated with Tm. (A) Dependence of kfast on actin or actin-Tm concentration. The data were fit to the equation: kfast = kfast(max)/(1 + Kact,fast/[actin]) + k0 to obtain values for kfast(max) and kfast(max)/Kact,fast (Table 1). (B) Dependence of kslow on actin or actin-Tm concentration. The data were fit to the equation: kslow = kslow(max)/(1 + Kact,slow/[actin]) + k0 to obtain values for kslow(max) and kslow(max)/Kact,slow (Table 1). k0 = 0.1 s−1 was measured in the absence of actin. kfast is associated with the attachment of actin to myosin-ADP-Pi (M-ADP-Pi), followed by product dissociation, and kslow is associated with the attachment of actin to myosin-ATP, followed by hydrolysis and product dissociation. (C) Scheme showing the steps of the actomyosin ATPase cycle, with M-ADP-Pi binding to A-Tm, followed by product dissociation. Assay conditions: 5 mM MOPS, pH 7, 2 mM MgCl2, 1 mM DTT, 20 mM KAc. Temperature: 20 °C.
Excimer Fluorescence and Light-Scattering of Actin-[N-(1-pyrenyl)iodoacetamide]-Tm in the Presence of Myosin S1.
To determine the cooperativity of the transition of actin-Tm filaments from the closed to open state by myosin S1, we measured the excimer fluorescence and light scattering (LS) intensity of actin-(PIA-Tm) during titrations with myosin S1 in the absence of nucleotide. The increase in excimer fluorescence of Tm labeled with [N-(1-pyrenyl)iodoacetamide] at Cys190 (PIA-Tm) reports a myosin S1-induced change in state of Tm in actin-(PIA-Tm) filaments and is interpreted as a measure of the fraction of actin-(PIA-Tm) in the S1-induced open state (31). The increase in LS intensity measures the amount of S1 bound to actin-(PIA-Tm) and saturates at a 1:1 stoichiometry. However, the increase in excimer fluorescence change is complete before the actin filament is saturated with S1, providing a measure of the regulatory unit that responds to S1 binding (Fig. 4 and Fig. S3). The calculated apparent cooperative unit size (n) is the average number of actin subunits trapped in the open state by the binding of one S1 (32). For actin-(PIA-WT), the half-maximal excimer fluorescence change corresponds to a S1 occupancy of about 0.1 or 1 S1 bound per seven actin subunits and n = 6–7, as previously reported (32). The apparent cooperative unit for actin-(PIA-P3-2-EKDE) is 2, corresponding to a S1 occupancy of 0.3–0.4 or 2–3 S1s bound per seven actin subunits. A smaller functional unit may make it more difficult for myosin S1 to switch actin-(PIA-P3-2-EKDE) filaments to the open state, causing inhibition of velocity, as seen in the motility assays. Actin-(PIA-P6-2-KEE) was similar to actin-(PIA-WT) (n = 6–7), inconsistent with the results of the motility and kinetic analyses. Given a poor understanding of the mechanism of thin filament cooperativity, we do not know if the discrepancies reflect a local versus a global response of the probe or directionality in the initiation and communication of information in the thin filament.
Fig. 4.
Excimer fluorescence and LS change of actin-(PIA-Tm) during titrations with myosin S1 in the absence of nucleotide. 3 μM F-actin and 0.5 µM PIA-Tm were combined and titrations carried out by addition of myosin S1 and monitoring the LS at 350 nm and the PIA excimer fluorescence at 485 nm after each addition of S1 in the same cuvette. The fraction S1 bound was calculated from the change in LS, and the fractional fluorescence change was calculated from the change in excimer fluorescence, normalized relative to the values at saturation (Fig. S3). The binding of S1 to actin-(PIA-Tm) saturates at a 1:1 stoichiometry of S1 to actin, but the excimer fluorescence change of PIA in response to S1 binding is complete before the actin filament is saturated with S1. The data for each Tm are from two independent experiments. The apparent cooperative unit size (n) was obtained by comparing the experimental curves with model curves for different n (dashed lines), using the equation, fopen = 1−(1 − fbound)n (32). Assay conditions: 25 mM imidazole, pH 7.6, 25 mM KCl, 4 mM MgCl2, 5 mM DTT.
Discussion
We have shown that mutation of evolutionarily conserved surface residues in the second halves of Tm periodic repeats 3–6 affects myosin regulation of the actin filament and the cooperative switch to the open state. We suggest that each of Tm’s periodic repeats includes a site in the first half for actin binding in the closed state and a site in the second half for cooperative activation by myosin to the open state. The extent of contribution to these two functions varies from site to site. In vitro motility assays showed that the velocities of actin-Tm(mutant) filaments were inhibited compared with actin-Tm(WT), inferring that the conserved surface residues in the second-half of periods are required for actomyosin regulation. The P2-2-EKD mutant was an exception to this general mechanism. Mutations in periods 3 and 6 (P3-2-EKDE and P6-2-KEE) showed the largest inhibition (≥80%) of filament velocity compared with WT, even in the presence of Tn + Ca2+. The P6-2-KEE mutant did not fully activate filament velocity even in the presence of NEM-S1. The rate of product dissociation from actomyosin was twofold slower for P3-2-EKDE and 1.5-fold slower for P6-2-KEE, compared with WT Tm, which may explain the inhibition of velocity shown by these mutants in the motility assays. The inhibition of regulation showed by the Tm mutants in motility assays is independent of the presence of Tn, indicating the underlying mechanism of myosin regulation primarily depends on Tm. Mutations in different Tm periods may affect different aspects of the regulatory mechanism. Our results also show that specific residues of Tm are involved in regulation, because mutation of residues in P2-2-EKD and P3-1-EE mutants did not affect motility. The results agree with previous studies reporting that deletion of P3–P6 had an inhibitory effect on actomyosin (18, 19) and that P3 was important for the cooperative activation of the actin filament (20, 21).
The inhibition of myosin regulation by Tm mutants suggests that the mutated residues in P3–P6 are required to shift the equilibrium of actin-Tm filaments to the open state and may be binding sites for actin and myosin in the open or activated state. The results provide a basis for establishing models for binding of Tm to the actin filament in the different regulatory states and the residues involved in switching it from one state to another. We have constructed a molecular model of Tm bound to F-actin in the open state by docking high-resolution Tm structures (33, 34) to a proposed binding site in a 8 Å rigor actin-Tm-myosin S1 complex determined by cryo-EM (10). Because of the necessity of applying the helical symmetry of actomyosin to achieve maximum resolution, the molecular ends of Tm as well as the axial and rotational position of Tm with respect to actomyosin are not defined in the EM structure. The Tm molecule in the pseudoatomic model of the rigor actin-Tm-myosin S1 complex showed electrostatic interactions of Tm residues in the first half of P5 with both actin and myosin (10), and fewer interactions of Tm residues in the second-half of the period (Fig. 5A). However, based on experimental data from the present study, first-half residues (P3-1-EE) do not affect actomyosin regulation. Therefore, we made a minor modification in the axial and rotational alignment of the Tm in the model [Protein Data Bank (PDB) ID 4A7F] by positioning high-resolution crystal structures of Tm fragments (PDB ID 2B9C and 2D3E) that include P3–P6 to optimize electrostatic interactions of the mutated residues in the second-half with charged residues on actin and myosin (Fig. 5 B and C, and Movie S1), but maintaining the azimuthal alignment of Tm on actin. Based on this model, the conserved residues in Tm mutated in this study can interact with K238, E226, and D311 of actin, and D312, K295, and K300 of myosin (Fig. 5 B and C). Earlier studies have indicated the importance of actin residues K238 and D311 in Tm binding and regulation (35–38). Direct interactions of Tm with both actin and myosin has also been inferred by earlier EM structures and supported by biochemical studies (8, 9, 39).
Fig. 5.
Molecular model for actin-Tm-myosin interaction in the open state. The model was constructed by docking high-resolution Tm crystal structures (33, 34) in a 8 Å rigor actin-Tm-myosin S1 complex determined by cryo-EM (10). A minor modification was made in the axial and rotational alignment of Tm in the EM model by positioning crystal structures of Tm fragments to optimize electrostatic interactions of the mutated Tm residues in second-half of P3–P6 with charged residues on actin and myosin, while maintaining the azimuthal alignment. The radial distance between Tm and F-actin is 40 Å (10). Interactions of Tm with actin (green) and myosin (red) are shown in P3–P6: (A) EM structure of actin-Tm-myosin S1 (PDB ID 4A7F) (10) showing few interactions of the mutated Tm residues in P3–P4 with actin and myosin, and (B and C) crystal structure of Tm fragments including P3–P4 (PDB ID 2B9C) (B), and P5–P6 (PDB ID 2D3E) (C) (33, 34) positioned in the EM structure of actin-Tm-myosin S1 (PDB ID 4A7F), showing more optimal electrostatic interactions of mutated Tm residues with actin and myosin (Movie S1). The Tm molecule in the EM structure is shown in gray and the crystal structures of Tm that were positioned in the EM structure are shown in blue (B and C). The side-chains of the mutated Tm residues are shown in white and have blue labels. The side-chains of actin and myosin are labeled as A and M, respectively. Based on this model, in P3-2-EKDE (B), residues E114 and K118 can interact with K238, E226, D311 of actin, and K118, D121, and E122 can interact with D312, K295, K300 of myosin. In P4-2-EEEE (B), E150 and E156 can interact with K238 of actin, and E156, E163, and E164 with K295, K300 of myosin. E195 and K198 in P5-2-EKNK (C), and K233, E234, and E240 in P6-2-KEE (C) can interact with K238, E226, and D311 of actin and D312, K295, and K300 of myosin. The model was constructed using the University of California at San Francisco Chimera package (45), and is provided as a structural model for discussion of the results.
In our model of the closed state and others, the conserved residues in the first halves of Tm’s periods (red in Fig. 1) are close to actin residues D25, K326, K328, and P333 (23, 33, 40). The weak but specific electrostatic interactions with actin place Tm in the closed position in the absence of myosin, partially occupying the myosin binding site on actin. This position is consistent with Tm’s ability to inhibit activation of myosin by actin at low myosin concentrations (14, 29). Higher concentrations of myosin cross-bridges in the strong binding state compete with the weak Tm binding for actin and shift the Tm to the open state. In the open state, as seen in our modification of the rigor actin-Tm-myosin structure, residues in Tm’s second set of sites (blue in Fig. 1) are positioned to interact with both actin and myosin, as shown in Fig. 5 B and C. The multiple weak but specific electrostatic interactions between Tm, actin, and myosin allow dynamic regulation of the thin filament that must change its functional state rapidly and in a cooperative manner. The mutations in the first and second halves of periods also approximately correspond to the α- and β-bands of Tm that have been proposed to be actin binding sites in the inactivated and activated states, respectively (41).
In conclusion, evolutionarily conserved surface residues of Tm in the second halves of P3–P6 are important for actomyosin regulation. We have also shown that specific residues of Tm are involved in regulation and different periods have different effects on regulation. Weak electrostatic interactions of Tm with both actin and myosin coupled with multiple binding sites and end-to-end association of Tm result in a strong apparent actin affinity of both Tm and myosin in the active state, and cooperative activation that extends the length of the thin filament. Even with Tn + Ca2+, the Tm mutations impair the switch to open state, inferring that although Ca2+ is the trigger for activation, it is the properties of Tm (and myosin) that determine cooperative activation of the thin filament. This finding is consistent with reports that show that Tm can activate or inhibit actomyosin, depending on the Tm and myosin isoforms (4–7). The transition from the closed to open state requires the conserved Tm residues, even when Tn + Ca2+ is present. The evolutionarily conserved sites are important for actomyosin regulation, a universal function of Tm that has a conserved structural basis and mechanism.
Materials and Methods
The Tm mutants used in this study have been described previously (23). Details about actin, troponin, myosin, and myosin S1 are provided in SI Materials and Methods. The in vitro motility assays were done as in Winkelmann et al. (42), stopped-flow kinetics experiments were done as in Heeley et al. (12), and the excimer fluorescence and light-scattering experiments were done as in Ishii and Lehrer (31) (SI Materials and Methods).
Supplementary Material
Acknowledgments
We thank Brinda Desai, Yazan Alkhawam, and Richard Trent for technical assistance, and Stefan Raunser for providing the article (10) and coordinates of the rigor actin-Tm-myosin S1 complex prior to publication. This work was supported by National Institutes of Health Grant GM093065 (to S.E.H.-D.). We acknowledge the late Annemarie Weber's (1923-2012) discoveries that Ca2+ and myosin crossbridges can cooperatively activate the thin filament, which are a basis of our work.
Footnotes
The authors declare no conflict of interest.
This article is a PNAS Direct Submission.
This article contains supporting information online at www.pnas.org/lookup/suppl/doi:10.1073/pnas.1212754109/-/DCSupplemental.
References
- 1.Pollard TD, Cooper JA. Actin, a central player in cell shape and movement. Science. 2009;326(5957):1208–1212. doi: 10.1126/science.1175862. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Wang CL, Coluccio LM. New insights into the regulation of the actin cytoskeleton by tropomyosin. Int Rev Cell Mol Biol. 2010;281:91–128. doi: 10.1016/S1937-6448(10)81003-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Gunning P, O’Neill G, Hardeman E. Tropomyosin-based regulation of the actin cytoskeleton in time and space. Physiol Rev. 2008;88(1):1–35. doi: 10.1152/physrev.00001.2007. [DOI] [PubMed] [Google Scholar]
- 4.Clayton JE, Sammons MR, Stark BC, Hodges AR, Lord M. Differential regulation of unconventional fission yeast myosins via the actin track. Curr Biol. 2010;20(16):1423–1431. doi: 10.1016/j.cub.2010.07.026. [DOI] [PubMed] [Google Scholar]
- 5.Ostap EM. Tropomyosins as discriminators of myosin function. Adv Exp Med Biol. 2008;644:273–282. doi: 10.1007/978-0-387-85766-4_20. [DOI] [PubMed] [Google Scholar]
- 6.Fanning AS, Wolenski JS, Mooseker MS, Izant JG. Differential regulation of skeletal muscle myosin-II and brush border myosin-I enzymology and mechanochemistry by bacterially produced tropomyosin isoforms. Cell Motil Cytoskeleton. 1994;29(1):29–45. doi: 10.1002/cm.970290104. [DOI] [PubMed] [Google Scholar]
- 7.Lehrer SS, Morris EP. Comparison of the effects of smooth and skeletal tropomyosin on skeletal actomyosin subfragment 1 ATPase. J Biol Chem. 1984;259(4):2070–2072. [PubMed] [Google Scholar]
- 8.Vibert P, Craig R, Lehman W. Steric-model for activation of muscle thin filaments. J Mol Biol. 1997;266(1):8–14. doi: 10.1006/jmbi.1996.0800. [DOI] [PubMed] [Google Scholar]
- 9.Milligan RA, Flicker PF. Structural relationships of actin, myosin, and tropomyosin revealed by cryo-electron microscopy. J Cell Biol. 1987;105(1):29–39. doi: 10.1083/jcb.105.1.29. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Behrmann E, et al. Structure of the rigor actin-tropomyosin-myosin complex. Cell. 2012;150(2):327–338. doi: 10.1016/j.cell.2012.05.037. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.McKillop DF, Geeves MA. Regulation of the interaction between actin and myosin subfragment 1: Evidence for three states of the thin filament. Biophys J. 1993;65(2):693–701. doi: 10.1016/S0006-3495(93)81110-X. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Heeley DH, Belknap B, White HD. Maximal activation of skeletal muscle thin filaments requires both rigor myosin S1 and calcium. J Biol Chem. 2006;281(1):668–676. doi: 10.1074/jbc.M505549200. [DOI] [PubMed] [Google Scholar]
- 13.Greene LE, Eisenberg E. Cooperative binding of myosin subfragment-1 to the actin-troponin-tropomyosin complex. Proc Natl Acad Sci USA. 1980;77(5):2616–2620. doi: 10.1073/pnas.77.5.2616. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Lehrer SS, Morris EP. Dual effects of tropomyosin and troponin-tropomyosin on actomyosin subfragment 1 ATPase. J Biol Chem. 1982;257(14):8073–8080. [PubMed] [Google Scholar]
- 15.Eaton BL. Tropomyosin binding to F-actin induced by myosin heads. Science. 1976;192(4246):1337–1339. doi: 10.1126/science.131972. [DOI] [PubMed] [Google Scholar]
- 16.Trybus KM, Taylor EW. Kinetic studies of the cooperative binding of subfragment 1 to regulated actin. Proc Natl Acad Sci USA. 1980;77(12):7209–7213. doi: 10.1073/pnas.77.12.7209. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Bremel RD, Weber A. Cooperation within actin filament in vertebrate skeletal muscle. Nat New Biol. 1972;238(82):97–101. doi: 10.1038/newbio238097a0. [DOI] [PubMed] [Google Scholar]
- 18.Landis C, Back N, Homsher E, Tobacman LS. Effects of tropomyosin internal deletions on thin filament function. J Biol Chem. 1999;274(44):31279–31285. doi: 10.1074/jbc.274.44.31279. [DOI] [PubMed] [Google Scholar]
- 19.Hitchcock-DeGregori SE, Song Y, Greenfield NJ. Functions of tropomyosin’s periodic repeats. Biochemistry. 2002;41(50):15036–15044. doi: 10.1021/bi026519k. [DOI] [PubMed] [Google Scholar]
- 20.Kawai M, Lu X, Hitchcock-Degregori SE, Stanton KJ, Wandling MW. Tropomyosin period 3 is essential for enhancement of isometric tension in thin filament-reconstituted bovine myocardium. J Biophys. 2009;2009:380967. doi: 10.1155/2009/380967. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Oguchi Y, Ishizuka J, Hitchcock-DeGregori SE, Ishiwata S, Kawai M. The role of tropomyosin domains in cooperative activation of the actin-myosin interaction. J Mol Biol. 2011;414(5):667–680. doi: 10.1016/j.jmb.2011.10.026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Lehman W, et al. Tropomyosin and actin isoforms modulate the localization of tropomyosin strands on actin filaments. J Mol Biol. 2000;302(3):593–606. doi: 10.1006/jmbi.2000.4080. [DOI] [PubMed] [Google Scholar]
- 23.Barua B, Pamula MC, Hitchcock-DeGregori SE. Evolutionarily conserved surface residues constitute actin binding sites of tropomyosin. Proc Natl Acad Sci USA. 2011;108(25):10150–10155. doi: 10.1073/pnas.1101221108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Fraser ID, Marston SB. In vitro motility analysis of actin-tropomyosin regulation by troponin and calcium. The thin filament is switched as a single cooperative unit. J Biol Chem. 1995;270(14):7836–7841. doi: 10.1074/jbc.270.14.7836. [DOI] [PubMed] [Google Scholar]
- 25.VanBuren P, Palmiter KA, Warshaw DM. Tropomyosin directly modulates actomyosin mechanical performance at the level of a single actin filament. Proc Natl Acad Sci USA. 1999;96(22):12488–12493. doi: 10.1073/pnas.96.22.12488. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Nagashima H, Asakura S. Studies on co-operative properties of tropomyosin-actin and tropomyosin-troponin-actin complexes by the use of N-ethylmaleimide-treated and untreated species of myosin subfragment 1. J Mol Biol. 1982;155(4):409–428. doi: 10.1016/0022-2836(82)90479-x. [DOI] [PubMed] [Google Scholar]
- 27.Pemrick S, Weber A. Mechanism of inhibition of relaxation by N-ethylmaleimide treatment of myosin. Biochemistry. 1976;15(23):5193–5198. doi: 10.1021/bi00668a038. [DOI] [PubMed] [Google Scholar]
- 28.Williams DL, Jr, Greene LE, Eisenberg E. Cooperative turning on of myosin subfragment 1 adenosinetriphosphatase activity by the troponin-tropomyosin-actin complex. Biochemistry. 1988;27(18):6987–6993. doi: 10.1021/bi00418a048. [DOI] [PubMed] [Google Scholar]
- 29.Gordon AM, Homsher E, Regnier M. Regulation of contraction in striated muscle. Physiol Rev. 2000;80(2):853–924. doi: 10.1152/physrev.2000.80.2.853. [DOI] [PubMed] [Google Scholar]
- 30.Kad NM, Kim S, Warshaw DM, VanBuren P, Baker JE. Single-myosin crossbridge interactions with actin filaments regulated by troponin-tropomyosin. Proc Natl Acad Sci USA. 2005;102(47):16990–16995. doi: 10.1073/pnas.0506326102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Ishii Y, Lehrer SS. Excimer fluorescence of pyrenyliodoacetamide-labeled tropomyosin: A probe of the state of tropomyosin in reconstituted muscle thin filaments. Biochemistry. 1990;29(5):1160–1166. doi: 10.1021/bi00457a010. [DOI] [PubMed] [Google Scholar]
- 32.Geeves MA, Lehrer SS. Dynamics of the muscle thin filament regulatory switch: The size of the cooperative unit. Biophys J. 1994;67(1):273–282. doi: 10.1016/S0006-3495(94)80478-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Brown JH, et al. Structure of the mid-region of tropomyosin: Bending and binding sites for actin. Proc Natl Acad Sci USA. 2005;102(52):18878–18883. doi: 10.1073/pnas.0509269102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Nitanai Y, Minakata S, Maeda K, Oda N, Maéda Y. Crystal structures of tropomyosin: Flexible coiled-coil. Adv Exp Med Biol. 2007;592:137–151. doi: 10.1007/978-4-431-38453-3_13. [DOI] [PubMed] [Google Scholar]
- 35.El-Saleh SC, Thieret R, Johnson P, Potter JD. Modification of Lys-237 on actin by 2,4-pentanedione. Alteration of the interaction of actin with tropomyosin. J Biol Chem. 1984;259(17):11014–11021. [PubMed] [Google Scholar]
- 36.Gerson JH, Bobkova E, Homsher E, Reisler E. Role of residues 311/312 in actin-tropomyosin interaction. In vitro motility study using yeast actin mutant e311a/r312a. J Biol Chem. 1999;274(25):17545–17550. doi: 10.1074/jbc.274.25.17545. [DOI] [PubMed] [Google Scholar]
- 37.Saeki K, Sutoh K, Wakabayashi T. Tropomyosin-binding site(s) on the Dictyostelium actin surface as identified by site-directed mutagenesis. Biochemistry. 1996;35(46):14465–14472. doi: 10.1021/bi961292c. [DOI] [PubMed] [Google Scholar]
- 38.Szilagyi L, Lu RC. Changes of lysine reactivities of actin in complex with myosin subfragment-1, tropomyosin and troponin. Biochim Biophys Acta. 1982;709(2):204–211. doi: 10.1016/0167-4838(82)90462-9. [DOI] [PubMed] [Google Scholar]
- 39.Tao T, Lamkin M. Crosslinking of tropomyosin to myosin subfragment-1 in reconstituted rabbit skeletal thin filaments. FEBS Lett. 1984;168(1):169–173. doi: 10.1016/0014-5793(84)80229-x. [DOI] [PubMed] [Google Scholar]
- 40.Li XE, et al. Tropomyosin position on F-actin revealed by EM reconstruction and computational chemistry. Biophys J. 2011;100(4):1005–1013. doi: 10.1016/j.bpj.2010.12.3697. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.McLachlan AD, Stewart M. The 14-fold periodicity in alpha-tropomyosin and the interaction with actin. J Mol Biol. 1976;103(2):271–298. doi: 10.1016/0022-2836(76)90313-2. [DOI] [PubMed] [Google Scholar]
- 42.Winkelmann DA, Bourdieu L, Ott A, Kinose F, Libchaber A. Flexibility of myosin attachment to surfaces influences F-actin motion. Biophys J. 1995;68(6):2444–2453. doi: 10.1016/S0006-3495(95)80426-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Whitby FG, Phillips GN., Jr Crystal structure of tropomyosin at 7 Angstroms resolution. Proteins. 2000;38(1):49–59. [PubMed] [Google Scholar]
- 44.Greenfield NJ, et al. Solution NMR structure of the junction between tropomyosin molecules: Implications for actin binding and regulation. J Mol Biol. 2006;364(1):80–96. doi: 10.1016/j.jmb.2006.08.033. [DOI] [PubMed] [Google Scholar]
- 45.Pettersen EF, et al. UCSF Chimera—A visualization system for exploratory research and analysis. J Comput Chem. 2004;25(13):1605–1612. doi: 10.1002/jcc.20084. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.