Abstract
Purpose of Review
The regulatory lipids are a class of bioactive lipids which regulate various important biological processes. Profiling these regulatory lipids is an attractive method to understand the role of these metabolites. This is especially true since most of these regulatory lipids are derived from several important pharmacological targets: cyclooxygenase (COX), lipoxygenase (LOX), and cytochrome P450 enzymes. This review highlights the development of methods to profile these regulatory lipids and the recent publications employing these profiling methods.
Recent findings
The recent development of methods for the profiling of regulatory lipids target two different directions: to expand coverage for discovery studies (fingerprinting) and to make the quantitative method more accurate, sensitive, and faster for diagnostic or more detailed studies. Recent applications of these profiling methods including assessment of in vivo drug engagement, pathways crosstalk, and possible mechanisms for side effects of a withdrawn anti-inflammatory drug-Rofecoxib are also reviewed here.
Summary
The profiling of regulatory lipids is a useful tool for many investigations. The breadth of coverage, throughput limits with detection, and reproducibility of quantitation are being improved. The resulting data will assist with fundamental investigation, disease biomarker discovery, drug discovery and drug development.
Keywords: Regulatory lipids, profiling, Lipidomics, oxylipins, eicosanoids
Introduction
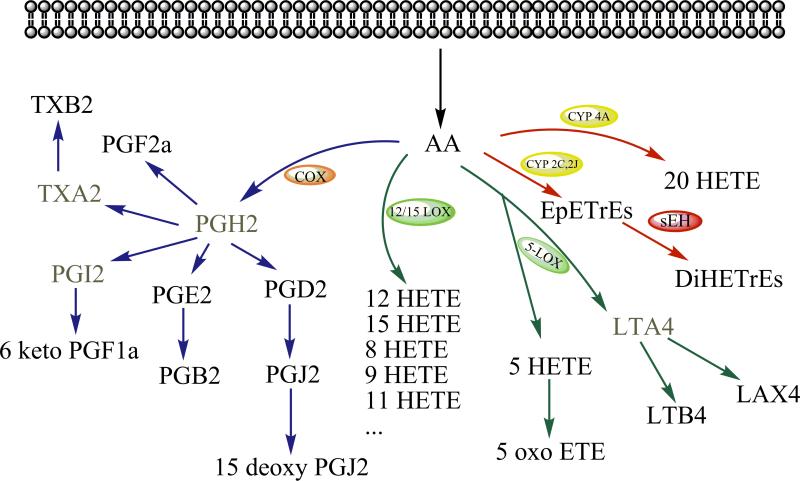
Techniques for omic analyses have continued to expand and improve during the last decade. They continue to evolve along with genomics, proteomics, transcriptomics, and metabolomics. As a very important part of the metabolome, the lipidome, aimed at mapping all of the lipids of a cell, is of increasing importance in omics evolution [1]. Many significant efforts have been made to promote research in this emerging field. Analysis of nutritional and structural lipids has been reviewed in detail elsewhere [2-7]. The regulatory lipids are referred to as lipid chemical mediators which regulate various important biological processes such as cell proliferation, apoptosis, tissue repair, blood clotting, blood vessel permeability, inflammation, immune cell behavior, and other biological functions, and which act at low concentrations to do so. Examples of regulatory lipids include the eicosanoids and the lipids derived from other polyunsaturated fatty acids (PUFAs). Eicosanoids are oxylipins produced from arachidonic acid and include prostaglandins (PGs), thromboxanes (TXs), leukotrienes (LTs), hydroxyeicosatetranoic acids (HETEs), epoxyeicosatrienoic acids (EpETrEs), and lipoxins (LXs). Figure 1 shows some of the regulatory lipids derived from arachidonic acid. There are targeted assays for some of the well investigated regulatory lipids such as PGE2 and LTs. However, there is cross talk among regulatory lipids as well as cross talk among regulatory lipids and chemokines, cytokines and other chemical mediators. Thus, it is vital to have a comprehensive profiling method for these regulatory lipids.
Figure 1.
Regulatory lipids from three main branches of the arachidonic acid cascade. The ones derived from other PUFAs were not shown here but are metabolized in a parallel manner. PL, phospholipid. The compounds in gray font are the unstable metabolites. Adapted from Yang J. et al. paper [8].
The analytical technologies for this lipid profiling have been developed by several research groups. This review will highlight current progress on the analytical methods and several important biological applications utilizing regulatory lipid profiling methods.
Analytical Methods
Regulatory lipids are products of a few fatty acid species and represent the addition of oxygen resulting in molecules with generally similar structures, chemistries, and physical properties. Many of these oxidized lipids or oxylipins are isomers, which makes their identification and quantitation in a single biological sample a challenging task. In addition, most compounds are present at low concentrations, but they can vary in concentration by more than 3 orders of magnitude. Some of the methods to measure subsets of these oxylipins include Enzyme-linked Immunosorbent assays (ELISA)[9] and other immunoassays, gas chromatography (GC)[10] or GC/mass spectrometry (GC/MS)[11], Liquid chromatography (LC)/UV, LC/MS[12], and LC/MS/MS[8, 13-22]. LC/MS/MS currently is the most powerful tool because of its specificity and sensitivity. Unfortunately, most of the published methods [11, 13-19, 21] only analyze a small portion of the known regulatory lipids.
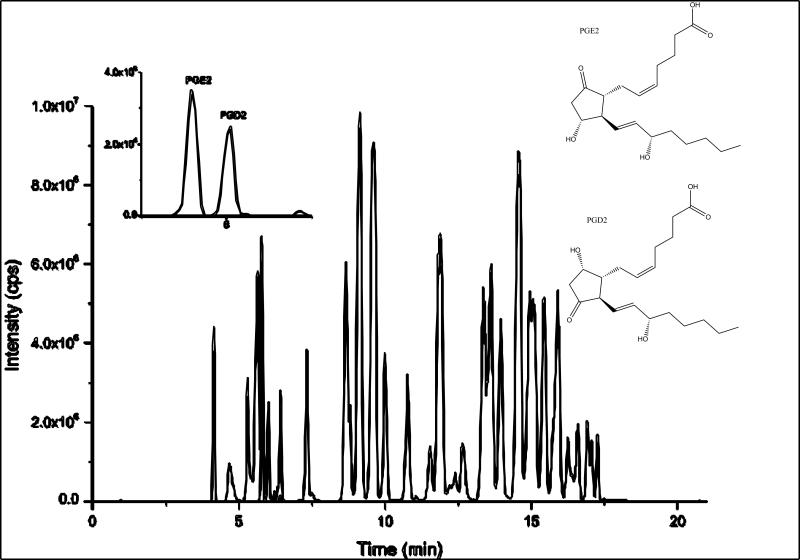
In a recent paper [8], a novel sensitive method was reported for the detection and quantification of the regulatory lipids that employs solid phase extraction (SPE), HPLC separation, and electrospray ionization (ESI)-MS/MS in multiple reaction-monitoring (MRM) mode. This method is quantitative for products of multiple enzymatic pathways, which allows a more complete assessment of localized and systemic pathological changes. The LC separations were carefully optimized to separate the critical separation pairs while keeping the total run relatively short. Figure 2 shows the typical separation and one of the critical separation pairs. In 21 minutes, 39 regulatory lipids were quantified along with eight corresponding internal standards.
Figure 2.
Typical separation of regulatory lipid profiling shown by the standard total ion chromatogram (TIC). The extracted ion chromatogram (XIC) in the insert illustrates one of the critical separation pairs, which share the identical MS/MS transition and similar structure. Adapted from Yang J. et al. paper [8].
Using two different types of internal standards, the extraction recovery and the instrument variability could be adjusted and higher accuracy measurements could be realized for trace regulatory lipids. An internal standard (type II internal standard) was chosen to monitor the injection variation and eight type I internal standards were selected to compensate for the differences in the extraction and ionization efficiency due to differences in chemical structure and chromatographic elution. When commercially available, deuterated internal standards were used. The validation shows this method meets exacting criteria for several important parameters. Linearity of each compound was greater than 0.999 over the calibration range of 0.07 to 32 pg (20 pM-10 nM) injected on the column. The accuracy and precision of the method were established by examining the reproducibility of several quality control samples over extended times and conditions. New methods for oxylipin analysis will continually be needed for broader coverage with better quantitation and sensitivity. Currently, well over 100 biologically active oxylipins have been identified, and the number continues to grow indicating that broader coverage of structures is needed. For example multiple regulatory lipids in the ω-3 series appear biologically important. On the other hand, the basal concentrations of several oxylipins are below the detection limit of even the most sensitive methods. However, even with careful optimization and more powerful instrumentation, there will be a tradeoff between the number of analytes monitored and the sensitivity. In addition, the chirality of many oxylipins is critical to biological activity, but most current methods do not address the chirality problem. Although conceptually straightforward, the analysis of chirality of some analytes will come at the expense of other parameters such as the number of lipids detected. Coverage, sensitivity, accuracy, precision, and speed must of course be weighed against cost of analysis. For any biological problem, there will be tradeoffs among these drivers, requiring analytical chemists to adjust their procedures for the goals of the project, develop new techniques to obtain more information from existing instrumentation, and adapt their methods to improved instrumentation. For studies in which there is a high degree of control over the experimental system and a desire to capture as much information as possible, the method with broader coverage is an obviously attractive approach. For diagnostic work or more defined studies, greater accuracy, quicker and sensitive quantitation methods are essential. [9]
There are several efforts to broaden the number of metabolites analyzed [20, 23]. One example is the development of an eicosanoid profiling method for 104 unique lipid species [23]. This method covers almost all of the known eicosanoids. However, it suffers from the problem of low sensitivity. The limit of detection (LOD) of the reported method is much worse than the limit of quantitation (LOQ) of the above method described at the beginning of this chapter. This is illustrated by an LOD of PGE2 for their method of 1 versus 0.07 pg. Due to increasing evidence showing that the regulatory lipids derived from ω -3 fatty acids play important roles in biological processes [24-27], another important addition involves the expansion of analytical methods to include the ω-3 derived regulatory lipids. The Hammock lab has already added 38 regulatory lipids derived from ω-3 fatty acids to the previous method. The improved method is targeted to 77 regulatory lipids without compromising detection sensitivity by taking new features (scheduled MRM) from the software and careful optimization (unpublished data).
An example of targeting several regulatory lipids is shown by Ferreorp-Vera et al [28]. The approach is based on a hyphenated system composed of a SPE workstation on-line coupled to a LC-triple quadrupole-tandem mass spectrometer. Nine regulatory lipids were selected to be monitored. Due to the reducing complexity, the analysis was accelerated by online SPE.
Applications of Regulatory Lipid Profiling
As mentioned above, the regulatory lipids play important roles in physiological and pathological process such as the inflammatory process. Thus, profiling these regulatory lipids is an attractive way to uncover the mystery behind these pathological phenomena. This is especially true for the eicosanoids where the arachidonate cascade is the target of over 75% of the world's pharmaceuticals.
The following examples highlight applications of lipid profiling from the simple but very important assessment of in vivo drug engagement to a complicated example of crosstalk among the different pathways. This is followed by a discovery study using the profiling method-the Rofecoxib story. Finally, an investigation related to the expansion to the ω-3 derived regulatory lipids is discussed.
The Assessment of in vivo Drug Engagement
Although pharmacokinetic studies can monitor drug concentrations as a function of time to provide information on absorption, distribution, metabolism and excretion (ADME), drug levels do not address the drug in the in vivo system. Analytical methods to target engagement provide an important link between plasma drug levels and in vivo efficacy.
Because most of the pharmacological targets are related to the arachidonate pathways, one of the obvious applications of regulatory profiling methods is to assess in vivo drug engagement. A newly discovered pharmacological target-soluble epoxide hydrolase (sEH) is chosen as an example here. sEH has proven to be a therapeutic target for acute inflammation, pain, and cardiovascular diseases [29]. sEH is an α/β hydrolase fold enzyme that converts EpETrEs to their less biologically active corresponding vicinal diols, dihydroxyeicosatrienoic acids (DiHETrEs). Endogenous P450-derived EpETrEs possess anti-inflammatory properties via inhibition of NF- κ B activation [30]. The biological activity and molecular signaling mechanisms of EpETrEs have been reviewed comprehensively elsewhere [31, 32]. By using a quantitative method to measure eicosanoids, changes in eicosanoid profiles can be assessed and used as valuable biomarkers of sEH inhibitors (sEHI) [8, 33]. Simplified biomarkers for sEHI effects are the dramatic increase in EpETrEs levels and/or increased ratios of EpETrEs to their corresponding diols.
Li et al. found a significantly decreased EpETrEs/DiHETrEs ratio, which represents an inflammatory state, in a murine model of myocardial infarction. The administration of sEHI increased EpETrEs/DiHETrEs ratios and was highly effective in the prevention of progressive cardiac remodeling post myocardial infarction [34]. The EpETrEs also are analgesic. Inceoglu et al. revealed that with increased EpETrEs /DiHETrEs ratios, sEHIs lead to anti-hyperalgesia by at least 2 spinal mechanisms, the induction of the COX2 gene and the regulation of an acute neurosteroid-producing gene, StARD1 [35].
Crosstalk among Pathways
Since the regulatory lipids are synthesized from related pathways as shown in Figure 1, a change in any pathway will presumably affect the adjacent pathways by shuttling of starting materials and intermediate metabolites. A well-known example is aspirin-induced asthma (AIA). In AIA, inhibition of COX is associated with release of cysteinylleukotrienes (cys-LTs), which have emerged as important mediators of asthma, and may be particularly prominent in aspirin-induced respiratory reactions [36]. This is thought to occur by mass action following the inhibition of the COX enzymes. Profiling the substrates and the metabolites (the regulatory lipids) of these pathways provides a good tool for studying crosstalk between these pathways from the metabolite levels. Combined with protein level and message level information, the biological mechanisms underlying these biological processes can be uncovered.
As an illustration of greater complexity in crosstalk, Schmelzer K. et al. [37] demonstrated that regulatory lipid profiles shifted after combination therapy of nonsteroidal anti-inflammatory drugs (NSAIDs) and sEHIs. Coadministration of sEHIs and NSAIDs produced synergistic decreases in COX pathway metabolites (PGD2 and PGE2) 6 hours after lipopolysaccharide (LPS) exposure while elevating the concentrations of EpETrEs. The COX-2 inhibitors increased the flow of arachidonic acid (AA) through the P450 pathway and led to increased EpETrEs and DiHETrEs in murine plasma after exposure to LPS. However, the administration of sEHI decreased PGE2 and PGD2 production instead of increasing them by mass shunting. Protein analysis (western blotting) of COX2 showed that inflammation induced hepatic COX2 expression was suppressed by the sEHIs, which explains the dramatic decrease in PGE2 and PGD2 production after sEH inhibition. These data predict synergistic interactions in reducing inflammation between a transcriptional down regulator of induced COX-2 (sEHI) and an enzyme inhibitor (COXIB or NSAID).
Not only did the lipid profiling method demonstrate the dramatic down regulation of PGE2 by sEHI, likely through a translational mechanism, but it showed a dramatic increase in EpETrEs with COX inhibitors such as aspirin. Thus, one could argue that the beneficial effects of aspirin and other NSAIDs come from the increase in anti-inflammatory lipid chemical mediators like EpETrEs as well as the decrease in inflammatory mediators like PGE2.
In a recent study by Liu J.Y. et al [38], crosstalk was investigated in the same model system. Significant crosstalk among branches of the AA cascade was found during selective modulation of COX, LOX or sEH. Again, profiling of regulatory lipids was employed to discover these interactions. Inhibition of the COX pathway significantly reduced the levels of sEH-produced DiHETrEs and the LOX products 5- HETE and 15-HETE. This effect may be due to feedback regulation of the anti-inflammatory effect from each individual inhibitor. Blockade of the LOX pathway by the 5-LOX activation protein (FLAP) inhibitor MK886 significantly decreased the production of COX products TXB2 and PGE2 while increasing the production of DiHETrEs. The inhibition of sEH suppresses the action of COX and LOX pathways by reducing the production of PGE2 and 5-HETE. This work suggests that metabolism of AA by COX, LOX and CYP450 family enzymes do not just follow a simple mass flow rule but involve a complex crosstalk among chemical mediators.
The Rofecoxib Story
Rofecoxib is a NSAID that was withdrawn from the market by Merck in 2004 due to its side effect of a higher risk for cardiovascular events and stoke in arthritic patients [39]. One of the proposed explanations is that rofecoxib can reduce the production of the platelet aggregation inhibitor prostacyclin I2 but not platelet activator thromboxane A2, which can increase the risk for cardiovascular diseases such as hypertension, myocardial infarction, and heart failure [40-43]. Thus, regulatory lipids profiles that include quantify PGI2 and TXA2 should be able to provide the evidence for this hypothesis.
In a recently published paper [44], instead of monitoring AA generated regulatory lipids only from COX enzymes, Liu and co-workers broadly quantified the representative regulatory lipids derived from ARA and linoleic acid mediated by COXs, CYP450s, and LOXs. Unexpectedly, they found a 120-fold increase of the eicosanoid 20-Hydroxyeicosatetraenoic acid (20-HETE) after oral administration of rofecoxib for 3 months. This increase was correlated with a significantly shorter tail bleeding time in a murine model. The decreased bleeding time may be one of the adverse cardiovascular events caused by rofecoxib. 20-HETE is a potent vasoconstrictor and plays important roles in cardiovascular function. This work suggested that regulatory lipid profiling is a promising tool to gain a comprehensive understanding of biological processes and that 20-HETE may potentially be used as a marker of rofecoxib induced cardiovascular adverse events.
ω-3 Fatty Acids-derived regulatory lipids
The early human diet was most likely rich in stearic acid and the ω-6 fatty acid oleic acid (18:1) with moderate amounts of ω-3 fats. Diet probably varied dramatically through divergent pathways of cultural evolution. Until the 1960s the American diet was largely stearate and oleate based with an increasing trend toward PUFA and particularly linoleic acid (18:2). A series of events led to a significant alteration in the American diet at this time with a massive increase in the proportion of linoleic acid and a decrease in ω-3 fats with little or no scientific driver [45]. Society seldom invests in clinical trials to evaluate the effects of diet since it is difficult for one organization to profit from the results. In this case, the food industry inadvertently increased the ratio of omega-6 fatty acids because of the availability and desirability of grain-based oils. However, due to increasing evidence that inflammation underlies many of today's diseases, and the involvement of omega-6 fatty acids, particularly AA, in the production of pro-inflammatory eicosanoids, there is currently another dietary revolution where ω-3 lipids will be supplemented. Low levels of ω-3s in the American diet come from a variety of plants with soybean oil being a major contributor. However, these PUFAs were dramatically reduced by classical breeding and genetic engineering due to their instability in foods. In addition, terrestrial plants provide only the precursor omega-3 fatty acid, alpha-linolenic acid, whereas it is the longer-chain derivatives eicosapentaenoic acid (EPA) and docosahexaenoic acid (DHA) that are precursors for oxylipins. Cold water fish such as salmon, tuna, anchovies, and sardines are rich in long-chain ω-3 fats EPA and DHA having concentrated them from diatoms. However, there are now many sources of varying quality.
Several ω-3 trials suggest that ω-3 fatty acids, the main contents of fish oil, have beneficial effects in many diseases[46] such as asthma[47], Alzheimer's disease[48], depression[49], inflammatory bowel disease[50], diabetes[51], cardiovascular diseases[52], and cancer[53]. Most ω-3 trials, however, lack the power to draw strong conclusions. The mechanisms of these beneficial effects are under investigation. The proposed mechanisms include among others the lipid raft theory[54], competition with AA to produce less inflammatory regulatory lipids, and production of anti-inflammatory regulatory lipids[25, 27, 55] from ω-3 starting materials.
A recent paper published by Morisseau C. et al [27] reported naturally occurring monoepoxides of EPA and DHA that acted as bioactive antihyperalgesic lipids in a rat pain model. By profiling the regulatory lipids in rat brain and spinal cord tissues, significantly more EpDPEs (from DHA) were found than EpETEs (from EPA). The ω-3 EpDPEs were in similar quantities to the EpETrEs although their plasma levels were much lower. These data imply that the epoxides derived from ω-3 fatty acids might act similarly to ω-6 EpETrEs, which were proven antihyperalgesic in a previous rat pain model [35]. A kinetic study showed that human sEH prefers ω-3 EpDPEs over ω-6 EpETrEs in terms of Kcat and KM. Furthermore, an in vivo rat model demonstrated that these epoxides are antihyperalgesic with both peripheral and central actions on the nervous system [27].
Conclusions
In the past two years, analytical efforts in several laboratories have broadened the coverage of the regulatory lipids while maintaining high quantitative accuracy. This makes it feasible to profile these important regulatory lipids in various systems in vivo. One trend has been a move towards quantitative methods. Other trends of analytical method development include 1) to increase the number of regulatory lipids analyzed to gain more information in a single run for discovery studies, and 2) to speed up the analytical process with higher detection ability in confirmation or diagnosis studies like in vivo drug engagement studies. In addition, a more systemic way to interpret the data will be a very important issue since the synergistic and antagonistic effects of these regulatory lipids generally exist.
Recent applications utilizing these methods for profiling regulatory lipids range from fundamental biochemical studies to clinical research. Due to the important roles these regulatory lipids play in inflammatory diseases, this profiling strategy will be one of the powerful new tools for investigating inflammatory diseases and pharmacological investigations. As more new regulatory lipids and new biological functions of the current lipids are discovered, regulatory lipid profiling will contribute more to these fields.
Key points.
Profiling regulatory lipids is an attractive method to understand the role of these metabolites, which regulate various important biological processes.
The recent development of methods for the profiling of regulatory lipids target two different directions: to expand coverage for discovery studies (fingerprinting) and to make the quantitative method more accurate, sensitive, and faster for diagnostic or more detailed studies.
Recent applications utilizing these methods for profiling regulatory lipids range from fundamental biochemical studies to clinical research.
Due to the important roles these regulatory lipids play in inflammatory diseases, this profiling strategy will be one of the powerful new tools for investigating inflammatory diseases and pharmacological investigations.
Acknowledgements
The authors’ work was supported in part by NIEHS SBRP Grant p42 ES004699, NIEHS Grant R37 ES02710, and NIH/NIEHS Grant R01 ES013933. Partial support was provided by the American Asthma Association #09-0269. J.Y. was supported by the Elizabeth Nash Memorial fellowship from the Cystic Fibrosis Research Inc. B.D.H. is a George and Judy Marcus Senior Fellow of the American Asthma Foundation. The authors want to thank Dr. Angela M. Zivkovic for revising the manuscript.
References and Recommended reading
Papers of particular interest, published within the annual period of review, have been highlighted as:
• of special interest
•• of outstanding interest
- 1.Dennis EA. Lipidomics joins the omics evolution. Proc Natl Acad Sci U S A. 2009;106(7):2089–90. doi: 10.1073/pnas.0812636106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2••.Wolf C, Quinn PJ. Lipidomics: practical aspects and applications. Prog Lipid Res. 2008;47(1):15–36. doi: 10.1016/j.plipres.2007.09.001. [This is a thorough review regarding on the lipidomics study.] [DOI] [PubMed] [Google Scholar]
- 3.Han X. Lipidomics: developments and applications. J Chromatogr B Analyt Technol Biomed Life Sci. 2009;877(26):2663. doi: 10.1016/j.jchromb.2009.07.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Hu C, et al. Analytical strategies in lipidomics and applications in disease biomarker discovery. J Chromatogr B Analyt Technol Biomed Life Sci. 2009;877(26):2836–46. doi: 10.1016/j.jchromb.2009.01.038. [DOI] [PubMed] [Google Scholar]
- 5.Ivanova PT, et al. Lipidomics: a mass spectrometry based systems level analysis of cellular lipids. Curr Opin Chem Biol. 2009;13(5-6):526–31. doi: 10.1016/j.cbpa.2009.08.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Proschogo N, Gaus K, Jessup W. Taking aim at cell lipids: shotgun lipidomics and imaging mass spectrometry push the boundaries. Curr Opin Lipidol. 2009;20(6):522–3. doi: 10.1097/MOL.0b013e32833301cb. [DOI] [PubMed] [Google Scholar]
- 7.Bou Khalil M, et al. Lipidomics era: accomplishments and challenges. Mass Spectrom Rev. 2010;29(6):877–929. doi: 10.1002/mas.20294. [DOI] [PubMed] [Google Scholar]
- 8••.Yang J, et al. Quantitative profiling method for oxylipin metabolome by liquid chromatography electrospray ionization tandem mass spectrometry. Anal Chem. 2009;81(19):8085–93. doi: 10.1021/ac901282n. [A method paper included the most important regulatory lipids profiling with the high sensitivity and accuracy.] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Wiest MM, Watkins SM. Biomarker discovery using high-dimensional lipid analysis. Curr Opin Lipidol. 2007;18(2):181–6. doi: 10.1097/MOL.0b013e3280895d82. [DOI] [PubMed] [Google Scholar]
- 10.Newman JW, Hammock BD. Optimized thiol derivatizing reagent for the mass spectral analysis of disubstituted epoxy fatty acids. J Chromatogr A. 2001;925(1-2):223–40. doi: 10.1016/s0021-9673(01)00998-0. [DOI] [PubMed] [Google Scholar]
- 11.Werner K, et al. Characterization and identification of cytochrome P450 metabolites of arachidonic acid released by human peritoneal macrophages obtained from the pouch of Douglas. Prostaglandins Leukot Essent Fatty Acids. 2002;67(6):397–404. doi: 10.1054/plef.2002.0449. [DOI] [PubMed] [Google Scholar]
- 12.Deems R, et al. Detection and quantitation of eicosanoids via high performance liquid chromatography-electrospray ionization-mass spectrometry. Methods Enzymol. 2007;432:59–82. doi: 10.1016/S0076-6879(07)32003-X. [DOI] [PubMed] [Google Scholar]
- 13.Takabatake M, et al. Simultaneous quantification of prostaglandins in human synovial cell-cultured medium using liquid chromatography/tandem mass spectrometry. Prostaglandins Leukot Essent Fatty Acids. 2002;67(1):51–6. doi: 10.1054/plef.2002.0381. [DOI] [PubMed] [Google Scholar]
- 14.Rinne S, et al. Fast and simple online sample preparation coupled with capillary LC-MS/MS for determination of prostaglandins in cell culture supernatants. J Sep Sci. 2007;30(12):1860–9. doi: 10.1002/jssc.200700064. [DOI] [PubMed] [Google Scholar]
- 15•.Hishinuma T, et al. Simultaneous quantification of seven prostanoids using liquid chromatography/tandem mass spectrometry: the effects of arachidonic acid on prostanoid production in mouse bone marrow-derived mast cells. Prostaglandins Leukot Essent Fatty Acids. 2007;76(6):321–9. doi: 10.1016/j.plefa.2007.04.005. [An example which shows the online SPE coupled LC/MS/MS for quicker analysis.] [DOI] [PubMed] [Google Scholar]
- 16.Newman JW, Watanabe T, Hammock BD. The simultaneous quantification of cytochrome P450 dependent linoleate and arachidonate metabolites in urine by HPLC-MS/MS. Journal of Lipid Research. 2002;43(9):1563–78. doi: 10.1194/jlr.d200018-jlr200. [DOI] [PubMed] [Google Scholar]
- 17.Cao H, et al. An improved LC-MS/MS method for the quantification of prostaglandins E(2) and D(2) production in biological fluids. Anal Biochem. 2008;372(1):41–51. doi: 10.1016/j.ab.2007.08.041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Masoodi M, et al. Simultaneous lipidomic analysis of three families of bioactive lipid mediators leukotrienes, resolvins, protectins and related hydroxy-fatty acids by liquid chromatography/electrospray ionisation tandem mass spectrometry. Rapid Commun Mass Spectrom. 2008;22(2):75–83. doi: 10.1002/rcm.3331. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Serhan CN. Mediator lipidomics. Prostaglandins Other Lipid Mediat. 2005;77(1-4):4–14. doi: 10.1016/j.prostaglandins.2004.09.016. [DOI] [PubMed] [Google Scholar]
- 20.Masoodi M, et al. Comprehensive lipidomics analysis of bioactive lipids in complex regulatory networks. Anal Chem. 2010;82(19):8176–85. doi: 10.1021/ac1015563. [DOI] [PubMed] [Google Scholar]
- 21.Miller TM, et al. Rapid, simultaneous quantitation of mono and dioxygenated metabolites of arachidonic acid in human CSF and rat brain. J Chromatogr B Analyt Technol Biomed Life Sci. 2009;877(31):3991–4000. doi: 10.1016/j.jchromb.2009.10.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22••.Buczynski MW, Dumlao DS, Dennis EA. Thematic Review Series: Proteomics. An integrated omics analysis of eicosanoid biology. Journal of Lipid Research. 2009;50(6):1015–38. doi: 10.1194/jlr.R900004-JLR200. [A great review described the eicasonoids in details.] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23••.Blaho VA, et al. Lipidomic analysis of dynamic eicosanoid responses during the induction and resolution of Lyme arthritis. Journal of Biological Chemistry. 2009;284(32):21599–612. doi: 10.1074/jbc.M109.003822. [Over 104 eicosanoids were listed in the method although just a small part of them were detected. The method covers the most comprehensive subsets of the regulatory lipids.] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Serhan CN, et al. Resolvins: a family of bioactive products of omega-3 fatty acid transformation circuits initiated by aspirin treatment that counter proinflammation signals. J Exp Med. 2002;196(8):1025–37. doi: 10.1084/jem.20020760. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25•.Serhan CN, Chiang N. Endogenous pro-resolving and anti-inflammatory lipid mediators: a new pharmacologic genus. Br J Pharmacol. 2008;153(Suppl 1):S200–15. doi: 10.1038/sj.bjp.0707489. [Great review on the pro-resolving and anti-inflamatory lipids derived from AA, DHA, EPA.] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Gleissman H, et al. Docosahexaenoic acid metabolome in neural tumors: identification of cytotoxic intermediates. Faseb Journal. 2010;24(3):906–15. doi: 10.1096/fj.09-137919. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Morisseau C, et al. Naturally occurring monoepoxides of eicosapentaenoic acid and docosahexaenoic acid are bioactive antihyperalgesic lipids. Journal of Lipid Research. 2010;51(12):3481–90. doi: 10.1194/jlr.M006007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Ferreiro-Vera C, et al. Automated targeting analysis of eicosanoid inflammation biomarkers in human serum and in the exometabolome of stem cells by SPE-LC-MS/MS. Anal Bioanal Chem. 2010 doi: 10.1007/s00216-010-4400-6. [DOI] [PubMed] [Google Scholar]
- 29.Imig JD, Hammock BD. Soluble epoxide hydrolase as a therapeutic target for cardiovascular diseases. Nat Rev Drug Discov. 2009;8(10):794–805. doi: 10.1038/nrd2875. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Node K, et al. Anti-inflammatory properties of cytochrome P450 epoxygenase-derived eicosanoids. Science. 1999;285(5431):1276–1279. doi: 10.1126/science.285.5431.1276. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Michaelis UR, Fleming I. From endothelium-derived hyperpolarizing factor (EDHF) to angiogenesis: Epoxyeicosatrienoic acids (EETs) and cell signaling. Pharmacology & Therapeutics. 2006;111(3):584–595. doi: 10.1016/j.pharmthera.2005.11.003. [DOI] [PubMed] [Google Scholar]
- 32.Spector AA, Norris AW. Action of epoxyeicosatrienoic acids on cellular function. American Journal of Physiology-Cell Physiology. 2007;292(3):C996–C1012. doi: 10.1152/ajpcell.00402.2006. [DOI] [PubMed] [Google Scholar]
- 33.Inceoglu B, et al. Soluble epoxide hydrolase inhibition reveals novel biological functions of epoxyeicosatrienoic acids (EETs). Prostaglandins Other Lipid Mediat. 2007;82(1-4):42–9. doi: 10.1016/j.prostaglandins.2006.05.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Li N, et al. Beneficial effects of soluble epoxide hydrolase inhibitors in myocardial infarction model: Insight gained using metabolomic approaches. Journal of Molecular and Cellular Cardiology. 2009;47(6):835–845. doi: 10.1016/j.yjmcc.2009.08.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Inceoglu B, et al. Soluble epoxide hydrolase and epoxyeicosatrienoic acids modulate two distinct analgesic pathways. Proceedings of the National Academy of Sciences of the United States of America. 2008;105(48):18901–18906. doi: 10.1073/pnas.0809765105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Szczeklik A. Mechanism of aspirin-induced asthma. Allergy. 1997;52(6):613–9. doi: 10.1111/j.1398-9995.1997.tb01039.x. [DOI] [PubMed] [Google Scholar]
- 37••.Schmelzer KR, et al. Enhancement of antinociception by coadministration of nonsteroidal anti-inflammatory drugs and soluble epoxide hydrolase inhibitors. Proc Natl Acad Sci U S A. 2006;103(37):13646–51. doi: 10.1073/pnas.0605908103. [This study use the profiling method shows the crosstalk between COX and sEH. It also shows coadministration of NSAID and sEHI could enhance the antinociception effect.] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38••.Liu JY, et al. Inhibition of soluble epoxide hydrolase enhances the anti-inflammatory effects of aspirin and 5-lipoxygenase activation protein inhibitor in a murine model. Biochem Pharmacol. 2010;79(6):880–7. doi: 10.1016/j.bcp.2009.10.025. [This study shows the complicated crosstalk among the three branches-COX, LOX and P450.] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Mukherjee D, Nissen SE, Topol EJ. Risk of cardiovascular events associated with selective COX-2 inhibitors. Jama-Journal of the American Medical Association. 2001;286(8):954–959. doi: 10.1001/jama.286.8.954. [DOI] [PubMed] [Google Scholar]
- 40.Cheng Y, et al. Role of prostacyclin in the cardiovascular response to thromboxane A(2). Science. 2002;296(5567):539–541. doi: 10.1126/science.1068711. [DOI] [PubMed] [Google Scholar]
- 41.Hippisley-Cox J, Coupland C. Risk of myocardial infarction in patients taking cyclo-oxygenase-2 inhibitors or conventional non-steroidal anti-inflammatory drugs: population based nested case-control analysis. British Medical Journal. 2005;330(7504):1366–1369. doi: 10.1136/bmj.330.7504.1366. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Johnson AG, et al. Nonsteroidal Antiinflammatory Drugs and Hypertension in the Elderly - a Community-Based Cross-Sectional Study. British Journal of Clinical Pharmacology. 1993;35(5):455–459. doi: 10.1111/j.1365-2125.1993.tb04169.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Page J, Henry D. Consumption of NSAIDs and the development of congestive heart failure in elderly patients - An underrecognized public health problem. Archives of Internal Medicine. 2000;160(6):777–784. doi: 10.1001/archinte.160.6.777. [DOI] [PubMed] [Google Scholar]
- 44••.Liu JY, et al. Metabolic profiling of murine plasma reveals an unexpected biomarker in rofecoxib-mediated cardiovascular events. Proc Natl Acad Sci U S A. 2010;107(39):17017–22. doi: 10.1073/pnas.1011278107. [This study is a excellent example which use the profiling method to discovery the new biomarker of the drug toxicity. Combined with other biological tool, 20-HETE was proven to be a potential biomarker for the rofecoxib-mediated cardiovascular events.] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Taubes G. Nutrition. The soft science of dietary fat. Science. 2001;291(5513):2536–45. doi: 10.1126/science.291.5513.2536. [DOI] [PubMed] [Google Scholar]
- 46.Yashodhara BM, et al. Omega-3 fatty acids: a comprehensive review of their role in health and disease. Postgrad Med J. 2009;85(1000):84–90. doi: 10.1136/pgmj.2008.073338. [DOI] [PubMed] [Google Scholar]
- 47.Reisman J, et al. Treating asthma with omega-3 fatty acids: where is the evidence? A systematic review. BMC Complement Altern Med. 2006;6:26. doi: 10.1186/1472-6882-6-26. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Huang TL. Omega-3 fatty acids, cognitive decline, and Alzheimer's disease: a critical review and evaluation of the literature. J Alzheimers Dis. 2010;21(3):673–90. doi: 10.3233/JAD-2010-090934. [DOI] [PubMed] [Google Scholar]
- 49.Liperoti R, et al. Omega-3 polyunsaturated fatty acids and depression: a review of the evidence. Curr Pharm Des. 2009;15(36):4165–72. doi: 10.2174/138161209789909683. [DOI] [PubMed] [Google Scholar]
- 50.Turner D, et al. Maintenance of remission in inflammatory bowel disease using omega-3 fatty acids (fish oil): a systematic review and meta-analyses. Inflamm Bowel Dis. 2011;17(1):336–45. doi: 10.1002/ibd.21374. [DOI] [PubMed] [Google Scholar]
- 51.Pouwer F, et al. Fat food for a bad mood. Could we treat and prevent depression in Type 2 diabetes by means of omega-3 polyunsaturated fatty acids? A review of the evidence. Diabet Med. 2005;22(11):1465–75. doi: 10.1111/j.1464-5491.2005.01661.x. [DOI] [PubMed] [Google Scholar]
- 52.Balk EM, et al. Effects of omega-3 fatty acids on serum markers of cardiovascular disease risk: a systematic review. Atherosclerosis. 2006;189(1):19–30. doi: 10.1016/j.atherosclerosis.2006.02.012. [DOI] [PubMed] [Google Scholar]
- 53.MacLean CH, et al. Effects of omega-3 fatty acids on cancer risk: a systematic review. JAMA. 2006;295(4):403–15. doi: 10.1001/jama.295.4.403. [DOI] [PubMed] [Google Scholar]
- 54.Yaqoob P. The nutritional significance of lipid rafts. Annu Rev Nutr. 2009;29:257–82. doi: 10.1146/annurev-nutr-080508-141205. [DOI] [PubMed] [Google Scholar]
- 55.Serhan CN. Systems approach with inflammatory exudates uncovers novel anti-inflammatory and pro-resolving mediators. Prostaglandins Leukot Essent Fatty Acids. 2008;79(3-5):157–63. doi: 10.1016/j.plefa.2008.09.012. [DOI] [PMC free article] [PubMed] [Google Scholar]