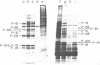Abstract
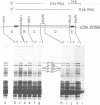
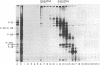
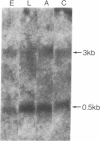
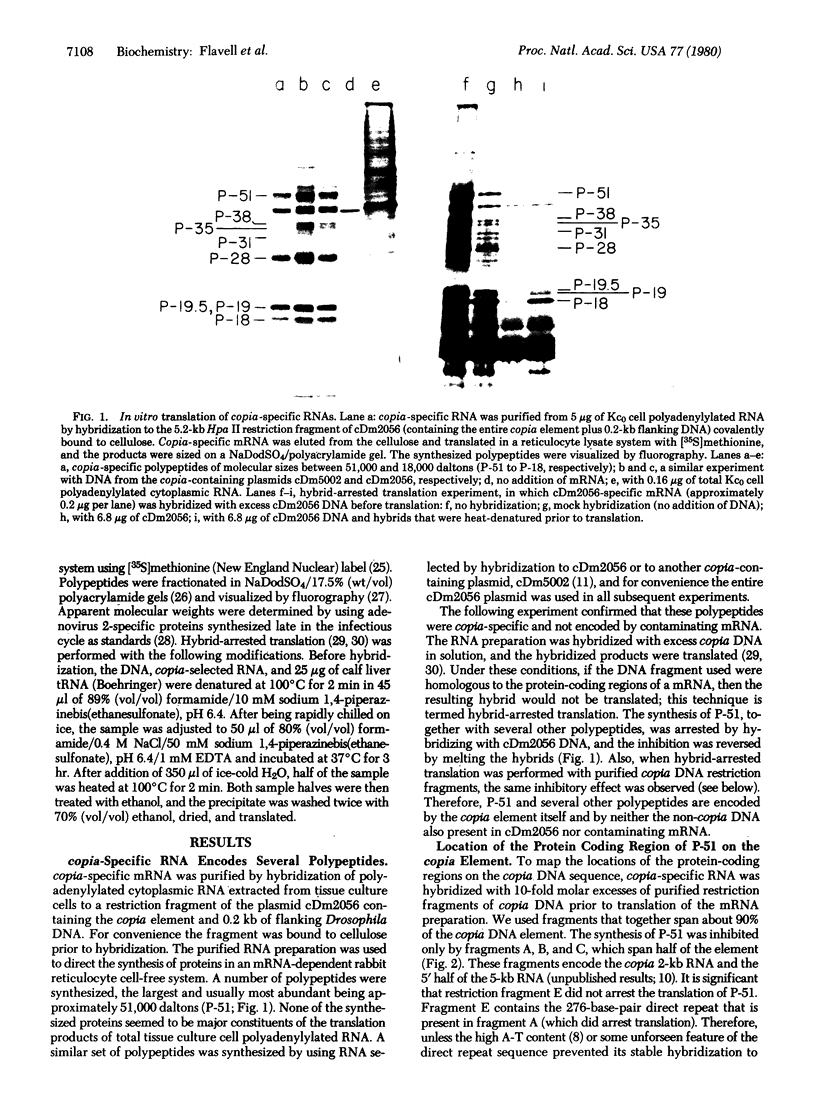
copia-specific RNA was isolated from Drosophila melanogaster tissue culture cells by hybridization of cytoplasmic polyadenylylated RNA to copia DNA immobilized on cellulose. The purified RNA was translated in reticulocyte lysates. One major polypeptide of approximately 51,000 daltons was synthesized in addition to several others between 18,000 and 38,000 daltons. The 51,000-dalton polypeptide and several of the others are encoded by mRNAs of about 2000 nucleotides. The approximate locations on the copia element of the coding sequences for the 51,000-dalton polypeptide and several other proteins were determined by hybrid-arrested translation with copia restriction fragments. The relative abundance of copia-specific RNA was determined at various stages of the Drosophila life cycle. The level of copia-specific RNA is modulated during development of the organism, with the highest level occurring during the larval stages.
Full text
PDF



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Alwine J. C., Kemp D. J., Stark G. R. Method for detection of specific RNAs in agarose gels by transfer to diazobenzyloxymethyl-paper and hybridization with DNA probes. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5350–5354. doi: 10.1073/pnas.74.12.5350. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bonner W. M., Laskey R. A. A film detection method for tritium-labelled proteins and nucleic acids in polyacrylamide gels. Eur J Biochem. 1974 Jul 1;46(1):83–88. doi: 10.1111/j.1432-1033.1974.tb03599.x. [DOI] [PubMed] [Google Scholar]
- Calos M. P., Miller J. H. Transposable elements. Cell. 1980 Jul;20(3):579–595. doi: 10.1016/0092-8674(80)90305-0. [DOI] [PubMed] [Google Scholar]
- Carlson M., Brutlag D. One of the copia genes is adjacent to satellite DNA in Drosophila melanogaster. Cell. 1978 Nov;15(3):733–742. doi: 10.1016/0092-8674(78)90259-3. [DOI] [PubMed] [Google Scholar]
- Dhar R., McClements W. L., Enquist L. W., Vande Woude G. F. Nucleotide sequences of integrated Moloney sarcoma provirus long terminal repeats and their host and viral junctions. Proc Natl Acad Sci U S A. 1980 Jul;77(7):3937–3941. doi: 10.1073/pnas.77.7.3937. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dunsmuir P., Brorein W. J., Jr, Simon M. A., Rubin G. M. Insertion of the Drosophila transposable element copia generates a 5 base pair duplication. Cell. 1980 Sep;21(2):575–579. doi: 10.1016/0092-8674(80)90495-x. [DOI] [PubMed] [Google Scholar]
- Farabaugh P. J., Fink G. R. Insertion of the eukaryotic transposable element Ty1 creates a 5-base pair duplication. Nature. 1980 Jul 24;286(5771):352–356. doi: 10.1038/286352a0. [DOI] [PubMed] [Google Scholar]
- Fincham J. R., Sastry G. R. Controlling elements in maize. Annu Rev Genet. 1974;8:15–50. doi: 10.1146/annurev.ge.08.120174.000311. [DOI] [PubMed] [Google Scholar]
- Finnegan D. J., Rubin G. M., Young M. W., Hogness D. S. Repeated gene families in Drosophila melanogaster. Cold Spring Harb Symp Quant Biol. 1978;42(Pt 2):1053–1063. doi: 10.1101/sqb.1978.042.01.106. [DOI] [PubMed] [Google Scholar]
- Flavell A. J., Cowie A., Legon S., Kamen R. Multiple 5' terminal cap structures in late polyoma virus RNA. Cell. 1979 Feb;16(2):357–371. doi: 10.1016/0092-8674(79)90012-6. [DOI] [PubMed] [Google Scholar]
- Fyrberg E. A., Kindle K. L., Davidson N., Kindle K. L. The actin genes of Drosophila: a dispersed multigene family. Cell. 1980 Feb;19(2):365–378. doi: 10.1016/0092-8674(80)90511-5. [DOI] [PubMed] [Google Scholar]
- Gafner J., Philippsen P. The yeast transposon Ty1 generates duplications of target DNA on insertion. Nature. 1980 Jul 24;286(5771):414–418. doi: 10.1038/286414a0. [DOI] [PubMed] [Google Scholar]
- Hastie N. D., Held W. A. Analysis of mRNA populations by cDNA.mRNA hybrid-mediated inhibition of cell-free protein synthesis. Proc Natl Acad Sci U S A. 1978 Mar;75(3):1217–1221. doi: 10.1073/pnas.75.3.1217. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Heffron F., McCarthy B. J., Ohtsubo H., Ohtsubo E. DNA sequence analysis of the transposon Tn3: three genes and three sites involved in transposition of Tn3. Cell. 1979 Dec;18(4):1153–1163. doi: 10.1016/0092-8674(79)90228-9. [DOI] [PubMed] [Google Scholar]
- Kleckner N. Translocatable elements in procaryotes. Cell. 1977 May;11(1):11–23. doi: 10.1016/0092-8674(77)90313-0. [DOI] [PubMed] [Google Scholar]
- Laemmli U. K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature. 1970 Aug 15;227(5259):680–685. doi: 10.1038/227680a0. [DOI] [PubMed] [Google Scholar]
- Levis R., Dunsmuir P., Rubin G. M. Terminal repeats of the Drosophila transposable element copia: nucleotide sequence and genomic organization. Cell. 1980 Sep;21(2):581–588. doi: 10.1016/0092-8674(80)90496-1. [DOI] [PubMed] [Google Scholar]
- MCCLINTOCK B. Controlling elements and the gene. Cold Spring Harb Symp Quant Biol. 1956;21:197–216. doi: 10.1101/sqb.1956.021.01.017. [DOI] [PubMed] [Google Scholar]
- Miller J. S., Ricciardi R. P., Roberts B. E., Paterson B. M., Mathews M. B. Arrangement of messenger RNAs and protein coding sequences in the major late transcription unit of adenovirus 2. J Mol Biol. 1980 Oct 5;142(4):455–488. doi: 10.1016/0022-2836(80)90258-2. [DOI] [PubMed] [Google Scholar]
- Noyes B. E., Stark G. R. Nucleic acid hybridization using DNA covalently coupled to cellulose. Cell. 1975 Jul;5(3):301–310. doi: 10.1016/0092-8674(75)90105-1. [DOI] [PubMed] [Google Scholar]
- Paterson B. M., Roberts B. E., Kuff E. L. Structural gene identification and mapping by DNA-mRNA hybrid-arrested cell-free translation. Proc Natl Acad Sci U S A. 1977 Oct;74(10):4370–4374. doi: 10.1073/pnas.74.10.4370. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Paucha E., Smith A. E. The sequences between 0.59 and 0.54 map units on SV40 DNA code for the unique region of small t antigen. Cell. 1978 Nov;15(3):1011–1020. doi: 10.1016/0092-8674(78)90285-4. [DOI] [PubMed] [Google Scholar]
- Pelham H. R., Jackson R. J. An efficient mRNA-dependent translation system from reticulocyte lysates. Eur J Biochem. 1976 Aug 1;67(1):247–256. doi: 10.1111/j.1432-1033.1976.tb10656.x. [DOI] [PubMed] [Google Scholar]
- Potter S. S., Brorein W. J., Jr, Dunsmuir P., Rubin G. M. Transposition of elements of the 412, copia and 297 dispersed repeated gene families in Drosophila. Cell. 1979 Jun;17(2):415–427. doi: 10.1016/0092-8674(79)90168-5. [DOI] [PubMed] [Google Scholar]
- Rothstein S. J., Jorgensen R. A., Postle K., Reznikoff W. S. The inverted repeats of Tn5 are functionally different. Cell. 1980 Mar;19(3):795–805. doi: 10.1016/s0092-8674(80)80055-9. [DOI] [PubMed] [Google Scholar]
- Rubin G. M., Hogness D. S. Effect of heat shock on the synthesis of low molecular weight RNAs in drosophilia: accumulation of a novel form of 5S RNA. Cell. 1975 Oct;6(2):207–213. doi: 10.1016/0092-8674(75)90011-2. [DOI] [PubMed] [Google Scholar]
- Shimotohno K., Mizutani S., Temin H. M. Sequence of retrovirus provirus resembles that of bacterial transposable elements. Nature. 1980 Jun 19;285(5766):550–554. doi: 10.1038/285550a0. [DOI] [PubMed] [Google Scholar]
- Strobel E., Dunsmuir P., Rubin G. M. Polymorphisms in the chromosomal locations of elements of the 412, copia and 297 dispersed repeated gene families in Drosophila. Cell. 1979 Jun;17(2):429–439. doi: 10.1016/0092-8674(79)90169-7. [DOI] [PubMed] [Google Scholar]
- Wensink P. C., Finnegan D. J., Donelson J. E., Hogness D. S. A system for mapping DNA sequences in the chromosomes of Drosophila melanogaster. Cell. 1974 Dec;3(4):315–325. doi: 10.1016/0092-8674(74)90045-2. [DOI] [PubMed] [Google Scholar]