The authors combine in vivo DMS modification with metabolic labeling with 4-thiouracil in order to follow kinetically the structural changes occurring during the maturation of RNA–protein complexes. They apply the method successfully to the pre-ribosomal particle assembly.
Keywords: ribosome synthesis, yeast, RNA structure, 4-thiouracil, ribonucleoprotein
Abstract
Pre-ribosomal particles undergo numerous structural changes during maturation, but their high complexity and short lifetimes make these changes very difficult to follow in vivo. In consequence, pre-ribosome structure and composition have largely been inferred from purified particles and analyzed in vitro. Here we describe techniques for kinetic analyses of the changes in pre-ribosome structure in living cells of Saccharomyces cerevisiae. To allow this, in vivo structure probing by DMS modification was combined with affinity purification of newly synthesized 20S pre-rRNA over a time course of metabolic labeling with 4-thiouracil. To demonstrate that this approach is generally applicable, we initially analyzed the accessibility of the region surrounding cleavage site D site at the 3′ end of the mature 18S rRNA region of the pre-rRNA. This revealed a remarkably flexible structure throughout 40S subunit biogenesis, with little stable RNA–protein interaction apparent. Analysis of folding in the region of the 18S central pseudoknot was consistent with previous data showing U3 snoRNA–18S rRNA interactions. Dynamic changes in the structure of the hinge between helix 28 (H28) and H44 of pre-18S rRNA were consistent with recently reported interactions with the 3′ guide region of U3 snoRNA. Finally, analysis of the H18 region indicates that the RNA structure matures early, but additional protection appears subsequently, presumably reflecting protein binding. The structural analyses described here were performed on total, affinity-purified, newly synthesized RNA, so many classes of RNA and RNA–protein complex are potentially amenable to this approach.
INTRODUCTION
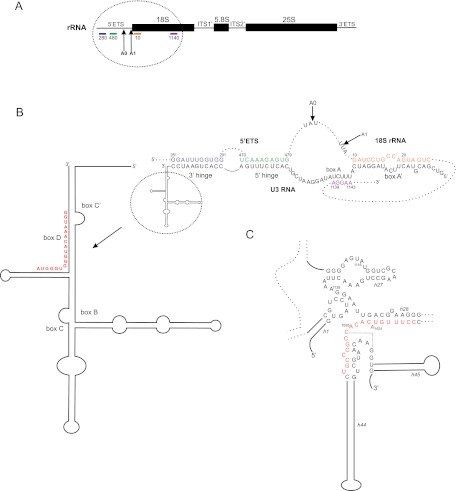
Ribosome synthesis in the yeast Saccharomyces cerevisiae is a complex, multistep pathway, involving more than 200 proteins and 75 small nucleolar RNAs (snoRNAs). The pre-rRNA is composed of the small subunit rRNA (18S) and large subunit rRNAs (25S and 5.8S), which are separated by internal transcribed spacers (ITS1 and ITS2) and flanked by external transcribed spacers (5′-ETS and 3′-ETS) (Fig. 1A). The nucleolar pre-rRNA is initially packaged with ribosomal proteins and ribosome synthesis factors and modified at many sites, most of which are selected by base-pairing with snoRNAs. A complex processing pathway then generates the mature rRNAs (Fig. 1B). During the early steps of pre-ribosome assembly, the U3 snoRNA base-pairs with the 5′-ETS and 18S regions of the pre-rRNA. These U3 interactions do not direct RNA modification but are required for the initial cleavage events at sites A0 and A1 in the 5′-ETS and at A2 in ITS1. Cleavage at site A2 splits the pre-ribosome into pre-40S and pre-60S complexes, which are independently exported to the cytoplasm, where the final maturation steps take place. Cytoplasmic processing of the 20S pre-rRNA to 18S rRNA, by cleavage of site D by the PINc endonuclease Nob1, is the last known event in maturation of the 40S ribosomal subunit (Udem and Warner 1973; Fatica et al. 2003; Pertschy et al. 2009), and takes place when the pre-40S particles have associated with the translation initiation factor eIF5b and mature 60S subunits (Lebaron et al. 2012; Strunk et al. 2012).
FIGURE 1.
Yeast pre-rRNA processing pathway. (A) Schematic representation of the yeast pre-rRNA showing hybridization probe locations. (B) Simplified schematic of the yeast pre-rRNA processing pathway.
Genetic analyses and mass spectrometry have identified many proteins required for rRNA processing and pre-ribosome assembly, and a range of techniques including cross-linking, electron microscopy (EM), and crystallography are revealing the details of pre-ribosome structure. Despite this, many unanswered questions remain about the structural rearrangement of rRNA and the many transient RNA–protein interactions that must occur during ribosome biogenesis. Several biochemical approaches based on the chemical modifications of RNA can potentially provide information about RNA structure within RNP complexes (Stern et al. 1988; Balzer and Wagner 1998; Liebeg and Waldsich 2009; Weeks 2010). However, most provide information about the complex at steady state and are less suited for analyses of dynamic structural changes in RNA and short-lived interactions. In an attempt to overcome these problems, we established an approach based on the combination of 4-thiouracil (4TU) labeling of nascent RNA in vivo (Cleary 2008; Zeiner et al. 2008) combined with DMS modification in vivo (Ares and Igel 1990b; King et al. 2003). DMS modification provides information about RNA structure and accessibility in living cells (Ares and Igel 1990a; Zaug and Cech 1995; Waldsich et al. 2002). DMS modifies the N1 position of adenosine and, to some extent, N3 of cytidine, unless they are involved in hydrogen-bond formation. The modification of uridine residues has also been observed, but the chemistry of U modification remains unclear (Wells et al. 2000). Following RNA extraction, incorporated 4TU residues can be biotinylated in vitro, allowing affinity purification of the RNA and conventional Northern and primer extension analyses. In this way, in vivo 4-thiouracil labeling can be used to determine the kinetics of RNA processing and identify intermediate products (Cleary 2008).
Here we demonstrate that the combination of these techniques is feasible and use this approach to address changes in the structure of maturing pre-40S ribosomal subunits.
RESULTS
Cleavage site D and the surrounding nucleotides are accessible throughout rRNA maturation
Incorporation of 4TU into RNA in wild-type yeast was low, whereas incorporation of 4-thio UTP during transcription run-on in permeablized cells appeared efficient (data not shown), strongly indicating that uptake was limiting. In an attempt to increase uptake, we expressed the uridine transporter Fui1 from a plasmid, and this resulted in enhanced incorporation of 4TU into RNA. For these studies, strain BY4741 had the URA3 gene restored by precise reintegration at the URA3 locus, to allow growth in medium lacking uracil, and was transformed with a multicopy plasmid overexpressing Fui1 under the control of its own promoter (pTA1). While this work was underway, the use of a different permease, the human equilibriative nucleoside transporter (hENT1), to enhance uptake of the related compound 4-thiouradine was reported independently (Miller et al. 2011).
The functionality of labeled ribosomes was not directly tested, but yeast cells labeled with 4TU or 4SU show no growth defects (data not shown) or changes in rates of mRNA accumulation (Miller et al. 2011; Sun et al. 2012), indicating the modified RNAs are fully functional.
The experimental design for purification and analysis of in vivo–modified yeast RNA is outlined in Figure 2A. To isolate newly synthesized pre-rRNA, 4-thiouracil (4TU) labeling was performed for short time courses of 2–7 min. During the final minute of these periods, DMS was immediately added to the yeast culture to modify the RNA. Ice-cold β-mercaptoethanol was then added, to simultaneously quench both DMS modification and 4TU uptake. To ensure complete quenching of the reactivity of DMS, isoamyl alcohol was also added. A range of conditions for DMS modification and quenching were assessed (data not shown), and the protocol given in Materials and Methods appeared optimal. Total RNA was extracted, the thio-ketone groups present in 4TU labeled RNA were reduced to sulfydryl groups by treatment with tris(2-carboxyethyl)phosphine (TCEP), and then biotinylated using HPDP-biotin. Biotinylated RNA was purified on streptavidin magnetic beads to isolate newly synthesized RNA, and sites of DMS modification were detected by primer extension.
FIGURE 2.
DMS modification in vivo and purification of newly synthesized rRNAs. (A) Schematic representation of the labeling and purification procedure, described in the text. (B) Northern analyses of newly synthesized rRNAs. One percent of total RNA and 100% of IP RNA were used for analysis. RNAs were separated on a 1.5% BPTE agarose gel and transferred to nylon membrane. Oligo probes that were used to monitor intermediates of rRNA processing and 18S rRNA are indicated.
The ability of this approach to monitor the processing of newly synthesized 20S pre-rRNA was assessed by Northern hybridization (Fig. 2B). After labeling for 2 min, 20S pre-rRNA was detected but not mature 18S rRNA, consistent with data based on metabolic radiolabeling of pre-rRNA in yeast with [3H] uracil (Kos and Tollervey 2010). Approximately 70% of nascent pre-rRNA transcripts are cleaved cotranscriptionally at sites A0–A2 to release 20S pre-RNA. This nascent transcript cleavage is responsible for the 20S detected at the 2-min time point. Subsequent maturation of 20S to 18S rRNA requires export to the cytoplasm and takes approximately a further 2 min (Kos and Tollervey 2010). Particularly at early time points, a major fraction of the labeled RNA is in the form of nascent transcripts, which generate the strong lane background.
To test whether chemical modification of the newly synthesized RNA can provide information regarding pre-rRNA structure, we analyzed the region surrounding cleavage site D. Cleavage of site D by the endonuclease Nob1 is the final step in maturation of 20S pre-rRNA to 18S rRNA (Fatica et al. 2003; Pertschy et al. 2009). Based on analysis at steady state, it was reported that the region flanking site D is accessible to chemical modification in pre-40S particles, indicating there are no stable structured elements in this region (Lamanna and Karbstein 2009, 2011; Granneman et al. 2010). However, it remained possible that more transient RNA structures or protein associations were formed during ribosome synthesis. To monitor potential structural rearrangements surrounding site D during pre-rRNA processing, DMS modification of newly synthesized RNA was performed. After 2 min of 4TU labeling, the DMS modification pattern was similar to the pattern obtained at steady state (Fig. 3A). All positions neighboring site D were modified with the same intensity as in total RNA (Fig. 3C,D), except for subtle differences in the reactivity of nucleotides 10 and 13, located at the 3′ side of site D.
FIGURE 3.
Accessibility of the region flanking cleavage site D to DMS modification during rRNA processing. (A) DMS modification in vivo. The 4TU labeling was performed for 2, 3, 5, and 7 min, with DMS added to the culture for the final minute. 4TU labeling and DMS modification were quenched together by addition of ice-cold β-mercaptoethanol and isoamyl alcohol. RNA was extracted and purified as described in Materials and Methods. Primer extension products were resolved on 12% PAA/7M UREA gel. The results of primer extension of total RNA and newly synthesized RNAs (4TU RNA) are presented on the autoradiogram. The lane described as “no 4TU” represents unlabeled RNA, which was mock-purified. The result from one experiment is presented on two autoradiograms. The first nucleotide at the 5′ side of the D cleavage site is numbered as position 1. The D cleavage site is indicated by the arrow. (Dotted line along the autoradiogram) The region that was quantified. Two adenosines in H45 dimethylated by Dim1 in vivo are marked by “M.” (B) DMS modification of RNA presented in purified pre-40S complex. The purification was performed using Tsr1-HTP (His6-TEV-ProtA) or PTH-Nob1 (ProtA-TEV-His6) as bait. The experiment was performed as described by Granneman et al. (2010). Primer 103 (complementary to ITS1) was used for primer extension through the site D region. (C) Semi-quantitative analysis of the DMS modification of total RNA relative to newly synthesized RNA. The columns represent averages, with error bars showing standard deviations from three independent in vivo experiments. In vitro structure probing was repeated twice. Due to degradation in position 11 of RNA probed in vitro, the intensity of modification was not measured. (D) DMS-modified nucleotides are marked on the secondary structure model of the 5′ end of ITS1 and 3′ end of the 18S rRNA region. The surrounding site D is accessible to modification in total RNA as well as in newly synthesized pre-rRNA. There are no statistically significant changes between the labeling of total RNA relative to newly synthesized RNA, so all positions are marked by gray dots.
To compare the structure of the region surrounding site D in vivo with the structure present in the pre-40S complex, the DMS modification was performed in vitro using tagged forms of the 40S ribosome synthesis factor Tsr1 or Nob1 as bait (Fig. 3B). Semi-quantitative analysis revealed minor changes in reactivity compared with modifications in vivo. However, all positions were modified, showing that the region surrounding the site D site is flexible. Stronger modification was observed close to site D, especially at position A2, adjacent to cleavage site (Fig. 3C; Supplemental Fig. S1). This indicates that single-stranded RNA is protruded in order to expose site D and facilitate the cleavage within the pre-40S particle. Overall, the structure of the 20S pre-rRNA appears very similar in the Tsr1–HTP and Nob1–HTP complexes (Fig. 3A; Supplemental Fig. S1). In vivo and in vitro A21 is not reactive, and only weak modifications are observed in the tract of four adenosine residues at positions 20–23 (indicated in Fig. 3A).
The observed modification pattern of newly synthesized, 4TU-selected RNA reflects pre-rRNA folding at early steps of rRNA biogenesis, whereas 20S pre-rRNA present in the purified pre-40S particle represents later steps in maturation. We therefore conclude that the region around site D is unstructured during pre-rRNA processing.
DMS modification of newly synthesized RNA is consistent with U3 snoRNA–rRNA interactions in the central region of the pre-18S rRNA
The U3 snoRNA makes base-paired interactions with the pre-rRNA at two sites in the 5′-ETS (see Fig. 4A,B; Beltrame and Tollervey 1995; Dutca et al. 2011) and in the loop of helix 1 (H1) at the 5′ end of the 18S rRNA (Hughes et al. 1987; Sharma and Tollervey 1999). In the mature 40S ribosome, these H1 loop nucleotides form a conserved long-range interaction, the “central pseudoknot,” with 18S rRNA nucleotides 1140–1142, and U3 was also proposed to base-pair with these nucleotides (see Fig. 4B; Sharma and Tollervey 1999). Mutational analysis was unable to confirm the interaction between U3 and the 3′ side of the 18S pseudoknot (Sharma and Tollervey 1999), but structure probing was consistent with this base-pairing (Dutca et al. 2011). To assess the kinetics of this potential interaction, we probed the central region of 18S rRNA in 4TU-labeled samples. The DMS modifications pattern of newly synthesized RNA supports the presence of the pre-rRNA–U3 interaction in early pre-ribosomes (Fig. 5; Supplemental Fig. S2). Strongly decreased reactivity of A1139 at the 5′ side of the pseudoknot was observed in newly synthesized RNA, consistent with U3 base-pairing (Fig. 4B). In strains depleted for U3 snoRNA, A1139 was reported to be fully accessible for DMS modification (Dutca et al. 2011). At later times of 4TU labeling, the reactivity of A1139 was increased, consistent with the accumulation of the mature pseudoknot structure of the 18S rRNA (Fig. 5; Supplemental Fig. S2). The reactivity of A1138, which lies adjacent to nucleotides involved in the U3 interaction, is unchanged during the time course. Lack of modification of A1142 can be explained if, as expected, the pre-rRNA interaction with U3 is rapidly followed by formation of the 18S pseudoknot. No strong modifications were observed in the surrounding central regions of 18S rRNA (Fig. 5B), consistent with previous data (Dutca et al. 2011).
FIGURE 4.
Interactions between U3 snoRNA and the pre-rRNA. (A) U3 interacts with the 5′ ETS at sites around positions +280 and +480 (Beltrame and Tollervey 1995; Dutca et al. 2011), and with the 18S rRNA around +10 and +1140 (Hughes et al. 1987; Sharma and Tollervey 1999; Kudla et al. 2011). (B) Sites of U3 binding along the pre-rRNA. Colors match the rRNA regions highlighted in panel A. (C) U3 interactions detected by CLASH within the 18S rRNA region. (B,C, red) The U3 and rRNA sequences involved in the interaction.
FIGURE 5.
Changes in base-pairing in the central part of 18S rRNA revealed by different access to DMS modification in newly synthesized RNA. (A) DMS modification in vivo of the central part of 18S rRNA. The 4TU labeling was performed for 2, 3, and 7 min, with DMS added to the culture for the final minute. DMS modification and RNA purification were performed as described in the legend of Figure 3 and the text. Four sequencing lanes are presented. Primer 101 (complementary to 18S H31) was used for primer extension through the central pseudoknot region. The nucleotide positions are marked on the right side of the gel. (Dotted line along the autoradiogram) The region that was quantified. The three “Total RNA” lanes are technical replicates. (B) Semi-quantitative analysis of the DMS modification of total RNA relative to newly synthesized RNAs. The bars represent averages, with error bars showing standard deviations based on biological replicates from three independent experiments. (C) Changes in the intensity of DMS modifications in newly synthesized RNA compared with total RNA displayed on the secondary structure model of the analyzed part of rRNA. The modifications whose intensity is below 2.5% are not presented on the structure. (Red) rRNA sequence involved in interactions with the U3 snoRNA.
Analysis of new interactions between U3 snoRNA and 18S rRNA
U3 is a member of the box C/D class of snoRNAs, and the characteristic proteins (Snu13, Nop1, Nop56, and Nop58) bind to the 3′ region of U3, which carries homologs of the C, C′, D, and D′ boxes. The interactions described above involve the 5′ region of the U3 snoRNA (see Fig. 4), whereas in all other characterized members of the large box C/D family, the pre-rRNA binding sites lie immediately adjacent to the D and D′ boxes. The mode of U3 binding therefore appeared to be different from all other box C/D snoRNAs. However, a recent analysis using cross-linking and sequencing of hybrids (CLASH) revealed further potential U3/18S rRNA interactions (see Fig. 4B,C; Kudla et al. 2011). This identified base-pairing between 18S sequences in the H28–H44 hinge region close to the 3′ side of the pseudoknot and a sequence in U3 that flanks the 3′ D box. This suggested that additional U3 interactions might aid pre-rRNA folding to bring together the 5′ and 3′ sides of the central pseudoknot, which involves only 3 bp between sequences that are separated by >1100 nt in the primary sequence. We therefore examined the structure of the central region of 18S including the newly identified U3 interaction site. To test the proposed U3–pre-RNA interaction, we initially analyzed the predicted secondary structure folding in silico. Two alternative models can be drawn, with similar free energy values (Fig. 6).
FIGURE 6.
Potential U3 snoRNA–pre-rRNA interactions. Secondary structure models of U3–pre-rRNA chimeras. Changes in the intensity of DMS modifications in newly synthesized RNA after 4 min of labeling compared with total RNA displayed on the secondary structure model of the analyzed part of rRNA. Based on the CLASH data, part of helix 3 and helix 5 of U3 snoRNA can interact with the region close to the 3′ side of the pseudoknot of 18S rRNA (Kudla et al. 2011). Alternative models for the structure of the U3 snoRNA–rRNA chimera, as predicted by the RNAStructure program. The predicted stable stem formed between the 3′ region of the rRNA and U3 snoRNA is supported by the lack of modification at C1636 and C1637. Higher reactivity of A1635 fits with both chimera structure models. Based on both the structure predictions and DMS modification patterns, the central region of the chimera is less stable with the potential for structural rearrangements.
In vivo structure probing revealed a strong primer extension stop at position C1639 (Fig. 7A; Supplemental Fig. S3) generated by endogenous RNA 2′-O-methylation, which made it difficult to perform accurate quantitative analysis. However, band intensities were measured, and the scanned lanes are presented on the graphs. DMS patterns at steady state were consistent with the secondary structure model of mature rRNA. Modification was observed at A1631, located in a bulge, and C1634, located in a single-stranded region (Figs. 6, 7A; Supplemental Fig. S3).
FIGURE 7.
Testing potential U3 snoRNA–pre-rRNA interactions. (A) Comparison of DMS modifications patterns of total RNA and newly synthesized RNAs. The nucleotide positions are indicated on the side of the gel. A primer extension stop generated by natural modification of C1639 is indicated (1639C-2′-O-methyl). The graph presents the intensity of bands in each lane: (black line) total RNA; (red line) 2 min; (green line) 4 min; (blue line) 6 min of 4TU labeling. (B) DMS modification of RNA present in purified pre-40S complex. The purification was performed using Tsr1-HTP as bait. 18S and 20S rRNAs were recovered as described for panel C. The graph presents the intensity of bands in the lane showing modification of 20S rRNA. Primer 104 (complementary to the apical loop of 18S rRNA helix 44) was used in all experiments. All experiments were repeated twice. (C) Isolation of 18S and 20S rRNA. RNA was extracted from purified pre-40S complexes. DMS modified (+) and unmodified (−) RNA was separated on a 2% agarose, BPTE gel. The gel was stained by SYBR to visual the RNA, the 18S and 20S regions were excised form the cut gel, and RNA was eluted.
C1636 was protected in total RNA, and the 80S ribosome crystal structure (Ben-Shem et al. 2010, 2011) indicates that it is involved in tertiary interaction with G1767, located in the junction between H44 and H45. However, C1637 and A1635 were also unmodified in total RNA, but these positions are unpaired and make no mapped ribosomal protein interactions. Both nucleotides, A1635 and C1637, are very close to the decoding center, so it is conceivable that interactions with the translational machinery results in the observed lack of DMS modification in total RNA. The 80S crystal structure was derived from ribosomes purified from non-translating cells, and it is very likely that these are not completely identical to the ribosomes present in actively growing cells.
For newly synthesized pre-rRNA, detected after 2 and 4 min of 4TU labeling, modification was strongest at position A1635 (Fig. 7A; Supplemental Fig. S3). After longer times of labeling, the reactivity of nucleotides in the H28/H44 hinge was substantially different. All positions were modified, with the strongest modifications observed at positions C1632 and A1635 (Fig. 7A; Supplemental Fig. S3). To obtain further details concerning the structure of the H28/H44 hinge, chemical probing was performed in vitro on pre-40S complexes purified using Trs1-HTP as bait. The 20S pre-rRNA was modified in vitro, extracted, and resolved on agarose gels to separate 18S rRNA from 20S pre-rRNA (Fig. 7C). The DMS modification pattern of isolated 20S rRNA resembles the DMS pattern for newly synthesized, in vivo rRNA after 6 min of labeling, except for C1634, which is not reactive in 20S pre-rRNA in vitro (Fig. 7A,B). These results provided strong evidence that structures present at early and late steps of rRNA biogenesis can be distinguished. The DMS pattern obtained in vivo after 6 min of labeling shows the combined reactivity of nucleotides in 20S pre-rRNA and mature 18S rRNA, since both are recovered at this time point.
The primer-extension signal after 2 min of labeling was faint, so we compared the DMS pattern obtaining after 4 min of labeling with modification patterns observed at steady state. At steady state, the strongest modification was observed at position C1634, whereas A1635 is the most reactive in newly synthesized, 4TU-labeled RNA. This observation is consistent with either chimera structure model, since this position is bulged in both. The central nucleotides of the U3–pre-RNA chimera are different in the two proposed models. Model 1 includes a 3-nt bulge, whereas model 2 includes two single bulged nucleotides separated by a 2-nt stem (Fig. 6). The lack of reactivity of C1634 in newly synthesized RNA (Fig. 7A) fits with model 2, in which this nucleotide is base-paired (Fig. 6). The lack of modification at A1633 in both steady-state and newly synthesized RNA also supports model 2 (Fig. 6). However, model 2 would also predict higher DMS modification of A1631 and C1632 relative to A1633 and C1634 on RNA extracted at early time points of 4TU labeling, which is not observed.
We envisage that the U3–pre-RNA interaction is dynamic and some discrepancy between DMS modification pattern and structure model results from rearrangements in base-pairing.
Our results indicate that the H28/H44 hinge region is repeatedly refolded during rRNA biogenesis. DMS modification in vivo provides experimental data that support the interaction between the U3 snoRNA guide region and the pre-RNA.
Changes in the structure of helix 18 during maturation
Figures 6 and 7 showed that the structural neighborhood on the 3′ side of the central pseudoknot is rearranged during rRNA processing. We therefore tested whether the structure 5′ to the central pseudoknot sequence, helix 18 (H18), is also refolded during pre-ribosome maturation.
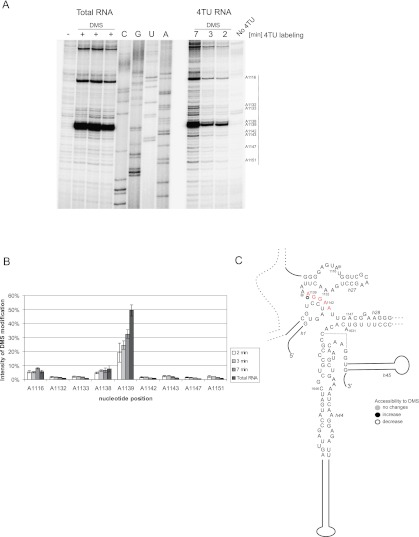
The DMS modification pattern of H18 in newly synthesized rRNA (Fig. 8A,B) is in good agreement with the tertiary structure derived from the yeast ribosome crystal structure (Fig. 8C,E; Ben-Shem et al. 2010). Strongly modified nucleotides at positions C554, A567, A570, and A579 (Fig. 8A,B) are all located in single-stranded regions (Fig. 8C,E). The crystal structure of the yeast ribosome also revealed further details about the pseudoknot in H18. Base-pairing was previously proposed between three nucleotides, G(552)GC, located in a bulge, and nucleotides G(571)CC, located in the apical loop of the stem (Fig. 8C,E). The crystal structure showed that C554 can stabilize this pseudoknot by stacking interactions, but the N3 position is not involved in hydrogen-bond formation (Fig. 8E). The reactivity seen for C554 in our analysis is therefore consistent with the crystal structure. Cytidines C572 and C573, which are base-paired within the pseudoknot (Fig. 8C), were unmodified in both the purified nascent RNA and total RNA. Position A579 is modified in newly synthesized rRNA and in RNA at steady state. We therefore conclude that the global structure of H18 is similar in newly synthesized and mature 18S rRNA, and the pseudoknot is formed at an early step of subunit maturation.
FIGURE 8.
Structure of helix 18. (A) In vivo DMS modification of helix 18 (H18) of rRNA. The 4TU labeling was performed for 2, 4, and 6 min. Lane C is an additional control with 6 min of 4TU labeling without DMS addition. The results from one experiment are visualized on two autoradiograms. Primer 102 (complementary to 18S H21) was used for primer extension analyses of the H18 region. (B) The graphs present the intensity of bands in each lane: newly synthesized RNA: (red line) 2 min; (green line) 4 min; (blue line) 6 min of 4TU labeling and total RNA (black line on the lower graph). (C) Secondary structure model of H18. Nucleotides involved in the pseudoknot are boxed. DMS modified positions, C554, A567, A570, and A579, which are discussed in the text, are marked on the structure. (D) The same positions are indicated on the tertiary structure of H18 (Ben-Shem et al. 2010). (E) Structure of the pseudoknot with C554 highlighted in blue, which takes part in the stacking interaction. The PyMol program was used to present tertiary structure of H18 (data from NDB ID: NA0650) (Ben-Shem et al. 2010, 2011).
However, in vivo chemical modification revealed some clear differences between newly synthesized rRNA and total RNA (Fig. 8A). Positions A567 and A570 were substantially more accessible to modification in newly synthesized RNA than in total RNA, which is predominately mature 18S (Fig. 8A,B). Decreased reactivity at A555 and A556 was also seen in total RNA. The protection of A567 and A570 in total RNA may reflect the assembly of proteins, particularly Rps23, onto 18S rRNA in this region (Supplemental Fig. S4). Site-directed mutagenesis of the rRNA showed that H18 is a functional element of the ribosome decoding center, acting together with H34 and part of H44 (O'Connor et al. 1997).
In vivo chemical modifications of newly synthesized rRNA showed that the final structure of H18 is formed early in pre-rRNA processing. The DMS modification pattern of newly synthesized rRNA is generally consistent with crystal structure of H18 in mature yeast ribosomes. However, the increased protection in total RNA likely reflects the assembly of proteins with this region.
DISCUSSION
The determination of the in vivo structure of large, highly dynamic, RNA–protein complexes such as pre-ribosomes has always been very challenging. For this reason, most analyses have been performed in vitro using purified complexes. This allows a range of structure probing reagents to be brought to bear on the problem but has the disadvantage that labile factors and structures may be lost during purification. Moreover, multiple states of the in vivo complex are not resolved and will be analyzed together. In vivo structure probing is feasible using DMS-induced methylation, which predominately modifies solvent-accessible A and C residues. Among other applications, this approach has been used on yeast pre-ribosomes to assess the structure surrounding cleavage site D at the 3′ end of the 18S rRNA (Lamanna and Karbstein 2009; Pertschy et al. 2009; Granneman et al. 2010) and interactions between U3 snoRNA and the central pseudoknot region (Dutca et al. 2011). While these analyses were undoubtedly informative, they provided an ensemble view of pre-ribosomes all along the maturation pathway. To address this problem, we provide a tool for kinetic analysis of in vivo assembly of pre-ribosomes and other complex RNPs. The analysis of pre-rRNA processing by metabolic labeling is a long-established approach. This has traditionally involved the use of [3H]-uracil to allow newly synthesized molecules to be specifically identified. Initially, pulse-chase labeling was used; however, pulse labeling without chase is more readily analyzed kinetically, with short-lived intermediates being assessed during the approach to steady state (Kos and Tollervey 2010). In the present study, we adopted a related method: metabolic pulse-only labeling, but using 4-thiouracil to follow the newly synthesized population. In vitro biotinylation of 4-thiouracil-labeled RNA allows its purification on streptavidin beads under very stringent conditions, with a consequent high degree of enrichment.
Using this technique, we were able to resolve changes in pre-ribosome structure that take place on a minute time scale. There are, of course, limitations to this approach. Principal among these is the requirement for suitably positioned A or C residues, which show clearly distinguished alterations in accessibility. This limitation was seen, for example, in the case of the proposed U3-18S interactions, where both the mature 18S and the U3-18S stems protect the same nucleotides, while the opposite side of the 18S stem lacks potentially reactive nucleotides. Clear differences, however, were seen in the H18 region of the decoding center, which plays a major role in aa-tRNA recognition in the ribosomal A-site (Ogle et al. 2003). Here the RNA stem and pseudoknot structure appears to form early, but protection of single-stranded nucleotides was greatly increased over time, indicative of protein binding. In cases in which the RNA population is heterogeneous, we predominately observe the modification pattern for the most abundant species, as is the case for many structural analyses. However, the method described here is an advance over previous, steady-state, approaches because RNA populations that are dominant at different time points can potentially be distinguished (as shown, e.g., in Fig. 5). In addition, the method can be used to assess whether the modification pattern reflects a population of different RNA structures. This was seen for the central pseudoknot (Fig. 5) and for the hinge region between H28 and H44 (Figs. 6, 7). In analyses at steady state, such information is lost.
Because the total 4TU-labeled RNA population is recovered following DMS treatment, these analyses can, in principal, be applied to any in vivo RNA or RNA–protein complex, most of which are less well characterized than are the pre-ribosomes. The time course of labeling is limited to ∼1 min resolution, but this should be sufficient for most analyses, particularly in higher eukaryotes.
MATERIALS AND METHODS
Strains and media
S. cerevisiae strain BY4741 with URA3 restored expressing the uridine permease Fui1 from the pAT1 plasmid was used in all in vivo DMS modification experiments. Cells were grown in SD −Ura −Leu at 30°C. S. cerevisiae strain D1091 (BY4741 with Trs1-HTP∷URA3) and strain BY4741 with the pADH-HTP-Nob1 plasmid were used to purify pre-40S complex (Granneman et al. 2010). Cells were grown in YPD at 30°C. The FUI1 gene, including the entire coding region plus 474 bases upstream and 191 bases downstream, was amplified from S. cerevisiae genomic DNA using primers Fui1_F1_Xho1 and Fui1_R1. The PCR primers used introduced novel XmaI and SacII sites at the 5′ and 3′ ends of FUI1, respectively. These sites were used to introduce the PCR fragment into the 2μ plasmid pRS426 at its multiple cloning site. The insert and plasmid were mixed in a 5:1 ratio and ligated using T4 DNA ligase overnight at 16°C. The FUI1 insert was verified by sequencing. To restore the URA3 gene in BY4741, S. cerevisiae URA3 was amplified by PCR from strain pSG100 using primers URA3_int1 and URA3_int 2. The resulting PCR product was used to transform BY4741, restoring the URA3 gene at its endogenous locus by homologous recombination. Correct URA3 insertion was confirmed by PCR.
Chemical modification in vivo and in vitro
DMS modification in vivo was performed followed the published protocol (Ares and Igel 1990b; King et al. 2003). Cells were grown to OD600 = 0.5–0.6, then 4-thiouracil labeling was performed for the times indicated, with DMS added during the final minute of labeling. Respectively after each time point of 4-thiouracil uptake, 25 mL of culture was taken, and 500 μL of DMS, freshly diluted 1∷2 in 95% ethanol, was added to the culture. RNA was modified for 1 min at 25°C with shaking. The reaction was quenched by 20 mL of 0.7 M ice-cold β-mercaptoethanol and 10 mL of water-saturated isoamyl alcohol. The cells were centrifuged, and the pellet was washed once more with 15 mL of 0.7 M cold β-mercaptoethanol and then twice with cold water.
Pre-40S complexes were prepared by one-step purification on IgG Sepharose beads as previously described (Granneman et al. 2010). DMS, diluted freshly 1∷10 in 95% ethanol, was added to the pre-40S complex presented on IgG Sepharose beads to the final concentration of 5% (5% of 1∷10 DMS solution). The reaction was stopped by adding an equal volume of a mix containing 0.5 M β-mercaptoethanol and 1.5 M sodium acetate (pH 5.3). RNA was immediately extracted by GTC phenol mix (4 M guanidine thiocyanate, 0.05 M Tris at pH 8.0, 0.01 M EDTA, 2% sarcosyl, 1% β-mercaptoethanol, 50% [v/v] phenol) (Sambrook et al. 1989).
Isolation of 18S and 20S RNA
RNA was resolved on 2% agarose, BPTE—10 mM PIPES (2-[4-(2-sulfoethyl)piperazin-1-yl]ethanesulfonic acid, 30 mM Bis-Tris, 1 mM EDTA at pH 8.0—gels (Sambrook et al. 1989). Electrophoresis was performed for 20 h at 60 V. The gel was stained by SYBR to visual RNA, and then 18S rRNA and 20S rRNA were cut and eluted. RNA was eluted using QG buffer (QIAGEN) and incubated at 42°C until dissolved. Then RNA was purified on columns (MiniElute-spin column 50; QIAGEN). RNA was precipitated using 3 volumes of cold ethanol and 2 μL of 20 mg/mL glycogen for 20 min at −20°C.
4-Thiouracil (4TU) labeling and purification
Cells containing the pAT1 plasmid with uridine permease Fui1 were grown in SD −Ura −Leu to OD600 = 0.6–0.75. 4-Thiouracil was added to the culture to a final concentration of 100 μM.
4-Thiouracil uptake and DMS reactivity
DMS and 4TU were quenched by 20 mL of 0.7 M ice-cold β-mercaptoethanol and 10 mL of water-saturated isoamyl alcohol. The next wash was performed using only 15 mL of 0.7 M ice-cold β-mercaptoethanol. The cell pellet was washed twice with cold water. RNA was extracted by hot phenol extraction at 60°C, and thio-ketone groups (C=S) of RNA were reduced to sulfhydryl groups (C—SH) by treatment with TCEP, tris(2-carboxyethyl)phosphine. RNA was biotinylated with a 0.4 mM final concentration of HPDP-biotin diluted in dimethylformamid, and then newly synthesized RNAs were recovered by binding to streptavidin magnetic beads. RNA was eluted from the beads twice in 150 μL of 0.1 M β-mercaptoethanol. Fifty percent of the total amount of recovered RNA was used for one primer extension reaction.
Reduction step
Total RNA was reduced with 20 μL of TECP agarose slurry (Pierce) in 50 mM Tris-HCl (pH 7.8), 1 mM EDTA in a total volume of 110 μL for 2 h at 16°C.
Biotinylation of RNA
RNA was biotinylated with 25 μL of HPDP-biotin (Pierce) (4 mM in dimethylformamid stock) in a total volume of 250 μL for 3 h at 25°C in the dark.
RNA was extracted once with chloroform/isoamyl alcohol and precipitated with 3 volumes of cold ethanol for 1–3 h at −20°C.
Purification of newly synthesized RNA
Newly synthesized RNA was bound to 50 μL of streptavidin magnetic beads (Roche) that were preblocked with 200 μg/mL glycogen (Roche) in RBS buffer (10 mM Tris at pH 7.5, 100 mM NaCl, 2.5 mM MgCl2, 0.4% Triton X-100) for 0.5 h at 25°C. Binding RNA to beads was performed for 1 h at 25°C. Then beads were washed three times with RBS buffer, twice with buffer (10 mM Tris-HCl at pH 7.5, 1 mM EDTA at pH 8.0, 1 M NaCl), and twice with water.
Elution
Newly synthesized RNA was eluted twice with 150 μL of 0.1 M β-mercaptoethanol.
Precipitation
RNA was precipitated with 0.1 volume of 3 M NaOAc (pH 5.2), 3 volumes of cold ethanol, and 2 μL of 20 mg/mL glycogen overnight at −20°C. Following precipitation, RNA was resuspended in 8 μL of water. Fifty percent of the total amount of RNA was used for primer extension, which is sufficient for five separate reactions.
Primer extension and Northern blotting
DMS modification positions were mapped by primer extension. The primer extension reaction was performed using a 0.5 mM final concentration of each dNTP in the presence of 50 units of reverse transcriptase (SuperScript III, Invitrogen) for 50 min at 50°C. The reaction was stopped by addition of an equal volume of loading dye and heating for 3 min at 85°C. One microgram of total RNA, 50% of total amount of newly synthesized RNAs, and ∼100 ng of purified RNA were used for each primer extension reaction. Semi-quantitative analysis of gel band intensities was performed using the adaptive image deconvolution algorithm (AIDA) (Hom et al. 2007). The intensity of the line was measured, and the percentage of each band in this line was calculated. The intensities were then compared for relevant bands (value of percentage) in total RNA versus newly synthesized RNA. The sequencing ladder was generated using 1–5 μg of unmodified total RNA. The final concentration of dideoxy terminating nucleotides (ddNTP) was 0.25 mM. Primer extension products were resolved on a 12% PAA/7 M urea gel. The primers used for primer extension reactions are listed in Table 1.
TABLE 1.
Oligonucleotides used in this work

Northern hybridization was preformed as described (Tollervey 1987). One microgram of total RNA and 100% of the total amount of newly synthesized RNAs were resolved on 1% agarose, BPTE—10 mM PIPES (2-[4-(2-sulfoethyl)piperazin-1-yl]ethanesulfonic acid), 30 mM Bis-Tris, 1 mM EDTA at pH 8.0—gels and then transferred on Hybond N+ membranes, which were hybridized using probes labeled with [γ-32P]ATP and directed against different intermediates of rRNA processing (Table 1).
SUPPLEMENTAL MATERIAL
Supplemental material is available for this article.
ACKNOWLEDGMENTS
The Wellcome Trust supported A.S., D.T. (077248), A.T. (086574), and D.B. (087551). W.W. and D.B. were supported by the EU UniCellSys network. Work in the Wellcome Trust Centre for Cell Biology is supported by Wellcome Trust core funding (092076). We thank the Tollervey laboratory for support and advice, particularly Claudia Schneider.
Footnotes
Article published online ahead of print. Article and publication date are at http://www.rnajournal.org/cgi/doi/10.1261/rna.034751.112.
REFERENCES
- Ares M Jr, Igel AH 1990a. Mutations define essential and nonessential U2 RNA structures. Mol Biol Rep 14: 131–132 [DOI] [PubMed] [Google Scholar]
- Ares M Jr, Igel AH 1990b. Lethal and temperature-sensitive mutations and their suppressors identify an essential structural element in U2 small nuclear RNA. Genes Dev 4: 2132–2145 [DOI] [PubMed] [Google Scholar]
- Balzer M, Wagner R 1998. A chemical modification method for the structural analysis of RNA and RNA–protein complexes within living cells. Anal Biochem 256: 240–242 [DOI] [PubMed] [Google Scholar]
- Beltrame M, Tollervey D 1995. Base-pairing between U3 and the pre-ribosomal RNA is required for 18S rRNA synthesis. EMBO J 14: 4350–4356 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ben-Shem A, Jenner L, Yusupova G, Yusupov M 2010. Crystal structure of the eukaryotic ribosome. Science 330: 1203–1209 [DOI] [PubMed] [Google Scholar]
- Ben-Shem A, Garreau de Loubresse N, Melnikov S, Jenner L, Yusupova G, Yusupov M 2011. The structure of the eukaryotic ribosome at 3.0 Å resolution. Science 334: 1524–1529 [DOI] [PubMed] [Google Scholar]
- Cleary MD 2008. Cell type–specific analysis of mRNA synthesis and decay in vivo with uracil phosphoribosyltransferase and 4-thiouracil. Methods Enzymol 448: 379–406 [DOI] [PubMed] [Google Scholar]
- Dutca LM, Gallagher JE, Baserga SJ 2011. The initial U3 snoRNA:pre-rRNA base pairing interaction required for pre-18S rRNA folding revealed by in vivo chemical probing. Nucleic Acids Res 39: 5164–5180 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fatica A, Oeffinger M, Dlakic M, Tollervey D 2003. Nob1p is required for cleavage of the 3′ end of 18S rRNA. Mol Cell Biol 23: 1798–1807 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Granneman S, Petfalski E, Swiatkowska A, Tollervey D 2010. Cracking pre-40S ribosomal subunit structure by systematic analyses of RNA–protein cross-linking. EMBO J 29: 2026–2036 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hom EFY, Marchis F, Lee TK, Haase S, Agard DA, Sedat JW 2007. AIDA: An adaptive image deconvolution algorithm with application to multi-frame and three-dimensional data. J Opt Soc Am A Opt Image Sci Vis 24: 1580–1600 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hughes JMX, Konings DAM, Cesareni G 1987. The yeast homologue of U3 snRNA. EMBO J 6: 2145–2155 [DOI] [PMC free article] [PubMed] [Google Scholar]
- King TH, Liu B, McCully RR, Fournier MJ 2003. Ribosome structure and activity are altered in cells lacking snoRNPs that form pseudouridines in the peptidyl transferase center. Mol Cell 11: 425–435 [DOI] [PubMed] [Google Scholar]
- Kos M, Tollervey D 2010. Yeast pre-rRNA processing and modification occur cotranscriptionally. Mol Cell 37: 809–820 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kudla G, Granneman S, Hahn D, Beggs J, Tollervey D 2011. Mapping in vivo RNA–RNA interactions by crosslinking, ligation and sequencing of hybrids. Proc Natl Acad Sci 108: 10010–10015 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lamanna AC, Karbstein K 2009. Nob1 binds the single-stranded cleavage site D at the 3′-end of 18S rRNA with its PIN domain. Proc Natl Acad Sci 106: 14259–14264 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lamanna AC, Karbstein K 2011. An RNA conformational switch regulates pre-18S rRNA cleavage. J Mol Biol 405: 3–17 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lebaron S, Schneider C, van Nues RW, Swiatkowska A, Walsh D, Bottcher B, Granneman S, Watkins NJ, Tollervey D 2012. Proofreading of pre-40S ribosome maturation by a translation initiation factor and 60S subunits. Nat Struct Mol Biol 19: 744–753 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liebeg A, Waldsich C 2009. Probing RNA structure within living cells. Methods Enzymol 468: 219–238 [DOI] [PubMed] [Google Scholar]
- Miller C, Schwalb B, Maier K, Schulz D, Dumcke S, Zacher B, Mayer A, Sydow J, Marcinowski L, Dolken L, et al. 2011. Dynamic transcriptome analysis measures rates of mRNA synthesis and decay in yeast. Mol Syst Biol 7: 458 doi: 10.1038/msb.2010.112 [DOI] [PMC free article] [PubMed] [Google Scholar]
- O'Connor M, Thomas CL, Zimmermann RA, Dahlberg AE 1997. Decoding fidelity at the ribosomal A and P sites: Influence of mutations in three different regions of the decoding domain in 16S rRNA. Nucleic Acids Res 25: 1185–1193 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ogle JM, Carter AP, Ramakrishnan V 2003. Insights into the decoding mechanism from recent ribosome structures. Trends Biochem Sci 28: 259–266 [DOI] [PubMed] [Google Scholar]
- Pertschy B, Schneider C, Gnadig M, Schafer T, Tollervey D, Hurt E 2009. RNA helicase Prp43 and its co-factor Pfa1 promote 20 to 18 S rRNA processing catalyzed by the endonuclease Nob1. J Biol Chem 284: 35079–35091 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sambrook J, Fritsch EF, Maniatis T 1989. Molecular cloning: A laboratory manual, 2nd ed. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY [Google Scholar]
- Sharma K, Tollervey D 1999. Base pairing between U3 small nucleolar RNA and the 5′ end of 18S rRNA is required for pre-rRNA processing. Mol Cell Biol 19: 6012–6019 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stern S, Moazed D, Noller HF 1988. Structural analysis of RNA using chemical and enzymatic probing monitored by primer extension. Methods Enzymol 164: 481–489 [DOI] [PubMed] [Google Scholar]
- Strunk BS, Novak MN, Young CL, Karbstein K 2012. A translation-like cycle is a quality control checkpoint for maturing 40S ribosome subunits. Cell 150: 111–121 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun M, Schwalb B, Schulz D, Pirkl N, Etzold S, Larivière L, Maier KC, Seizl M, Tresch A, Cramer P 2012. Comparative dynamic transcriptome analysis (cDTA) reveals mutual feedback between mRNA synthesis and degradation. Genome Res 22: 1350–1359 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tollervey D 1987. A yeast small nuclear RNA is required for normal processing of pre-ribosomal RNA. EMBO J 6: 4169–4175 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Udem SA, Warner JR 1973. The cytoplasmic maturation of a ribosomal precursor ribonucleic acid in yeast. J Biol Chem 248: 1412–1416 [PubMed] [Google Scholar]
- Waldsich C, Grossberger R, Schroeder R 2002. RNA chaperone StpA loosens interactions of the tertiary structure in the td group I intron in vivo. Genes Dev 16: 2300–2312 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weeks KM 2010. Advances in RNA structure analysis by chemical probing. Curr Opin Struct Biol 20: 295–304 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wells SE, Hughes JM, Igel AH, Ares M Jr 2000. Use of dimethyl sulfate to probe RNA structure in vivo. Methods Enzymol 318: 479–493 [DOI] [PubMed] [Google Scholar]
- Zaug AJ, Cech TR 1995. Analysis of the structure of Tetrahymena nuclear RNAs in vivo: Telomerase RNA, the self-splicing rRNA intron, and U2 snRNA. RNA 1: 363–374 [PMC free article] [PubMed] [Google Scholar]
- Zeiner GM, Cleary MD, Fouts AE, Meiring CD, Mocarski ES, Boothroyd JC 2008. RNA analysis by biosynthetic tagging using 4-thiouracil and uracil phosphoribosyltransferase. Methods Mol Biol 419: 135–146 [DOI] [PubMed] [Google Scholar]