Abstract
We have examined the alternative splicing of the Drosophila melanogaster prospero twintron, which contains splice sites for both the U2- and U12-type spliceosome and generates two forms of mRNA, pros-L (U2-type product) and pros-S (U12-type product). We find that twintron splicing is developmentally regulated: pros-L is abundant in early embryogenesis while pros-S displays the opposite pattern. We have established a Kc cell in vitro splicing system that accurately splices a minimal pros substrate containing the twintron and have examined the sequence requirements for pros twintron splicing. Systematic deletion and mutation analysis of intron sequences established that twintron splicing requires a 46-nucleotide purine-rich element located 32 nucleotides downstream of the U2-type 5′ splice site. While this element regulates both splicing pathways, its alteration showed the severest effects on the U2-type splicing pathway. Addition of an RNA competitor containing the wild-type purine-rich element to the Kc extract abolished U2-type splicing and slightly repressed U12-type splicing, suggesting that a trans-acting factor(s) binds the enhancer element to stimulate twintron splicing. Thus, we have identified an intron region critical for prospero twintron splicing as a first step towards elucidating the molecular mechanism of splicing regulation involving competition between the two kinds of spliceosomes.
Introns are removed from pre-mRNA by two transesterification steps catalyzed by a large multicomponent complex known as the spliceosome. Five small nuclear RNAs, U1, U2, U4/U6, and U5, and more than 60 polypeptides form the active U2-type spliceosome (5). Spliceosome assembly proceeds in an ordered fashion that is directed by the recognition of conserved sequence motifs within the pre-mRNA by small nuclear RNAs and protein factors. These sequences allow efficient and accurate removal of introns and are located at the 5′ and 3′ splice sites as well as the branch point (43).
A small subset of introns is removed via a unique and divergent spliceosome (U12-type), whose composition and splice site recognition signals differ (22, 56). U11, U12, and U4atac/U6atac are the functional analogues of U1, U2, and U4/U6 small nuclear RNPs in the U12-type spliceosome; U5 is the only small nuclear RNP shared by both types of spliceosomes (22, 32, 34, 61). As in its U2-type counterpart, interactions of conserved sequence elements at the 5′ and 3′ splice sites and branch point with the components of the U12-type spliceosome direct the correct recognition and removal of introns from the pre-mRNA.
In addition to the intrinsic quality of the splice sites themselves (19), splice site selection can depend on other properties of the pre-mRNA, such as exon sequences and relative splice site proximity (28, 49), RNA secondary structure (38, 58), exon size (27, 51), and intronic sequences (4). For many pre-mRNAs, the splicing reaction produces only a single product from the pre-mRNA transcript. However, other genes undergo alternative splicing, a process in which splice sites in a single primary transcript are differently paired to generate two or more mRNAs encoding multiple protein isoforms with slightly altered or opposing functional properties (18). It was recently estimated that nearly 60% of all human genes undergo at least one alternative splicing event (36).
A remarkable variety of alternative splicing patterns has so far been observed, involving differential 5′ or 3′ splice site selection, alternative exon selection, and intron retention (18). In many instances, alternative splicing is regulated in a developmental or tissue-specific manner. Such complex patterns have been suggested to derive from numerous distinct mechanisms (72). For many examples of alternatively spliced genes, exonic (21, 31, 67) or intronic (3, 9, 11, 20, 26, 29, 52, 68) cis-acting RNA sequences that positively or negatively regulate splice site choice have been identified. Binding of specific proteins to these regulatory sequences, in turn, dictates the specificity and efficiency of splicing resulting in promotion or repression of each splicing event (20).
The Drosophila melanogaster prospero (pros) pre-mRNA provides a rare and unusual example of alternative splicing. The prospero gene locus encodes a homeodomain-containing neuronal transcription factor, Prospero, which is involved in control of axon outgrowth and in cell fate specification in the developing Drosophila central nervous system (14, 40, 66). The second intron of pros pre-mRNA contains two complete sets of splice sites, an arrangement referred to as twintron. One set of splice sites is of the U2-type (GT-AG termini), whose usage leads to a production of the pros-L mRNA isoform. The U2-type splice sites are nested inside an intron defined by a second set of splice sites of the U12 type (with AT-AC termini), whose usage leads to the production of the pros-S mRNA isoform, 87 nucleotides shorter than pros-L (Fig. 1A). Excision of the larger flanking U12-type intron thus removes 29 amino acids with 24 amino acids upstream and 5 amino acids within the N terminus of the homeodomain, a region demonstrated to influence the interaction of the Prospero protein with Drosophila Deformed and mouse Hoxa-5 proteins (25, 46).
FIG. 1.
Splicing of the prospero pre-mRNA is temporally regulated during Drosophila embryogenesis and early larval development. (A) Schematic of the alternatively spliced second intron of D. melanogaster prospero. The U2-type intron is 730 nucleotides long. The U12-type intron contains 59 and 28 nucleotides (dotted boxes) from the pros-L open reading frame, flanking the U2-type intron. The sequences encoding five amino acids of the homeodomain (HD) that are altered by alternative splicing are shown as horizontally lined and checkered boxes. The positions of primers for reverse transcription-PCR analysis are indicated. (B) Input RNA levels correlate with reverse transcription-PCR product levels. In vitro-transcribed RNAs, derived from pros-L and pros-S cDNA, were mixed at various ratios, reverse transcribed, and PCR amplified with the pros-specific primers. The DNA amplicons were then electrophoretically separated, and the quantities of pros-L and pros-S amplicons were measured by PhosphorImager analysis. (C) Stage-specific splicing of pros pre-mRNA during Drosophila embryogenesis and early larval stages. Total RNA was extracted from wild-type embryos collected at different times, reverse transcribed, and PCR amplified. The DNA amplicons were resolved on 12% denaturing polyacrylamide gels. (D) Quantification of the pros-L and pros-S mRNA products at various times throughout Drosophila embryogenesis and early larval stages. For each time point, the percentage of the pros-L (or pros-S) product was calculated by dividing the radioactivity in the pros-L (or pros-S) band by the radioactivity in the pros-L plus the pros-S band. pros-S is denoted by squares and pros-L by triangles.
Fundamental to understanding the alternative splicing of the prospero twintron is an elucidation of the mechanism(s) involved in splice site selection. The existence of two competing splicing pathways within one intron raises the question of whether the two splicing pathways are coregulated or separately controlled through different mechanisms. We began by analyzing the temporal splicing profile of the two alternatively spliced pros mRNA isoforms and found that pros-L predominates during the first half of Drosophila embryogenesis while pros-S is more abundant at later stages. We also set out to identify possible regulatory elements within the twintron. The splicing of pros minigene constructs containing both sets of splice sites was analyzed both in vitro and in vivo. The results show that twintron splicing depends on a 46-nucleotide intronic sequence, called the purine-rich element, located 32 nucleotides downstream of the U2-type 5′ splice site. Its deletion or modification decreases or completely abolishes U2-type splicing. Lastly, we find that the same element is involved in the control of U12-type splicing, since mutation of the element induces a marked decrease in the splicing efficiency of that pathway. Several hypotheses to explain how this element exerts its regulatory functions are discussed.
MATERIALS AND METHODS
Drosophila embryo RNA preparation.
Synchronized Drosophila embryos were collected from a wild-type Canton S population cultured at 25°C on apple juice collection plates coated with a paste of brewer's yeast at 3-h intervals, the first 1-h collection being discarded. The embryos were washed, homogenized by tissue-tearer, and total RNA extracted by Trizol reagent according to the manufacturer's directions.
Minigene constructs.
All DNA constructs were created with standard cloning procedures. For cloning the wild-type pros minigene, two DNA fragments of pros were PCR amplified from wild-type total Drosophila genomic DNA as follows. Fragment I, nucleotides 4069 to 4296 (accession AF190403), and fragment II, nucleotides 4798 to 4984, had XhoI and XbaI sites introduced at the 5′ and 3′ termini, respectively. Fragment I included 118 nucleotides of exon 2 and 109 nucleotides of downstream intronic sequence, while fragment II included 63 nucleotides of exon 3 and 123 nucleotides of upstream intronic sequence. The resulting 413-nucleotide wild-type pros construct was cloned between the XhoI and XbaI sites of the pBluescript (SK+) plasmid (Statagene) downstream of a T7 promoter.
Several pros intron deletion and replacement mutants were constructed, denoted Δx-y for deletions and Rx-y for mutations, where x indicates the nucleotide at which the deletion or mutation starts and y indicates the last nucleotide deleted or mutated. The deletion and replacement mutant minigenes were generated with the QuickChange Mutagenesis system (Stratagene).
Plasmids pGEM-2V6 and pGEM-2TB/R (50), containing the Drosophila ftz and tra genes, respectively, were a generous gift from Emmanuel Laborier and Donald Rio (UC Berkeley).
For generation of in vivo splicing constructs, each in vitro pros XhoI-XbaI fragment was PCR amplified from pBluescript (SK+) and inserted into the SpeI and XhoI sites of the Drosophila expression vector pMT/V5-HisC (Invitrogen) downstream of a metallothionein promoter and upstream of a simian virus 40 polyadenylation signal. All plasmid constructs were verified by restriction enzyme mapping and DNA sequencing (Keck DNA sequencing lab, Yale University).
In vitro transcription.
prospero splicing substrate constructs were linearized by cleaving at the 3′ restriction site with XbaI to create templates for in vitro transcription by T7 RNA polymerase, performed essentially as described by Tarn and Steitz (61) with [α-32P]UTP and 1 μg of linearized plasmid. After RQ1 DNase treatment (Promega), RNA transcripts were gel purified on 5% denaturing polyacrylamide gels and quantified by Cerenkov counting.
Competitor RNA fragments were transcribed under the same conditions except that reactions contained 400 μM each ribonucleoside triphosphate and 0.5 μCi of [α-32P]UTP (Amersham). The trace-labeled competitor RNA fragments were gel purified and quantified by Cerenkov counting.
Nuclear extract preparations.
The Drosophila Kc cell line was grown in suspension at 25°C in standard D22 medium, supplemented with 50 μg/ml streptomycin and penicillin (Sigma). Cells were split to ≈2.5 × 106 cells/ml daily and harvested at 4 to 5 × 106 cells/ml. Nuclear extracts were prepared essentially as described (50) except that the packed nuclei were resuspended in 2.5 ml of buffer C per 1010 cells to give a more concentrated nuclear extract. Human HeLa cells were grown in suspension in RPMI (Gibco-BRL) containing 10% fetal bovine serum, and nuclear extract was prepared by the protocol of Dignam et al. (13).
Splicing reactions.
Each splicing reaction was performed in a total volume of 10 μl, containing 1 fmol of the substrate RNA (≈20,000 cpm), 30% Kc nuclear extract, 2% polyvinyl alcohol, 3 mM ATP, 3 mM MgCl2, 20 mM creatine phosphate, 20 mM HEPES, pH 7.6, and 1-U/μl RNase inhibitor (Roche). Reactions were incubated at 25°C for 3 h. Increasing the time to 5 h did not improve splicing efficiency; rather it increased background, probably because of RNA degradation. The concentrations of 2′-O-methyl oligonucleotides U2b (complementary to 20 out of 23 nucleotides of Drosophila U2 small nuclear RNA [nucleotides 27 to 49] [15]) and U12a (complementary to 16 out of 18 nucleotides of Drosophila U12 small nuclear RNA [nucleotides 11 to 28]) (46) were 4.8 μM and 0.5 μM, respectively. The nuclear extract was preincubated at 25°C with the various 2′-O-methyl oligonucleotides for 15 min prior to substrate addition.
Reverse transcription-PCR.
After the splicing reaction was complete, the reaction was treated with proteinase K, extracted with phenol-chloroform-isoamyl alcohol, and RNAs were isolated by ethanol precipitation. Reverse trancription was primed by the PR324 oligonucleotide (0.5 μg/μl; 5′-CCTTGCGCAGG-3′) complementary to the pros exon 3 sequences nucleotides 280 to 290 (Fig. 2A) (nucleotides 4974 to 4984, accession AF190403). The reaction was performed in 33 μl with the RTG First Strand Synthesis kit (Amersham) at 37°C for 2 h.
FIG. 2.
RNA substrates used for in vitro and in vivo splicing assays. (A) Sequence of the wild-type pros minigene construct, which includes parts of exons 2 and 3 and a shortened intron 2. The sequence was derived from D. melanogaster genomic sequence nucleotides 4069 to 4296 and nucleotides 4798 to 4984 (accession AF190403) and is numbered relative to the U2-type 5′ splice site. The splice site consensus sequences are underlined; U12-type splice sites are in dark red; U2-type splice sites are in blue; the purine-rich element (PRE) is green. Residues mutated in R106-135 are shown in yellow; those in the R136-177 mutation are shown in purple. (B) Summary of deletion and replacement mutations used to delineate pros twintron splicing regulation in vitro and in vivo. The sequence of the purine-rich element of the wild-type pros substrate is shown on top in green. Sequences that were kept intact are shown in green, and those that were mutated are in purple. Heterologous SR protein binding sequences are shown in orange; dashes represent deletions. Minigenes are named by designating the nucleotides deleted (Δ) or replaced (R) within the intron.
cDNA obtained by reverse transcription was amplified with 0.1-μg/μl primers complementary to the sequences in pros exon 2 (PR 514; 5′-AGGATCGCCAGTCCGAGT-3′; nucleotides −118 to −100 [Fig. 2A] or nucleotides 4069 to 4084, accession AF190403) and exon 3 (PR323; 5′-GGTGCATCGGTGTCAATGTA-3′; nucleotides 262 to 281 [Fig. 2A] or nucleotides 4956 to 4975, accession AF190403), in a final volume of 100 μl with 2.5 U of AmpliTaq Gold polymerase (Perkin-Elmer), 50 mM Tris (pH 8.3), 75 mM KCl, 7.5 mM dithiothreitol, 10 mM MgCl2, 0.08-mg/ml bovine serum albumin, and 2.4 mM each deoxynucleoside triphosphate.
To quantitate the amplification products, [γ-32P]ATP 5′ end-labeled primer PR514 (200,000 cpm/μl) was added to the standard 100-μl PCR. Since pilot experiments showed that the PCR amplification rates of the endogenous pros transcripts and of the minigene-derived pros transcripts were linear between the 22nd and 32nd cycles, we used the following cycle parameters for PCR amplification: an initial denaturation at 95°C for 10 min, followed by 30 cycles at 95°C for 1 min, 60°C for 1 min, and 72°C for 2 min. Amplicons (5% of the PCR mixture) were separated on a 12% polyacrylamide gel, dried and exposed. Bands were quantitated with a Molecular Dynamics PhosphorImager. Each quantification was performed on at least 3 experiments. Representative reverse transcription-PCR splicing assays are presented in the figures.
Quantification of each band was corrected by a corresponding background value. In embryos, the percent pros-L inclusion for all time points was calculated as [cpm pros-L/(cpm pros-L + cpm pros-S)] × 100. For in vitro splicing assays, the percent pros-L spliced was calculated as [cpm pros-L/cpm pre-mRNA] × 100. Calculations for pros-S were done similarly to pros-L.
For the in vitro splicing competition assay, 1 μl of trace-labeled RNA competitor (1 to 300 fmol) was added together with 1 fmol of pre-mRNA to the splicing mix. Splicing reactions were carried out as described above.
Cell culture and DNA transfection.
Schneider (S2) cells were grown at 25°C in Shields and Sang M3 medium (Sigma) supplemented with 12.5% fetal bovine serum (Gibco-BRL) and 5% penicillin/streptomycin. Schneider cells were transiently transfected in 35-mm dishes with 1.5 μg of plasmid DNA and 50 μl of Lipofectin reagent (Invitrogen) per dish, according to the manufacturer's protocol. Expression was induced 24 h after transfection by adding 0.7 mM cupric sulfate. Cells were harvested 24 h postinduction and total cellular RNA was isolated with Trizol reagent according to the manufacturer's instructions. Contaminating genomic DNA was removed by treating with RQ DNase I (Promega) in 5 mM MgCl2 and 50 mM Tris, pH 8.0, at 37°C for 1 h.
RNase protection assay.
The template for the RNase protection probe (185 nucleotides) was subcloned from the wild-type pros pMT/V5-HisC construct into the pBluescript SK+ vector (Stratagene) in the reverse orientation with restriction sites XbaI and EcoRI. It contains antisense sequences of the entire downstream exon (63 nucleotides), 45 nucleotides of the 3′-most intron sequence, and 108 nucleotides of the pMT-C vector sequence downstream of the 3′ exon; it was designed such that the same probe could be used for all pros mutant constructs. The probe was transcribed with [32P]UTP and T7 RNA polymerase. Some transfections were also assayed by reverse transcription-PCR (data not shown).
RNase protection assays were performed as follows: 3 μg of total RNA from transfected cells was hybridized to an excess (20,000 cpm) of probe in 30 μl of hybridization mixture: 80% formamide, 0.4 M NaCl, 40 mM PIPES, pH 6.4, 1 mM EDTA at 40°C overnight. RNase digestion was carried out in 300 μl at 30°C for 30 min, with 0.07-U/μl T1 (Calbiochem) and 0.03-μg/μl RNase A. The reaction was stopped by proteinase K digestion, followed by phenol-chloroform-isoamyl alcohol extraction and ethanol precipitation. Protected probe fragments were analyzed on denaturing 8% polyacrylamide gels and were designed to yield three fragments: a fragment of 137 nucleotides for pros-L, a fragment of 108 nucleotides for pros-S, and a fragment of 197 nucleotides for pre-mRNA. Quantitation was done with a Molecular Dynamics PhosphorImager: the percent pros-L spliced was calculated as [cpm pros-L/cpm pre-mRNA] × 100. Each minigene was transfected independently at least four times, and each RNA sample was analyzed at least three times. Results presented are thus the averages of several independent determinations.
RESULTS
Specificity of prospero splicing changes during early development.
The D. melanogaster prospero gene encodes an essential homeodomain protein expressed mostly during embryogenesis from the alternatively spliced pros-S and pros-L mRNAs (Fig. 1A) (14, 66). To ask whether prospero twintron splicing is temporally regulated, we examined the relative levels of pros spliced isoforms during Drosophila embryogenesis and early larval development. Reverse transcription-PCR was performed on total RNA with a single pair of PCR primers since the U2-type intron is nested within the U12-type intron (see Fig. 1A for primer locations). Separation of the radioactively labeled products by denaturing polyacrylamide gel electrophoresis yielded two major bands of 84 and 171 bp (Fig. 1C), corresponding to the U2-type splicing product (pros-S) and the U12-type product (pros-L), respectively. The identity of these bands was confirmed by comparison with the mobility of DNA fragments reverse transcribed and amplified from cloned pros constructs (see below). The quantitative nature of the assay was confirmed by observing that the levels of reverse transcription-PCR amplicons after 18 to 30 cycles correspond to input levels of in vitro-transcribed RNAs synthesized from cloned pros-L and pros-S cDNAs when subjected to serial dilution (Fig. 1B). Thus, the reverse transcription-PCR assay accurately measures the relative amounts of RNA that have undergone alternative splicing and was used throughout this study.
The developmental profile revealed temporal specificity in the appearance of the two alternatively spliced prospero mRNA isoforms. Both isoforms were detected at all time points tested, but showed significant changes during the course of embryonic and larval development (Fig. 1C). Figure 1D plots the data as the percentage of each spliced product, calculated as an average from a minimum of five independent experiments. Roughly 60% of prospero pre-mRNA is spliced via the U2-type splicing pathway to yield pros-L mRNA in the first 9 h of embryogenesis (Fig. 1C and Fig. 1D). Later, pros-L levels continuously decline to less than 20% of total spliced mRNA, while pros-S levels gradually increase to more than 80% for the remainder of the developmental time studied (Fig. 1C and Fig. 1D, 15 to 36 h). About halfway through embryogenesis (12 h; Fig. 1C and Fig. 1D, 9 to 12 h), the levels of the two spliced isoforms are similar. Although a maternal contribution of prospero mRNA cannot be formally excluded, it is unlikely to have impact beyond the first 2.5 h of embryogenesis when zygotic transcription peaks (C. Hashimoto, personal communication). These data suggest that the choice of splicing pathway is developmentally regulated and strictly controlled during embryogenesis.
Design of prospero minigenes and in vitro splicing assay.
We defined the minimum pros RNA sequence required to direct both types of splicing in a homologous Drosophila Kc nuclear extract. A minigene substrate (wild-type pros) that contains shortened exons 2 (from 234 nucleotides to 118 nucleotides) and 3 (from 132 nucleotides to 63 nucleotides) and a shortened intron 2 (from 729 nucleotides to 232 nucleotides) but conserves both sets of splice sites and their spacing with respect to one another was constructed (Fig. 2A). It includes the splicing enhancer element, the purine-rich element, defined in this study (Fig. 2A, green). This wild-type pros pre-mRNA construct was processed so poorly in both Drosophila Kc and human HeLa nuclear extracts that 32P-labeled spliced products could not be detected under a variety of conditions tested (data not shown), even though Drosophila ftz and tra pre-mRNAs both spliced efficiently (data not shown), as previously reported (50). Therefore, the use of reverse transcription-PCR was necessary to assess pros spliced products generated in Drosophila Kc nuclear extract.
Reverse transcription-PCR analysis of the in vitro splicing of the parental wild-type pros substrate (Fig. 3) demonstrated that the twintron was recognized correctly to generate both pros-L and pros-S spliced RNAs, resulting from the U2- and U12-type splicing pathways, respectively. Maximal splicing efficiency was obtained with 3 mM MgCl2 and 30 mM KCl (data not shown), where about 2.5% of pre-mRNA became spliced after 3 h at 24°C, compared with 6% of the human P120 U12-type splicing substrate that becomes spliced after 4 h in a HeLa extract (61). A quantitative analysis of the PCR products revealed that pros-S splicing is 10-fold greater than pros-L. Since the nuclear extract used was prepared from Kc cells, which are derived from late-stage Drosophila embryos (55), and because the in vitro splicing pattern of the wild-type pros substrate was similar to that of the full-length pros pre-mRNA in Drosophila late-stage embryos (Fig. 1), we conclude that the wild-type pros minigene substrate contains sequences necessary to permit a study of twintron splicing regulation.
FIG. 3.
In vitro splicing of prospero substrates in Drosophila Kc nuclear extract. (A) Reverse transcription-PCR analysis of unspliced (unspl; lane 1) and wild-type pros (lane 2) in Kc nuclear extract in the presence of the U12a or U2b 2′-O-methyl blocking oligonucleotides (lanes 3 and 4). Lanes 5 and 6 show in vitro-transcribed RNAs derived from pros-L and pros-S cDNAs, which were reverse transcribed with oligonucleotide PR324 followed by PCR amplification with primers PR514 and PR323. Amplified DNA fragments were resolved on 12% polyacrylamide gels. The pre-mRNA and the spliced products, pros-L and pros-S, are indicated. (B) Splicing of the pros twintron requires sequences within intron 2. The pros splicing substrates listed in Fig. 2B were spliced in vitro, reverse transcription-PCR amplified, and analyzed as in A. (C and D) Graphical representation of the data in B showing the percentage of spliced product for each pros substrate tested, calculated as percent pros-L/pre-mRNA (left-hand scale) and percent pros-S/pre-mRNA (right-hand scale). Measurements were made directly from the gels by PhosphorImager analysis. Each value is an average of at least three independent splicing reactions; error bars indicate the standard deviation associated with each average value. Black bars, pros-L products of U2-type splicing; gray bars, pros-S products of U12-type splicing.
The identities of the in vitro spliced products were inferred from their electrophoretic mobility (Fig. 3A, lanes 5 and 6), from 2′-O-methyl oligonucleotide blocking experiments (Fig. 3A, lanes 3 and 4) and from the results of splice site mutations (Fig. 3B, lanes 1 and 13; Fig. 3C, MUTATAC and MUTGUAG). When the nuclear extract was preincubated with a 2′-O-methyl oligonucleotide, U2b, complementary to the branch point binding region of U2 small nuclear RNA, U2-type splicing was blocked (15), as seen by the absence of pros-L product, while wild-type levels of pros-S spliced product were observed (Fig. 3A, lane 4). Conversely, blocking the competing U12-type splicing pathway with a 2′-O-methyl oligonucleotide, U12a, directed against the comparable region of U12 small nuclear RNA (15) produced wild-type levels of pros-L spliced product but no pros-S product (Fig. 3A, lane 3).
As previously observed with the U12-dependent P120 intron, the oligonucleotide complementary to U2 small nuclear RNA actually stimulates the level of in vitro U12-type splicing (61). Likewise, mutations in the U2-type splice sites produced wild-type levels of pros-S spliced product but no pros-L product (Fig. 3B, lane 13; Fig. 3C, MUTGUAG), whereas mutations in the U12-type splice sites produced the opposite effect (Fig. 3B, lane 1; Fig. 3C, MUTATAC). Thus, the two splicing pathways operate independently; blocking one splicing pathway by mutation does not affect the other. In addition to the pros-L and pros-S cDNA products, we frequently observed variable amounts of nonspecific bands running between the two; these were not present in embryo reverse transcription-PCR preparations (compare Fig. 1C, lanes 1 through 12, with Fig. 3B, lanes 1 though 13) and thus most likely are amplified from RNAs present in the nuclear extract.
Defining sequence elements required for prospero twintron splicing in vitro.
Given the unusual structure of twintron, we anticipated that pros splicing would be dependent upon cis-acting sequences in addition to its two sets of splice sites. Most striking is a purine-rich sequence in the 5′ part of the intron, containing 11 GA repeats flanked by more heterogeneous purine-rich sequences (Fig. 2A). To examine the possible role of this intronic sequence in pros splicing, replacements (Rx-y) and deletions (Δx-y; numbering here and henceforth refers to the distance downstream from the 5′ U2-type splice site of the pros twintron) were introduced into the twintron substrate, as depicted in Fig. 2B, and their splicing patterns were examined in Kc nuclear extract. We calculated the percentage of each spliced product relative to the remaining pre-mRNA. These data, from at least four experiments, are plotted in Fig. 3C and D.
The R106-135 mutation replaces nucleotides that are pyrimidine rich in the middle of the intron with random sequence (Fig. 2A, yellow). The R106-135 substrate reproducibly showed no significant decrease in the splicing of pros twintron or in the ratio of pros-L to pros-S (Fig. 3C, R106-135) compared to wild-type pros (Fig. 3B, lane 2; Fig. 3C, wild-type pros). Similarly, the R136-177 mutation, which replaces 3′ intron sequences (Fig. 2A, purple), retains substantial levels of both splicing pathways (Fig. 3C, R136-177). A substrate containing a 10-nucleotide deletion immediately upstream of the purine-rich region, Δ22-32, also showed no effect on splicing levels (Fig. 3B, lane 8; Fig. 3C, Δ22-32). These results indicate that intronic regions outside of the purine-rich region contribute little or nothing to pros twintron splicing regulation.
Similar analyses of the purine-rich intronic region suggested that multiple copies of GA repeats might have an important enhancer function (Fig. 3B, lanes 4 and 5). To test this hypothesis, we first replaced the purine-rich region with a random sequence (39% purine) in construct R32-77 (Fig. 2B); this largely abolished the major-class splicing pathway and caused a marked decrease in minor-class splicing (Fig. 3B, lane 4; Fig. 3C, R32-77). Deletions of the intronic purine-rich region which remove the 11 GA repeats and an additional 19 or 24 nucleotides of flanking purine-rich sequence (constructs Δ32-77 and Δ37-77, Fig. 2B) showed a marked reduction in splicing levels compared to wild-type pros (Fig. 3B, lane 3; Fig. 3C, Δ32-77; Fig. 3D, Δ37-77) although the reduction was not as great as that of the R32-77 and R38-77 replacement constructs (Fig. 3B, lane 4; Fig. 4B, lane 2; Fig. 3C, R32-77; Fig. 4C, R38-77). These results suggest that the 46-nucleotide purine-rich region contains a sequence or structure that functions as an essential activating element in twintron splicing. We therefore designated the purine-rich region an intronic enhancer and refer to it as the purine-rich element.
FIG.4.
First 20 nucleotides of the purine-rich element are necessary for pros in vitro splicing. (A) Purine substitutions that substitute a heterologous sequence for the purine-rich element while preserving the spacing found in the parent substrate, wild-type pros. The wild-type purine-rich element sequence is underlined. (B) The substrates in A were spliced in Kc nuclear extract, followed by reverse transcription-PCR. The pre-mRNA and the spliced products (pros-L and pros-S) are indicated. (C) Graphic representation of pros-L and pros-S spliced products. Each percentage, calculated as described for Fig. 3C and D, is from a minimum of three independent experiments.
A purine-rich element downstream of the major 5′ splice site enhances splicing of both pathways.
To further define the role of the purine-rich element, we made additional constructs deleting or mutating parts of the element (Fig. 2B). Constructs Δ40-77 and Δ42-77 had no significant effect on pros twintron splicing (Fig. 3D), while construct R32-36+Δ37-77, in which the first five nucleotides of the purine-rich element are replaced by a nonpurine sequence and the remainder of the purine-rich element is deleted, showed a marked decrease in pros-L spliced product (90% decrease) as well as in pros-S product (45% decrease) (Fig. 3D). Replacing the first 6 or 38 nucleotides with a heterologous non-purine-rich sequence, constructs R32-37 and R32-69, did not rescue splicing efficiency either (Fig. 3B, lane 7; Fig. 3C, R32-37 and R32-69). These experiments confirm that essential regulatory sequences are positioned within the purine-rich element (nucleotides 32 to 77 of intron 2) and that they are required for the splicing of both types of intron, although the effects are more pronounced on the U2-type splicing pathway (pros-L) than on the U12-type (pros-S).
Since the sequence downstream of the purine-rich element contains several GTA repeats flanked by purines and is itself quite purine rich (71.4% versus 93.5% for the purine-rich element), this region may substitute for the purine-rich element in deletion and substitution mutants. To test this hypothesis, we constructed two substrates, Δ32-105 and Δ40-105, that contained complete or partial deletions of the purine-rich element together with a deletion in the GTA-repeat region immediately downstream (Fig. 2B). Splicing analysis revealed the largest negative effects on both splicing pathways seen so far (Fig. 3B, lanes 11 and 12), with splicing levels of pros-L and pros-S diminished to roughly 10% for Δ32-105 (Fig. 3B, lane 11; Fig. 3D). Keeping the first 8 nucleotides of the purine-rich element, in Δ40-105, raised the U2-type splicing level to roughly 20% to 30% (Fig. 3B, lane 12; Fig. 3D). Thus, the first eight nucleotides of the purine-rich element appear to be a primary determinant of enhancer activity, with the downstream purine-rich region participating in splicing regulation of pros when the purine-rich element is deleted, but not when it is replaced. Taken together with the results of the Δ32-77 and Δ37-77 substrates, these data suggest that the purine-rich element functions as a bipartite element in which nucleotides 32 to 37 and 77 to 105 influence the overall efficiency of both splicing pathways, whereas nucleotides 37 to 77 primarily influence the ratio of pros-S to pros-L.
Purine-rich element is not replaceable by a heterologous splicing enhancer.
The prospero purine-rich element is comprised of a 46-nucleotide-long stretch of purines interrupted by only three C residues. It therefore resembles the GAR (R = purine) repeats present in many previously characterized exonic splicing enhancers (26, 31, 60, 69). To ask whether the purine-rich element can be replaced by a known purine-rich enhancer element competent to facilitate splicing of other introns by binding to a known SR protein, we chose two such elements and cloned them into a substrate lacking the entire purine-rich element (see Fig. 2B, constructs R-SR and R-ASF/SF2). The first sequence is an enhancer found in the human fibronectin alternate ED1 exon, while the second was identified as an ASF/SF2 binding sequence by SELEX (59). When placed in the context of the pros purine-rich element, neither completely restored pros-L splicing to wild-type levels (Fig. 3B, lanes 9 and 10; Fig. 3D, R-SR and R-ASF/SF2), suggesting that specific elements within the purine-rich element or the spacing of repeats are important for full enhancement of pros twintron splicing.
First 20 nucleotides of the purine-rich element enhance splicing.
To define the regulatory sequences within the pros purine-rich element more precisely, mutant constructs were made in which 5-nucleotide portions of the region were kept intact and the rest of the purine-rich element replaced with mutant sequences, thereby maintaining the spacing (Fig. 4A). When tested for their ability to activate prospero splicing in vitro (Fig. 4B), we observed that constructs R38-77 and R42-77, which contained only the first 6 or 10 nucleotides of the purine-rich element, respectively, decreased splicing to ≈70% for the pros-S spliced product and to ≈15% for pros-L product (Fig. 4B, lanes 2 and 3; Fig. 4C, R38-77 and R42-77). When the first 15 nucleotides of the purine-rich element were kept intact (construct R47-77), splicing proceeded with higher efficiency, but still significantly below wild-type levels (pros-S at ≈80% and pros-L at ≈45%) (Fig. 4B, lane 4; Fig. 4C, R47-77).
We also examined RNA substrates retaining the first 20 to 40 nucleotides of the purine-rich element and found that they were spliced at levels similar to the wild-type pros substrate (Fig. 4B, lanes 5 to 9). Here, pros-S was produced at ≈95% and pros-L at 85 to 90% (Fig. 4C, R52-77, R57-77, R62-77, R67-77, and R72-77). Conversely, the R32-36+Δ37-77 RNA substrate, which has the first 5 nucleotides of the purine-rich element replaced, is spliced even less efficiently than the R38-77 substrate (Fig. 3D, R32-36+Δ37-77; Fig. 4C, R38-77). From these analyses, we conclude that the activating sequence essential for pros twintron splicing is located in the 5′ half of the purine-rich element, 32 to 52 nucleotides downstream of the U2-type 5′ splice site. This region includes the eight nucleotides identified as important above and contains six copies of the GA repeat at its 3′ end.
trans-acting factors mediate enhancement of pros splicing via the purine-rich element.
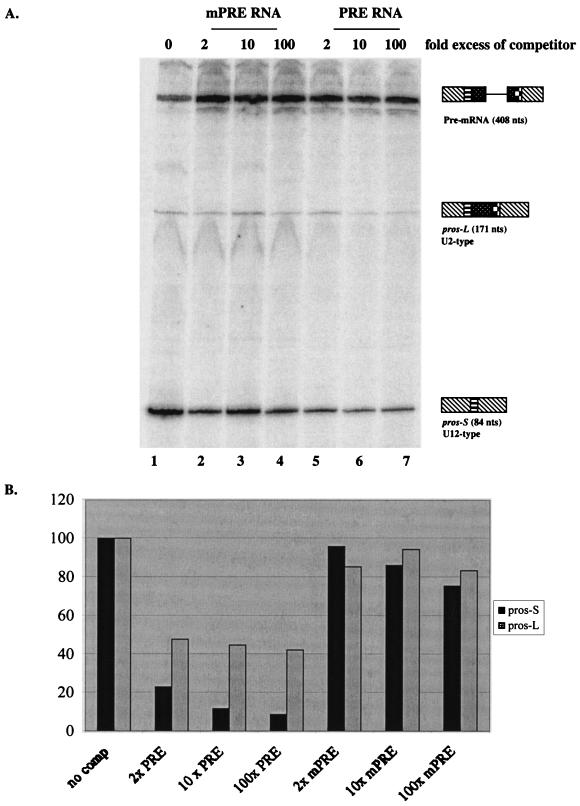
To ask whether the purine-rich element interacts with trans-acting factors to enhance pros twintron splicing, we carried out competition experiments with increasing concentrations of a 46-nucleotide RNA containing the entire purine-rich element (purine-rich element RNA). Partial inhibition (to 25%) of U2-type splicing was observed with as little as twofold molar excess, while almost complete inhibition (to 10%) was observed with 100-fold excess of purine-rich element RNA (Fig. 5A, lanes 5 to 7; Fig. 5B). For U12-type splicing, addition of a twofold molar excess of purine-rich element RNA resulted in a 50% decrease in splicing (Fig. 5A, lane 5; Fig. 5B), while addition of a 100-fold molar excess further decreased splicing to about 40% (Fig. 5A, lane 7; Fig. 5B). In contrast, competition with a non-purine-rich RNA (mPRE RNA), containing sequences that did not substitute for the purine-rich element element in in vitro splicing experiments (Fig. 2B, R32-77; Fig. 3B, lane 4), did not significantly reduce the splicing efficiency of wild-type pros RNA even at the highest concentration tested (Fig. 5A, lane 4). These results are consistent with the idea that the RNA competitor sequesters, by mass action, one or more trans-acting factors required for splicing enhancement of the pros twintron.
FIG. 5.
Inhibition of in vitro splicing by competition with purine-rich element RNA. (A) Kc nuclear extract was incubated with pre-mRNA and increasing amounts of an RNA containing the purine-rich element sequence (lanes 5 to 7) or a nonspecific sequence (Fig. 2B, R32-77; mPRE, lanes 2 to 4) that does not substitute for the purine-rich element in in vitro splicing assays. After splicing, the RNAs were analyzed by reverse transcription-PCR as above. The molar excess of competitor is indicated at the top of each lane. The wild-type pros pre-mRNA and products, pros-L and pros-S, are indicated. (B) Graphic representation of the data in A showing the percentage of pros-L and pros-S spliced products in the presence of RNA competitors. Each value was calculated as percent pros-L (or pros-S)/pre-mRNA; the value of wild-type pros with no competitor was set at 100%.
Purine-rich element enhances prospero splicing in vivo.
The results reported in the preceding sections identified an intronic purine-rich element that acts as an enhancer of pros splicing in vitro. To determine whether the purine-rich element affects pros splicing in vivo as it does in vitro, we transiently transfected Drosophila S2 cells with minigenes carrying the same intronic sequence variants as analyzed in vitro. The 413-bp prospero minigene substrate used in in vitro splicing was PCR amplified from the pBS vector and subcloned into a Drosophila expression vector, pMT/V5-HisC, downstream of the metallothionein promoter and upstream of the simian virus 40 polyadenylation signal (Fig. 6A). The Schneider (S2) cell line was used in these assays because Drosophila Kc cells could not be efficiently transfected under a variety of conditions tested (data not shown). After transient transfection and induction with copper sulfate, total RNA was isolated and the splicing pattern of the minigene-derived mRNA molecules monitored by RNase protection (Fig. 6B). Because the S2 cell line is derived from blood precursor cells and pros is primarily expressed in developing neurons and in the midgut, we did not expect or detect any endogenous pros expression (data not shown).
FIG.6.
Splicing of prospero substrates in Drosophila S2 cells. (A) Organization of the pros minigene used for transfection. The line underneath the construct shows the probe used in the RNase protection assay. Vector sequences are not drawn to scale. (B) Representative gels of RNase-protected fragments analyzing the products of transient transfection. The pros-S product is always detected as a doublet, often seen in RNase protection analyses. (C and D) The bar graphs show the levels of pros-L splicing for each substrate tested. Error bars indicate the standard deviation of the measurements from four to six in vivo splicing experiments. The percent of pros-L was defined as [(total counts in the pros-L product)/(total counts in the pre-mRNA)].
Similar to in vitro splicing patterns, in vivo analyses identified reverse transcription-PCR amplicons of two spliced products (pros-L and pros-S), as well as of the pre-mRNA, based on electrophoretic mobility compared to cDNA plasmid standards cloned into the pMT/V5-HisC vector and transfected into S2 cells (data not shown). In addition, U2- and U12-type splice site mutations cloned into the same vector revealed that interrupting one splicing pathway did not impede the other splicing pathway in vivo (Fig. 6C, mATAC and mGUAG). In strong contrast to the in vitro results, in vivo splicing of wild-type pros in S2 cells produced a small amount of pros-S (2% of pre-mRNA) and a large amount of pros-L product (300% of pre-mRNA). Nevertheless, splicing of the pros twintron was greatly reduced for minigenes lacking the purine-rich element (Δ32-77, Fig. 6B, lane 3; Fig. 6C) or where the purine-rich element was mutated (R32-77, Fig. 6B, lane 2; Fig. 6C). Therefore, the purine-rich element plays a similar role in pros splicing activation in vivo as it does in vitro.
The results of RNase protection analyses of RNAs obtained from transfected S2 cells are summarized in Fig. 6C, and representative splicing patterns are shown in Fig. 6B. Because the splicing efficiency of the U12-type splicing pathway was very low and thus difficult to quantify accurately, only the results for U2-type spliced products are discussed. As in vitro, mutations outside of the purine-rich element (Δ22-32, R106-135, and R136-177) did not significantly affect splicing efficiency in vivo (Fig. 6C). Likewise, deletion mutations within the purine-rich element (R32-36+Δ37-77 and Δ32-105) severely reduced U2-type splicing (Fig. 6C), while replacement mutants within the purine-rich element, constructs R32-37 and R32-69 reduced the pros-L splicing levels by 80% (Fig. 6D). In contrast with constructs Δ40-77 and Δ42-77, where the first eight or ten nucleotides of the purine-rich element were intact, the levels of pros-L were similar to that of wild-type pros (Fig. 6C), as in vitro. Likewise, double deletions encompassing the purine-rich element and downstream GTA repeat region showed a severe splicing phenotype, with the pros-L levels decreased by 95% when compared to wild-type pros (Δ32-105 and Δ40-105, Fig. 6B, lanes 5 and 6; Fig. 6C), again similar to the splicing phenotype observed in vitro. Lastly, the chimeric constructs R-SR and R-ASF, where the purine-rich element was substituted with heterologous purine-rich enhancer sequences, did not splice better in vivo (Fig. 6B, lane 4; Fig. 6D; data not shown) than in vitro (Fig. 3B), underscoring the sequence-specific functioning of the purine-rich element.
We also tested minigene replacement mutants in which increasing lengths of the purine-rich element were kept intact while replacing the rest with a nonpurine sequence to maintain the total length of the intron (Fig. 6B, lanes 9 to 16). Analysis of these additional mutants showed that the first 20 nucleotides of the purine-rich element are required for minimal pros-L splicing in vivo and the first 30 nucleotides are necessary for spliced levels equivalent to those of wild-type pros (Fig. 6B, lanes 12 to 14; Fig. 6D, R38-77 through R67-77), in good agreement with the in vitro data.
Overall, the in vivo analyses support the in vitro results. The S2 cell line, like the Kc nuclear extract, specifically requires the purine-rich element for splicing of the pros twintron, suggesting that a trans-acting factor(s) recognizing the purine-rich element is present in a variety of cell types.
DISCUSSION
Regulation of prospero twintron splicing is important for determining the balance between pros-S and pros-L, two isoforms encoded by the Drosophila melanogaster pros gene that arise from usage of the U12- and U2-type splicing pathways, respectively. In this study, we show that these two isoforms are expressed in a stage-specific manner; the U2-type spliced product predominates in the early stages of Drosophila embryogenesis, while the U12-type spliced product is abundant during the second half of embryo development. We have also identified a 46-nucleotide cis-acting element, the purine-rich element, located 32 nucleotides downstream of the 5′ U2-type splice site that enhances pros twintron splicing both in vitro and in vivo. Its deletion or mutation has two effects, the most noticeable being the complete inhibition of the U2-type splicing pathway and the second a more subtle downregulation of the U12-type splicing pathway. Based on competition data, we conclude that the purine-rich element acts by binding a trans-acting factor(s) which remains to be identified.
Implications of differential temporal expression of prospero isoforms.
Stage- and tissue-specific pre-mRNA splicing events are important aspects of gene regulation that allow an organism to produce related proteins from a single gene (18). Alternative splicing of the prospero twintron produces two isoforms, pros-L and pros-S mRNAs, via the usage of two distinct splicing pathways, the U2- and U12-type, respectively. Removal of the larger U12-type intron excises 29 amino acids encoded by pros-L (removed by the U2-type pathway), five of which are part of the N terminus of the pros homeodomain.
We began by investigating the stage-specific profile of prospero pre-mRNA splicing in the Drosophila embryo and early larva and found that one form (pros-L) predominates in early embryogenesis while the other (pros-S) predominates in middle to late embryogenesis and early larval development. Similar stage-specific splicing has been reported for another alternatively spliced Drosophila homeotic gene, ultrabithorax (ubx), where the RNA levels have been shown to fluctuate during Drosophila embryonic development with the long ubx isoforms (Ia and Ib) predominating during the first 7.5 h and a short isoform (IV) predominating from 18 to 22.5 h of Drosophila embryogenesis (35, 44).
What functional differences might derive from the structural differences between the proteins encoded by the two pros mRNAs? Pros protein has been reported to regulate multiple target genes in different lineages and different stages of neuronal development by repressing or activating their transcription (10, 66). It is therefore possible that the two pros isoforms interact with different coactivators in different tissues, and at different developmental stages. Recent structural studies on the Pros-L C-terminal domain (encompassing the homeodomain and Prospero domain) suggested three regions of potential Prospero-DNA contacts, among them, the N-terminal arm of the homeodomain appears to contact the minor groove of the DNA (53). The Pros homeodomain is essential for the sequence-specific DNA binding function of Prospero and its N terminus has been shown to associate with two homeobox proteins, Engrailed (12) and Deformed (25). Because the Pros-S homeodomain contains a different N terminus, it is tempting to speculate that the two Pros proteins exhibit different DNA-binding capacities that lead to target gene selectivity during neurogenesis in a tissue or stage-specific manner. As a result, production of various amounts of the two protein isoforms would be expected at different stages of development or in different tissues.
Two additional examples of twintrons in the second intron of the prospero gene are found in D. pseudoobscura and D. virilis. Sequence alignment further reveals a homologous purine-rich element in a similar location in D. pseudoobscura; it is located 40 nucleotides downstream of the 5′ U2-type splice site and contains the same number of GA repeats flanked on both sides by purines. The 5′ half of the purine-rich region is 100% conserved between the two species (apart from three missing residues, which could be due to sequencing error) while the 3′ end is less conserved. D. virilis (which is more diverged from D. melanogaster than D. pseudoobscura [48]) also contains a purine-rich region located 28 nucleotides downstream of its U2-type 5′ splice site, but this region contains only five GA repeats. These observations support the functional importance of the purine-rich element as a regulatory cis-element in twintron splicing.
Additional Prospero orthologs, called Prox1, have been identified in Caenorhabditis elegans (6), chicken (64), mouse (64), and human (73). Their protein expression patterns indicate that they are critical for the development and function of the developing central nervous system, eye, and midgut (64). While Prox1 is very well conserved in vertebrates (64), including 102 amino acids at the 3′ terminus of the Prox1 homeodomain which are 62% identical to corresponding residues of Drosophila Prospero (45), the first seven amino acids of the Prox 1 homeodomain do not share homology with Drosophila Prospero. Strikingly, all of the vertebrate prox 1 genes contain a single U12-type intron located near the N terminus of the homeobox in exactly the same position as the U12-type intron in Drosophila prospero (64). However, no alternatively spliced variants of Prox 1 have been identified (45), and we have found no hints of U2-type splicing signals in the second intron of mouse or human Prox 1. Furthermore, sequence analysis of human prospero has not revealed any pronounced intronic purine-rich sequences. This suggests that the twintron and its intronic purine-rich element are unique to Drosophila species, where they may play a special role in development.
Splicing of prospero is enhanced by an intronic purine-rich element.
To our knowledge, prospero provides the only example of a twintron whose splicing can proceed via either the U12- or U2-type pathway. Our documentation of temporal specificity in embryos lends further support to the idea that regulatory signals within the pros pre-mRNA act as sensors of developmental and/or tissue-specific cues. We generated a minimal splicing substrate and established an in vitro splicing system in Kc nuclear extract as a means to map and characterize cis-regulatory elements in a systematic way. Deletion and replacement mutant analysis of the intron led to identification of a 46-nucleotide purine-rich regulatory element, denoted the purine-rich element, that is located 32 nucleotides downstream of the U2-type 5′ splice site and is required for twintron splicing both in vitro and in vivo. Complete replacement of the purine-rich element impairs U2-type splicing both in vitro and in vivo. The presence of multiple GA repeats is consistent with a partite, redundant element structure, a common feature of many intronic regulatory sequences (26). Indeed, in vitro studies indicated that only the first 20 nucleotides of the purine-rich element are required for pros twintron splicing. Furthermore, replacements of the purine-rich element by purine-rich SR protein binding sites did not completely restore splicing, while the deletion mutants in which only the first 8 or 10 nucleotides of the purine-rich element were kept did.
Although in vitro splicing in Kc cell extract yielded high levels of pros-S and low levels of pros-L products, the same transcripts spliced in vivo in the Schneider (S2) cell line showed the opposite effect. The linearity of the transfection data of Patel et al. (47), obtained in S2 cells under the same conditions, excludes the possibility that the observed differences result from artificial saturation of the U12-type splicing machinery by high levels of pre-mRNA expressed from the transfected constructs. Although the basis of reversed levels of pros-S and pros-L in vivo compared to the in vitro system is unknown, reduction in pros-S levels and increase in pros-L levels similar to those in S2 cells are also observed when the pros substrate is spliced in vitro in HeLa nuclear extracts (data not shown).
A likely explanation for this discrepancy stems from the origin of the two cell lines. Although both were derived from late-stage Drosophila embryos, Kc cells are of hematopoietic origin (1), whereas the Schneider cell line is of macrophage origin (54). Antagonistic splicing factors have been shown to influence alternative splicing in vivo in a concentration-dependent manner (7, 70). For instance, in vivo variations in hnRNP A1 protein levels have been shown to influence 5′ splice site choice (70). Factors can also vary between cell types (16, 71), such as levels of ASF/SF2 differing nearly 20-fold between heart and testis (24) and levels of X16/SRp20 protein being high in pre-B cells and thymus but not abundant in other cells (2). Therefore, the enhancement of pros-L splicing observed in S2 cells could be due to various levels of hnRNP, SR proteins, or other splicing factors.
The location of the purine-rich element and its enhancer function resemble the situation with several previously characterized intronic enhancer elements (3, 4, 8, 9, 11, 17, 29, 30, 33, 52, 57). Intronic enhancer sequences are often purine-rich, but are diverse in sequence and distinct from exonic enhancers. For instance, purine-rich regions act as important regulatory signals for neural splicing of mammalian c-src, calcitonin/CGRP, and agrin (39, 41, 68). In the c-src gene, the intronic enhancer located just downstream of the N1 exon is required for the N1 exon to be positively selected (4); it assembles a complex of proteins, including hnRNP F, hnRNP H, and KSRP (41).
While intronic enhancers generally have a complex architecture, the prospero purine-rich element sequence appears distinct in that it is composed almost exclusively of purines. Furthermore, its deletion or modification had two effects: the almost complete abolition of major-class splicing and the down regulation of minor-class splicing. Specifically, the mutant in which the entire 46-nucleotide of the purine-rich element was replaced with a non-purine-rich sequence yielded no observable major class product, while deletions within purine-rich element decreased major class splicing to a lesser degree. The purine-rich region immediately downstream of the purine-rich element was able to partially replace portions of the purine-rich element in the deletion mutants (perhaps by being brought closer to the 5′ splice sites) but not in the replacement mutants. Thus, activating sequences in addition to the purine-rich element may exist in the vicinity of the pros twintron.
There are at least two possible mechanisms by which the prospero purine-rich element may exert its stimulatory effect: one is that it participates in formation of a secondary structure that improves the accessibility of splicing factors to the splice sites, and another is that the element serves as a target of a trans-acting factor(s). Competition experiments (Fig. 6B) showed that the stimulatory effect of the purine-rich element can be titrated by competitor purine-rich element RNA, strongly supporting the latter mechanism. We thus propose that the purine-rich element serves as a target sequence for the formation of a protein enhancer complex. The data further suggest that the purine-rich element functions as a bipartite element in which nucleotides 32 to 37 and 77 to 105 influence the overall splicing efficiency of both splicing pathways, while nucleotides 37 to 77 primarily influence the ratio of pros-S to pros-L. Future research will focus on the factors that bind the purine-rich element and flanking sequences, their binding locations and how they function to influence spliceosome assembly and promote twintron splicing in general.
Acknowledgments
We thank C. Hashimoto, Y. Huang, R. Lytle, L. Otake, and K. Tycowski for comments on the manuscript; members of the Steitz laboratory for helpful discussions; and L. Raymond for assistance with the manuscript. We acknowledge C. Hashimoto for help with Drosophila embryo collections and D. Rio and E. Labourier for invaluable help in making the Kc nuclear extracts.
This work was supported by National Institutes of Health grant GM26154 to J.A.S., who is an investigator of the Howard Hughes Medical Institute.
REFERENCES
- 1.Andres, A. J., and P. Cherbas. 1992. Tissue-specific ecdysone responses: regulation of the Drosophila genes Eip28/29 and Eip40 during larval development. Development 116:865-876. [DOI] [PubMed] [Google Scholar]
- 2.Ayane, M., U. Preuss, G. Kohler, and P. J. Nielsen. 1991. A differentially expressed murine RNA encoding a protein with similarities to two types of nucleic acid binding motifs. Nucleic Acids Res. 19:1273-1278. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Balvay, L., D. Libri, M. Gallego, and M. Y. Fiszman. 1992. Intronic sequence with both negative and positive effects on the regulation of alternative transcripts of the chicken beta tropomyosin transcripts. Nucleic Acids Res. 20:3987-3992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Black, D. L. 1992. Activation of c-src neuron-specific splicing by an unusual RNA element in vivo and in vitro. Cell 69:795-807. [DOI] [PubMed] [Google Scholar]
- 5.Burge, C. B., T. Tuschl, and P. A. Sharp. 1999. The nature of modern RNA suggests a prebiotic RNA, p. 525-560. In R. F. Gesteland, T. Cech, and J. F. Atkins (ed.), RNA world. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y.
- 6.Burglin, T. R. 1994. A Caenorhabditis elegans prospero homologue defines a novel domain. Trends Biochem. Sci. 19:70-71. [DOI] [PubMed] [Google Scholar]
- 7.Caceres, J. F., S. Stamm, D. M. Helfman, and A. R. Krainer. 1994. Regulation of alternative splicing in vivo by overexpression of antagonistic splicing factors. Science 265:1706-1709. [DOI] [PubMed] [Google Scholar]
- 8.Carlo, T., R. Sierra, and S. M. Berget. 1996. An intron splicing enhancer containing a G-rich repeat facilitates inclusion of a vertebrate micro-exon. RNA 2:342-353. [PMC free article] [PubMed] [Google Scholar]
- 9.Chan, R. C., and D. L. Black. 1995. Conserved intron elements repress splicing of a neuron-specific c-src exon in vitro. Mol. Cell. Biol. 15:6377-6385. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Chu-Lagraff, Q., D. M. Wright, L. K. McNeil, and C. Q. Doe. 1991. The prospero gene encodes a divergent homeodomain protein that controls neuronal identity in Drosophila. Development Suppl. 2:79-85. [PubMed] [Google Scholar]
- 11.Del Gatto, F., A. Plet, M. C. Gesnel, C. Fort, and R. Breathnach. 1997. Multiple interdependent sequence elements control splicing of a fibroblast growth factor receptor 2 alternative exon. Mol. Cell. Biol. 17:5106-5116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Desplan, C., J. Theis, and P. H. O'Farrell. 1988. The sequence specificity of homeodomain-DNA interaction. Cell 54:1081-1090. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Dignam, J. D., R. M. Lebovitz, and R. G. Roeder. 1983. Accurate transcription initiation by RNA polymerase II in a soluble extract from isolated mammalian nuclei. Nucleic Acids Res. 11:1475-1489. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Doe, C. Q., Q. Chu-Lagraff, D. M. Wright, and M. P. Scott. 1991. The prospero gene specifies cell fates in the Drosophila central nervous system. Cell 65:451-464. [DOI] [PubMed] [Google Scholar]
- 15.Frilander, M. J., and J. A. Steitz. 1999. Initial recognition of U12-dependent introns requires both U11/5′ splice-site and U12/branchpoint interactions. Genes Dev. 13:851-863. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Fu, X. 1995. The superfamily of arginine/serine-rich splicing factors. RNA 1:663-680. [PMC free article] [PubMed] [Google Scholar]
- 17.Gontarek, R. R., M. T. McNally, and K. Beemon. 1993. Mutation of an RSV intronic element abolishes both U11/U12 small nuclear RNP binding and negative regulation of splicing. Genes Dev. 7:1926-1936. [DOI] [PubMed] [Google Scholar]
- 18.Graveley, B. R. 2001. Alternative splicing: increasing diversity in the proteomic world. Trends Genet. 17:100-107. [DOI] [PubMed] [Google Scholar]
- 19.Green, M. R. 1991. Biochemical mechanisms of constitutive and regulated pre-mRNA splicing. Annu. Rev. Cell Biol. 7:559-599. [DOI] [PubMed] [Google Scholar]
- 20.Guil, S., R. Gattoni, M. Carrascal, et al. 2003. Roles of hnRNP A1, SR proteins, and p68 helicase in cHras alternative splicing regulation. Mol. Cell. Biol. 23:2927-2941. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Guo, M., P. C. H. Lo, and S. M. Mount. 1993. Species-specific signals for the splicing of a short Drosophila intron in vitro. Mol. Cell. Biol. 13:1104-1118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Hall, S. L., and R. A. Padgett. 1994. Conserved sequences in a class of rare eukaryotic nuclear introns with non-consensus splice sites. J. Mol. Biol. 239:357-365. [DOI] [PubMed] [Google Scholar]
- 23.Hall, S. L., and R. A. Padgett. 1996. Requirement of U12 small nuclear RNA for in vivo splicing of a minor class of eukaryotic nuclear pre-mRNA introns. Science 271:1716-1718. [DOI] [PubMed] [Google Scholar]
- 24.Hanamura, A., J. F. Caceres, A. Mayeda, B. R. Franza, and A. R. Krainer. 1998. Regulated tissue-specific expression of antagonistic pre-mRNA splicing factors. RNA 4:430-444. [PMC free article] [PubMed] [Google Scholar]
- 25.Hassan, B., L. Li., K. A. Bremer, W. Chang, J. Pinsonneault, and H. Vaessin. 1997. Prospero is a panneural transcription factor that modulates homeodomain protein activity. Proc. Natl. Acad. Sci. USA 94:10991-10996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Hastings, M. L., C. L. Wilson, and S. H. Munroe. 2001. A purine-rich intronic element enhances alternative splicing of thyroid hormone receptor mRNA. RNA 7:859-874. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Hawkins, J. D. 1988. A survey on intron and exon lengths. Nucleic Acids Res. 16:9893-9908. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Hedley, M. L., and T. Maniatis. 1991. Sex-specific splicing and polyadenylation of dsx pre-mRNA requires a sequence that binds specifically to tra-2 protein in vitro. Cell 65:579-586. [DOI] [PubMed] [Google Scholar]
- 29.Huh, G. S., and R. O. Hynes. 1993. Elements regulating an alternatively spliced exon of the rat fibronectin gene. Mol. Cell. Biol. 13:5301-5314. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Hui, J. G. Reither, and A. Bindereif. 2003. Novel functional role of CA repeats and hnRNP L in RNA stability. RNA 9:931-937. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Humphrey, M. B., J. Bryan, T. A. Cooper, and S. M. Berget. 1995. A 32-nucleotide exon-splicing enhancer regulates usage of competing 5′ splice sites in a differential internal exon. Mol. Cell. Biol. 15:3979-3988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Incorvaia, R., and R. A. Padgett. 1998. Base pairing with U6atac small nuclear RNA is required for 5′ splice site activation of U12-dependent introns in vivo. RNA 4:709-718. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Kawamoto, S. 1996. Neuron-specific alternative splicing of nonmuscle myosin II heavy chain-B pre-mRNA requires a cis-acting intron sequence. J. Biol. Chem. 271:17613-17616. [PubMed] [Google Scholar]
- 34.Kolossova, I., and R. A. Padgett. 1997. U11 small nuclear RNA interacts in vivo with the 5′ splice site of U12-dependent (AU-AC) pre-mRNA introns. RNA 3:227-233. [PMC free article] [PubMed] [Google Scholar]
- 35.Kornfeld, K., R. B. Saint, P. A. Beachy, et al. 1989. Structure and expression of a family of ultrabithorax mRNAs generated by alternative splicing and polyadenylation in Drosophila. Genes Dev. 3:243-258. [DOI] [PubMed] [Google Scholar]
- 36.Lander, E. S., et al. 2001. Initial sequencing and analysis of the human genome. Nature 409:860-921. [DOI] [PubMed] [Google Scholar]
- 37.Lavigueur, A., H. La Branche, A. R. Kornblihtt, and B. Chabot. 1993. A splicing enhancer in the human fibronectin alternate ED1 exon interacts with SR proteins and stimulates U2 small nuclear RNP binding. Genes Dev. 7:2405-2417. [DOI] [PubMed] [Google Scholar]
- 38.Libri, D., A. Piseri, and M. Y. Fiszamn. 1991. Tissue-specific splicing in vivo of the beta-tropomyosin gene: dependence on an RNA secondary structure. Science 252:1842-1845. [DOI] [PubMed] [Google Scholar]
- 39.Lou, H., R. F. Gagel, and S. M. Berget. 1996. An intron enhancer recognized by splicing factors activates polyadenylation. Genes Dev. 10:208-219. [DOI] [PubMed] [Google Scholar]
- 40.Matsuzaki, F., K., H. Koizumiama, T. Yoshioka, and Y. Nabeshima. 1992. Cloning of the Drosophila prospero gene and its expression in ganglion mother cells. Biochem. Biophys. Res. Commun. 182:1326-1332. [DOI] [PubMed] [Google Scholar]
- 41.Min, H., R. C. Chan, and D. L. Black. 1995. The generally expressed hnRNP F is involved in a neural-specific pre-mRNA splicing event. Genes Dev. 9:2659-2671. [DOI] [PubMed] [Google Scholar]
- 42.Min, H., C. V. Turck, J. M. Nikolic, and D. L. Black. 1997. A new regulatory protein, KSRP, mediates exon inclusion through an intronic splicing enhancer. Genes Dev. 11:1023-1036. [DOI] [PubMed] [Google Scholar]
- 43.Moore, M. J., C. C. Query, and P. A. Sharp. 1993. Splicing of precursors to mRNAs by the spliceosome, p. 303-357. In R. F. Gesteland, T. Cech, and J. F. Atkins (ed.), RNA world. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y.
- 44.O'Connor, M. B., R. Binari, L. A. Perkins, and W. Bender. 1988. Alternative RNA products from the Ultrabithorax domain of the bithorax complex. EMBO J. 7:435-445. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Oliver, G., B. Sosa-Pineda, S. Geisendorf, P. Spana, E. C. Q. Doe, and P. Gruss. 1993. Prox 1, a prospero-related homeobox gene expressed during mouse development. Mech. Dev. 44:3-16. [DOI] [PubMed] [Google Scholar]
- 46.Otake, L. R., P. Scamborova, C. Hashimoto, and J. A. Steitz. 2002. The divergent U12-type spliceosome is required for pre-mRNA splicing and is essential for development in Drosophila. Mol. Cell 9:439-446. [DOI] [PubMed] [Google Scholar]
- 47.Patel, A. A., M. McCarthy, and J. A Steitz. 2002. The splicing of U12-type introns can be a rate-limiting step in gene expression. EMBO J. 21:3804-3815. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Powell, J. R., and E. N. Moriyama, 1997. Evolution of codon usage bias in Drosophila. Proc. Natl. Acad. Sci. USA 94:7784-7790. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Reed, R., and T. Maniatis. 1986. A role for exon sequences and splice-site proximity in splice site selection. Cell 46:681-690. [DOI] [PubMed] [Google Scholar]
- 50.Rio, D. C. 1988. Accurate and efficient pre-mRNA splicing in Drosophila cell-free extracts. Proc. Natl. Acad. Sci. USA 85:2904-2908. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Robberson, B. L., G. J. Cote, and S. M. Berget. 1990. Exon definition may facilitate splice site selection in RNAs with multiple exons. Mol. Cell. Biol. 10:84-94. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Ryan, K. J., and T. A. Cooper. 1996. Muscle-specific splicing enhancers regulate inclusion of the cardiac troponin T alternative exon in embryonic skeletal muscle. Mol. Cell. Biol. 16:4014-4023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Ryter, J. M., C. Q. Doe, and B. W. Matthews. 2002. Structure of the DNA binding region of prospero reveals a novel homeo-prospero domain. Structure 10:1541-1549. [DOI] [PubMed] [Google Scholar]
- 54.Samokovlis, C., D. A. Kimbrel, P. Kylsten, A. Engstrom, and D. Hultmark. 1990. The immune response in Drosophila: pattern of cecropin expression and biological activity. EMBO J. 9:2969-2976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Schneider, I. and A. B. Blumenthal. 1978. Drosophila cell tissue and culture, p. 265-315. In M. Ashburner and T. R. F. Wright (ed.), The genetics and biology of Drosophila, vol. 2a. Academic Press, New York, N.Y.
- 56.Sharp, P. A., and C. B. Burge. 1997. Classification of introns: U2 type or U12 type. Cell 91:875-879. [DOI] [PubMed] [Google Scholar]
- 57.Sirand-Pugnet, P., P. Durosay, E. Brody, and J. Marie. 1995. An intronic (A/U)GGG repeat enhances the splicing of an alternative intron of the chicken beta-tropomyosin pre-mRNA. Nucleic Acids Res. 23:3501-3507. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Solnick, D. 1985. Alternative splicing caused by RNA secondary structure. Cell 43:667-676. [DOI] [PubMed] [Google Scholar]
- 59.Tacke, R., and J. L. Manley. 1995. The human splicing factors ASF/SF2 and SC35 possess distinct, functionally significant RNA binding specificities. EMBO J. 14:3540-3551. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Tanaka, K., A. Watakabe, and Y. Shimura. 1994. Polypurine sequences within a downstream exon function as a splicing enhancer. Mol. Cell. Biol. 14:1347-1354. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Tarn, W. Y., and J. A. Steitz. 1996. Highly diverged U4 and U6 small nuclear RNAs required for splicing rare AT-AC introns. Science 273:1824-1832. [DOI] [PubMed] [Google Scholar]
- 62.Tarn, W. Y., and J. A. Steitz. 1996. A novel spliceosome containing U11, U12, and U5 small nuclear RNPs excises a minor class (AT-AC) intron in vitro. Cell 84:801-811. [DOI] [PubMed] [Google Scholar]
- 63.Tarn, W. Y., and J. A. Steitz. 1997. Pre-mRNA splicing: the discovery of a new spliceosome doubles the challenge. Trends Biochem. Sci. 22:132-137. [DOI] [PubMed] [Google Scholar]
- 64.Tomarev, S. I., O. Sundin, S. Banerjee-Basu, et al. 1996. Chicken homeobox gene Prox 1 related to Drosophila prospero is expressed in the developing lens and retina. Dev. Dyn. 206:354-367. [DOI] [PubMed] [Google Scholar]
- 65.Tomarev, S. I., R. D. Zinovieva, B. Chang, and N. L. Hawes. 1998. Characterization of the mouse Prox1 gene. Biochem. Biophys. Res. Commun. 248:684-689. [DOI] [PubMed] [Google Scholar]
- 66.Vaessin, H., E. Grell, E. Wolff, E. Bier, L. Y. Jan, and Y. N. Jan. 1991. Prospero is expressed in neuronal precursors and encodes a nuclear protein that is involved in the control of axonal outgrowth in Drosophila. Cell 67:941-953. [DOI] [PubMed] [Google Scholar]
- 67.Watakabe, A., K. Tanaka, and Y. Shimura. 1993. The role of exon sequences in splice site selection. Genes Dev. 7:407-418. [DOI] [PubMed] [Google Scholar]
- 68.Wei, N., C. Q. Lin, E. F. Modafferi, W. A. Gomes, and D. L. Black. 1997. A unique intronic splicing enhancer controls the inclusion of the agrin Y exon. RNA 3:1275-1288. [PMC free article] [PubMed] [Google Scholar]
- 69.Xu, R., J. Cheng, and T. A. Cooper. 1993. The cardiac troponin T alternative exon contains a novel purine-rich positive splicing element. Mol. Cell. Biol. 13:3660-3674. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Yang, X., M. R. Bani, S. J. Lu, S. Rowan, Y. Ben-David, and B. Chabot. 1994. The A1 and A1B proteins of heterogeneous nuclear ribonucleoparticles modulate 5′ splice site selection in vivo. Proc. Natl. Acad. Sci. USA 91:6924-6928. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Zahler, A. M., K. M. Neugebauer, J. A. Stolk, and M. B. Roth. 1993. Distinct functions of SR proteins in alternative pre-mRNA splicing. Science 260:219-222. [DOI] [PubMed] [Google Scholar]
- 72.Zhang, L., M. Ashiya, T. G. Sherman, and P. J. Grabowski. 1996. Essential nucleotides direct neuron-specific splicing of gamma 2 pre-mRNA. RNA 2:682-698. [PMC free article] [PubMed] [Google Scholar]
- 73.Zinovieva, R. D., M. K. Duncan, T. R. Johnson, et al. 1996. Structure and chromosomal localization of the human homeobox gene Prox 1. Genomics 35:517-522. [DOI] [PubMed] [Google Scholar]