Summary
Leaf primordia form around the shoot apical meristem, which consists of indeterminate stem cells. Upon initiation of leaf development, adaxial-abaxial patterning is crucial for appropriate lateral expansion, via cellular proliferation, and the formation of flat symmetric leaves. Many genes that specify such patterning have been identified, but regulation by upstream factors of the expression of relevant effector genes remains poorly understood. In Arabidopsis thaliana, ASYMMETRIC LEAVES2 (AS2) and AS1 play important roles in repressing transcription of class 1 KNOTTED1-like homeobox (KNOX) genes and leaf abaxial-determinant effector genes. We report here a mutation, designated enhancer of asymmetric leaves2 and asymmetric leaves1 (eal), that is associated with efficient generation of abaxialized filamentous leaves on the as2 or as1 background. Levels of transcripts of many abaxial-determinant genes, including ETTIN (ETT)/AUXIN RESPONSE FACTOR3 (ARF3), and all four class 1 KNOX genes were markedly elevated in as2 eal shoot apices. Rudimentary patterning in as2 eal leaves was suppressed by the ett mutation. EAL encodes BOBBER1 (BOB1), an Arabidopsis ortholog of eukaryotic NudC domain proteins. BOB1 was expressed in plant tissues with division potential and bob1 mutations resulted in lowered levels of transcripts of some cell-cycle genes and decreased rates of cell division in shoot and root apices. Coordinated cellular proliferation, supported by BOB1, and repression of all class 1 KNOX genes, ETT/ARF3 by AS2 (AS1) and BOB1 might be critical for repression of the indeterminate state and of aberrant abaxialization in the presumptive adaxial domain of leaf primordia, which might ensure the formation of flat symmetric leaves.
Keywords: ASYMMETRIC LEAVES2 (AS2), AUXIN RESPONSE FACTOR (ARF), Arabidopsis thaliana, KNOX, NudC, Leaf polarity
Introduction
Leaf primordia are clusters of cells in a determinate state at the periphery of the shoot apical meristem (SAM), which contains aggregates of indeterminate stem cells. As each leaf grows, its morphology becomes established along three axes, the proximal-distal, adaxial-abaxial and medial-lateral axes. Adaxial-abaxial patterning at the initial stage, occurring in regions adjacent to the SAM, is critical for the lateral expansion of the lamina along the medial-lateral axis for formation of flat symmetric leaves (Steeves and Sussex, 1989; Waites and Hudson, 1995; Tsukaya, 2006; Iwakawa et al., 2007; Szakonyi et al., 2010; Moon and Hake, 2010). Mechanisms that repress stem cell identity and control initial patterning for establishment of adaxial-abaxial polarity in the leaf primordium are obviously critical to plant development.
In Arabidopsis thaliana, three members of the family of class III homeodomain-leucine zipper (HD-ZIPIII) genes, namely PHABULOSA (PHB), PHAVOLUTA (PHV) and REVOLUTA (REV) are expressed in the leaf adaxial domain and determine adaxial cell fate (McConnell and Barton, 1998; McConnell et al., 2001; Emery et al., 2003) and levels of their transcripts are negatively regulated by microRNA165 (miR165) and miR166 (Bao and Barton, 2004; Mallory et al., 2004). Mutation of any HD-ZIPIII gene that confers resistance to miR165/166-mediated degradation of the corresponding transcript results in formation of radialized leaves with adaxial identity or trumpet-shaped leaves (McConnell et al., 2001; Emery et al., 2003; Mallory et al., 2004; Zhong and Ye, 2004). Genes in the YABBY (YAB) and KANADI (KAN) families promote the specification of leaf abaxial fate (Sawa et al., 1999; Siegfried et al., 1999; Bowman and Smyth, 1999; Kerstetter et al., 2001; Eshed et al., 2001; Kumaran et al., 2002; Eshed et al., 2004; Wu et al., 2008; Sarojam et al., 2010). In addition, two functionally redundant genes, ETTIN/AUXIN RESPONSE FACTOR3 (ETT/ARF3) and ARF4, whose transcripts are degraded by a trans-acting small interfering RNA (ta-siRNA), designated tasiR-ARF, in the adaxial domain of the leaf primordium to limit their specific expression to the inner region and the abaxial domain of leaves (Montgomery et al., 2008; Chitwood et al., 2009; Schwab et al., 2009), play important roles in lateral growth, as well as in specification of the abaxial fate and heteroblasty of leaves (Pekker et al., 2005; Hunter et al., 2006). The results of investigations of these effectors support the hypothesis that the specification of adaxial-abaxial polarity is tightly coupled with lateral expansion (Waites and Hudson, 1995). However, upstream factors that control the expression of such direct effectors in polarity-controlling pathways remain to be identified.
The ASYMMETRIC LEAVES2 (AS2) and AS1 genes of A. thaliana are key regulators of the formation of the flat symmetric leaves. AS2 and AS1 encode, respectively, a plant-specific nuclear protein with an AS2/LOB domain (Iwakawa et al., 2002; Shuai et al., 2002) and a nuclear protein with a myb domain (Byrne et al., 2000; Sun et al., 2002). The two proteins have been reported to form a complex (Xu et al., 2003; Yang et al., 2008) and effects of overexpression of AS2 are reported to depend on AS1 (Iwakawa et al., 2007). Mutations in these genes are associated with pleiotropic abnormalities in leaves along the three developmental axes (Rédei and Hirono, 1964; Tsukaya and Uchimiya, 1997; Byrne et al., 2000; Ori et al., 2000; Semiarti et al., 2001; Iwakawa et al., 2002). AS2 and AS1 repress the transcription of class 1 KNOTTED-like homeobox (KNOX) genes, namely, BREVIPEDICELLUS (BP)/KNAT1, KNAT2 and KNAT6, and with the exception of SHOOT-MERISTEMLESS (STM) (Byrne et al., 2000; Ori et al., 2000; Semiarti et al., 2001), which are exclusively expressed around the SAM and play roles in maintaining an indeterminate cell state (Hake et al., 2004). In addition, the AS1-AS2 complex directly represses the transcription of BP and KNAT2 (Guo et al., 2008). Some of the pleiotropic abnormalities, including short leaves, of as2 and as1 plants have been attributed to ectopic expression of BP, KNAT2 and KNAT6 (Ikezaki et al., 2010). Furthermore, AS2 and AS1 repress levels of transcripts of ETT/ARF3, KAN2 and YAB5 genes in shoot apices (Iwakawa et al., 2007), suggesting that AS2 and AS1 might be involved in adaxial development. However, genetic evidence in support of this suggestion remains to be demonstrated.
Although it has been proposed that AS2 and AS1 are involved in the establishment of adaxial polarity, abnormalities related to adaxial defects in leaves are not obvious in as2 and as1 single mutants. However, defects in polarity do develop in as2 and as1 leaves under certain growth conditions and, also, in conjunction with mutation of members of certain groups of genes (Qi et al., 2004; Inagaki et al., 2009; see Introduction of Kojima et al., 2011). These genes include several that mediate the biogenesis of tasiR-ARF (see above). Other relevant genes belong to several different groups: those for ribosome biogenesis; chromatin modification; the genome stability; and cell proliferation. While these observations do suggest genetic interactions between AS2 (also AS1) and each of these enhancer genes, our understanding of the mechanism of regulation of the expression of polarity-related effectors by AS2/AS1 is still limited.
In the present study, we report a novel enhancer mutation in A. thaliana that causes marked defects in adaxial development, with generation of abaxialized filamentous leaves, on the as2 or the as1 background. In doubly mutant in AS2 (or AS1) and the enhancer gene, levels of transcripts of many abaxial-determining genes, including ETT/ARF3 and all four class 1 KNOX genes rose markedly. Furthermore, introduction of mutation of ETT/ARF3 into the double mutant significantly suppressed the formation of filamentous leaves, with formation of flat symmetric leaves. We propose that maintenance of low levels of transcripts of ETT/ARF3 and of all class 1 KNOX genes by AS2/AS1 and the enhancer gene must be critical. AS2/AS1 and the enhancer might temporally and spatially control the developmental transition from the indeterminate meristematic state to the determinate state in the shoot apex that might be required for the formation of flat symmetric leaves.
Results
The establishment of adaxial cell fate in as2-1 eal-1 and as1-1 eal-1 double-mutant leaves was defective
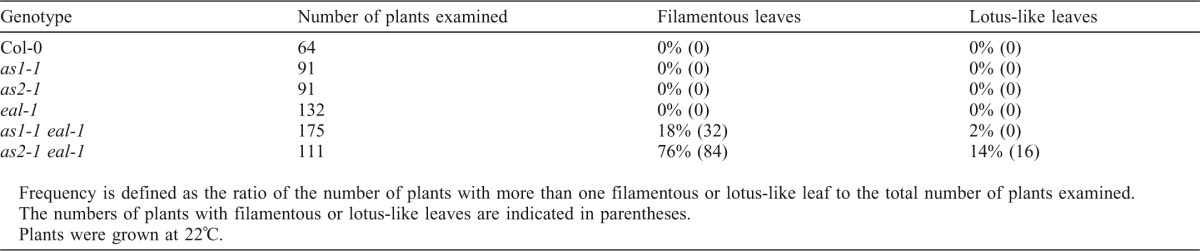
In a genetic screening for enhancers of the effect of the as1-1 mutation, we identified several mutations that generated filamentous leaves both on the as1-1 and on the as2-1 mutant background. We named one of these mutations enhancer of asymmetric leaves2 and asymmetric leaves1-1 (eal-1). The eal-1 single mutant formed flat, symmetric and pointed leaves, while the wild type and the as1, as2 and eal-1 single mutants did not have filamentous leaves (Fig. 1A–D, Table 1). In contrast to the single mutants, 18% and 2% of as1-1 eal-1 double-mutant plants had filamentous leaves and lotus-like leaves, respectively. The as2-1 eal-1 double-mutant plants produced filamentous leaves and lotus-like leaves at efficiencies of 76% and 14%, respectively (Fig. 1F–J, Table 1). Despite phenotypic similarity between the as2-1 eal-1 and as1-1 eal-1 double mutants, the frequency of formation of filamentous leaves by as2-1 eal-1 plants was much higher than that by as1-1 eal-1 plants. Therefore, as described bellow, we focused on the as2-1 eal-1 double mutant.
Fig. 1. The as1 eal and as2 eal double mutants generated filamentous leaves.
(A) Wild-type Col-0, (B) as1-1, (C) as2-1, (D) eal-1, (E) eal-1/eal-3 trans-heterozygote (TH), (F,G) as1-1 eal-1, and (H–J) as2-1 eal-1 plants. (G,I) High-magnification views of shoot apices, corresponding to boxed regions designated g and i in F and H, respectively. (J) SEM of as2 eal, showing filamentous leaves. Arrowheads indicate filamentous leaves and arrows indicate lotus-like leaves. Scale bars: 10 mm in A-E and H; 5 mm in F; 2 mm in G and I; 500 µm in J.
Table 1. Frequencies of plants with filamentous and lotus-like leaves.
Using transverse sections of leaves, we analyzed vascular patterns in as2-1 eal-1 leaves and in corresponding single-mutant leaves (Fig. 2A–F). In wild-type, as2-1 and eal-1 plants, xylem and phloem tissues were similarly located on the adaxial and abaxial sides, respectively, of the vascular bundles (Fig. 2A–C). In the as2-1 eal-1 filamentous leaves, neither phloem nor xylem cells were obvious (Fig. 2D,E). In as2-1 eal-1 double-mutant lotus-like leaves, phloem tissue was observed around xylem tissue (Fig. 2F).
Fig. 2. Filamentous leaves were abaxialized.
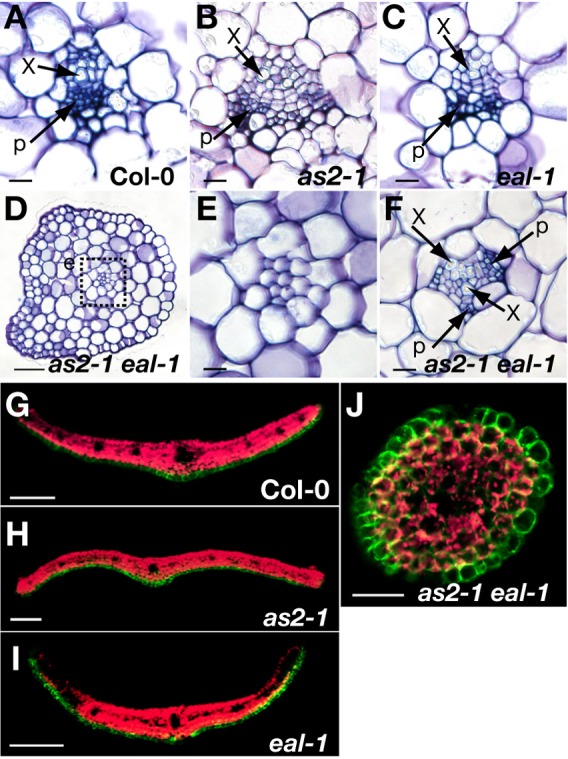
(A–F) Defects in vascular patterning in as2 eal leaves. Transverse sections of rosette leaves of (A) wild-type Col-0, (B) as2-1, (C) eal-1 and (D-F) as2-1 eal-1 plants. Sections from filamentous leaves (D,E) and lotus-like leaves (F) of the as2 eal double mutant. (E) High-magnification view of the central region of leaves, which corresponds to the boxed region designated e in D. x, xylem; p, phloem. (G–J) Expression of FILp:GFP in transverse sections of developing leaves of (G) Col-0, (H) as2-1, (I) eal-1 and (J) as2-1 eal-1 plants. Green, signals due to GFP; red, autofluorescence. Scale bars: 10 µm in A-D and F; 50 µm in E and J; 100 µm in G-I.
We investigated patterns of expression of cDNA for green fluorescent protein (GFP) under the control of the FIL promoter (the cDNA was designated FILp:GFP), which is expressed in abaxial cells of leaf primordia (Watanabe and Okada, 2003). We detected signals due to GFP only on the abaxial sides of wild-type, as2-1 and eal-1 leaves (Fig. 2G–I). By contrast, signals due to GFP were detected over the either surface of the filamentous leaves of the as2-1 eal-1 double mutant (Fig. 2J), suggesting a defect in adaxialization.
Levels of transcripts of polarity-determining and class 1 KNOX genes were elevated in as2-1 eal-1 shoot apices
We performed real-time RT-PCR using RNA from the shoot apices of wild type, as2-1, eal-1 and as2-1 eal-1 double-mutant plants. We quantified transcripts of three families of transcription-related genes that are involved in the determination of adaxial-abaxial polarity, namely, KAN genes; FIL/YAB genes; and ARF genes, which separately and redundantly specify abaxial cell fate; genes in the HD-ZIP III family (PHB, PHV, and REV), which specify adaxial cell fate; and all the genes in the class 1 KNOX gene family, which are expressed in the SAM and its periphery in wild-type plants.
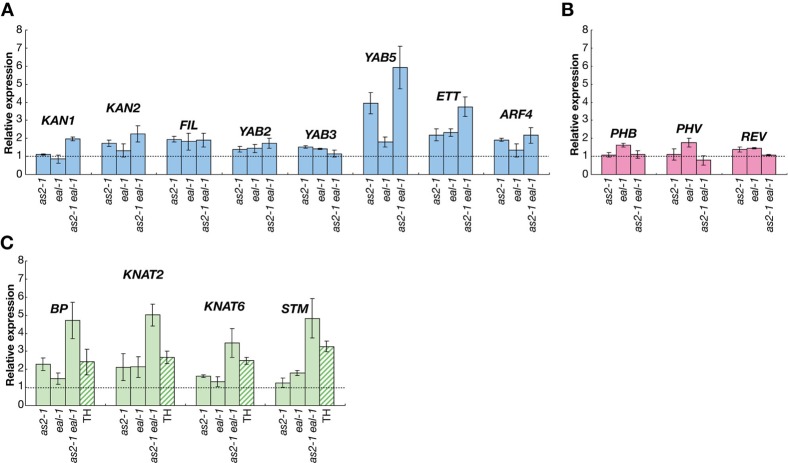
As shown in Fig. 3A, levels of transcripts of a number of genes that are involved in the establishment of abaxial cell fate (KAN1, KAN2, YAB5 and ETT/ARF3) were higher in the as2-1 eal-1 double mutant than in the wild type and in the corresponding single mutants. Levels of transcripts of ETT/ARF3, FIL and YAB5 were elevated in the eal-1 mutant, while those of KAN2, FIL, ETT/ARF3 and YAB5 were elevated in the as2-1 mutant. However, levels of HD-ZIP III transcripts in the double mutant were not significantly different from those in the wild type (Fig. 3B). These results suggest that the filamentous leaves of as2-1 eal-1 plants had accentuated abaxialized features. Levels of transcripts of all class 1 KNOX genes (BP, KNAT2, KNAT6 and STM) were also much higher in the as2-1 eal-1 double mutant than in the wild type and in the as2-1 and eal-1 single-mutant plants.
Fig. 3. Levels of transcripts of genes involved in the determination of leaf polarity and of class 1 KNOX genes.
Levels of transcripts of genes that are involved in (A) abaxialization and (B) adaxialization of leaves; those of (C) class1 KNOX genes; and those of corresponding genes in wild-type Col-0 plants were measured by quantitative real-time RT-PCR (qRT-PCR). Total RNA was extracted from shoot apices of 18-day-old wild-type, as2-1, eal-1 and as2-1 eal-1 plants and of 15-day-old Col-0 and eal-1/eal-3 trans-heterozygotes, designated TH. Each value from 18-day-old plants was normalized by reference to the level of POLYUBIQUITIN 10 (UBQ10, At4g05320) transcripts and those from 15-day-old plants were normalized by reference to the level of ß-6-tubulin (At5G12250) transcripts. The values from wild-type plants were set arbitrarily at 1.0. Bars indicate the s.d. among more than three biological replicates.
Since eal-1 is a weak allele and other eal alleles were embryonic-lethal (see below), we generated trans-heterozygotes (eal-1/eal-3; TH). The eal-1/eal-3 plants had a dwarf phenotype, with pointed leaves that were smaller than those of eal-1 plants (Fig. 1E). We examined the effects of the single mutation in the EAL gene on levels of transcripts of class 1 KNOX genes in shoot apices. As shown in Fig. 3C, levels of transcripts of all four class 1 KNOX genes were markedly elevated in eal-1/eal-3 (TH) shoot apices.
ETT/ARF3 was involved in the polarity defects in as2-1 eal-1 leaves
Since the transcript level of ETT/ARF3 was elevated in as2-1 eal-1, we examined effects of a mutation of ETT/ARF3 on phenotypes of as2-1 eal-1. We introduced the ett-13 mutation, into the as2-1 eal-1 double mutant to generate the as2-1 eal-1 ett-13 triple mutant. As shown in Figs 4A and 4B, the phenotype of eal-1 ett-13 mutant plant was similar to that of the eal-1 plant. Most of our as2-1 eal-1 ett-13 triple mutants (79%) had symmetrically expanded leaves and no filamentous or lotus-like leaves (Fig. 4C,D). Thus, the polarity defects of the as2-1 eal-1 double mutant were efficiently suppressed by the ett-13 mutation, indicating that increase in the level of ETT/ARF3 transcripts was responsible for the adaxial defect.
Fig. 4. The ett mutation efficiently suppressed the abnormal phenotype of as2-1 eal-1 leaves.
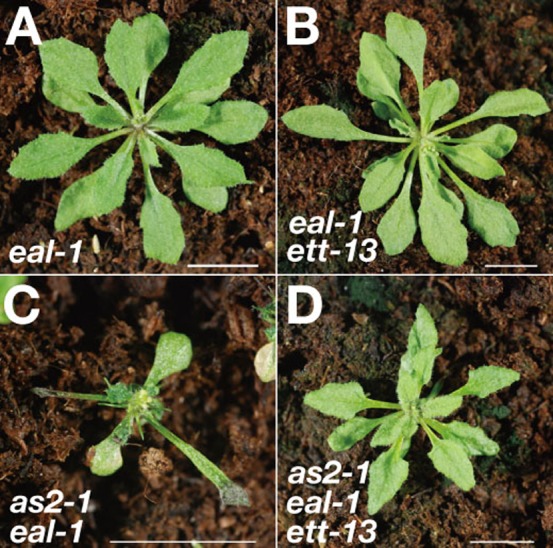

Gross morphology of (A) eal-1, (B) eal-1 ett-13, (C) as2-1 eal-1 and (D) as2-1 eal-1 ett-13 triple-mutant plants. Scale bars: 10 mm.
The eal-1 mutation was located in the BOBBER1 (BOB1) gene that encodes a homolog of the NudC protein of Aspergillus nidulans
We identified the eal mutation as a mutation in the BOB1 gene, which encodes 304 amino acid residues. BOB1 is homologous to nuclear distribution gene C (nudC) of Aspergillus nidulans (Fig. 5A, supplementary material Fig. S1) (Jurkuta et al., 2009). The eal-1 mutation, which was identical to the bob1-3 mutation (Perez et al., 2009), was generated by a G-to-A base substitution that caused the replacement of Gly by Glu at position 141 (Fig. 5A).
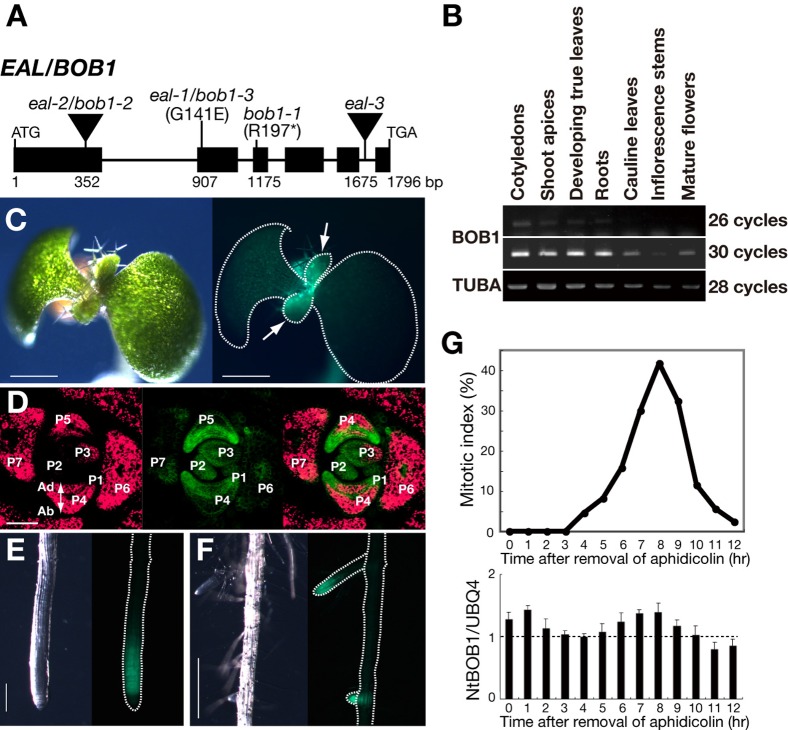
Fig. 5. The EAL gene encodes BOBBER1, a homolog of NudC.
(A) Exon-intron (black rectangle-thin line) organization between the initiation and termination codons of the EAL/BOB1 gene and sites of point mutations in eal-1/bob1-3 and bob1-1 and of T-DNA insertions in eal-2/bob1-2 and eal-3. (B) Accumulation of BOB1 transcripts in wild-type Col-0 as determined by RT-PCR. Transcripts were detected in the cotyledons, developing true leaves and shoot apices of 10-day-old plants; in the third and fourth rosette leaves and roots of 19-day-old plants; and in the cauline leaves, inflorescence stems and mature flowers of 40-day-old plants. (C–F) The expression of BOB1::GFP, driven by the BOB1 promoter, in eal-1 plants. (C,E,F) Panels include a bright-field image and a fluorescence image. (D) The expression of BOB1::GFP in a transverse section of an 11-day-old vegetative shoot apex. Expression in leaf primordia at different growth stages. P1-P7 indicate stages of development; the larger the number, the older the primordium. (E) Expression at the tip of a primary root and (F) a lateral root of 6- or 8-day-old plants. Green, signals due to GFP; red, autofluorescence. Ad, adaxial side; Ab, abaxial side. (G) Patterns of accumulation of the NtBOB1 transcript during progression of the cell cycle in BY-2 cells that had been synchronized with aphidicolin. Mitotic indices are shown in the upper panel. Levels of transcripts at indicated times were determined by qRT-PCR (lower panel). Scale bars: 500 µm in C and F; 100 µm in D; 200 µm in E.
We obtained two T-DNA insertion lines, eal-2 (SALK_001125) and eal-3 (GK_406_D03) (Fig. 5A, supplementary material Fig. S1). The eal-2 mutation was the same as bob1-2 (Jurkuta et al., 2009), while eal-3 was a new allele of BOB1. The embryonic development of eal-3 homozygotes was arrested at the globular stage, and this phenotype is similar to that of bob1-1 and bob1-2 homozygotes (supplementary material Fig. S2) (Jurkuta et al., 2009; Perez et al., 2009).
BOB1 was expressed in tissues with cell-division potential
BOB1 transcripts accumulated in tissues, such as shoot apices, developing rosette leaves and roots, that contain division-competent cells (Fig. 5B). To prepare a functional reporter construct, we cloned the genomic DNA that contained the 2,181-bp 5′-upstream region and the 1,796-bp coding region of BOB1. We fused this genomic DNA, in frame, to the GFP reporter gene at the last codon of the BOB1 gene to create the fusion gene pBOB1::BOB1::GFP, which we then introduced into the eal-1 mutant. The fusion gene restored a normal phenotype and complemented the eal-1 mutation in all transgenic lines (supplementary material Fig. S3).
In the aerial parts of the transgenic plants, signals due to GFP were most abundant in shoot apices with developing leaves (Fig. 5C), which contain strongly division-competent cells. We also examined patterns of expression of BOB1::GFP in transverse sections of shoot apices (Fig. 5D). Although the detected signals throughout leaf primordia at early stages (P1-P3), the intensities of signals in central regions and the epidermis were stronger than those from other cells at later stages (P4 and P5) (Fig. 5D). As the primordia grew, the intensity of signals fell rapidly (P6 and P7). We also observed signals due to GFP at the tips exclusively of primary and lateral roots (Fig. 5E,F). These observations suggest a potential role for BOB1 in cell-division ability and/or in maintenance of cells in an indeterminate state.
Using a BOB1 homolog of Nicotiana tabacum (NtBOB1), we quantified levels of NtBOB1 transcripts by real-time RT-PCR during progression of the cell cycle in synchronized tobacco BY-2 cells. Fig. 5G shows that accumulation of the NtBOB1 transcript exhibited two peaks that effectively overlapped progression through the S and M phase, respectively. However, the relative heights of the peaks were small. Thus, it seems likely that transcription of NtBOB1 is controlled, at least to a small extent, by the cell cycle.
BOB1 complemented defects in colony growth and the movement of nuclei in the nudC3 mutant of Aspergillus nidulans
Colonies of the nudC3 mutant of A. nidulans exhibit temperature-sensitive growth, and the migration of nuclei that normally occurs prior to the formation of cell plates during cytokinesis is also temperature-sensitive (Osmani et al., 1990; Chiu and Morris, 1995; Chiu and Morris, 1997). We examined whether BOB1 might allow nudC3 mutant cells to grow normally at an elevated temperature with normal movement of nuclei using BOB1 cDNAs driven by the alcA promoter of A. nidulans (Fig. 6A).
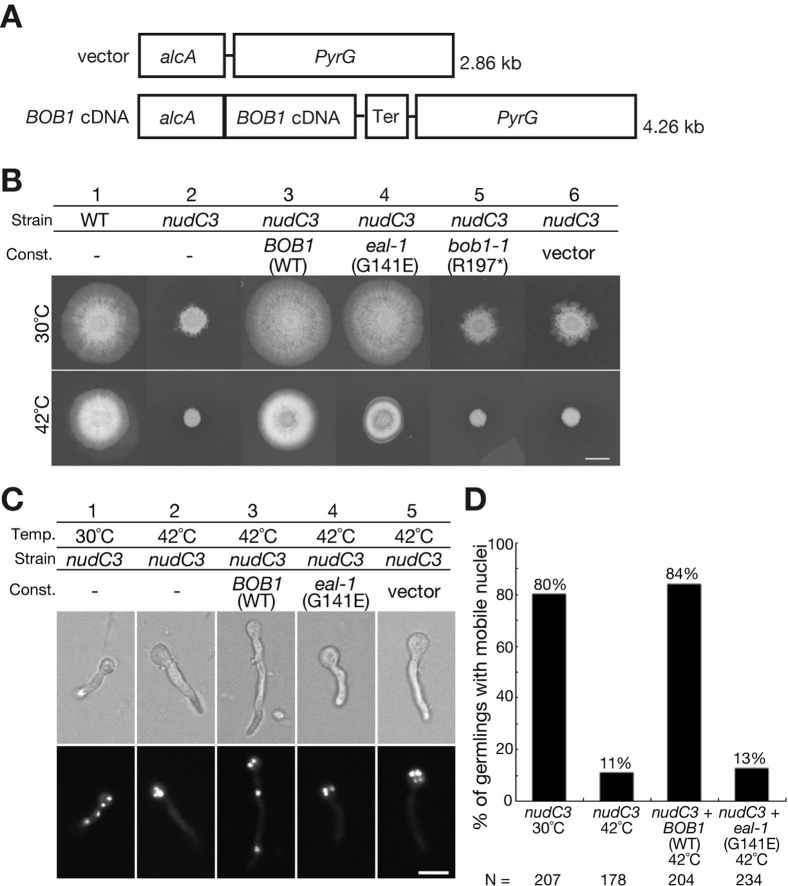
Fig. 6. The BOB1 gene of Arabidopsis rescues the temperature-sensitive nudC3 mutation of in A. nidulans.
(A) Schematic representation of the DNA constructs used for complementation tests. The vector included the inducible/repressible alcA promoter, the pyrG gene of A. nidulans and the terminator sequence (Ter) of Aspergillus oryzae Taka-amylase A. The wild-type and mutant BOB1 cDNA (eal-1 or bob1-1) were cloned, separately, downstream of the alcA promoter. The unmodified vector was used as a negative control. (B) Complementation of the growth defect of the nudC3 mutant by BOB1. Cells of non-transformed wild-type A. nidulans (column 1), the nudC3 mutant (column 2) and nudC3 harboring the BOB1 (column 3), eal-1 (column 4), bob1-1 (column 5) or the vector (column 6) construct were incubated at 30°C or 42°C for 4 days. Scale bar: 10 mm. (C) Complementation of the defect in the movement of nuclei by BOB1. Spores from the nudC3 mutant were germinated and the mutant was grown at 30°C (column 1) and at 42°C (columns 2-5) for 10 hours. The germlings were stained with DAPI. Scale bar: 10 µm. (D) The overall rates of movement of nuclei. Germlings into which nuclei had moved under the indicated growth conditions were counted. N represents the number of germlings scored under each growth condition.
Even at the permissive temperature (30°C), nudC3 colonies were smaller than wild-type colonies (Fig. 6B, columns 1 and 2). At the restrictive temperature (42°C), the nudC3 colonies were much smaller than the wild-type colonies. Wild-type BOB1 cDNA fully reversed the defects in colony growth of nudC3 cells at both temperatures (Fig. 6B, column 3). Moreover, eal-1 cDNA also reversed the growth defects at 30°C and 42°C, but eal-1 cDNA was slightly less effective than BOB1 cDNA (Fig. 6B, column 4). As anticipated, bob1-1 cDNA failed to reverse the growth defects at 30°C and 42°C (Fig. 6B, columns 2, 5 and 6).
As shown in Fig. 6C,D, BOB1 cDNA rescued the defect in nuclear migration in nudC3 cells at 42°C, and 84% of nuclei in nudC3 cells that had been transformed with BOB1 cDNA were normally distributed. By contrast, eal-1 cDNA yielded only the background level of movement of nuclei.
Mutations in BOB1 affected the progression of the cell cycle in shoot and root apices
Using cross sections of shoot apices, we investigated the structure of the shoot apical meristem (SAM) in wild-type, as2-1, eal-1, as2-1 eal-1 and eal-1/eal-3 trans-heterozygous (TH) plants. The number of cells in the L1 and L2 layers in the region of the SAM was depressed in the eal-1 and as2 eal-1 mutants (Fig. 7A,B). In addition, the structure of the as2-1 eal-1 SAM was disorganized. The number of cells in the SAM was reduced still further in the eal-1/eal-3 SAM, suggesting reduced cell-division ability in the eal-1/eal-3 SAM.
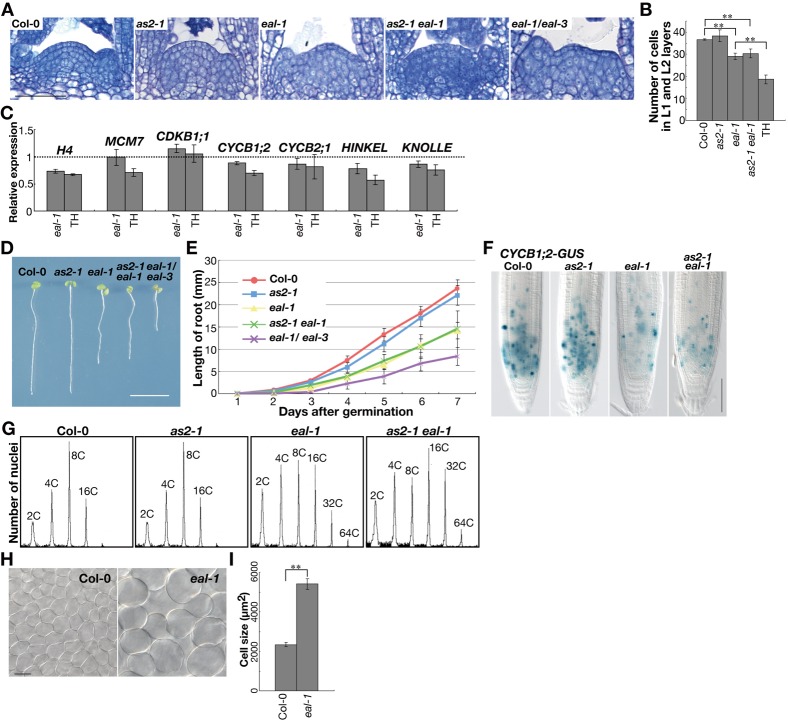
Fig. 7. Mutation of the BOB1 gene negatively affects cell division.
(A) Toluidine blue-stained sections of shoot apices from 9-day-old wild-type Col-0, as2-1, eal-1, as2-1 eal-1 and eal-1/eal-3 trans-heterozygote (TH) plants. (B) Number of cells in L1 and L2 layers of meristem domes in 9-day-old wild-type, as2-1, eal-1, as2-1 eal-1 and eal-1/eal-3 trans-heterozygote plants. Bars indicate the s.e.m. among three plants. (C) Expression of transcripts, as determined by qRT-PCR. Relative levels of expression of cell cycle-related genes in the shoot apices of 15-day-old eal-1/eal-3 (TH) plants, relative to those of Col-0. Each value was normalized by reference to levels of transcripts of ß-6-tubulin (At5G12250). Bars indicate the s.d. among more than three biological replicates. (D) Seedlings of 7-day-old Col-0, as2-1, eal-1, as2-1 eal-1 and eal-1/eal-3 (TH) plants. (E) Root growth of Col-0, as2-1, eal-1 and as2-1 eal-1 plants from 1 to 7 days after germination. Averages ± s.d. for roots of more than 10 plants of each genotype are shown. The root length was measured using ImageJ software (http://rsb.info.nih.gov/ij/). (F) Patterns of staining for GUS activity in primary root tips of 6-day-old Col-0, as2-1, eal-1 and as2-1 eal-1 plants that harbored CYCB1;2-GUS. (G) Flow cytometric analysis of ploidy using nuclei prepared from first and second leaves of 18-day-old Col-0, as2-1, eal-1 and as2-1 eal-1 plants. (H) Palisade cells in first leaves of 21-day-old Col-0 and eal-1 plants. (I) Quantitative analysis of cell size in the first leaves of Col-0 and eal-1 plants. Bars indicate the s.d. Scale bars: 50 µm in A; 10 mm in D; 100 µm in F; and 10 µm in H. Significant differences were determined by Student's t-test. **P<0.01 (B,I).
We quantified transcripts of genes that are involved in progression of the cell cycle in shoot apices of wild-type, eal-1, and eal-1/eal-3 plants. As shown in Fig. 7C, in eal-1, levels of transcripts of histone H4, MINICHROMOSOME MAINTENANCE-7 (MCM7), CYCLIN B1;2 (CYCB1;2), HINKEL and KNOLLE genes were slightly depressed, and the extent of such depression was much greater in eal-1/eal-3 plants. The eal-1 mutant had short roots and the eal-1/eal-3 mutant had even shorter roots (Fig. 7D). The extent of root elongation in eal-1 (60%) and eal-1/eal-3 (30-40%) plants was significantly lower than in wild-type plants (Fig. 7E). To examine the efficiency of cell division, we monitored the expression of CYCB1;2-ß-glucuronidase (GUS), in which the CYCB1;2 genomic sequence containing the promoter region was fused to the GUS gene, which we used as a marker of the G2-M transition. In eal-1 and as2-1 eal-1 seedlings, the intensity and number of signals due to GUS were markedly lower than those in the wild type (Fig. 7F). These results suggest that cell-division ability is reduced in the eal-1 mutant.
We measured the ploidy of nuclei in the first two leaves of 18-day-old wild-type, as2-1, eal-1 and as2-1 eal-1 plants by flow cytometry (Fig. 7G). In wild-type and as2-1 leaves, patterns of ploidy from 2C to 16C were similar. However, the eal-1 mutant contained nuclei with higher ploidy (from 32C and 64C). The ploidy in the as2-1 eal-1 double mutant was even higher. Palisade cells of the eal-1 mutant were also more than twice the size of those of wild-type plants (Fig. 7H,I). These results indicate that mutations in BOB1 resulted in early entry into the endocycle and that increases in ploidy were exacerbated by the as2-1 mutation.
Discussion
Our present study showed that the eal-1 mutation in the BOB1 gene enhances defects in the adaxial development of as2 leaves, converting flat leaves to abaxialized filamentous leaves. This effect is attributable to elevated levels of transcripts of the ETT/ARF3 gene (Figs 1–5). The BOB1 gene, in cooperation with the AS2 (AS1) gene, plays a crucial developmental role via repression of both the indeterminate state and the abaxial fate of cells in the presumptive adaxial domain of leaf primordia after the commitment to leaf initiation around the shoot apical meristem, promoting the establishment of the adaxial polarity of leaves (Fig. 8). Thus, repressive activity is required for the formation of flat symmetric leaves with appropriate adaxial-abaxial polarity.
Fig. 8. Roles of AS1, AS2 and BOBBER1 in shoot apices during the formation of flat symmetric leaves.
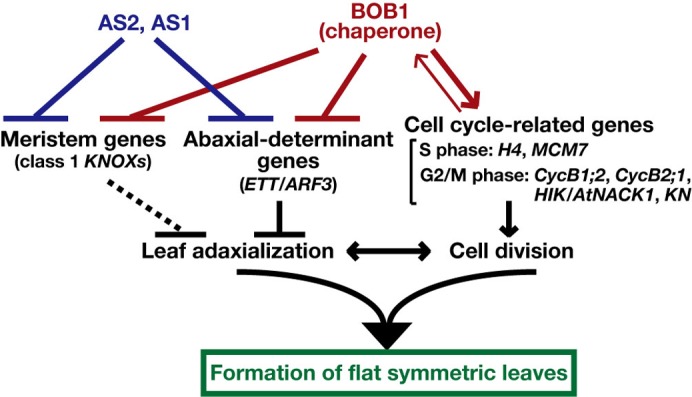
The AS1-AS2 complex and BOB1 act independently to repress the levels of transcripts of leaf abaxial-determinant genes, which include ETT/ARF3, and of “meristem” genes, namely class1 KNOX genes. The relationship between class 1 KNOX genes and the differentiation of the leaf adaxial side of leaves remains to be clarified. BOB1 acts positively to regulate cell division for the formation of flat symmetric leaves via regulation of the transcription of cell cycle-related genes. The balance between leaf adaxialization and cell division is critical for the formation of flat symmetric leaves.
BOB1 is involved in the repression of transcription of genes involved in leaf abaxialization via an unknown pathway that is independent of the AS2 (AS1) pathway
How do BOB1 and AS2 (AS1) act together to regulate the repression of levels of ETT/ARF3 transcripts? AS2 transcripts are detected in the adaxial region of leaf primordia and AS1 transcripts are detected in the inner region of leaves that includes the vasculature (Iwakawa et al., 2007). Regions of expression of BOB1 in leaf primordia are overlapped sites at which AS2 and AS1 transcripts were found (Fig. 5). The levels of the ETT/ARF3 transcript in shoot apices of as2-1 and eal-1 were higher than in the wild type, while that in the as2-1 eal-1 double mutant was even higher still (Fig. 3A). Let us consider the following two possibilities; BOB1 might act on the repression of transcription of the ETT/ARF3 gene via an unidentified pathway that might be independent of the AS2 (AS1)-mediated pathway (Fig. 8). Alternatively, both BOB1 and AS2 (AS1) might function in the same pathway. The overlap among regions of expression of the BOB1, AS2 and AS1 genes in the shoot apex (Fig. 5C,D) (Byrne et al., 2000; Iwakawa et al., 2007) supports both possibilities. However, our bob1 and as2 (as1) mutants had completely different morphology (Fig. 1), suggesting that the second possibility is unlikely. Similarly, it seems plausible that AS2 and BOB1 might repress the expression of class 1 KNOX genes via two independent pathways.
BOB1 is involved in progression of the cell cycle and in the extent of expression of class 1 KNOX genes in the shoot apex, acting in cooperation with AS2 (AS1)
We have described six lines of evidence in support of the hypothesis that BOB1 might be involved in cell division. Mutations in BOB1 caused (1) a reduction in the number of cells in shoot apices; (2) a delay of root growth, due, perhaps, to slow cell division; (3) a decrease in levels of transcripts of genes involved in the cell cycle; and (4) early transition to the endocycle, which might occur during the progression of G2 phase (Fig. 7). In addition, we observed (5) two peaks in levels of transcripts of a tobacco homolog of BOB1, during the cell cycle, namely, during the S and M phases; and (6) elevated levels of expression of BOB1 in leaf primordia at early developmental stages and in primary and lateral root tips, where the potential for cell division is high (Fig. 5). These results imply that mutations in BOB1 result in a decrease in the rate of progression of the cell cycle and/or in the frequency of onset of cell division around shoot and root apical meristems. Such impaired cell division in bob1 tissues might be due to lower levels of the transcripts of cell cycle-related genes, as described above. Although the wild-type BOB1 gene plays an inhibitory role in the expression of ETT/ARF3 and class 1 KNOX genes (Fig. 3A), it acts positively to maintain appropriate levels of transcripts of cell cycle-related genes to ensure the proper progression of the cell-division cycle in shoot and root apices (Figs 7C, 8).
Our observation that levels of transcripts of all four class 1 KNOX genes were significantly elevated in the eal-1/eal-3 shoot apex (Fig. 3C; TH) are consistent with previous reports of the expansion of the domain of expression of the STM gene in bob1 embryos (Jurkuta et al., 2009). In addition, levels of transcripts of all class 1 KNOX genes were even higher and the expression domain of the BP gene was even more extensive in petioles at the as2-1 eal-1 shoot apex than at the shoot apex of the corresponding single mutants (Fig. 3C, supplementary material Fig. S4). These results predict that the size of the population of cells in the indeterminate state is enhanced in the shoot apex of the double mutant. Effects of the expansion of the expression domain of class 1 KNOX genes in the shoot apex on development of the adaxial domain remains to be examined.
Relationship between the molecular function and developmental role of BOB1
In A. nidulauns, the nudC gene is required for regulation of the movement of nuclei during the asexual reproductive cycle, as well as for deposition of the cell wall, colony growth and viability (Osmani et al., 1990; Chiu et al., 1997). Homologs of NudC from mammals, Drosophila, Caenorhabditis elegans and Arabidopsis complement nudC3 mutation of A. nidulans, restoring the normal movement of nuclei and colony growth (Fig. 6) (Miller et al., 1999; Morris et al., 1997; Cunniff et al., 1997; Dawe et al., 2001). These observations suggest that the function of NudC in the movement of nuclei is evolutionarily conserved in eukaryotes.
Cappello et al. (2011) showed that mammalian NudC is required for the migration of nuclei and of neurons during neocortical development of the brain. In addition, suppression of the expression of genes homologous to nudC led to defects in cytokinesis and chromosome congression during karyokinesis in cultured non-neural cells (Aumais et al., 2003; Nishino et al., 2006; Zhang et al., 2002; Zhou et al., 2003; Zhou et al., 2006). Thus, mammalian homologs of NudC appear to be involved both in organogenesis and in cell division. BOB1 might play similar roles in both phenomena.
Several homologs of NudC, including BOBBER1, have been shown to act as molecular chaperones in vitro, and a role as chaperone is the only known molecular function of these homologs (Faircloth et al., 2009; Perez et al., 2009; Zhu et al., 2010). The bob1-3 (identical to eal-1) mutation does not affect such chaperone activity in vitro (Perez et al., 2009), but this mutation did abolish the migration of nuclei, in A. nidulans (Fig. 6). The mutation also acted as an effective enhancer of defects in adaxial development in as2 leaves and depressed the efficiency of cell division in shoot and root apices (Figs 1, 2). Thus, we found no obvious correlation between the chaperone activity of eal-1 protein and the developmental and cellular abnormalities in eal-1 mutant plants. These observations suggest that BOBBER1 not only acts as a molecular chaperone but also has some other unidentified function. However, the observed absence of a correlation between the activity as a chaperone and defects in leaf development and cell division in eal-1 plants might be explained by the assumption that the developmental effects of the eal-1 mutation are greater in vivo than the effects of the eal-1 mutation on chaperone activity in vitro.
BOB1 acts together with AS2 (AS1) as a modulator in the formation of flat leaves
As discussed above, BOB1 plays positive roles in cell division, as well as in the establishment of adaxial polarity in leaves. Waites and Hudson (1995) proposed that the cell proliferation required for the lateral growth of the leaf lamina might be tightly coupled with the establishment of adaxial-abaxial polarity in the leaf primordium. As proposed in Fig. 8, the balance between leaf adaxialization and cell division might be coordinately controlled to ensure the development of flat symmetric leaves. Since BOB1 is involved in both the stimulation of cell division and the establishment of adaxialization, it might act, together with AS2 (AS1), as a crucial factor that appropriately modulates cell division and development.
Materials and Methods
Plant materials and growth conditions
Seeds of Arabidopsis thaliana ecotype Col-0 (CS1092), as2-1 (CS3117), as1-1 (CS3374) and eal-2 (SALK_001125) were obtained from the Arabidopsis Biological Resource Center (ABRC; Ohio State University, Columbus, OH, USA). Seeds of the eal-3 (GK_476D03) mutant were obtained from the Nottingham Arabidopsis Stock Center (NASC; University of Nottingham, Loughborough, UK). Seeds of ett-13 (SALK_040513) mutant and CYCB1;2-GUS were kindly provided by Dr Yuval Eshed (Weizmann Institute of Science, Israel) and Dr Masaki Ito (Nagoya University, Japan), respectively. Growth conditions for plants were described previously (Semiarti et al., 2001).
Isolation and mapping of the EAL gene
To isolate the eal-1 mutant, seeds of as1-1 were mutagenized by 0.03% ethyl methanesulfonate and approximately 5,500 M2 plants from 201 M1 stocks were screened on the basis of leaf phenotype. We crossed the eal-1 mutant with wild-type Col-0 and the segregation rate of F2 plants suggested that the enhancement of the as1 phenotype was due to single recessive locus. To map the EAL locus, we crossed the as1-1 eal-1 double mutant with wild-type Landsberg erecta (Ler). Samples of genomic DNA from 1,306 F2 plants were analyzed in terms of simple sequence length polymorphism and cleaved-amplified polymorphism markers, which were generated on the basis of polymorphism data provided by Cereon (http://www.Arabidopsis.org/Cereon). The EAL locus was mapped to an approximately 30-kb region on chromosome 5, which contains eleven predicted genes. We compared the genomic sequences of the wild type and the eal-1 mutant and found that the eal-1 mutant harbored a mutation at the 19th nucleotide of the second exon of At5g53400. For genotyping of eal-1, we amplified genomic DNAs by the polymerase chain reaction (PCR) using the primers listed in supplementary material Table S1 and digested the DNA products with the restriction enzyme XmnI (New England Biolabs, Beverly, MA, USA).
Construction of pBOB1::BOB1::GFP
A 4.1-kb fragment of genomic DNA from a wild-type plant (Col-0), encompassing the 2.3-kb upstream sequence of BOB1 and the 1.8-kb coding region of BOB1, was amplified by PCR with a pair of primers (supplementary material Table S2). The resultant DNA fragment was inserted into pENTR/D-TOPO (Invitrogen, Carlsbad, CA, USA), and then the product was introduced into the binary vector pGWB4 (Nakagawa et al., 2007) with the GFP coding sequence. We transformed the eal-1 mutant using Agrobacterium tumefaciens strain GV3101 that harbored the plasmid with the incorporated DNA fragment, as described by Bechtold and Pelletier (1998).
RT-PCR
Manipulation of DNA and qRT-PCR were performed as described by Iwakawa et al. (2007). Primer sets for RT-PCR and qRT-PCR are listed in supplementary material Table S3. Results were normalized by reference to results for UBQ10 (At4g05320), ß-6-tubulin (At5G12250) or α-tubulin (At5G19770).
Fluorescence microscopy
We crossed as2-1 eal-1 and FILp:GFP (Watanabe and Okada, 2003; Ueno et al., 2007). Shoot apices containing leaf primordia were embedded in 5% agar and then agarose blocks were sliced into sections with a vibratome. Fluorescence was observed with a confocal laser scanning microscope (LSM510 META; Carl Zeiss Inc., Oberkochen, Germany). We observed fluorescence images of tissues derived from plants that harbored pBOB1::BOB1::GFP with a stereomicroscope (SteREO Lumar.V12; Carl Zeiss Inc., Jena, Germany).
Detection of GUS activity and histological analysis
The activity of GUS in roots was detected as described by Iwakawa et al. (2007). Plant tissues were prepared for thin sectioning as described by Ueno et al. (2007). Scanning electron microscopy (SEM) was performed as described previously (Semiarti et al., 2001).
Measurements of ploidy
Ploidy analysis was performed as described by Suzuki et al. (2005).
Strains of Aspergillus nidulans, growth conditions and transformation
Aspergillus nidulans strains FGSC A773 (pyrG89; wA3; pyroA4) and FGSC A779 (nudC3 pyrG89 pabaA1; wA2; nicA2) were obtained from the Fungal Genetics Stock Center (FGSC; University of Missouri, Kansas City, MO, USA). The strains were grown at a permissive temperature of 30°C or at a restrictive temperature of 42°C in standard minimal medium (Rowlands and Turner, 1973) supplemented with appropriate requirements. The minimal medium for A773 was supplemented with 0.0002% pyridoxine HCl, 0.12% uridine and 0.11% uracil. The minimal medium for A779 was supplemented with 0.0002% niacin, 0.0002% p-aminobenzoic acid, 0.12% uridine and 0.11% uracil (Morris et al., 1997). The promoter of alcA, the gene for alcohol dehydrogenase I of A. nidulans, was fused to full-length BOB1 cDNA and the fused construct and the PyrG coding sequence were subcloned into pBluescript II KS+ (Stratagene, La Jolla, CA, USA). The eal-1 and bob1-1 mutations in BOB1 cDNA were induced with a QuikChange Multi Site-Directed Mutagenesis Kit (Stratagene, La Jolla, CA, USA). A. nidulans was transformed as described by Makita et al. (2009). Transformants were grown in liquid medium or on 1.4% agar-solidified medium, which were the minimal medium for A779 without uridine and uracil.
DAPI staining of nuclei
To stain nuclei of developing germlings, conidia of A779 and the transformants were incubated in 10 ml of each liquid standard minimal medium for 10 hours. Then, we added 4′,6-diamidino-2-phenylindole (DAPI) and Triton X-100 to final concentrations of 0.1 µg/ml and 0.1%, respectively, and incubated cells for a further 10 minutes. We observed DAPI fluorescence under an Olympus microscope (BX51TRF; Olympus, Tokyo, Japan).
BY-2 cells and synchronization
Maintenance of tobacco suspension-cultured BY-2 cells and synchronization of the BY-2 cell cycle at the G1/S boundary were performed as described previously (Nishihama et al., 2002). RNA was extracted from BY-2 cells with an RNeasy kit (QIAGEN, Hilden, Germany) and poly(A)+ RNA was isolated with Dynabeads® (Dynal Biotech, Lake Success, NY, USA). Reverse transcription was performed with a First-Strand cDNA Synthesis Kit (GE Healthcare, Buckinghamshire, UK). Primer sets for qRT-PCR are listed in supplementary material Table S1. Results were normalized by reference to results for NtUBQ4.
Supplementary Material
Acknowledgments
The authors thank Dr Masaki Ito (Nagoya University, Japan) for helpful discussions and providing seeds of CYCB1;2-GUS and primer sets for qRT-PCR and Dr Ken Kosetsu (Nagoya University, Japan) for providing the cDNA samples of synchronized BY-2 cells. They thank Dr Yuval Eshed (Weizmann Institute of Science, Israel) for helpful discussions and providing seeds of the ett-13 mutant. They also thank Drs. Yoko Matsumura, Michiko Sasabe (Nagoya University, Japan), Ayami Nakagawa and Mayumi Iwasaki (Chubu University, Japan), and other members of Kobayashi and Machida laboratory for their encouragements and supports. This work was supported, in part, by a Grant-in-Aid for Scientific Research on Priority Areas (no. 19060003) from the Ministry of Education, Culture, Sports, Science and Technology of Japan (MEXT). N.I. was supported by a Grant-in-Aid for the Global Center of Excellence Program, awarded to the Division of Biological Science of Nagoya University from MEXT.
References
- Aumais J. P., Williams S. N., Luo W., Nishino M., Caldwell K. A., Caldwell G. A., Lin S. H., Yu-Lee L. Y. (2003). Role for NudC, a dynein-associated nuclear movement protein, in mitosis and cytokinesis. J. Cell Sci. 116, 1991–2003 10.1242/jcs.00412 [DOI] [PubMed] [Google Scholar]
- Bao N., Lye K. W., Barton M. K. (2004). MicroRNA binding sites in Arabidopsis class III HD-ZIP mRNAs are required for methylation of the template chromosome. Dev. Cell 7, 653–662 10.1016/j.devcel.2004.10.003 [DOI] [PubMed] [Google Scholar]
- Bechtold N., Pelletier G. (1998). In planta Agrobacterium-mediated transformation of adult Arabidopsis thaliana plants by vacuum infiltration. Methods Mol. Biol. 82, 259–266 [DOI] [PubMed] [Google Scholar]
- Bowman J. L., Smyth D. R. (1999). CRABS CLAW, a gene that regulates carpel and nectary development in Arabidopsis, encodes a novel protein with zinc finger and helix-loop-helix domains. Development 126, 2387–2396 [DOI] [PubMed] [Google Scholar]
- Byrne M. E., Barley R., Curtis M., Arroyo J. M., Dunham M., Hudson A., Martienssen R. A. (2000). Asymmetric leaves1 mediates leaf patterning and stem cell function in Arabidopsis. Nature 408, 967–971 10.1038/35050091 [DOI] [PubMed] [Google Scholar]
- Cappello S., Monzo P., Vallee R. B. (2011). NudC is required for interkinetic nuclear migration and neuronal migration during neocortical development. Dev. Biol. 357, 326–335 10.1016/j.ydbio.2011.07.001 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chitwood D. H., Nogueria F. T., Howell M. D., Montgomery T. A., Carrington J. C., Timmermans M. C. (2009). Pattern formation via small RNA mobility. Genes Dev. 23, 549–554 10.1101/gad.1770009 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chiu Y. H., Morris N. R. (1995). Extragenic suppressors of nudC3, a mutation that blocks nuclear migration in Aspergillus nidulans. Genetics 141, 453–464 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chiu Y. H., Morris N. R. (1997). Genetic and molecular analysis of a tRNALeu missense suppressor of nudC3, a mutation that blocks nuclear migration in Aspergillus nidulans. Genetics 145, 707–714 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chiu Y. H., Xiang X., Dawe A. L., Morris N. R. (1997). Deletion of nudC, a nuclear migration gene of Aspergillus nidulans, causes morphological and cell wall abnormalities and is lethal. Mol. Biol. Cell 8, 1735–1749 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cunniff J., Chiu Y. H., Morris N. R., Warrior R. (1997). Characterization of DnudC, the Drosophila homolog of an Aspergillus gene that functions in nuclear motility. Mech. Dev. 66, 55–68 10.1016/S0925-4773(97)00085-3 [DOI] [PubMed] [Google Scholar]
- Dawe A. L., Caldwell K. A., Harris P. M., Morris N. R., Caldwell G. A. (2001). Evolutionarily conserved nuclear migration genes required for early embryonic development in Caenorhabditis elegans. Dev. Genes Evol. 211, 434–441 10.1007/s004270100176 [DOI] [PubMed] [Google Scholar]
- Emery J. F., Floyd S. K., Alvarez J., Eshed Y., Hawker N. P., Izhaki A., Baum S. F., Bowman J. L. (2003). Radial patterning of Arabidopsis shoots by class III HD-ZIP and KANADI genes. Curr. Biol. 13, 1768–1774 10.1016/j.cub.2003.09.035 [DOI] [PubMed] [Google Scholar]
- Eshed Y., Baum S. F., Perea J. V., Bowman J. L. (2001). Establishment of polarity in lateral organs of plants. Curr. Biol. 11, 1251–1260 10.1016/S0960-9822(01)00392-X [DOI] [PubMed] [Google Scholar]
- Eshed Y., Izhaki A., Baum S. F., Floyd S. K., Bowman J. L. (2004). Asymmetric leaf development and blade expansion in Arabidopsis are mediated by KANADI and YABBY activities. Development 131, 2997–3006 10.1242/dev.01186 [DOI] [PubMed] [Google Scholar]
- Faircloth L. M., Churchill P. F., Caldwell G. A., Caldwell K. A. (2009). The microtubule-associated protein, NUD-1, exhibits chaperone activity in vitro. Cell Stress Chaperones 14, 95–103 10.1007/s12192-008-0061-1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guo M., Thomas J., Collins G., Timmermans M. C. (2008). Direct repression of KNOX loci by the ASYMMETRIC LEAVES1 complex of Arabidopsis. Plant Cell 20, 48–58 10.1105/tpc.107.056127 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hake S., Smith H. M., Holtan H., Magnani E., Mele G., Ramirez J. (2004). The role of knox genes in plant development. Annu. Rev. Cell Dev. Biol. 20, 125–151 10.1146/annurev.cellbio.20.031803.093824 [DOI] [PubMed] [Google Scholar]
- Hunter C., Willmann M. R., Wu G., Yoshikawa M., de la Luz Gutiérrez-Nava M., Poethig S. R. (2006). Trans-acting siRNA-mediated repression of ETTIN and ARF4 regulates heteroblasty in Arabidopsis. Development 133, 2973–2981 10.1242/dev.02491 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ikezaki M., Kojima M., Sakakibara H., Kojima S., Ueno Y., Machida C., Machida Y. (2010). Genetic networks regulated by ASYMMETRIC LEAVES1 (AS1) and AS2 in leaf development in Arabidopsis thaliana: KNOX genes control five morphological events. Plant J. 61, 70–82 10.1111/j.1365-313X.2009.04033.x [DOI] [PubMed] [Google Scholar]
- Inagaki S., Nakamura K., Morikami A. (2009). A link among DNA replication, recombination, and gene expression revealed by genetic and genomic analysis of TEBICHI gene of Arabidopsis thaliana. PLoS Genetics 5, e1000613 10.1371/journal.pgen.1000613 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Iwakawa H., Ueno Y., Semiarti E., Onouchi H., Kojima S., Tsukaya H., Hasebe M., Soma T., Ikezaki M., Machida C. et al. (2002). The ASYMMETRIC LEAVES2 gene of Arabidopsis thaliana, required for formation of a symmetric flat leaf lamina, encodes a member of a novel family of proteins characterized by cysteine repeats and a leucine zipper. Plant Cell Physiol. 43, 467–478 10.1093/pcp/pcf077 [DOI] [PubMed] [Google Scholar]
- Iwakawa H., Iwasaki M., Kojima S., Ueno Y., Soma T., Tanaka H., Semiarti E., Machida Y., Machida C. (2007). Expression of the ASYMMETRIC LEAVES2 gene in the adaxial domain of Arabidopsis leaves represses cell proliferation in this domain and is critical for the development of properly expanded leaves. Plant J. 51, 173–184 10.1111/j.1365-313X.2007.03132.x [DOI] [PubMed] [Google Scholar]
- Jurkuta R. J., Kaplinsky N. J., Spindel J. E., Barton M. K. (2009). Partitioning the apical domain of the Arabidopsis embryo requires the BOBBER1 NudC domain protein. Plant Cell 21, 1957–1971 10.1105/tpc.108.065284 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kerstetter R. A., Bollman K., Taylor R. A., Bomblies K., Poethig R. S. (2001). KANADI regulates organ polarity in Arabidopsis. Nature 411, 706–709 10.1038/35079629 [DOI] [PubMed] [Google Scholar]
- Kojima S., Iwasaki M., Takahashi H., Imai T., Matsumura Y., Fleury D., Van Lijsebettens M., Machida Y., Machida C. (2011). ASYMMETRIC LEAVES2 and Elongator, a histone acetyltransferase complex, mediate the establishment of polarity in leaves of Arabidopsis thaliana. Plant Cell Physiol. 52, 1259–1273 10.1093/pcp/pcr083 [DOI] [PubMed] [Google Scholar]
- Kumaran M. K., Bowman J. L., Sundaresan V. (2002). YABBY polarity genes mediate the repression of KNOX homeobox genes in Arabidopsis. Plant Cell 14, 2761–2770 10.1105/tpc.004911 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Makita T., Katsuyama Y., Tani S., Suzuki H., Kato N., Todd R. B., Hynes M. J., Tsukagoshi N., Kato M., Kobayashi T. (2009). Inducer-dependent nuclear localization of a Zn(II)2Cys6 transcriptional activator, AmyR, in Aspergillus nidulans. Biosci. Biotechnol. Biochem. 73, 391–399 10.1271/bbb.80654 [DOI] [PubMed] [Google Scholar]
- Mallory A. C., Reinhart B. J., Jones-Rhoades M. W., Tang G., Zamore P. D., Barton M. K., Bartel D. P. (2004). MicroRNA control of PHABULOSA in leaf development: importance of pairing to the microRNA 5′ region. EMBO J. 23, 3356–3364 10.1038/sj.emboj.7600340 [DOI] [PMC free article] [PubMed] [Google Scholar]
- McConnell J. R., Barton M. K. (1998). Leaf polarity and meristem formation in Arabidopsis. Development 125, 2935–2942 [DOI] [PubMed] [Google Scholar]
- McConnell J. R., Emery J., Eshed Y., Bao N., Bowman J., Barton M. K. (2001). Role of PHABULOSA and PHAVOLUTA in determining radial patterning in shoots. Nature 411, 709–713 10.1038/35079635 [DOI] [PubMed] [Google Scholar]
- Miller B. A., Zhang M. Y., Gocke C. D., De Souza C., Osmani A. H., Lynch C., Davies J., Bell L., Osmani S. A. (1999). A homolog of the fungal nuclear migration gene nudC is involved in normal and malignant human hematopoiesis. Exp. Hematol. 27, 742–750 10.1016/S0301-472X(98)00074-5 [DOI] [PubMed] [Google Scholar]
- Montgomery T. A., Howell M. D., Cuperus J. T., Li D., Hansen J. E., Alexander A. L., Chapman E. J., Fahlgren N., Allen E., Carrington J. C. (2008). Specificity of ARGONAUTE7-miR390 interaction and dual functionality in TAS3 trans-acting siRNA formation. Cell 133, 128–141 10.1016/j.cell.2008.02.033 [DOI] [PubMed] [Google Scholar]
- Moon J., Hake S. (2010). How a leaf gets its shape. Curr. Opin. Plant Biol. 14, 1–7 10.1016/j.pbi.2010.08.012 [DOI] [PubMed] [Google Scholar]
- Morris S. M., Anaya P., Xiang X., Morris N. R., May G. S., Yu-Lee L. (1997). A prolactin-inducible T cell gene product is structurally similar to the Aspergillus nidulans nuclear movement protein NUDC. Mol. Endocrinol. 11, 229–236 10.1210/me.11.2.229 [DOI] [PubMed] [Google Scholar]
- Nakagawa T., Suzuki T., Murata S., Nakamura S., Hino T., Maeo K., Tabata R., Kawai T., Tanaka K., Niwa Y. et al. (2007). Improved gateway binary vectors: high-performance vectors for creation of fusion constructs in transgenic analysis of plants. Biosci. Biotechnol. Biochem. 71, 2095–2100 10.1271/bbb.70216 [DOI] [PubMed] [Google Scholar]
- Nishihama R., Soyano T., Ishikawa M., Araki S., Tanaka H., Asada T., Irie K., Ito M., Terada M., Banno H. et al. (2002). Expansion of the cell plate in plant cytokinesis requires a kinesin-like protein/MAPKKK complex. Cell 109, 87–99 10.1016/S0092-8674(02)00691-8 [DOI] [PubMed] [Google Scholar]
- Nishino M., Kurosawa Y., Evans R., Lin S. H., Brinkley B. R., Yu-Lee L. Y. (2006). NudC Is Required for Plk1 Targeting to the Kinetochore and Chromosome Congression. Curr. Biol. 16, 1414–1421 10.1016/j.cub.2006.05.052 [DOI] [PubMed] [Google Scholar]
- Ori N., Eshed Y., Chuck G., Bowman J. L., Hake S. (2000). Mechanisms that control knox gene expression in the Arabidopsis shoot. Development 127, 5523–5532 [DOI] [PubMed] [Google Scholar]
- Osmani A. H., Osmani S. A., Morris N. R. (1990). The molecular cloning and identification of a gene product specifically required for nuclear movement in Aspergillus nidulans. J. Cell Biol. 111, 543–551 10.1083/jcb.111.2.543 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pekker I., Alvarez J. P., Eshed Y. (2005). Auxin response factors mediate Arabidopsis organ asymmetry via modulation of KANADI activity. Plant Cell 17, 2899–2910 10.1105/tpc.105.034876 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Perez D. E., Hoyer J. S., Johnson A. I., Moody Z. R., Lopez J., Kaplinsky N. J. (2009). BOBBER1 is a noncanonical Arabidopsis small heat shock protein required for both development and thermotolerance. Plant Physiol. 151, 241–252 10.1104/pp.109.142125 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Qi Y., Sun Y., Xu L., Xu Y., Huang H. (2004). ERECTA is required for protection against heat-stress in the AS1/AS2 pathway to regulate adaxial-abaxial leaf polarity in Arabidopsis. Planta 219, 270–276 10.1007/s00425-004-1248-z [DOI] [PubMed] [Google Scholar]
- Rédei G. P., Hirono Y. (1964). Linkage studies. Arabidopsis Inf. Serv. 1, 9 [Google Scholar]
- Rowlands D. J., Turner G. (1973). Nuclear and extranuclear inheritance of oligomycin resistance in Aspergillus nidulans. Mol. Gen. Genet. 126, 201–206 10.1007/BF00267531 [DOI] [PubMed] [Google Scholar]
- Sarojam R., Sappl P. G., Goldshmidt A., Efroni I., Floyd S. K., Eshed Y., Bowman J. L. (2010). Differentiating Arabidopsis shoots from leaves by combined YABBY activities. Plant Cell 22, 2113–2130 10.1105/tpc.110.075853 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sawa S., Watanabe K., Goto K., Kanaya E., Morita E. H., Okada K. (1999). FILAMENTOUS FLOWER, a meristem and organ identity gene of Arabidopsis, encodes a protein with a zinc finger and HMG-related domains. Genes Dev. 13, 1079–1088 10.1101/gad.13.9.1079 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schwab R., Maizel A., Ruiz-Ferrer V., Garcia D., Bayer M., Crespi M., Voinnet O., Martienssen R. A. (2009). Endogenous tasiRNAs mediate non-cell autonomous effects on gene regulation in Arabidopsis thaliana. PLoS ONE 4, e5980 10.1371/journal.pone.0005980 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Semiarti E., Ueno Y., Tsukaya H., Iwakawa H., Machida C., Machida Y. (2001). The ASYMMETRIC LEAVES2 gene of Arabidopsis thaliana regulates formation of a symmetric lamina, establishment of venation and repression of meristem-related homeobox genes in leaves. Development 128, 1771–1783 [DOI] [PubMed] [Google Scholar]
- Shuai B., Reynaga-Peña C. G., Springer P. S. (2002). The LATERAL ORGAN BOUNDARIES gene defines a novel, plant-specific gene family. Plant Physiol. 129, 747–761 10.1104/pp.010926 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Siegfried K. R., Eshed Y., Baum S. F., Otsuga D., Drews G. N., Bowman J. L. (1999). Members of the YABBY gene family specify abaxial cell fate in Arabidopsis. Development 126, 4117–4128 [DOI] [PubMed] [Google Scholar]
- Steeves T. A., Sussex I. M. (1989). Patterns in Plant Development, 2nd edn Cambridge: Cambridge University Press [Google Scholar]
- Sun Y., Zhou Q., Zhang W., Fu Y., Huang H. (2002). ASYMMETRIC LEAVES1, an Arabidopsis gene that is involved in the control of cell differentiation in leaves. Planta 214, 694–702 10.1007/s004250100673 [DOI] [PubMed] [Google Scholar]
- Suzuki T., Nakajima S., Inagaki S., Hirano-Nakakita M., Matsuoka K., Demura T., Fukuda H., Morikami A., Nakamura K. (2005). TONSOKU is expressed in S phase of the cell cycle and its defect delays cell cycle progression in Arabidopsis. Plant Cell Physiol. 46, 736–742 10.1093/pcp/pci082 [DOI] [PubMed] [Google Scholar]
- Szakonyi D., Moschopoulos A., Byrne M. E. (2010). Perspectives on leaf dorsoventral polarity. J. Plant Res. 123, 281–290 10.1007/s10265-010-0336-3 [DOI] [PubMed] [Google Scholar]
- Tsukaya H. (2006). Mechanism of leaf-shape determination. Annu. Rev. Plant Biol. 57, 477–496 10.1146/annurev.arplant.57.032905.105320 [DOI] [PubMed] [Google Scholar]
- Tsukaya H., Uchimiya H. (1997). Genetic analyses of the formation of the serrated margin of leaf blades in Arabidopsis: combination of a mutational analysis of leaf morphogenesis with the characterization of a specific marker gene expressed in hydathodes and stipules. Mol. Gen. Genet. 256, 231–238 10.1007/s004380050565 [DOI] [PubMed] [Google Scholar]
- Ueno Y., Ishikawa T., Watanabe K., Terakura S., Iwakawa H., Okada K., Machida C., Machida Y. (2007). Histone deacetylases and ASYMMETRIC LEAVES2 are involved in the establishment of polarity in leaves of Arabidopsis. Plant Cell 19, 445–457 10.1105/tpc.106.042325 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Watanabe K., Okada K. (2003). Two discrete cis elements control the abaxial side-specific expression of the FILAMENTOUS FLOWER gene in Arabidopsis. Plant Cell 15, 2592–2602 10.1105/tpc.015214 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Waites R., Hudson A. (1995). Phantastica: a gene required for dorsoventrality in leaves of Antirrhinum majus. Development 121, 2143–2154 [Google Scholar]
- Wu G., Lin W. C., Huang T., Poethig R. S., Springer P. S., Kerstetter R. A. (2008). KANADI1 regulates adaxial-abaxial polarity in Arabidopsis by directly repressing the transcription of ASYMMETRIC LEAVES2. Proc. Natl. Acad. Sci. USA 105, 16392–16397 10.1073/pnas.0803997105 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu L., Xu Y., Dong A., Sun Y., Pi L., Xu Y., Huang H. (2003). Novel as1 and as2 defects in leaf adaxial-abaxial polarity reveal the requirement for ASYMMETRIC LEAVES1 and 2 and ERECTA functions in specifying leaf adaxial identity. Development 130, 4097–4107 10.1242/dev.00622 [DOI] [PubMed] [Google Scholar]
- Yang J. Y., Iwasaki M., Machida C., Machida Y., Zhou X., Chua N. H. (2008). ßC1, the pathogenicity factor of TYLCCNV, interacts with AS1 to alter leaf development and suppress selective jasmonic acid response. Genes Dev. 22, 2564–2577 10.1101/gad.1682208 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang M. Y., Huang N. N., Clawson G. A., Osmani S. A., Pan W., Xin P., Razzaque M. S., Miller B. A. (2002). Involvement of the fungal nuclear migration gene nudC human homolog in cell proliferation and mitotic spindle formation. Exp. Cell Res. 273, 73–84 10.1006/excr.2001.5414 [DOI] [PubMed] [Google Scholar]
- Zhong R., Ye Z. H. (2004). Amphivasal vascular bundle 1, a gain-of-function mutation of the IFL1/REV gene, is associated with alterations in the polarity of leaves, stems and carpels. Plant Cell Physiol. 45, 369–385 10.1093/pcp/pch051 [DOI] [PubMed] [Google Scholar]
- Zhou T., Aumais J. P., Liu X., Yu-Lee L. Y., Erikson R. L. (2003). A role for Plk1 phosphorylation of NudC in cytokinesis. Dev. Cell 5, 127–138 10.1016/S1534-5807(03)00186-2 [DOI] [PubMed] [Google Scholar]
- Zhou T., Zimmerman W., Liu X., Erikson R. L. (2006). A mammalian NudC-like protein essential for dynein stability and cell viability. Proc. Natl. Acad. Sci. USA 103, 9039–9044 10.1073/pnas.0602916103 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhu X. J., Liu X., Jin Q., Cai Y., Yang Y., Zhou T. (2010). The L279P mutation of nuclear distribution gene C (NudC) influences its chaperone activity and lissencephaly protein 1 (LIS1) stability. J. Biol. Chem. 285, 29903–29910 10.1074/jbc.M110.105494 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.