Abstract
The planar cell polarity (PCP) signaling pathway governs collective cell movements duringvertebrate embryogenesis, and certain PCP proteins are also implicated in the assembly ofcilia. The septins are cytoskeletal proteins controlling behaviors such as cell division and migration. Here, we identified control of septin localization by the PCP protein Fritz as a crucial control point for both collective cell movement and ciliogenesis in Xenopus embryos. We also linked mutations in human Fritz to Bardet-Biedl and Meckel-Gruber syndromes, a notable link given that other genes mutated in these syndromes also influence collective cell movement and ciliogenesis. These findings shed light on the mechanisms by which fundamental cellular machinery, such as the cytoskeleton, is regulated during embryonic development and human disease.
A major unresolved question in embryonic tissue morphogenesis concerns how developmental signals are translated into changes in cell behavior (1). As cells become specified, different cell types initiate specific, yet varied, behaviors. For example, cells in the notochord and spinal cord of vertebrates interdigitate in a planar polarized manner forming a longer, narrower array (2, 3). This collective cell movement, convergent extension (CE), drives gastrulation and elongates the embryonic axis (Fig. 1A) (4–7). By contrast, multiciliated epithelial cells undergo a radical process of cellular morphogenesis, developing dozens of apical motile cilia (8, 9). Proteins in the Wnt/PCP pathway act in both cell types (6, 7, 10–18), but how they regulate such dissimilar cellular behaviors remains unknown.
Fig. 1.

Fritz and Septins control convergent extension. (A) Cell intercalation (black, white, and gray cells) drives CE, elongating the body axis and closing the blastopore (bp) during gastrulation.(B) Control embryo with closed blastopore (whitecircle). (C) Sibling Fritz morphants fail to close the blastopore (red).(D) Coinjection of GFP-Fritz mRNA rescued blastopore closure (blue). (E) Control embryo. (F) Sibling sept7 morphants fail to close the blastopore (red). (G) Sept7 mRNA rescues closure of blastopore (blue). (H) Quantification of blastopore closure (mean ± SEM; n values for each column, left to right, are as follows: 21, 20, 28, 18, 11, 8, 16, 12, and 18). (I) Still frame from movie of control Keller explant; green arrows indicate mediolaterally aligned cells, alignment of all cells is quantified in the inset diagram. (J) Xdd1, a PCP-specific dominant-negative Dishevelled, disrupts elongation and alignment (red arrows indicate misoriented cells). (K) Fritz morphant cells display defective elongation, but normal polarized alignment. (L) Sept2 morphant cells display defective elongation, but normal polarized alignment. (M) Control cells (movie 1); orange/yellow lines indicate positions of kymographs in m′ and m″. (N) Fritz morphant Keller explant; red arrows indicateintercellular spaces (movie 2). (O) Sept2 morphant Keller explant; red arrows indicate intercellular spaces (movie 3).
We investigated the vertebrate ortholog of the cytoplasmic WD40 repeat protein Fritz, which controls PCP in Drosophila (19). We first assessed the requirement for Fritz in Xenopus gastrulation, where PCP-mediated CE in the dorsal mesoderm drives the reorganization of the germ layers and closure of the blastopore. (Fig. 1A) (2, 4, 6, 20). Fritz was strongly expressed in these cells, and morpholino (MO)–mediated knockdown elicited defective gastrulation indicated by a persistent open blastopore and failure of axis elongation (Fig. 1, B to D, and H; and fig. S1, A and E to I).
We then explored the cell behavioral basis of these defects using explanted gastrula tissue in organotypic culture (“Keller” explants) (fig. S2) (2, 10). During normal CE, cells polarize and align their long axes mediolaterally, acquiring elongate morphologies as they interdigitate (Fig. 1, A and I, and fig. S2). Core PCP proteins govern both polarized alignment and cell elongation (Fig. 1J, and fig. S3)(10, 12). By contrast, Fritz morphant cells displayed significantly reduced cell elongation, whereas their polarized alignment was largely normal (Fig. 1K, and fig. S3). Moreover, in time-lapse movies, cell membranes in controls were relatively static, whereas Fritz morphant cell membranes undulated, frequently producing intercellular spaces that opened and closed rapidly between cells (Fig. 1, M and N, and movies S1 and S2). Thus, Fritz controls cell shape but not polarization during CE and thus executes a specific subset of PCP-mediated cell behaviors in Xenopus, consistent with data from Drosophila(19).
The instability of plasma membranes in Fritz morphants suggested a possible role for septins, as these cytoskeletal elements can brace plasma membranes and resist aberrant cell shape deformation (21–23). Sept2 and sept7 were expressed strongly in cells during CE, and MO-mediated knockdown of either resulted in defective gastrulation (Fig. 1, E to H, and fig. S4). A cell-permeable inhibitor of septin dynamics, forchlorfenuron (FCF) (24, 25), also elicited gastrulation defects (Fig. 2, A to C and G). Moreover, cells in septin morphants displayed reduced cell elongation but normal polarization, similar to Fritz morphants (Fig. 1L and fig. S3). Finally, time-lapse imaging revealed membrane undulations in septin morphants similar to those in Fritz morphants (Fig. 1O, movies S3 and S4, and fig. S5).
Fig. 2.
Functional interaction between Fritz and septins. (A) Control embryo withclosed blastopore. (B) Sibling embryo treated with 100 μM FCF. (C) Sibling treated with 200μM FCF. (D) Sibling injected with a low dose of Fritz MO. (E) Sibling Fritz morphant treated with 100μM FCF. (F) Sibling Fritz morphant treated with 200 μM FCF. (G) Quantification of blastopore closure (mean ± SEM; n = nine embryos per column). (H) Sept2-GFP concentrated at the cortex (red in h′) of cells engaged in CE. (I) Cortical sept2-GFP concentration was reduced in Fritz morphants. (J) Sept2-GFP localization was quantified as the ratio of cortical versus cytoplasmic pixel intensities. (K)Cortical/cytoplasmic Sept2-GFP ratio was reduced significantly in Fritz morphants; the ratio of coexpressed membrane-RFP is unchanged (mean ± SEM; n = 123 cells for control, 150 for morphant).
These similar phenotypes led us to test for functional interactions between Fritz and septins. First, application of FCF exacerbated significantly the gastrulation defects in low-dose Fritz morphants (Fig. 2, D to G). Second, sept2 was associated consistently with Fritz in coimmuno-precipitations from Xenopus embryos (fig. S6). Finally, septins act at the cortex in cultured cells (22, 23), and both sept2 fused to green fluorescent protein (sept2-GFP) and Fritz-GFP were concentrated at the cell cortex during CE (Fig. 2H and fig. S1J). Moreover, the ratio of cortical-to-cytoplasmic sept2-GFP was consistently reduced in Fritz morphants compared with controls (Fig. 2, I to K). These data suggest that Fritz and septins collaborate during PCP-mediated collective cell movements.
In addition to their role in CE, vertebrate orthologs of certain Drosophila PCP proteins control ciliogenesis (13–18). At organogenesis stages, Fritz was expressed in ciliated cell types, and Fritz morphants displayed defects in craniofacial morphogenesis and Hedgehog signaling similar to those associated with defective ciliogenesis (Fig. 3, A to F, and fig. S1, J to L) (13). Exploiting the multiciliated cells of the Xenopus epidermis (Fig. 3, G and H)(9, 13, 15, 16), we found that those lacking Fritz (Fritz knockdown) resulted in the development of fewer and shorter cilia (Fig. 3, G and H).
Fig. 3.
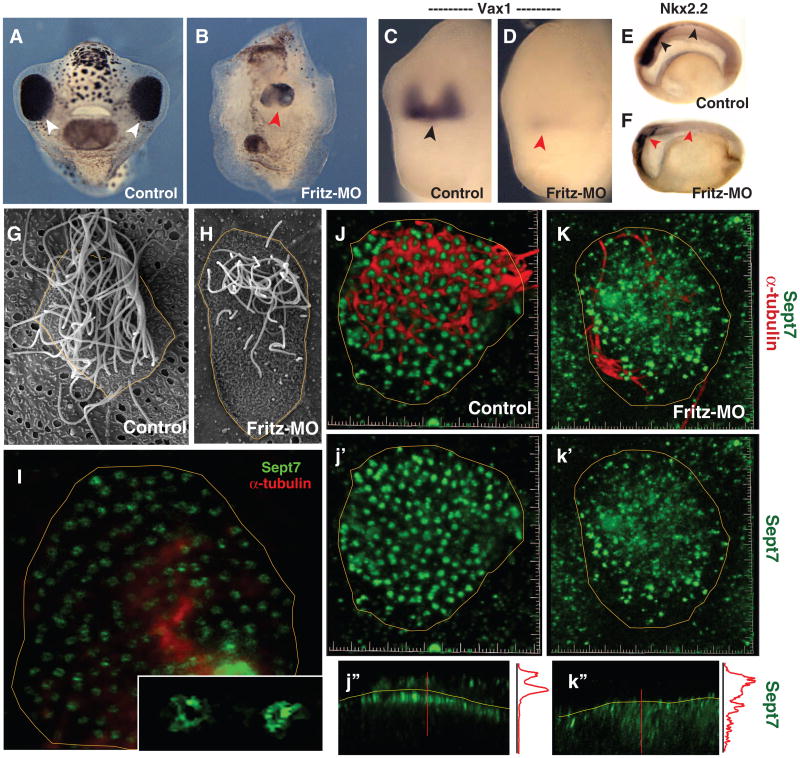
Fritz controls septin localization, ciliogenesis, and Hedgehog signaling.(A) Control tadpole (anterior view; arrowheads indicate eyes).(B) Sibling Fritz morphant (red arrowhead indicates single, medial eye). (C) Vax1 expression in control embryo.(D) Vax1 expression is lost in a Fritz morphant. (E) Sagittal view of Nkx2.2 expression in the spinal cord of a control embryo, black arrowheads. (F) Nkx2.2 expression is lost in Fritz morphant, red arrowheads. (G) A single multiciliated cell from the Xenopus epidermis. (H) Multiciliated cells in Fritz morphants display fewer and shorter cilia. (I) Sept7 in ring-like structures (inset) at the base of cilia in a confocal slice of a multiciliated cell. (J) Sept7 structures (green; j′) are highly ordered in a stack from a control multiciliated cell: cilia are visible in red. (K) Sept7 structures (green; k′) are disorganized in a Fritz morphant multiciliated cell: few cilia are visible in red. (j″) Z-projection reveals tight association of Sept7 with the apical surface (yellow line); intensity plot is shown at right (red line). (k″) Z-projection reveals loss of sept7 from the apical surface.
No common mechanism as yet explains the role of PCP proteins in ciliogenesis and collective cell movement, but because Fritz functions with septins during CE, we tested the possibility that these proteins also act together during ciliogenesis. Indeed, sept7 and Fritz were localized along ciliary axonemes and in distinct foci near basal bodies (Fig. 3, I and J, and fig. S1, J and K). These sept7 foci were most prominent, and frequently appeared ringlike, in single optical sections of the apical surface of a multiciliated cell viewed en face (Fig. 3I, inset). These sept7 structures were consistent with the septin-based diffusion barrier at the base of cilia (26, 27) and also consistent in size with septin-based rings that self-assemble in vitro (28).
In Fritz morphants, disrupted ciliogenesis was associated with defects in the size and shape of individual sept7 structures, as well as their positioning at the apical surface (Fig. 3, J and K). The mean area of Sept7 structures in Fritz morphant multiciliated cells was reduced significantly compared with controls (fig. S7A), and individual sept7 structures displayed heterogeneous pixel intensities in Fritz morphants, which suggested a defect in their structure (fig. S7B). Finally, both sept7 and sept2 were restricted to the apical surface and axonemes of control multiciliated cells but were detected consistently throughout the cytoplasm of Fritz morphant cells (Fig. 3, j″ and k″, and fig. S8, A to D). Consistent with a link between Fritz, septins, and ciliogenesis, knockdown of Sept7 or Sept2 elicited defects in ciliogenesis (Fig. 4, A and B, and fig. S8, E and F). Finally, septin knockdown also disrupted expression of Hedgehog target genes in the midline, althoughNotch targets were unaffected (Fig. 4, C and D, and figs. S8, G and H, and S9).
Fig. 4.

Septins govern ciliogenesis and Hedgehog signaling in developing embryos.(A)Control Xenopus multiciliated cells (B) Cilia are shorter and fewer in number sept7 morphants (C) Sagittal view of the Hedgehog target gene Nkx2.2 in the spinal cord of a control embryo marked by black arrowheads. (D) Nkx2.2 expression is lost in sept2 and sept7 morphant embryos; red arrowheads. (E) Table of mutations in human Fritz (C2ORF86) in BBS and MKS patients. In only two cases (*, **) were mutations present in trans with mutations in a known BBS- or MKS-associated gene.
Fritz thus acts together with septins to control both CE and ciliogenesis in Xenopus. Human genes associated with Meckel-Gruber (MKS) and Bardet-Biedl (BBS) syndromes likewise affect both processes (29–32), so we asked whether mutationsin Fritz might contribute to these disorders. We found a significant enrichment of nonsynonymous coding changes in human Fritz (C2orf86) in MKS and BBS patients as compared with controls (6 alleles in 192 patients vs. 0 in 384 controls; P < 0.015; Fig. 4E and fig. S10A). In the MKS cohort, we did not identify alleles sufficient to explain the phenotype, which suggested that these changes might interact in trans with primary MKS loci [e.g., (30, 33)].For the BBS cohort, we found two heterozygous missense alleles that were absent from 384 ethnically matched controls, HapMap, and 1000 genomes. Also, two of these changes map to the same surface-exposed face of the predicted β-propeller structure of the Fritzprotein (fig. S10, B and C).
Notably, we also found a homozygous Fritz mutation that segregated with the disorder in a BBS family (Fig. 4E). Both parents and an unaffected sib are heterozygous carriers (fig. S10D), and this position is invariant in Fritz, absent from 384 controls, HapMap, and 1000 genomes. Although we are cautious in interpreting data from a single family, this result suggests that Fritz loss-of-function mutations may be sufficient to cause BBS, and corroborates other studies linking BBS proteins to ciliogenesis and CE (29–32).
Here, we link Fritz to the septin cytoskeleton in both ciliogenesis and collective cell movement to provide insights into both PCP signaling and the function and regulation of septins during vertebrate embryogenesis. An important challenge will be to understand the broader protein network underlying Fritz-dependent septin functions in vivo. Intriguing candidates are the previously identified BBS and MKS proteins and also IFT88, as these proteins are implicated in both ciliogenesis and CE (29, 34).
Acknowledgments
We thank P. Abitua for technical assistance; E. Spiliotis for the antibodies; and A. Ewald, B. Mitchell, E. Brooks, and K. Smith for critical discussions and reading. This work was supported by grants from the Uehara Memorial Foundation (A.S.); National Institute of General Medcial Science (NIH), March of Dimes, Burroughs Wellcome Fund, the Sandler Program for Asthma Research, and the Texas Advanced Research Program (J.B.W.); National Institute of Child Health and Human Development and National Institute of Digestive Diseases and Kidney Disease (NIH) (N.K.). N.K. is the George W. Brumley Professor. J.B.W. is an Early Career Scientist of the Howard Hughes Medical Institute.
Footnotes
Supporting Online Material:www.sciencemag.org/cgi/content/full/science.1191184/DC1, Materials and Methods, Figs S1 to S8, References, Movies S1 to S4
References and Notes
- 1.Trinkaus JP. Cell Into Organs—The Forces That Shape the Embryo. Prentice-Hall; Englewood Cliffs, NJ: 1969. [Google Scholar]
- 2.Shih J, Keller R. Development. 1992;116:901. doi: 10.1242/dev.116.4.901. [DOI] [PubMed] [Google Scholar]
- 3.Keller R, Shih J, Sater A. Dev Dyn. 1992;193:199. doi: 10.1002/aja.1001930302. [DOI] [PubMed] [Google Scholar]
- 4.Keller R. Science. 2002;298:1950. doi: 10.1126/science.1079478. [DOI] [PubMed] [Google Scholar]
- 5.Wallingford JB, Harland RM. Development. 2002;129:5815. doi: 10.1242/dev.00123. [DOI] [PubMed] [Google Scholar]
- 6.Ewald AJ, Peyrot SM, Tyszka JM, Fraser SE, Wallingford JB. Development. 2004;131:6195. doi: 10.1242/dev.01542. [DOI] [PubMed] [Google Scholar]
- 7.Wang J, et al. Development. 2006;133:1767. doi: 10.1242/dev.02347. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Vladar EK, Stearns T. J Cell Biol. 2007;178:31. doi: 10.1083/jcb.200703064. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Hayes JM, et al. Dev Biol. 2007;312:115. doi: 10.1016/j.ydbio.2007.09.01. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Wallingford JB, et al. Nature. 2000;405:81. doi: 10.1038/35011077. [DOI] [PubMed] [Google Scholar]
- 11.Heisenberg CP, et al. Nature. 2000;405:76. doi: 10.1038/35011068. [DOI] [PubMed] [Google Scholar]
- 12.Jessen JR, et al. Nat Cell Biol. 2002;4:610. doi: 10.1038/ncb828. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Park TJ, Haigo SL, Wallingford JB. Nat Genet. 2006;38:303. doi: 10.1038/ng1753. [DOI] [PubMed] [Google Scholar]
- 14.Oishi I, Kawakami Y, Raya Á, Callol-Massot C, Izpisúa Belmonte JC. Nat Genet. 2006;38:1316. doi: 10.1038/ng1892. [DOI] [PubMed] [Google Scholar]
- 15.Park TJ, Mitchell BJ, Abitua PB, Kintner C, Wallingford JB. Nat Genet. 2008;40:871. doi: 10.1038/ng.104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Gray RS, et al. Nat Cell Biol. 2009;11:1225. doi: 10.1038/ncb1966. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Zeng H, Hoover AN, Liu A. Dev Biol. 2010;339:418. doi: 10.1016/j.ydbio.2010.01.003. [DOI] [PubMed] [Google Scholar]
- 18.Tissir F, et al. Nat Neurosci. 2010;13:700. doi: 10.1038/nn.2555. [DOI] [PubMed] [Google Scholar]
- 19.Collier S, Lee H, Burgess R, Adler P. Genetics. 2005;169:2035. doi: 10.1534/genetics.104.033381. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Materials and methods are available in the supporting material on Science Online.
- 21.Weirich CS, Erzberger JP, Barral Y. Nat Rev Mol Cell Biol. 2008;9:478. doi: 10.1038/nrm2407. [DOI] [PubMed] [Google Scholar]
- 22.Tooley AJ, et al. Nat Cell Biol. 2009;11:17. doi: 10.1038/ncb1808. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Tanaka-Takiguchi Y, Kinoshita M, Takiguchi K. Curr Biol. 2009;19:140. doi: 10.1016/j.cub.2008.12.030. [DOI] [PubMed] [Google Scholar]
- 24.Hu Q, Nelson WJ, Spiliotis ET. J Biol Chem. 2008;283:29563. doi: 10.1074/jbc.M804962200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Iwase M, Okada S, Oguchi T, Toh-e A. Genes Genet Syst. 2004;79:199. doi: 10.1266/ggs.79.199. [DOI] [PubMed] [Google Scholar]
- 26.Hu Q, et al. Science. 2010;329:436. doi: 10.1126/science.1191054. published online 17 June 2010, 10.1126/science.1191054. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Caudron F, Barral Y. Dev Cell. 2009;16:493. doi: 10.1016/j.devcel.2009.04.003. [DOI] [PubMed] [Google Scholar]
- 28.Kinoshita M, Field CM, Coughlin ML, Straight AF, Mitchison TJ. Dev Cell. 2002;3:791. doi: 10.1016/s1534-5807(02)00366-0. [DOI] [PubMed] [Google Scholar]
- 29.Ross AJ, et al. Nat Genet. 2005;37:1135. doi: 10.1038/ng1644. [DOI] [PubMed] [Google Scholar]
- 30.Leitch CC, et al. Nat Genet. 2008;40:443. doi: 10.1038/ng.97. [DOI] [PubMed] [Google Scholar]
- 31.Gerdes JM, et al. Nat Genet. 2007;39:1350. doi: 10.1038/ng.2007.12. [DOI] [PubMed] [Google Scholar]
- 32.Smith UM, et al. Nat Genet. 2006;38:191. doi: 10.1038/ng1713. [DOI] [PubMed] [Google Scholar]
- 33.Khanna H, et al. Nat Genet. 2009;41:739. doi: 10.1038/ng.366. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Jones C, et al. Nat Genet. 2008;40:69. doi: 10.1038/ng.2007.54. [DOI] [PubMed] [Google Scholar]