Abstract
Background
The rat lungworm Angiostrongylus cantonensis can cause eosinophilic meningoencephalitis in humans. This nematode’s main definitive hosts are rodents and its intermediate hosts are snails. This parasite was first described in China and currently is dispersed across several Pacific islands, Asia, Australia, Africa, some Caribbean islands and most recently in the Americas. Here, we report the genetic variability among A. cantonensis isolates from different geographical locations in Brazil using mitochondrial cytochrome c oxidase subunit I (COI) gene sequences.
Methods
The isolates of A. cantonensis were obtained from distinct geographical locations of Brazil. Genomic DNAs were extracted, amplified by polymerase reaction, purified and sequenced. A partial sequence of COI gene was determined to assess their phylogenetic relationship.
Results
The sequences of A. cantonensis were monophyletic. We identified a distinct clade that included all isolates of A. cantonensis from Brazil and Asia based on eight distinct haplotypes (ac1, ac2, ac3, ac4, ac5, ac6, ac7 and ac8) from a previous study. Interestingly, the Brazilian haplotype ac5 is clustered with isolates from Japan, and the Brazilian haplotype ac8 from Rio de Janeiro, São Paulo, Pará and Pernambuco states formed a distinct clade. There is a divergent Brazilian haplotype, which we named ac9, closely related to Chinese haplotype ac6 and Japanese haplotype ac7.
Conclusion
The genetic variation observed among Brazilian isolates supports the hypothesis that the appearance of A. cantonensis in Brazil is likely a result of multiple introductions of parasite-carrying rats, transported on ships due to active commerce with Africa and Asia during the European colonization period. The rapid spread of the intermediate host, Achatina fulica, also seems to have contributed to the dispersion of this parasite and the infection of the definitive host in different Brazilian regions.
Keywords: Rattus norvegicus, Achatina fulica, Eosinophilic meningoencephalitis, Molecular phylogeny, Cytochrome c oxidase subunit I, Brazil
Background
Angiostrongylus cantonensis (Chen, 1935) is a nematode that lives in the right ventricle and pulmonary arteries of rats. Rodents such as Rattus rattus and Rattus norvegicus are considered the most important definitive hosts [1]. This nematode is the most common cause of eosinophilic meningoencephalitis in humans [2,3]. The rat lungworm was first described in China infecting these same rodents. Currently, the nematode is dispersed across several Pacific islands, Asia, Australia, Africa, some Caribbean islands and most recently in the Americas [4-6]. The transmission of this nematode has been linked to dispersal of invasive organisms [7]. In particular, the introduction of Achatina fulica in Brazil and Pomacea canaliculata (Lamarck, 1822) in China are examples of the importance of exotic snails in the spread of this helminthiasis [5,8,9]. Similarly, the nematode may have A. fulica as one of its main intermediate hosts in Brazil [10,11]. Currently, this mollusk is experiencing the explosive phase of invasion, since it has been found in 25 of the 26 Brazilian states and in the Federal District [12]. Humans are accidentally infected by eating raw and undercooked snails that contain the third stage larvae (L3) [13]. The infection can also occur by eating animals that act as paratenic hosts, such as shrimps, crabs, lizards, frogs and terrestrial planarians or through vegetables contaminated by the mucus of infected snails [4]. More recently, the presence of A. fulica naturally infected by L3 larvae of A. cantonensis in different states of Brazil, such as Espírito Santo, São Paulo, Pernambuco and Santa Catarina, was confirmed by experimental infection in R. norvegicus[14]. Cases of human eosinophilic meningoencephalitis have also been reported in the states of Espírito Santo, Pernambuco and São Paulo [5,15,16].
The morphological heterogeneity, such as bursal rays characteristics among Brazilian isolates of A. cantonensis, was reported by Maldonado et al. (2010) [14], who have suggested this occurred as a result of distinct entry of the parasite into the country. Recent studies using sequencing of the mitochondrial protein-coding gene cytochrome c oxidase subunit I (COI) to distinguish A. cantonensis isolates confirmed the presence of three geographical isolates in Asia [17]. Subsequently, Simões et al. (2011) [18], using COI data, observed that A. cantonensis from Rio de Janeiro, Brazil yield a single haplotype, which formed a clade with low genetic distance to the Chinese isolates. Interestingly, Tokiwa et al. (2012) [7], analyzing a great number of geographical isolates from the Asian continent, proposed that Rio de Janeiro isolates are more similar to A. cantonensis isolated from Japan.
In the present study, we analyzed A. cantonensis worms from Brazil using the COI gene to assess the genetic variability of different geographical isolates as well as to determine the phylogenetic relationship among the Brazilian isolates.
Methods
Geographical isolates
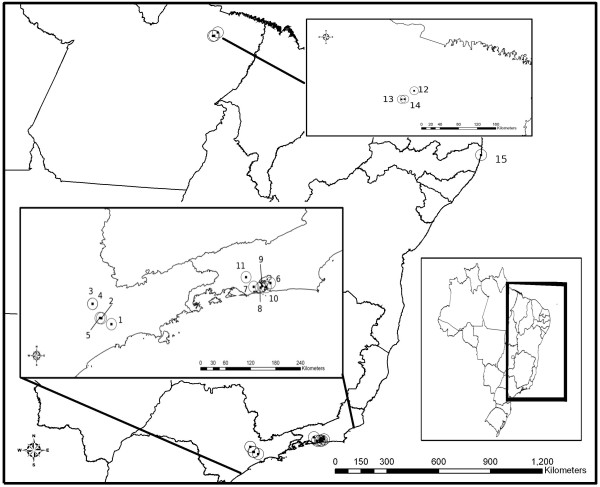
The isolates of A. cantonensis were obtained from distinct geographical locations in Brazil (Figure 1), after experimental infection of R. norvegicus with L3 larvae recovered from A. fulica naturally infected obtained from the National Reference Laboratory of Medical Malacology, Oswaldo Cruz Foundation, or through naturally infected R. norvegicus and R. rattus (Table 1). Collection permits for rodents were obtained from the animal use ethics committee of Oswaldo Cruz Foundation (FIOCRUZ) (CEUA no. LW 24/10).
Figure 1.
Map of Brazilian locations where A. cantonensis isolates were collected. (1) São Mateus; (2) Freguesia do Ó; (3) Jundiaí; (4) Vila Rami; (5) Pirituba; (6) São Gonçalo; (7) Encantado; (8) Túnel Noel Rosa; (9) Caju; (10) Niterói; (11) Queimados; (12) Marituba; (13) Jurunas; (14) Guamá; (15) Olinda.
Table 1.
Angiostrongylus cantonensis samples isolated from São Paulo (SP), Rio de Janeiro (RJ), Pará (PA) and Pernambuco (PE) used in the present study
| Geographic locations | Worm stage | Origins | Date of collection | Geographic coordinates | CHIOC | GenBank accession | Haplotype |
|---|---|---|---|---|---|---|---|
| São Mateus – SP |
Adult |
Achatina fulica |
2011 |
23°35′40.18”S, 46°28′28.39”W |
35788 |
JX471064 |
ac8 |
| Freguesia do Ó – SP |
Adult |
Achatina fulica |
2010 |
23°29′00.77”S, 46°41′22.91”W |
35832 |
JX471058 |
ac8 |
| Jundiaí – SP |
Adult |
Achatina fulica |
2011 |
23°12′54.44”′S, 46°52′49.69”W |
35789 |
JX471062 |
ac8 |
| Vila Rami – SP |
Adult |
Achatina fulica |
2011 |
23°12′52.43”S, 46°52′59.14”W |
35834 |
JX471067 |
ac8 |
| Pirituba – SP |
Adult |
Achatina fulica |
2010 |
23°28′44.40”S, 46°43′23.24”W |
35724 |
JX471054 |
ac8 |
| São Gonçalo – RJ |
Adult |
Rattus norvegicus |
2011 |
22°49′30”S, 43°02′30”W |
35703 |
JX471066 |
ac8 |
| Encantado – RJ |
Adult |
Achatina fulica |
2010 |
22°53′45”S, 43°18′08”W |
35723 |
JX471057 |
ac8 |
| Túnel Noel Rosa – RJ |
Adult |
Achatina fulica |
2010 |
22°54′35.87”S, 43°15′12.97”W |
35721 |
JX471068 |
ac8 |
| Caju – RJ |
Adult |
Rattus rattus |
2011 |
22°52′58.13”S, 43°13′07.35”W |
35831 |
JX471055 |
ac9 |
| Niterói – RJ |
Adult |
Achatina fulica |
2010 |
22°53′55.11”S, 43°07′53.97”W |
35829 |
JX471059 |
ac5 |
| Queimados – RJ |
Adult |
Achatina fulica |
2010 |
22°42′55.20”S, 43°34′06”W |
35725 |
JX471060 |
ac5 |
| Marituba – PA |
Adult |
Rattus rattus |
2010 |
01°36′60.7”S, 48°34′49.4”W |
35722 |
JX471065 |
ac8 |
| Jurunas – PA |
Adult |
Rattus rattus |
2010 |
01°47′34.3”S, 48°49′24.8”W |
35833 |
JX471063 |
ac8 |
| Guamá – PA |
Adult |
Rattus rattus |
2010 |
01°47′24.2”S, 48°45′59.15”W |
35830 |
JX471061 |
ac8 |
| Olinda – PE | Adult | Achatina fulica | 2009 | 08°00′32.25”S, 34°51′07.95”W | 35661 | JX471056 | ac8 |
Experimental infection
The snails were individually minced and digested in a 0.7% HCl solution for 6 h. The digested samples were then placed in a Baermann apparatus and allowed to sediment overnight. The L3 nematode larvae obtained from digested snails were administered orally to 3-month-old R. norvegicus (Wistar strain) rats (100 L3/animal). Thirty-five days after administration of the larvae, the rodents were euthanized using a CO2 chamber and adult worms were collected from the pulmonary arteries, washed in physiologic solution and fixed in 70% ethanol or frozen for molecular analysis. The specimens from each isolate were cleared and mounted as temporary slides in lactophenol solution and examined under a light microscope. Taxonomic identification of the nematodes was based on morphological parameters obtained from previous studies [12,14].
Molecular and phylogenetic analysis
Genomic DNA samples were extracted using the Qiagen QIAamp DNA Mini Kit, according to the manufacturer’s protocol. The extracted DNA was stored at 4°C until use. The DNA amplification by polymerase reaction was conducted using the previously described primers COI_F 5’ TTTTTTGGGCATCCTGAGGTTTAT 3’ and COI_R 5’ TAAAGAAAGAACATAATGAAAATG 3’ for a partial region of the COI gene [19,20]. The reaction mixture was prepared in a total volume of 50 μL containing 16.2 μL of water, 5 μL of 10 x PCR buffer (Tris-HCl, KCl), 2.5 μL of MgCl2 (2.5 mM), 5 μL of dNTP mix (10 mM each), 10 μL of each primer (0.2 mM), 0.3 μL of Taq DNA polymerase (1.5U) and 1 μL of sample DNA. The thermocycler was programmed to incubate the samples for 5 min at 94°C, followed by 40 cycles at 94°C for 30s, 55°C for 30s, 72°C for 1 min and final extension at 72°C for 5 min. The reaction products were separated by electrophoresis on 1.0% agarose gel, stained with ethidium bromide and visualized under ultraviolet light. Amplified products were purified using the QIAquick PCR Purification Kit (Qiagen). Sequencing reactions were performed using an ABI PrismDyeTerminator Cycle Sequencing Core Kit (Applied Biosystems, USA) as described by Genomic Platform-DNA Sequencing (PDTIS/FIOCRUZ). A partial sequence of the COI gene was determined to assess their phylogenetic relationship. All sequences determined in this study have been deposited in the GenBank database: six sequences of A. cantonensis from the state of Rio de Janeiro; five sequences from the state of São Paulo; three sequences from the state of Pará and one sequence from the state of Pernambuco (Table 1).
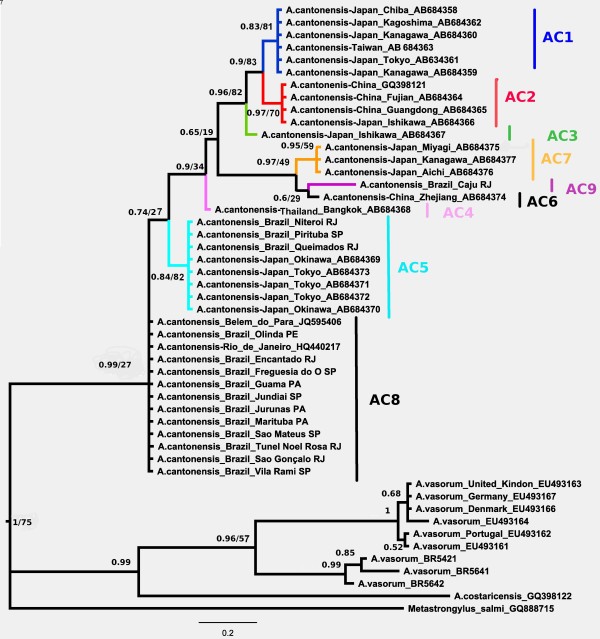
Alignment and editing of sequences were performed using Clustal W in MEGA version 5 [21,22]. The nucleotide variation and p-distance were calculated using the resultant alignment in the MEGA version 5 software. The aligned sequences were subjected to neighbor-joining (NJ) analysis, also performed using the MEGA version 5 software, and Bayesian inference (BI), which was performed using the MrBayes 3.2.0 program [23]. The evolutionary model applied to BI was chosen using the Bayesian information criterion (BIC), Akaike information criterion (AIC) and corrected Akaike information criterion (AICc), by which BIC and AICc indicated choice of the same evolutionary model, and were calculated on topologies optimized by maximum likelihood as implemented by the MrAIC program [24-26]. NJ bootstrap values were estimated using 1000 replicates with TrN93+G distances and the BI was performed with the HKY+G model of sequence evolution. Different evolutionary models were used in both trees since the HKY+G model considers the frequency of transitional changes between purines and pyrimidines as equal. As there is no analytical form to estimate HKY+G distances, we used TrN93 model [27]. The posterior probabilities (BPP) were estimated using Markov chain Monte Carlo (MCMC) analysis, which was run for 10,000,000 generations with data sampling every 500 generations, discarding the first 1000 sampled trees as burn-in. A BLAST search (http://blast.ncbi.nlm.nih.gov/Blast.cgi) was performed to clarify any similarities with the sequences obtained and previously published sequences. COI sequences from Angiostrongylus spp. were obtained from the GenBank as follows: A. cantonensis from Japan, China, Taiwan, Thailand and Brazil; A. vasorum from the UK and A. costaricensis. Sequences of Metastrongylus salmi were used as outgroup. Haplotypes for A. vasorum isolates from Brazil (A. vasorum 5421, 5641, and 5642) were reconstructed from published information [20] (Figure 2).
Figure 2.
Bayesian tree with GenBank accession numbers of Angiostrongylus spp. using 360 bp of mitochondrial COI gene. Values of Bayesian posterior probabilities (left) and neighbor-joining bootstrap values (right) are represented at the nodes.
Results
The COI partial sequences of A. cantonensis were determined from 15 geographic isolates from Brazil and presented a length varying from 440 to 460 bp. We aligned 360 bp of COI gene to compare it with other sequences previously available in the GenBank database. The sequences analyzed revealed that in 338 bp (93.9%) the positions of the nucleotides were monomorphic or invariable, while 22 sites (6.1%) were variable, of which four sites were parsimony-informative, and the amino acid sequences did not reveal any variability in 120 encoded amino acids, generating three different haplotypes of COI (ac5, ac8 and ac9) based on eight distinct haplotypes, as mencioned by Tokiwa et al. (2012) [7].
The evolutionary parameters kappa, pi(A), pi(C), pi(G), pi(T) and alpha were estimated through MCMC sampling with lower and upper limits and presented the following values of average and variance respectively: kappa 7.7 / 6.0; pi(A) 0.20 / 0.0002; pi(C) 0.05 / 0.00008; pi(G) 0.30 / 0.0004; pi(T) 0.45 / 0.0006 and alpha 0.15 / 0.0002.
The phylogenetic trees inferred using the two methods showed a similar topology but with some minor differences in six nodes involving A. vasorum and A. cantonensis from Túnel Noel Rosa (Rio de Janeiro). The Bayesian tree, presented with a condensed root and posterior probabilities at nodes (BPP) and bootstrap values for NJ, can be found in Figure 2 and showed higher nodal support valued than the NJ tree rooted on M. salmi based on TrN93 distances (Additional file 1: Figure S2). The bootstrap values of the branches with different topologies mentioned above, were not included in Bayesian tree. The Angiostrongylus species were grouped into two major clades. The sequences of A. cantonensis were monophyletic. The clade corresponding to A. cantonensis was supported with low bootstrap values of 27% for NJ and high posterior probability values of 0.99 for BI. Within A. cantonensis, the geographical isolates corresponding to Brazilian haplotypes ac5 and ac8 were the first to branch, with low bootstrap values of 27% for NJ and posterior probability values of 0.74 for BI. The Brazilian haplotype ac9 was clustered in a distinct clade with Chinese haplotype ac6 and Japanese haplotype ac7, presenting bootstrap values of 49% for NJ and high posterior probability values of 0.97 for BI.
Only one A. cantonensis isolate, Caju from the state of Rio de Janeiro, named haplotype ac9, formed a clade with haplotype ac6 from China (Zhejiang) and with haplotype ac7 from Japan (Miyagi, Kanagawa and Aichi). The isolates from Queimados and Niterói (state of Rio de Janeiro) and Pirituba (state of São Paulo) constituted a clade with isolates from Japan (Okinawa and Tokyo), corresponding to haplotype ac5. The haplotype ac8 corresponded to the other Brazilian isolates from the Southeast region: Jundiaí, Freguesia do Ó, São Mateus, Vila Rami (state of São Paulo), Túnel Noel Rosa, Encantado, São Gonçalo and Rio de Janeiro city (state of Rio de Janeiro) [GenBank: HQ440217]; North region: Guamá, Jurunas, Marituba and Belém (state of Pará) [GenBank: JQ595406]; and Northeast region: Olinda (state of Pernambuco), forming a distinct clade.
The closely related species had interspecific p-distance values, ranging from 12.2% (between A. cantonensis and A. vasorum) to 19.0% (between A. cantonensis and A. costaricensis). Intraspecific distance values among A. cantonensis ranged from 0.8% (between haplotypes ac1 and ac2) to 6.4% (between haplotypes ac5 and ac9) (Table 2).
Table 2.
p-distance values of haplotypes of Angiostrongylus cantonensis, Angiostrongylus costaricensis and Angiostrongylus vasorum based on mitochondrial COI gene
| ac1 | ac2 | ac3 | ac4 | ac5 | ac6 | ac7 | ac8 | ac9 | ACO | AV42 | AV21 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
A. cantonensis ac1 |
- |
|
|
|
|
|
|
|
|
|
|
|
|
A. cantonensis ac2 |
0. 008 |
- |
|
|
|
|
|
|
|
|
|
|
|
A. cantonensis ac3 |
0.011 |
0.011 |
- |
|
|
|
|
|
|
|
|
|
|
A. cantonensis ac4 |
0.020 |
0.020 |
0.014 |
- |
|
|
|
|
|
|
|
|
|
A. cantonensis ac5 |
0.038 |
0.038 |
0.032 |
0.017 |
- |
|
|
|
|
|
|
|
|
A. cantonensis ac6 |
0.051 |
0.051 |
0.044 |
0.035 |
0.054 |
- |
|
|
|
|
|
|
|
A. cantonensis ac7 |
0.051 |
0.051 |
0.044 |
0.035 |
0.048 |
0.011 |
- |
|
|
|
|
|
|
A. cantonensis ac8 |
0.038 |
0.038 |
0.032 |
0.017 |
0.011 |
0.054 |
0.048 |
- |
|
|
|
|
|
A. cantonensis ac9 |
0.054 |
0.054 |
0.048 |
0.044 |
0.064 |
0.020 |
0.026 |
0.057 |
- |
|
|
|
|
A. costaricensis GQ398122 |
0.176 |
0.189 |
0.190 |
0.190 |
0.190 |
0.185 |
0.176 |
0.172 |
0.167 |
- |
|
|
|
A. vasorum BR5642 |
0.155 |
0.151 |
0.147 |
0.134 |
0.130 |
0.142 |
0.146 |
0.126 |
0.130 |
0.169 |
- |
|
|
A. vasorum BR5421 |
0.156 |
0.152 |
0.147 |
0.135 |
0.143 |
0.142 |
0.146 |
0.127 |
0.131 |
0.169 |
0.008 |
- |
| A. vasorum BR5641 | 0.146 | 0.142 | 0.138 | 0.130 | 0.138 | 0.142 | 0.145 | 0.122 | 0.126 | 0.178 | 0.023 | 0.017 |
The nucleotide variation between the clade that includes haplotypes ac5 and ac8 consisted of four mutational steps, while the mutational steps between clades containing haplotype ac9 were 21 and 19 from haplotype ac5 and ac8, respectively. In addition, the mutational steps between haplotype ac9 and haplotypes ac6 and ac7 were seven and nine, respectively (Table 3 and Figure 3).
Table 3.
Variable nucleotide positions within the mitochondrial COI gene from different haplotypes of Angiostrongylus cantonensis
| |
Nucleotide position |
|||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 022 | 025 | 031 | 034 | 073 | 076 | 100 | 109 | 118 | 130 | 136 | 160 | 169 | 181 | 187 | 193 | 211 | 229 | 232 | 235 | 244 | 268 | 280 | 289 | 292 | 295 | 298 | 301 | 304 | 310 | 313 | 358 | |
| Halplotypes | ||||||||||||||||||||||||||||||||
| ac1 |
A |
G |
A |
A |
A |
A |
T |
T |
T |
G |
G |
G |
A |
T |
G |
G |
G |
A |
G |
A |
A |
T |
A |
T |
T |
G |
A |
C |
G |
A |
G |
A |
| ac2 |
. |
. |
. |
. |
. |
. |
. |
. |
. |
. |
A |
. |
. |
. |
. |
. |
. |
. |
. |
. |
. |
. |
. |
C |
. |
. |
. |
G |
. |
. |
. |
. |
| ac3 |
. |
. |
. |
G |
. |
. |
. |
. |
. |
. |
A |
. |
. |
. |
. |
. |
. |
. |
. |
. |
. |
C |
. |
. |
. |
. |
. |
T |
. |
. |
. |
. |
| ac4 |
. |
. |
G |
G |
. |
G |
. |
. |
. |
. |
A |
. |
. |
. |
. |
. |
A |
. |
. |
. |
. |
. |
. |
. |
G |
. |
. |
T |
. |
. |
. |
. |
| ac5 |
. |
. |
G |
G |
. |
G |
. |
C |
C |
. |
A |
. |
. |
. |
. |
. |
A |
G |
. |
. |
. |
. |
G |
. |
G |
. |
G |
T |
A |
. |
. |
. |
| ac6 |
G |
T |
G |
G |
G |
. |
A |
. |
. |
. |
A |
. |
G |
C |
. |
A |
A |
. |
A |
G |
. |
. |
. |
. |
G |
A |
. |
T |
. |
. |
. |
G |
| ac7 |
G |
T |
G |
G |
G |
. |
A |
. |
. |
A |
A |
. |
G |
. |
. |
A |
A |
. |
A |
. |
. |
. |
G |
. |
G |
A |
. |
T |
. |
. |
. |
G |
| ac8 |
. |
. |
G |
G |
. |
G |
. |
. |
C |
. |
A |
A |
. |
. |
. |
. |
A |
G |
. |
. |
. |
. |
G |
. |
G |
. |
G |
T |
. |
G |
. |
. |
| ac9 | G | T | G | G | G | . | A | . | . | . | A | . | G | . | T | A | A | . | A | G | G | . | . | . | . | . | . | T | . | G | A | G |
Figure 3.
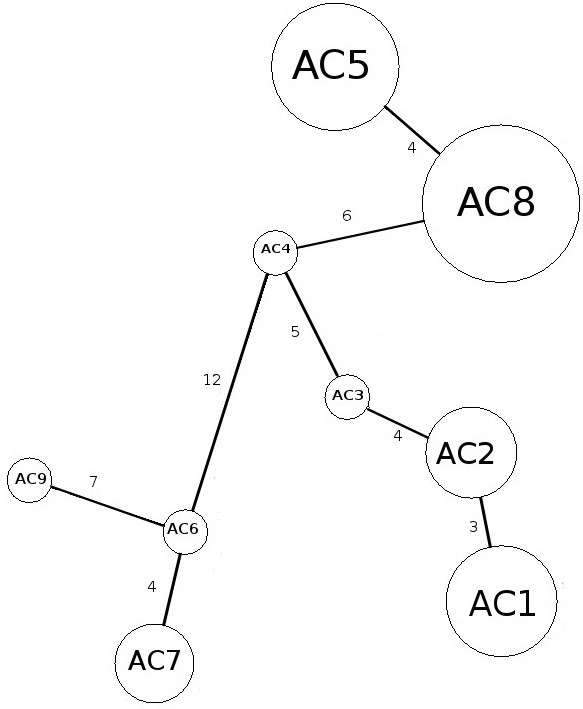
Haplotype network in the form of a minimum spanning tree based on partial COI gene sequence.
Discussion
In the present study, Brazilian isolates of A. cantonensis were analyzed using mitochondrial COI gene sequences. This allowed evaluation of variability in A. cantonensis isolates from different geographical locations in Brazil. All sequences from Brazil were monophyletic with sequences from Asia. Tokiwa et al. (2012) [7], distinguished eight different haplotypes, named ac1 to ac8. Most sequences from Brazilian samples were either ac5 or ac8. Moreover, we described a new haplotype named ac9, monophyletic with Chinese haplotype ac6.
The intraspecific variation observed among the Brazilian isolates ranged from 0.8% to 6.4%. These values are in agreement with the findings of Blouin (2002) [28], which showed that the level of mtDNA sequence variation among nematode individuals of the same species is lower than 10%.
The data observed in this study showed that the A. cantonensis isolate from Caju (state of Rio de Janeiro) is restricted to the port area and could have entered the country through trade from Asia. The factor that might have prevented dispersal of haplotype ac9 to other places in the country is the absence of the main intermediate host, A. fulica, at the site where the rats were trapped.
Similarly, the Brazilian isolates from Pirituba (state of São Paulo), Queimados and Niterói (state of Rio de Janeiro), which correspond to haplotype ac5 from Japan, are believed to have entered through Rio de Janeiro or São Paulo also from the Asian continent. This hypothesis is also considered for the most abundant Brazilian haplotype (ac8), showing the possible spread from the arrival area to the Southeast, Northeast and North regions, probably through the giant African snail, A. fulica.
Likewise, Araujo (1967) [29] showed in a study on helminth fauna in Rattus norvegicus in the city of São Paulo that all rats captured were parasitized by 1 to 11 species of helminths. Interestingly, the helminth fauna lacked species of the genus Angiostrongylus. Moreover, Pessôa and Martins (1982) [30] reported that J.E. Alicata did not find A. cantonensis infection in rodents collected in the Brazilian state of Bahia, suggesting the recent introduction of the parasite in the country.
A. fulica has been considered a snail pest in tropical and subtropical regions where it has been introduced. In Brazil, this exotic snail was introduced in the state of Paraná in the 1980s, probably brought from Indonesia for commercial purposes that were not successful (escargot farming). The high reproductive capacity and the tendency for people to release snails into the wild are the probable reasons for the rapid invasion of this species [8,11]. This snail is currently found in most Brazilian states. Factors such as its voracious feeding habits contribute to the extermination of the native snail fauna, reducing the available resources and increasing competition for physical space. The absence of natural pathogens also contributes to the high dispersion of these snails [31].
The increased presence of A. cantonensis in the country is likely a result of the rapid spread of its intermediate host, A. fulica, contributing to the dispersion of this parasite and infection of the definitive host [12]. This phenomenon is described as one of the primary causes of the spread of eosinophilic meningoencephalitis [14].
The genetic variation observed among Brazilian isolates supports the hypothesis that the appearance of A. cantonensis in Brazil is a result of multiple introductions of parasite-carrying rats and the snails that act as intermediate hosts. These were likely transported on ships due to trade with Africa and Asia during the period of European colonization [8,14] and dispersed via human transport, becoming endemic in port areas [7]. At the present moment a phylogeographic study of A. cantonensis is essential to locate the geographical origin of these introductions, especially of haplotypes ac8 and ac9.
Conclusions
In summary, we studied the molecular variation of A. cantonensis isolates from different geographical locations in Brazil based on COI DNA sequences. This study showed that four Brazilian isolates are clustered with isolates from Japan, China and Thailand (haplotypes ac5 and ac9), and 11 Brazilian isolates form a distinct clade (haplotype ac8). In addition, haplotype ac9 represent a new A. cantonensis haplotype. The COI gene appears as a good marker for differentiating geographical isolates of A. cantonensis. The phylogenetic features of this nematode help to understand how phylogeography can influence the transmission dynamics of this parasite.
Abbreviations
COI: Cytochrome c oxidase subunit I; L3: Third stage larvae; PDTIS: Genomic Platform-DNA Sequencing; NJ: Neighbor-joining; BI: Bayesian inference; BIC: Bayesian information criterion; AIC: Akaike information criterion; AICc: Corrected Akaike information criterion; TrN+G: Tamura-Nei model with Gamma distributed; HKY+G: Hasegawa-Kishino-Yano model with Gamma distributed; BPP: Posterior probabilities; MCMC: Markov chain Monte Carlo.
Competing interests
The authors declare that they have no competing interests.
Authors’ contributions
TCCM participated in the study design, carried out laboratory experiments, participated in data analysis and participated in drafting the manuscript. ROS participated in the study design and participated in drafting the manuscript. APMO participated in the laboratory experiments. SCT participated in the study design and drafting the manuscript. CFN participated in the field work. AJS participated in the study design and drafting the manuscript. PCE participated in the study design, participated in data analysis and participated in drafting the manuscript. AMJ participated in the study design, carried out laboratory experiments, participated in data analysis and participated in drafting the manuscript. All authors read and approved the final version of the manuscript.
Authors’ information
Arnaldo Maldonado Júnior has a fellowship from the National Council for Scientific and Technological Development (CNPq).
Supplementary Material
Figure S2. Neighbor-joining tree using 360 bp of mitochondrial COI gene.
Contributor Information
Tainá CC Monte, Email: taina.monte@ioc.fiocruz.br.
Raquel O Simões, Email: raquel83@ioc.fiocruz.br.
Ana Paula M Oliveira, Email: apmartin@ioc.fiocruz.br.
Clodoaldo F Novaes, Email: cfnovaes@gmail.com.
Silvana C Thiengo, Email: sthiengo@ioc.fiocruz.br.
Alexandre J Silva, Email: abs8@cdc.gov.
Pedro C Estrela, Email: estrela@ioc.fiocruz.br.
Arnaldo Maldonado Júnior, Email: maldonad@ioc.fiocruz.br.
Acknowledgements
We are grateful to Oswaldo Cruz Institute/FIOCRUZ, CAPES (Program of Basic Parasitology – grant 3031/2011) and CNPq, which supported this work and also thank the Genomic Platform-DNA Sequencing (PDTIS/FIOCRUZ). PCE was funded by CNPq-PDJ-FIOCRUZ grant 500091/2012-2. We also thank Franklyn Enrique Samudio Acosta for valuable suggestions and Nilson J Melo, Gerônimo N Costa and Jorge M Silva for technical support.
References
- Lunn JA, Lee R, Smaller J, Mackay BM, King T, Hunt GB, Martin P, Krockenberger MB, Spielman D, Malik R. Twenty two cases of canine neural angiostrongylosis in eastern Australia (2002-2005) and a review of the literature. Parasit Vectors. 2012;70:1–18. doi: 10.1186/1756-3305-5-70. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Graeff-Teixeira C, Silva ACA, Yoshimura K. Update on eosinophilic meningoencephalitis and its clinical relevance. Clin Microbiol Rev. 2009;22:322–348. doi: 10.1128/CMR.00044-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen CY, Kuo CC, Lo CP, Huang MY, Wang YM, Wang WY. Eosinophilic meningoencephalitis caused by Angiostrongylus cantonensis. QJM. 2011. [DOI] [PubMed]
- Foronda P, Lopez-Gonzales M, Miquel J, Torres J, Segovia M, Abreu-Costa N, Casanova JC, Valladares B, Mas-Coma S, Bargues MD, Feliu C. Finding of Parastrongylus cantonensis (chen,1935) in Rattus rattus in tenerife, Canary Islands (Spain) Acta Trop. 2010;114:123–127. doi: 10.1016/j.actatropica.2010.02.004. [DOI] [PubMed] [Google Scholar]
- Maldonado AJ, Simões R, Thiengo S. In: Zoonosis. Volume 1. 1. Morales-Lorenzo J, editor. InTech; 2012. Angiostrongyliasis in the Americas; pp. 303–320. [Google Scholar]
- Morassutti AL, Graeff-Teixeira C. Interface molecules of Angiostrongylus cantonensis: their role in parasite survival and modulation of host defenses. Int J Inflam. 2012;2012:1–6. doi: 10.1155/2012/512097. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tokiwa T, Harunari T, Tanikawa T, Komatsu N, Koizumi N, Tung K-C, Suzuki J, Kadosaka T, Takada N, Kumagai T, Akao N, Ohta N. Phylogenetic relationships of rat lungworm, Angiostrongylus cantonensis, isolated from different geographical regions revealed widespread multiple lineages. Parasitol Int. 2012;61:431–436. doi: 10.1016/j.parint.2012.02.005. [DOI] [PubMed] [Google Scholar]
- Thiengo S, Faraco F, Salgado N, Cowie R, Fernandez M. Rapid spread of an invasive snail in Soth America: the giant African snail, Achatina fulica, in Brasil. Biol Invasions. 2007;9:693–702. doi: 10.1007/s10530-006-9069-6. [DOI] [Google Scholar]
- Lv S, Zhang Y, Steinmann P, Yang G-J, Yank K, Zhou X-N, Utzinger J. The emergence of angiostrongyliasis in the People’s Republic of China: the interplay between invasive snails, climate change and transmission dynamics. Freshwater Biol. 2011;56:717–734. doi: 10.1111/j.1365-2427.2011.02579.x. [DOI] [Google Scholar]
- Graeff-Teixeira C. Expansion of Achatina fulica in Brazil and potential increased risk for angiostrongyliasis. Trans R Soc Trop Med Hyg. 2007;101:743–744. doi: 10.1016/j.trstmh.2007.03.012. [DOI] [PubMed] [Google Scholar]
- Zanol J, Fernandez MA, Oliveira APM, Russo CAM, Thiengo SC. O caramujo exótico invasor Achatina fulica (Stylommatophora, Mollusca) no Estado do Rio de Janeiro (Brasil): situação atual. Biota Neotrop. 2010;10:447–451. doi: 10.1590/S1676-06032010000300038. [DOI] [Google Scholar]
- Thiengo SC, Maldonado A, Mota EM, Torres EJ, Caldeira R, Carvalho OS, Oliveira AP, Simões RO, Fernandez MA, Lanfredi RM. The giant African snail Achatina fulica as natural intermediate host of Angiostrongylus cantonensis in Pernambuco, northeast Brazil. Acta Trop. 2010;115:194–199. doi: 10.1016/j.actatropica.2010.01.005. [DOI] [PubMed] [Google Scholar]
- Martin-Alonso A, Foronda P, Quispe-Ricalde MA, Feliu C, Valladares B. Seroprevalence of Angiostrongylus cantonensis in Wild Rodents from the Canary Islands. PLoS One. 2011;6:1–5. doi: 10.1371/journal.pone.0027747. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maldonado AJ, Simões RO, Oliveira APM, Motta EM, Fernandez MA, Pereira ZM, Monteiro SS, Torres EJL, Thiengo SC. First report of Angiostrongylus cantonensis (Nematoda: Metastrongylidae) in Achatina fulica (Mollusca: Gastropoda) from Southeast and South Brazil. Mem Inst Oswaldo Cruz. 2010;105:938–941. doi: 10.1590/s0074-02762010000700019. [DOI] [PubMed] [Google Scholar]
- Caldeira RL, Mendonca CLF, Gouveia CO, Lenzi HL, Graeff-Teixeira C, Lima WS. First record of molluscs naturally infected with Angiostrongylus cantonensis (Chen, 1935) (Nematoda: Metastrongylidae) in Brazil. Mem Inst Oswaldo Cruz. 2007;102:887–889. doi: 10.1590/S0074-02762007000700018. [DOI] [PubMed] [Google Scholar]
- Lima AR, Mesquita SD, Santos SS, Aquino ER, Rosa LRS, Duarte FS. Alicata disease: neuroinfestation by Angiostrongylus cantonensis in Recife, Pernambuco, Brazil. Arq Neuropsiquiatr. 2009;67:1093–1096. doi: 10.1590/S0004-282X2009000600025. [DOI] [PubMed] [Google Scholar]
- Eamsobhana P, Lim PE, Solano G, Zhang H, Gan X, Yong HS. Molecular differentiation of Angiostrongylus taxa (Nematoda: Angiostrongylidae) by cytochrome c oxidase subunit I (COI) gene sequences. Acta Trop. 2010;116:152–156. doi: 10.1016/j.actatropica.2010.07.005. [DOI] [PubMed] [Google Scholar]
- Simões RO, Monteiro FA, Sánchez E, Thiengo SC, Garcia JS, Costa-Neto SF, Luque JL, Maldonado A. Endemic Angiostrongyliasis, Rio de Janeiro, Brazil. Emerg Infect Dis. 2011;17:1331–1333. doi: 10.3201/eid1707.101822. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bowles J, Blair D, McManus DP. Genetic variants within the genus Echinococcus identified by mitochondrial DNA sequencing. Mol Biochem Parasitol. 1992;54:165–173. doi: 10.1016/0166-6851(92)90109-W. [DOI] [PubMed] [Google Scholar]
- Jefferies R, Shaw SE, Viney ME, Morgan ER. Angiostrongylus vasorum from South America and Europe represent distinct lineages. Parasitology. 2009;136:107–115. doi: 10.1017/S0031182008005258. [DOI] [PubMed] [Google Scholar]
- Thompson J, Higgins D, Gibson T. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994;22:4673–4680. doi: 10.1093/nar/22.22.4673. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S. MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance and maximum parsimony methods. Mol Biol Evol. 2011;28:2731–2739. doi: 10.1093/molbev/msr121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ronquist F, Teslenko M, van der Mark P, Ayres DL, Darling A, Höhna S, Larget B, Liu L, Suchard MA, Huelsenbeck JP. MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol. 2012;61:539–542. doi: 10.1093/sysbio/sys029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Akaike H. A new look at the statistical model identification. IEEE Trans Autom Control. 1974;19:716–723. doi: 10.1109/TAC.1974.1100705. [DOI] [Google Scholar]
- Burnham KP, Anderson DR. Multimodel inference: understanding AIC and BIC in model selection. Sociol Method Res. 2004;33:261–304. doi: 10.1177/0049124104268644. [DOI] [Google Scholar]
- Nylander JAA. MrAIC.pl. Program distributed by the author. Uppsala: Evolutionary Biology Centre, Uppsala University; 2004. [Google Scholar]
- Tamura K, Nei M. Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Mol Biol Evol. 1993;10:512–526. doi: 10.1093/oxfordjournals.molbev.a040023. [DOI] [PubMed] [Google Scholar]
- Blouin MS. Molecular prospecting for cryptic species of nematodes: mitochondrial DNA versus internal transcribed spacer. Int J Parasitol. 2002;32:527–531. doi: 10.1016/S0020-7519(01)00357-5. [DOI] [PubMed] [Google Scholar]
- Araujo P. Helmintos de Rattus norvegicus (Berkenhout,1769) da cidade de São Paulo. Rev Fac Farm Bioquím S. Paulo. 1967;5:141–159. [Google Scholar]
- Pessôa SB, Martins AV. Pessôa Parasitologia Médica, tenth ed. Rio de Janeiro, Rio de Janeiro: Guanabara Koogan; 1982. [Google Scholar]
- Ohlweiler FP, Guimarães MCA, Takahashi FY, Eduardo JM. Current distribution of Achatina fulica Bowdich, 1822 in the State of São Paulo including records of Aelurostrongylus abstrusus (Nematoda) larvae infestation. Rev Inst Med Trop S. Paulo. 2010;52:211–214. doi: 10.1590/S0036-46652010000400009. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Figure S2. Neighbor-joining tree using 360 bp of mitochondrial COI gene.