Abstract
The ability to perceive noxious stimuli is critical for an animal's survival in the face of environmental danger, and thus pain perception is likely to be under stringent evolutionary pressure. Using a neuronal-specific RNAi knock-down strategy in adult Drosophila, we recently completed a genome-wide functional annotation of heat nociception that allowed us to identify α2δ3 as a novel pain gene. Here we report construction of an evolutionary-conserved, system-level, global molecular pain network map. Our systems map is markedly enriched for multiple genes associated with human pain and predicts a plethora of novel candidate pain pathways. One central node of this pain network is phospholipid signaling, which has been implicated before in pain processing. To further investigate the role of phospholipid signaling in mammalian heat pain perception, we analysed the phenotype of PIP5Kα and PI3Kγ mutant mice. Intriguingly, both of these mice exhibit pronounced hypersensitivity to noxious heat and capsaicin-induced pain, which directly mapped through PI3Kγ kinase-dead knock-in mice to PI3Kγ lipid kinase activity. Using single primary sensory neuron recording, PI3Kγ function was mechanistically linked to a negative regulation of TRPV1 channel transduction. Our data provide a systems map for heat nociception and reinforces the extraordinary conservation of molecular mechanisms of nociception across different species.
Author Summary
Nociception is the perception of noxious, potentially damaging stimuli; and this pain or its equivalent behavioral readout is evolutionarily conserved from fruit flies to humans. Using genetic techniques in the fruit fly, we have been able to evaluate the potential functional contribution of every gene in the fruit fly genome for a role in avoidance of high noxious temperatures (heat pain-like responses). Using this functional genomics data set, we have developed a conserved network map of heat pain/nociception that predicts numerous conserved genes and pathways as novel pain pathways, including phospholipid signaling. Studies in multiple mutant mice confirmed a role for lipid signaling in pain perception, and more specifically we identify the critical lipid kinase (PI3Kγ) as a negative regulator of TRPV1 (receptor for noxious heat and capsaicin, the active component in chili peppers) signaling. This finding shows that our fly-based genetic pain network map is a valuable tool for the discovery of novel “nociception genes” in mammals.
Introduction
Although studies in inbred mouse strains and in human twin cohorts have indicated that pain has a strong genetic component [1]–[5], with an estimated heritability of ∼50%, little is known about the specific genes involved in regulating pain sensitivity across phyla. Further, conservation of gene function between species and across evolutionary time acts as a useful tool to develop an understanding of core genetic mechanisms relative to more specialized programs and how they influence behavior [6]. Drosophila is an excellent model organism for characterizing genetic regulators of behavior such as nociception [7]. Use of Drosophila genetics has highlighted a conserved role for multiple genes in the detection and avoidance of noxious heat [8], [9], and recent work on mechanosensation suggests the genetics of this process is likely also highly conserved across phyla [10]. We have previously reported a global in vivo RNAi screen for avoidance of noxious heat in Drosophila, and identification of hundreds of novel genes required in the adult fly, for its manifestation [8]. To interrogate this resource, we have now constructed a global systems network of heat pain. Our goal was to identify potentially conserved genes and pathways involved in pain perception, to provide a tool for focusing research on key pain molecular pathways.
One pathway highlighted in this global systems network was phosphatidylinositol signaling. Phosphatidylinositol signaling is a second messenger cascade involving the sequential phosphorylation of phosphatidylinositol 4-phosphate (PIP) to generate phosphatidylinositol 4,5-bisphosphate (PIP2) via PIP5' kinases (phosphatidylinositol-5-OH kinase; PIP5K), and then phosphorylation of PIP2 via PI3 kinases (phosphatidylinositol-3-OH kinase; PI3K) to generate phosphatidylinositol 3,4,5-trisphosphate (PIP3). In mammalian systems phospholipid signaling has been implicated in regulating pain perception [11]–[13], TRPV1 function [14]–[21], and itch [22], but how phosphatidylinositol signaling is involved in mammalian nociception is controversial, with data suggesting for example that PIP2 may either increase or decrease TRPV1 function [23], [24]. Based on the suggested involvement of phospholipid signaling in our conserved functional pain network, and in context of the controversial role for phosphatidylinositol signaling in pain perception, we employed genetic approaches to evaluate the role of phosphatidylinositol signaling in mammalian nociception.
Results/Discussion
To construct a global systems network of heat pain we first identified potential mouse and human orthologs of fly candidate pain genes (Figure S1). Of the 580 candidate fly thermal nociception genes we had previously identified [8], 399 had human orthologs (Table S1), many of which are known mammalian pain genes (Table S2). Gene ontology (GO) analyses of the human and mouse orthologs of the fly thermal nociception hits showed a marked enrichment of genes involved in neurotransmission and secretion, housekeeping systems such as mitochondrial structure, ATP synthesis, metabolism, or calcium signaling (Figure S2A–S2C and Table S3). We next generated an interaction map based on first-degree binding partners for the fly thermal nociception hits (Table S4). All binding partners were identified in yeast-2-hybrid screens and reported in the biomolecular interaction network database BIND, i.e. binding partners experimentally confirmed to interact with the candidate genes. Among the first degree binding partners in this interaction network, we found fly allatostatin C receptor 1, which has homology with mammalian opioid receptors, a fly homolog of Lmx1b, which regulates central serotonergic responses to opioids [25], a fly gene related to the nuclear factor-erythroid 2-related factor 2, which has anti-nociceptive effects [26] by inducing upregulation of heme oxygenase-1, and tyrosine hydroxylase, the enzyme required for dopamine and catecholamine production [27]. Thus, our genome-wide functional screen for thermal nociception in flies has generated a human gene network that includes orthologs of several known mammalian pain genes, in addition to numerous uncharacterized genes and pathway not previously associated with pain perception.
To construct a mammalian systems map of thermal nociception, we performed an enrichment analysis (using KEGG pathways and Broad Institute C2 gene sets) on the mouse and human orthologs of the fly pain genes and their first degree binding partners (Figure S1; Table S5 and Table S6). We found significant enrichment of genes (hypergeometric enrichment >90%) involved in mitochondria, metabolism, calcium signaling, inflammation, cell adhesion, RNA processing, and neurotransmission. Finally, to generate a comprehensive conserved network map of thermal nociception, the KEGG pathways from Drosophila, mouse and human were combined with relevant gene sets from the C2 annotations to create a global putative “nociception network” (Figure 1; Table S7). From this combined systems network, we identified gene sets or pathways known to play key roles in many major neural systems.
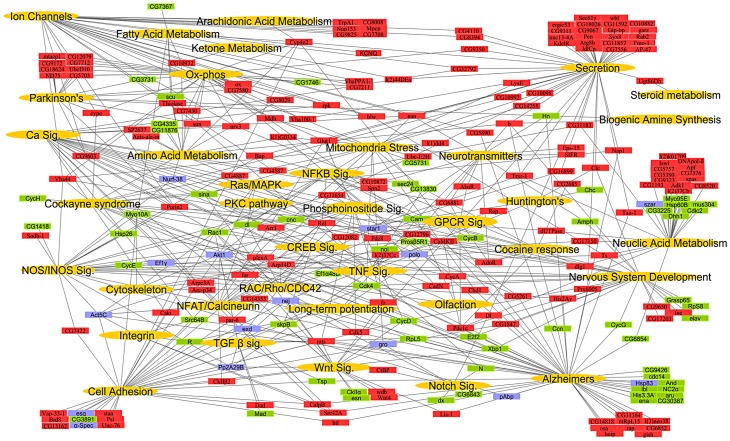
Figure 1. A global network map of thermal nociception.
The systems network includes data from significantly enriched Drosophila KEGG pathways and GO processes, mouse and human KEGG pathways and C2 gene sets. Pathways, processes and gene sets that share a role in a biological process were pooled into functional classes while the underlying genes that constitute them are depicted with a connection to their respective functional class. Functional classes (gold), genes representing direct hits with a thermal nociception phenotype (red), their first degree binding partners (green), and developmental lethal genes (blue) represent the nodes in the network. Only select KEGG pathways, biological processes and C2 gene sets were used to build systems map. For the entire list of individual pathways, gene sets, and processes see Tables S5, S6, S7.
The connectivity of the entire systems map remains intact even after omitting all binding partners (Figure S3), i.e., expansion of the “pain” network map by including binding partners does not introduce a bias. Moreover, all except one pathway in the network map has >50% representation from the Drosophila heat pain hits alone. To test whether this approach can predict mammalian pain genes, we measured the overlap of the direct candidate pain hits and their binding partners with previous rat microarray expression profiling data generated from pain studies [28] and all “pain” annotated genes from OMIM (Online Mendelian Inheritance in Man, NCBI). Intriguingly, we found a 38.55% overlap of our direct pain hits and a 42.33% overlap of their binding partners with the microarray and OMIM data for pain. In contrast, 100 random gene lists gave a maximum of 6% overlapping genes and a minimum of 0% (Table 1; Table S8). Thus, our hypothesis-free systems map is markedly enriched for genes known to be associated with rodent and human pain. Considering the complexity of the neuronal network involved in generating nociception in the periphery and CNS, these pathways may operate across several neurons; however, our genome-wide functional fly pain screen and in silico data mining provide a road map for conserved molecular components and pathways putatively involved in heat nociception globally across phyla.
Table 1. Comparison of systems map with microarray and OMIM data for pain.
| Genes in system map | Genes matching with pain report in OMIM | Genes matching with pain report in Microarray data | Common to OMIM & Microarray | Total matched genes (Microarray+OMIM) | |||||
| Number of genes in System map | Percentage of genes in System map | Number of genes in OMIM | Percentage of genes in OMIM | Number of genes in Microarray data | Percentage of genes in Microarray data | Number of genes common to OMIM & Microarray | Number of genes in Microarray+OMIM | Percentage of genes in Microarray+OMIM | |
| Pain | 166 | 68.03% | 40 | 24.10% | 34 | 20.48% | 10 | 64 | 38.55% |
| Binding Partner | 78 | 31.97% | 20 | 25.64% | 23 | 29.48% | 10 | 33 | 42.31% |
A comparison of genes in the global network map of thermal nociception with genes with a pain annotation in the OMIM database (http://www.ncbi.nlm.nih.gov/omim) and previous rat microarray expression profiling data generated from pain studies [28] showed the global network map of thermal nociception is enriched for genes implicated in mammalian pain diseases.
We next wanted to validate whether this “nociception network” has the power to identify conserved pathways involved in nociception, and if this pathway information can then help to pinpoint key mammalian pain genes. To this end we focused our first efforts on phosphatidylinositol signaling, one of the major nodes predicted from the pain systems map, and a heat pain precedented pathway. Phosphatidylinositol signaling has been implicated in heat nociception and regulation of TRPV1 by multiple groups [11], [13], [16], [18], however its precise role has been controversial [16], [18], [23], and the specific participation of different phospholipid kinases has never been evaluated genetically, which we now decided to do.
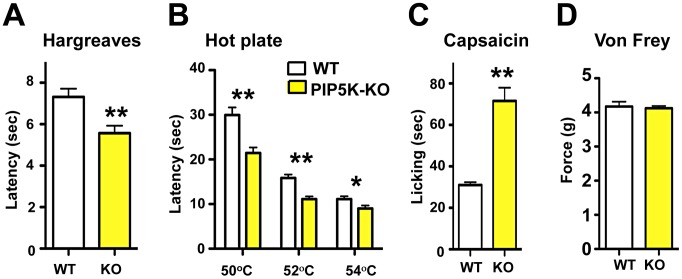
Phosphatidylinositol signaling involves the generation of PIP2 via PIP5K, and then phosphorylation of PIP2 via PI3K to generate PIP3. PIP5Kα is highly expressed in the nervous system but no neuronal function for this kinase has been established [29]. PIP5Kα mutant mice are viable and exhibit an exaggerated anaphylactic immune reaction in response to Fc-receptor engagement [30]. We find that PIP5Kα mutant mice exhibit a significant hyper-responsiveness to radiant heat (Figure 2A) and contact heat, when compared to littermate controls (Figure 2B). In mammals, TRPV1 is the prototypical noxious heat thermo-receptor, and is also the receptor for capsaicin, the active ingredient in chili peppers [31]. We therefore tested whether PIP5Kα mutant mice exhibit exaggerated TRPV1 agonist responses. Indeed, following capsaicin injection, PIP5Kα mutant mice display heightened reactivity compared to littermate controls (Figure 2C) but exhibited no difference in mechanical pain threshold using the von Frey test (Figure 2D).
Figure 2. PI5Kα signaling controls thermal and capsaicin nociception in vivo.
(A) Thermal pain thresholds of wild type (WT) and PIP5Kα −/− (KO) littermates in response to radiant heat (Hargreaves; n = 6 for WT; n = 6 for KO mice). (B) PIP5Kα KO mice also show enhanced thermal sensitivity in the hot plate assay (n = 12 for WT; n = 12 for KO mice) and (C) an exaggerated capsaicin-evoked behavioral response (n = 12 for WT; n = 9 for KO mice). (D) PIP5Kα KO mice exhibited normal mechanical pain (force threshold latency) as assessed by the von Frey test (n = 9 for WT; n = 9 for KO mice). All data are presented as mean +/− sem. *p<0.05; **p<0.01 (t-test).
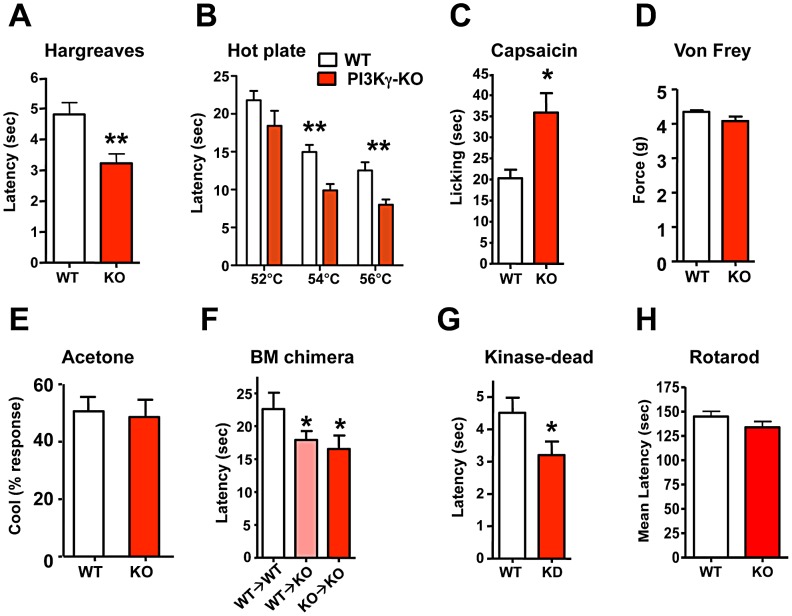
PI3Kγ, the only G-protein coupled PI3K, is expressed in TRPV1-positive peripheral sensory neurons in both rats and mice [32], [33] and has been implicated in morphine-induced peripheral analgesia [32] and morphine tolerance [33]. Use of non-specific inhibitors, like wortmannin, has suggested a role for PI3' kinases in producing NGF-mediated TRPV1 sensitization [21],[34]. We therefore tested thermal nociception in PI3Kγ (p110γ) mutant mice [35] and found that these mice, like the PIP5Kα null mice, also exhibit an exaggerated behavioral response to radiant heat plantar stimulation using the Hargreaves test (Figure 3A). This enhanced pain sensitivity was confirmed using the hot plate assay (Figure 3B). PI3Kγ mutant mice also exhibit an enhanced pain response to a capsaicin challenge (Figure 3C). Similar to PIP5Kα null mice, the mechanical pain threshold using the von Frey test (Figure 3D), and the behavioral responses to acetone application (a cooling sensation) (Figure 3E) were comparable between control and PI3Kγ−/− littermates. Of note we found a similar thermal hyperalgesia phenotype in a second independent PI3Kγ−/− mutant mouse strain [36] (not shown). Thus, genetic loss of PI3Kγ and PIP5Kα results in enhanced pain responses to heat and capsaicin, providing evidence that phosphatidylinositol signaling acts as a negative regulator of heat pain perception and TRPV1 reactivity in vivo.
Figure 3. PI3Kγ signaling controls thermal and capsaicin nociception in vivo.
(A) PI3Kγ −/− (KO) mice show enhanced thermal pain thresholds in response to radiant heat (Hargreaves test) as compared to wild type (WT) littermates (n = 22 for WT; n = 27 for KO mice). (B) PI3Kγ KO mice exhibit enhanced thermal sensitivity in the hot plate assay (n = 13 for WT; n = 9 for KO mice) and (C) an exaggerated capsaicin-evoked behavioral response (n = 9 for WT; n = 13 for KO mice). (D) Normal mechanical pain (force threshold latency) in PI3Kγ KO mice as assessed by the von Frey test (n = 21 for WT; n = 16 for KO mice). (E) PI3Kγ KO mice exhibit comparable cold sensitivity as assessed by acetone application (n = 21 for WT; n = 16 for KO mice). (F) Thermal nociception in PI3Kγ KO mice reconstituted with wild type (WT→KO) or PI3Kγ −/− (KO→KO) bone marrow. WT mice reconstituted with WT bone marrow are shown as controls. Mice were assayed using the hot plate assay at 54°C. n>6 for each group. (G) Thermal nociception thresholds in response to radiant heat (Hargreaves test) in littermate control and PI3Kγ kinase-dead (KD) knock-in mice (n = 19 for WT; n = 23 for KD mice). (H) Control and PI3Kγ KO mice exhibit similar coordination in the Rotarod test (n = 11 for WT and n = 12 for KO mice). All data are presented as mean +/− sem. *p<0.05; **p<0.01 (t-test).
Since PIP5Kα and PI3Kγ cooperate to sequentially generate PIP3, and both mutant mice exhibit a similar hypersensitivity phenotype, we focused further on the role of PI3Kγ and PIP3 generation in setting the threshold for heat pain perception. PI3Kγ is highly expressed in haematopoietic cells and functions as key mediator of inflammatory cell migration to the site of injury [35], [37]. We therefore tested PI3Kγ mutant mice for potential defects in inflammation-induced pain sensitization, i.e. thermal hyperalgesia. PI3Kγ−/− and control mice developed comparable levels of thermal hyperalgesia (Figure S4A and S4B) following plantar CFA injection. CFA-induced inflammation, as determined by paw swelling, was also comparable between mutant and control mice (Figure S4C). To further exclude a potential role of haematopoietic cells, we transplanted wild type bone marrow into PI3Kγ mutant mice (WT→KO) and PI3Kγ mutant bone marrow into wild type mice (KO→WT). The presence of a wild type haematopoietic system did not rescue the enhanced sensitivity to thermal pain in the PI3Kγ mutant background (Figure 3F), i.e. the requirement for PI3Kγ in thermal sensing maps to non-haematopoeitic cells. PI3Kγ has been shown to act both in a kinase-dependent fashion, through conversion of PIP2 to PIP3, and in a kinase-independent manner [38]. We therefore tested the behavioral response of PI3Kγ kinase-dead (KD) knock-in mice. PI3Kγ KD mice exhibited a heightened reaction to noxious heat with a reduced thermal nociception latency (Figure 3G), comparable to the enhanced heat pain responses observed in complete PI3Kγ mutant mice. Thus, the kinase activity generating PIP3 modulates heat pain. We also assessed the general neurological phenotypes of PI3Kγ−/− mice, all of which appeared normal (Figure 3H; Figure S5A–S5G). Furthermore, the overall morphology and histology of the central nervous system appeared normal in PI3Kγ−/− mice. These data demonstrate that generation of PIP3 through PI3Kγ negatively regulates pain sensitivity in vivo.
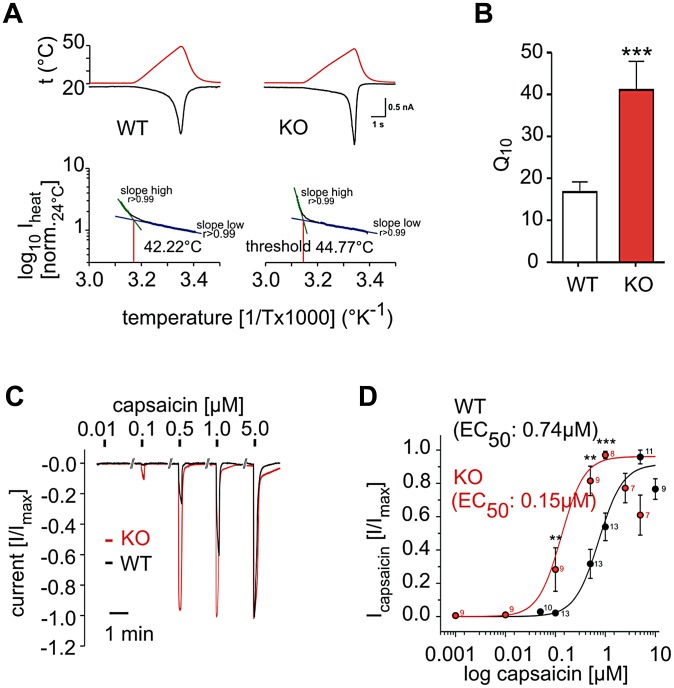
To test if the phosphatidylinositol signaling pathway acts in primary sensory nociceptors, we employed electrophysiology on isolated wild type and PI3Kγ−/− dorsal root ganglion (DRG) neurons. PI3Kγ−/− DRG neurons responded to a thermal ramp (Figure 4A) with a significantly increased steepness in the inward current response to increasing temperature when compared to control neurons. This translated into a substantial increase in the Q10 value (Figure 4B), a measure of temperature-dependent rate change in channel conductivity, indicating that PI3Kγ−/− DRG cells exhibit massive hyper-activation in response to noxious heat, although initiation of this response occurs at a slightly elevated temperature (44.77°C for PI3Kγ−/− vs 42.22°C for wild type DRG neurons, Figure 4A). Since we also observed an enhanced response to capsaicin in PI3Kγ−/− mice in vivo, we directly tested TRPV1 reactivity to capsaicin in sensory neurons in vitro. In accordance with our behavioral data, isolated PI3Kγ−/− DRG neurons exhibited augmented sensitivity to capsaicin (Figure 4C and 4D). Thus, PI3Kγ functions as a negative regulator of TRPV1 responses in nociceptive neurons.
Figure 4. PI3Kγ acts in DRG neurons as a negative regulator of thermal and TRPV1 responses.
(A) Representative temperature response ramps and Arrhenius plots for heat-activated currents measured in single DRG neurons isolated from wild type (WT) and PI3Kγ mutant (KO) mice. For temperature response ramps, red lines depict temperature ramps and black lines depict inward current. (B) Q10 as a measure of the rate of inward current changes in response to temperature. n = 37 for isolated WT; n = 9 for PI3Kγ KO DRG neurons. (C,D) Capsaicin sensitivity of DRG neurons isolated from WT and PI3Kγ KO mice. (C) Representative capsaicin responses from a single DRG neuron. (D) Dose-response curves to different concentrations of capsaicin. The capsaicin EC50 is indicated. Numbers indicate numbers of single neurons tested with the indicated capsaicin doses at the respective data points. Electrophysiology data was generated by single neuron patch clamping. Data are presented as mean +/− sem. ** p<0.01, *** p<0.001 (Mann-Whitney u-test).
Our data provide a conserved functional systems network map for pain behaviour. This network map revealed many pathways and gene sets previously reported to be involved in mammalian nociception, including multiple genes annotated as candidate pain genes in the human OMIM database. Thus, our systems approach, starting from a functional whole genome fly screen and bioinformatic construction of a conserved pain network map, has the power to identify regulators of mammalian nociception. Our network pointed to a key role for phosphatidylinositol signaling in noxious heat nociception. Positive as well as negative regulatory functions of phosphatidylinositol signaling on the thermal nociceptive sensor TRPV1 have been reported [15], [16], [19], [20], [23], [24]. Our results provide genetic data that the phosphatidylinositol signaling pathway is relevant to heat pain sensitivity in vivo. In particular, we find that the lipid kinases PIP5α and PI3Kγ are involved in regulating heat nociception responses by acting as negative modulators of thermal pain perception and TRPV1 activity.
Our data reinforce the extraordinary evolutionary conservation of the neurobiological mechanisms of nociception, from its manifestation as an acute damage avoidance response in simple organisms like flies to the complex sensation of pain in mammals. When used in conjunction with additional complimentary approaches (e.g. published literature, gene expression profiling, or genetic association studies), this systems network map should be a valuable tool to further pinpoint and prioritize novel candidate nociception genes in mammals.
Materials and Methods
Bioinformatics analysis
Identification of fly orthologs in mouse and human was done using pre-computed orthology predictions [39]. Gene Ontology (GO) analysis was performed using GOstat. Binding partner identification was done using GeneSpring GX. Hypergeometric tests were used to identify over-represented gene lists (BROAD Institute) and pathways (KEGG) amongst the pain hits and to generate a conserved systems map. Pain genes and binding partners in the system map that have been annotated as pain genes in the Online Mendelian Inheritance in Man database or by our previous Microarray experiments [28] were also identified.
Mouse behavioral tests
PI3Kγ (p110γ) knock out [35], [36], kinase dead PI3Kγ knock-in [38], and PIP5Kα mutant mice [30] and have been previously described. Thermal and mechanical sensitivities were assessed using the Hargreaves, hot plate, and von Frey tests. Capsaicin behavior was assessed over 5 minutes following intraplantar injection of capsaicin.
Single DRG neuron recordings
Lumbar dorsal root ganglia (DRG) were harvested as previously reported [40], [41]. Patch-clamp recordings were performed using the whole-cell voltage-clamp configuration of the patch-clamp technique as previously described [40], [41].
Ethics statement
All mice were bred and maintained according to an ethical animal license protocol complying with the current Austrian law.
Detailed Materials and Methods are available in Text S1.
Supporting Information
Flow charts for global bioinformatics analyses. (A) Flow chart indicating the steps in the conversion of Drosophila candidate thermal nociception genes into mouse and human orthologs. (B) Flow chart indicating KEGG and C2 gene set analysis in Drosophila, mice, and humans. Data from all three organisms were pooled to generate a global network map. See Methods for details.
(PDF)
Conserved regulators of thermal nociception. GO enrichment analysis for human (A) and mouse (B) orthologs of Drosophila thermal nociception hits. GO terms were pooled into functional categories for data representation. Numbers indicate the counts of GO terms included in each functional category. These GO terms are grouped according to their parent terms - cellular components (CC), biological processes (BP), and molecular functions (MF). (C) Global C2 data set for mammalian orthologs. Functional classification of C2 gene sets (MsigDb, Broad institute) found enriched in mouse and human orthologs of our primary fly candidate thermal nociception genes and their first degree binding partners. The numbers indicate statistically significant C2 genes sets grouped into each functional category.
(PDF)
A global network map of thermal nociception based on primary hits without binding partners. The systems network includes data from the significantly enriched Drosophila KEGG pathways and GO processes, mouse and human KEGG pathways and C2 gene sets based on analysis of primary screening hits without binding partners. Pathways, processes and gene sets that share a role in a biological process were pooled into functional classes while the underlying genes that constitute them are depicted with a connection to their respective functional class. Functional classes (brown), genes representing hits with pain phenotype (red), and developmental lethal genes (blue) are shown in the network. For the entire list of individual pathways and gene sets see Tables S5, S6, S7.
(PDF)
PI3Kγ KO mice show no difference in inflammatory pain responses. (A) PI3Kγ KO mice exhibit significant enhancement of baseline thermal nociception sensitivity but a comparable degree of thermal hyperalgesia following intraplantar CFA challenge. (B) A sensitization ratio (Baseline/CFA latency) shows similar CFA-induced sensitization in wild type (WT) and PI3Kγ KO mice 10 days after CFA injection. (C) WT and PI3Kγ KO mice exhibit similar paw swelling in response to CFA over the course of the experiment (data shown was recorded at day 8). Data are presented as mean +/− sem. * P<0.05, ** P<0.01 by Student's t test.
(PDF)
Basic behavioral analysis of PI3Kγ KO mice. PI3Kγ expressing control and PI3Kγ knock-out (KO) mice exhibit similar behavioral responses in multiple paradigms. (A) PI3Kγ KO mice exhibit intact accuracy in a test for skilled reaching and (B) a slight decrease in cages crosses over a 24 hour period; but this trend failed to reach statistical significance. (C) In an open field test, control and PI3Kγ KO mice showed the same activity over a ten minute period and (D) similar levels of anxiety-related defecation. (E–G) No differences were detected between control and PI3Kγ KO mice in three models of learning: (E) Water maze, (F) T-maze, and (G) passive avoidance learning. (n = 11 for control and n = 12 for KO for the above tests). Data are presented as mean +/− sem. In all experiments there we no significant differences (t-test).
(PDF)
Predicted mammalian orthologs for Drosophila pain genes and binding partners. To identify orthologs between Drosophila and mouse or Drosophila and human, we used pre-computed orthology predictions obtained from Compara r49, Homologene (03/08), Inparanoid v6.1, Orthomcl v2 [39]. Both One-to-one and many to many mappings were observed between the Drosophila and the mammalian genes.
(XLS)
Drosophila candidate pain genes with known mammalian pain gene orthologs. Drosophila candidate pain genes were compared to known mouse pain genes, and overlap is shown. Unique Drosophila (CG number) and mouse (Entrez ID) identifiers are provided. In some cases, multiple independent transgenic RNAi fly lines targeting a particular Drosophila gene were found to exhibit a nociception defect, in which case all transformant ids are provided for a given Drosophila CG number. In some instances, a single candidate Drosophila pain gene was homologous to multiple known mammalian pain genes. Mammalian pain genes were identified using the Pain Genes Database [42].
(XLS)
GO analysis for mammalian orthologs of Drosophila candidate pain genes. GO enrichment for all mouse and human orthologs corresponding to Drosophila candidate pain genes. Significant GO ids, corresponding GO terms, functional category, Biological Process (BP), Molecular Function (MF) and Cellular Component (CC), gene count, total gene count, and P value have been included. Since GO terms that lie at level 4 or below in the GO hierarchy tree convey more biological information, we excluded terms with more than 500 genes. Further, if both parent and several child terms are found significant, we manually curated the data and retained the term that contained the maximum overlapping genes with the pain hit list. These terms are highlighted in the mouse and the human lists separately. From these selected GO terms, those that denote related functions in biological systems were manually clubbed together and functionally annotated. This analysis is included in the column labeled “Manually assigned functional category”. A P-value of 0.1 was considered significant. “Count” and “Total” indicate the numbers of hits among all genes assigned to a defined GO term.
(XLS)
Interactions between candidate pain genes and first degree binding partners. All interactions involving mammalian orthologs of Drosophila pain hit genes and their first degree binding partners are listed. Each row corresponds to a biological interaction between two proteins. Every interaction has two participating nodes: Source and Target. Gene symbols and CG IDs for all source and target nodes are listed. Classifications of all source and target nodes into pain hits (pain), binding partners (BP), and developmentally lethal phenotypes are provided. Potential mosue and human orthologs are presented for hits and binding partners. Both One-to-one and many to many mappings were observed between the Drosophila and the mammalian genes.
(XLS)
Mammalian KEGG analysis. Enrichment of KEGG pathways for mouse and human orthologs corresponding to Drosophila candidate pain genes. The complete analysis for 202 mouse (top), and 209 human (bottom) KEGG pathways are shown. For each pathway, the KEGG database ID, pathway name, KEGG gene sets, CG/Entrez IDs for the mapped genes, numbers of genes assigned to each KEGG pathway, numbers of hits and their binding partners that map to the KEGG pathways, percentages of genes mapped among the total KEGG gene sets, and hypergeometric probabilities are listed. KEGG pathway ID numbers are presented and can be used to obtain complete list of genes included (see http://www.genome.jp/kegg/). Pathways that are over 90% enriched were manually annotated into functional groups and combined with C2 gene sets for visual representation in Figure 1 and Table S7.
(XLS)
Mammalian C2 gene set analysis. Analysis for 687 mouse (top) and 680 human (bottom) C2 gene sets found significant at 99% probability (hypergeometric test). For each gene set, the C2 gene set name, gene symbols, counts for the total number of genes, pain hit orthologs, and percentages of genes mapped among the total C2 gene sets are given. C2 gene set names are presented and can be used to obtain complete list of genes included (see http://www.broadinstitute.org/gsea/msigdb/index.jsp). For constructing the systems map of nociception and uniform functional annotation as in the KEGG pathways, a unique list of significant C2 gene sets were combined from mouse and humans and manually grouped into similar functional categories, as the KEGG pathways. Combined information is provided in Table S7.
(XLS)
Combined KEGG and C2 data for mammalian orthologs of Drosophila candidate pain genes and their binding partners. (A) The table represents a combined list of select significant GO terms, KEGG pathways and C2 gene sets from Drosophila, mouse, and human, used to construct the combined systems map shown in Figure 1. Selected significant KEGG pathways and C2 gene sets were manually grouped into uniform functional categories as shown in the column “Gene set categories”. Gene symbols, Entrez IDs and Drosophila CG IDs were extracted for each pathway or gene set, as appropriate. (B) Percentage of original pain hits and lethals that contribute to each of the significant enrichment of each gene set or pathway.
(XLS)
Overlap of pain systems map with pain OMIM and microarray data. A comparison of genes in the global network map of thermal nociception with genes with a pain annotation in the OMIM database (http://www.ncbi.nlm.nih.gov/omim) and previous rat microarray expression profiling data generated from pain studies [28] showed the global network map of thermal nociception is enriched for genes implicated in mammalian pain. Included are the Drosophila gene CGID, mammalian gene symbol, the genes status as a direct hit (pain) or predicted binding partner (BP), and if this gene was identified as significantly regulated in the dorsal horn (DH) or dorsal root ganglia (DRG) following experimentally induced chronic pain. Genes with a pain annotation in the OMIM database are also identified.
(XLS)
Acknowledgments
We thank all members of our laboratories for helpful discussions and excellent technical support.
Funding Statement
JMP is supported by grants from IMBA, the Austrian Ministry of Sciences, the Austrian Academy of Sciences, GEN-AU (AustroMouse), ApoSys, and an EU ERC Advanced Grant. GGN is supported by a Mary Curie IIF Fellowship, EuroThymaide, and Australian National Health and Medical Research Council project grants (APP1026310, APP1029672, and APP1028887 and an NHMRC CDF fellowship APP1034654). MC is supported by the U.S. National Institutes of Health (R01-NS074430-01A1). CJW is supported by NIH NS039518 and NS038253. MK is supported by FWF (grants 184440 and 205620 and graduate school SPIN) and IFTZ-B14 of MUI. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
- 1. Livshits G, Popham M, Malkin I, Sambrook PN, Macgregor AJ, et al. (2011) Lumbar disc degeneration and genetic factors are the main risk factors for low back pain in women: the UK Twin Spine Study. Ann Rheum Dis 70: 1740–1745. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Lariviere WR, Wilson SG, Laughlin TM, Kokayeff A, West EE, et al. (2002) Heritability of nociception. III. Genetic relationships among commonly used assays of nociception and hypersensitivity. Pain 97: 75–86. [DOI] [PubMed] [Google Scholar]
- 3. Mogil JS, Wilson SG, Bon K, Lee SE, Chung K, et al. (1999) Heritability of nociception I: responses of 11 inbred mouse strains on 12 measures of nociception. Pain 80: 67–82. [DOI] [PubMed] [Google Scholar]
- 4. Nielsen CS, Stubhaug A, Price DD, Vassend O, Czajkowski N, et al. (2008) Individual differences in pain sensitivity: genetic and environmental contributions. Pain 136: 21–29. [DOI] [PubMed] [Google Scholar]
- 5. Hartvigsen J, Nielsen J, Kyvik KO, Fejer R, Vach W, et al. (2009) Heritability of spinal pain and consequences of spinal pain: a comprehensive genetic epidemiologic analysis using a population-based sample of 15,328 twins ages 20–71 years. Arthritis Rheum 61: 1343–1351. [DOI] [PubMed] [Google Scholar]
- 6. Reaume CJ, Sokolowski MB (2011) Conservation of gene function in behaviour. Philos Trans R Soc Lond B Biol Sci 366: 2100–2110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Tracey WD Jr, Wilson RI, Laurent G, Benzer S (2003) painless, a Drosophila gene essential for nociception. Cell 113: 261–273. [DOI] [PubMed] [Google Scholar]
- 8. Neely GG, Hess A, Costigan M, Keene AC, Goulas S, et al. (2010) A genome-wide Drosophila screen for heat nociception identifies alpha2delta3 as an evolutionarily conserved pain gene. Cell 143: 628–638. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Neely GG, Keene AC, Duchek P, Chang EC, Wang QP, et al. (2011) TrpA1 regulates thermal nociception in Drosophila. PLoS ONE 6: e24343 doi:10.1371/journal.pone.0024343. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Kim SE, Coste B, Chadha A, Cook B, Patapoutian A (2012) The role of Drosophila Piezo in mechanical nociception. Nature 483: 209–212. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Pereira PJ, Lazarotto LF, Leal PC, Lopes TG, Morrone FB, et al. (2011) Inhibition of phosphatidylinositol-3 kinase gamma reduces pruriceptive, inflammatory, and nociceptive responses induced by trypsin in mice. Pain 152: 2861–2869. [DOI] [PubMed] [Google Scholar]
- 12. Xu Q, Fitzsimmons B, Steinauer J, O'Neill A, Newton AC, et al. (2011) Spinal phosphinositide 3-kinase-Akt-mammalian target of rapamycin signaling cascades in inflammation-induced hyperalgesia. J Neurosci 31: 2113–2124. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Pezet S, Marchand F, D'Mello R, Grist J, Clark AK, et al. (2008) Phosphatidylinositol 3-kinase is a key mediator of central sensitization in painful inflammatory conditions. J Neurosci 28: 4261–4270. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Yao J, Qin F (2009) Interaction with phosphoinositides confers adaptation onto the TRPV1 pain receptor. PLoS Biol 7: e46 doi:10.1371/journal.pbio.1000046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Liu B, Zhang C, Qin F (2005) Functional recovery from desensitization of vanilloid receptor TRPV1 requires resynthesis of phosphatidylinositol 4,5-bisphosphate. J Neurosci 25: 4835–4843. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Prescott ED, Julius D (2003) A modular PIP2 binding site as a determinant of capsaicin receptor sensitivity. Science 300: 1284–1288. [DOI] [PubMed] [Google Scholar]
- 17. Ufret-Vincenty CA, Klein RM, Hua L, Angueyra J, Gordon SE (2011) Localization of the PIP2 sensor of TRPV1 ion channels. J Biol Chem 286: 9688–9698. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Kim AY, Tang Z, Liu Q, Patel KN, Maag D, et al. (2008) Pirt, a phosphoinositide-binding protein, functions as a regulatory subunit of TRPV1. Cell 133: 475–485. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Klein RM, Ufret-Vincenty CA, Hua L, Gordon SE (2008) Determinants of molecular specificity in phosphoinositide regulation. Phosphatidylinositol (4,5)-bisphosphate (PI(4,5)P2) is the endogenous lipid regulating TRPV1. J Biol Chem 283: 26208–26216. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Stein AT, Ufret-Vincenty CA, Hua L, Santana LF, Gordon SE (2006) Phosphoinositide 3-kinase binds to TRPV1 and mediates NGF-stimulated TRPV1 trafficking to the plasma membrane. J Gen Physiol 128: 509–522. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Zhuang ZY, Xu H, Clapham DE, Ji RR (2004) Phosphatidylinositol 3-kinase activates ERK in primary sensory neurons and mediates inflammatory heat hyperalgesia through TRPV1 sensitization. J Neurosci 24: 8300–8309. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Lee B, Descalzi G, Baek J, Kim JI, Lee HR, et al. (2011) Genetic enhancement of behavioral itch responses in mice lacking phosphoinositide 3-kinase-gamma (PI3Kgamma). Mol Pain 7: 96. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Lukacs V, Thyagarajan B, Varnai P, Balla A, Balla T, et al. (2007) Dual regulation of TRPV1 by phosphoinositides. J Neurosci 27: 7070–7080. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Chuang HH, Prescott ED, Kong H, Shields S, Jordt SE, et al. (2001) Bradykinin and nerve growth factor release the capsaicin receptor from PtdIns(4,5)P2-mediated inhibition. Nature 411: 957–962. [DOI] [PubMed] [Google Scholar]
- 25. Zhao ZQ, Gao YJ, Sun YG, Zhao CS, Gereau RWt, et al. (2007) Central serotonergic neurons are differentially required for opioid analgesia but not for morphine tolerance or morphine reward. Proc Natl Acad Sci U S A 104: 14519–14524. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Rosa AO, Egea J, Lorrio S, Rojo AI, Cuadrado A, et al. (2008) Nrf2-mediated haeme oxygenase-1 up-regulation induced by cobalt protoporphyrin has antinociceptive effects against inflammatory pain in the formalin test in mice. Pain 137: 332–339. [DOI] [PubMed] [Google Scholar]
- 27. Hnasko TS, Sotak BN, Palmiter RD (2005) Morphine reward in dopamine-deficient mice. Nature 438: 854–857. [DOI] [PubMed] [Google Scholar]
- 28. Costigan M, Befort K, Karchewski L, Griffin RS, D'Urso D, et al. (2002) Replicate high-density rat genome oligonucleotide microarrays reveal hundreds of regulated genes in the dorsal root ganglion after peripheral nerve injury. BMC Neurosci 3: 16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Volpicelli-Daley LA, Lucast L, Gong LW, Liu L, Sasaki J, et al. (2010) Phosphatidylinositol-4-phosphate 5-kinases and phosphatidylinositol 4,5-bisphosphate synthesis in the brain. J Biol Chem 285: 28708–28714. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Sasaki J, Sasaki T, Yamazaki M, Matsuoka K, Taya C, et al. (2005) Regulation of anaphylactic responses by phosphatidylinositol phosphate kinase type I {alpha}. J Exp Med 201: 859–870. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Caterina MJ, Schumacher MA, Tominaga M, Rosen TA, Levine JD, et al. (1997) The capsaicin receptor: a heat-activated ion channel in the pain pathway. Nature 389: 816–824. [DOI] [PubMed] [Google Scholar]
- 32. Cunha TM, Roman-Campos D, Lotufo CM, Duarte HL, Souza GR, et al. (2010) Morphine peripheral analgesia depends on activation of the PI3Kgamma/AKT/nNOS/NO/KATP signaling pathway. Proc Natl Acad Sci U S A 107: 4442–4447. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Konig C, Gavrilova-Ruch O, von Banchet GS, Bauer R, Grun M, et al. (2010) Modulation of mu opioid receptor desensitization in peripheral sensory neurons by phosphoinositide 3-kinase gamma. Neuroscience 169: 449–454. [DOI] [PubMed] [Google Scholar]
- 34. Bonnington JK, McNaughton PA (2003) Signalling pathways involved in the sensitisation of mouse nociceptive neurones by nerve growth factor. J Physiol 551: 433–446. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Sasaki T, Irie-Sasaki J, Jones RG, Oliveira-dos-Santos AJ, Stanford WL, et al. (2000) Function of PI3Kgamma in thymocyte development, T cell activation, and neutrophil migration. Science 287: 1040–1046. [DOI] [PubMed] [Google Scholar]
- 36. Hirsch E, Katanaev VL, Garlanda C, Azzolino O, Pirola L, et al. (2000) Central role for G protein-coupled phosphoinositide 3-kinase gamma in inflammation. Science 287: 1049–1053. [DOI] [PubMed] [Google Scholar]
- 37. Hawkins PT, Stephens LR (2007) PI3Kgamma is a key regulator of inflammatory responses and cardiovascular homeostasis. Science 318: 64–66. [DOI] [PubMed] [Google Scholar]
- 38. Patrucco E, Notte A, Barberis L, Selvetella G, Maffei A, et al. (2004) PI3Kgamma modulates the cardiac response to chronic pressure overload by distinct kinase-dependent and -independent effects. Cell 118: 375–387. [DOI] [PubMed] [Google Scholar]
- 39. Kuzniar A, van Ham RC, Pongor S, Leunissen JA (2008) The quest for orthologs: finding the corresponding gene across genomes. Trends Genet 24: 539–551. [DOI] [PubMed] [Google Scholar]
- 40. Obreja O, Biasio W, Andratsch M, Lips KS, Rathee PK, et al. (2005) Fast modulation of heat-activated ionic current by proinflammatory interleukin 6 in rat sensory neurons. Brain 128: 1634–1641. [DOI] [PubMed] [Google Scholar]
- 41. Obreja O, Rathee PK, Lips KS, Distler C, Kress M (2002) IL-1 beta potentiates heat-activated currents in rat sensory neurons: involvement of IL-1RI, tyrosine kinase, and protein kinase C. Faseb J 16: 1497–1503. [DOI] [PubMed] [Google Scholar]
- 42. Lacroix-Fralish ML, Ledoux JB, Mogil JS (2007) The Pain Genes Database: An interactive web browser of pain-related transgenic knockout studies. Pain 131: 3 e1–4. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Flow charts for global bioinformatics analyses. (A) Flow chart indicating the steps in the conversion of Drosophila candidate thermal nociception genes into mouse and human orthologs. (B) Flow chart indicating KEGG and C2 gene set analysis in Drosophila, mice, and humans. Data from all three organisms were pooled to generate a global network map. See Methods for details.
(PDF)
Conserved regulators of thermal nociception. GO enrichment analysis for human (A) and mouse (B) orthologs of Drosophila thermal nociception hits. GO terms were pooled into functional categories for data representation. Numbers indicate the counts of GO terms included in each functional category. These GO terms are grouped according to their parent terms - cellular components (CC), biological processes (BP), and molecular functions (MF). (C) Global C2 data set for mammalian orthologs. Functional classification of C2 gene sets (MsigDb, Broad institute) found enriched in mouse and human orthologs of our primary fly candidate thermal nociception genes and their first degree binding partners. The numbers indicate statistically significant C2 genes sets grouped into each functional category.
(PDF)
A global network map of thermal nociception based on primary hits without binding partners. The systems network includes data from the significantly enriched Drosophila KEGG pathways and GO processes, mouse and human KEGG pathways and C2 gene sets based on analysis of primary screening hits without binding partners. Pathways, processes and gene sets that share a role in a biological process were pooled into functional classes while the underlying genes that constitute them are depicted with a connection to their respective functional class. Functional classes (brown), genes representing hits with pain phenotype (red), and developmental lethal genes (blue) are shown in the network. For the entire list of individual pathways and gene sets see Tables S5, S6, S7.
(PDF)
PI3Kγ KO mice show no difference in inflammatory pain responses. (A) PI3Kγ KO mice exhibit significant enhancement of baseline thermal nociception sensitivity but a comparable degree of thermal hyperalgesia following intraplantar CFA challenge. (B) A sensitization ratio (Baseline/CFA latency) shows similar CFA-induced sensitization in wild type (WT) and PI3Kγ KO mice 10 days after CFA injection. (C) WT and PI3Kγ KO mice exhibit similar paw swelling in response to CFA over the course of the experiment (data shown was recorded at day 8). Data are presented as mean +/− sem. * P<0.05, ** P<0.01 by Student's t test.
(PDF)
Basic behavioral analysis of PI3Kγ KO mice. PI3Kγ expressing control and PI3Kγ knock-out (KO) mice exhibit similar behavioral responses in multiple paradigms. (A) PI3Kγ KO mice exhibit intact accuracy in a test for skilled reaching and (B) a slight decrease in cages crosses over a 24 hour period; but this trend failed to reach statistical significance. (C) In an open field test, control and PI3Kγ KO mice showed the same activity over a ten minute period and (D) similar levels of anxiety-related defecation. (E–G) No differences were detected between control and PI3Kγ KO mice in three models of learning: (E) Water maze, (F) T-maze, and (G) passive avoidance learning. (n = 11 for control and n = 12 for KO for the above tests). Data are presented as mean +/− sem. In all experiments there we no significant differences (t-test).
(PDF)
Predicted mammalian orthologs for Drosophila pain genes and binding partners. To identify orthologs between Drosophila and mouse or Drosophila and human, we used pre-computed orthology predictions obtained from Compara r49, Homologene (03/08), Inparanoid v6.1, Orthomcl v2 [39]. Both One-to-one and many to many mappings were observed between the Drosophila and the mammalian genes.
(XLS)
Drosophila candidate pain genes with known mammalian pain gene orthologs. Drosophila candidate pain genes were compared to known mouse pain genes, and overlap is shown. Unique Drosophila (CG number) and mouse (Entrez ID) identifiers are provided. In some cases, multiple independent transgenic RNAi fly lines targeting a particular Drosophila gene were found to exhibit a nociception defect, in which case all transformant ids are provided for a given Drosophila CG number. In some instances, a single candidate Drosophila pain gene was homologous to multiple known mammalian pain genes. Mammalian pain genes were identified using the Pain Genes Database [42].
(XLS)
GO analysis for mammalian orthologs of Drosophila candidate pain genes. GO enrichment for all mouse and human orthologs corresponding to Drosophila candidate pain genes. Significant GO ids, corresponding GO terms, functional category, Biological Process (BP), Molecular Function (MF) and Cellular Component (CC), gene count, total gene count, and P value have been included. Since GO terms that lie at level 4 or below in the GO hierarchy tree convey more biological information, we excluded terms with more than 500 genes. Further, if both parent and several child terms are found significant, we manually curated the data and retained the term that contained the maximum overlapping genes with the pain hit list. These terms are highlighted in the mouse and the human lists separately. From these selected GO terms, those that denote related functions in biological systems were manually clubbed together and functionally annotated. This analysis is included in the column labeled “Manually assigned functional category”. A P-value of 0.1 was considered significant. “Count” and “Total” indicate the numbers of hits among all genes assigned to a defined GO term.
(XLS)
Interactions between candidate pain genes and first degree binding partners. All interactions involving mammalian orthologs of Drosophila pain hit genes and their first degree binding partners are listed. Each row corresponds to a biological interaction between two proteins. Every interaction has two participating nodes: Source and Target. Gene symbols and CG IDs for all source and target nodes are listed. Classifications of all source and target nodes into pain hits (pain), binding partners (BP), and developmentally lethal phenotypes are provided. Potential mosue and human orthologs are presented for hits and binding partners. Both One-to-one and many to many mappings were observed between the Drosophila and the mammalian genes.
(XLS)
Mammalian KEGG analysis. Enrichment of KEGG pathways for mouse and human orthologs corresponding to Drosophila candidate pain genes. The complete analysis for 202 mouse (top), and 209 human (bottom) KEGG pathways are shown. For each pathway, the KEGG database ID, pathway name, KEGG gene sets, CG/Entrez IDs for the mapped genes, numbers of genes assigned to each KEGG pathway, numbers of hits and their binding partners that map to the KEGG pathways, percentages of genes mapped among the total KEGG gene sets, and hypergeometric probabilities are listed. KEGG pathway ID numbers are presented and can be used to obtain complete list of genes included (see http://www.genome.jp/kegg/). Pathways that are over 90% enriched were manually annotated into functional groups and combined with C2 gene sets for visual representation in Figure 1 and Table S7.
(XLS)
Mammalian C2 gene set analysis. Analysis for 687 mouse (top) and 680 human (bottom) C2 gene sets found significant at 99% probability (hypergeometric test). For each gene set, the C2 gene set name, gene symbols, counts for the total number of genes, pain hit orthologs, and percentages of genes mapped among the total C2 gene sets are given. C2 gene set names are presented and can be used to obtain complete list of genes included (see http://www.broadinstitute.org/gsea/msigdb/index.jsp). For constructing the systems map of nociception and uniform functional annotation as in the KEGG pathways, a unique list of significant C2 gene sets were combined from mouse and humans and manually grouped into similar functional categories, as the KEGG pathways. Combined information is provided in Table S7.
(XLS)
Combined KEGG and C2 data for mammalian orthologs of Drosophila candidate pain genes and their binding partners. (A) The table represents a combined list of select significant GO terms, KEGG pathways and C2 gene sets from Drosophila, mouse, and human, used to construct the combined systems map shown in Figure 1. Selected significant KEGG pathways and C2 gene sets were manually grouped into uniform functional categories as shown in the column “Gene set categories”. Gene symbols, Entrez IDs and Drosophila CG IDs were extracted for each pathway or gene set, as appropriate. (B) Percentage of original pain hits and lethals that contribute to each of the significant enrichment of each gene set or pathway.
(XLS)
Overlap of pain systems map with pain OMIM and microarray data. A comparison of genes in the global network map of thermal nociception with genes with a pain annotation in the OMIM database (http://www.ncbi.nlm.nih.gov/omim) and previous rat microarray expression profiling data generated from pain studies [28] showed the global network map of thermal nociception is enriched for genes implicated in mammalian pain. Included are the Drosophila gene CGID, mammalian gene symbol, the genes status as a direct hit (pain) or predicted binding partner (BP), and if this gene was identified as significantly regulated in the dorsal horn (DH) or dorsal root ganglia (DRG) following experimentally induced chronic pain. Genes with a pain annotation in the OMIM database are also identified.
(XLS)