Background: Sumoylation is recognized by proteins with SUMO-interacting motifs.
Results: SUMO-interacting motifs were identified through a computational string search and validated in SUMO binding assays.
Conclusion: Arkadia, FLASH, C5orf25, and SOBP all contain clustered SIMs.
Significance: These proteins contain distinct SUMO binding structures responsible for the recognition of diverse forms of sumoylation.
Keywords: Post-translational Modification, Sumo, Sumoylation, Ubiquitin, Ubiquitylation, Arkadia/RNF111, RNF4, SUMO Binding Domain, SUMO-interacting Motif, String Search
Abstract
Polysumoylation is a crucial cellular response to stresses against genomic integrity or proteostasis. Like the small ubiquitin-like modifier (SUMO)-targeted ubiquitin ligase RNF4, proteins with clustered SUMO-interacting motifs (SIMs) can be important signal transducers downstream of polysumoylation. To identify novel polySUMO-binding proteins, we conducted a computational string search with a custom Python script. We found clustered SIMs in another RING domain protein Arkadia/RNF111. Detailed biochemical analysis of the Arkadia SIMs revealed that dominant SIMs in a SIM cluster often contain a pentameric VIDLT ((V/I/L/F/Y)(V/I)DLT) core sequence that is also found in the SIMs in PIAS family E3s and is likely the best-fitted structure for SUMO recognition. This idea led to the identification of additional novel SIM clusters in FLASH/CASP8AP2, C5orf25, and SOBP/JXC1. We suggest that the clustered SIMs in these proteins form distinct SUMO binding domains to recognize diverse forms of protein sumoylation.
Introduction
Protein sumoylation is a covalent modification on lysine residues by small ubiquitin-like modifier (SUMO)3 family proteins. It occurs through a biochemical pathway similar to ubiquitylation (ubiquitination), symbolized as an E1-E2-E3 catalytic cascade. Like ubiquitylation, sumoylation modifies many proteins in diverse cellular processes yet has distinct functional consequences (1, 2). Sumoylation promotes a protein-protein interaction through the physical contact between SUMO and the SUMO-interacting motif (SIM), a β-strand consisting of hydrophobic (in the pattern of (V/I/L)(V/I/L)X(V/I/L), (V/I/L)X(V/I/L)(V/I/L), or (V/I/L)X(V/I/L)X(V/I/L)) and acidic residues (3–7). As all SIMs are believed to dock onto the same site on SUMO, they constitute a single class of SUMO binding motif. This is in contrast to the multitude of structures recognizing ubiquitin at several surface sites (8).
SUMO-interacting motifs regulate the activity and specificity of sumoylation and are found in both the catalytic cascade (e.g. the PIAS family SUMO ligases) and the substrates (e.g. Bloom's syndrome helicase) (4, 7, 9–11). The other prominent role of SIMs is to transduce the biological effects of sumoylation (3). For example, upon monosumoylation of human thymine DNA glycosylase, an intramolecular interaction between its SIM and the conjugated SUMO causes an allosteric inhibition of the enzyme's activity (12–14). A SIM in promyelocytic leukemia protein (PML) may facilitate the formation of PML nuclear bodies by promoting the self-assembly of sumoylated PML (15, 16). The SUMO-modified PML bodies in turn recruit other SIM-containing proteins such as Daxx, thymine DNA glycosylase, and RNF4 (14, 17–20). The SUMO-SIM interaction thus is crucial for the functional consequences of protein sumoylation.
To explore how the SUMO signal is recognized through SUMO-binding proteins, we previously identified a family of SUMO-targeted ubiquitin ligases (STUbLs), including RNF4 and its fission yeast homologs, Rfp1 and Rfp2 (21). Interestingly, RNF4 family proteins all contain multiple SIMs positioned closely as a cluster. This has raised the possibility that clustered SIMs are attuned to specifically recognize polySUMO chains (20, 22). Although the significance of polysumoylation has been recognized, its function is yet to be well defined (23). Through RNF4-family STUbLs, polySUMO may be a signal for ubiquitylation and proteasome-dependent degradation (18, 20, 24, 25). Likewise, additional SUMO-binding proteins with clustered SIMs may exist and mediate biological effects of polysumoylation other than protein turnover.
We learned from our work that SIM-containing proteins defy a common homology-based identification; although BLAST analysis revealed the similarity between Rfp2 and the mammalian RNF4 based on the sequences of their C-terminal RING domains, it failed to highlight the homology in their SIMs even though this is obvious to the naked eye (21). However, we reasoned from this experience that SIMs could still be computationally identified through a simple motif scan. Similar approaches have been successful in the identification of consensus phosphorylation sites (hence the substrates) of protein kinases (26) as well as other short linear motifs for protein-protein interaction (27). In fact, an in silico search with a limited scope has revealed atypical SIMs similar to the one in CoREST1 (in the pattern of (V/I/L)X(V/I/L)X(V/I/L)) (6). Here we report our discovery of novel SUMO-binding proteins through a systematic computational string search. In particular, we have searched for proteins containing multiple SIMs with sequence similarity to those in RNF4. We identified four mammalian proteins, Arkadia/RNF111, FLASH/CASP8AP2, C5orf25, and SOBP/JXC1, all containing clustered SIMs.
EXPERIMENTAL PROCEDURES
Reagents
The cDNAs of human and mouse Arkadia cDNAs (BC060862 and BC069835, respectively), human C5orf25 (BC037298), and mouse sine oculis-binding protein (SOBP) cDNA (BC059851) were obtained from the Mammalian Gene Collection and were further PCR-amplified for subcloning into appropriate expression vectors as indicated in the paper. Human FLASH cDNA was a gift from Prof. Odd Gabrielsen (University of Oslo). cDNAs of RNF4 and fission yeast Rfp1 and Rfp2 has been described previously (21). Mutants were generated through PCR amplification (Phusion, New England Biolabs) or PCR-based site-directed mutagenesis (QuikChange, Agilent). Recombinant tri-SUMO proteins (FLAG-3xSUMO1 and FLAG-3xSUMO2) are the fusion of an N-terminal FLAG-tagged, full-length SUMO moiety followed by two copies of SUMOΔN mutant (SUMO1ΔN17 or SUMO2ΔN11, respectively) and were purified as His8 fusion proteins. Anti-GST/anti-GFP bivalent antiserum was raised in rabbit against a GST-GFP fusion protein. Anti-FLAG (M2) and anti-panSUMO antibodies were purchased from Sigma and Abgent, respectively. Poly-SUMO2 chains were purchased from BostonBiochem, Inc. All recombinant proteins were expressed in Escherichia coli strain BL21 (DE3) and purified through affinity purification using a His or GST tag following routine protocols. Human embryonic kidney 293T cells were maintained in Dulbecco's modified Eagle's medium supplemented with 10% fetal bovine serum.
String Search
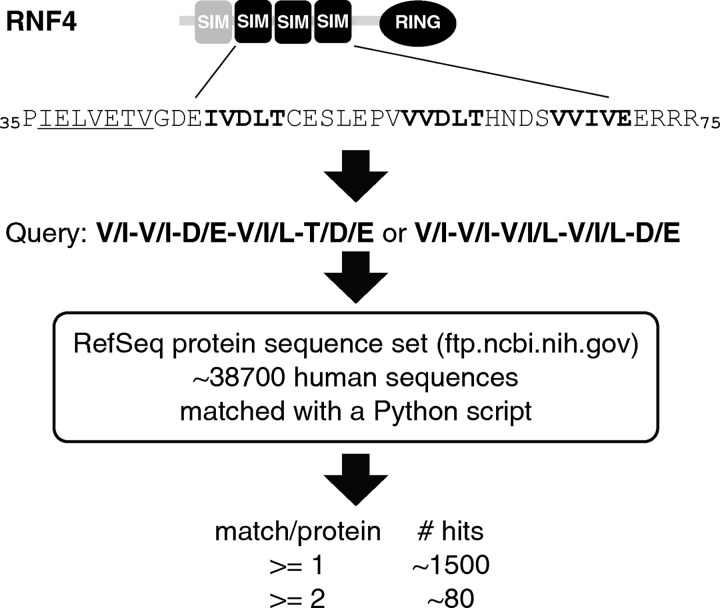
A naïve string search script was written in Python 3 (see supplemental Scripts S1 and S2) to collect protein sequences that each contains more than one SIM-like motifs. Reference proteome sequence files were retrieved from NCBI or UniProt. Using (V/I)(V/I)(D/E)(V/I/L)(T/D/E) and (V/I)(V/I)(V/I/L)(V/I/L)(D/E) as query sequences (as in supplemental Script S1), we obtained about 1500 single hits and about 80 multiple hits (supplemental Table S1). Using (V/I/L/F/Y)(V/I)DLT as the query sequence (supplemental Script S2), we obtained about 200 total hits (supplemental Table S2). Both lists include redundant records or splicing isoforms.
In Vitro GST Pulldown Assay
Glutathione-agarose beads (10 μl bed volume) were mixed with 10 μg of GST fusion protein in 800 μl of binding buffer (50 mm Tris-HCl pH 7.5, 400 mm NaCl, 1% Nonidet P-40, 0.1% sodium deoxycholate, 0.5% BSA) at 4 °C for 30 min followed by the addition of 20 μg of His-FLAG-3xSUMO1 (or 3xSUMO2). The mixture was incubated at 4 °C for 1 h. The beads were washed 3 times with the binding buffer without BSA and boiled in SDS-PAGE sample buffer. The bead-bound GST-fusion proteins and FLAG-3xSUMO were visualized with anti-GST and anti-FLAG immunoblot.
Immunoprecipitation
293T cells growing in 60-mm dishes were transfected using the calcium phosphate precipitation method. For all immunoprecipitation assays the cells were treated with bortezomib (100 nm) for 16 h before cell lysis. Two days after transfection, total cell lysates were prepared with 600 μl per 60-mm dish lysis buffer (50 mm Tris-HCl, pH 8.0, 400 mm NaCl, 0.5% Nonidet P-40, 0.1% sodium deoxycholate, with protease inhibitor mixture, and 2.5 mg/ml of N-ethylmaleimide) assisted with sonication bursts (3 × 10 s using a microtip on a Branson Sonifer 450 with output control set at 1.5). Anti-GFP immunoprecipitation was carried out by incubating the cleared lysate with 2 μl of anti-GFP serum for 1 h at 4 °C followed by another 1-h incubation with protein A-agarose beads. For anti-FLAG immunoprecipitation, a 2-h incubation with anti-FLAG (M2)-conjugated agarose beads (Sigma) was applied instead. The immune complex-bound beads were washed three times with the lysis buffer and analyzed through SDS-PAGE and immunoblotting.
RESULTS
String Search Identified Proteins Containing Multiple Putative SIMs
We conducted our string search in the form of a bioinformatics exercise; we wrote a Python script to run a “naïve string search” against a reference proteome data set, such as a NCBI RefSeq file for human proteins (Fig. 1 and supplemental Script S1). As the consensus sequence of a SIM would match to a very large number of proteins, we only aimed for proteins with SIM-like sequences resembling those in RNF4. In particular, considering that all known SIM structures are β-strands, we have noticed that SIM2, -3, and -4, but not SIM1 in RNF4, favor a β-strand conformation as predicted by PELE, a collection of secondary structure prediction algorithms hosted on Biology Workbench (supplemental Fig. S1A). Prompted by this observation and also by the experimental evidence arguing for a dispensable SIM1 (20), we restricted our search criterion to sequences based on those of SIM2, 3, and 4 of RNF4, and thus conducted the string search using degenerate sequences (V/I)(V/I)(D/E)(V/I/L)(T/D/E) and (V/I)(V/I)(V/I/L)(V/I/L)(D/E) as a start. This search resulted in ∼80 proteins with at least two matches. Besides RNF4 itself and the hypothetical product of an RNF4 pseudogene (Fig. 1 and supplemental Table S1), the list contained three previously reported SIM-containing proteins: chromatin assembly factor 1 subunit A (CAF1p150/CHAF1A), Smad-interacting zinc finger protein 1 (SIZN1, also known as zinc finger CCHC domain containing protein 12 or ZCCHC12), and androgen receptor-interacting protein 4 (ARIP4, or RAD54-like protein 2) (supplemental Table S1 and Fig. S1B) (28–30). While our algorithm did not automatically detect clustered SIMs, a list of 80 proteins as such was short enough for us to visually inspect each protein. As a result, we identified Arkadia/RNF111 with closely located SIM-like motifs as well as a C-terminal RING domain, making it a potential novel SUMO-targeted ubiquitin ligase (Fig. 2A). Notably, secondary structure prediction by PELE strongly suggested that all the SIM-like motifs in Arkadia form β-strands, consistent with the idea that SIMs are acidic β-strands (supplemental Fig. S1A).
FIGURE 1.
Identification of RNF4-like SUMO-interacting motifs through a computational string search. A diagram describes the computational identification of putative SIM-containing proteins based on SIM2, −3, and −4 in RNF4.
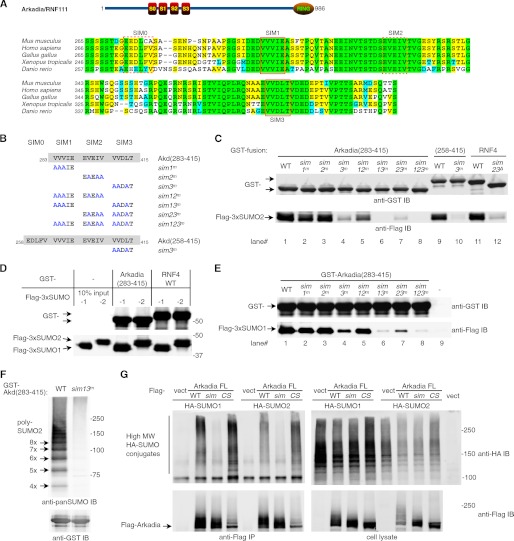
FIGURE 2.
Validation of SUMO-interacting motifs in Arkadia/RNF111. A, shown is the domain structure of Arkadia (top) and an alignment of the predicted SUMO binding regions of Arkadia proteins from mouse (Mus musculus NP_291082), human (Homo sapiens NP_060080), frog (Xenopus tropicalis NP_001072805), chicken (Gallus gallus NP_001186680), and zebrafish (Danio rerio XP_001922708). Solid line rectangles highlight the two motifs (SIM1 and SIM3) matching the initial search string; dashed line rectangles highlight two motifs (SIM0 and SIM2) found through visual inspection. B, shown is a list of Arkadia mutants used for testing SUMO binding. Mutation in each SIM replaced three core hydrophobic residues (Val, Ile, or Leu) with Ala. C, shown is the contribution of Arkadia SIMs to SUMO binding. Various GST fusion proteins as indicated were tested for their association with a FLAG-tagged tri-SUMO2 (FLAG-3xSUMO2) in a GST pulldown assay. The glutathione-agarose bead-bound proteins were detected with anti-GST and anti-FLAG antibodies in an immunoblot. The GST-RNF4 and its sim23Δ, an internal deletion mutant lacking both SIM2 and 3, were used as positive and negative controls, respectively. D, shown is a comparison of the interaction of Arkadia and RNF4 with two major SUMO isoforms. A GST pulldown assay was used to compare the association of Arkadia SBD or RNF4 with either tri-SUMO1 or tri-SUMO2. E, shown is the contribution of Arkadia SIMs to SUMO binding. Various GST fusion proteins as indicated were tested for their association with a FLAG-tagged tri-SUMO1 (FLAG-3xSUMO1) in a GST pulldown assay conducted essentially as in C. IB, immunoblot. F, shown is SIM-dependent association of Arkadia with polySUMO chains. The GST pulldown assay was carried out essentially as in C but against the polySUMO2 ladder. The bead-bound proteins were analyzed with anti-panSUMO or anti-GST immunoblots as indicated. Detectable polySUMO2 species are labeled with the calculated length of SUMO2 units. G, shown is SIM-dependent association between Arkadia and high molecular weight sumoylated proteins in cultured cells detected through coimmunoprecipitation. FLAG-tagged full-length Arkadia in the indicated forms (WT; sim, sim13m; CS, RING domain mutant C971S) were coexpressed with HA-tagged SUMO1 or SUMO2 in 293T cells. Proteins in the anti-FLAG immune complex and in the cell lysate were analyzed in a 4–12% Bis-Tris SDS-PAGE followed by immunoblotting using anti-FLAG and anti-HA antibodies as indicated.
Arkadia Has a SUMO Binding Domain with Three SIMs
Our computational search identified two motifs, 300VVVIE304 and 382VVDLT386 in human Arkadia. Both motifs are surrounded by acidic residues as in a typical SIM. Interestingly, further inspection of the sequence in this region revealed two additional motifs, 274EEDLFV279 and 325EVEIVTV331 (Fig. 2A). The first motif, 274EEDLFV279, is also composed of hydrophobic and acidic residues, but it is not similar to any known SIMs and is not strictly conserved among Arkadia homologs. The second motif, 325EVEIVTV331, which would not have been identified by the string search, partially resembles the SIM1 in RNF4. Sequence alignment indicates that 300VVVIE304, 325EVEIVTV331, and 382VVDLT386 are identical among all Arkadia proteins found in vertebrates and hence are designated as SIM1, SIM2, and SIM3. The motif 274EEDLFV279 upstream of SIM1 is designated as SIM0 (Fig. 2A).
To validate the SUMO binding activity of this region in Arkadia, we carried out a number of protein binding assays. We first isolated an Arkadia fragment that covers the predicted SIM1-SIM3 (283–415) and conducted a GST pulldown against a linear tri-SUMO2 protein, which serves as a polySUMO mimetic. Specifically, to determine whether the SUMO binding was SIM-mediated, mutations were introduced in each SIM-like motif individually and in all four possible combinations (sim1m, sim2m, sim3m, sim12m, sim13m, sim23m, and sim123m) (Fig. 2, B and C). We found that among the three motifs, SIM3 is the most critical, as mutating SIM3 resulted in more significant loss of SUMO binding than mutating SIM1 or SIM2 (Fig. 2C, lane 4). In contrast, mutating SIM2 alone appeared to have no obvious impact on SUMO binding. The contribution of SIM1 to SUMO binding is intermediate between SIM2 and SIM3. Mutating SIM1 and SIM3 together totally abolished SUMO binding as did mutation of all three SIMs (Fig. 2C, lanes 6 and 8). Moreover, we found no difference between the fragment covering SIM1–3 (283–415) and a longer one including the upstream SIM0 (258–415) (Fig. 2C, compare lanes 1 to 9 and lanes 4 to 10), indicating that SIM0 is not functionally significant. In conclusion, Arkadia contains a SIM cluster that recognizes polySUMO chains. Hereinafter, we refer to this unique SUMO-binding structure as a SUMO binding domain (SBD).
Although the precise role of SUMO1 in polySUMO chain formation is still being investigated (31, 32), recent identification of an inverted sumoylation consensus motif, (D/E)XK, raised the possibility that SUMO1-containing or SUMO1-exclusive polySUMO chains can form in vivo with Lys-17 (as in 14GDKKEG19) of SUMO1 serving as the branching site (33). To determine whether the Arkadia SBD has differential affinity toward SUMO isoforms (SUMO1 versus SUMO2/3), we compared linear tri-SUMO1 with linear tri-SUMO2 in parallel binding assays (Fig. 2, D and E). We found that both the Arkadia SBD and the full-length RNF4 protein associated with either tri-SUMO proteins equally well (Fig. 2D), suggesting that the SBDs from Arkadia and RNF4 are not selective against the two major SUMO isoforms. Moreover, when the same set of Arkadia SIM mutants were tested against tri-SUMO1, we obtained a binding profile nearly identical to that with tri-SUMO2 (compare Fig. 2, C and E). Therefore, among the three SIMs in Arkadia SBD, the same hierarchy of SUMO affinity exists regardless of SUMO isoforms, with SIM3 being the most critical and SIM2 being the weakest SIM for both SUMO1 and SUMO2.
In additional assays, we confirmed that the Arkadia SBD could specifically associate (a) with a ladder of authentic polySUMO2 chains in vitro (Fig. 2F), (b) with both EGFP-SUMO1 and EGFP-SUMO2 monomeric fusion proteins in a bead halo binding assay (supplemental Fig. S2A) (34), (c) with a smear of high molecular weight SUMO conjugates in cultured cells (supplemental Fig. S2B), and (d) with SUMO1 in a yeast two-hybrid assay (supplemental Fig. S2C), all in a SIM1/3-dependent manner. Finally, to ascertain the SUMO binding activity of full-length Arkadia proteins, we immunoprecipitated FLAG-tagged, full-length Arkadia from 293T cells coexpressing an HA-tagged SUMO1 or SUMO2 and found that high molecular weight SUMO1 or SUMO2 conjugates specifically associated with wild-type Arkadia and that this association was dependent on an intact SIM cluster but independent of the RING domain (Fig. 2G). We conclude that Arkadia is a SUMO-binding protein capable of recognizing both SUMO1 and SUMO2 through its SUMO binding domain.
Dominant SIMs in an SBD Are Formed by the VIDLT Motif
Our mutational analysis of the Arkadia SBD suggests that individual SIMs in a SIM cluster contribute differentially to SUMO binding (Fig. 2, C and E, and supplemental Fig. S2C). In fact, other clustered SIMs that we encountered previously, i.e. the ones in RNF4 and its fission yeast homologs Rfp1 and Rfp2, also display a two-tiered hierarchy in SUMO binding (supplemental Fig. S3) (20). The SIM3 in Arkadia, SIM2, and SIM3 together in RNF4, SIM2 in Rfp1, and SIM1 in Rfp2 are all essential for their SBDs to interact with SUMO and can provide significant SUMO binding affinity in the absence of the rest of the SIMs in each SBD. We thus call them “dominant SIMs.” In contrast, the rest of the SIMs in each SBD all play an “accessory” role, as their mutation caused minimal or no reduction of SUMO binding (Fig. 2C and supplemental Figs. S2C and S3). We conclude that the two-tiered SUMO affinity is a common phenomenon with clustered SIMs; each SUMO binding domain contains at least one dominant SIM that is more critical for SUMO binding than the rest of the SIMs. We suggest that the combined SUMO recognition by dominant and accessory SIMs provides an optimal avidity effect for targeting complex polySUMO structures.
Most dominant SIMs (except the SIM2 in Rfp1) contain a VIDLT core motif similar to the singular SIMs in PIAS family SUMO E3 ligases (Fig. 3A) (21), indicating that 1) such a high degree of sequence conservation implies a requirement for specific structural features in a dominant SIM and 2) the SUMO affinity of individual SIMs in an SBD largely relies on their particular sequence. These notions prompted us to conduct a more detailed mutational analysis to further ascertain the contribution of each residue of 382VVDLT386 in Arkadia SIM3. We replaced each of the five residues with Ala and found that indeed all five residues are essential for SUMO binding; in contrast, Val-387, the following C-terminal hydrophobic residue, is dispensable (Fig. 3B). Our observation thus complements that of Yuan Chen and co-workers (35), where they show that only a small set of amino acids is allowed in the PIAS1 SIM for high affinity SUMO binding. Together our results indicate that a precise structure is needed for maximum SUMO affinity of the VIDLT-type SIMs, which accepts only limited perturbations.
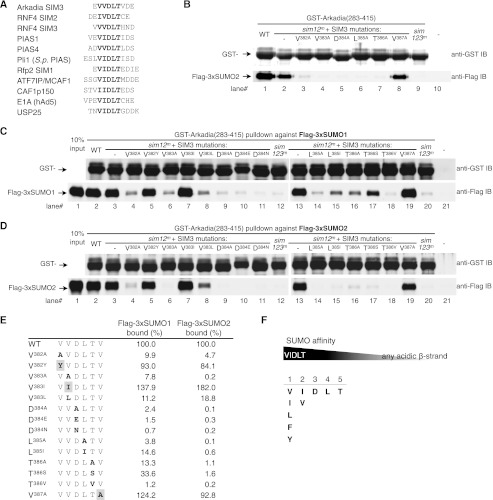
FIGURE 3.
Dominant SIMs are made of VIDLT motif. A, dominant SIMs Arkadia and RNF4 resemble those in PIAS family SUMO ligases. The list shows the alignment of VIDLT-type SIMs in Arkadia, RNF4, PIAS1, PIAS4 (7), Pli1, Rfp2, ATF7IP/MCAF1 (75), CAF1p150 (30), E1A of human adenovirus 5 (76), and USP25 (77). B, all the five core residues in Arkadia SIM3 (VVDLT) are essential for SUMO2 binding. The SIM3 residues Val-382, Val-383, Asp-384, Leu-385, Thr-386, and Val-387 were each replaced with Ala in a sim12m background as indicated. Binding assays were conducted as described in Fig. 2C. IB, immunoblots. C, a rigid structure of VIDLT-type SIMs allowing minimum variations for full strength SUMO1 binding. Additional Arkadia SIM3 mutants, V382Y, V383I, V383L, D384E, D384N, L385I, T386S, and T386V were generated in a sim12m background as indicated. Binding assays were performed as described in Fig. 2C. Examples of purified GST fusion proteins are shown in supplemental Fig. S4. D, a rigid structure of VIDLT-type SIMs allowing minimum variations for full strength SUMO2 binding. Binding assays were performed in parallel to C. E, shown is a summary of single-residue mutations tested in C and D. Mutated residues are indicated in bold; mutations that retained wild-type affinity for SUMO are highlighted. Quantitative densitometry obtained from C and D measures the bound tri-SUMO (FLAG-3xSUMO1 or −3xSUMO2) to the respective GST fusion proteins and is shown as the percentage relative to that of wild-type SIM3. F, shown is a diagram indicating a sequence-dependent SUMO affinity of all putative SIMs (upper panel) and the limited sequence space for the VIDLT-type SIMs (bottom panel).
To further determine the prevalence of this structural rigidity among the VIDLT-type SIMs and to dissect the contribution of VIDLT residues to the recognition of different SUMO isoforms, we introduced more single-residue mutations in Arkadia SIM3; we replaced the 382VVDLT386 pentamer with YVDLT, VIDLT, VLDLT, VVELT, VVNLT, VVDIT, VVDLS, and VVDLV. Some of the substitutions, such as Val to Leu, Asp to Glu, or Leu to Ile, etc., are subtle and would otherwise be considered homologous in common sequence comparisons. We tested these mutants in parallel assays for their physical interaction with tri-SUMO1 and tri-SUMO2. We found that except for YVDLT and VIDLT, all other variants showed significant loss of affinity toward both SUMO1 and SUMO2, again indicating that a precise structure is needed for maximum SUMO affinity of the VIDLT-type SIMs (Fig. 3, C–E). Nevertheless, we did observe that certain mutations at Leu-385 and Thr-386 preserved significant residual affinity toward SUMO1 but not SUMO2 (Fig. 3, C and D, lanes 15–17). We suggest that this reflects a subtle difference between SUMO1 and SUMO2 in the shape of their SIM docking sites at the atomic level, which accounts for the selectivity between certain SIMs (of lesser SUMO affinity) and the different SUMO isoforms (5, 35). It is also remarkable that the change from VVDLT to VIDLT enhanced SUMO binding (Fig. 3, C and D, comparing lanes 3 with 7; Fig. 3E), suggesting VIDLT is indeed the most fitted SIM.
Together, our data suggest that not all SIMs are identical in the spectrum of SUMO affinity and the VIDLT-type SIMs are perhaps best fitted for SUMO binding (Fig. 3F). Within the VIDLT pentamer, structural flexibility is allowed at the first position, where a bulkier hydrophobic side chain, such as Phe, Tyr, or Leu, in addition to Val or Ile, may also confer full affinity toward SUMO; in the second position, only Ile or Val is allowed; the rest of the pentamer, i.e. DLT, is essential (Fig. 3F).
Novel SUMO-binding Proteins with Clustered VIDLT-type SIMs
Our data together with the recent findings by Yuan Chen and co-workers (35), strongly argue that aromatic residues Phe or Tyr may also be allowed in the first position of a pentameric VIDLT-type SIM even though they are rarely seen in reported SIMs. We thus ran another focused string search for proteins containing multiple (V/I/L/F/Y)(V/I)DLT motifs (Fig. 4A, supplemental Script S2 and Table S2). This iteration led to only four hits: RNF4, human FLICE-associated huge protein (FLASH, also known as caspase 8-associated protein 2, CASP8AP2), uncharacterized chromatin 5 open reading frame 25 (C5orf25), and sine oculis-binding protein (SOBP, also known as Jackson circler protein 1, JXL1) (Figs. 4A and supplemental Fig. S5). As with the first string search, our algorithm did not automatically detect clustered SIMs, yet strikingly, the predicted SIMs are all clustered in these four proteins (Fig. 4B and Fig. 5). Thus, both the sequence and the clustered presence of these motifs are highly indicative of important SUMO-binding structures.
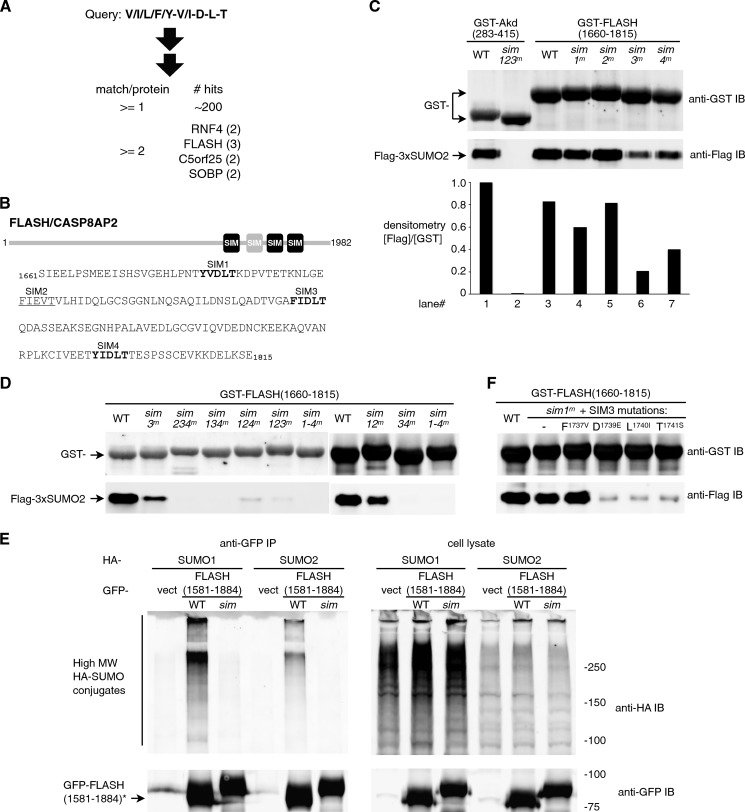
FIGURE 4.
FLASH/CASP8AP2 is a novel SUMO-binding protein with clustered SIMs. A, an alternative string search against (V/I/L/F/Y)(V/I)DLT led to the identification of putative SUMO-binding proteins with clustered VIDLT-type SIMs. B, predicted SIMs in human FLASH/CASP8AP2 (NP_001131140) are shown. Top panel, the relative position of predicted SIMs in FLASH; middle panel, the sequence of the FLASH residues 1661–1815 with the predicted SIMs (1683YVDLT, 1700FIEVT, 1737FIDLT, and 1794YIDLT) highlighted. C, validation of the FLASH SUMO binding domain is shown. FLASH-(1661–1815) and its mutants as indicated were purified as GST fusion proteins and tested for their association with FLAG-3xSUMO2. Analogous to the mutational analysis with Arkadia shown in Fig. 2B, single SIM mutations (sim1m, sim2m, sim3m, sim4m) were generated by replacing all three core hydrophobic residues (Tyr/Phe, Val/Ile, and Leu) with Ala in each of the four putative SIMs. Binding assays were carried out as in Fig. 2C. The GST-Arkadia-(283–415) and its sim123m mutant were used as positive and negative controls, respectively. Dual color immunoblot (anti-GST and anti-FLAG) on a single membrane was processed through the Odyssey Infrared Imaging System. Quantitative densitometry is shown in the bar graph below as the ratio of each FLAG-3xSUMO2 band to its corresponding GST band. IB, immunoblot. D, shown is the contribution of individual SIMs in FLASH SUMO binding domain. The FLASH mutants with only one (sim234m, sim134m, sim124m, sim123m) or none (sim1–4m) of the four SIMs left intact as well as two double SIM mutants (sim12m, sim34m) were tested for SUMO binding as in C. Examples of purified GST-fusion proteins are shown in supplemental Fig. S4. E, shown is the SIM-dependent association between the FLASH SBD and high molecular weight sumoylated proteins in cultured cells. EGFP-tagged fragment of FLASH (residues 1581–1884, either wild-type or sim134m mutant) were coexpressed with HA-tagged SUMO1 or SUMO2 in 293T cells as indicated. Immunoprecipitation (IP) was carried out with anti-GFP antiserum and protein A-agarose beads. Proteins in the precipitated immune complex and in the cell lysate were analyzed in an 8% Tris-glycine SDS-PAGE followed by immunoblotting using anti-GFP and anti-HA antibodies as indicated. The stacking gel was preserved during immunoblotting in order to detect high molecular weight SUMO conjugates that could not enter the resolving gel. vect, EGFP alone; sim, sim134m. *, it is unclear why the sim134m fragment showed a different migration (as also seen with some other SIM mutants in this study); this may reflect a conformational change due to the loss of intact SIMs. F, a precise structure of FLASH SIM3 allows minimum variations for full strength SUMO binding. Single-residue replacements in FLASH SIM3, F1737V, D1739E, L1740I, and T1741S were generated as indicated in a sim1m background. Binding assays were performed as in C and D.
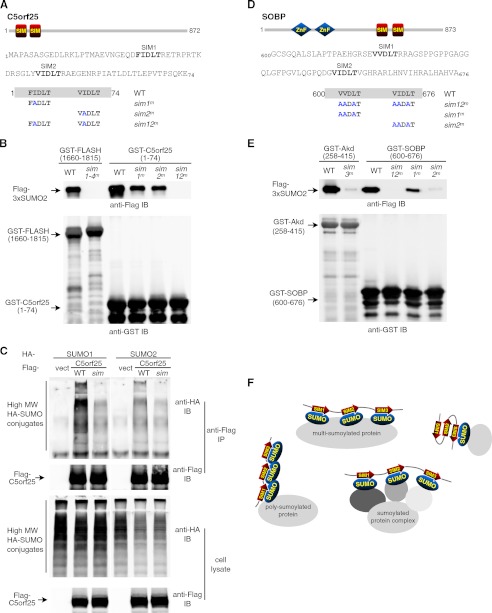
FIGURE 5.
C5orf25 and SOBP each contains an SBD with two VIDLT-type SIMs. A, predicted SIMs in the N terminus of C5orf25 are shown. Top panel, relative position of predicted SIMs in the protein; middle panel, sequence of the N-terminal region (1–74) of C5orf25 with the two SIMs (26FIDLT and 45VIDLT) highlighted; bottom panel, SIM mutants of C5orf25 were generated by replacing a single hydrophobic residue in each or both of the two putative SIMs (sim1m, I27A; sim2m, I46A). B, validation of the C5orf25 SUMO binding domain. C5orf25-(1–74) and its mutants as indicated were purified as GST fusion proteins and tested for their association with FLAG-3xSUMO2. Binding assays were carried out as in Fig. 2C. The GST-FLASH-(1660–1885) and its sim1–4m mutant were used as positive and negative controls, respectively. Examples of purified GST-fusion proteins are shown in supplemental Fig. S4. IB, immunoblots. C, SIM-dependent association between C5orf25 and high molecular weight sumoylated proteins in cultured cells detected through coimmunoprecipitation. FLAG-tagged full-length C5orf25 in indicated forms (WT; sim, sim12m) were coexpressed with HA-tagged SUMO1 or SUMO2 in 293T cells. Immunoprecipitation was carried out with anti-FLAG (M2)-conjugated agarose beads. Proteins in the precipitated immune complex and in the cell lysate were analyzed in an 8% Tris-glycine SDS-PAGE followed by immunoblotting using anti-FLAG and anti-HA antibodies as indicated. The stacking gel was preserved during immunoblotting to detect high molecular weight SUMO conjugates that could not enter the resolving gel. D, shown are predicted SIMs in SOBP. Top panel, the relative position of predicted SIMs in SOBP; middle panel, the sequence of SOBP residues 600–676 with the two SIMs (620VVDLT and 653VIDLT) highlighted; bottom panel, SIM mutants of SOBP were generated by replacing three hydrophobic residue in each or both of the two putative SIMs (sim1m, V620A/V621A/L623A; sim2m, V653A/I654A/L656A). E, SOBP SIMs show differential contributions to SUMO binding. Wild-type and mutant forms of SOBP fragment 600–676 as indicated were purified as GST fusion proteins. The binding assay was performed essentially as described in Fig. 2C. GST-Arkadia-(258–415) and its sim3m mutant (see Fig. 2C) were included as positive and negative controls. F, a diagram illustrating diverse forms of protein sumoylation that can be recognized by SUMO binding domains containing clustered SIMs.
In human FLASH, the string search identified three closely spaced motifs, 1683YVDLT1687, 1737FIDLT1741, and 1794YIDLT1798, designated as SIM1, SIM3, and SIM4 (Fig. 4B). We also designated 1700FIEVT1704 as SIM2 and predicted that SIM2 would serve as a negative control in our validation even though it is similar to the other three perfectly matched motifs. We validated the SUMO binding of these putative SIMs by carrying out SUMO binding assays against tri-SUMO2 using a GST fusion of FLASH-(1661–1815). We found that the wild-type fragment bound to SUMO as robustly as the Arkadia SUMO binding domain. Except for SIM2, mutating SIM1, −3, or 4 alone all caused reduction of SUMO binding to a certain degree, with sim3m resulting in the most severe loss followed by sim4m (Fig. 4C). Next, we tested triple-SIM mutants with only one SIM left intact and found that all triple-SIM mutants showed diminished SUMO association (Fig. 4D), among which sim124m, with an intact SIM3, preserved the most significant amount of residual SUMO affinity, just as predicted by the binding result with single-SIM mutants. Moreover, sim134m and sim234m are incapable of SUMO binding as is the quadruple mutant sim1–4m, indicating that SIM3 and SIM4 together contribute to the majority of the SUMO binding affinity of FLASH-(1661–1815), whereas SIM2 is indeed insignificant as expected. This is further confirmed by the result of two double-SIM mutants, sim12m and sim34m, with the former mutant retaining robust SUMO binding, whereas the latter being completely inactive (Fig. 4D). Last, a slightly larger FLASH fragment (1581–1884) containing the SUMO binding domain could specifically associate with high molecular weight SUMO1 or SUMO2 conjugates in cultured cells in a SIM1/3/4-dependent manner (Fig. 4E). We thus identify FLASH as a novel SUMO-binding protein. Its SBD contains a cluster of three functional SIMs, all with a distinct signature led by the aromatic residues Phe or Tyr.
As with the Arkadia SIM3, we also examined the structural rigidity of the most critical SIM, SIM3 (1737FIDLT1741) in FLASH by testing single-residue mutations (at a smaller scale, though); we replaced the FIDLT pentamer with VIDLT, FIELT, FIDIT, and FIDLS in a sim1m background (with SIM4 kept intact for a proper dynamic range in detecting the SUMO binding). Consistently, we found that whereas the VIDLT mutant was fully functional, the other three mutations resulted in a significantly diminished SIM3 (Fig. 4F). Our data suggest that FIDLT indeed belongs to the VIDLT-type SIMs with a highly conserved atomic structure and confirm that motifs with the (V/I/L/F/Y)(V/I)DLT sequence represent SIMs with high SUMO affinity.
In C5orf25, we designated the two identified motifs, 26FIDLT30 and 45VIDLT49, as SIM1 and SIM2 (Fig. 5A) and tested the GST fusion proteins of C5orf25-(1–74) and its sim1m, sim2m, and sim12m mutants in the tri-SUMO2 binding assays. We found that SIM1 and SIM2 contributed equally and together are essential for SUMO-binding (Fig. 5B). Consistently, the association of full-length C5orf25 with high molecular weight SUMO1 or SUMO2 conjugates in 293T cells required the simultaneous presence of its SIM1 and SIM2, suggesting that C5orf25 contains an N-terminal SBD with two VIDLT-type SIMs (Fig. 5C).
In SOBP, we isolated a 77-amino acid fragment (SOBP 600–676) containing the predicted SIM1 (620VVDLT624) and SIM2 (653VIDLT657) and its mutant forms (sim12m, sim1m, and sim2m) as GST fusion proteins (Fig. 5D). In the SUMO binding assay, essentially parallel to previous ones, we found that the wild-type fragment bound to tri-SUMO in a similar fashion to the Arkadia SBD. Unlike C5orf25, here the two SOBP SIMs appeared to function differently; whereas the simultaneous mutation of both SIM1 and SIM2 abolished SUMO binding, mutating SIM2 resulted in a more significant reduction of SUMO association than mutating SIM1 (Fig. 5E). Together, we conclude that a functional SBD with two VIDLT-type SIMs exists in both C5orf25 and SOBP.
In conclusion, a detailed structural analysis of the Arkadia SBD has led us to the computational identification of additional novel SBDs in FLASH, C5orf25, and SOBP. All three SBDs contain clustered VIDLT-type SIMs, indicating that these SBDs, like the ones in RNF4 and Arkadia, are crucial SUMO binding structures responsible for the recognition of diverse forms of sumoylation (Fig. 5F).
DISCUSSION
SIM Prediction
We have successfully identified novel SUMO-binding proteins using a simple computational search based on a single criterion; that is, the sequence similarity to existing SIMs. As all known structures of SUMO-bound SIMs are β-strands (5, 7, 9, 12), we have used secondary structure prediction to add confidence to our SIM search (supplemental Fig. S1A). Consistently, we note that besides Val and Ile, Thr is also frequently associated with known SIMs, most likely because as β-branched amino acids, Thr, Val, and Ile all have the greatest propensity to form a β-strand (36, 37). Therefore, for future improvements, secondary structure prediction algorithms may be combined with the string search to achieve a more accurate SIM prediction. A more comprehensive protocol for computational SIM prediction has also been described recently (38). In either case, it may be difficult to balance accuracy and thoroughness in such a motif scan. For example, without filtering, certain giant proteins such as Titin were found in our search presumably because of the abundance of β-strands in their structure. As another example, the two SIM-like motifs found in small CTD phosphatase 3 (supplemental Table S1) are buried inside the protein according to its crystal structure (PDB 2HHL) (39) and are, therefore, unlikely to be exposed for SUMO interaction unless the protein is unfolded. Above all, a SIM prediction is more meaningful when closely tied to careful experimental validation.
Various Flavors of SIM Sequences
The three SIMs in Arkadia come in different flavors: SIM1 (VVVIE) resembles RNF4 SIM4 (VVIVD), SIM2 (VEIVTV) is similar to RNF4 SIM1 (IELVET), and SIM3 (VVDLT) is almost identical to both SIM2 (IVDLT) and SIM3 (VVDLT) in RNF4. Arkadia SIM1 and SIM3 also match the previous designation of SIM subtypes, SIM-a and SIM-b, respectively, whereas SIM2 appears to be rather a bidirectional SIM than a pure SIM-r (40, 41). Differences in sequence may contribute to the selectivity of individual SIMs toward different SUMO isoforms (5, 35). Consistently, some of our Arkadia SIM3 mutants showed better residual interaction with SUMO1 than with SUMO2 (Fig. 3). On the other hand, with the intact Arkadia or RNF4 SBD, we did not observe any binding preference toward either SUMO1 or SUMO2 (Figs. 2 and 3, and supplemental Fig. S2A), indicating that the Arkadia SIMs together can recognize both SUMO1 and SUMO2-conjugated proteins. Moreover, the dominant SIMs in both RNF4 and Arkadia are the VIDLT-type (or the SIM-b subtype) SIMs that are also found in the PIAS SIMs, which display a high affinity to both SUMO1 and SUMO2/3 (Figs. 2 and 3) (5, 35), again suggesting that clustered SIMs with a VIDLT-type SIM can potently target both SUMO1 and SUMO2/3 isoforms.
It is striking that the VIDLT pentamer maintains a rigid structure; even a subtle variation would lower its affinity for SUMO, except for the first position where it can accommodate aromatic residues Phe and Tyr (Fig. 3) (35). This indicates that not all SIMs (or the permutations of SIM consensus sequence) are equal; some of our single-residue mutations in Arkadia SIM3 resulted in dramatic reduction of SUMO binding yet still matched the SIM consensus sequence. We believe that the VIDLT-type SIMs make the best fitted structure for the interaction with all SUMO isoforms and are likely preserved as a key feature of the sumoylation system during evolution (35). Likewise, chemical mimetics of VIDLT may disrupt overall SUMO-SIM interaction and impair the activity of sumoylation essential for living cells, e.g. during their recovery from genotoxic stress (37). We further speculate that the dominant SIMs drive a “zipping-up” process during the recognition between clustered SIMs and multiple SUMO moieties, which likely initiates in a dynamic manner before adopting a stable configuration.
Significance of Clustered SIMs
The tandem array of SIMs in RNF4 family proteins is thought to specifically recognize polySUMO chains (20). It is worth noting that the spacing from SIM1 to SIM2 in Rfp1 or Rfp2 is ∼20 amino acids because a similar spacing is also found between the tandem ubiquitin-interacting motifs in a number of polyubiquitin-binding proteins (42). This is also about the same distance between RNF4′s SIM1 and SIM3 or its SIM2 and SIM4 and also between the SIM1 and SIM2 in C5orf25. This particular spacing may contribute to the optimal recognition of polySUMO chains, as with the tandem ubiquitin-interacting motifs for polyubiquitin chains (42, 43). Although a structural model is lacking for the interaction between a SIM cluster and a polySUMO chain, one can speculate that two adjacent SUMO units form physical contacts with either the SIM1/3 or the SIM2/4 pair in a stable RNF4-polySUMO chain complex.
Besides polySUMO interaction, clustered SIMs may also target other forms of sumoylation, such as a protein or protein complex that is monosumoylated at multiple sites (Fig. 5F). The SBDs in Arkadia and FLASH may be particularly fitted for this kind of SUMO binding, where the SIMs are further separated and presumably more flexible. The avidity effect due to multiple SIM-SUMO contacts may thus provide sufficient affinity and specificity for the formation of sumoylation-regulated protein complexes, which would be distinct from those involving a singular SIM and often a secondary, SUMO-independent physical contact (44, 45). Furthermore, it is also conceivable that clustered SIMs form a folded β-sheet (Fig. 5F). It then follows that there may exist a dynamic balance between a folded, stand-alone β-sheet and an open, less ordered conformation with multiple extended SIMs in complex with poly/multi-SUMO. Given its unique structure, we propose that a SIM cluster or a tandem array of closely located SIMs be designated as an SBD. More detailed structural and functional characterization will be needed for us to understand how SIM-containing proteins interpret the SUMO signal.
Potential Biological Function of SBDs
Arkadia is known to act as a RING domain ubiquitin ligase to promote transcriptional activation downstream of the TGFβ signaling pathway, especially during early embryonic development in vertebrates (46–49). Arkadia may also influence the activity of other signaling pathways, as Arkadia was found to interact with Axin1, a canonical component of the Wnt pathway (50). The identification of Arkadia as a SUMO-binding protein in the TGFβ pathway raises the possibility that Arkadia may interact with a sumoylated protein specific to this signaling pathway, such as Smad4 or SnoN (51–54), or with a poly- or multisumoylated transcriptional repressor complex (6, 55). We will describe in a separate paper that the combined activity of SUMO binding and RING domains in Arkadia provides a novel SUMO-targeted ubiquitin ligase specific to the TGFβ pathway.4
The SUMO binding domain of FLASH is in a region previously found to interact with the tandem death effector domains (DEDs) of procaspase-8 and was named “DED-related domain” (DRD) (56). However, the sequence similarity between FLASH and CED-4/Apaf-1, its alleged functional homologs, has been disputed (57). Thus, the precise role of FLASH in apoptosis is still unclear. We note that Ubc9, the SUMO-conjugating enzyme (E2), was identified in addition to FLASH in the initial yeast two-hybrid screen against the procaspase-8 DEDs and that the FLASH SBD itself has also been shown to form a yeast two-hybrid interaction with Ubc9 and PIAS1 (56, 58, 59). It may not be coincidental that both SUMO1 itself (also known as Sentrin) and TTRAP/TDP2 and Daxx, two SIM-containing proteins, were initially found as death domain-binding proteins also through yeast two-hybrid screens against death receptors Fas/CD95 or TNFR/CD40 (5, 19, 60–63). As both death domain and DED belong to the death domain superfamily and form oligomers (78), perhaps using an oligomerization domain as bait in a yeast two-hybrid screen is prone to hitting the SUMO pathway components and SUMO-binding proteins. This raises an intriguing possibility that protein oligomerization triggers sumoylation and may also explain the early identification of Ubc9 (UbE2I) and SUMO1 from a two-hybrid screen against Rad52, a protein also forming homotypic polymers (64–67). More recently, FLASH was shown to localize to Cajal bodies and function in transcriptional control or RNA processing (68–71). It is conceivable that the SBD in FLASH contributes to sumoylation-regulated assembly of nuclear complexes such as Cajal bodies or is responsible for the communication between Cajal body and sumoylated nuclear entities such as PML bodies (3, 70, 72).
SOBP is a nuclear zinc finger protein whose molecular functions are largely unknown. Recessive mutations of mouse Sobp gene cause defective patterning of sensory epithelium and deformed organ of Corti in the inner ear, resulting in a deafness phenotype known as Jackson circular (jc) (73). An SOBP truncating mutation lying immediately after the two SIMs has also been associated with defects in the development of the nervous system in humans (74). SOBP may act as a critical transcription factor for neural development in mammals. Sumoylation is well known for its role in transcriptional repression through modification of certain transcription factors and SIM-mediated assembly of nuclear protein complex (55). Future studies will resolve how sumoylation regulates the function of SOBP through its SUMO binding domain.
In summary, we have found a number of SUMO-binding proteins with unique SIM compositions through a data mining approach. In contrast to most SUMO-binding proteins, which often have a singular, stand-alone SIM, the proteins described here all contain SUMO binding domains with clustered SIMs and are likely crucial components for SUMO recognition in diverse biological processes.
Acknowledgments
We thank Professor Odd S. Gabrielsen (University of Oslo) for the human FLASH cDNA. We thank members of the Hunter laboratory for helpful discussions, Yijing Liu and Kate Wright for technical assistance in molecular cloning, Suzy Simon for assistance in picking Mammalian Gene Collection cDNA clones, and Jill Meisenhelder for the anti-GST/anti-GFP bivalent antiserum.
This work was supported, in whole or in part, by National Institutes of Health Grants CA14195, CA80100, and CA82683 (to T. H.).

This article contains supplemental Figs. S1–S5, Tables S1 and S2, and Scripts 1 and 2.
H. Sun, Y. Liu, and T. Hunter, manuscript in preparation.
- SUMO
- small ubiquitin-like modifier
- SIM
- SUMO-interacting motif
- SBD
- SUMO-binding domain
- PML
- promyelocytic leukemia protein
- SOBP
- sine oculis-binding protein
- DED
- death effector domains.
REFERENCES
- 1. Creton S., Jentsch S. (2010) SnapShot. The SUMO system. Cell 143, 848–848 [DOI] [PubMed] [Google Scholar]
- 2. Meulmeester E., Melchior F. (2008) Cell biology. SUMO. Nature 452, 709–711 [DOI] [PubMed] [Google Scholar]
- 3. Matunis M. J., Zhang X. D., Ellis N. A. (2006) SUMO. The glue that binds. Dev. Cell 11, 596–597 [DOI] [PubMed] [Google Scholar]
- 4. Minty A., Dumont X., Kaghad M., Caput D. (2000) Covalent modification of p73α by SUMO-1. Two-hybrid screening with p73 identifies novel SUMO-1-interacting proteins and a SUMO-1 interaction motif. J. Biol. Chem. 275, 36316–36323 [DOI] [PubMed] [Google Scholar]
- 5. Hecker C. M., Rabiller M., Haglund K., Bayer P., Dikic I. (2006) Specification of SUMO1- and SUMO2-interacting motifs. J. Biol. Chem. 281, 16117–16127 [DOI] [PubMed] [Google Scholar]
- 6. Ouyang J., Shi Y., Valin A., Xuan Y., Gill G. (2009) Direct binding of CoREST1 to SUMO-2/3 contributes to gene-specific repression by the LSD1/CoREST1/HDAC complex. Mol. Cell 34, 145–154 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Song J., Durrin L. K., Wilkinson T. A., Krontiris T. G., Chen Y. (2004) Identification of a SUMO binding motif that recognizes SUMO-modified proteins. Proc. Natl. Acad. Sci. U.S.A. 101, 14373–14378 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Seet B. T., Dikic I., Zhou M. M., Pawson T. (2006) Reading protein modifications with interaction domains. Nat. Rev. Mol. Cell Biol. 7, 473–483 [DOI] [PubMed] [Google Scholar]
- 9. Reverter D., Lima C. D. (2005) Insights into E3 ligase activity revealed by a SUMO-RanGAP1-Ubc9-Nup358 complex. Nature 435, 687–692 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Yang S. H., Sharrocks A. D. (2010) The SUMO E3 ligase activity of Pc2 is coordinated through a SUMO interaction motif. Mol. Cell Biol. 30, 2193–2205 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Zhu J., Zhu S., Guzzo C. M., Ellis N. A., Sung K. S., Choi C. Y., Matunis M. J. (2008) Small ubiquitin-related modifier (SUMO) binding determines substrate recognition and paralog-selective SUMO modification. J. Biol. Chem. 283, 29405–29415 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Baba D., Maita N., Jee J. G., Uchimura Y., Saitoh H., Sugasawa K., Hanaoka F., Tochio H., Hiroaki H., Shirakawa M. (2005) Crystal structure of thymine DNA glycosylase conjugated to SUMO-1. Nature 435, 979–982 [DOI] [PubMed] [Google Scholar]
- 13. Hardeland U., Steinacher R., Jiricny J., Schär P. (2002) Modification of the human thymine-DNA glycosylase by ubiquitin-like proteins facilitates enzymatic turnover. EMBO J. 21, 1456–1464 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Takahashi H., Hatakeyama S., Saitoh H., Nakayama K. I. (2005) Noncovalent SUMO-1 binding activity of thymine DNA glycosylase (TDG) is required for its SUMO-1 modification and colocalization with the promyelocytic leukemia protein. J. Biol. Chem. 280, 5611–5621 [DOI] [PubMed] [Google Scholar]
- 15. Boddy M. N., Howe K., Etkin L. D., Solomon E., Freemont P. S. (1996) PIC 1, a novel ubiquitin-like protein which interacts with the PML component of a multiprotein complex that is disrupted in acute promyelocytic leukaemia. Oncogene 13, 971–982 [PubMed] [Google Scholar]
- 16. Shen T. H., Lin H. K., Scaglioni P. P., Yung T. M., Pandolfi P. P. (2006) The mechanisms of PML-nuclear body formation. Mol. Cell 24, 331–339 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Häkli M., Karvonen U., Jänne O. A., Palvimo J. J. (2005) SUMO-1 promotes association of SNURF (RNF4) with PML nuclear bodies. Exp. Cell Res. 304, 224–233 [DOI] [PubMed] [Google Scholar]
- 18. Lallemand-Breitenbach V., Jeanne M., Benhenda S., Nasr R., Lei M., Peres L., Zhou J., Zhu J., Raught B., de Thé H. (2008) Arsenic degrades PML or PML-RARα through a SUMO-triggered RNF4/ubiquitin-mediated pathway. Nat. Cell Biol. 10, 547–555 [DOI] [PubMed] [Google Scholar]
- 19. Lin D. Y., Huang Y. S., Jeng J. C., Kuo H. Y., Chang C. C., Chao T. T., Ho C. C., Chen Y. C., Lin T. P., Fang H. I., Hung C. C., Suen C. S., Hwang M. J., Chang K. S., Maul G. G., Shih H. M. (2006) Role of SUMO-interacting motif in Daxx SUMO modification, subnuclear localization, and repression of sumoylated transcription factors. Mol. Cell 24, 341–354 [DOI] [PubMed] [Google Scholar]
- 20. Tatham M. H., Geoffroy M. C., Shen L., Plechanovova A., Hattersley N., Jaffray E. G., Palvimo J. J., Hay R. T. (2008) RNF4 is a poly-SUMO-specific E3 ubiquitin ligase required for arsenic-induced PML degradation. Nat. Cell Biol. 10, 538–546 [DOI] [PubMed] [Google Scholar]
- 21. Sun H., Leverson J. D., Hunter T. (2007) Conserved function of RNF4 family proteins in eukaryotes. Targeting a ubiquitin ligase to SUMOylated proteins. EMBO J. 26, 4102–4112 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Bruderer R., Tatham M. H., Plechanovova A., Matic I., Garg A. K., Hay R. T. (2011) Purification and identification of endogenous polySUMO conjugates. EMBO Rep. 12, 142–148 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Ulrich H. D. (2008) The fast-growing business of SUMO chains. Mol. Cell 32, 301–305 [DOI] [PubMed] [Google Scholar]
- 24. Liew C. W., Sun H., Hunter T., Day C. L. (2010) RING domain dimerization is essential for RNF4 function. Biochem. J. 431, 23–29 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Plechanovová A., Jaffray E. G., McMahon S. A., Johnson K. A., Navrátilová I., Naismith J. H., Hay R. T. (2011) Mechanism of ubiquitylation by dimeric RING ligase RNF4. Nat. Struct. Mol. Biol. 18, 1052–1059 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Yaffe M. B., Leparc G. G., Lai J., Obata T., Volinia S., Cantley L. C. (2001) A motif-based profile scanning approach for genome-wide prediction of signaling pathways. Nat. Biotechnol. 19, 348–353 [DOI] [PubMed] [Google Scholar]
- 27. Weatheritt R. J., Gibson T. J. (2012) Linear motifs: lost in (pre)translation. Trends Biochem. Sci. 37, 333–341 [DOI] [PubMed] [Google Scholar]
- 28. Cho G., Lim Y., Golden J. A. (2009) SUMO interaction motifs in Sizn1 are required for promyelocytic leukemia protein nuclear body localization and for transcriptional activation. J. Biol. Chem. 284, 19592–19600 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Ogawa H., Komatsu T., Hiraoka Y., Morohashi K. (2009) Transcriptional suppression by transient recruitment of ARIP4 to sumoylated nuclear receptor Ad4BP/SF-1. Mol. Biol. Cell 20, 4235–4245 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Uwada J., Tanaka N., Yamaguchi Y., Uchimura Y., Shibahara K., Nakao M., Saitoh H. (2010) The p150 subunit of CAF-1 causes association of SUMO2/3 with the DNA replication foci. Biochem. Biophys. Res. Commun. 391, 407–413 [DOI] [PubMed] [Google Scholar]
- 31. Tatham M. H., Jaffray E., Vaughan O. A., Desterro J. M., Botting C. H., Naismith J. H., Hay R. T. (2001) Polymeric chains of SUMO-2 and SUMO-3 are conjugated to protein substrates by SAE1/SAE2 and Ubc9. J. Biol. Chem. 276, 35368–35374 [DOI] [PubMed] [Google Scholar]
- 32. Matic I., van Hagen M., Schimmel J., Macek B., Ogg S. C., Tatham M. H., Hay R. T., Lamond A. I., Mann M., Vertegaal A. C. (2008) In vivo identification of human small ubiquitin-like modifier polymerization sites by high accuracy mass spectrometry and an in vitro to in vivo strategy. Mol. Cell. Proteomics 7, 132–144 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Matic I., Schimmel J., Hendriks I. A., van Santen M. A., van de Rijke F., van Dam H., Gnad F., Mann M., Vertegaal A. C. (2010) Site-specific identification of SUMO-2 targets in cells reveals an inverted SUMOylation motif and a hydrophobic cluster SUMOylation motif. Mol. Cell 39, 641–652 [DOI] [PubMed] [Google Scholar]
- 34. Tanaka N., Saitoh H. (2010) A real-time SUMO binding assay for the analysis of the SUMO-SIM protein interaction network. Biosci. Biotechnol. Biochem. 74, 1302–1305 [DOI] [PubMed] [Google Scholar]
- 35. Namanja A. T., Li Y. J., Su Y., Wong S., Lu J., Colson L. T., Wu C., Li S. S., Chen Y. (2012) Insights into high affinity small ubiquitin-like modifier (SUMO) recognition by SUMO-interacting motifs (SIMs) revealed by a combination of NMR and peptide array analysis. J. Biol. Chem. 287, 3231–3240 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Street A. G., Mayo S. L. (1999) Intrinsic β-sheet propensities result from van der Waals interactions between side chains and the local backbone. Proc. Natl. Acad. Sci. U.S.A. 96, 9074–9076 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Li Y. J., Perkins A. L., Su Y., Ma Y., Colson L., Horne D. A., Chen Y. (2012) Gold nanoparticles as a platform for creating a multivalent poly-SUMO chain inhibitor that also augments ionizing radiation. Proc. Natl. Acad. Sci. U.S.A. 109, 4092–4097 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Vogt B., Hofmann K. (2012) Bioinformatical detection of recognition factors for ubiquitin and SUMO. Methods Mol. Biol. 832, 249–261 [DOI] [PubMed] [Google Scholar]
- 39. Almo S. C., Bonanno J. B., Sauder J. M., Emtage S., Dilorenzo T. P., Malashkevich V., Wasserman S. R., Swaminathan S., Eswaramoorthy S., Agarwal R., Kumaran D., Madegowda M., Ragumani S., Patskovsky Y., Alvarado J., Ramagopal U. A., Faber-Barata J., Chance M. R., Sali A., Fiser A., Zhang Z. Y., Lawrence D. S., Burley S. K. (2007) Structural genomics of protein phosphatases. J. Struct. Funct. Genomics 8, 121–140 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Song J., Zhang Z., Hu W., Chen Y. (2005) Small ubiquitin-like modifier (SUMO) recognition of a SUMO binding motif. A reversal of the bound orientation. J. Biol. Chem. 280, 40122–40129 [DOI] [PubMed] [Google Scholar]
- 41. Miteva M., Keusekotten K., Hofmann K., Praefcke G. J., Dohmen R. J. (2010) Sumoylation as a signal for polyubiquitylation and proteasomal degradation. Subcell. Biochem. 54, 195–214 [DOI] [PubMed] [Google Scholar]
- 42. Sims J. J., Cohen R. E. (2009) Linkage-specific avidity defines the lysine 63-linked polyubiquitin-binding preference of rap80. Mol. Cell 33, 775–783 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Song A. X., Zhou C. J., Peng Y., Gao X. C., Zhou Z. R., Fu Q. S., Hong J., Lin D. H., Hu H. Y. (2010) Structural transformation of the tandem ubiquitin-interacting motifs in ataxin-3 and their cooperative interactions with ubiquitin chains. PLoS ONE 5, e13202. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Armstrong A. A., Mohideen F., Lima C. D. (2012) Recognition of SUMO-modified PCNA requires tandem receptor motifs in Srs2. Nature 483, 59–63 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Bergink S., Jentsch S. (2009) Principles of ubiquitin and SUMO modifications in DNA repair. Nature 458, 461–467 [DOI] [PubMed] [Google Scholar]
- 46. Episkopou V., Arkell R., Timmons P. M., Walsh J. J., Andrew R. L., Swan D. (2001) Induction of the mammalian node requires Arkadia function in the extraembryonic lineages. Nature 410, 825–830 [DOI] [PubMed] [Google Scholar]
- 47. Koinuma D., Shinozaki M., Komuro A., Goto K., Saitoh M., Hanyu A., Ebina M., Nukiwa T., Miyazawa K., Imamura T., Miyazono K. (2003) Arkadia amplifies TGF-β superfamily signalling through degradation of Smad7. EMBO J. 22, 6458–6470 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48. Levy L., Howell M., Das D., Harkin S., Episkopou V., Hill C. S. (2007) Arkadia activates Smad3/Smad4-dependent transcription by triggering signal-induced SnoN degradation. Mol. Cell Biol. 27, 6068–6083 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49. Niederländer C., Walsh J. J., Episkopou V., Jones C. M. (2001) Arkadia enhances nodal-related signalling to induce mesendoderm. Nature 410, 830–834 [DOI] [PubMed] [Google Scholar]
- 50. Liu W., Rui H., Wang J., Lin S., He Y., Chen M., Li Q., Ye Z., Zhang S., Chan S. C., Chen Y. G., Han J., Lin S. C. (2006) Axin is a scaffold protein in TGF-β signaling that promotes degradation of Smad7 by Arkadia. EMBO J. 25, 1646–1658 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51. Hsu Y. H., Sarker K. P., Pot I., Chan A., Netherton S. J., Bonni S. (2006) Sumoylated SnoN represses transcription in a promoter-specific manner. J. Biol. Chem. 281, 33008–33018 [DOI] [PubMed] [Google Scholar]
- 52. Lee P. S., Chang C., Liu D., Derynck R. (2003) Sumoylation of Smad4, the common Smad mediator of transforming growth factor-β family signaling. J. Biol. Chem. 278, 27853–27863 [DOI] [PubMed] [Google Scholar]
- 53. Lin X., Liang M., Liang Y. Y., Brunicardi F. C., Feng X. H. (2003) SUMO-1/Ubc9 promotes nuclear accumulation and metabolic stability of tumor suppressor Smad4. J. Biol. Chem. 278, 31043–31048 [DOI] [PubMed] [Google Scholar]
- 54. Lin X., Liang M., Liang Y. Y., Brunicardi F. C., Melchior F., Feng X. H. (2003) Activation of transforming growth factor-β signaling by SUMO-1 modification of tumor suppressor Smad4/DPC4. J. Biol. Chem. 278, 18714–18719 [DOI] [PubMed] [Google Scholar]
- 55. Gill G. (2005) Something about SUMO inhibits transcription. Curr. Opin. Genet. Dev. 15, 536–541 [DOI] [PubMed] [Google Scholar]
- 56. Imai Y., Kimura T., Murakami A., Yajima N., Sakamaki K., Yonehara S. (1999) The CED-4-homologous protein FLASH is involved in Fas-mediated activation of caspase-8 during apoptosis. Nature 398, 777–785 [DOI] [PubMed] [Google Scholar]
- 57. Koonin E. V., Aravind L., Hofmann K., Tschopp J., Dixit V. M. (1999) Apoptosis. Searching for FLASH domains. Nature 401, 662–663 [DOI] [PubMed] [Google Scholar]
- 58. Alm-Kristiansen A. H., Lorenzo P. I., Molvaersmyr A. K., Matre V., Ledsaak M., Saether T., Gabrielsen O. S. (2011) PIAS1 interacts with FLASH and enhances its co-activation of c-Myb. Mol. Cancer 10, 21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59. Alm-Kristiansen A. H., Norman I. L., Matre V., Gabrielsen O. S. (2009) SUMO modification regulates the transcriptional activity of FLASH. Biochem. Biophys. Res. Commun. 387, 494–499 [DOI] [PubMed] [Google Scholar]
- 60. Okura T., Gong L., Kamitani T., Wada T., Okura I., Wei C. F., Chang H. M., Yeh E. T. (1996) Protection against Fas/APO-1- and tumor necrosis factor-mediated cell death by a novel protein, sentrin. J. Immunol. 157, 4277–4281 [PubMed] [Google Scholar]
- 61. Ryu S. W., Chae S. K., Kim E. (2000) Interaction of Daxx, a Fas binding protein, with sentrin and Ubc9. Biochem. Biophys. Res. Commun. 279, 6–10 [DOI] [PubMed] [Google Scholar]
- 62. Yang X., Khosravi-Far R., Chang H. Y., Baltimore D. (1997) Daxx, a novel Fas-binding protein that activates JNK and apoptosis. Cell 89, 1067–1076 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63. Pype S., Declercq W., Ibrahimi A., Michiels C., Van Rietschoten J. G., Dewulf N., de Boer M., Vandenabeele P., Huylebroeck D., Remacle J. E. (2000) TTRAP, a novel protein that associates with CD40, tumor necrosis factor (TNF) receptor-75, and TNF receptor-associated factors (TRAFs) and that inhibits nuclear factor-κB activation. J. Biol. Chem. 275, 18586–18593 [DOI] [PubMed] [Google Scholar]
- 64. Singleton M. R., Wentzell L. M., Liu Y., West S. C., Wigley D. B. (2002) Structure of the single-strand annealing domain of human RAD52 protein. Proc. Natl. Acad. Sci. U.S.A. 99, 13492–13497 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65. Kagawa W., Kurumizaka H., Ishitani R., Fukai S., Nureki O., Shibata T., Yokoyama S. (2002) Crystal structure of the homologous-pairing domain from the human Rad52 recombinase in the undecameric form. Mol. Cell 10, 359–371 [DOI] [PubMed] [Google Scholar]
- 66. Shen Z., Pardington-Purtymun P. E., Comeaux J. C., Moyzis R. K., Chen D. J. (1996) Associations of UBE2I with RAD52, UBL1, p53, and RAD51 proteins in a yeast two-hybrid system. Genomics 37, 183–186 [DOI] [PubMed] [Google Scholar]
- 67. Shen Z., Pardington-Purtymun P. E., Comeaux J. C., Moyzis R. K., Chen D. J. (1996) UBL1, a human ubiquitin-like protein associating with human RAD51/RAD52 proteins. Genomics 36, 271–279 [DOI] [PubMed] [Google Scholar]
- 68. Alm-Kristiansen A. H., Saether T., Matre V., Gilfillan S., Dahle O., Gabrielsen O. S. (2008) FLASH acts as a co-activator of the transcription factor c-Myb and localizes to active RNA polymerase II foci. Oncogene 27, 4644–4656 [DOI] [PubMed] [Google Scholar]
- 69. Barcaroli D., Bongiorno-Borbone L., Terrinoni A., Hofmann T. G., Rossi M., Knight R. A., Matera A. G., Melino G., De Laurenzi V. (2006) FLASH is required for histone transcription and S-phase progression. Proc. Natl. Acad. Sci. U.S.A. 103, 14808–14812 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70. Barcaroli D., Dinsdale D., Neale M. H., Bongiorno-Borbone L., Ranalli M., Munarriz E., Sayan A. E., McWilliam J. M., Smith T. M., Fava E., Knight R. A., Melino G., De Laurenzi V. (2006) FLASH is an essential component of Cajal bodies. Proc. Natl. Acad. Sci. U.S.A. 103, 14802–14807 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71. Yang X. C., Burch B. D., Yan Y., Marzluff W. F., Dominski Z. (2009) FLASH, a proapoptotic protein involved in activation of caspase-8, is essential for 3' end processing of histone pre-mRNAs. Mol. Cell 36, 267–278 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72. Milovic-Holm K., Krieghoff E., Jensen K., Will H., Hofmann T. G. (2007) FLASH links the CD95 signaling pathway to the cell nucleus and nuclear bodies. EMBO J. 26, 391–401 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73. Chen Z., Montcouquiol M., Calderon R., Jenkins N. A., Copeland N. G., Kelley M. W., Noben-Trauth K. (2008) Jxc1/Sobp, encoding a nuclear zinc finger protein, is critical for cochlear growth, cell fate, and patterning of the organ of corti. J. Neurosci. 28, 6633–6641 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74. Birk E., Har-Zahav A., Manzini C. M., Pasmanik-Chor M., Kornreich L., Walsh C. A., Noben-Trauth K., Albin A., Simon A. J., Colleaux L., Morad Y., Rainshtein L., Tischfield D. J., Wang P., Magal N., Maya I., Shoshani N., Rechavi G., Gothelf D., Maydan G., Shohat M., Basel-Vanagaite L. (2010) SOBP is mutated in syndromic and nonsyndromic intellectual disability and is highly expressed in the brain limbic system. Am. J. Hum. Genet. 87, 694–700 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75. Uchimura Y., Ichimura T., Uwada J., Tachibana T., Sugahara S., Nakao M., Saitoh H. (2006) Involvement of SUMO modification in MBD1- and MCAF1-mediated heterochromatin formation. J. Biol. Chem. 281, 23180–23190 [DOI] [PubMed] [Google Scholar]
- 76. Yousef A. F., Fonseca G. J., Pelka P., Ablack J. N., Walsh C., Dick F. A., Bazett-Jones D. P., Shaw G. S., Mymryk J. S. (2010) Identification of a molecular recognition feature in the E1A oncoprotein that binds the SUMO conjugase UBC9 and likely interferes with polySUMOylation. Oncogene 29, 4693–4704 [DOI] [PubMed] [Google Scholar]
- 77. Meulmeester E., Kunze M., Hsiao H. H., Urlaub H., Melchior F. (2008) Mechanism and consequences for paralog-specific sumoylation of ubiquitin-specific protease 25. Mol. Cell 30, 610–619 [DOI] [PubMed] [Google Scholar]
- 78. Park H. H., Lo Y. C., Lin S. C., Wang L., Yang J. K., Wu H. (2007) The death domain superfamily in intracellular signaling of apoptosis and inflammation. Annu. Rev. Immunol. 25, 561–586 [DOI] [PMC free article] [PubMed] [Google Scholar]