Abstract
All peroxisomal proteins are nuclear encoded, synthesized on free cytosolic ribosomes, and posttranslationally targeted to the organelle. We have used an in vitro assay to reconstitute protein import into pumpkin (Cucurbita pepo) glyoxysomes, a class of peroxisome found in the cotyledons of oilseed plants, to study the mechanisms involved in protein transport across peroxisome membranes. Results indicate that ATP hydrolysis is required for protein import into peroxisomes; nonhydrolyzable analogs of ATP could not substitute for this requirement. Nucleotide competition studies suggest that there may be a nucleotide binding site on a component of the translocation machinery. Peroxisomal protein import also was supported by GTP hydrolysis. Nonhydrolyzable analogs of GTP did not substitute in this process. Experiments to determine the cation specificity of the nucleotide requirement show that the Mg2+ salt was preferred over other divalent and monovalent cations. The role of a putative protonmotive force across the peroxisomal membrane was also examined. Although low concentrations of ionophores had no effect on protein import, relatively high concentrations of all ionophores tested consistently reduced the level of protein import by approximately 50%. This result suggests that a protonmotive force is not absolutely required for peroxisomal protein import.
Peroxisomes are ubiquitous organelles involved in a variety of important cellular processes, including the degradation of hydrogen peroxide and the β-oxidation of fatty acids (for review, see Olsen and Harada, 1995). The single peroxisomal membrane surrounds a dense matrix, sometimes containing paracrystalline inclusions (Frederick et al., 1975). New organelles are thought to arise by the fission of preexisting peroxisomes, with the subsequent incorporation of additional membrane lipids and peroxisomal proteins. There are several classes of peroxisomes found in higher plants (Kindl, 1992; Gietl, 1996). Glyoxysomes are abundant in the cotyledons of most plants and are involved in lipid mobilization to provide nutrients during germination and seedling growth (Trelease, 1984). In leaves, peroxisomes contain enzymes such as GLO that are necessary for photorespiration (Ogren, 1984).
All peroxisomal proteins must be nuclear encoded because peroxisomes do not contain DNA. Peroxisomal proteins are synthesized on free, cytosolic ribosomes and imported posttranslationally into the organelle. The majority of matrix proteins, including GLO, are targeted to peroxisomes via a carboxyl-terminal tripeptide comprising the amino acids Ser-Lys-Leu or conserved variants (Gould et al., 1987; McNew and Goodman, 1996). Other peroxisomal proteins, such as malate dehydrogenase and thiolase, use an amino-terminal targeting signal (Gietl, 1990; Swinkels et al., 1991; Preisig-Müller and Kindl, 1993). Considerable genetic and biochemical evidence indicates the involvement of a proteinaceous receptor in peroxisomal protein import (Wolins and Donaldson, 1994; Rachubinski and Subramani, 1995; Dodt and Gould, 1996; McNew and Goodman, 1996; Brickner et al., 1997). However, a detailed description of the targeting mechanism and an understanding of receptor localization remain elusive. Protein import into yeast peroxisomes may also use other proteins (Dodt and Gould, 1996), including a cytosolic ATPase (Yahraus et al., 1996), a membrane-bound docking factor (Elgersma et al., 1996; Erdmann and Blobel, 1996; Gould et al., 1996), an N-ethylmaleimide-sensitive factor (Wendland and Subramani, 1993), and a molecular chaperone such as hsp70 (Walton et al., 1994).
Protein trafficking requires the investment of energy. The hydrolysis of ATP and/or GTP is required for protein transport into mitochondria (Pfanner et al., 1990), chloroplasts (Keegstra et al., 1989; Theg et al., 1989; Kessler et al., 1994), nuclei (Powers and Forbes, 1994), and the ER (Walter and Johnson, 1994). ATP hydrolysis is known to be necessary for peroxisomal protein import (Imanaka et al., 1987; Wendland and Subramani, 1993; Horng et al., 1995; Brickner et al., 1997). A careful analysis of the energy requirements for protein binding to chloroplasts revealed that other NTPs can support this process, although less efficiently than ATP (Olsen et al., 1989). Exactly where and how the energy is being used during translocation is currently unknown. It is likely that NTP hydrolysis induces conformational changes in the translocation machinery or in the targeted protein (perhaps by interaction with an energy-dependent molecular chaperone), thereby facilitating the protein's entry into the organelle (Pfanner et al., 1990).
A PMF is required, in addition to ATP, for the export of proteins from bacteria (Yamane et al., 1987; Wong and Buckley, 1989); a pH gradient is needed to transport some proteins across thylakoid membranes (Cline et al., 1992; Theg and Scott, 1993); and the electrical component of the PMF facilitates the import of some proteins into mitochondria (Pfanner and Neupert, 1986). Addition of inhibitors or ionophores that collapse the PMF abolishes protein transport across each of these membranes. A PMF may exist across the peroxisomal membrane. Some researchers have found that ionophores inhibit protein import into peroxisomes (Bellion and Goodman, 1987), whereas others observed no effect on peroxisomal protein import (Imanaka et al., 1987; Wendland and Subramani, 1993). An ATPase, analogous to the V-class H+-ATPase found on vacuolar membranes, may be present on the peroxisomal membrane (Douma et al., 1987; del Valle et al., 1988; Wolvetang et al., 1990; Whitney and Bellion, 1991). ATP hydrolysis on the cytosolic face would make the peroxisomal matrix acidic (Nicolay et al., 1987), thus establishing a PMF that could be used to drive protein translocation. Alternatively, nonspecific pores may allow small ions and metabolites to diffuse freely across the membrane, thereby dissipating an electrochemical or pH gradient (for review, see van den Bosch et al., 1992).
We have extensively characterized the energy requirements for protein import into glyoxysomes using an optimized in vitro assay to reconstitute the transport event. First, we examined the ability of various NTPs to support the import of GLO into isolated pumpkin (Cucurbita pepo) glyoxysomes. The competence of nonhydrolyzable analogs of ATP and GTP to substitute for the NTP requirement was also analyzed. Next, we briefly investigated the cation specificity of the nucleotide requirement for protein import. Finally, in an effort to characterize the role that a putative PMF might play, we explored the effect of ionophores on GLO import. Ionophores were chosen that collapsed the pH gradient, the electrical gradient, or both components of the PMF. A thorough study of the energetics of protein import into peroxisomes is necessary to facilitate the understanding of the molecular mechanisms involved in the transport process.
MATERIALS AND METHODS
All NTPs, NTP analogs, ionophores, and general chemicals were purchased from Sigma. Sephadex G-25–80 (fine) was purchased from Pharmacia. Pumpkin (Cucurbita pepo var Half Moon) seeds were purchased from Petoseed Co., Inc. (Saticoy, CA). Redi-vue [35S]Met (specific activity, 43.5 TBq/mmol) was purchased from Amersham.
Preparation of Radiolabeled GLO
The plasmid pGLOZf, containing a cDNA insert for the entire coding region of the peroxisomal enzyme GLO in the pGEM7Zf(+) vector (Promega), was linearized with HindIII and transcribed with SP6 RNA polymerase as described by Brickner et al. (1997). Radiolabeled GLO was synthesized in a cell-free, wheat germ lysate system in the presence of [35S]Met. To test the NTP requirements for protein import, free nucleotides were removed from the translation products by size-exclusion chromatography using Sephadex G-25–80 (fine) as described by Olsen et al. (1989). The efficiency of the translation reaction was assessed by TCA precipitation of the proteins onto glass fiber filters, followed by ethanol washes and quantitation in a liquid scintillation counter (model LS 6800, Beckman). Standard import reactions contained radiolabeled GLO equivalent to 0.5 × 106 to 1.0 × 106 TCA-precipitable counts (usually 3–15 μL).
Isolation of Pumpkin Glyoxysomes
Pumpkin seeds were germinated in damp vermiculite for 5 to 6 d at 28 to 30°C in complete darkness. For each experiment approximately 40 g of cotyledons was harvested manually in dim light. Glyoxysomes were isolated as described by Brickner et al. (1997). The organelle isolation buffer included 10 mm azide for all experiments except the cation specificity and the ionophore experiments (see below).
In Vitro Import Assays
Standard import reactions contained glyoxysomes (80–500 μg of protein), radiolabeled GLO, 5 mm ATP (Mg2+ salt), and import buffer (25 mm Mes-KOH, pH 6.0, 0.5 m Suc, 10 mm KCl, 1 mm MgCl2, sometimes also with 10 mm NaN3; see below) in a final volume of 200 μL. All import reactions were initiated by the addition of translation products and incubated at 26°C for 30 min. After import, samples were treated with 10 μg/mL proteinase K for 30 min on ice to digest translation products not protected by the glyoxysomal membrane. Protease treatment was stopped by the addition of the inhibitor PMSF (1 mm final concentration). Intact glyoxysomes were reisolated on a 0.7 m Suc cushion and centrifuged at 8500g for 15 min in a refrigerated microcentrifuge. The pellets were resuspended in SDS-PAGE sample buffer, heated at 80 to 90°C for 2 to 5 min, and stored at −20°C until further analysis. Radioactive proteins were analyzed by SDS-PAGE and visualized by fluorography.
The Mg2+ salt of ATP was used in most standard import reactions, except that Na2ATP was used for the cation specificity experiments (see Fig. 2). Other import reactions were supplemented with additional cations as indicated in the legend to Figure 2. Import reactions containing other NTPs and/or NTP analogs were supplemented with equimolar MgCl2 in addition to the 1 mm MgCl2 present in the import buffer. The presence of Mg2+ by itself had no effect on import (data not shown).
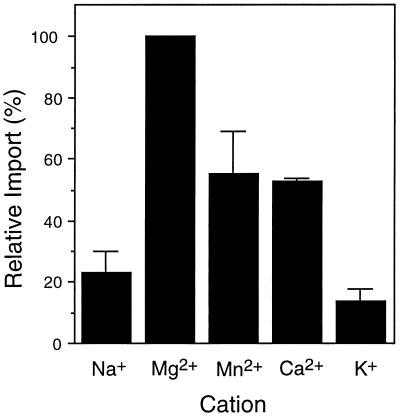
Figure 2.
Import of GLO into isolated glyoxysomes requires Mg2+. Standard import reactions were performed as described in Figure 1 and Methods except that 5 mm Na2ATP was present in each reaction instead of MgATP. All other import reactions were supplemented with additional cations (supplied by MgCl2, MnCl2, CaCl2, or KCl) at 5 mm final concentration. The level of import observed with 5 mm MgCl2 (and 5 mm Na2ATP) added to the import reaction was set at 100% for comparison with import levels in the presence of the other cations. The values presented are the average ± se of two experiments.
Glyoxysomes were preincubated with ionophores and 5 mm ATP for 20 min at room temperature before radiolabeled GLO was added to start the import reaction. The endogenous NTPs were not removed from the radiolabeled GLO used in the ionophore experiments. Modified import buffer (containing only 25 mm Mes-KOH, pH 6.0, and 0.5 m Suc; i.e. lacking the 10 mm KCl, 1 mm MgCl2, and azide) was used in the cation specificity experiments.
Azide (10 mm NaN3) was included in the organelle isolation buffer and import buffer in all experiments except the cation specificity (see Fig. 2) and the ionophore experiments (see Fig. 7). The presence of azide during organelle isolation and in import reactions did not affect import levels (data not shown). The purpose of the azide was to inhibit any ATP synthesis by the small amounts of mitochondria that may have been present in the glyoxysome preparation (Brickner et al., 1997), and thus remove any endogenous ATP supplied by the mitochondria. Azide was not included in the cation-specificity and ionophore experiments because the exact level of ATP present was not critical. In addition, we wanted to control precisely the amount of Na present in the import reactions when testing for cation effects on import.
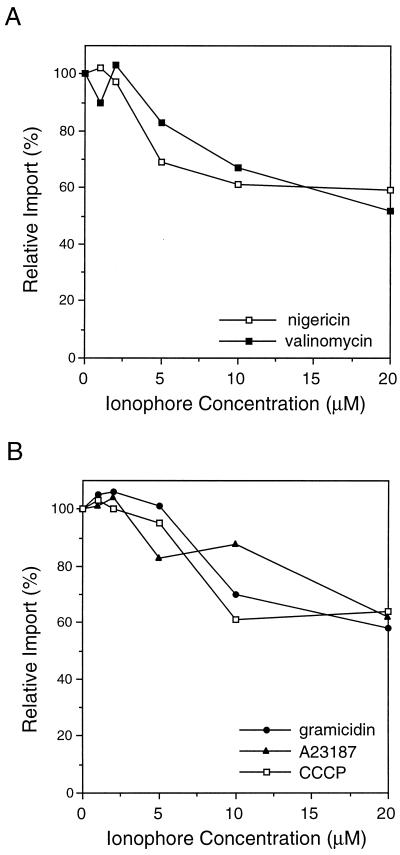
Figure 7.
Ionophores reduce the level of glyoxysomal protein import. To determine whether a PMF plays a role in peroxisomal protein import, isolated glyoxysomes were preincubated for 20 min at room temperature with various ionophores (1–20 μm final concentration). Import reactions were performed and analyzed as described in Figure 1. The level of protease-resistant GLO present in samples that had no ionophore added was set as 100% relative import for comparison with the ionophore-treated samples. A representative experiment is presented. A, GLO import into glyoxysomes in the presence of nigericin or valinomycin, ionophores that collapse a single component of the PMF. B, GLO import into glyoxysomes in the presence of ionophores that collapse the total PMF.
Each of the ionophore stock solutions was prepared in ethanol; dilutions of the stocks were used in the import reactions presented in Figure 7. Ethanol by itself, equivalent to the highest concentration present in the ionophore treatments (approximately 0.4%), had no effect on the level of GLO import (data not shown). Import reactions testing the effect of the Ca2+ ionophore A23187 also contained 10 mm CaCl2; 10 mm CaCl2 was also present in the controls and all import samples testing the Ca2+ ionophore A23187; 10 mm KCl was included in the controls and import samples testing valinomycin.
Quantitation
Levels of protein import were quantified by rehydrating manually excised, radioactive gel slices in 30% hydrogen peroxide overnight at 50°C. Standard scintillation cocktail for aqueous samples was added and samples were counted on a liquid scintillation counter (LS 6800, Beckman). Unless noted otherwise, the amount of protease-protected GLO detected after 30 min of import in the presence of 5 mm ATP was set at 100% relative import for comparison with the other treatments. We routinely got import efficiencies of 5 to 15% of the added radiolabeled protein imported into the isolated glyoxysomes in control samples. This corresponds to an average import of 3.2 × 105 molecules GLO/μg of glyoxysomes. The actual numbers for the experiments presented here are as follows: Table I, 5 mm ATP control, 3.4 × 105 molecules GLO/μg glyoxysomes; Figure 1, 5 mm ATP control, 3.2 × 105 molecules GLO/μg glyoxysomes; Figure 2, 5 mm Mg2+ supplementing the 5 mm ATP(Na+) salt, 5.8 × 105 molecules GLO/μg glyoxysomes; Figure 3, 5 mm ATP control, 6.2 × 105 molecules GLO/μg glyoxysomes; Figure 4, 5 mm GTP control, 1.9 × 105 molecules GLO/μg glyoxysomes; Figure 5, 5 mm ATP control, 1.7 × 105 molecules GLO/μg glyoxysomes; Figure 6, 5 mm ATP control, 1.4 × 105 molecules GLO/μg glyoxysomes; and Figure 7, no-ionophore control, 1.8 × 105 molecules GLO/μg glyoxysomes.
Table I.
NTPs are capable of supporting import of GLO into isolated glyoxysomes
| NTP | Percent Relative Importa |
|---|---|
| None | 22 ± 4 |
| ATP | 100 |
| GTP | 83 ± 11 |
| CTP | 76 ± 12 |
| UTP | 77 ± 11 |
| TTP | 51 ± 3 |
| ITP | 27 ± 4 |
Standard import assays were performed as described in Figure 1, except that each reaction contained 5 mm NTP, as indicated. The values presented are the average (± se) of three experiments.
The import level observed with 5 mm ATP was set at 100% for comparison with the other NTPs.
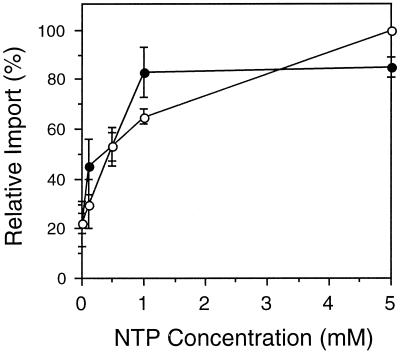
Figure 1.
Import of GLO into glyoxysomes is energy dependent. To characterize the NTP dependence of protein transport, increasing concentrations of ATP (○) or GTP (•) were added to standard import reactions (see Methods). Before addition to the import reactions, GLO translation products were desalted on a Sephadex G-25 column to remove endogenous nucleotides and other small molecules. The amount of radiolabeled GLO that remained protease protected after import in the presence of 5 mm ATP was set as 100% relative import for comparison with the other samples. The average ± se of three independent experiments is shown.
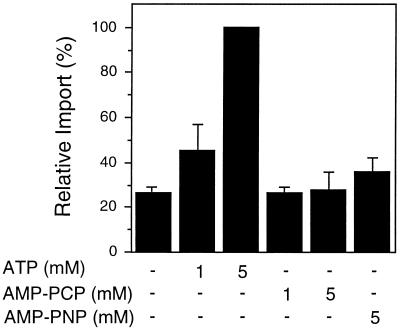
Figure 3.
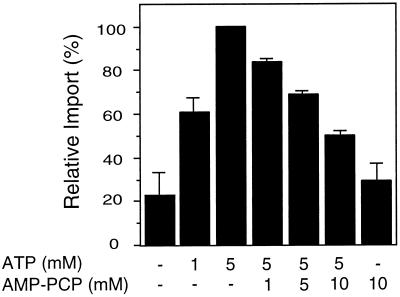
Nonhydrolyzable ATP analogs cannot support GLO import. To determine whether ATP hydrolysis is required for peroxisomal protein import, increasing amounts of ATP, AMP-PCP, or AMP-PNP (nonhydrolyzable analogs of ATP) were added to standard import reactions. Before addition to the import reactions, GLO translation products were desalted on a Sephadex G-25 column to remove endogenous nucleotides and other small molecules (see Methods). The amount of GLO imported into glyoxysomes in the presence of 5 mm ATP was set at 100% for comparison with the other samples in the same experiment. The average ± se of three independent experiments is shown.
Figure 4.
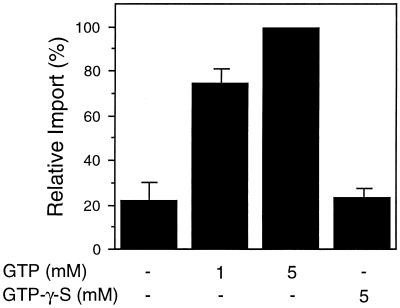
GTP hydrolysis is required for peroxisomal protein import. To determine whether the hydrolysis of GTP was required for peroxisomal protein import, increasing amounts of either GTP or GTP-γ-S were added to NTP-depleted import reactions, as described in Figure 1. The average ± se of two independent experiments is shown.
Figure 5.
AMP-PCP competes with ATP during protein import. Isolated glyoxysomes were incubated with increasing amounts of ATP and challenged with the nonhydrolyzable ATP analog, AMP-PCP, at the concentrations indicated. Subsequent import reactions were performed as described in Figure 1. The results shown are the average ± se of three separate experiments.
Figure 6.
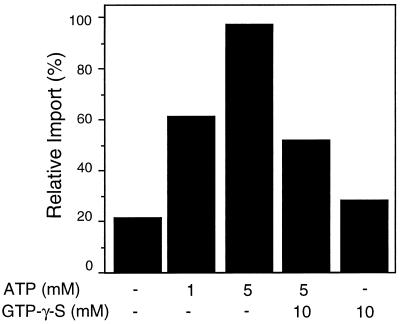
GTP-γ-S inhibits ATP-dependent protein import. To examine the effects of GTP-γ-S on ATP-dependent import, ATP and/or excess GTP-γ-S were preincubated with isolated glyoxysomes for 5 min. Protein import was initiated by the addition of radiolabeled GLO proteins. The amount of radiolabeled GLO imported in the presence of 5 mm ATP was set as 100% relative import. Because of minor differences in the ways in which each replicate experiment was performed, a representative experiment is presented.
RESULTS
NTP Requirement for Peroxisomal Protein Import
The energy requirements for the import of the leaf peroxisomal protein GLO were assessed using an in vitro import assay system described by Brickner et al. (1997). Glyoxysomes were isolated from dark-grown pumpkin cotyledons and incubated with radiolabeled GLO at 26°C for 30 min in the presence of various NTPs. The amount of protein imported into the organelle was measured by protease resistance; all protease-resistant GLO was assumed to be protected by the peroxisomal membrane.
As shown in Figure 1, the amount of GLO imported increased with higher concentrations of ATP or GTP. Maximal levels of import were observed at 5 mm ATP; no additional GLO was imported in the presence of 10 mm ATP (Brickner et al., 1997). Low levels of protein import (approximately 18–22%, relative to import at 5 mm ATP) were observed even when no exogenous NTP was added to the import reaction. This background level of import has been observed by others (Imanaka et al., 1987; Horng et al., 1995) and may be mediated by NTP bound to a component of the isolated organelles, to translation products, or to a cytosolic factor present in the wheat germ lysate; free NTPs were removed by the desalting procedure (Olsen et al., 1989). GTP supported GLO import nearly as well as ATP. Maximal GTP-dependent import was achieved at a lower NTP concentration; only 1 mm GTP was needed, compared with 5 mm ATP. This result suggests that GTP itself can be used as an energy source for protein import. It does not distinguish between a single energy-dependent process, which can use either ATP or GTP to drive import, and two separate energy-requiring steps, one dependent on ATP and the other dependent on GTP. Finally, the addition of both ATP and GTP to the same import reaction stimulated import only slightly more than the level in the control samples (data not shown).
In addition to ATP and GTP, other NTPs were assessed for their ability to support peroxisomal protein import (Table I). Standard import reactions were performed in the presence of 5 mm NTP; the radiolabeled GLO translation products were desalted to remove ATP added to the translation reaction or present in the wheat germ lysate (see Methods). With the exception of ITP, all NTPs tested in our in vitro protein import system could serve as an alternative energy source for peroxisomal protein import, although much less efficiently than ATP.
Cation Specificity for Peroxisomal Protein Import
Standard import reactions were performed in the presence of 5 mm ATP, provided as the Mg2+ salt. The disodium salt of ATP did not support GLO import unless equimolar levels of MgCl or Mg2+ acetate were also added to the reactions (Fig. 2). There was no difference in the level of protein import in reactions containing the Mg2+ salt of ATP compared with reactions containing disodium ATP supplemented with equimolar Mg2+ salts (data not shown); Mg2+ alone, i.e. without ATP, had no effect on import (data not shown). To determine whether other cations could substitute for Mg2+ during protein import, various salts were added to import reactions in the presence of disodium ATP. As seen in Figure 2, import in the presence of the divalent cations Mn2+ and Ca2+ was slightly better than with Na alone, but was still significantly less than import in reactions supplemented with Mg2+. K+ did not increase protein import at all above the disodium ATP background level.
NTP Hydrolysis Is Required for Peroxisomal Protein Import
Hydrolysis of the high-energy bonds of ATP may cause a conformational change in a component of the translocation machinery or in the translocating protein itself to facilitate protein entry into the organelle. To determine whether the hydrolysis of ATP is required for peroxisomal protein import, we incubated isolated glyoxysomes and radiolabeled GLO with the nonhydrolyzable ATP analogs AMP-PCP and AMP-PNP. Figure 3 shows that these nonhydrolyzable analogs of ATP did not substitute as an energy source for glyoxysomal protein import. Thus, ATP hydrolysis is required at some step during glyoxysomal protein import.
Because GTP was also capable of supporting peroxisomal protein import (Fig. 1), the necessity of GTP hydrolysis was examined. GTP-γ-S, a nonhydrolyzable analog of GTP, was added to standard import reactions in the absence of other NTPs, but was unable to increase the import of GLO to greater than background levels (Fig. 4). Therefore, it appears that the hydrolysis of GTP may also provide energy to drive peroxisomal proteins across the membrane.
Two components of the chloroplast protein import machinery have been shown to bind GTP (Kessler et al., 1994). There are at least two proteins involved in peroxisome biogenesis or function that have ATP-binding domains (Swartzman et al., 1996; Yahraus et al., 1996). One approach to the determination of whether the NTP hydrolysis requirement for peroxisomal protein import is caused by direct binding of the NTP by a protein factor (rather than an indirect metabolic role for NTP) is the use of nucleotide competition experiments. For these experiments an increasing concentration of the nonhydrolyzable ATP analog AMP-PCP was preincubated with ATP and isolated glyoxysomes. The import reaction was then initiated by the addition of radiolabeled GLO protein. As shown in Figure 5, the amount of protease-protected GLO decreased with increasing amounts of nonhydrolyzable ATP analog even in the presence of ATP. This competitive interaction between ATP and AMP-PCP indicates that there may be a discrete nucleotide binding site on some component of the translocation apparatus, or on another factor that is required for import of proteins into peroxisomes.
To assess the effect of a nonhydrolyzable GTP analog on ATP-dependent protein import levels, NTP competition experiments were performed using GTP-γ-S as the competitor. Competition was established by challenging an import reaction containing ATP with an excess of GTP-γ-S (Fig. 6). The addition of 10 mm GTP-γ-S significantly reduced the level of GLO imported into glyoxysomes in the presence of 5 mm ATP. This suggests that ATP and GTP are either competing for a common nucleotide binding site on some component required for protein translocation, or that both NTPs are required at different steps of protein import, one of which is inhibited by the binding of GTP-γ-S.
Effects of Ionophores on Peroxisomal Protein Import
To determine whether a PMF contributes to protein import, a variety of ionophores was added to standard in vitro import reactions. Isolated glyoxysomes were preincubated for 20 min at room temperature with either nigericin (a H+/K+ antiporter) to collapse the pH gradient, valinomycin (a K+ uniporter) to dissipate the membrane potential, or gramicidin (a channel former), CCCP (a H+ uniporter), A23187 (a Ca2+/H+ antiporter), or nigericin and valinomycin together to collapse both components of the PMF. Each ionophore reduced the level of GLO import to roughly 50 to 60% of the control import reaction when present at 10 to 20 μm (Fig. 7). Nigericin and valinomycin are typically used at micromolar to submicromolar concentrations (Pfanner and Neupert, 1986; Nicolay et al., 1987; Cline et al., 1992; Theg and Scott, 1993). In our experiments, low concentrations (1 and 2 μm) of nigericin and valinomycin had little or no effect on peroxisomal protein import (Fig. 7A). However, when the total PMF is collapsed by 10 to 20 μm CCCP, which is within the active range (Bellion and Goodman, 1987; Imanaka et al., 1987; Nicolay et al., 1987; Cline et al., 1992; Theg and Scott, 1993; Wendland and Subramani, 1993), peroxisomal protein import was inhibited by approximately 40 to 50% (Fig. 7B). This suggests that although a PMF across the peroxisomal membrane is not absolutely required for protein import, it may be involved indirectly in protein translocation into peroxisomes.
DISCUSSION
Energy is required to transport polypeptides through phospholipid membranes. At least three possible sources for that energy have been proposed: (a) energy released by induced conformational changes caused by the initial interaction between the protein and the membrane; (b) energy released by the hydrolysis of high-energy bonds found in ATP and other NTPs; and (c) a PMF consisting of a transmembrane electric potential and a pH gradient (Pugsley, 1989). Different protein import systems use each of these sources to varying degrees. We have addressed only the latter two in this report. Our in vitro assay for protein import into peroxisomes allows us to biochemically manipulate the exact reaction conditions. For most experiments, we first removed any endogenous NTPs from the GLO translation products and then added back different forms of energy to examine their effects on protein translocation.
We and others have shown that peroxisomal protein import is an ATP-dependent process (Imanaka et al., 1987; Soto et al., 1993; Horng et al., 1995; Brickner et al., 1997), but it is not yet known where in the targeting and translocation pathway the energy is required or how the energy is used. One possibility is that ATP is required by cytosolic chaperones or other cytosolic factors to ensure proper targeting of the protein to peroxisomes. In fact, the cytoplasmic ATPase PXAAA1 from humans apparently stabilizes a soluble receptor required for peroxisomal protein import (Yahraus et al., 1996). The Mg2+ salt of ATP has been shown to be specifically required by some chaperones for optimal function (Miernyk, 1997). The cation specificity experiments also showed that Mg2+ is the preferred salt for protein import into peroxisomes (Fig. 2). However, Mg2+ likely has multiple functions in this pathway. For instance, a yeast peroxisomal, membrane-associated, proton-translocating ATPase is also Mg2+ dependent (Douma et al., 1987).
A second possible role for NTPs during peroxisomal protein import is suggested by the nucleotide binding sites identified on several peroxisome-associated proteins (Verheyden et al., 1992; Swartzman et al., 1996) as well as on two components of the chloroplast translocation machinery (Kessler et al., 1994). When isolated glyoxysomes were preincubated with ATP and an excess of a nonhydrolyzable NTP analog, subsequent GLO import was significantly decreased (Figs. 5 and 6), suggesting that nucleotide binding to an unidentified factor may be important for peroxisomal protein transport.
Both ATP and GTP were able to support GLO import in our in vitro assay system (Figs. 1, 4–6). GTP-binding proteins are known to be involved in several other transport pathways (Balch, 1990; Pfeffer, 1992; Kessler et al., 1994) and three small GTP-binding proteins have been identified in rat peroxisomal membranes (Verheyden et al., 1992). Thus, it is possible that GTP has a direct role in peroxisomal protein import. Although Wendland and Subramani (1993) did not observe a similar involvement of GTP-hydrolyzing proteins when they examined peroxisomal protein import in permeabilized mammalian cells, they used only 100 μm GTP-γ-S in the presence of 1 mm ATP and an ATP-regenerating system. The concentration of the GTP analog may have been too low to observe a clear effect on the localization of the microinjected peroxisomal protein.
It is possible that ATP and GTP are both necessary for protein import, but at distinct energy-requiring steps. When a nonhydrolyzable analog of GTP was added to import reactions in the presence of 5 mm ATP, the level of GLO import was lower than in reactions containing ATP only, but higher than the import seen with GTP-γ-S alone (Fig. 6). This reduction in the amount of GLO imported in an ATP-dependent manner indicates that there is either a common NTP-binding site for which ATP and GTP compete, or that the addition of GTP-γ-S inhibits a separate, GTP-requiring import step. At this time we are unable to distinguish between these two options. It is likely that ATP (and possibly GTP as well) has multiple roles and acts at several different steps along the import pathway. This will make it more complicated to determine exactly how each nucleotide is involved in peroxisomal protein import.
Some protein transport systems also require a PMF for maximal efficiency of translocation (Pfanner and Neupert, 1986; Yamane et al., 1987; Wong and Buckley, 1989; Cline et al., 1992; Theg and Scott, 1993), but it is not clear whether a PMF is present across the peroxisomal membrane. Our results, using a broad range of ionophores, suggest that a PMF is involved in peroxisomal protein transport, although it is not absolutely required (Fig. 7). Each of the ionophores tested consistently reduced the amount of GLO imported to approximately 50 to 60% of control import (in the absence of ionophores), indicating that these ionophores inhibited protein import but did not abolish the process. It appears that this inhibition is not caused by decreased efficiency of import; GLO import in the presence of ionophores does not recover even after longer reaction times (data not shown). It may be that a PMF-requiring factor is depleted in the presence of ionophores such that translocation into the matrix is compromised. Alternatively, the pH of the peroxisome matrix may be important for optimal protein import; addition of ionophores (except valinomycin) would make it difficult to maintain a constant pH in the matrix, resulting in lower levels of peroxisomal protein import. A proton-ATPase on the peroxisome membrane may be responsible for generating and maintaining the acidic matrix of yeast peroxisomes (Douma et al., 1987; Nicolay et al., 1987; del Valle et al., 1988; Waterham et al., 1990); however, neither an ATPase nor an acidic matrix has yet been described for plant peroxisomes.
There are two additional factors that must be considered. First, it is important to note that low concentrations of nigericin or valinomycin did not inhibit GLO import in vitro (Fig. 7A). There was slightly greater inhibition of import when nigericin and valinomycin were added together, but the results were not strictly additive (data not shown). Nicolay et al. (1987) found that 1 to 2 μm nigericin and valinomycin together was sufficient to dissipate the pH gradient across yeast peroxisomal membranes. GLO import into plant peroxisomes was clearly inhibited by 5 to 10 μm nigericin (Fig. 7A). When used at 10 μm, neither valinomycin nor nigericin inhibited in vitro protein transport into rat liver peroxisomes, although no quantitation of these results was presented (Imanaka et al., 1987). In most systems, nigericin and valinomycin are expected to be active at very low concentrations, i.e. 1 μm or less (Pfanner and Neupert, 1986; Cline et al., 1992; Theg and Scott, 1993). Concentrations of nigericin or valinomycin greater than 10 μm may have nonspecific or surfactant effects (Reed, 1979).
Second, there may be some differences in the ways in which peroxisomes from different organisms respond to ionophores. When the total PMF was collapsed by 10 μm CCCP, protein import into plant peroxisomes was inhibited (Fig. 7B). Wendland and Subramani (1993) found that 10 μm CCCP does not inhibit import into peroxisomes in semipermeabilized mammalian cells; Imanaka et al. (1987) observed no inhibition of in vitro protein transport into rat liver peroxisomes in the presence of 10 μm CCCP. Neither group presented any quantitation of these results. CCCP may have more effect on yeast peroxisomes, although it is difficult to know for sure because higher concentrations of CCCP have been used. The assembly of alcohol oxidase is prevented by 25 μm CCCP, leading the authors to conclude that a PMF is required for the import and maturation of this yeast protein (Bellion and Goodman, 1987). Using a 31P NMR assay, Nicolay et al. (1987) found that the pH gradient across yeast peroxisomal membranes is destroyed by 2 mm CCCP.
Thus, it may be that a single component of the PMF alone does not affect peroxisomal protein import in plants; neither nigericin nor valinomycin at low (active) concentrations inhibited GLO import (Fig. 7A). However, even 10 μm CCCP showed maximal inhibition of GLO import (Fig. 7B), suggesting that collapsing the total PMF decreases protein import into peroxisomes in vitro. It seems unlikely that the PMF is directly providing necessary energy for the translocation event. A secondary role for a PMF, such as in maintaining a pH gradient across the membrane, may be responsible for the consistent level of inhibition we observed in the presence of each ionophore. It is not possible to conclude anything definitive based solely on results provided by experiments with a single ionophore at a single concentration. Variables such as the size of the compartment and the magnitude of the pH gradient across the membrane may also influence the behavior of individual ionophores (Reed, 1979). We have tested the effects on peroxisomal protein import of a wide range of ionophores at many concentrations, and we conclude that a PMF appears to play a role in peroxisomal protein import, perhaps through an indirect effect, but that the energy of a PMF is not absolutely required for protein translocation.
In a continuing effort to understand the cellular requirements and mechanisms of higher plant peroxisomal protein import, we have extensively characterized the energy requirements for GLO import into glyoxysomes in vitro. We have firmly established that energy from NTPs is required during protein translocation in peroxisomes. However, future studies are needed to define exactly where and how this energy is being used. Detailed investigations may lead to mechanistic models that include energy-requiring cytosolic factors and/or peroxisome-specific factors that are necessary to translocate peroxisomal proteins through the lipid bilayer.
ACKNOWLEDGMENTS
We thank Jason Brickner, Wendy Crookes, Yan Lin, and Aaron Liepman for many helpful discussions. Olivia Bottum and Jessica McHie provided excellent technical assistance. Drs. Charles Yocum and Eran Pichersky contributed useful advice and comments during the preparation of the manuscript.
Abbreviations:
- A23187
calcinomycin
- AMP-PCP
methylene adenosine 5′-triphosphate
- AMP-PNP
5′-adenylylimidodiphosphate
- CCCP
carbonyl cyanide m-chloro-phenylhydrazone
- GLO
glycolate oxidase
- NTP
nucleotide triphosphate
- PMF
protonmotive force
Footnotes
This work was funded by a grant from the U.S. Department of Agriculture to L.J.O. D.G.B. was supported in part by a fellowship from the Cellular Biotechnology Training Program (National Institutes of Health grant no. GM08353).
LITERATURE CITED
- Balch WE. Small GTP-binding proteins in vesicular transport. Trends Biochem Sci. 1990;15:473–477. doi: 10.1016/0968-0004(90)90301-q. [DOI] [PubMed] [Google Scholar]
- Bellion E, Goodman JM. Proton ionophores prevent assembly of a peroxisomal protein. Cell. 1987;48:165–173. doi: 10.1016/0092-8674(87)90367-9. [DOI] [PubMed] [Google Scholar]
- Brickner DG, Harada JJ, Olsen LJ. Protein transport into higher plant peroxisomes. In vitro assay provides evidence for receptor involvement. Plant Physiol. 1997;113:1213–1221. doi: 10.1104/pp.113.4.1213. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cline K, Ettinger WF, Theg SM. Protein-specific energy requirements for protein transport across or into thylakoid membranes. J Biol Chem. 1992;267:2688–2696. [PubMed] [Google Scholar]
- del Valle R, Soto U, Necochea C, Leighton F. Detection of an ATPase activity in rat liver peroxisomes. Biochem Biophys Res Commun. 1988;156:1353–1359. doi: 10.1016/s0006-291x(88)80781-2. [DOI] [PubMed] [Google Scholar]
- Dodt G, Gould SJ. Multiple PEX genes are required for proper subcellular distribution and stability of Pex5p, the PTS1 receptor: evidence that PTS1 protein import is mediated by a cycling receptor. J Cell Biol. 1996;135:1763–1774. doi: 10.1083/jcb.135.6.1763. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Douma AC, Veenhuis M, Sulter GJ, Harder W. A proton-translocating adenosine triphosphatase is associated with the peroxisomal membrane of yeasts. Arch Microbiol. 1987;147:42–47. doi: 10.1007/BF00492903. [DOI] [PubMed] [Google Scholar]
- Elgersma Y, Kwast L, Klein A, Voorn-Brouwer T, van den Berg M, Metzig B, America T, Tabak HF, Distel B. The SH3 domain of the Saccharomyces cerevisiae peroxisomal membrane protein Pex13p functions as a docking site for Pex5p, a mobile receptor for the import of PTS1 containing proteins. J Cell Biol. 1996;135:97–109. doi: 10.1083/jcb.135.1.97. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Erdmann R, Blobel G. Identification of Pex13p, a peroxisomal membrane receptor for the PTS1 recognition factor. J Cell Biol. 1996;135:111–121. doi: 10.1083/jcb.135.1.111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Frederick SE, Gruber PJ, Newcomb EH. Plant microbodies. Protoplasma. 1975;84:1–29. [Google Scholar]
- Gietl C. Glyoxysomal malate dehydrogenase from watermelon is synthesized with an amino-terminal transit peptide. Proc Natl Acad Sci USA. 1990;87:5773–5777. doi: 10.1073/pnas.87.15.5773. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gietl C. Protein targeting and import into plant peroxisomes. Physiol Plant. 1996;97:599–608. [Google Scholar]
- Gould SJ, Kalish JE, Morrell JC, Bjorkman J, Urquhart AJ, Crane DI. An SH3 protein in the peroxisome membrane is a docking factor for the PTS1 receptor. J Cell Biol. 1996;135:85–95. doi: 10.1083/jcb.135.1.85. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gould SJ, Keller G-A, Subramani S. Identification of a peroxisomal targeting signal at the carboxy terminus of firefly luciferase. J Cell Biol. 1987;105:2923–2931. doi: 10.1083/jcb.105.6.2923. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Horng JT, Behari R, Burke ECA, Baker A. Investigation of the energy requirement and targeting signal for the import of glycolate oxidase into glyoxysomes. Eur J Biochem. 1995;230:157–163. doi: 10.1111/j.1432-1033.1995.tb20546.x. [DOI] [PubMed] [Google Scholar]
- Imanaka T, Small GM, Lazarow PB. Translocation of acyl-CoA oxidase into peroxisomes requires ATP hydrolysis but not a membrane potential. J Cell Biol. 1987;105:2915–2922. doi: 10.1083/jcb.105.6.2915. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Keegstra K, Olsen LJ, Theg SM. Chloroplastic precursors and their transport across the envelope membranes. Annu Rev Plant Physiol Plant Mol Biol. 1989;40:471–501. [Google Scholar]
- Kessler F, Blobel G, Patel HA, Schnell DJ. Identification of two GTP-binding proteins in the chloroplast protein import machinery. Science. 1994;266:1035–1039. doi: 10.1126/science.7973656. [DOI] [PubMed] [Google Scholar]
- Kindl H. Plant peroxisomes: recent studies on function and biosynthesis. Cell Biochem Funct. 1992;10:153–158. doi: 10.1002/cbf.290100304. [DOI] [PubMed] [Google Scholar]
- McNew JA, Goodman JM. The targeting and assembly of peroxisomal proteins: some old rules do not apply. Trends Biochem Sci. 1996;21:54–58. [PubMed] [Google Scholar]
- Miernyk JA. The 70 kDa stress-related proteins as molecular chaperones. Trends Plant Sci. 1997;2:180–187. [Google Scholar]
- Nicolay K, Veenhuis M, Douma AC, Harder W. A 31P NMR study of the internal pH of yeast peroxisomes. Arch Microbiol. 1987;147:37–41. doi: 10.1007/BF00492902. [DOI] [PubMed] [Google Scholar]
- Ogren WL. Photorespiration: pathways, regulation, and modification. Annu Rev Plant Physiol. 1984;35:415–442. [Google Scholar]
- Olsen LJ, Harada JJ. Peroxisomes and their assembly in higher plants. Annu Rev Plant Physiol Plant Mol Biol. 1995;46:123–146. [Google Scholar]
- Olsen LJ, Theg SM, Selman BR, Keegstra K. ATP is required for the binding of precursor proteins to chloroplasts. J Biol Chem. 1989;264:6724–6729. [PubMed] [Google Scholar]
- Pfanner N, Neupert W. Transport of F1-ATPase subunit β into mitochondria depends on both a membrane potential and nucleoside triphosphates. FEBS Lett. 1986;209:152–156. doi: 10.1016/0014-5793(86)81101-2. [DOI] [PubMed] [Google Scholar]
- Pfanner N, Rassow J, Guiard B, Sollner T, Hartl F-U, Neupert W. Energy requirements for unfolding and membrane translocation of precursor proteins during import into mitochondria. J Biol Chem. 1990;265:16324–16329. [PubMed] [Google Scholar]
- Pfeffer SR. GTP-binding proteins in intracellular transport. Trends Cell Biol. 1992;2:41–46. doi: 10.1016/0962-8924(92)90161-f. [DOI] [PubMed] [Google Scholar]
- Powers MA, Forbes DJ. Cytosolic factors in nuclear import: what's importin? Cell. 1994;79:931–934. doi: 10.1016/0092-8674(94)90024-8. [DOI] [PubMed] [Google Scholar]
- Preisig-Müller R, Kindl H. Thiolase mRNA translated in vitro yields a peptide with a putative N-terminal presequence. Plant Mol Biol. 1993;22:59–66. doi: 10.1007/BF00038995. [DOI] [PubMed] [Google Scholar]
- Pugsley AP (1989) Basic principles and techniques. In Protein Targeting. Academic Press, San Diego, CA, pp 13–44
- Rachubinski RA, Subramani S. How proteins penetrate peroxisomes. Cell. 1995;83:525–528. doi: 10.1016/0092-8674(95)90091-8. [DOI] [PubMed] [Google Scholar]
- Reed PW. Ionophores. Methods Enzymol. 1979;55:435–454. doi: 10.1016/0076-6879(79)55058-7. [DOI] [PubMed] [Google Scholar]
- Soto U, Pepperkok R, Ansorge W, Just WW. Import of firefly luciferase into mammalian peroxisomes in vivo requires nucleoside triphosphates. Exp Cell Res. 1993;205:66–75. doi: 10.1006/excr.1993.1059. [DOI] [PubMed] [Google Scholar]
- Swartzman EE, Viswanathan MN, Thorner J. The PAL1 gene product is a peroxisomal ATP-binding cassette transporter in the yeast Saccharomyces cerevisiae. J Cell Biol. 1996;132:549–563. doi: 10.1083/jcb.132.4.549. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Swinkels BW, Gould SJ, Bodnar AJ, Rachubinski RA, Subramani S. A novel, cleavable peroxisomal targeting signal at the amino-terminus of the rat 3-ketoacyl-CoA thiolase. EMBO J. 1991;10:3255–3262. doi: 10.1002/j.1460-2075.1991.tb04889.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Theg SM, Bauerle C, Olsen LJ, Selman BR, Keegstra K. Internal ATP is the only energy requirement for the translocation of precursor proteins across chloroplastic membranes. J Biol Chem. 1989;264:6730–6736. [PubMed] [Google Scholar]
- Theg SM, Scott SV. Protein import into chloroplasts. Trends Cell Biol. 1993;3:186–190. doi: 10.1016/0962-8924(93)90212-j. [DOI] [PubMed] [Google Scholar]
- Trelease RN. Biogenesis of glyoxysomes. Annu Rev Plant Physiol. 1984;35:321–347. [Google Scholar]
- van den Bosch H, Schutgens RBH, Wanders RJA, Tager JM. Biochemistry of peroxisomes. Annu Rev Biochem. 1992;61:157–197. doi: 10.1146/annurev.bi.61.070192.001105. [DOI] [PubMed] [Google Scholar]
- Verheyden K, Fransen M, Van Veldhoven PP, Mannaerts GP. Presence of small GTP-binding proteins in the peroxisomal membrane. Biochim Biophys Acta. 1992;1109:48–54. doi: 10.1016/0005-2736(92)90185-o. [DOI] [PubMed] [Google Scholar]
- Walter P, Johnson AE. Signal sequence recognition and protein targeting to the endoplasmic reticulum membrane. Annu Rev Cell Biol. 1994;10:87–119. doi: 10.1146/annurev.cb.10.110194.000511. [DOI] [PubMed] [Google Scholar]
- Walton PA, Wendland M, Subramani S, Rachubinski RA, Welch WJ. Involvement of 70 kD heat-shock proteins in peroxisomal import. J Cell Biol. 1994;125:1037–1046. doi: 10.1083/jcb.125.5.1037. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Waterham HR, Keizer-Gunnick I, Goodman JM, Harder W, Veenhuis M. Immunocytochemical evidence for the acidic nature of peroxisomes in methylotrophic yeasts. FEBS Lett. 1990;262:17–19. doi: 10.1016/0014-5793(90)80142-6. [DOI] [PubMed] [Google Scholar]
- Wendland M, Subramani S. Cytosol-dependent peroxisomal protein import in a permeabilized cell system. J Cell Biol. 1993;120:675–685. doi: 10.1083/jcb.120.3.675. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Whitney AB, Bellion E. ATPase activities in peroxisome-proliferating yeast. Biochim Biophys Acta. 1991;1058:345–355. doi: 10.1016/s0005-2728(05)80130-6. [DOI] [PubMed] [Google Scholar]
- Wolins NE, Donaldson RP. Specific binding of the peroxisomal protein targeting sequence to glyoxysomal membranes. J Biol Chem. 1994;269:1149–1153. [PubMed] [Google Scholar]
- Wolvetang EJ, Tager JM, Wanders RJA. Latency of the peroxisomal enzyme acyl-CoA:dihydroxyacetonephosphate acyltransferase in digitonin-permeabilized fibroblasts: the effect of ATP and ATPase inhibitors. Biochem Biophys Res Commun. 1990;170:1135–1143. doi: 10.1016/0006-291x(90)90511-k. [DOI] [PubMed] [Google Scholar]
- Wong KR, Buckley JT. Science. 1989;246:654–656. doi: 10.1126/science.2814486. [DOI] [PubMed] [Google Scholar]
- Yahraus T, Braverman N, Dodt G, Kalish JE, Morrell JC, Moser HW, Valle D. The peroxisome biogenesis disorder group 4 gene, PXAAA1, encodes a cytoplasmic ATPase required for the stability of the PTS1 receptor. EMBO J. 1996;15:2914–2923. [PMC free article] [PubMed] [Google Scholar]
- Yamane K, Ichihara S, Mizushima S. In vitro translocation of protein across Escherichia coli membrane vesicles requires both the proton motive force and ATP. J Biochem. 1987;262:2358–2362. [PubMed] [Google Scholar]