Abstract
New series of customizable diastereomeric cis- and trans-monocyclic enol-phosphonate analogs to Cyclophostin and Cyclipostins were synthesized. Their potencies and mechanisms of inhibition toward six representative lipolytic enzymes belonging to distinct lipase families were examined. With mammalian gastric and pancreatic lipases no inhibition occurred with any of the compounds tested. Conversely, Fusarium solani Cutinase and lipases from Mycobacterium tuberculosis (Rv0183 and LipY) were all fully inactivated. Best inhibitors displayed a cis conformation (H and OMe) and exhibited higher inhibitory activities than the lipase inhibitor Orlistat towards same enzymes. Our results have revealed that chemical group at the γ-carbon of the phosphonate ring strongly impacts the inhibitory efficiency, leading to a significant improvement in selectivity toward a target lipase over another. The powerful and selective inhibition of microbial (fungal and mycobacterial) lipases suggests that these 7-membered monocyclic enol-phosphonates should provide useful leads for the development of novel and highly selective antimicrobial agents.
Keywords: cyclic phosphonates, microbial lipase, selective inhibition
INTRODUCTION
Lipases and their inhibitors have potential applications in the fields of chemistry,1 biotechnology2 and medicine. In the later domain, for instance, inhibiting digestive lipases to reduce fat absorption has become the main pharmacological approach to the treatment of obesity during the last decade.3 The best example is the FDA-approved anti-obesity drug Orlistat (marketed as Xenical by Roche and Alli by GlaxoSmithKline), which reacts with the nucleophilic serine residue from the catalytic triad of pancreatic and gastric lipases and inhibits these enzymes within the gastrointestinal tract.4,5 Orlistat is also known to inhibit various other enzymes, like fatty acid synthase,6 providing this compound with potential applications in the treatment of Alzheimer’s disease,7 type II diabetes,8 as well as anti-tumoral6 and anti-mycobacterial9–11 activities. However, this lack of selectivity could also lead to undesirable side-effects, which might affect its current clinical use.
In this context, designing and synthesizing selective inhibitors of various animal and microbial lipases represent attractive and useful probes to reveal the catalytic mechanisms of these lipases, and especially to better understand specific enzyme-substrate interactions.12 Such inhibitors also represent attractive leads for developing specific treatments towards various organisms involving lipolytic enzymes as essential metabolic enzymes or virulence factors.13,14 Phosphonate analogs of biologically active phosphates have been shown to be extremely useful tools in investigating mechanistic details of various enzymatic systems.15,16 Their success is usually attributed to the non-hydrolyzability of a P-C bond (phosphonate analog) when compared to the P-O bond of the corresponding phosphate leading to enhanced compound lifetime in vivo.17,18 Moreover, phosphonates are known to mimic in both their charge distribution and geometry the first transition state occurring during enzymatic carboxylester hydrolysis, with the formation of an irreversible covalent bond between the nucleophilic Oγ of the active site serine and the phosphorus atom.19
Cyclophostin (Scheme 1), a bicyclic enol-phosphate isolated from a fermentation solution of Streptomyces lavendulae (strain NK901093),20 was identified as an irreversible inhibitor of acetylcholinesterase (AChE) with IC50 values in the nanomolar range.16,20,21 The unusual bicyclic enol-phosphate moiety is also found in a second family of structurally related natural products, named the Cyclipostins22 (Scheme 1). The Cyclipostins possess a core structure similar to that of Cyclophostin, but are phosphate esters of long chain lipophilic alcohols of various lengths and structures. All identified Cyclipostins have been described to be potent inhibitors of hormone-sensitive lipase (HSL),22 and have also been reported to inhibit the growth of various mycobacteria including Mycobacterium smegmatis, Mycobacterium phlei, Nocardia abcessus and Corynebacterium diphteriae.23 The minimum inhibitory concentrations (MIC) obtained were similar or even lower than those of the well-known antibiotics Rifampicin or Penicillin G. These recent results strongly suggest that Cyclophostin and Cyclipostins compounds can inhibit serine hydrolases produced by these organisms, including mycobacterial lipases.
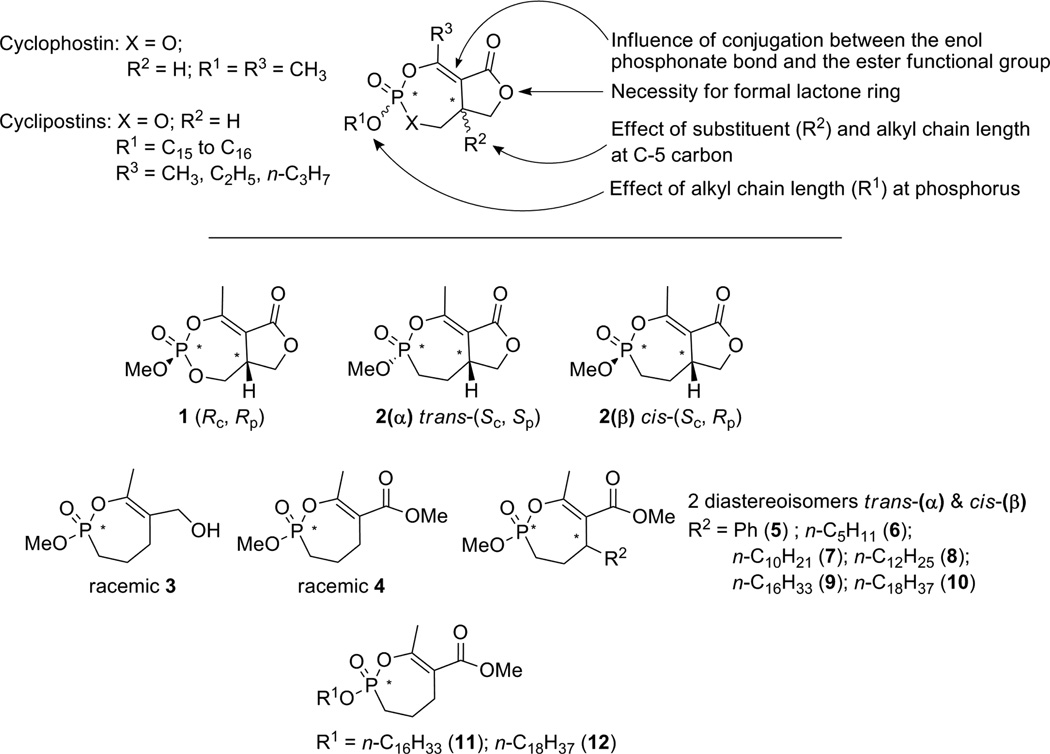
Scheme 1.
Rationale for synthesis of Cyclophostin and Cyclipostins analogs, and related structures investigated: Cyclophostin molecule (1)21 and phosphonate analogs (2);15 monocyclic phosphonate analogs to either Cyclophostin (3–4,16 5–10) or Cyclipostins (11–12).
In the present article, we report the synthesis and first mechanistic study of a new series of cyclic enol phosphonate analogs of both Cyclophostin and Cyclipostins (Scheme 1) as potential inhibitors of lipases. Three mammalian digestive lipases (human pancreatic lipase, HPL; dog gastric lipase, DGL; and guinea pig pancreatic lipase-related protein 2, GPLRP2) and three microbial lipases (Fusarium solani Cutinase, Mycobacterium tuberculosis Rv0183 and LipY) were chosen as representative enzyme targets. HPL and DGL, the main lipases involved in the digestion of dietary lipids, are sn-1,3-regioselective and sn-3-stereoselective lipases, respectively, acting on both triacylglycerols and diacylglycerols.24 GPLRP2 belongs to the pancreatic lipase gene family,25 but differs from HPL by its kinetic and structural properties.26 More precisely, the lid domain controlling the access to the active site, as in the case of HPL and DGL,27 is missing in GPLRP2 and its catalytic serine is therefore easily accessible to the solvent.25 GPLRP2 exhibits lipase, phospholipase A1 and galactolipase activities,28–30 and its main physiological function is the hydrolysis of dietary galactolipids29 and phospholipids.25 Among the three microbial lipases investigated, Cutinase is able to hydrolyze a wide range of substrates (fatty acid esters, triacylglycerols and phospholipids), and possesses, like GPLRP2, the advantage to exhibit an active site always accessible to the solvent.31–33 Mycobacterium tuberculosis Rv0183 is a monoglyceride lipase implicated in the architecture of the membrane and in the degradation of extracellular monoacylglycerols;14,34,35 while LipY is a triacylglycerol lipase belonging to the HSL family, involved in triacylglycerols degradation during persistence36 and in host lipids degradation during reactivation of the bacteria.37 Both LipY and Rv0183, which may contribute to the growth and pathogenicity of Mycobacterium tuberculosis, therefore appear as potential drug targets against tuberculosis.
These six lipases were then chosen as representative models belonging to different lipase families (Table 1). They have distinct structural properties (with or without a lid controlling the access to the active site for instance) and display various substrate specificities. Such diversity, covering a wide range of lipolytic enzymes, will then allow us to access structure-activity relationships between these targeted enzymes and the diastereoisomeric cyclic enol-phosphonate inhibitors analogous to Cyclipostins and Cyclophostin.
Table 1.
Biochemical properties of the six lipases tested
| Enzyme | Lipase gene family | Presence of a “lid” | Main substrates |
|---|---|---|---|
| DGL | Acid lipase | Yes - (PDB ID: 1K8Q) a | Triacylglycerols Diacylglycerols |
| HPL | Pancreatic lipase | Yes - (PDB ID: 1LPB) b | Triacylglycerols Diacylglycerols |
| GPLRP2 | Pancreatic lipase | No - (PDB ID: 1GPL) c | Triacylglycerols Phospholipids Galactolipids |
| Cutinase | Fungal lipase | No - (PDB ID: 1XZL) d | Cutin Triacylglycerols Phospholipids Galactolipids |
| Rv0183 | Monoglyceride lipase | Structural data unavailable | Monoacylglycerols |
| LipY | Hormone-sensitive lipase | Structural data unavailable | Triacylglycerols |
In this context, the inhibitory power and selectivity of this new series of organophosphorous compounds have been evaluated on the basis of the following structural properties: ❶ effect of removal of conjugation between the enol-phosphonate bond and the ester functional group; ❷ requirement of the formal lactone ring; ❸ effect of substituent R2 at C-5 carbon atom, and ❹ effect of substituent R1 at phosphorus atom (Scheme 1). To reach this goal, new synthetic routes were developed in order to easily provide a library of enol-phosphonate compounds bearing different chemical groups.
RESULTS
Synthesis of Cyclophostin and Cyclipostins phosphonate analogs
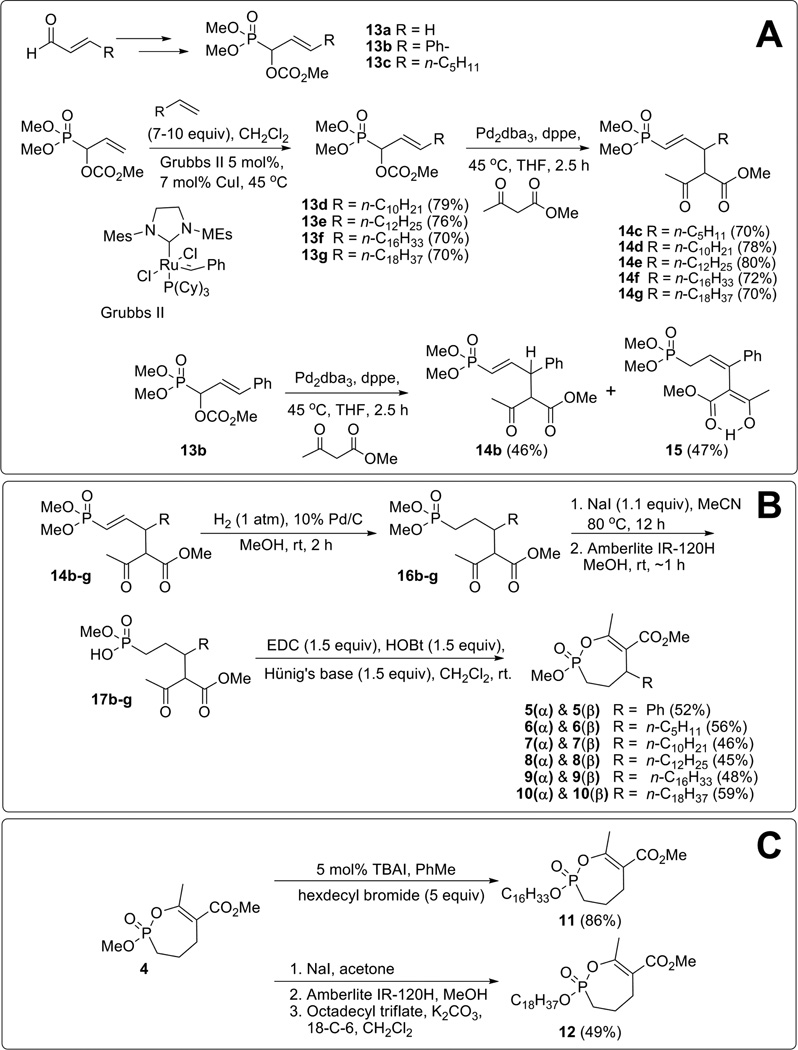
The syntheses of racemic Cyclophostin 1,21 racemic bicyclic phosphonate analogs 2(α)/2(β),15 and unsubstituted monocyclic analogs 3 and 416 have already been reported. In general, the methods employed in the synthesis of 2(α)/2(β) and 4 were adapted to the synthesis of the novel substituted monocyclic phosphonate analogs 5–10. The precursor carbonates 13a–13c were prepared following literature methods38 (Scheme 2A). Additional carbonates 13d–13g were prepared by the cross metathesis reaction of the parent acrolein-derived carbonate 13a with terminal alkenes using Grubbs second generation catalyst with copper(I) iodide as co-catalyst.38,39 The carbonates 13d–13g were formed in good yields (70–79%) as mixture of E and Z isomers in ratios varying from 9:1 to 14:1. Palladium(0)-catalyzed reaction of the carbonates 13c–13g with methyl acetoacetate gave the vinyl phosphonates 14c–g (Scheme 2A) in good yields (70–80%). The products were formed as diastereoisomeric mixtures, but with exclusively E alkene geometry. However, the reaction of carbonate 13b produced both the vinyl phosphonate 14b and the allyl phosphonate 15, which is the result of the migration of double bond into conjugation with the phenyl group. Interestingly, spectroscopic analysis (NMR and IR) and X-ray crystallography showed that the allyl phosphonate 15 exists predominantly in the enol-form of β-keto ester. Both vinyl 14b and allyl phosphonates 15 were taken on to the next step since, after hydrogenation, they should yield the same alkyl phosphonate.
Scheme 2.
Synthesis of novel monocyclic enol-phosphonate analogs of either Cyclophostin (5–10) or Cyclipostins (11–12).
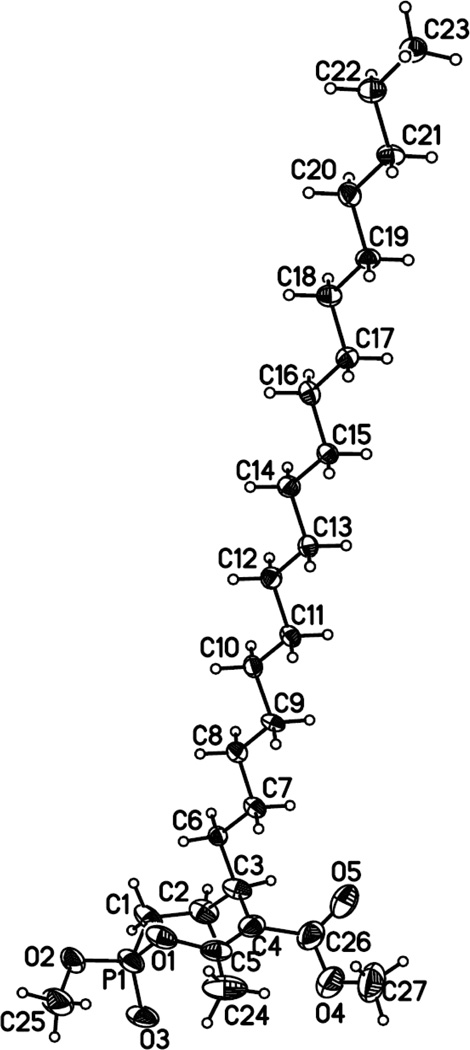
The treatment of a methanolic solution of the vinyl phosphonates 14b–g (or 15) with hydrogen gas over 10% palladium on carbon gave saturated phosphonates 16a–g in quantitative yields (Scheme 2B). As expected, hydrogenation of either 14 or 15 gave the saturated phosphonate 16b. The phosphonates 16b–16g were mono-demethylated using sodium iodide (1.1 equiv) in refluxing acetonitrile to give the corresponding sodium salts, which were subsequently converted to monophosphonic acids 17a–g by treatment with Amberlite IR-120H resin in methanol. The monophosphonic acids 17a–g were cyclized, without further purification, using a combination of EDC, HOBt and Hünig’s base (i-Pr2NEt) to give the substituted monocyclic phosphonates 5–10 as diastereoisomeric mixtures in 45–59% yields over the three steps. As previously demonstrated in the synthesis of the racemic bicyclic phosphonate analogs of Cyclophostin 2(α) and 2(β),15 compounds 5 to 10 were best described by the relationship between the OMe on phosphorus and the H-substituent on C-5 carbon atom as being either in trans (α-isomer) or cis (β-isomer) relationship. The two racemic diastereoisomers (α and β) were further separated by careful silica gel column chromatography (Scheme 2B). A single crystal X-ray diffraction structure of 10(α) was further obtained at an edge resolution of ~ 0.97 Å (Tables S1 to S5) thus confirming these stereochemical assignments (trans-α and cis-β). As shown in Figure 1, the obtained structure clearly exhibits a trans relationship between the methoxy (numbered O2-C25) and the hydrogen atom located on the carbon numbered C3.
Figure 1.
Projection view of 10(α) with 30% thermal ellipsoids (disorder atoms omitted for clarity). This crystal structure has been deposited at the Cambridge Crystallographic Data Centre and allocated the deposition number CCDC 873939.
The initial approach to forming long chain phosphonate esters (Scheme 2C) from the monocyclic phosphonate 4 involved de-methylation and then re-alkylation in three separate reaction steps. Thus, the phosphonate 4 was treated with NaI in acetone to give the sodium salt, which was protonated by treatment with Amberlite IR-120H in methanol. The phosphonic acid was then re-alkylated using potassium carbonate, 18-Crown-6 and octadecyl triflate40 to give phosphonate 12 in 49% yield over three steps. However, we have discovered more recently that trans-esterification reaction of the phosphonate methyl ester 4 (and others) could be performed in a single pot reaction.21 The phosphonate 4 was then treated with a 5-fold excess of hexadecyl bromide in toluene with a catalytic amount of nBu4NI (TBAI) (5 mol%) leading to the hexadecyl phosphonate 11 in 86% yields. Final products 1–12 were fully analyzed and characterized by NMR, HRMS and HPLC.
Lipase inhibition
The 18 new chiral organophosphorus compounds synthesized (i.e. compounds 1, 3 to 5, 11 and 12, as well as trans-(α) and cis-(β) isomers of 2 and 6 to 10; Scheme 1) were further tested for their inhibitory activity toward HPL, DGL, GPLRP2, Cutinase, Rv0183 and LipY. The pH-stat technique41 was used to measure lipase activities and quantify the inhibitory power, defined here as the inhibitor molar excess leading to 50% lipase inhibition (xI50 value).42 Lipase stereopreference was also assessed through the Diastereoselectivity Index (DI)43 between the two diastereoisomers trans-(α) & cis-(β) as defined in Eq. 1 (see Experimental Section).
Inhibition results using cyclic enol-phosphorus compounds
A significant inhibition was obtained with the three microbial lipases tested (Table 2). Cutinase was strongly inactivated by most of the organophosphorous compounds and the efficiency of the inhibition was found to be dependant on the chemical structure of each inhibitor. First, synthetic racemic Cyclophostin 1 (xI50 = 0.96) was 4- to 10-fold more potent than its two homologous phosphonates 2(α) (xI50 = 4.49) and 2(β) (xI50 = 8.77). With these two latter compounds, one can also notice that Cutinase exhibited a slight stereopreference for the trans isomer 2(α) (DI = 32.3%). Regarding the monocyclic enol-phosphonates analogous to Cyclophostin molecules, compounds 3 and 5, bearing respectively an alcohol instead of an ester function and a phenyl group at C-5 position of the oxaphosphepine cycle, did not inhibit Cutinase. The following classification based on the decreasing inhibitory potency was then obtained: 10(β) (xI50 = 1.18) > 4 (xI50 = 1.39) ≅ 6(β) (xI50 = 1.42) ≥ 7(β) (xI50 = 1.98) ≅ 8(β) (xI50 = 2.00) > 9(β) (xI50 = 2.54). In all cases xI50 values were indicative of a very potent inhibition. In addition, Cutinase showed a strong stereopreference for the (β) isomers having a cis conformation. The higher diastereoselectivity (DI > 70.0%) was then found for compounds 6(β), 7(β) and 10(β) bearing C5, C10 and C16 aliphatic alkyl chains, respectively. Finally, the Cyclipostins analogs 11 (xI50 = 0.56) and 12 (xI50 = 0.95) as well as Cyclophostin 1 (xI50 = 0.96) were found to be the best inhibitors of Cutinase. Based on the absence of inhibition observed with compounds 3 and 5 (xI50 > 100), a free or cyclic (lactone) ester group seems therefore required to provide the enol-phosphonates with an inhibitory activity.
Table 2.
Inhibition of Cutinase, Rv0183 and LipY by cyclic enol-phosphorous compounds and Orlistat after 30-min incubation period. a
| Compound | Cutinase | Rv0183 | LipY | |||
|---|---|---|---|---|---|---|
| xI50 | DI | xI50 | DI | xI50 | DI | |
| 1 | 0.96 | - | 5.22 | - | > 100 | - |
| 2(α) | 4.49 | 32.3% | 41.6 | 1.9% | > 100 | NA |
| 2(β) | 8.77 | 43.2 | > 100 | |||
| 3 | > 100 | - | NI | - | NI | - |
| 4 | 1.39 | - | 30.5 | - | > 100 | - |
| 5 | > 100 | - | 16.2 | - | NI | - |
| 6(α) | 26.2 | 89.7% | 3.79 | 16.7% | > 100 | NA |
| 6(β) | 1.42 | 2.70 | > 100 | |||
| 7(α) | 14.6 | 76.2% | 3.57 | 51.9% | 25.8 | 80.3% |
| 7(β) | 1.98 | 1.13 | 2.81 | |||
| 8(α) | 3.63 | 29.0% | 2.44 | 36.3% | 6.14 | 28.0% |
| 8(β) | 2.00 | 5.23 | 3.46 | |||
| 9(α) | 4.43 | 27.2% | > 100 | > 97.0% | 3.47 | 70.5% |
| 9(β) | 2.54 | 1.32 | 0.60 | |||
| 10(α) | 7.26 | 72.1% | > 100 | > 93.0% | 7.47 | 81.3% |
| 10(β) | 1.18 | 3.15 | 0.77 | |||
| 11 | 0.56 | - | NI | - | 0.83 | - |
| 12 | 0.95 | - | NI | - | 0.59 | - |
| Orlistat | 2.52 | - | > 200 | - | 7.07 | - |
xI50 values were determined as described in Experimental Section. Diastereoselectivity Index (DI) between trans-(α) and cis-(β) isomers was calculated for compounds 2 and 6 to 10 using Eq. 1. Results are expressed as mean values of at least two independent assays (CV% < 5.0%). NA = not applicable. NI = no significant inhibition observed.
Concerning Rv0183, the monoglyceride lipase from Mycobacterium tuberculosis,34 Cyclophostin 1 displayed a xI50 value of 5.22 which was nearly 8-fold lower than that of its homologous phosphonates 2(α, β) (mean xI50 = 42.4 ±0.8). Low lipophilic compounds 2(α, β), 4 and 5 also exhibited rather weak inhibitory power with xI50 values ranging from 16.2 to 43.2. Compound 3, which has an alcohol instead of an ester function, was completely inactive toward this lipase. On the contrary, the more lipophilic enol-phosphonates 6 to 10 strongly inhibited Rv0183, with xI50 values in the 1.1 to 5.2 range. Surprisingly, Cyclipostins analogs 11 and 12 were not inhibitors of Rv0183, suggesting that the positioning of the C16–C18 alkyl chains on the phosphorous atom may be responsible for this lack of inhibition. Rv0183 displayed a low diastereoselectivity (DI = 17–52%) between trans-(α) and cis-(β) isomers when using compounds 6 to 8, suggesting that both conformations where similarly recognized when short (C5) and medium (C10–C12) alkyl chain lengths were present at C-5 carbon atom. By contrast, only cis-(β) isomers of compounds 9 and 10 bearing long (C16–C18) alkyl chains were found to be active against the Rv0183 monoglyceride lipase. These findings highlight the fact that the ester function is necessary but not sufficient to inhibit Rv0183. This monoglyceride lipase therefore displayed a strong chemopreference for lipophilic Cyclophostin enol-phosphonate analogs, and a high stereopreference for the cis-(β) isomers bearing long C16–C18 alkyl chain length. Best inhibitors were found to be the Cyclophostin enol-phosphonate analogs 7(β) (xI50 = 1.13) and 9(β) (xI50 = 1.32).
Compounds 1, 2, 4 and 6 exhibited very weak inhibitory properties toward LipY, with 14% to 36% inhibition at a high xI value of 40. No inhibition of LipY was obtained with compounds 3 and 5. Conversely, LipY displayed an unambiguous chemopreference for the most lipophilic phosphonates 7(β) (xI50 = 2.81), 8(β) (xI50 = 3.46), 9(β) (xI50 = 0.60) and 10(β) (xI50 = 0.77), all bearing medium to long alkyl chain lengths. Moreover, and as expected for a member of the HSL family,22 LipY was strongly inactivated by the two monocyclic enol-phosphonates 11 (xI50 = 0.83) and 12 (xI50 = 0.59) analogous to Cyclipostins. Interestingly, no significant differences in xI50 values were observed between the Cyclophostin analogs 9(β) and 10(β) (mean xI50 = 0.69 ±0.08), and phosphonates 11 and 12 (mean xI50 = 0.71 ±0.12). Such result suggests that the presence of a long C16–C18 lipophilic alkyl chain is essential for high inhibition to take place, whatever its positioning: either on the C-5 carbon atom of the oxaphosphepine cycle (i.e., in 9(β) and 10(β)) or on the phosphorus atom (i.e., in 11 and 12). Like Cutinase and Rv0183, LipY exhibited a high diastereoselectivity (28.0 ≤ DI ≤ 81.3%) for the cis-(β) isomers. The best inhibitors of LipY were then Cyclipostin analogs 11 (xI50 = 0.83) and 12 (xI50 = 0.59) as well as Cyclophostin analogs 9(β) (xI50 = 0.60) and 10(β) (xI50 = 0.77).
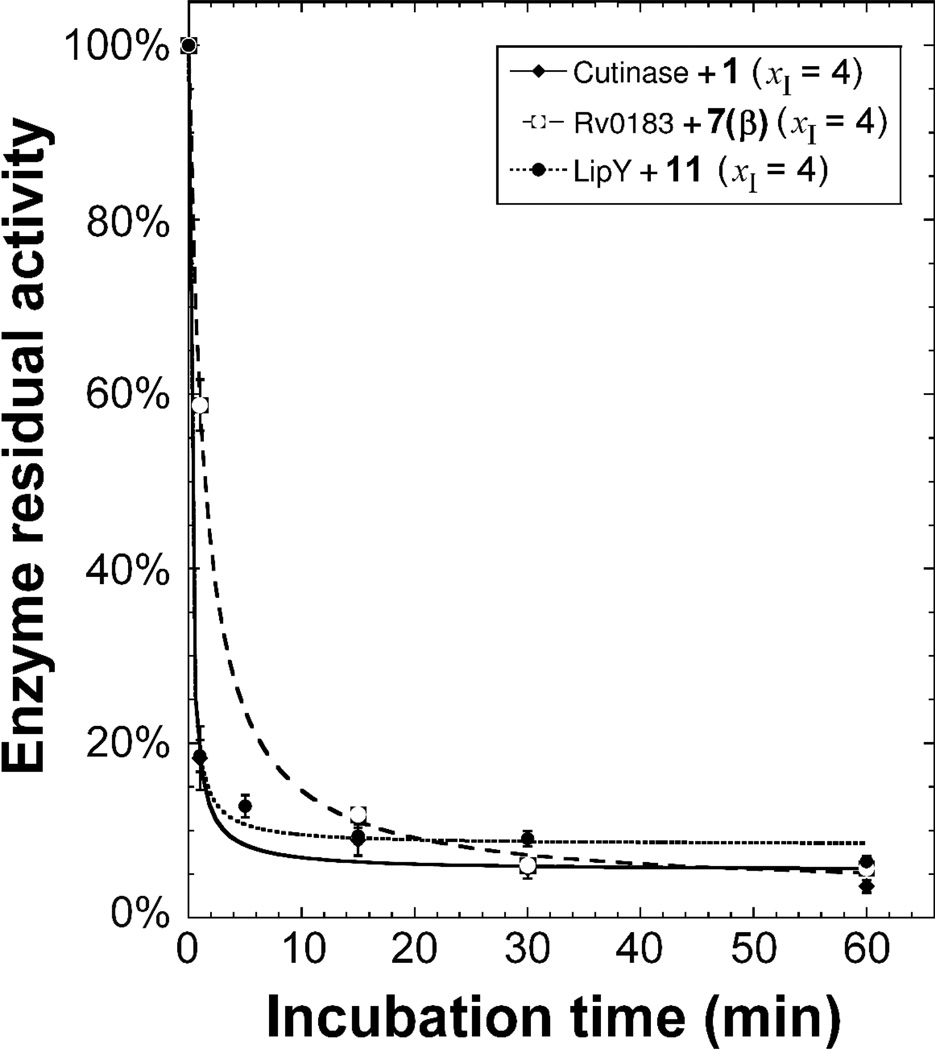
The influence of the incubation time on the levels of inhibition of Cutinase, Rv0183 and LipY by inhibitors 1, 7(β) and 11 was then investigated. As depicted in Figure 2, the residual activity of all three enzymes decreased rapidly and reached a plateau value at around 6.0% residual lipase activity after approximately 30 min of incubation. From these inhibition curves, the values of the half-inactivation time (t1/2) were determined and found to be 0.18 min for 1 acting on Cutinase, 1.44 min for 7(β) acting on Rv0183 and 0.16 min for 11, acting on LipY. Such values thus reflect an extremely high rate of inhibition of each lipase by the corresponding phosphonate compounds.
Figure 2.
Effect of the incubation time on the inhibition level of Cutinase (♦), Rv0183 (□) and LipY (●) by compounds 1, 7(β) and 11, respectively. Each enzyme was pre-incubated with the corresponding phosphonate compound at a fixed inhibitor molar excess xI = 4, and the residual activity was measured over a 60-min. period at various time intervals using the pH-stat technique. Results are expressed as mean values of at least two independent assays (CV% < 5.0%).
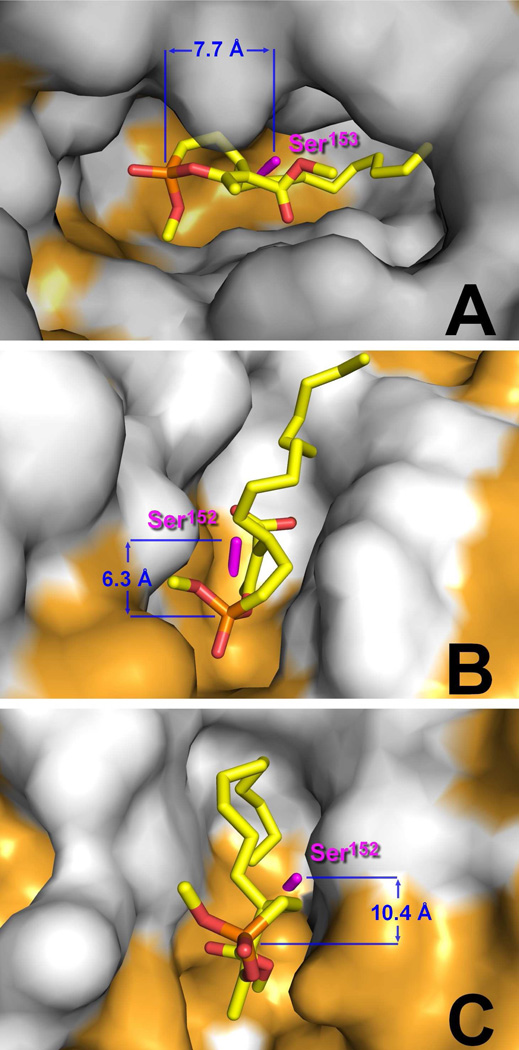
In order to compare the inhibitory activity of these organophosphorous compounds with that of Orlistat, the same kinetic parameters were also measured with this inhibitor and each of the tested lipases. The inhibitory effects of Orlistat towards Cutinase (xI50 = 2.52), LipY (xI50 = 7.07) and Rv0183 (xI50 > 200) was found to be 2.6- to 177-fold lower than that of compounds 1 (xI50 = 0.96), 11 (xI50 = 0.83) and 7(β) (xI50 = 1.13) acting on the same lipases, respectively. Conversely, while Orlistat is a potent inhibitor of mammalian lipases, i.e. HPL (xI50 = 18.4), GPLRP2 (xI50 = 0.53) and DGL (xI50 = 5.60); these lipases are not inhibited by all the enol-phosphorous compounds tested, even by using high inhibitor molar excess (xI = 200) and after 2 hours of incubation period. To further investigate such absence of inhibition experimentally observed with these cyclic enol-phosphorus compounds, in silico molecular docking experiments were conducted on 7(β) with the reported crystal structures of DGL,44 HPL45 and GPLRP225 (Figure 3). In all cases, the best scoring-positions obtained (i.e. lowest energy complex) showed that the reactive oxaphosphepine ring adopted a non-productive orientation. The phosphorus atom of 7(β) was indeed found in an opposite and unfavorable position preventing the occurrence of any reactions with the catalytic serine ([Ser-Oγ / P] distances > 6.0 Å) and thus the formation of a covalent bond.
Figure 3.
In silico molecular docking of compound 7(β) into crystallographic structures of (A) DGL (PDB: 1K8Q), (B) HPL (PDB: 1LPB) and (C) GPLRP2 (PDB: 1GPL) in a van der Waals surface representation. Hydrophobic residues (alanine, leucine, isoleucine, valine, tryptophan, tyrosine, phenylalanine, proline and methionine) are highlighted in white. The side chain of catalytic serine residue is shown as sticks and colored in magenta. Compound 7(β) is represented in atom color-code sticks (carbon, yellow; phosphorus, orange, oxygen, red). Structures were drawn using the PyMOL Molecular Graphics System (Version 1.3, Schrödinger, LLC). The [Ser-Oγ / P] calculated distances are indicated in blue.
Mass spectrometry analysis of inhibitor–modified enzymes
To investigate whether the inhibition with cyclic enol-phosphonates occurring with microbial lipases is due to the formation of a covalent bond with the catalytic serine, mass spectrometry analyses were further conducted on Cutinase, Rv0183 and LipY inhibited by compounds 4, 7(β) and 11, respectively. Samples of the lipase-inhibitor complexes were then subjected to MALDI-TOF mass spectrometry analyses. The obtained m/z mass values are summarized in Table 3.
Table 3.
Mass spectrometry analysis of lipases inhibited by monocyclic enol-phosphonate inhibitors a
| Species |
m/z native lipase (❶) |
m/z inhibited lipase (❷) |
Mass modifications (❷−❶) |
|---|---|---|---|
| Cutinase | 22,187.9 Da | 22,420.8 Da | +232.9 Da |
| Rv0183 | 30,177.4 Da | 30,494.2 Da | +376.8 Da |
| LipY | 47,145.6 Da | 47,589.4 Da | +443.8 Da |
Cutinase, Rv0183 and LipY were pre-incubated for 30 min at 25 °C with compound 4 (Mw, 234.2 Da), 7(β) (Mw, 374.4 Da) and 11 (Mw, 444.3 Da) respectively, at a xI value of 40, prior to MALDI-TOF mass spectrometry analysis.
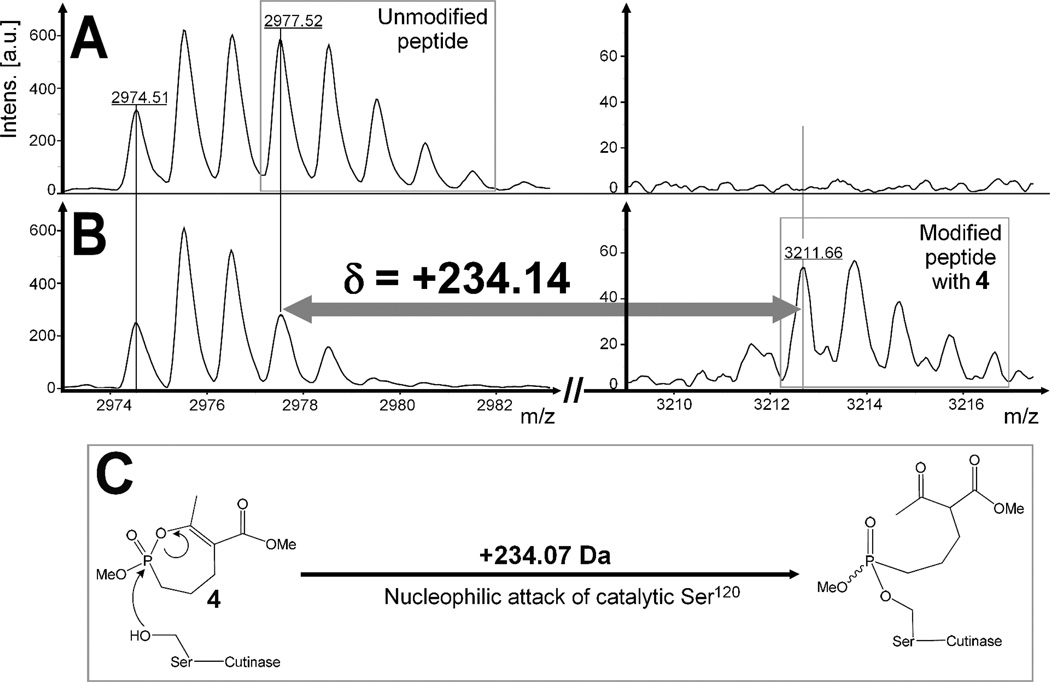
In presence of inhibitor, a shift in the molecular mass of each inhibited protein was observed, corresponding to the molecular mass of the inhibitor and reflecting the covalent binding of 4 (Mw, 234.2 Da) to Cutinase, of 7(β) (Mw, 374.4 Da) to Rv0183, as well as of 11 (Mw, 444.3 Da) to LipY. Moreover, in order to confirm the mechanism of action of these phosphonate compounds and consequently their specific covalent binding via a nucleophilic attack of the catalytic serine residue to the phosphorus center; as initially argued with human AChE;16 tryptic digestion of the Cutinase-4 complex was performed. The peptide mass fingerprint (PMF)46 was then established by MALDI-TOF mass spectrometry. Peptide C109-R138 (C109PDATLIAGGYS120QGAALAAASIEDLDSAIR138) containing the catalytic Ser120 residue, was detected in the native Cutinase as a multiplet at a molecular mass of around 2,977.5 Da (Figure 4A). This catalytic peptide appeared concomitantly with a second multiplet peptide (V169-R196) detected at a molecular mass of around 2,974.5 Da. When Cutinase was complexed with compound 4, a mass increase of +234.14 Da was obtained for the catalytic peptide only (Figure 4B), while multiplet peptide V169-R196 remained unchanged. The disappearance of 2,977.5 Da peptide in the inhibited lipase followed by the appearance of a new one at m/z of 3,211.7 Da, confirmed that a covalent bond had been formed between the enol-phosphonate 4 and the catalytic serine residue of Cutinase, in accordance with the reaction mechanism depicted in Figure 4C.
Figure 4.
Peptide mass fingerprint spectra of the digested Cutinase, before (upper panels, A) and after (lower panels, B) a 30-min incubation period with compound 4 at an inhibitor molar excess xI = 40; and (C) the corresponding confirmed reaction mechanism with the active site Ser120. (A–B): Zoom on the region of interest. Left panels, region of the unmodified peptide C109-R138 (C109PDATLIAGGYS120QGAALAAASIEDLDSAIR138) containing the catalytic Ser120 and detected at 2,977.52 Da. This multiplet peak is overlaid with a first one corresponding to peptide V169-R196, detected at 2,974.51 Da. Right panels, region in which a new peptide was expected to occur resulting from the covalent binding of compound 4 to the catalytic serine. Mass shifts were calculated as the difference between the experimental m/z of the modified peptide and the theoretical m/z of the unmodified catalytic peptide. The expected exact mass shift is +234.07 Da.
DISCUSSION
Cyclophostin and the Cyclipostins are known to be very potent natural organophosphate-based inhibitors of AChE20 and HSL,22 respectively. Variation of the nature of the alkyl group at the phosphate center, methyl vs. long lipophilic alkyl groups, leads to a dramatic switch of the enzyme selectivity by these organophosphates. This particular fact provides a unique opportunity to investigate the enzyme selectivity of this class of compounds or their analogs, based on structure modification of the parent molecule. In that way, phosphonates have garnered a reputation for being excellent structural mimic of natural phosphates. It is also well known that for phosphonate compounds, the mode of inhibition involves the formation of a non-hydrolysable covalent bond between the phosphorus and the catalytic Oγ serine residue of the enzyme active site.19 For this purpose, novel synthetic routes were proposed and a full library of new monocyclic enol-phosphonate compounds, analogous to either Cyclophostin or Cyclipostins (Schemes 1 and 2), has been synthesized.
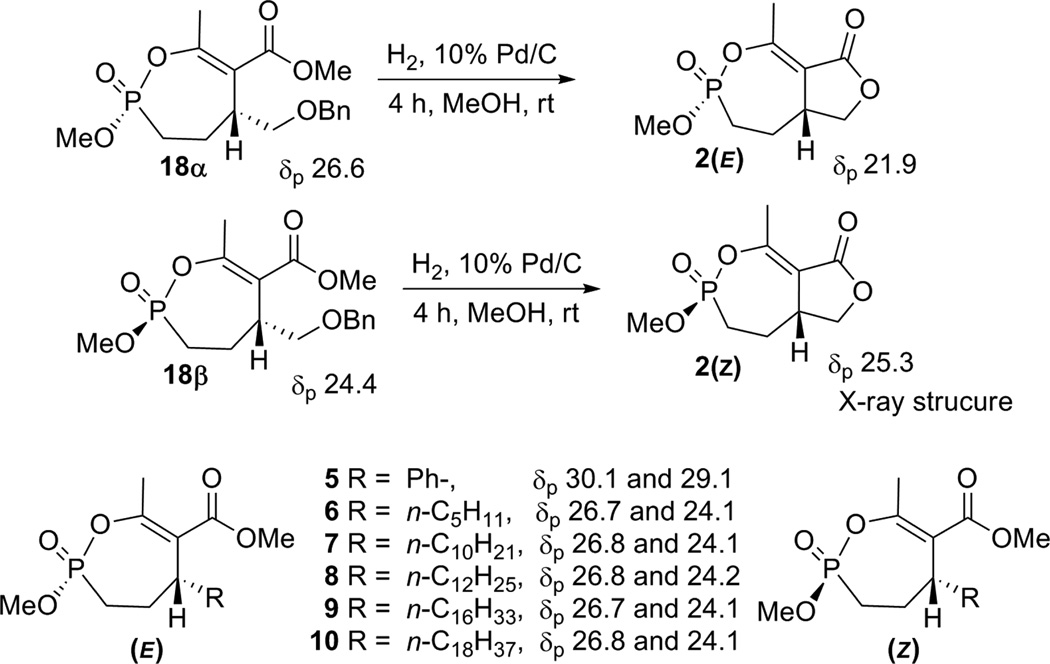
The strategy applied; involving palladium(0)-catalyzed addition of methyl acetoacetate to phosphono allylic carbonates and selective mono-demethylation and cyclization; allowed for the custom synthesis of monocyclic phosphonates substituted at the phosphorous atom or at the C-5 position. The monocyclic phosphonates 6–10 were formed as mixtures of diastereoisomers which were further easily separated and purified. The fast eluting (α) isomer had a signal in the 31P NMR spectrum at 26.8 ppm, whereas the slower eluting (β) isomer had a signal at 24.2 ppm, consistent with all examples (6–10) studied. We had earlier demonstrated in the synthesis of the racemic bicyclic phosphonate analogs of cyclophostin 2(α) and 2(β)15 that the monocyclic phosphonate isomer 18β with the upfield shift (24.2 ppm) had the phosphonate methoxy and the 5-hydrogen in a cis conformation (5-alkyl substituent is therefore trans to the methoxy) (Scheme 3). Consequently, the isomer 18α with the down field shift (26.6 ppm) had the phosphonate methoxy and the 5-hydrogen substituent trans to each other.
Scheme 3.
Relative Stereochemistry of the monocyclic phosphonates 5–10.
From these experimental observations, combined with the obtained single crystal X-ray diffraction structure of 10(α) (Figure 1), it was then possible to assign without any ambiguity the trans-α and cis-β relative stereochemistry of the monocyclic phosphonates 5–10.
Overall, the chemical pathways developed here make possible the synthesis of large libraries of fully customizable enol-phosphorous compounds with controlled cis or trans diastereochemistry.
Based on their structural properties previously depicted in Scheme 1, these organophosphorous compounds have been tested for their abilities to inhibit six lipolytic enzymes, including mammalian digestive lipases (HPL, DGL, and GPLRP2) and three microbial lipases (Cutinase, Rv0183 and LipY). In view of the specific mechanism of action of these lipolytic enzymes,27 the Michaelis-Menten–Henri model no longer applies47 and therefore the Km, Ki and IC50 values often estimated for lipases are irrelevant when insoluble substrates and/or inhibitors are involved. The use of more appropriate kinetic constants such as the inhibitor molar excess leading to 50% lipase inhibition, i.e. xI50 value,42 is thus recommended when assessing the inhibitory potency of insoluble inhibitors, such as these cyclic enol-phosphorus derivatives. Thereby, a xI50 value of 0.5 is synonymous with a 1:1 stoichiometric ratio between the inhibitor and the lipase, and is therefore the highest level of inhibitory activity that can be achieved. Regarding lipase stereopreference, this parameter has also been studied on the basis of the Diastereoselectivity Index (DI)43 between the two diastereoisomers trans-(α) & cis-(β) (see Eq. 1 in Experimental Section).
Regarding the three microbial lipases, Cutinase, Rv0183 and LipY were all inhibited by the cyclic enol-phosphorous compounds. The fused butyrolactone did not appear to be necessary for activity since the monocyclic diastereomers showed similar or even better xI50 values than those of the bicycles 1 and 2(α, β). However, the presence of ester functionality in conjugation with the enol-phosphate or phosphonate bond was found essential to the biological activity. This effect was well illustrated when comparing 3 with 4. These two compounds only differ at the exocyclic functionality: the alcohol in 3 is a poor leaving group and 3 is an extremely poor inhibitor, while compound 4 with the ester group is more potent. This phenomenon, which depicts the electron withdrawing contribution of the ester moiety in increasing the leaving group characteristics of the enol from the P–center, is in line with mass spectrometry analyses confirming that the inhibition is due to phosphorylation of the catalytic serine residue for each enzyme, as depicted in Figure 4C. Such findings demonstrate that the enol-phosphonate really acts as a leaving group on reaction with the active site serine, leading to the formation of a stoichiometric covalent enzyme-inhibitor complex. The fact that comparable results had also been observed when testing the inhibitory properties of the phosphonate analogs of Cyclophostin (i.e., compounds 1 to 4) against the housefly AChE (CSMA strain) therefore suggest a similar mode of action of this enzyme.15,16,21 The trans-2(α) diastereoisomer of bicyclic phosphonate analogs of Cyclophostin was also found to be almost 10 times more potent than the cis-2(β) stereoisomer when tested against human AChE.15 Based on these previous experimental kinetic data, it was suggested that the diastereopreference could be due to some specific initial contacts between the inhibitor and the active site amino acid residues, which ultimately led to a successful inhibition.16 In the present study, LipY was barely affected by the bicyclic molecules 1 and 2(α, β), while Rv0183 did not discriminate between the two conformations (DI = 1.9%) and Cutinase exhibited a slight stereopreference (DI = 32.3%) for the trans isomer 2(β). These results illustrate the difference in selectivity exerted by the three lipases. Indeed, although the best monocyclic phosphonate inhibitors all displayed a cis conformation (i.e. (β) isomers), the group on the γ-carbon of the phosphonate ring significantly affect and modify the biological activity of the monocyclic analogs. Such structural changes lead to more selective inhibitors towards the microbial lipases tested. Rv0183 reacts indeed selectively with Cyclophostin analogs bearing medium to long alkyl chains; while LipY is only fully inactivated by highly lipophilic compounds. Cutinase, however, is found less selective since it reacts with most of the organophosphorous compounds without any real chemopreference (Table 2). Such a high and stringent chemoselectivity exerted by Rv0183 and LipY, or conversely the lower one in the case of Cutinase, is consistent with the respective substrate specificity of these three lipolytic enzymes.32–34,36,37
Comparisons with the potent but non selective lipase inhibitor Orlistat were very instructive. Indeed, the inhibitory power exerted by compounds 11, 12 and 7(β) on Cutinase, LipY and Rv0183, respectively, was 4.5 to 176 times higher than that of Orlistat acting on the same lipases. Moreover, with xI50 values as low as 0.56 for 11 acting on Cutinase, which is close to the lowest achievable value (the theoretical minimum value being 0.50); these phosphorous compounds react in a quasi-stoichiometric way with these enzymes. By contrast, the enol-phosphonates had no effects on HPL, DGL and GPLRP2; while these mammalian lipases were strongly inhibited by Orlistat. In silico molecular docking experiments with 7(β) suggest that these 7-membered ring phosphorus molecules can enter the active site of HPL, DGL and GPLRP2, but cannot interact with the catalytic serine residue. In each case, the lipophilic alkyl chains of the best scoring-positions of 7(β) would be stabilized by hydrophobic residues located near the active site (Figure 3, white color). In this binding mode, however, reactive oxaphosphepine ring of 7(β) would adopt a non-productive orientation: the phosphorus atom is indeed found in an opposite position making it impossible to react with the catalytic serine.
Taken together, all these findings suggest that this series of enol-phosphonates 6–12 may be selective inhibitors of these families of microbial lipolytic enzymes; i.e. Cutinase, monoglyceride lipase and hormone sensitive lipase families.
It has been shown recently that the use of Orlistat could significantly reduce the growth of Mycobacterium smegmatis10 a non-pathogen Mycobacterium species, and of Mycobacterium bovis BCG9 which is closely related to the pathogen Mycobacterium tuberculosis. In addition, some recent papers pointed out the fact that Orlistat could also target potentially essential enzymes involved either in the mycobacteria cell wall biosynthesis such as the Cutinase-like Rv3802c,11 or in the hydrolysis of stored triacylglycerols during persistence and reactivation phases such as LipY.36 Fighting tuberculosis is still challenging, not only in third world countries but also in the so-called industrialized world, due to the impressive recrudescence of this major infectious disease. In that context, our encouraging results on the selective inhibition of Cutinase as well as of pure microbial lipolytic enzymes from Mycobacterium tuberculosis suggest that this new class of phosphonate analogs of Cyclophostin and Cyclipostins could be potential scaffolds and may provide useful leads for the further development of more selective anti-microbial agents, including anti-mycobacterial compounds. In particular, thanks to our synthetic routes, we have clearly showed that modulation of the lipophilicity and/or functionalization at the C-5 carbon atom or at the phosphorus can be exploited either to attenuate or increase the affinity of one inhibitor to target a specific microbial lipase.
CONCLUSION
This work provided the first fundamental premise for the understanding of the structure-activity relationships of monocyclic enol-phosphonate analogs in connection with the inhibition of several lipases having distinct substrate specificities. Moreover, the major interest in this new family of phosphonate inhibitors is that their chemical structure is flexible at will. The strategy developed here; involving palladium(0)-catalyzed addition of methyl acetoacetate to phosphono allylic carbonates followed by selective mono-demethylation and cyclization; allows for the synthesis of a large variety of fully customizable monocyclic enol-phosphonates substituted at the C-5 position. Furthermore, the methyl phosphonate ester can be easily trans-esterified to give long chain ester compounds analogous to Cyclipostins. This flexibility makes it possible to design selective inhibitors for a given enzyme. In order to shed more light on the influence of the chirality at both phosphorus and carbon centers on lipase inhibition, further experiments are currently under way and will be reported in due course.
EXPERIMENTAL SECTION
Chemistry - Synthesis of compounds 1 to 12
General Experimental
All reactions were carried out in oven dried glassware under an atmosphere of argon unless otherwise noted. 1H, 13C and 31P NMR spectra were recorded at 300, 75 and 121 MHz respectively. 1H NMR spectra are referenced to CDCl3 (7.27 ppm), 13C NMR spectra are referenced to the center line of CDCl3 (77.23 ppm) and 31P NMR spectra are referenced to external H3PO4. Coupling constants, J, are reported in Hz. Analytical thin-layer chromatography (TLC) analyses were performed on silica gel plates 60PF254. Visualization was accomplished with UV light, KMnO4 solution or iodine. The synthesis of compounds racemic 1,21 racemic 2(α) and 2(β),15 3 and 416 as well as carbonates 13a, 13b and 13c38 have already been reported. Target compounds were purified by chromatography and have purity of ≥ 95% as determined by HPLC analysis conducted on an Agilent 1100 system, using a Zorbax Eclipse XBD-C18 column. Elution was performed at a flow rate of 2 mL.min−1 using a gradient of H2O/TFA/CH3CN 95:0.1:5 to H2O/TFA/CH3CN 5:0.1:95 over 6 min; or using a gradient of H2O/TFA/CH3CN 95:0.1:5 to 100% CH3CN over 5 min and then hold at 100% CH3CN for 3 min. The compounds were detected using a diode array and ELS. The HPLC data was supported by careful analysis of the 1H, 13C and particularly the 31P NMR spectra.
General procedure for synthesis of the carbonates 13 via cross metathesis reaction.39
To a solution of carbonate 13a (1.0 mmol) and alkene (5.0–10.0 mmol) in CH2Cl2 (2 mL) was added Grubbs 2nd generation catalyst (5 mol %) followed by CuI (7 mol %). After stirring the solution for 2 min at room temperature, the septum was replaced by a condenser connected to an argon bubbler and the reaction flask was placed in an oil bath preheated to 45°C. After completion of the reaction (TLC and 31P NMR analysis), the solvent was removed under reduced pressure to give the crude product. Purification of the crude product by column chromatography (SiO2, 20–50% EtOAc in hexanes) gave the carbonate 13 (E:Z ≥10:1).
(±) (E)-1-(dimethoxyphosphoryl)tridec-2-en-1-yl methyl carbonate (13d)
To a solution of carbonate 13a (1.12 g, 5.00 mmol) and 1-dodecene (11.1 mL, 50.0 mmol) in dry CH2Cl2 (10 mL) was added Grubbs 2nd generation catalyst (0.21 g, 0.25 mmol) followed by CuI (0.07 g, 0.35 mmol) to give 13d (1.43 g, 79%). IR (neat, NaCl) 1756 cm−1; 1H NMR (CDCl3) δ 5.97 (1H, ddt, JHH = 15.2, 6.7 Hz, JHP = 3.7 Hz), 5.55 (1H, ddd, JHH = 15.2, 7.6 Hz, JHP = 5.2 Hz), 5.45 (1H, dd, JHH = 8.0 Hz, JHP = 12.4 Hz), 3.81 (d, 3H, JHP = 10.6 Hz), 3.81 (3H, s), 3.8 (d, 3H, JHP = 10.6 Hz), 2.08 (2H, ddd, JHH = 13.9, 6.6 Hz, JHP = 3.4 Hz), 1.37 (2H, t, JHH = 6.7 Hz), 1.25 (16H, br s), 0.87 (3H, t, JHH = 6.6 Hz); 13C NMR (CDCl3) δ 155.0 (d, JCP = 9.9 Hz), 139.4 (d, JCP = 12.5 Hz), 120.3 (d, JCP = 3.8 Hz), 73.3 (d, JCP = 170.2 Hz), 55.5, 54.0 (d, JCP = 7.2 Hz), 32.6, 32.1, 29.8, 29.6, 29.5, 29.3, 28.8 (d, JCP = 2.4 Hz), 22.9, 14.3; 31P NMR (CDCl3) δ 20.8, 20.5 (E:Z = 10:1); HRMS (FAB, NBA, MH+) calcd. for C17H34O6P: 365.2092, found 365.2103.
(±) (E)-1-(dimethoxyphosphoryl)pentadec-2-en-1-yl methyl carbonate (13e)
To a solution of carbonate 13a (1.12 g, 5.00 mmol) and 1-tetradecene (12.7 mL, 50.0 mmol) in dry CH2Cl2 (10 mL) was added Grubbs 2nd generation catalyst (0.21 g, 0.25 mmol) followed by CuI (0.07 g, 0.35 mmol) to give 13e (1.5 g, 76%). IR (neat, NaCl) 1755 cm−1; 1H NMR (CDCl3) δ 5.95 (1H, ddt, JHH = 15.2, 6.7 Hz, JHP = 3.8 Hz), 5.55 (1H, ddd, JHH = 15.2, 7.6 Hz, JHP = 5.2 Hz), 5.45 (1H, dd, JHH = 7.8Hz, JHP = 12.4 Hz), 3.81 (d, 3H, JHP = 10.6 Hz), 3.81 (3H, s), 3.80 (d, 3H, JHP = 10.6 Hz), 2.08 (2H, ddd, JHH = 13.9, 6.7 Hz, JHP = 3.5 Hz), 1. 37 (2H, t, JHH = 6.7), 1.24 (18H, br s), 0.87 (3H, t, JHH = 6.6 Hz); 13C NMR (CDCl3) δ 154.9 (d, JCP = 9.8 Hz), 139.3 (d, JCP = 12.5 Hz), 120.3 (d, JCP = 3.8 Hz), 73.2 (d, JCP = 170.3 Hz), 55.5, 54.0 (d, JCP = 7.2 Hz), 32.6, 32.1, 29.9, 29.8, 29.7, 29.6, 29.5, 29.4, 29.3, 28.8 (d, JCP = 2.4 Hz), 22.9, 14.3; 31P NMR (CDCl3) δ 20.9, 20.5 (E:Z = 8:1); HRMS (FAB, NBA, MH+) calcd for C19H37O6PNa: 415.2225, found 415.2227.
(±) (E)-1-(dimethoxyphosphoryl)nonadec-2-en-1-yl methyl carbonate (13f)
To a solution of carbonate 13a (1.12 g, 5.00 mmol) and 1-octadecene (8.0 mL, 25 mmol) in dry CH2Cl2 (10 mL) was added Grubbs 2nd generation catalyst (0.21 g, 0.25 mmol) followed by CuI (0.07 g, 0.35 mmol) to give 13f (1.57g, 70%). IR (neat, NaCl), 1755 cm−1; 1H NMR (CDCl3) δ 5.96 (1H, ddt, JHH = 14.6, 7.1 Hz, JHP = 3.8 Hz), 5.55 (1H, ddd, JHH = 7.7, 15.2 Hz, JHP = 5.2 Hz), 5.44 (1H,dd, JHH = 7.8 Hz, JHP = 12.3 Hz), 3.82 (3H, s), 3.81 (d, 3H, JHP = 10.6 Hz), 3.81 (d, 3H, JHP = 10.6 Hz), 2.1 (2H, ddd, JHH = 7, 13.7 Hz, JHP = 3.2 Hz), 1.37 (2H, t, JHH = 6.5 Hz), 1.25 (26H, br s), 0.87 (3H, t, JHH = 6.44 Hz); 13C NMR (CDCl3) δ 155 (d, JCP = 9.8 Hz), 139.3 (d, JCP = 12.4 Hz), 120.3 (d, JCP = 3.8 Hz), 73.3 (d, JCP = 170.3 Hz), 55.5, 54.0 (d, JCP = 7.3 Hz), 32.6, 32,1, 29.9, 29.8, 29.7, 29.6, 29.5, 29.3, 28.8, 28.7, 22.9, 14.4; 31P NMR (CDCl3) δ 20.8, 20.5 (E:Z = 10:1); HRMS (FAB, NBA, MNa+) calcd for C23H45O6PNa: 471.2851, found 471.2860.
(±) (E)-1-(dimethoxyphosphoryl)henicos-2-en-1-yl methyl carbonate (13g)
To a solution of carbonate 13a (0.67 g, 3 mmol) and 1-eicocene (4.21 mL, 15 mmol) in dry CH2Cl2 (6 mL) was added Grubbs 2nd generation catalyst (0.13 g, 0.15 mmol) followed by CuI (0.04 g, 0.21 mmol) to give 13g (1.1 g, 70%). IR (neat, NaCl) 1755 cm−1; 1H NMR (CDCl3) δ 5.97 (1H, ddt, JHH = 14.9, 6.7 Hz, JHP = 3.8 Hz), 5.56 (1H, ddd, JHH = 15.1, 7.7 Hz, JHP = 5.3 Hz), 5.46 (1H, dd, JHH = 7.8 Hz, JHP = 12.2 Hz), 3.82 (3H, s), 3.81 (d, 3H, JHP = 10.6 Hz), 3.81 (d, 3H, JHP = 10.6 Hz), 2.1 (2H, JHH = 6.7, 13.7 Hz, JHP = 3.4 Hz), 1.39 (2H, t, JHH = 6.7 Hz), 1.26 (30 H, br s), 0.88 (3H, t, JHH = 6.7 Hz); 13C NMR (CDCl3) δ 155.0 (d, JCP = 9.8 Hz), 139.4 (d, JCP = 12.5 Hz), 120.3 (d, JCP = 3.8 Hz), 73.3 (d, JCP = 170.3 Hz), 55.5, 54.0 (d, JCP = 7.1 Hz), 32.6, 32,1, 30, 29.9, 29.8, 29.6, 29.5, 29.3, 28.8 (d, JCP = 2.4Hz), 22.9, 14.4; 31P NMR (CDCl3) δ 20.8, 20.5 (E:Z = 10:1); HRMS (FAB, NBA, MH+) calcd. for C25H49O6PNa: 499.3164, found 499.3170.
General procedure for synthesis of vinyl phosphonates 14
To a stirring solution of carbonate 13 (1 mmol) and methyl acetoacetate (3 mmol) in dry THF (2.5 mL) was added Pd2(dba)3 (0.02 mmol) followed by dppe (0.5 mmol) and the resulting mixture was stirred at room temperature for 3–4 min. The septum was replaced by a condenser connected to an argon bubbler and the reaction flask was placed in an oil bath preheated to 45 °C. After completion of the reaction (TLC and 31P NMR analysis), the reaction mixture was partitioned between brine and EtOAc. The organic layer was collected and the aqueous layer was extracted with additional EtOAc. The combined organic layers were dried over anhydrous Na2SO4 and evaporated under reduced pressure to give crude product. Purification of the crude by column chromatography (SiO2, 20–70% EtOAc in hexanes) gave the vinyl phosphonate 14.
(±) (E)-methyl 2-acetyl-5-(dimethoxyphosphoryl)-3-phenylpent-4-enoate (14b) and (2Z,3Z)-methyl 5-(dimethoxyphosphoryl)-2-(1-hydroxyethylidene)-3-phenylpent-3-enoate (15)
To a solution of carbonate 13b (0.75 g, 2.5 mmol) and methyl acetoacetate (0.87 mL, 7.6 mmol) in THF (6 mL) was added Pd2(dba)3 (0.05 g, 0.05 mmol) and dppe (0.05 g, 0.13 mmol) to give 14b as 1:1 diastereoisomeric mixture (0.32 g, 46%). Pale yellow solid; IR (neat), 1742, 1714 cm−1; 1H NMR (CDCl3) δ 7.26 (m, 5H), 6.73 (1H, m), 5.75 (1H, m), 4.28 (1H, m), 4.1(1H, d, JHH = 11.27 Hz), 3.66 (6H, m). 3.61 (3H, s), 2.14 (3H, s); 13C NMR (CDCl3) δ 200.4 (200.3), 167.7 (167.3), 152.3 (152.26) (d, JCC = 5.2 Hz), 137.9 (137.6), 129.1 (128.9), 128.2 (128.1), 128.1, 127.8 (127.6), 117.8 (117.5) (d, JCP = 185.4 Hz), 64.1 (63.5), 52.7 (52.5), 52.3 (d, JCP = 5.7 Hz), 49.3 (48.9) (d, JCP = 22.2 Hz), 30.4 (30.1); 31P NMR (CDCl3) δ 20.87, 20.1 ppm; HRMS (FAB, NBA, MH+) calcd. for C16H21O6P: 340.1153, found 341.1157 and 15 (0.34 g, 47%). White solid; IR (neat) 1646, 1611 cm−1; 1H NMR (CDCl3) δ 13.02, (1H, s), 7.37 – 7.25 (5H, m), 6.18 (1H, JHH = 7.6 Hz), 3.76 (6H, d, = 10.87 Hz). 3.71 (3H, s), 2.7 (2H, dd, JHH = 7.6 Hz, JHP = 22.6 Hz), 1.88 (3H, s); 13C NMR (CDCl3) δ 175.5, 172.9, 140.7, 138.2, 128.9, 128.0, 126.3, 121.1, 99.4, 53.0 (d, JCP = 11.43 Hz), 52.2, 27.9 (d, JCP = 139.88 Hz), 19.8; 31P NMR (CDCl3) δ 30.46; HRMS (FAB, NBA, MH+) calcd. for C16H22O6P: 340.1153, found 341.1150
(±) (E)-methyl 2-acetyl-3-(2-(dimethoxyphosphoryl)vinyl)octanoate (14c)
To a solution of carbonate 14c (0.29 g, 1.0 mmol) and methyl acetoacetate (0.33 mL, 3.0 mmol) in dry THF (2 mL) was added Pd2(dba)3 (0.02 g, 0.02 mmol) and dppe (0.02 g, 0.05 mmol) to give 14c as 1:1 mixture of diastereoisomers (1.17 g, 70%). Pale yellow oil; IR (neat, NaCl) 1743, 1717; 1H NMR (CDCl3) δ 6.55 (0.5H, ddd, JHH = 17.13, 9.5 Hz, JHP = 21.5 Hz), 6.45 (0.5H, ddd, JHH = 17.13, 9.5 Hz, JHP = 21.5 Hz), 5.68 (0.5H, dd, JHH = 17.13, JHP = 21.5 Hz), 5.61 (0.5H, ddd, JHH = 17.13 Hz, JHP = 21.5 Hz), 3.67 (9H, m), 3.51 (1H, 2d, JHH = 9.4, 9.3 Hz), 2.95 (1H, 2q, JHH = 9.4 and 9.4 Hz), 2.21 (1.5H, s), 2.15 (1.5H, s), 1.28 (8H, m); 0.83 (3H, t, JHH = 6.44 Hz); 13C NMR (CDCl3) δ 201.3, 168.6 (168.4), 153.4 (153.3) (d, JCP = 4.6 Hz), 118.8 (118.7) (d, JCP =184.5 Hz), 63.8 (63.7) (d, JCP = 1.0 Hz), 52.7 (52.5), 52.4 (d, JCP = 5.6 Hz), 44.0 (43.9) (d, JCP = 21.8 Hz), 31.8 (31.6), 31.5 (d, JCP = 1.3 Hz), 30.1 (29.9), 26.8 (26.7), 22.5, 14.0; 31P NMR (CDCl3) δ 20.7, 20.6; HRMS (FAB, NBA, MH+) calcd. for C15H27O6PNa: 357.1442, found 357.1450.
(±) (E)-methyl 2-acetyl-3-(2-(dimethoxyphosphoryl)vinyl)tridecanoate (14d)
To the stirring solution of carbonate 13d (0.182 g, 0.50 mmol) and methyl acetoacetate (0.17 mL, 1.5 mmol) in dry THF (1mL) was added Pd2(dba)3 (0.01 g, 0.01 mmol) followed by dppe (0.10 g, 0.025 mmol) to give 14d as a 1:1 mixture of diastereoisomers (0.16 g, 78%). Pale yellow oil; IR (neat, NaCl), 1743, 1719 cm−1; 1H NMR (CDCl3) δ 6.59 (0.5H, ddd, JHH = 17.1, 9.6 Hz, JHP = 21.5 Hz), 6.49 (0.5H, ddd, JHH = 17.1, 9.6 Hz, JHP = 21.5 Hz), 5.7 (0.5H, dd, JHH = 17.1 Hz, JHP = 21.5 Hz), 5.65 (0.5H, dd, JHH = 17.1Hz, JHP = 21.5 Hz), 3.67 (9H, m), 3.54 (1H, 2d, JHH = 9.4 and 9.2 Hz), 2.98 (1H, 2q, JHH = 9.3 Hz), 2.24 (1.6H, s), 2.18 (1.4H, s), 1.23 (30H, br s), 0.87 (3H, t, JHH = 6.7 Hz); 13C NMR (CDCl3) δ 201.5, 168.7 (168.5), 153.5 (d, JCP = 9.1 Hz), 118.8 (d, JCP = 171.7 Hz), 63.9 (63.8), 52.8 (52.7), 52.5 (d, JCP = 5.4 Hz), 44.3 (44), 32.1, 31.9 (31.8), 30.3 (30.1), 29.8, 29.6, 29.5, 27.3 (27.20), 22.9, 14.3; 31P NMR (CDCl3) δ 20.7, 20.6; HRMS (FAB, NBA, MH+) calcd. for C20H38O6P: 405.2406, found 405.2403.
(±) (E)-methyl 2-acetyl-3-(2-(dimethoxyphosphoryl)vinyl)pentadecanoate (14e)
To a stirring solution of carbonate 13e (1.26 g, 3.20 mmol) and methyl acetoacetate (1.05 mL, 9.60 mmol) in dry THF (6.5 mL) was added Pd2(dba)3 (0.06 g, 0.64 mmol) followed by dppe (0.06 g, 0.16 mmol) to give 14e as a 1:1 mixture of diastereoisomers (1.13 g, 80%). Pale yellow oil; IR (neat, NaCl) 1743, 1719 cm−1; 1H NMR (CDCl3) δ 6.59 (0.5H, ddd, JHH = 17.1, 9.5 Hz, JHP = 21.5 Hz), 6.49 (0.5H, ddd, JHH = 17.1, 9.5 Hz, JHP = 21.5 Hz), 5.72 (0.5H, dd, JHH = 17.1Hz, JHP = 21.5 Hz), 5.65 (0.5H, dd, JHH = 17.1Hz, JHP = 21.5 Hz), 3.71 (9H, m), 3.5 (1H, 2d, JHH = 9.4 and 9.2 Hz), 2.98 (1H, 2q, JHH = 9.1 and 9.3 Hz, JHH = 1.6 Hz), 2.21 (1.5H, s), 2.15 (1.5H, s), 1.22 (22H, br s), 0.88 (3H, t, JHH = 6.7 Hz); 13C NMR (CDCl3) δ 201.5, 168.7 (168.6), 153.5 (d, JCP = 4.4 Hz), 118.9 (185) (d, JCP = 184.8 Hz), 63.9 (63.8), 52.6 (52.7), 52.5 (d, JCP = 5.6 Hz), 44.3 (44), 32.1, 31.9 (31.8), 30.3 (30.1), 29.9, 29.8, 29.7, 29.6, 29.5, 27.3 (27.2), 22.9, 14.3; 31P NMR (CDCl3) δ 20.7, 20.6; HRMS (FAB, MH+) calcd. for C22H42O6P: 433.2718, found 433.2720.
(±) (E)-methyl 2-acetyl-3-(2-(dimethoxyphosphoryl)vinyl)nonadecanoate (14f)
To a stirring solution of carbonate 13f (0.45 g, 1.0 mmol) and methyl acetoacetate (0.64 mL, 3.0 mmol) in dry THF (2mL) was added Pd2(dba)3 (0.02g, 0.02 mmol) followed by dppe (0.02g, 0.05 mmol) to give 14f as a 1:1 mixture of diastereoisomers (0.36 g, 72%). Waxy solid; IR (neat, NaCl), 1743, 1637 cm−1; 1H NMR (CDCl3) δ 6.59 (0.5H, ddd, JHH = 17.1, 9.5 Hz, JHP = 21.5 Hz), 6.45 (0.5H, ddd, JHH = 17.1, 9.4 Hz, JHP = 21.5 Hz), 5.72 (0.5H, dd, JHH = 17.1 Hz, JHP = 21.5 Hz), 5.65 (0.5H, dd, JHH = 17.1 Hz, JHP = 21.5 Hz), 3.71 (9H, m), 3.54 (1H, 2d, JHH = 9.4 and 9.2 Hz), 2.97 (1H, 2q, JHH = 9.1 and 9.3 Hz), 2.21 (1.5H, s), 2.15 (1.5H, s), 1.22 (30H, br s), 0.85 (3H, t, JHH = 6.66 Hz); 13C NMR (CDCl3) δ 201.3, 168.6 (168.5), 153.4 (153.3) (d, JCP = 8.4 Hz), 118.8 (118.6) (d, JCP = 184.3 Hz), 63.8 (63.7), 52.7 (52.5), 52.7 (d, JCP = 5.6 Hz), 44.2 (43.9) (d, JCP = 2.2 Hz), 32.1, 31.8 (31.7), 30.2 (30), 29.9, 29.8, 29.7, 29.6, 29.5, 29.4, 27.2 (27.1), 22.8, 14.3; 31P NMR (CDCl3) δ 20.7, 20.6; HRMS (FAB, NBA, MH+) calcd. for C26H50O6P: 485.3345, found 489.3340.
(±) (E)-methyl 2-acetyl-3-(2-(dimethoxyphosphoryl)vinyl)henicosanoate (14g)
To a stirring solution of carbonate 13g (0.48 g, 1.0 mmol) and methyl acetoacetate (0.33 mL, 3.0 mmol) in dry THF (2 mL) was added Pd2(dba)3 (0.02 g, 0.02 mmol) followed by dppe (0.02 g, 0.05 mmol) to give 14g as a 1:1 mixture of diastereoisomers (0.35 g, 70%). Pale yellow oil; IR (neat, NaCl) 1742, 1713 cm−1; 1H NMR (CDCl3) δ 6.56 (0.5H, ddd, JHH = 17.1, 9.5 Hz, JHP = 21.4 Hz), 6.45 (0.5H, ddd, JHH = 17.1, 9.5 Hz, JHP = 21.5 Hz), 5.67 (0.5H, dd, JHH = 17.1 Hz, JHP = 21.5 Hz), 5.60 (0.5H, dd, JHH = 17.1 Hz, JHP = 21.5 Hz), 3.67 (9H, m), 3.54 (1H, 2d, JHH = 9.4 and 9.2 Hz), 2.97 (1H, 2q, JHH = 9.4 and 9.2 Hz), 2.21 (1.5H, s), 2.15 (1.5H, s), 1.21 (34H, br s), 0.84 (3H, t, JHH = 6.6 Hz); 13C NMR (CDCl3) δ 201.3, 168.6 (168.4), 153.4 (153.3) (d, JCP = 4.6 Hz), 118.8 (118.6) (d, JCP = 184.6 Hz), 63.8 (63.7), 52.7 (52.5), 52.4 (d, JCP = 5.5 Hz), 44.2 (43.9) (d, JCP = 2.3 Hz), 32.0, 31.8 (31.7), 30.2 (30), 29.9, 29.8, 29.7, 29.6, 29.5, 29.4, 27.2 (27.1), 22.8, 14.3; 31P NMR (CDCl3) δ 20.7, 20.6; HRMS (FAB, NBA, MH+) calcd. for C26H50O6P: 485.3345, found 489.3340.
General procedure for hydrogenation of vinyl phosphonate
To a solution of vinyl phosphonate 14 (1 mmol) in methanol (3 mL) was added 10% palladium on charcoal (0.07 g). The solution was first flushed with argon and then with hydrogen gas. A reservoir of hydrogen was provided by a balloon. After completion of the reaction (31P NMR analysis), the reaction mixture was filtered through Celite which was washed with additional CH2Cl2. The solvent was evaporated under reduced pressure to give the saturated phosphonate 16.
(±) Methyl 2-acetyl-5-(dimethoxyphosphoryl)-3-phenylpentanoate (16b)
To a solution of 14b (or, 15) (0.29 g, 0.84 mmol) in methanol (3 mL) was added palladium on charcoal (0.09 g) and hydrogen gas to give the saturated phosphonate 14b as a 1:1 mixture of diastereoisomers (0.30 g, 100%). Colorless oil; IR (neat, NaCl) 1745, 1717 cm−1; 1H NMR (CDCl3) δ 7.34 (m, 5H), 4.28(1H, m), 4.1(1H, d, JHH = 11.27 Hz), 3.66 (6H, m), 3.61 (3H, s), 2.14 (3H, s); 13C NMR (CDCl3) δ 201.8, 168.8 (168.2), 139.5 (139.2), 129.1, 128.8, 128.4, 127.7, 127.6, 66.6 (65.8), 52.8, 52.5 (d, JCP = 6.5 Hz), 46.1 (45.6) (d, JCP = 19.1 Hz), 30.3 (30.0), 26.9 (26.8) (d, JCP = 16.4 Hz), 23.5 (23.4), 21.6 (d, JCP = 8.7 Hz); 31P NMR (CDCl3) δ 34.6 ppm; HRMS (FAB, MH+) calcd. for C16H24O6P: 343.1310, found 343.1311.
(±) Methyl 2-acetyl-3-(2-(dimethoxyphosphoryl)ethyl)octanoate (16c)
To a solution of 14c (0.650 g, 1.96 mmol) in methanol (5 mL) was added palladium on charcoal (0.21 g) and hydrogen gas to give the saturated phosphonate 16c as a 1:1 mixture of diastereoisomers (0.65 g, 100%). Colorless oil; IR (neat, NaCl) 1739, 1716 cm−1; 1H NMR (CDCl3) δ 3.73 (9H, m), 3.45 (1H, d, JHH = 8.3 Hz), 2.26 (1H, br s), 2.23 (4H, s), 1.67 (m, 4H), 1.25 (br, 8H), 0.87 (t, 3H, JHH = 6.7 Hz); 13C NMR (CDCl3) δ 202.9, 169.7 (169.69), 63.2(63.1), 52.6, 52.5 (d, JCP = 6.5 Hz), 38.1 (d, JCP = 17.2a Hz), 32.0 (d, JCP = 7.1 Hz), 30.6 (30.4), 29.6, 26.1, 22.7, 14.2; 31P NMR (CDCl3) δ 35.0, 34.9; HRMS (FAB, NBA, MH+) calcd. for C15H29O6P: 336.1780, found 337.1779.
(±) Methyl 2-acetyl-3-(2-(dimethoxyphosphoryl)ethyl)tridecanoate (16d)
To the solution of 14d (0.44 g, 1.08 mmol) in methanol (5 mL) was added palladium on charcoal (0.01 g) and hydrogen gas to give the saturated phosphonate 16d as a 1:1 mixture of diastereoisomers (0.44 g, 100%). Colorless oil IR (neat, NaCl) 1737, 1712 cm−1; 1H NMR (CDCl3) δ 3.73 (6H, d, JHD = 10.8 Hz), 3.73 (3 H, app d), 3.45(1H, d, JHH = 8.3 Hz), 2.26 (1H, br s), 2.23 (3H, s), 1.76 – 1.6 (4H, m), 1.25 (18 H, br s), 0.88 (3H, t, JHH = 6.44 Hz); 13C NMR (CDCl3) δ 202.9, 169.7, 63.2, 52.6, 52.5, 38.2 (38), 32.1, 30.5 (d, JCP = 10.5 Hz), 30, 29.9, 29.8, 29.7, 29.6, 29.5, 26.5, 22.9, 14.3; 31P NMR (CDCl3) δ 35.0, 34.9; HRMS (FAB+) calcd for C20H39O6PNa: 429.2381, found 429.2351.
(±) Methyl 2-acetyl-3-(2-(dimethoxyphosphoryl)ethyl)pentadecanoate (16e)
To a solution of 14e (0.33 g, 0.757 mmol) in methanol (4 mL) was added palladium on charcoal (0.08 g) and hydrogen gas to give the saturated phosphonate 16e as a 1:1 mixture of diastereoisomers (0.33 g, 100%). Colorless oil; IR (neat, NaCl) 1736, 1712 cm−1; 1H NMR (CDCl3) δ 3.73 (6H, JHP = 10.8 Hz), 3.73 (3H, app d), 3.46 (1H, d, JHH = 8.3,Hz), 2.27 (1H, br s), 2.23 (3H, s), 1.77 – 1.6 (4H, m), 1.25 (20 H, br s), 0.88 (3H, t, JHH = 6.7 Hz); 13C NMR (CDCl3) δ 203, 169.7, 63.2, 52.6 (d, JCP = 7.1 Hz), 52.5, 38.2 (38), 32.1, 30.6 (d, JCP = 10.6 Hz), 30, 29.9, 29.8, 29.7, 29.6, 29.5, 26.5, 22.9 14.3; 31P NMR (CDCl3) δ 35.0, 34.9; HRMS (FAB, NBA, MH+) calcd for C22H44O6P: 435.2875, found 435.2868.
(±) Methyl 2-acetyl-3-(2-(dimethoxyphosphoryl)ethyl)nonadecanoate (16f)
To a solution of 14f (0.22 g, 0.45 mmol) in methanol (3 mL) was added palladium on charcoal (0.05 g) and hydrogen gas to give the saturated phosphonate 16f as a 1:1 mixture of diastereoisomers (0.22 g, 100%). Colorless oil; IR (neat, NaCl) 1736, 1718 cm−1; 1H NMR (CDCl3) 3.72 (6H, d, JHP = 10.7 Hz), 3.71 (3H, app d), 3.44(1H, d, JHH = 8.9Hz), 2.24 (1H, br s), 2.21 (3H, s), 1.73 - 1.58 (4H, m), 1.23 (30H, br s), 0.86 (3H, t, JHH = 6.7 Hz); 13C NMR (CDCl3) δ 203, 169.7, 63.1, 52.6 (d, JCP = 7.1 Hz), 52.5, 38.3, 38, 32.1, 30.5 (d, JCP = 10.2 Hz), 29.9, 29.7, 29.6, 29.5, 26.5, 23.5, 22.9, 20.7, 14.3; 31P NMR (CDCl3) δ 35, 34.9; HRMS (FAB, NBA, MH+) calcd for C26H52O6P: 491.3501, found 491.3507.
(±) Methyl 2-acetyl-3-(2-(dimethoxyphosphoryl)ethyl)henicosanoate (16g)
To a solution of 14g (0.28 g, 0.55 mmol) in methanol (5 mL) was added palladium on charcoal (0.06 g, 0.06 mmol) and hydrogen gas to give the saturated phosphonate 16g as a 1:1 mixture of diastereoisomers (0.29 g, 100%). Colorless oil; IR (neat, NaCl) 1735, 1712 cm−1; 1H NMR (CDCl3) 3.73 (6H, d, JHP = 10.8 Hz), 3.73 (3H, app d), 3.46 (1H, d, JHH = 8.3 Hz), 2.26 (1H, br s), 2.23 (3H, s), 1.78 - 1.55 (4H, m), 1.25 (34H, br s), 0.88 (3H, t, JHH = 6.7 Hz); 13C NMR (CDCl3) δ 203, 169.7, 63.2, 52.6 (d, JCP = 6.2 Hz), 52.5, 38.2 (38), 32.1, 30.5, 29.9, 29.8, 29.7, 29.6, 26.5, 22.9, 14.3; 31P NMR (CDCl3) δ 35.0, 34.9; HRMS (FAB, NBA, MH+) calcd for C26H57O6PNa: 519.3814, found 519.3818.
General procedure for the synthesis of monocylic phosphonate analogs (5(α, β)-10(α, β))
To a solution of alkyl phosphonate 16 (1 mmol) in acetonitrile (2.6 mL) was added NaI (1.1 equiv) and the resulting mixture was heated at reflux. After completion of the reaction (31P NMR analysis), the solvent was removed under reduced pressure to give mono-sodium salt as solid. The sodium salt was suspended in methanol, Amberlite IR-120H resin (prewashed with methanol) was added and resulting mixture was shaken in an orbit shaker. After completion of the reaction (31P NMR analysis), the resin was removed by filtration and washed with additional methanol. The solvent was removed under reduced pressure to give free monophosphonic acid as a red viscous liquid. The monophosphonic acid (1 mmol) was dissolved in freshly distilled CH2Cl2 (6 mL) EDC (1.5 mmol) and HOBt (1.5 mmol) were added, followed by Hunig’s base (i-Pr2NEt) (1.5 mmol). After the completion of the reaction (31P NMR analysis), the reaction mixture was diluted with additional CH2Cl2 and the resulting solution was washed with saturated NaHCO3. The organic layer was separated and dried over anhydrous Na2SO4, then evaporated under reduced pressure to give crude product. Purification of the crude product by chromatography (SiO2, 10–60% EtOAc in hexanes) gave the product as mixture of diastereomers which were separated by further careful column chromatography.
(±) Methyl 2-methoxy-7-methyl-5-phenyl-2,3,4,5-tetrahydro-1,2-oxaphosphepine-6-carboxylate 2-oxide (5(α) & 5(β))
A solution of phosphonate 16b (0.5 g, 1.46 mmol) in acetonitrile (5 mL) was treated as described above to give the monocyclic phosphonate as a 1:1 mixture of diastereoisomers (0.28 g, 52%). Further chromatographic separation of the diastereoisomers gave 5(α) (95:5 ratio with 5(β)). IR (neat, NaCl) 1719 cm−1; 1H NMR (CDCl3) δ 7.27(m, 5H), 4.28(1H, t, JHH = 7 Hz), 3.86 (3H, d, JHP = 11.15 Hz). 3.51 (3H, s), 2.25 (3H, s), 2.19 (1H, m), 2.05 (1H, m), 1.98 (1H, t, JHH = 6 Hz); 13C NMR (CDCl3) δ 168.97, (168.78), 155.65 (d, JCP = 7.28Hz), (154.66); 141.35, (141.12); (129.04), 129.00; (128.10), 127.91; (127.25), 127.15; 121.44 (d, JCP = 5.1 Hz), 53.07 (d, JCP = 6.9Hz), 52.71 (d, JCP = 6.9 Hz), 52.33, 45.78 (45.63), 28.53 (d, JCP = 7.04 Hz), 23.16 (d, JCP = 133.1 Hz), 23.04 (d, JCP = 133.1 Hz), 22 (21.76); 31P NMR (CDCl3) δ 30.1; HRMS (EI+) calcd. for C15H19O5P: 310.0970, found 310.0975.
(±) Methyl 2-methoxy-7-methyl-5-pentyl-2,3,4,5-tetrahydro-1,2-oxaphosphepine-6-carboxylate 2-oxide (6(α) & 6(β))
To a solution of 16c (0.52 g, 1.55 mmol) in acetonitrile (5 mL) was treated as described above to give the monocyclic phosphonates 6 as a 1:1.25 mixture of diastereoisomers (0.27 g, 56%). Further chromatographic separation of the diastereoisomers gave 6(α) as pale yellow oil. IR (neat, NaCl) 1717 cm−1; 1H NMR (CDCl3) δ 3.82 (d, 3H, JHP = 11.18 Hz), 3.73 (3H, s), 2.97 (1H, m), 2.22 (3H, d, JHH = 1.28 Hz), 2.12–1.81 (4H, m), 1.61 (1H, m), 1.46 (1H, m), 1.25 (6H, br s), 0.87 (3H, t, JHH = 6.5 Hz); 13C NMR (CDCl3) δ 169.34, 156.1 (d, JCP = 7.28 Hz), 123.55 (d, JCP = 5.1 Hz), 52.39 (d, JCP = 7.1 Hz), 52.26, 37.63, 32.21, 31.1, 27.7, 25.31 (d, JCP = 6.8 Hz), 22.93, 22.3 (d, JCP = 134.3 Hz), 21.67, 14.44; 31P NMR (CDCl3) δ 26.8; HRMS (FAB, NBA, MH+) calcd. for C14H26O5P: 305.1517, found 305.1523; and 6(β) as pale yellow oil. IR (neat, NaCl) 1717 cm−1; 1H NMR (CDCl3) δ 3.82 (3H, d, JHP = 11.18 Hz), 3.75 (3H, s), 2.88 (1H, m), 2.18 (3H, d, JHH = 1.3 Hz), 2.14-1.87 (4H, m), 1.63 (1H, m), 1.47 (1H, m), 1.25 (6H, br s), 0.87 (3H, t, JHH = 6.5 Hz); 13C NMR (CDCl3) δ 169.51, 155.27 (d, JCP = 9.3 Hz), 123.44 (d, JCP = 4.73 Hz), 52. 95 (d, JCP = 6.67 Hz), 52.32, 37.5, 32.13, 31.2, 30.1, 27.65, 25.51 (d, JCP = 7.91 Hz), 22.91, 22.64 (d, JCP = 133.33Hz), 21.51 (d, JCP = 1.87 Hz), 14.43; 31P NMR (CDCl3) δ 24.2; HRMS (FAB, NBA, MH+) calcd. for C14H26O5P: 305.1517, found 305.1523.
(±) Methyl 5-decyl-2-methoxy-7-methyl-2,3,4,5-tetrahydro-1,2-oxaphosphepine-6-carboxylate 2-oxide (7(α) & 7(β))
To a solution of alkyl phosphonate 16d (0.47 g, 1.16 mmol) in acetonitrile (3 mL) was treated as described above to give the monocyclic phosphonates 7 as a 1:1.25 mixtures of diastereoisomers (0.21 g, 46%). Further chromatographic separation of the diastereoisomers gave 7(α) as pale yellow oil. IR (neat, NaCl) 1719 cm−1; 1H NMR (CDCl3) δ 3.82 (3H, d, JHP = 11.2 Hz), 3.75 (3H, s), 2.99 (1H, m), 2.23 (3H, d, JHH = 1.6 Hz), 2.14 –1.87 (4H, m), 1.62 (1H, m), 1.48 (1H, m), 1.25 (16H, br s), 0.88 (3H, t, JHH = 6.7 Hz); 13C NMR (CDCl3) δ 169.2 (d, JCP = 1.9 Hz), 155.9 (d, JCP = 7.3 Hz), 123.4 (d, JCP = 5.3 Hz), 52.2 (d, JCP = 7.1 Hz), 52.1, 37.4, 32.1, 30.9, 29.9, 29.8, 29.7, 29.5, 27.9, 25.1 (d, JCP = 6.9 Hz), 22.9, 22.1 (d, JCP = 134.4Hz), 21.51, 14.3; 31P NMR (CDCl3) δ 26.8; HRMS (FAB, NBA, MNa+) calcd. for C19H35O5PNa: 397.2119, found 397.2125; and 7(β) also as pale yellow oil. IR (neat, NaCl) 1719 cm−1; 1H NMR (CDCl3) δ 3.83 (3H, d, JHP = 11.1 Hz), 3.76 (3H, s), 2.88 (1H, m), 2.18 (3H, d, JHH = 1.3 Hz), 2.26–1.87 (4H, m), 1.63 (1H, m), 1.45 (1H, m), 1.25 (16H, br s), 0.88 (3H, t, JHH = 6.7 Hz); 13C NMR (CDCl3) δ 169.3 (d, JCP = 1.7 Hz), 155.1 (d, JCP = 9.4 Hz), 123.3 (d, JCP = 4.6 Hz), 52.8 (d, JCP = 6.7 Hz), 52.1, 37.3, 32.1, 31.1, 29.9, 29.8, 29.7, 29.5, 27.8, 25.3 (d, JCP = 7.8 Hz), 22.9, 22.5 (d, JCP = 133.4 Hz), 21.3 (d, JCP = 1.9 Hz), 14.3; 31P NMR (CDCl3) δ 24.2; HRMS (FAB, NBA, MH+) calcd. for C19H35O5PNa: 397.2119, found 397.2115.
(±) Methyl 5-dodecyl-2-methoxy-7-methyl-2,3,4,5-tetrahydro-1,2-oxaphosphepine-6-carboxylate 2-oxide (8(α) & 8(β))
To a solution of alkyl phosphonate 16e (0.32 g, 0.74 mmol) in acetonitrile (2 mL) was treated as described above to give the monocyclic phosphonates 8 as a 1:1.25 mixtures of diastereomers (0.14 g, 45%) which were further separated by column chromatography to give 8(α) as pale yellow oil. IR (neat, NaCl) 1719 cm−1; 1H NMR (CDCl3) δ 3.83 (3H, d, JHP = 11.2 Hz), 3.75 (3H, s), 2.99 (1H, m), 2.23 (3H, d, JHH = 1.6 Hz), 2.14 – 1.87 (4H, m), 1.63 (1H, m), 1.47 (1H, m), 1.25 (20H, br s), 0.89 (3H, t, JHH = 6.7 Hz); 13C NMR (CDCl3) δ 169.2, 155.9 (d, JCP = 7.1 Hz), 123.4 (d, JCP = 5.3 Hz), 52.2 (d, JCP = 7.8 Hz), 52.1, 37.4, 32.1, 30.9, 29.9, 29.8, 29.7, 29.6, 27.9, 25.1 (d, JCP = 6.8 Hz), 22.9, 22.6 (d, JCP = 134.3Hz), 21.5, 14.3; 31P NMR (CDCl3) δ 26.8; HRMS (FAB, NBA, MH+) calcd. for C21H40O5P:403.2613, found 403.2613; and 8(β) as pale yellow oil. IR (neat, NaCl) 1719 cm−1; 1H NMR (CDCl3) δ 3.83 (3H, d, JHP = 11.2 Hz), 3.76 (3H, s), 2.88 (1H, m), 2.19 (3H, d, JHH = 1.2 Hz), 2.14 –1.87 (4H, m), 1.63 (1H, m), 1.47 (1H, m), 1.25 (6H, br s), 0.87 (3H, t, JHH = 6.5 Hz); 13C NMR (CDCl3) δ 169.4, 155.1 (d, JCP = 9.4 Hz), 123.3 (d, JCP = 4.3 Hz), 52.8 (d, JCP = 6.7 Hz), 52.2, 37.3, 32.1, 31.0, 29.9, 29.8, 29.7, 29.6, 27.8, 25.3 (d, JCP = 7.7 Hz), 22.9, 22.4 (d, JCP = 133.5Hz), 21.3 (d, JCP = 2.0 Hz), 14.3; 31P NMR (CDCl3) δ 24.2; HRMS (FAB, NBA, MNa+) calcd. for C21H39O5PNa: 425.2439, found 425.2427.
(±) Methyl 5-hexadecyl-2-methoxy-7-methyl-2,3,4,5-tetrahydro-1,2-oxaphosphepine - 6-carboxylate 2-oxide (9(α) & 9(β))
To a solution of alkyl phosphonate 16f (1.05 g, 2.14 mmol) in acetonitrile (5 mL) was treated as described above to give the monocyclic phosphonates 9 as a 1:1.25 mixtures of diastereomers (0.45 g, 46%) which were further separated by column chromatography to give 9(α) as white solid. IR (neat, NaCl) 1720 cm−1; 1H NMR (CDCl3) δ 3.82 (3H, d, JHP = 11.2 Hz), 3.74 (3H, s), 2.98 (1H, m), 2.23 (3H, d, JHH = 1.5 Hz), 2.14 – 1.88 (4H, m), 1.62 (1H, m), 1.45 (1H, m), 1.25 (30 H, br s), 0.88 (3H, t, JHH = 6.7 Hz); 13C NMR (CDCl3) δ 169.2 (d, JCP = 1.8 Hz), 155.9 (d, JCP = 7.3 Hz), 123.4 (d, JCP = 5.3 Hz), 52.2 (d, JCP = 7.1 Hz), 52.1, 37.4 (d, JCP = 1.5 Hz), 32.1, 30.9, 30.0, 29.9, 29.8, 29.7, 29.6, 27.9, 25.1 (d, JCP = 6.8 Hz), 22.9, 22.1 (d, JCP = 134.4 Hz), 21.5 (d, JCP = 1.3 Hz), 14.3; 31P NMR (CDCl3) δ 26.8; HRMS (FAB, NBA, MH+) calcd. for C25H48O5P: 459.3239, found 459.3239; and 9(β) as white solid. IR (neat, NaCl) 1720 cm−1; 1H NMR (CDCl3) δ 3.82 (3H, d, JHP = 11.1 Hz), 3.76 (3H, s), 2.88 (1H, m), 2.18 (3H, s), 2.11(1H, m), 1.99 (2H, m), 1.89 (1H, m), 1.63 (1H, m), 1.49 (1H, m), 1.25 (30H, br s), 0.89 (3H, t, JHH = 6.6 Hz); 13C NMR (CDCl3) δ 169.3 (d, JCP = 1.7 Hz), 155.1 (d, JCP = 9.5 Hz), 123.3 (d, JCP = 4.6 Hz), 52.8 (d, JCP = 6.7 Hz), 52.2, 37.3, 32.1, 31.0, 30.0, 29.9, 29.8, 29.7, 29.6, 29.3, 27.8, 25.3 (d, JCP = 7.9 Hz), 22.9, 22.4 (d, JCP = 133.4 Hz), 21.3 (d, JCP = 1.9 Hz), 14.3; 31P NMR (CDCl3) δ 24.1; HRMS (FAB, NBA, MH+) calcd. for C25H48O5P: 459.3239, found 459.3244.
(±) Methyl 2-methoxy-7-methyl-5-octadecyl-2,3,4,5-tetrahydro-1,2-oxaphosphepine-6-carboxylate 2-oxide (10(α) & 10(β))
To a solution of alkyl phosphonate 9g (0.29 g, 0.56 mmol) in acetonitrile (2 mL) was treated as described above to give the monocyclic phosphonates 10 as a 1:1.25 mixtures of diastereomers (0.16 g, 59%) which were further separated by column chromatography to give 10(α) as white solid. IR (neat, NaCl) 1719 cm−1; 1H NMR (CDCl3) δ 3.83 (3H, d, JHP = 11.18 Hz), 3.73 (3H, s), 2.99 (1H, m), 2.24 (3H, d, JHH = 1.6 Hz), 2.14 – 1.88 (4H, m), 1.63 (1H, m), 1.47 (1H, m), 1.26 (32 H, br s), 0.89 (3H, t, JHH = 6.7 Hz); 13C NMR (CDCl3) δ 169.2, 155.9 (d, JCP = 7.5 Hz), 123.4 (d, JCP = 5.3 Hz), 52.2 (d, JCP = 7.1 Hz), 52.1, 37.4, 32.1, 30.9, 30.0, 29.9, 29.8, 29.7, 27.9, 25.5 (d, JCP = 6.9 Hz), 22.9, 22.1 (d, JCP = 134.5 Hz), 21.5, 14.3; 31P NMR (CDCl3) δ 26.8; HRMS (FAB, NBA, MH+) calcd. for C27H51O5PNa: 509.3371, found 509.3375; and 10(β) as white solid. IR (neat, NaCl) 1718 cm−1; 1H NMR (CDCl3) δ 3.83 (3H, d, JHP = 11.1 Hz), 3.76 (3H, s), 2.88 (1H, m), 2.19 (3H, d, JHH = 1.2 Hz), 2.11(1H, m), 2.01 (2H, m), 1.89 (1H, m)1.64 (2H, m), 1.50 (2H, m), 1.26 (26H, br s), 0.88 (3H, t, JHH = 6.6 Hz); 13C NMR (CDCl3) δ 169.3, 155.1 (d, JCP = 9.3 Hz), 123.3 (d, JCP = 4.7 Hz), 52.8 (d, JCP = 6.7 Hz), 52.2, 37.3, 32.1, 31.0, 30, 29.9, 29.7, 29.6, 27.8, 25.3 (d, JCP = 8 Hz), 22.9, 22.4 (d, JCP = 133.3 Hz), 21.3 (d, JCP = 2.1 Hz), 14.3; 31P NMR (CDCl3) δ 24.2; HRMS (FAB, MH+) calcd. for C27H51O5PNa: 509.3371, found 509.3363.
Methyl 7-methyl-2-(hexadecyloxy)-2,3,4,5-tetrahydro-1,2-oxaphosphepine-6-carboxylate 2-oxide (11)
To a solution of monocyclic phosphonate analog 4 (0.024 g, 0.10 mmol) and 1-bromohexadecane (0.15 g, 0.50 mmol) in dry toluene (0.5 mL) was added nBu4NI (TBAI) (0.003 mmol, 0.002 g, 5 mol %) at room temperature. The mixture was heated to reflux. After completion of reaction (TLC and 31P NMR analysis), the solvent was removed under reduced pressure and the crude product was purified by column chromatography (SiO2, 10–30% EtOAc in hexane) to give phosphonate analog 11 (0.037g, 86 %) as a waxy solid. IR (neat, NaCl), 1717 cm−1; 1H NMR (CDCl3) δ 4.12 (2H, m), 3.73 (3H, s), 2.65 (1H, m), 2.47 (1H, m), 2.31 (3H, s), 2.18 – 1.89 (4H, m), 1.68 (2H, p, JHH = 6.9 Hz), 1.37 – 1.24 (26H, br s), 0.86 (3H, t, JHH = 6.6 Hz); 13C NMR (CDCl3) δ 168.22 (d, JCP = 1.7 Hz), 159.43 (d, JCP = 7.8 Hz), 119.20 (d, JCP = 4.6 Hz), 66.24 (d, JCP = 7.0 Hz), 52.00, 32.08, 30.57 (d, JCP = 6.0 Hz), 29.85, 29.81, 29.80, 29.71, 29.66, 29.52, 27.68, 26.42, 26.78 (d, JCP = 133.6 Hz), 26.41 (d, JCP = 2.7 Hz), 25.61, 22.85, 21.29 21.67 (d, JCP = 7.5 Hz), 21.12 (d, JCP = 1.6 Hz), 14.44; 31P NMR (CDCl3) δ 23.1; HRMS (FAB, NBA, MH+) calcd. For C24H46O5P: 445.3083, found 445.3076.
Methyl 7-methyl-2-(octadecyloxy)-2,3,4,5-tetrahydro-1,2-oxaphosphepine-6-carboxylate 2-oxide (12)
The monocyclic phosphonate 4 (0.064 g, 0.26 mmol) was dissolved in dry acetone (0.5 mL) and NaI (0.044 g, 0.29 mmol) was added. The yellow solution was heated at reflux overnight. After completion of the reaction (31P NMR analysis) the solvent was removed under reduced pressure. The dark orange residue was dissolved in dry methanol (10 mL) and pre-rinsed Amberlite IR-120H resin (0.320 g) was added. The mixture was shaken on an orbital shaker for 45 min, filtered and concentrated under reduced pressure to yield a light orange oil. The oil was dissolved in dry CH2Cl2 (0.20 mL) and then potassium carbonate (0.040 g, 0.29 mmol) and 18-crown-6 (0.002 g, 0.08 mmol) were added, followed by the addition of n-octadecyl triflate (0.149 g, 0.369 mmol)40 in dry CH2Cl2 (0.30 mL) dropwise. After 48 h, the reaction was quenched with deionized water and extracted three times with CH2Cl2. The combined organic extracts was dried over Na2SO4 and concentrated under reduced pressure. The crude product was purified by column chromatography (SiO2, 7–15% EtOAc in hexanes) to give 12 (0.061 g, 49%) as a white waxy solid. IR (neat, ATR), 1712, 1646 cm−1; 1H NMR (CDCl3) δ 4.13 (2H, m), 3.72 (3H, s), 2.65 (1H, m), 2.47 (1H, m), 2.30 (3H, s), 2.20 – 1.83 (4H, m), 1.67 (2H, p, JHH = 6.8 Hz), 1.30 – 1.16 (28H, br s), 0.85 (3H, t, JHH = 6.6 Hz); 13C NMR (CDCl3) δ 168.19, 159.414 (d, JCP = 7.8 Hz), 119.18 (d, JCP = 4.6 Hz), 66.22 (d, JCP = 7.2 Hz), 51.98, 32.07, 30.56 (d, JCP = 6.2), 29.84, 29.81, 29.78, 29.70, 29.65, 29.51, 29.28, 26.78 (d, JCP = 134.3 Hz), 26.39 (d, JCP = 2.7 Hz), 25.61, 22.84, 21.33, 21.12, 14.27; 31P NMR (CDCl3) δ 23.7; HRMS (FAB, NBA, MH+) calcd. For C26H49O5P: 473.3396, found 473.3385.
X-ray Structure Determination of compound 10(α)
Crystals of appropriate dimension were obtained by slow evaporation of acetone. A crystal with approximate dimensions 0.01×0.03×0.16 mm3 was mounted on a Mitgen cryoloop in a random orientation. Intensity data were collected at 150K on a D8 goniostat at 70 mm crystal to detector distance equipped with a Bruker APEXII CCD detector at Beamline 11.3.1 at the Advanced Light Source (Lawrence Berkeley National Laboratory) using synchrotron radiation tuned to λ = 1.2399 Å. For data collection frames were measured for a duration of 1-s for low angle data and 3-s for high angle data at 0.3° intervals of ω with a maximum 2θ value of ~ 60°. The data frames were collected using the program APEX2 and processed using the program SAINT routine within APEX2 (Bruker Analytical X-Ray, Madison, WI, 2010). The data were corrected for absorption and beam corrections based on the multi-scan technique as implemented in SADABS (Bruker Analytical X-Ray, Madison, WI, 2010). Crystal data and intensity data collection parameters are listed below. Structure solution and refinement were carried out using the SHELXTL-PLUS software package.48 The structure was solved by direct methods and refined successfully in the space group, P21/c. Full matrix least-squares refinement was carried out by minimizing Σw(Fo2-Fc2)2. The non-hydrogen atoms were refined anisotropically to convergence. All hydrogen atoms were treated using appropriate riding model (AFIX m3). Complete listings of positional and isotropic displacement coefficients for hydrogen atoms, anisotropic displacement coefficients for the non-hydrogen atoms are listed as supplementary material (Tables S1 to S5). The obtained crystal structure has been deposited at the Cambridge Crystallographic Data Centre and allocated the deposition number CCDC 873939.
Crystal data and structure refinement for compound 10(α).
| Empirical formula | C27H51O5P | |
| Formula weight | 486.65 | |
| Temperature | 150(2) K | |
| Wavelength | 1.23990 Å | |
| Crystal system | Monoclinic | |
| Space group | P21/c | |
| Unit cell dimensions | a = 10.4218(10) Å | a= 90°. |
| b = 5.6365(5) Å | b= 91.908(5)°. | |
| c = 48.570(5) Å | g = 90°. | |
| Volume | 2851.5(5) Å3 | |
| Z | 4 | |
| Density (calculated) | 1.134 Mg/m3 | |
| Absorption coefficient | 0.571 mm−1 | |
| F(000) | 1072 | |
| Crystal size | 0.16 × 0.03 × 0.01 mm3 | |
| Theta range for data collection | 2.93 to 39.46° | |
| Index ranges | −10≤h≤10, −5≤k≤5, −49≤l≤49 | |
| Reflections collected | 111131 | |
| Independent reflections | 3094 [R(int) = 0.0813] | |
| Completeness to theta = 39.46° | 96.1% | |
| Absorption correction | Semi-empirical from equivalents | |
| Max. and min. transmission | 0.9704 and 0.7626 | |
| Refinement method | Full-matrix least-squares on F2 | |
| Data / restraints / parameters | 3094 / 47 / 320 | |
| Goodness-of-fit on F2 | 1.105 | |
| Final R indices [I>2sigma(I)] | R1 = 0.1522, wR2 = 0.3741 | |
| R indices (all data) | R1 = 0.1740, wR2 = 0.3822 | |
| Largest diff. peak and hole | 0.398 and −0.399 e.Å−3 | |
In vitro biological evaluation
Reagents
Orlistat, glyceryl tributyrate (tributyrin, TC4), 1-oleoyl-rac-glycerol (1-monoolein, MC18), bovine serum albumin (BSA), sodium taurodeoxycholate (NaTDC), calcium chloride dihydrate (CaCl2, 2H2O), benzamidine, 4-morpholineethanesulfonic acid (MES) were purchased from Sigma-Aldrich-Fluka Chimie (St-Quentin-Fallavier, France). Sodium chloride (NaCl) and Tris-(hydroxymethyl)-aminomethane (Tris) were purchased from VWR International (Fontenay-sous-Bois, France) and from Euromedex (Mundolsheim, France), respectively. All organic solvents were purchased from Carlo Erba Reactifs-SDS (Val de Reuil, France) and were of HPLC grade. For protein digestion, trypsin of sequencing grade was purchased from Sigma-Aldrich-Fluka Chimie (St-Quentin-Fallavier, France).
Lipases
All lipolytic enzymes were produced and purified to homogeneity at our laboratory. Recombinant Human pancreatic lipase (HPL) was produced in the yeast Pichia pastoris and purified as described previously.49 Recombinant guinea pig pancreatic lipase-related protein 2 (GPLRP2) was expressed in Aspergillus orizae and purified as described in Hjorth et al.50 Recombinant dog gastric lipase (DGL) was produced in transgenic maize by Meristem Therapeutics (Clermont-Ferrand, France)51 and purified as described in Roussel et al.44 Fusarium solani pisi Cutinase was produced and purified according to Lauwereys et al.52 The lipolytic enzymes from Mycobacterium tuberculosis, Rv0183 (monoglyceride lipase) and LipY (triacylglycerol lipase) were produced and purified as previously reported.34,53 Porcine pancreatic colipase (i.e. colipase) was purified from lipid-free porcine pancreatic powder (Laboratoire Industriel de Biologie - Soisy-sous-Montmorency, France) according to Fernandez et al.54
Lipase activity measurements using the pH-stat technique
Enzymatic activity was assayed at 37 °C by measuring the amount of free fatty acid (FFA) released from a mechanically stirred acylglycerols emulsions using 0.1 N NaOH with a pH-stat (Metrohm 718 STAT Titrino, Switzerland) adjusted to a fixed end point value.41 Except for Rv0183 acting on MC18 emulsions; all lipase activities were determined using TC4 specific assay emulsions: 0.5 mL TC4 was mixed with 14.5 mL buffer solution. The activities of HPL were determined using the standard assay solution for pancreatic lipase:55 0.3 mM Tris-HCl (pH 8.0), 150 mM NaCl, 2 mM CaCl2 and 4 mM NaTDC, in the presence of colipase added at a molar excess.56 The assay solution for GPLRP2 was 1 mM Tris-HCl (pH 8.0), 150 mM NaCl, 5 mM CaCl2, 2 mM NaTDC.50 In the case of DGL, the assay solution was 150 mM NaCl, 2 mM NaTDC, 2 µM BSA at pH 5.5.57 With Cutinase, the assay solution was 2.5 mM Tris-HCl (pH 8.0), 150 mM NaCl and 0.25 mM NaTDC. With LipY, the assay solution was 2.5 mM Tris-HCl (pH 7.5), 300 mM NaCl and 3 mM NaTDC. With regards to Rv0183, assays were performed with 100 µL MC18 added to 15 mL of 2.5 mM Tris-HCl (pH 8.0), 150 mM NaCl and 3 mM NaTDC.34 All experiments were at least performed in duplicate. Activities were expressed as international units: 1 U = 1 µmole FFA released per minute. The specific activities of HPL, GPLRP2, DGL, Cutinase, LipY and Rv0183, expressed in U per mg of pure enzyme, were found to be 8000 ±144, 2270 ±33, 340 ±21, 2270 ±63, 129 ±2 and 287 ±7 U/mg, respectively.
Inhibition assays
The lipase-inhibitor pre-incubation method was used to test, in aqueous medium and in the absence of substrate, the possible direct reactions between lipases and potential inhibitors.41 Each enzyme was then pre-incubated for 30 min at 25 °C with each inhibitor (5 mg/mL stock solution in isopropanol) at various inhibitor molar excess (xI = 2, 4, 20, 40, 100). Experiments with HPL were performed in the presence of a five-fold molar excess of colipase vs. HPL. Lipase incubations with inhibitors were performed in presence of 4 mM NaTDC in the case of HPL and DGL;58 2 mM NaTDC with GPLRP2 and Cutinase; and in absence of NaTDC with Rv0183 and LipY. Regarding the inhibition by Orlistat, 2 mM NaTDC were used in the case of Cutinase, Rv0183 and GPLRP2. During the inhibition experiments, samples of the incubation medium were collected after 30 min. incubation period and injected into the pH-stat vessel in order to measure the residual lipase activity at various inhibitor molar excess. The variation in the residual lipase activity was then used to determine the molar excess of the inhibitor which reduced the enzyme activity to 50% of its initial value (xI50).42 In addition, each enzyme was pre-incubated at 25 °C with best inhibitor compounds, at a fixed molar excess xI = 4 for one hour. Then, the residual enzyme activity was measured at various times in order to determine the half-inactivation time (t1/2).59 The xI value used in these latter experiments was chosen so that the average residual activities of Cutinase, Rv0183 and LipY obtained after 30 min of incubation were in the 10–15% range. In each case, control experiments were performed in the absence of inhibitor and with the same volume of isopropanol. It is worth noting that isopropanol at a final volume of less than 5% has no effect on the enzyme activity.
The lipase stereopreference for the compounds tested was also studied on the basis of the discrimination between the two diastereoisomers trans-(α) & cis-(β) of a same compound. A Diastereoselectivity Index (DI)43 was defined by Eq. 1:
| (Eq. 1) |
Protein digestion prior to MALDI-TOF mass spectrometry analysis
Protein digestion was performed with sequencing grade trypsin, in line with the manufacturer’s recommendations. Briefly, samples were treated as follows: 5–50 pmol of native and inhibited protein were diluted in 15 µL of 100 mM ammonium bicarbonate (pH 8.0) buffer and successively treated with 10 mM dithiothreitol (DTT) for 45 min at 56 °C, and 55 mM iodoacetamide (IAA) for 30 min at room temperature in the dark. Finally samples were incubated with 0.33 mg/mL trypsin solution in 25 mM ammonium bicarbonate (pH 8) buffer and digestions were performed overnight at 37 °C. The generated peptides were then dried on a SpeedVac (10 min) and acidified with 1 µL of 12.5% TFA in water before being analyzed by MALDI-TOF.
Matrix-Assisted Laser Desorption Ionization Time-of-Flight (MALDI-TOF) mass spectrometry analysis
Cutinase, Rv0183 and LipY were pre-incubated for 30 min at 25 °C with compounds 4, 7(β) and 11 (5 mg/mL stock solution in isopropanol), respectively, at a xI value of 40 in order to abolish any residual lipolytic activity. In each case, blank experiments were performed in absence of inhibitor. A saturated solution of α-cyano-4-hydroxycinnamic acid in acidified water (0.1% trifluoroacetic acid, TFA) and acetonitrile (30:70, v/v) was used as a matrix. Prior to the direct mass analysis of the entire lipases, aliquots of 1 µL of native and inhibited lipase solutions (corresponding to 20–100 pmol of protein) were mixed with 1 µL of a saturated solution of sinapinic acid in acidified water (0.1% TFA) and acetonitrile (60:40, v/v) and spotted onto the target. Samples were left to air dry at room temperature. MALDI-TOF analyses were performed on a Bruker Microflex II mass spectrometer (Daltonik, Deutchland). Mass spectra were acquired in the positive ion mode, using the FlexAnalysis software program (Bruker, Daltonik, Deutchland). Protein identifications were carried out using the MASCOT version 2.2.0 search engine (Matrix Science, London, UK) and the NCBI protein database. Theoretical and experimental peptide masses were obtained using the BioTools software program (Bruker, Daltonik, Deutchland).
Molecular docking
In silico molecular docking of lipase inhibitors present in the active site of lipases was performed with the Autodock Vina program.60 The PyMOL Molecular Graphics System (version 1.3, Schrödinger, LLC) was used as working environment with an in-house version of the AutoDock/Vina PyMOL plugin.61 The X-ray crystallographic structures of HPL, DGL and GPLRP2 (PDB entry codes: 1LPB,45 1K8Q,44 and 1GPL,25 respectively) available at the Protein Data Bank were used as receptors. Docking runs were performed after replacing the catalytic serine by a glycine residue to enable the ligand (i.e., the inhibitor) to adopt a suitable position corresponding to the pre-bound intermediate before the nucleophilic attack in the lipase active site. The box size used for the various receptors was chosen to fit the whole active site cleft and to allow non constructive binding positions. The model structure file was generated for each inhibitor molecule, and its geometry was refined using the Avogadro open-source program (Version 1.0.1. http://avogadro.openmolecules.net/).
Supplementary Material
ACKNOWLEDGEMENTS
Sadia Diomande and Vincent Delorme were funded by PhD fellowships from EGIDE and the Ministère de l’Enseignement Supérieur et de la Recherche, respectively. This work was supported by the CNRS, the Agence Nationale de la Recherche Française (ANR-07-PCVI-0009 and ANR MIEN 2009–00904 FOAMY_TUB) and by the LISA Carnot Institute (Convention ANR n°07-CARN-009-01). The compounds synthesis was supported by grant number R01-GM076192 from National Institute of General Medical studies. Authors would like to thank Dr. R. Lebrun, S. Lignon and R. Puppo (at the Proteomics platform of the Institut de Microbiologie de la Méditerranée, Marseille, France) for their precious help and support with the mass spectrometry experiments. Crystallographic data were collected through the SCrALS (Service Crystallography at Advanced Light Source) program at the Small-Crystal Crystallography Beamline 11.3.1 at the Advanced Light Source (ALS), Lawrence Berkeley National Laboratory. The ALS is supported by the U.S. Department of Energy, Office of Energy Sciences Materials Sciences Division, under contract DE-AC02-05CH11231.
ABBREVIATIONS USED
- AChE
acetylcholinesterase
- DAG
diacylglycerol
- DGL
dog gastric lipase
- GPLRP2
guinea pig pancreatic lipase-related protein 2
- HPL
human pancreatic lipase
- HSL
hormone-sensitive lipase
- MIC
minimal inhibitory concentration
- MAG
monoacylglycerol
- TAG
triacylglycerol
- xI
inhibitor molar excess related to 1 mol of enzyme
- xI50
inhibitor molar excess leading to 50% lipase inhibition
Footnotes
SUPPORTING INFORMATION AVAILABLE
Supplementary Tables S1 to S5 related to X-ray structure of compound 10(α) are available free of charge via the Internet at http://pubs.acs.org.
REFERENCE
- 1.Schmid RD, Verger R. Lipases: interfacial enzymes with attractive applications. Angew. Chem. Int. Ed. 1998;37:1608–1633. doi: 10.1002/(SICI)1521-3773(19980703)37:12<1608::AID-ANIE1608>3.0.CO;2-V. [DOI] [PubMed] [Google Scholar]
- 2.Vulfson E. Lipases: Their Structure, Biochemistry, and Application. Cambridge, UK: Cambridge University Press; 1994. Industrial applications of lipases; pp. 271–288. [Google Scholar]
- 3.Lengsfeld H, Beaumier-Gallon G, Chahinian H, De Caro A, Verger R, Laugier R, Carrière F. Physiology of Gastrointestinal Lipolysis and Therapeutical Use of Lipases and Digestive Lipase Inhibitors. In: Müller G, Petry S, editors. Lipases and phospholipases in drug development. Weinheim: Wiley-VCH; 2004. pp. 195–223. [Google Scholar]
- 4.Hadvàry P, Sidler W, Meister W, Vetter W, Wolfer H. The lipase inhibitor tetrahydrolipstatin binds covalently to the putative active site serine of pancreatic lipase. J. Biol. Chem. 1991;266:2021–2027. [PubMed] [Google Scholar]
- 5.Carrière F, Renou C, Ransac S, Lopez V, De Caro J, Ferrato F, De Caro A, Fleury A, Sanwald-Ducray P, Lengsfeld H, Beglinger C, Hadvary P, Verger R, Laugier R. Inhibition of gastrointestinal lipolysis by Orlistat during digestion of test meals in healthy volunteers. Am. J. Physiol. Gastrointest. Liver Physiol. 2001;281:G16–G28. doi: 10.1152/ajpgi.2001.281.1.G16. [DOI] [PubMed] [Google Scholar]
- 6.Yang PY, Liu K, Ngai MH, Lear MJ, Wenk MR, Yao SQ. Activity-Based Proteome Profiling of Potential Cellular Targets of Orlistat - An FDA-Approved Drug with Anti-Tumor Activities. J. Am. Chem. Soc. 2010;132:656–666. doi: 10.1021/ja907716f. [DOI] [PubMed] [Google Scholar]
- 7.Du J, Wang Z. Therapeutic potential of lipase inhibitor orlistat in Alzheimer's disease. Medical Hypotheses. 2009;73:662–663. doi: 10.1016/j.mehy.2009.04.046. [DOI] [PubMed] [Google Scholar]
- 8.Nelson RH, Miles JM. The use of orlistat in the treatment of obesity, dyslipidaemia and Type 2 diabetes. Expert Opin. Pharmacother. 2005;6:2483–2491. doi: 10.1517/14656566.6.14.2483. [DOI] [PubMed] [Google Scholar]
- 9.Kremer L, de Chastellier C, Dobson G, Gibson KJC, Bifani P, Balor S, Gorvel JP, Locht C, Minnikin DE, Besra GS. Identification and structural characterization of an unusual mycobacterial monomeromycolyl-diacylglycerol. Mol. Microbiol. 2005;57:1113–1126. doi: 10.1111/j.1365-2958.2005.04717.x. [DOI] [PubMed] [Google Scholar]
- 10.Dhouib R, Ducret A, Hubert P, Carriere F, Dukan S, Canaan S. Watching intracellular lipolysis in mycobacteria using time lapse fluorescence microscopy. Biochim. Biophys. Acta. 2011;1811:234–241. doi: 10.1016/j.bbalip.2011.01.001. [DOI] [PubMed] [Google Scholar]
- 11.West NP, Cergol KM, Xue M, Randall EJ, Britton WJ, Payne RJ. Inhibitors of an essential mycobacterial cell wall lipase (Rv3802c) as tuberculosis drug leads. Chem. Commun (Camb) 2011;47:5166–5168. doi: 10.1039/c0cc05635a. [DOI] [PubMed] [Google Scholar]
- 12.Hiratake J. Enzyme inhibitors as chemical tools to study enzyme catalysis: rational design, synthesis, and applications. Chem. Record. 2005;5:209–228. doi: 10.1002/tcr.20045. [DOI] [PubMed] [Google Scholar]
- 13.Jaeger K-E, Ransac S, Dijkstra BW, Colson C, Vanheuvel M, Misset O. Bacterial lipases. FEMS Microbiology Reviews. 1994;15:29–63. doi: 10.1111/j.1574-6976.1994.tb00121.x. [DOI] [PubMed] [Google Scholar]
- 14.Côtes K, Bakala N’Goma J, Dhouib R, Douchet I, Maurin D, Carrière F, Canaan S. Lipolytic enzymes in Mycobacterium tuberculosis. Appl. Microbiol. Biotechnol. 2008;78:741–749. doi: 10.1007/s00253-008-1397-2. [DOI] [PubMed] [Google Scholar]
- 15.Bandyopadhyay S, Dutta S, Spilling CD, Dupureur CM, Rath NP. Synthesis and Biological Evaluation of a Phosphonate Analog of the Natural Acetyl Cholinesterase Inhibitor Cyclophostin. J. Org. Chem. 2008;73:8386–8391. doi: 10.1021/jo801453v. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Dutta S, Malla RK, Bandyopadhyay S, Spilling CD, Dupureur CM. Synthesis and kinetic analysis of some phosphonate analogs of cyclophostin as inhibitors of human acetylcholinesterase. Bioorg. Med. Chem. 2010;18:2265–2274. doi: 10.1016/j.bmc.2010.01.063. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Engel R. Phosphonates as analogues of natural phosphates. Chem. Rev. 1977;77:349–367. [Google Scholar]
- 18.Metcalf WW, van der Donk WA. Biosynthesis of phosphonic and phosphinic acid natural products. Annu Rev biochem. 2009;78:65. doi: 10.1146/annurev.biochem.78.091707.100215. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Cavalier J-F, Buono G, Verger R. Covalent inhibition of digestive lipases by chiral phosphonates. Acc. Chem. Res. 2000;33:579–589. doi: 10.1021/ar990150i. [DOI] [PubMed] [Google Scholar]
- 20.Kurokawa T, Suzuki K, Hayaoka T, Nakagawa T, Izawa T, Kobayashi M, Harada N. Cyclophostin, acetylcholinesterase inhibitor from Streptomyces lavendulae. J. Antibiot. (Tokyo) 1993;46:1315–1318. doi: 10.7164/antibiotics.46.1315. [DOI] [PubMed] [Google Scholar]
- 21.Malla RK, Bandyopadhyay S, Spilling CD, Dutta S, Dupureur CM. The First Total Synthesis of (±)-Cyclophostin and (±)-Cyclipostin P: Inhibitors of the Serine Hydrolases Acetyl Cholinesterase and Hormone Sensitive Lipase. Org. Lett. 2011;13:3094–3097. doi: 10.1021/ol200991x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Vertesy L, Beck B, Bronstrup M, Ehrlich K, Kurz M, Muller G, Schummer D, Seibert G. Cyclipostins, novel hormone-sensitive lipase inhibitors from Streptomyces sp. DSM 13381. II. Isolation, structure elucidation and biological properties. J. Antibiot. (Tokyo) 2002;55:480–494. doi: 10.7164/antibiotics.55.480. [DOI] [PubMed] [Google Scholar]
- 23.Seibert G, Toti L, Wink J. Treating mycobacterial infections with cyclipostins. WO/2008/025449. [Google Scholar]
- 24.Carrière F, Rogalska E, Cudrey C, Ferrato F, Laugier R, Verger R. In vivo and in vitro studies on the stereoselective hydrolysis of tri- and diglycerides by gastric and pancreatic lipases. Bioorg. Med. Chem. 1997;5:429–435. doi: 10.1016/s0968-0896(96)00251-9. [DOI] [PubMed] [Google Scholar]
- 25.Withers-Martinez C, Carrière F, Verger R, Bourgeois D, Cambillau C. A pancreatic lipase with a phospholipase A1 activity: crystal structure of a chimeric pancreatic lipase-related protein 2 from guinea pig. Structure. 1996;4:1363–1374. doi: 10.1016/s0969-2126(96)00143-8. [DOI] [PubMed] [Google Scholar]
- 26.Thirstrup K, Carrière F, Hjorth S, Rasmussen PB, Woldike H, Nielsen PF, Thim L. One-step purification and characterization of human pancreatic lipase expressed in insect cells. FEBS Letters. 1993;327:79–84. doi: 10.1016/0014-5793(93)81044-z. [DOI] [PubMed] [Google Scholar]
- 27.Aloulou A, Rodriguez JA, Fernandez S, van Oosterhout D, Puccinelli D, Carrière F. Exploring the specific features of interfacial enzymology based on lipase studies. Biochim. Biophys. Acta. 2006;1761:995–1013. doi: 10.1016/j.bbalip.2006.06.009. [DOI] [PubMed] [Google Scholar]
- 28.De Caro J, Sias B, Grandval P, Ferrato F, Halimi H, Carriere F, De Caro A. Characterization of pancreatic lipase-related protein 2 isolated from human pancreatic juice. Biochim. Biophys. Acta. 2004;1701:89–99. doi: 10.1016/j.bbapap.2004.06.005. [DOI] [PubMed] [Google Scholar]
- 29.Eydoux C, De Caro J, Ferrato F, Boullanger P, Lafont D, Laugier R, Carrière F, De Caro A. Further biochemical characterization of human pancreatic lipase-related protein 2 expressed in yeast cells. J. Lipid Res. 2007;48:1539–1549. doi: 10.1194/jlr.M600486-JLR200. [DOI] [PubMed] [Google Scholar]
- 30.Amara S, Lafont D, Fiorentino B, Boullanger P, Carrière F, De Caro A. Continuous measurement of galactolipid hydrolysis by pancreatic lipolytic enzymes using the pH-stat technique and a medium chain monogalactosyl diglyceride as substrate. Biochim. Biophys. Acta. 2009;1791:983–990. doi: 10.1016/j.bbalip.2009.05.002. [DOI] [PubMed] [Google Scholar]
- 31.Longhi S, Mannesse M, Verheij HM, de Haas GH, Egmond M, Knoops-Mouthuy E, Cambillau C. Crystal structure of cutinase covalently inhibited by a triglyceride analogue. Protein Science. 1997;6:275–286. doi: 10.1002/pro.5560060202. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Egmond MR, de Vlieg J. Fusarium solani pisi cutinase. Biochimie. 2000;82:1015–1021. doi: 10.1016/s0300-9084(00)01183-4. [DOI] [PubMed] [Google Scholar]
- 33.Parker SK, Curtin KM, Vasil ML. Purification and Characterization of Mycobacterial Phospholipase A: an Activity Associated with Mycobacterial Cutinase? J. Bacteriol. 2007;189:4153–4160. doi: 10.1128/JB.01909-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Côtes K, Dhouib R, Douchet I, Chahinian H, De Caro A, Carriere F, Canaan S. Characterization of an exported monoglyceride lipase from Mycobacterium tuberculosis possibly involved in the metabolism of host cell membrane lipids. Biochem. J. 2007;408:417–427. doi: 10.1042/BJ20070745. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Saravanan P, Dubey VK, Patra S. Potential Selective Inhibitors against Rv0183 of Mycobacterium tuberculosis Targeting Host Lipid Metabolism. Chem. Biol. Drug Des. 2012;79:1056–1062. doi: 10.1111/j.1747-0285.2012.01373.x. [DOI] [PubMed] [Google Scholar]
- 36.Deb C, Daniel J, Sirakova T, Abomoelak B, Dubey V, Kolattukudy P. A Novel Lipase Belonging to the Hormone-sensitive Lipase Family Induced under Starvation to Utilize Stored Triacylglycerol in Mycobacterium tuberculosis. J. Biol. Chem. 2006;281:3866–3875. doi: 10.1074/jbc.M505556200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Mishra K, de Chastellier C, Narayana Y, Bifani P, Brown A, Besra G, Katoch V, Joshi B, Balaji K, Kremer L. Functional Role of the PE Domain and Immunogenicity of the Mycobacterium tuberculosis Triacylglycerol Hydrolase LipY. Infect. Immun. 2008;76:127–140. doi: 10.1128/IAI.00410-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Yan B, Spilling CD. Synthesis of Cyclopentenones via Intramolecular HWE and the Palladium-Catalyzed Reactions of Allylic Hydroxy Phosphonate Derivatives. J. Org. Chem. 2008;73:5385–5396. doi: 10.1021/jo8004028. [DOI] [PubMed] [Google Scholar]
- 39.He A, Yan B, Thanavaro A, Spilling CD, Rath NP. Synthesis of nonracemic allylic hydroxy phosphonates via alkene cross metathesis. J. Org. Chem. 2004;69:8643–8651. doi: 10.1021/jo0490090. [DOI] [PubMed] [Google Scholar]
- 40.Lainé C, Mornet E, Lemiègre L, Montier T, Cammas-Marion S, Neveu C, Carmoy N, Lehn P, Benvegnu T. Folate-Equipped Pegylated Archaeal Lipid Derivatives: Synthesis and Transfection Properties. Chem. Eur. J. 2008;14:8330–8340. doi: 10.1002/chem.200800950. [DOI] [PubMed] [Google Scholar]
- 41.Ransac S, Gargouri Y, Marguet F, Buono G, Beglinger C, Hildebrand P, Lengsfeld H, Hadváry P, Verger R. Covalent inactivation of lipases. Methods Enzymol. 1997;286:190–231. doi: 10.1016/s0076-6879(97)86012-0. [DOI] [PubMed] [Google Scholar]
- 42.Ransac S, Rivière C, Gancet C, Verger R, de Haas GH. Competitive inhibition of lipolytic enzymes. I. A kinetic model applicable to water-insoluble competitive inhibitors. Biochim. Biophys. Acta. 1990;1043:57–66. doi: 10.1016/0005-2760(90)90110-j. [DOI] [PubMed] [Google Scholar]
- 43.Cavalier J-F, Ransac S, Verger R, Buono G. Inhibition of human gastric and pancreatic lipases by chiral alkylphosphonates. A kinetic study with 1,2-didecanoyl-sn-glycerol monolayer. Chem. Phys. Lipids. 1999;100:3–31. doi: 10.1016/s0009-3084(99)00028-6. [DOI] [PubMed] [Google Scholar]
- 44.Roussel A, Miled N, Berti-Dupuis L, Riviere M, Spinelli S, Berna P, Gruber V, Verger R, Cambillau C. Crystal structure of the open form of dog gastric lipase in complex with a phosphonate inhibitor. J. Biol. Chem. 2002;277:2266–2274. doi: 10.1074/jbc.M109484200. [DOI] [PubMed] [Google Scholar]
- 45.Egloff M-P, Marguet F, Buono G, Verger R, Cambillau C, van Tilbeurgh H. The 2.46 Å resolution structure of the pancreatic lipase-colipase complex inhibited by a C11 alkyl phosphonate. Biochemistry. 1995;34:2751–2762. doi: 10.1021/bi00009a003. [DOI] [PubMed] [Google Scholar]
- 46.Pappin DJC, Hojrup P, Bleasby AJ. Rapid identification of proteins by peptide-mass fingerprinting. Curr. Biol. 1993;3:327–332. doi: 10.1016/0960-9822(93)90195-t. [DOI] [PubMed] [Google Scholar]
- 47.Verger R, de Haas GH. Interfacial enzyme kinetics of lipolysis. Annu. Rev. Biophys. Bioeng. 1976;5:77–117. doi: 10.1146/annurev.bb.05.060176.000453. [DOI] [PubMed] [Google Scholar]
- 48.Sheldrick GM. A short history of SHELX. Acta Crystallogr A. 2008;64:112–122. doi: 10.1107/S0108767307043930. [DOI] [PubMed] [Google Scholar]
- 49.Belle V, Fournel A, Woudstra M, Ranaldi S, Prieri F, Thome V, Currault J, Verger R, Guigliarelli B, Carriere F. Probing the Opening of the Pancreatic Lipase Lid Using Site-Directed Spin Labeling and EPR Spectroscopy. Biochemistry. 2007;46:2205–2214. doi: 10.1021/bi0616089. [DOI] [PubMed] [Google Scholar]
- 50.Hjorth A, Carrière F, Cudrey C, Wöldike H, Boel E, Lawson DM, Ferrato F, Cambillau C, Dodson GG, Thim L, Verger R. A structural domain (the lid) found in pancreatic lipases is absent in the guinea pig (phospho)lipase. Biochemistry. 1993;32:4702–4707. doi: 10.1021/bi00069a003. [DOI] [PubMed] [Google Scholar]
- 51.Gruber V, Berna P, Arnaud T, Bournat P, Clément C, Mison D, Olagnier B, Philippe L, Theisen M, Baudino S. Large-scale production of a therapeutic protein in transgenic tobacco plants: effect of subcellular targeting on quality of a recombinant dog gastric lipase. Molecular Breeding. 2001;7:329–340. [Google Scholar]
- 52.Lauwereys M, de Geus P, de Meutter J, Stanssens P, Matthyssens G. Cloning, expression and characterization of cutinase, a fungal lipolytic enzyme. GBF monogr. 1991;16:243–251. [Google Scholar]
- 53.Brust B, Lecoufle M, Tuaillon E, Dedieu L, Canaan S, Valverde V, Kremer L. Mycobacterium tuberculosis Lipolytic Enzymes as Potential Biomarkers for the Diagnosis of Active Tuberculosis. PLoS ONE. 2011;6:e25078. doi: 10.1371/journal.pone.0025078. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Fernandez S, Jannin V, Rodier J-D, Ritter N, Mahler B, Carriere F. Comparative study on digestive lipase activities on the self emulsifying excipient Labrasol(R), medium chain glycerides and PEG esters. Biochim. Biophys. Acta. 2007;1771:633–640. doi: 10.1016/j.bbalip.2007.02.009. [DOI] [PubMed] [Google Scholar]
- 55.Erlanson C, Borgström B. Tributyrin as a substrate for determination of lipase activity of pancreatic juice and small intestinal content. Scand. J. Gastroenterol. 1970;5:293–295. [PubMed] [Google Scholar]
- 56.Bezzine S, Ferrato F, Ivanova MG, Lopez V, Verger R, Carriere F. Human pancreatic lipase: colipase dependence and interfacial binding of lid domain mutants. Biochemistry. 1999;38:5499–5510. doi: 10.1021/bi982601x. [DOI] [PubMed] [Google Scholar]
- 57.Carrière F, Moreau H, Raphel V, Laugier R, Benicourt C, Junien JL, Verger R. Purification and biochemical characterization of dog gastric lipase. Eur. J. Biochem. 1991;202:75–83. doi: 10.1111/j.1432-1033.1991.tb16346.x. [DOI] [PubMed] [Google Scholar]
- 58.Marguet F, Cudrey C, Verger R, Buono G. Digestive lipases: Inactivation by phosphonates. Biochim. Biophys. Acta. 1994;1210:157–166. doi: 10.1016/0005-2760(94)90116-3. [DOI] [PubMed] [Google Scholar]
- 59.Mannesse MLM, Boots JWP, Dijkman R, Slotboom AJ, Vanderhijden HTWV, Egmond MR, Verheij HM, Dehaas GH. Phosphonate analogues of triacylglycerols are potent inhibitors of lipase. Biochim. Biophys. Acta. 1995;1259:56–64. doi: 10.1016/0005-2760(95)00145-3. [DOI] [PubMed] [Google Scholar]
- 60.Trott O, Olson AJ. AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 2010;31:455–461. doi: 10.1002/jcc.21334. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Seeliger D, de Groot BL. Ligand docking and binding site analysis with PyMOL and Autodock/Vina. J. Comput. Aided Mol. Des. 2010;24:417–422. doi: 10.1007/s10822-010-9352-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.