Abstract
Background
Diagnostic analysis of patients with developmental disorders has improved over recent years largely due to the use of microarray technology. Array methods that facilitate copy number analysis have enabled the diagnosis of up to 20% more patients with previously normal karyotyping results. A substantial number of patients remain undiagnosed, however.
Methods and Results
Using the Genome-Wide Human SNP array 6.0, we analyzed 35 patients with a developmental disorder of unknown cause and normal array comparative genomic hybridization (array CGH) results, in order to characterize previously undefined genomic aberrations. We detected no seemingly pathogenic copy number aberrations. Most of the vast amount of data produced by the array was polymorphic and non-informative. Filtering of this data, based on copy number variant (CNV) population frequencies as well as phenotypically relevant genes, enabled pinpointing regions of allelic homozygosity that included candidate genes correlating to the phenotypic features in four patients, but results could not be confirmed.
Conclusions
In this study, the use of an ultra high-resolution SNP array did not contribute to further diagnose patients with developmental disorders of unknown cause. The statistical power of these results is limited by the small size of the patient cohort, and interpretation of these negative results can only be applied to the patients studied here. We present the results of our study and the recurrence of clustered allelic homozygosity present in this material, as detected by the SNP 6.0 array.
Keywords: Developmental disorder, SNP array, Diagnostic yield
Background
The diagnostic yield of microarray comparative genomic hybridizations (array CGH) has already proven to exceed that of cytogenetic methods, except when it comes to balanced rearrangements. A consensus statement suggests that microarrays should be used as the first line of testing for developmental disorders of unknown cause [1]. However, as our previous study shows, approximately 80 % of patients with a developmental disorder of unknown cause (mental retardation and/or malformations and/or neurological disorders) remain undiagnosed even by array CGH analysis (44 K, 180 K, or 244 K) [2]. Thus, other methods are clearly needed to define the pathogenic mechanisms. It is plausible that small copy number variants (CNVs) may go undetected if the probe coverage is limited, as it may be in low-resolution arrays. By increasing the resolution, one would thus expect to detect increasingly smaller pathogenic CNVs.
The frequency of uniparental disomy (UPD) in newborns is reportedly ~0.029% [3]. Around 1,100 cases of whole chromosome UPD and some 120 reports on segmental UPD are described in the literature [4]. Some recessive diseases are expressed in children who have inherited the mutation form a single carrier parent [5]. The explanation for this is a meiotic or very early mitotic recombination event between parental homologous chromosomes causing segmental UPD of the genomic segment containing the mutation, and thereby causing a reduction to homozygosity.
As the market is flooded with new arrays, most having an increased resolution and a promise of ever higher detection rates, the question remains what the added value is of these ultra-high resolution arrays. To test this, we used an array with 1.8 million probes and an average resolution of 0.7Kb to analyze samples of 35 patients with developmental disorder of unknown cause, normal karyotype, and normal array CGH results by use of Agilent 44 K, 180 K, or 244 K platforms (Agilent Technologies, Santa Clara, CA, USA).
Methods
Participants
Patients with previous normal array CGH results were asked to participate in the project. All 35 patients were Finnish of origin and had mild to severe mental retardation, associated with dysmorphic features and/or congenital anomalies (Table 1). In addition, 16 patients also had epilepsy. For diagnostic purposes patients had previously been analyzed by whole-genome array CGH (Human Genome CGH Microarray, Agilent Technologies, Santa Clara, CA); 20 using the 244 K platform, 8 using a 180 K platform, and 7 using a 44 K platform. Informed consent was given by all participating families. Blood samples were collected from all patients and their parents. Ethical permission for this project was given by the Ethics Review Board of Helsinki and Uusimaa Hospital District.
Table 1.
Clinical characteristics of patients studied
| P. Nr. | Growth | Head and neck | Eyes and vision | Ears and hearing | Face | Cardiovascular | Genitourinary | Skeletal and limb defects | Neurologic | Other |
|---|---|---|---|---|---|---|---|---|---|---|
| 1. |
Obesity |
|
Astigmatism |
|
|
|
|
Postaxial polydactyly (one foot) Short meta-carpals V finger clino-dactyly |
ID Hypotonia |
Bardet-Biedl suspected |
| 2 |
|
|
|
|
|
|
|
|
Severe DD No walk/crawl No speech Epilepsy Drooling |
|
| 3. |
|
Dolicocephaly Narrow, prominent forehead Low, uneven hairline |
Epichantal folds |
|
|
|
|
|
DD Abnormal pons |
Hemangiomas |
| 4. |
Short stature |
|
|
|
Low nasal bridge |
|
Horseshoe kidney Anal atresia |
Small hands and feet |
ID |
Balanced t(X;13)(q28;q12) |
| 5. |
|
|
|
Simple ears |
Thick and straight eyebrows Broad nasal bridge Long philtrum Retrognathia |
|
|
|
Autism ID Epilepsy |
|
| 6. |
Short stature |
Microcephaly |
Blindness Optic nerve hypoplasia |
|
|
|
|
Scoliosis |
ID Epilepsy Severe hypotonia |
|
| 7. |
|
Hydrocephalus |
|
|
|
|
|
|
Brain malformation Severe DD |
|
| 8 |
|
Microcephaly |
Hypertelorism |
|
Small nose Low nasal bridge Tented upper lip |
|
|
|
Severe DD Severe epilepsy |
ATRX suspected |
| 9. |
|
|
Severe optic atrophy Impaired vision |
|
|
|
|
|
ID Epilepsy Cortical atrophy |
|
| 10. |
|
|
Mild hypertelorism |
Low-set ears |
Triangular face Small jaw High palate Thin upper lip |
|
|
Hyper-extensible joints |
ID Autistic features Intractable epilepsy |
Frax-dna, SCN1A, CLN8 4p-FISH normal |
| 11. |
|
|
Upslanting palpebal fissures |
Large earlobes |
Small jaw |
|
|
Clubfoot |
ID Autistic features Intractable epilepsy |
|
| 12. |
|
|
Epichantic folds |
Large earlobes |
Flat face |
|
|
Tapering fingers |
ID Autistic features ADHD |
Balanced t(2;9)(q13q22.3) de novo |
| 13. |
|
Macrocephaly |
|
|
|
|
|
|
Severe ID Hypotonia Autism Epilepsy |
Inv 2(p13p25) mat., DMPK mutation negative |
| 14. |
|
|
Severe myopia Cataracta |
|
Synophrys Curved eyebrows Upturned pinched nose Big mouth Full lips |
Atrial septum defect |
|
|
Severe DD Epilepsy |
|
| 15. |
|
Microcephaly |
Impaired vision |
|
|
Ventricular septum defect |
|
|
Epilepsy DD |
|
| 16. |
|
|
|
|
|
|
|
|
ID Beahvioural disturbances Autism |
No malformation or dysmorphism |
| 17. |
|
|
|
|
|
|
|
|
ID DD |
|
| 18. |
Growth retardation |
|
Hypertelorism |
|
|
Mild ventricular septum defect |
|
|
ID |
|
| 19. & 20. |
|
|
|
|
|
|
|
|
DD |
No structural defects |
| 21 |
|
Microcephaly |
Strabismus |
Missing lobuli |
Small nose Low nasal bridge Smooth philtrum Thin lips |
|
|
Proximal thumbs Pes planus |
ID Intractable epilepsy Ataxia |
|
| 22. |
Pre- and postnatal growth retardation |
|
|
|
Broad nasal root Short nose Bifid nasal tip |
|
Cryptorchidism Hypoplastic scrotum |
Scoliosis Syndactylies |
Slow motor development Hypotonia Expressive language disorder |
Congenital contractures Dimples |
| 23. |
|
|
|
|
Mild dysmorphism |
|
|
|
ID Epilepsy |
|
| 24. |
|
Microcephaly |
Hypertelorism Epicanthic folds Disorder of visual cortex |
Low-set and posteriorly rotated ears |
Micrognathia Cleft palate |
|
|
|
ID Epilepsy Hypoplastic cerebellar vermis |
Monozygotic twin, twin sister healthy |
| 25. |
Tall stature Advanced bone age |
|
Deep set eyes Hypotelorism Epicanthic folds Strabismus |
|
Short nose Anteverted nares Tented upper lip |
|
Cryptorchidism |
|
ID No speech Autism |
Glypican-3 and PHF6 mutation analyses negative |
| 26. |
Short stature |
|
|
|
|
|
|
|
ID Intractable epilepsy Tremor Myoclonias Distal spasticity |
|
| 27. |
Small for Gestational Age |
|
Downslanting palpebral fissures Strabismus |
|
Frontal bossing |
|
|
Exostosis (familial) Broad hallux Overriding toes Scoliosis |
ID Epilepsy |
Inguinal hernia |
| 28. |
|
|
Downslanting palpebral fissures |
Hearing impairment |
Coarse hair Thick eyebrows Thick lips Malposition of teeth |
Hypertrophic cardiomyopathy |
|
Hip dis-placement Long thin bones |
Normal intelligence |
|
| 29. |
|
|
|
|
|
|
|
|
Epilepsy ID Alternating hemiplegia of childhood |
|
| 30. |
Small for gestational age Prematurity Short stature |
Microcephaly |
Severe myopia Coloboma of papillae Optic atrophy Nystagmus Strabismus |
|
High palate Gum hypertrophy |
Coarctation of aorta |
Inguinal hernia |
|
ID Intractable epilepsy Hemiparesis (peri-ventricular leukomalacia) |
|
| 31. |
|
|
Central blindness Nystagmus |
|
|
|
|
|
ID Intractable epilepsy Hypotonia Distal spasticity |
|
| 32. |
Short stature |
|
|
|
|
|
|
|
Normal development |
Vomiting Feeding difficulties |
| 33. |
|
Neck fistula |
|
|
Dysmorphic malocclusion of teeth |
Uni-ventricular heart |
|
|
Brain atrophy Epilepsy |
Simian-crease Sinus pilonidalis |
| 34. |
|
|
|
|
|
|
|
|
ID Intractable epilepsy Hypotonia Distal spasticity |
|
| 35. | Dolicocephaly | Epichantic fold | Simple ears | Thin upper-lip Long philtrum Broad nasal bridge | ID Arnold Chiari malformation |
The table presents the clinical characteristics of the 35 patients studied. DD = developmental delay, ID = intellectual disability.
SNP array
DNA was extracted from blood samples according to standard protocols. Analysis by the Genome-wide human SNP array 6.0 was performed according to manufacturer protocols (Affymetrix, Santa Clara, CA, USA). In short; DNA was digested, ligated to adapters, and amplified by PCR. Samples were purified using magnetic beads and further fragmented and labelled with biotin. After hybridization arrays were washed and stained with streptavidin and anti-streptavidin antibodies and finally the arrays were scanned using the Affymetrix GeneChip scanner.
Analysis
Data was extracted from the scanned image using the Genotyping console software V.3.0.2, creating a CEL file. Areas containing CNVs and allelic homozygosity were detected using the Hidden-Markow-Model. The resulting data was analyzed using the Chromosome Analysis Suite software V.1.0.
Reference data
Data was extracted by comparison to a reference data set established from 90 Caucasian individuals, which had previously been analyzed using the SNP 6.0 array in the HapMap project (http://www.hapmap.org). As an additional in-house reference set, we used results of 54 individuals studied using the SNP 6.0 array, whereof 19 healthy normal relatives of the patients, and 35 unrelated patients with an unexplained developmental disorder. Sample identities were kept anonymous and the information was only used for reference purposes. These in-house reference sets were used to filter out polymorphic changes in the patient data studied here.
Selected CNVs of the patients were compared to a Finnish population cohort [6]. This population cohort data consist of CNVs detected, using whole-genome SNP analysis, in 2163 healthy Finnish individuals with PennCNV [7]. In the population data, low quality samples (N = 98) with Log R Ratio standard deviation of probe signal intensities > 0.35 or > 115 CNV calls were excluded. Only CNVs with three or more probes were included in the final population data. CNV calls of the study samples were clustered into CNV regions when individual CNVs overlapped by one or more base pairs.
Filtering relevant CNVs and potential UPDs
The first set of default filtering marked all duplications and deletions ≥0.7Kb and all allelic homozygosities ≥100Kb containing at least 10 markers to be included. This was based on the theoretical resolution of the array being 0.7Kb, in addition to information from the HapMap phase 1 study showing that approximately 70% of common haplotype blocks are ≤100Kb [8] . Allelic homozygosity was called by the analysis software where there was a stretch of homozygous SNPs in a chromosomal segment.
The second filtering was based on the comparison of all aberrations detected in the patients of this study (N = 35), an in-house patient reference set (N = 35) and an in-house normal reference set (N = 19) as well as the database of genomic variants (DGV) [9]. Aberrations of one patient that were not present in any of the other groups (potentially “unique”) were further processed by studying their genetic content and association to diseases and traits, as stated in publications or OMIM, and whether these correlated to the patient’s phenotype. If this did not yield a candidate aberration, all aberrations of a patient were reviewed based on only the associated OMIM disease, despite the frequency of similar changes in the reference sets. The CNVs that were picked out as potential candidates were further compared to CNV data from 2,065 healthy Finnish individuals. Only those aberrations that were present in less than 50 individuals of the Finnish population cohort were initially considered as potentially pathogenic.
Validation of candidate aberrations
Only aberrations that were considered to potentially associate with patient’s phenotype were attempted validation.
Microsatellite marker analysis
Potential segmental UPDs were analyzed by microsatellite marker analysis (chr15: D15S204, D15S124; chr6: D6S468, D6S2418; chr11: D11S4140; chr17: D17S578, D17S1832, D17S1828).
The markers were selected based on their location, and on information that they are highly polymorphic in the Caucasian population. Fragments were labelled with a fluorescent HEX label, and separated on an Applied Biosystem 3730XL (Life Technologies, Carlsbad, CA, USA) capillary electrophoresis instrument, according to manufacturer recommendations. Genotypes were called using Applied Biosystems GeneMapper 3.7 software.
Results
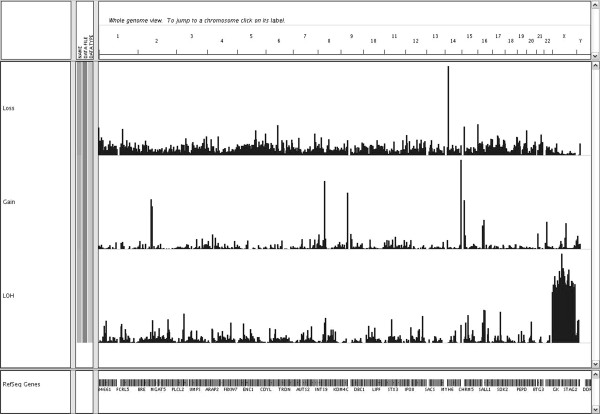
The immense amount of data created by the Genome-wide human SNP array 6.0 warrants filtering for clear interpretation. The Genotyping Console software identified between 200–1000 changes per patient (Figure 1), depending on the technical quality of the result. More changes were detected in samples with lesser quality. After filtering, based on the uniqueness of the CNVs or regions of homozygosity compared to the references, each patient presented 8–20 unique changes (≥90% CNVs) on average. In samples with lesser quality, ≥100 unique changes were detected. Further research on gene content and phenotypes previously mapped to these regions revealed 23 CNVs and 28 regions of homozygosity that putatively correlated with the clinical phenotype in 26 patients. Nine patients had no CNVs or regions of homozygosity spanning known genes or genes known to associate with a disease that correlated with the patient’s phenotype, and their results were thus considered normal. The associated phenotypes related to the aberration found in 26 patients were further evaluated by the patients’ clinicians, and the frequencies of the observed CNVs were monitored in the Finnish population cohort (n = 2,065). As a result, in four patients, a region of allelic homozygosity was considered a potential candidate for causation of their clinical state (Table 2). No CNVs were considered candidates after clinical evaluation.
Figure 1.
Frequency of CNVs and allelic homozygosity. The figure visualizes the frequency of copy number changes (loss and gain) and regions of allelic homozygosity (LOH) in 70 patients (patients of this study N = 35 and the in-house reference set N = 35) with developmental disorders of unknown cause as seen by the Integrative Genomics Viewer (IGV) software V.1.5 (The Broad Institute, Cambridge, MA, USA). The vertical bars show the percentage of patients that have a CNV in a particular area of a chromsome. The higher the bar, the higher the percentage, thus indicating as CNP.
Table 2.
Results from microsatellite marker analysis
| Patient | Chromosome location | Marker | Location start | Location stop | Patient | Mother | Father | Associated OMIMdisease(gene) |
|---|---|---|---|---|---|---|---|---|
| 1 |
15q23q24.1 |
D15S204 |
72300758 |
72300879 |
123/123 |
123/125 |
123/125 |
MIM #209900, Bardet-Biedl Syndrome (BBS4) |
| |
|
D15S124 |
73092468 |
73092572 |
104/106 |
106/106 |
104/106 |
|
| 17 |
6q16.3 |
D6S468 |
101630330 |
101630479 |
155/159 |
159/159 |
155/155 |
MIM #611092, Mental retardation (GRIK2) |
| |
|
D6S2418 |
101352425 |
101352639 |
222/230 |
222/248 |
230/238 |
|
| 32 |
11q13.4 |
DS11S4140 |
71945684 |
71945874 |
195/195 |
195/197 |
195/197 |
MIM #270400, Smith-Lemli-Opitz syndrome (DHCR7) |
| 28 |
17p13.2p13.1 |
D17S578 |
6824007 |
6824153 |
173/173 |
173/173 |
155/173 |
MIM #201475, AcylCoA dehydrogenase deficiency (ACADVL) |
| |
|
D17S1832 |
5972677 |
5972867 |
173/185 |
173/185 |
171/173/185/193 |
|
| D17S1828 | 3810467 | 3810673 | 220/220 | 214/220 | 214/220 |
The table presents the results of microsatellite marker analysis of 4 patients and their parents, suggesting biparental inheritance of the genomic segment, and thus exclude segmental UPD. The numbers in the patient/mother/father column represent the two markers detected. In all cases, the patient has likely inherited one marker from each parent.
We further attempted verification of four potential segmental UPDs by microsatellite marker analysis. In all the four cases we observed two distinct alleles with at least some of the more informative multiallelic markers, suggesting that the observed LOHs were most probably caused by the same allelic SNP haplotypes being inherited from both the parents (Table 2).
Further comparison, of all patient SNP array data to the normal data from unaffected individuals, revealed 21 regions of clustering (≥40 % frequency) of allelic homozygosity to specific locations of the genome (Table 3). However, no significant differences were detected between the frequencies of clustered regions in patients and the unaffected relatives in this small set of samples.
Table 3.
Regions of clustered allelic homozygosity
| Chromosome | Band | Appoximate range (Kb) | Frequency patients (n = 70) | Frequency normals (n = 19) |
|---|---|---|---|---|
|
1 |
p33-p32.3 |
48 700–53 300 |
47.7% |
47.7% |
|
1 |
q21.1-q21.2 |
145 800–148 500 |
49% |
50% |
|
2 |
q21.2-q21.3 |
134 334–136 693 |
42% |
58% |
|
3 |
p21.31-p21.1 |
46 500–52 500 |
71% |
68% |
|
4 |
p15.1 |
31 838–34 524 |
60% |
57% |
|
8 |
q22.2 |
99 200–101 200 |
47% |
63% |
|
8 |
p11.21-p11.1 |
41 870–43 270 |
49% |
47% |
|
8 |
q11.1-q11.21 |
47 040–49 000 |
46% |
68% |
|
10 |
p11.21 |
36 720–38 490 |
43% |
31% |
|
10 |
q22.2-q22.2 |
73 200–76 460 |
44% |
31% |
|
12 |
q21.32-q21.33 |
85 850–89 100 |
47% |
47% |
|
12 |
q24.11-q24.13 |
108 600–111 600 |
55% |
68% |
|
14 |
q23.3-q24.1 |
65 500–67 100 |
62% |
73% |
|
15 |
q12-q13.1 |
25 400–27 200 |
71% |
68% |
|
15 |
q15.1-q21.1 |
40 100–43 730 |
64% |
84% |
|
15 |
q23-q24.1 |
69 300–71 700 |
41% |
15% |
|
16 |
p11.2-p11.1 |
33 394–34 550 |
62% |
68% |
|
16 |
q11.2-q12.1 |
45 092–47 450 |
64% |
63% |
|
16 |
q21-q22.1 |
64 850–67 100 |
48% |
57% |
|
17 |
q22-q23.2 |
54 610–56 850 |
67% |
68% |
| 20 | q11.22-q11.23 | 31 910–35 500 | 68% | 42% |
The table presents the frequency (> 40%) of clustered regions of allelic homozygosity in the patient cohort (n = 70, including in-house reference) compared to the unaffected relatives (n = 19). Kilobase range according to Genome build 19 (NCBI 37).
Discussion
In our previous study of 150 patients with developmental disorders of unknown cause and a normal karyotype, we were able to identify a (potential) causative aberration in 18% of the patients, by using a 44 K or 244 K array CGH platform [2].
To determine whether, by increasing the resolution, any additional copy number changes or regions of UPD could be detected, we studied 35 patients with a normal array CGH result.
Allelic homozygosity is typically caused by linkage and co-segregation of certain blocks of DNA, termed haplotypes [8]. In a small founder population, such as the Finnish population, the founder effect increases the likelihood that the parents will have the same haplotype, and is as such not a segmental UPD [10]. True segmental UPD is typically due to a duplication in one chromosome and a reciprocal deletion in the other; or the fertilization of a disomic and monosomic gamete, somatic crossing over and subsequent trisomic rescue [11]. If the genomic segment harbours a recessive mutation, which subsequent to UPD will be present in two copies (reduction to homozygosity), it causes a recessive disease. Equally relevant, the segment can be preferentially imprinted, causing complete silencing, which is the equivalent of a deletion. Such presentations of recessive syndromes that are inherited from one normal parent are known in some 40 patients [12].
We were interested to see whether the regions of allelic homozygosity detected by the SNP array were in fact segmental UPDs and associated with an autosomal recessive disease. We were, however, unable to confirm these results and thus the SNP array did not yield more molecular diagnoses in this study of developmental disorders of unknown cause. This may be due to the fact that all patients had previously been studied by another high-resolution array, with 8.9Kb (244 K), 13Kb (180 K), and 35Kb (44 K) theoretical resolutions. Although several new CNVs, previously undetected by the array CGH platform, as well as regions of homozygosity were detected, the pathogenic relevance of these changes were considered insignificant in correlation to the patient’s phenotype. It is, however, possible that changes dismissed in this study are pathogenic by means of spanning genomic segments that do not directly involve disease genes, but rather their regulatory elements.
Our results differ from previously published studies using similar research settings. Bernardini et al. (2010), using the SNP 6.0 array platform with a 75Kb cut-off value for detected CNVs, reported potentially pathogenic CNVs in 6% of patients with normal array CGH result (44 K) [13]. Mannik et al. (2010), using another SNP array with a 50Kb resolution, reported a 23% detection rate in patients with a normal karyotype [14]. Bernardini et al. and Mannik et al. had higher detection rates than this study; perhaps as their first-line of array analysis was, at least partly, done by a lower-resolution method (35Kb and 50Kb respectively) compared to the first-line of detection in this study (8,9Kb, 13Kb and 35Kb). Also, it is important to note that the statistical power is limited by the small size of our patient cohort, and thus results are not entirely comparable with Bernardini and Mannik’s.
UPDs have not been reported in either of the above mentioned studies. However, in a study of 117 patients with a normal karyotype analysed using the 250 K SNP array (Affymetrix, Santa Clara, CA, USA), pathogenic CNVs were detected in 18 patients, and potentially pathogenic segmental UPDs ≥5 Mb in 5, verified by microsatellite marker analysis [15]. The presence of UPDs was also evaluated in another study of 120 patients, using a 500 K SNP array platform [16]. In that study they were unable to verify UPD in any of 121 detected regions of homozygosity in 72 patients with developmental disorder of unknown cause. In addition, in a study of 100 patients with developmental disorder, using a 500 K SNP array, two patients were found to have UPD, the clinical significance of which remained unclear [17]. Thus, UPDs are detectable using SNP arrays, but their clinical significance is difficult to interpret.
Conclusions
Although there is a clear added value of high-resolution arrays in various fields of genetics, it seems that there is a limit to how much the yield can be increased by increasing the theoretical resolution of the analysis platform. Despite the fact that the SNP array has increased probe spacing compared to the 44 K, 180 K, and 244 K array, and is able to detect more CNVs and regions of homozygosity, interpretation of the vast amount of data and pinpointing of pathogenic changes is difficult. One benefit of using a SNP based platform is the possibility to detect UPDs; however these are relatively rare findings.
This study had a limited number of patients, and so it can only be said that for this study group the optimal yield was conceived when using a resolution of approximately 9Kb [2]. Increasing the resolution beyond that did not confer more diagnoses. It must be emphasized, however, that a larger patient cohort needs to be studied in order to draw final conclusions on the added value of an ultra-high resolution array compared to others. Furthermore, as several reports have shown, some pathogenic aberrations span only a few exons and for detecting such small changes the sensitivity of the SNP 6.0 platform is adequate [18]. It is, however a challenge to filter results correctly and so for diagnostic purposes the choice of platform needs to be carefully considered. Patients with developmental disorders of unknown cause and normal array results may also harbour such small genomic changes (i.e. mutations and unbalanced rearrangements) that are difficult to interpret using microarrays and would require higher resolution methods, such as whole-exome sequencing. Interestingly, a recent study suggests that 80% of patients with a developmental disorder of unknown cause and normal array results can be diagnosed using whole-exome sequencing [19]. Only the future can tell.
Availability of supporting data
The data set supporting the results of this article is available in the CanGEM repository, http://www.cangem.org/browse.php.
Competing interests
The authors declare that they have no competing interests.
Authors’ contributions
LS set up the study, performed the laboratory work and analysis and drafted the manuscript. SA-M helped set up the study and assisted in clinical evaluation of results. TL, KA, MH, LM, and EK, contacted patients and assisted in clinical evaluation of results. IS performed bioinformatics filtering of the results. KK compared CNV findings with data from the Health2000 project. PL performed the microsatellite marker analysis. ML performed analysis for the Health2000 project. JS supervised the performance of the validation studies. SK helped set up and supervise the project. All authors assisted in drafting of the manuscript.
Pre-publication history
The pre-publication history for this paper can be accessed here:
Contributor Information
Linda Siggberg, Email: linda.siggberg@helsinki.fi.
Ala-Mello Sirpa, Email: sirpa.ala-mello@fimnet.fi.
Linnankivi Tarja, Email: tarja.linnankivi@hus.fi.
Avela Kristiina, Email: kristiina.avela@vaestoliitto.fi.
Scheinin Ilari, Email: ilari.scheinin@helsinki.fi.
Kristiansson Kati, Email: kati.kristiansson@thl.fi.
Lahermo Päivi, Email: paivi-lahermo@helsinki.fi.
Hietala Marja, Email: marja.hietala@tyks.fi.
Metsähonkala Liisa, Email: liisa.metsahonkala@hus.fi.
Kuusinen Esa, Email: esa.kuusinen@satshp.fi.
Laaksonen Maarit, Email: maarit.laaksonen@thl.fi.
Saarela Janna, Email: janna.saarela@helsinki.fi.
Knuutila Sakari, Email: sakari.knuutila@helsinki.fi.
Acknowledgements
We thank all the participating patients and their families. A thanks to Marketta Dalla Valle for her contribution of clinical information. The staff from the Genotyping Facilities at the Wellcome Trust Sanger Institute is acknowledged for genotyping of the Health 2000 study sample. This study was kindly supported by Rinnekoti Research Foundation, Medicinska Understödsföreningen Liv och Hälsa rf., and The Helsinki and Uusimaa Hospital District State Appropriations. K.K. was supported by the Orion-Farmos Research Foundation and the Academy of Finland (grant no. 125973).
References
- Miller DT, Adam MP, Aradhya S, Biesecker LG, Brothman AR, Carter NP, Church DM, Crolla JA, Eichler EE, Epstein CJ, Faucett WA, Feuk L, Friedman JM, Hamosh A, Jackson L, Kaminsky EB, Kok K, Krantz ID, Kuhn RM, Lee C, Ostell JM, Rosenberg C, Scherer SW, Spinner NB, Stavropoulos DJ, Tepperberg JH, Thorland EC, Vermeesch JR, Waggoner DJ, Watson MS, Martin LC, Ledbetter DH. Consensus statement: chromosomal microarray is a first-tier clinical diagnostic test for individuals with developmental disabilities or congenital anomalies. Am J Hum Genet. 2010;86:749–764. doi: 10.1016/j.ajhg.2010.04.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Siggberg L, Ala-Mello S, Jaakkola E, Kuusinen E, Schuit R, Kohlhase J, Bohm D, Ihnatius J, Knuutila S. Array CGH in molecular diagnosis of mental retardation - A study of 150 Finnish patients. Am J Med Genet A. 2010;152A:1398–1410. doi: 10.1002/ajmg.a.33402. [DOI] [PubMed] [Google Scholar]
- Robinson WP. Mechanisms leading to uniparental disomy and their clinical consequences. BioEssays. 2000;22:52–459. doi: 10.1002/(SICI)1521-1878(200005)22:5<452::AID-BIES7>3.0.CO;2-K. [DOI] [PubMed] [Google Scholar]
- Liehr T. Cytogenetic contribution to uniparental disomy (UPD) Mol Cytogenet. 2010;3:8. doi: 10.1186/1755-8166-3-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kotzot D. Complex and segmental uniparental disomy updated. J Med Genet. 2008;45:545–556. doi: 10.1136/jmg.2008.058016. [DOI] [PubMed] [Google Scholar]
- Health. THL - National Institute for Health and Welfare. 2000. Accessed 30.8.2010, 2010, at http://www.terveys2000.fi/indexe.html.
- Wang K, Li M, Hadley D, Liu R, Glessner J, Grant SF, Hakonarson H, Bucan M. PennCNV: An integrated hidden Markov model designed for high-resolution copy number variation detection in whole-genome SNP genotyping data. Genome Res. 2007;17:1665–1674. doi: 10.1101/gr.6861907. [DOI] [PMC free article] [PubMed] [Google Scholar]
- International HapMap Consortium. A haplotype map of the human genome. Nature. 2005;437:1299–1320. doi: 10.1038/nature04226. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Iafrate AJ, Feuk L, Rivera MN, Listewnik ML, Donahoe PK, Qi Y, Sherer SW, Lee C. Detection of large-scale variation in the human genome. Nat Genet. 2004;36:949–951. doi: 10.1038/ng1416. [DOI] [PubMed] [Google Scholar]
- Kristiansson K, Naukkarinen J, Peltonen L. Isolated populations and complex disease gene identification. Genome Biol. 2008;9:109. doi: 10.1186/gb-2008-9-8-109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Conlin LK, Thiel BD, Bonnemann CG, Medne L, Ernst LM, Zackai EH, Deardorff MA, Krantz ID, Hakonarson H, Spinner NB. Mechanisms of mosaicism, chimerism and uniparental disomy identified by single nucleotide polymorphism array analysis. Hum Mol Genet. 2010;19:1263–1275. doi: 10.1093/hmg/ddq003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Engel E. A fascination with chromosome rescue in uniparental disomy: Mendelian recessive outlaws and imprinting copyrights infringements. Eur J Hum Genet. 2006;14:1158–1169. doi: 10.1038/sj.ejhg.5201619. [DOI] [PubMed] [Google Scholar]
- Bernardini L, Alesi V, Loddo S, Novelli A, Bottillo I, Battaglia A, Digilio MC, Zampino G, Ertel A, Fortina P, Surrey S, Dallapiccola B. High-resolution SNP arrays in mental retardation diagnostics: how much do we gain? Eur J Hum Genet. 2010;18:178–185. doi: 10.1038/ejhg.2009.154. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mannik K, Parkel S, Palta P, Zilina O, Puusepp H, Esko T, Magi R, Noukas M, Veidenberg A, Nelis M, Metspalu A, Remm M, Ounap K, Kurg A. A parallel SNP array study of genomic aberrations associated with mental retardation in patients and general population in Estonia. Eur J Med Genet. 2011;54:136–143. doi: 10.1016/j.ejmg.2010.11.005. [DOI] [PubMed] [Google Scholar]
- Bruno DL, Ganesamoorthy D, Schoumans J, Bankier A, Coman D, Delatycki M, Gardner RJ, Hunter M, James PA, Kannu P, McGilivray G, Pachter N, Peters H, Rieubland C, Savarirayan R, Scheffer IE, Sheffield L, Tan T, White SM, Yeung A, Bowman Z, Ngo C, Choy KW, Cacheux V, Wong L, Amor DJ, Slater HR. Detection of cryptic pathogenic copy number variations and constitutional loss of heterozygosity using high resolution SNP microarray analysis in 117 patients referred for cytogenetic analysis and impact on clinical practice. J Med Genet. 2009;6:123–131. doi: 10.1136/jmg.2008.062604. [DOI] [PubMed] [Google Scholar]
- McMullan DJ, Bonin M, Hehir-Kwa JY, de Vries BB, Dufke A, Rattenberry E, Steehouwer M, Moruz L, Pfundt R, de Leeuw N, Riess A, Altug-Teber O, Enders H, Singer S, Grasshof U, Walter M, Walker JM, Lamb CV, Davison EV, Bruetin L, Riess O, Veltman JA. Molecular karyotyping of patients with unexplained mental retardation by SNP arrays: a multicenter study. Hum Mutat. 2009;30:1082–1092. doi: 10.1002/humu.21015. [DOI] [PubMed] [Google Scholar]
- Friedman J, Adam S, Arbour L, Armstrong L, Baross A, Birch P, Boerkoel C, Chan S, Chai D, Delaney AD, Flibotte S, Gibson WT, Langlois S, Lemyre E, Li HI, MacLeod P, Mathers J, Michaud JL, McGillivray BC, Patel MS, Qian H, Rouleau GA, Van Allen MI, Yong SL, Zahir FR, Eydoux P, Marra MA. Detection of pathogenic copy number variants in children with idiopathic intellectual disability using 500 K SNP array genomic hybridization. BMC Genomics. 2009;10:526. doi: 10.1186/1471-2164-10-526. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Behnecke A, Hinderhofer K, Bartsch O, Nümann A, Ipach ML, Damatova N, Haaf T, Dufke A, Riess O, Moog U. Intragenic deletions of IL1RAPL1: Report of two cases and review of the literature. Am J Med Genet A. 2011;155A:372–379. doi: 10.1002/ajmg.a.33656. [DOI] [PubMed] [Google Scholar]
- Vissers LE, de Ligt J, Gilissen C, Janssen I, Steehouwer M, de Vries P, van Lier B, Arts P, Wieskamp N, del Rosario M, van Bon BW, Hoischen A, de Vries BB, Bruner HG, Veltman JA. A de novo paradigm for mental retardation. Nat Genet. 2010;42:1109–1112. doi: 10.1038/ng.712. [DOI] [PubMed] [Google Scholar]