Abstract
Tyrosine hydroxylase is the rate-limiting step in the synthesis of dopamine and is tightly regulated. Previous studies have shown it to be covalently modified and potently inhibited by 3,4-dihydroxyphenylacetaldehyde (DOPAL), an endogenous neurotoxin via dopamine catabolism which is relevant to Parkinson’s disease. In order to elucidate the mechanism of enzyme inhibition, a source of pure, active tyrosine hydroxylase was necessary. The cloning and novel purification of human recombinant TH from Escherichia coli is described here. This procedure led to the recovery of ~23 mg of pure, active and stable enzyme exhibiting a specific activity of ~17 nmol/min/mg. The enzyme produced with this procedure can be used to delineate the tyrosine hydroxylase inhibition by DOPAL and its relationship to Parkinson’s disease. This procedure improves upon previous methods because the fusion protein gives rise to high expression and convenient affinity-capture, and the cleaved and highly purified hTH makes the product useful for a wider variety of applications.
Keywords: Human tyrosine hydroxylase, Escherichia coli, Cloning and purification
Introduction
Tyrosine hydroxylase (TH, EC 1.14.16.2) catalyzes the rate-limiting step in dopamine (DA) synthesis. Tyrosine is oxidized to L-DOPA, which is further metabolized to DA [1,2]. Enzyme activity is tightly regulated via many pathways. Phosphorylation of three key Ser residues can control activation, while high levels of substrate and DA lead to feed-back inhibition of TH [3,4]. Recent studies have demonstrated TH to be modified and inactivated by the monoamine oxidase metabolite of DA, 3,4-dihydroxyphenylacetaldehyde (DOPAL). Inhibition by DOPAL was revealed to have both a reversible and irreversible component, with TH showing partially recovered activity in competition assays [5].
These studies have implications for Parkinson’s disease (PD), due to the important role of TH in the production of L-DOPA and DA. Selective dopaminergic neuron death, and low levels of DA are key hallmarks of PD [6]. Inhibition of TH leads to decreased DA production, and elevated levels of DOPAL have been shown to cause cell death [7,8]. In order to complete mechanistic studies of DOPAL inhibition of TH and map adduct sites, a source of pure, active recombinant enzyme was required.
While there are a number of procedures in the literature using both PC-12 cells and Escherichia coli as a source for recombinant TH [4,9–12], these papers use outdated methods and involve arduous, lengthy procedures, excessive culture volumes and/or poor yields. Here, we describe a novel, efficient method for the cloning and purification of stable human recombinant TH with initial expression driven by fusion to bacterial maltose binding protein (MBP). This method will allow for the use of higher resolution techniques (i.e. mass spectrometry and high throughput screening) to study the modification of TH and the role this may play in PD.
Materials and methods
Materials
Lyophilized oligonucleotides were obtained from Integrated DNA Technologies (Coralville, IA) and resuspended in water. The human TH gene was purchased from Harvard PlasmID (Clone ID HsCD00378692) as a glycerol stock of pENTR223-transformed E. coli strain DH5α. DNA purification kits, QIAprep Spin Miniprep (cat# 27106) and QIAquick gel extraction (cat# 28704), and Ni–NTA resin (cat# 30230) were obtained from Qiagen (Valencia, CA). Restriction enzymes SalI (cat# R0138S) and BamHI (cat# R0136S), T4 ligase (cat# M0202S), amylose resin (cat# E8021L) were obtained from New England Biolabs (Ipswich, MA). DNaseI (cat# 10104159001) and E-64 (cat# 11585681001) were purchased Roche Applied Science (Indianapolis, IN). Phenylmethylsulfonyl fluoride (PMSF, cat# 28131) and bovine serum albumin fraction V (cat# 9048-46-8) were obtained from Research Products International (Mt. Prospect, IL). Hen egg white lysozyme (cat# 89833) and pepstatin a (cat# bp267125) were purchased from Thermo Scientific (Waltham, MA). Leupeptin (cat# 108975) was obtained from EMD Chemicals (Gibbstown, NJ). HiPrep Q FF 16/10 column (cat# 28-9365-43) and AKTApurifier fast protein liquid chromatography (FPLC) system were purchased from GE Healthcare (Piscataway, NJ). Sequencing was performed by the University of Iowa DNA Facility.
Molecular biology
TH-pENTR223-containing DH5α stock was streaked onto a plate of LB agar supplemented with 100 μg/mL spectinomycin and grown for 16 h at 37 °C. A single colony was selected and expanded into 8 mL LB broth supplemented with 100 μg/mL spectinomycin, and grown for 16 h at 37 °C. Bacteria were pelleted at 3600g for 10 min at 4 °C. pENTR223 plasmid was purified according to manufacturer’s instructions (Qiagen).
The open reading frame containing full-length human TH (EC 1.14.16.2, amino acids MPT…AIG) was amplified with primers containing a 5′ BamHI recognition site (5′-CGGATCCATGCCCACCCC CGACGCCACCAC-3′) and an in-frame 3′engineered stop codon followed by a SalI recognition site (5′-CCCGTCGACCTAGCCAATGGCACTCAGCGCA- 3′) using Platinum Pfx according to manufacturer’s instructions (Life Technologies, Carlsbad, CA). This fragment and pMALc2H10T vector [13] were gel-purified per manufacturer’s protocol (Qiagen), double-digested with BamHI and SalI according to manufacturer instructions (New England Biolabs). Digested TH and pMALc2H10T were gel-purified again (Qiagen) and ligated using T4 DNA ligase according to manufacturer’s instructions (New England Biolabs) using a 3:1 insert:vector ratio and incubation overnight at 16 °C. The ligated product was transformed into XL10-Gold bacteria according to manufacturer’s protocol (Stratagene, La Jolla, CA).
Following transformation, colonies were picked, miniprepped (Qiagen), and their DNA sequenced to ensure in-frame insertion of TH with forward and reverse with primers: pMAL5′sense GCGGTCGTCAGACTGTCGATGAAG, TH middle antisense GCGC TCCAGCAAAGCAAAGGCC, TH middle sense CCCAGTATATCCGCCACGCG, and pMAL3′antisense GGCGATTAAGTTGGGTAACGCC AGG.
Expression
pMALc2H10T–TH vector was transformed into BL21(DE3) E. coli, and a single colony was picked and grown in 10 mL in LB supplemented with 100 μg/mL ampicillin and shaken for 16 h at 37 °C. The culture was pelleted at 3600g for 10 min at 4 °C and resuspended in 500 μL LB to which was added 500 μL 80% (v/v) sterile glycerol and stored at −20 °C. To express MBP-TH, 5 μL of glycerol stock was expanded to a starter culture of 10 mL LB+amp for 8 h shaken at 37 °C, which was expanded to a seed culture of 200 mL LB+amp overnight shaken at 37 °C. On the morning of induction, the seed culture was expanded to 4 × 1 L LB+amp, grown to OD600 0.4 at 37 °C, cooled to 25 °C by shaking the flasks in a tub of ice, and grown to OD600 0.6 at 25 °C.
Protein expression induced with 250 μMβ-D-1-thiogalactopyranoside (IPTG) added as a 1000x stock freshly prepared in water. Simultaneously with IPTG addition, the medium was supplemented with 100 μMFeSO4 and 0.2% (w/v) glucose for enzyme stability and to prevent the production of bacterial amylases, respectively. Induction proceeded for 20 h at 25 °C, after which bacteria was pelleted at 3600g at 4 °C yielding 17 g wet weight of cells and resuspended in 30 mL TH buffer A at 4 °C (50 mM HEPES pH 6.8 100 mM NaCl 5 mM β-mercaptoethanol) supplemented with protease inhibitors (100 μg/mL PMSF, 1 μM Leupeptin, 1 μg/mL Pepstatin, and 1 μME-64) added to buffer immediately before use. The suspended pellet was snap frozen in liquid nitrogen and stored at −80 °C until purification.
Purification
The pellet was thawed at 37 °C, stirred on ice with 0.8 mg lysozyme per mL of pellet for 10 min, snap frozen, and thawed again. Other protocols may yield superior lysis and can be substituted at the user’s discretion. Following lysis, 10 μg/mL DNaseI was stirred into viscous mixture for 10 min on ice, and the lysate was centrifuged for 1 h at 36500 rpm (~100,000g) in a Sorvall T-647.5 rotor. The supernatant was filtered (0.45 μm) and loaded onto a 10 mL amylose resin column (New England Biolabs) equilibrated with buffer A. The resin was washed with 50 mL buffer A and eluted with 70 mL (5 mL fractions) buffer A containing 10 mM maltose. Following sodium dodecylsulfate polyacrylamide gel electrophoresis (SDS–PAGE) analysis, MBP–TH fusion protein-containing fractions were pooled and digested with TEV protease (10:1 M ratio TH:TEV) for 8 h at 4 °C on a rotating mixer. Digestion was monitored by SDS–PAGE to ensure complete cleavage, after which the sample was diluted 5-fold in TH buffer Q (50 mM HEPES, pH 6.8, 5 mM βME) and loaded onto a HiPrep Q FF 16/10 column (C.V. 21.1 mL) equilibrated with TH buffer Q on an AKTApurifier (GE Healthcare). TH buffer Q plus 1 M NaCl was used as eluent and is denoted TH buffer QB. The sample was eluted after a gradient of 0–15% QB over 1 C.V., 5 C.V. at 15% QB, a gradient of 15–30% QB over 5 C.V., and then 30–100% QB over 5 C.V. The first eluting peak, eluting at 13.5 mS/cm, contains pure MBP. The second peak, eluting at 22.5 mS/cm, contains 99%+pure TH by SDS–PAGE.
TEV protease expression and purification
BL21 (DE3) CodonPlus cells, picked from a single colony of a freshly transformed plate, containing pRK1043 (Addgene plasmid 8827) (MBP–TEV [S219 V]–Arg5) were grown in 6 L LB with ampicillin (100 μg/ml) and chloramphenicol (30 μg/ml) to OD600 0.55 at 37 °C. Expression was induced with 1 mM IPTG at 30 °C for 4 h. Cells were harvested by centrifugation. The ~25 g of wet cell paste was resuspended in 30 mL of TEV buffer A (50 mM NaPO4, pH 7.5, 500 mM NaCl, 5 mM βME) with protease inhibitors (see TH purification). Lysis procedure was performed as for TH except centrifuging for 30 min at 22500 rpm. Supernatant was filtered (0.45 μm), and loaded onto a 8 mL Ni–NTA (New England Biolabs) column equilibrated with 80 mL TEV buffer A. The column was washed with 120 mL buffer A with 25 mM imidazole and sample eluted in 8 fractions of 4 mL each. Fractions were analyzed by SDS–PAGE, and those containing TEV (~26 kDa) were pooled and dialyzed against 5 L TEV buffer B (25 mM TRIS, pH 8, 100 mM NaCl, 1 mM EDTA, 2 mM DTT) overnight. The cloudy suspension was centrifuged at 10000g at 4 °C in a fixed angle tabletop centrifuge for 1 min, and the supernatant loaded onto a HiPrep Q FF 16/10 column in an AKTApurifier FPLC (GE Healthcare) equilibrated with TEV buffer B. Following loading, TEV was eluted with a linear gradient from 0–100% TEV buffer B plus 1 mM NaCl over 12 C.V.; TEV elutes at approximately 44% at 99+% purity as determined by SDS– PAGE. The fractions were pooled, stirred at 4 °C adding glycerol dropwise until 10% (v/v), snap frozen as 126 1 mg aliquots, and stored at −80 °C until needed.
TH activity assay
Recombinant TH activity was determined as previously described [5] with some modifications. Briefly, 10 μg of recombinant protein was incubated with 50 μM tyrosine, 0.25 mM tetrahydrobiopterin, and 2.5 μM iron (II) sulfate in 10 mM sodium phosphate buffer (pH 6.8) at 37 °C for 20 min. At 5 min time points, aliquots were taken and placed in 5% (v/v) perchloric acid to stop the reaction. L-DOPA production was monitored using an Agilent 1200 Series Capillary HPLC system with a photodiode array detector set to absorbance 202 and 280 nm. Fifteen microliters of sample was injected and separated using a Phenomenex Luna C18 column. The mobile phase consisted of 97% 0.1% trifluoroacetic acid (v/v) in HPLC-grade water (A), and 3% acetonitrile (B) using an isocratic flow. L-DOPA had a retention time of approximately 7 min and peak area was converted into concentration using a standard curve.
Statistical analysis and Bradford assay
All linear regression and statistical analysis were performed using the software GraphPad Prism 5.0 (Graph Pad Software, San Diego, CA). Data for TH activity and L-DOPA formation was determined using linear regression. To determine protein concentration after each step in the purification, the microtiter plate Bradford assay was used with bovine serum albumin as the standard [14].
Results
TH purification
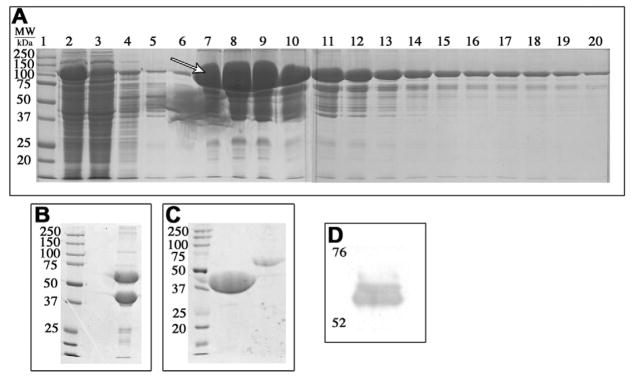
We used a two-step purification beginning with an amylose resin column to capture MBP–TH fusion proteins (see Fig. 1 for construct) from crude cell lysate (Fig. 2A). Following cleavage of the fusion protein with TEV protease (Fig. 2B), an anion exchange column was used to purify the TH from MBP (Fig. 2C). Fig. 2A depicts a typical gel acquired after use of an amylose column. The major band indicated by the arrows between 110–120 kDaMW is indicative of the MBP–TH construct. This binds readily with the amylose, and is then released in the presence of a competing concentration of maltose. Fractions in lanes 7–19 were collected and subjected to TEV protease digestion in order to cleave TH from the MBP. 2B is a representative gel after cleavage for ~8 h at 4 °C. The band at 56 kDa is TH, while the band at 44 kDa is the maltose-binding protein. Immediately following confirmation of cleavage, the sample was applied to a HiPrep Q FF 16/10 anionic exchange column and the 2 proteins were separated using a salt gradient. Fractions of each peak were collected and analyzed by SDS–PAGE. Fig. 2C shows SDS–PAGE analysis of the 2 peaks eluted, MBP is in lane 2, and TH is in lane 3. Examination of the 2nd peak (Lane 3 in Fig. 2C) via western blot using antibodies specific to TH, confirmed the identity of the protein as TH (Fig. 2D). While it may appear to be a subtle doublet in 2D (a ghost or blowout band) due to overloading of the lane [15], the 56 kDa band represents the pure TH enzyme protein isolation.
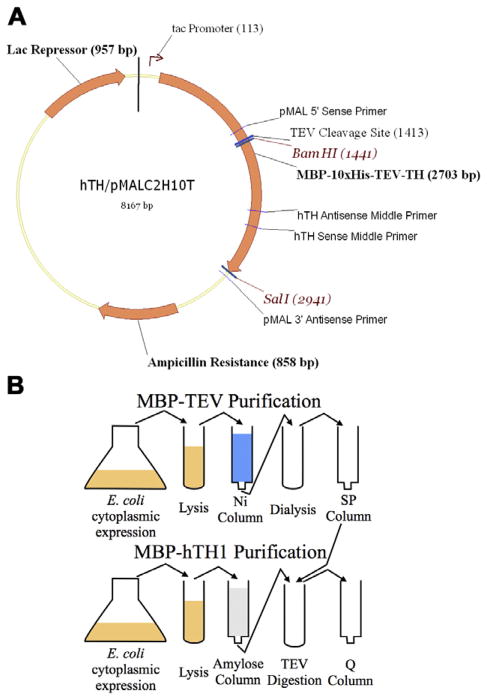
Fig. 1.
(A) hTH/pMALc2H10T construct used to express recombinant tyrosine hydroxylase. The TH insert size is 1500 bp, and the MBP–TH fusion protein coding sequence is 2703 bp. The BamHI and SalI restriction sites used to clone TH are unique in the vector and insert. (B) The scheme used to purify hTH consisting of two chromatography steps and an enzymatic digestion with TEV protease (purification also provided).
Fig. 2.
Typical gels representing the purification procedure. (A) After the crude sample is run through an amylose resin column and the MBP-bound TH is eluted off using maltose. Lane (1) MW marker, (2) load (crude lysate), (3) amylose column unbound sample flow-through, (4) wash, 5–20: eluted fractions. All fractions were analyzed by SDS–PAGE, and fractions containing the MBP–TH construct were further purified. (B) TH and MBP after cleavage by TEV for 8 h at 4C; TH and MBP are cleaved (56 and 44 kDa, respectively). (C) Post-HiPrep Q FF 16/10 column using a salt gradient for elution. MBP elutes first, followed by TH. (D) Western blot analysis for TH after the final purification step.
Enzyme characterization
TH activity was measured as previously described by Mexas, et al., using an HPLC coupled with a photodiode array detector (5). Data in Fig. 3 demonstrates TH exhibits time-dependent conversion of tyrosine to L-DOPA over a 20 min time course. When TH was incubated with 50 μM tyrosine, ~25 μM L-DOPA was formed. Table 1 summarizes the specific activities of each step in the purification as determined by an average of two 2 and one 4 L preparations. Tyrosine hydroxylase activity and purity (as assessed by SDS–PAGE and immunoblot) increases, indicating the purification process does not interfere or negatively affect the enzyme.
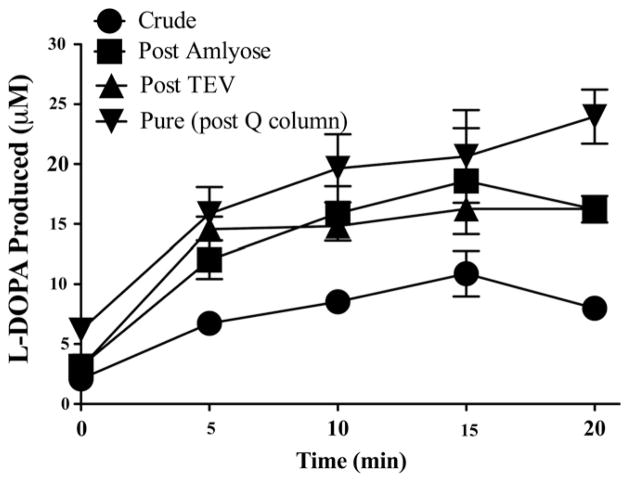
Fig. 3.
Activity of tyrosine hydroxylase at each step in the purification, as determined by L-DOPA production. ten micrograms of pure TH was incubated with tyrosine, iron, and cofactor. HPLC analysis determined the production of L-DOPA over a 20 min time course.
Table 1.
Tabular results of purification of a 2 L preparation of TH (values average of n = 3 independent cultures). Overall yield of purified, cleaved TH is 41% of total soluble cellular protein collecteda.
| Purification step | Total protein(mg) | Specific activity (nmol/min/mg) | Total enzyme units (nmol/min) | % Yieldb | Approximate purity (%) |
|---|---|---|---|---|---|
| Crude | 89.4 | 5.40 | 483 | 100 | 33 |
| Amylose | 42.6 | 12.2 | 520 | 108 | 75 |
| TEV | 26.0 | 12.2 | 317 | 66 | 75 |
| Q-column | 12.0 | 16.3 | 196 | 41 (73)c | 100 |
Wet weight cells 17 g.
Overall yield reported as fraction of TH at each purification step compared to the amount of TH in soluble crude cell lysate protein.
Parenthetical percent yield incorporates the mass difference of cleaved TH (56 kDa) versus the MBP–TH fusion protein (100 kDa).
Discussion
We have successfully cloned and purified the enzyme tyrosine hydroxylase in this fast and high-yielding protocol. The full-length cDNA of human cytosolic TH was cloned in frame downstream of E. coli MBP, which in addition to serving as an affinity tag for the first step of purification, enhances the solubility of passenger proteins [16], and was separated from MBP by the insertion of a TEV protease recognition site between MBP and TH [17]. Nakashima and coworkers attempted purification of hTH from an MBP fusion protein, but the cleavage proteins (factor Xa and enterokinase) used in that study cleaved non-specifically within TH due to the poor specificity of factor Xa and enterokinase [18]. Advantages to this study include completion of the purification process in one day, a high yield, a stable and 99%+pure enzyme, and highly active TH. Previous yields published indicate recovery of pure human TH can vary greatly, with some producing as low as 0.65 mg per liter of culture medium [9,11,12,19]. Our study recovered an average of 6 mg of pure TH per liter of culture medium, indicating a high-yield procedure as compared to other purification procedures. Characterization of TH included measuring enzyme activity during all stages of the purification process (crude, post-amylose, post-TEV, and pure), in order to ensure minimal loss of enzyme activity. TH exhibited a high turn-over of tyrosine using the HPLC method previously established [5]. Data in Fig. 3 demonstrates that compared to controls (i.e., pure enzyme), the preceding steps in the purification are significantly less active, indicating there is an increase in tyrosine turnover as the enzyme is purified.
The pure TH recovered in this study had a specific activity of ~16 nmol/min/mg protein. Previous literature values have reported specific activities ranging from 1 to 2700 nmol/min/mg for both rat and human recombinant TH [3,4,19,20]. These results indicate the activity of TH is highly variable, depending on the method of purification, and our results fall into this range. One confounding factor in comparing TH activity in this study to others is that the activity assay used in many other studies is the tritium release method, which detects tritiated water that is released in the conversion of tyrosine to L-DOPA [21]. In the case of this study, the formation of L-DOPA was monitored using an HPLC with photodiode array detection. Furthermore, the concentration of tyrosine and iron sulfate used in other studies varies greatly; reported values are from 20 to 200 μM [3,4,10,11] for tyrosine, and 0 to 13 μM [9,11,12,22] for iron sulfate. Our studies utilized 50 μM tyrosine and 2.5 μM iron sulfate and control experiments with several concentrations of tyrosine and iron sulfate (50–200 μM and 2.5–20 μM, respectively) yielded a similar specific activity to that that is described above (data not shown). These results indicate the concentration of tyrosine and iron sulfate used are acceptable for enzyme activity.
Overall, there are a large number of techniques and materials utilized in the purification of recombinant TH and the determination of specific activity, and the results presented here are well within the ranges previously reported. Furthermore, the enzyme obtained is pure tyrosine hydroxylase (as confirmed by SDS–PAGE and Western blot analysis), and highly functional, oxidizing tyrosine to L-DOPA. This novel cloning and purification method is able to rapidly and easily produce pure, stable and active human recombinant tyrosine hydroxylase, which can be used to further study the modification and inhibition of TH, and how this may play a role in PD.
Acknowledgments
We would like to thank the ORFeome Collaboration (Center for Cancer Systems Biology) for the production of the human tyrosine hydroxylase plasmid, and the Harvard Medical School PlasmID Database for providing the plasmid upon purchase. We would also like to thank Dr. John Tesmer (University of Michigan) for kindly providing the pMALc2H10T vector for our use and David Waugh (NCI, Macromolecular Crystallography Laboratory) via Addgene.org for the TEV construct used in this study. The project described was supported by ES15507 (J.A.D.) and T32GM067795 (C.A.H.) from NIH. Its contents are solely the responsibility of the authors and do not necessarily represent the official views of the NIH.
Contributor Information
Colin A. Higgins, Email: colin-higgins@uiowa.edu.
Lydia M. Vermeer, Email: lydia-mexas@uiowa.edu.
Jonathan A. Doorn, Email: jonathan-doorn@uiowa.edu.
References
- 1.Elsworth JD, Roth RH. Dopamine synthesis, uptake, metabolism, and receptors: relevance to gene therapy of Parkinson’s disease. Exp Neurol. 1997;144:4–9. doi: 10.1006/exnr.1996.6379. [DOI] [PubMed] [Google Scholar]
- 2.Nagatsu T, Levitt M, Udenfriend S. Tyrosine hydroxylase. The Initial step in norepinephrine biosynthesis. J Biol Chem. 1964;239:2910–2917. [PubMed] [Google Scholar]
- 3.Daubner SC, Lauriano C, Haycock JW, Fitzpatrick PF. Site-directed mutagenesis of serine 40 of rat tyrosine hydroxylase. Effects of dopamine and cAMP-dependent phosphorylation on enzyme activity. J Biol Chem. 1992;267:12639–12646. [PubMed] [Google Scholar]
- 4.Xu Y, Stokes AH, Roskoski R, Jr, Vrana KE. Dopamine, in the presence of tyrosinase, covalently modifies and inactivates tyrosine hydroxylase. J Neurosci Res. 1998;54:691–697. doi: 10.1002/(SICI)1097-4547(19981201)54:5<691::AID-JNR14>3.0.CO;2-F. [DOI] [PubMed] [Google Scholar]
- 5.Mexas LM, Florang VR, Doorn JA. Inhibition and covalent modification of tyrosine hydroxylase by 3,4-dihydroxyphenylacetaldehyde, a toxic dopamine metabolite. Neurotoxicology. 2011;40:471–477. doi: 10.1016/j.neuro.2011.03.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Andersen JK. Oxidative stress in neurodegeneration: cause or consequence? Nat Med. 2004;10(Suppl):S18–25. doi: 10.1038/nrn1434. [DOI] [PubMed] [Google Scholar]
- 7.Burke WJ, Li SW, Williams EA, Nonneman R, Zahm DS. 3,4- Dihydroxyphenylacetaldehyde is the toxic dopamine metabolite in vivo: implications for Parkinson’s disease pathogenesis. Brain Res. 2003;989:205–213. doi: 10.1016/s0006-8993(03)03354-7. [DOI] [PubMed] [Google Scholar]
- 8.Burke WJ. 3,4-dihydroxyphenylacetaldehyde: a potential target for neuroprotective therapy in Parkinson’s disease. Curr Drug Targets CNS Neurol Disord. 2003;2:143–148. doi: 10.2174/1568007033482913. [DOI] [PubMed] [Google Scholar]
- 9.Le Bourdelles B, Horellou P, Le Caer JP, Denefle P, Latta M, Haavik J, Guibert B, Mayaux JF, Mallet J. Phosphorylation of human recombinant tyrosine hydroxylase isoforms 1 and 2: an additional phosphorylated residue in isoform 2, generated through alternative splicing. J Biol Chem. 1991;266:17124–17130. [PubMed] [Google Scholar]
- 10.Wang YH, Citron BA, Ribeiro P, Kaufman S. High-level expression of rat PC12 tyrosine hydroxylase cDNA in Escherichia coli: purification and characterization of the cloned enzyme. Proc Natl Acad Sci USA. 1991;88:8779–8783. doi: 10.1073/pnas.88.19.8779. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Gahn LG, Roskoski R., Jr Tyrosine hydroxylase purification from rat PC12 cells. Protein Expres Purif. 1991;2:10–14. doi: 10.1016/1046-5928(91)90002-z. [DOI] [PubMed] [Google Scholar]
- 12.Haavik J, Le Bourdelles B, Martinez A, Flatmark T, Mallet J. Recombinant human tyrosine hydroxylase isozymes. Reconstitution with iron and inhibitory effect of other metal ions. Eur J Biochem. 1991;199:371–378. doi: 10.1111/j.1432-1033.1991.tb16133.x. [DOI] [PubMed] [Google Scholar]
- 13.Kristelly R, Gao G, Tesmer JJ. Structural determinants of RhoA binding and nucleotide exchange in leukemia-associated Rho guanine-nucleotide exchange factor. J Biol Chem. 2004;279:47352–47362. doi: 10.1074/jbc.M406056200. [DOI] [PubMed] [Google Scholar]
- 14.Olson BJ, Markwell J. Assays for determination of protein concentration. Curr Protoc Protein Sci. 2007;Chapter 3(Unit 3):4. doi: 10.1002/0471140864.ps0304s48. [DOI] [PubMed] [Google Scholar]
- 15.Alegria-Schaffer A, Lodge A, Vattem K. Performing and optimizing Western blots with an emphasis on chemiluminescent detection. Methods Enzymol. 2009;463:573–599. doi: 10.1016/S0076-6879(09)63033-0. [DOI] [PubMed] [Google Scholar]
- 16.Kapust RB, Waugh DS. Escherichia coli maltose-binding protein is uncommonly effective at promoting the solubility of polypeptides to which it is fused. Protein Sci. 1999;8:1668–1674. doi: 10.1110/ps.8.8.1668. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Kapust RB, Tozser J, Fox JD, Anderson DE, Cherry S, Copeland TD, Waugh DS. Tobacco etch virus protease: mechanism of autolysis and rational design of stable mutants with wild-type catalytic proficiency. Protein Eng. 2001;14:993–1000. doi: 10.1093/protein/14.12.993. [DOI] [PubMed] [Google Scholar]
- 18.Nakashima A, Mori K, Nagatsu T, Ota A. Expression of human tyrosine hydroxylase type I in Escherichia coli as a protease-cleavable fusion protein. Short communication. J Neural Transm. 1999;106:819–824. doi: 10.1007/s007020050202. [DOI] [PubMed] [Google Scholar]
- 19.Kuhn DM, Billingsley ML. Tyrosine hydroxylase: purification from PC-12 cells, characterization and production of antibodies. Neurochem Int. 1987;11:463–475. doi: 10.1016/0197-0186(87)90036-2. [DOI] [PubMed] [Google Scholar]
- 20.Kojima K, Mogi M, Oka K, Nagatsu T. Purification and immunochemical characterization of human adrenal tyrosine hydroxylase. Neurochem Int. 1984;6:475–480. doi: 10.1016/0197-0186(84)90117-7. [DOI] [PubMed] [Google Scholar]
- 21.Nagatsu T, Levitt M, Udenfriend S. A rapid and simple radioassay for tyrosine hydroxylase activity. Anal Biochem. 1964;9:122–126. doi: 10.1016/0003-2697(64)90092-2. [DOI] [PubMed] [Google Scholar]
- 22.Fitzpatrick PF. Steady-state kinetic mechanism of rat tyrosine hydroxylase. Biochemistry. 1991;30:3658–3662. doi: 10.1021/bi00229a010. [DOI] [PubMed] [Google Scholar]