Abstract
Background
The genus Eucalyptus consists of approximately 600 species and subspecies and has a physiological plasticity that allows some species to propagate in different regions of the world. Eucalyptus is a major source of cellulose for paper manufacturing, and its cultivation is limited by weather conditions, particularly water stress and low temperatures. Gene expression studies using quantitative reverse transcription polymerase chain reaction (qPCR) require reference genes, which must have stable expression to facilitate the comparison of the results from analyses using different species, tissues, and treatments. Such studies have been limited in eucalyptus.
Results
Eucalyptus globulus Labill, Eucalyptus urograndis (hybrid from Eucalyptus urophylla S.T. Blake X Eucalyptus grandis Hill ex-Maiden) and E. uroglobulus (hybrid from E. urograndis X E. globulus) were subjected to different treatments, including water deficiency and stress recovery, low temperatures, presence or absence of light, and their respective controls. Except for treatment with light, which examined the seedling hypocotyl or apical portion of the stem, the expression analyses were conducted in the apical and basal parts of the stem. To select the best pair of genes, the bioinformatics tools GeNorm and NormFinder were compared. Comprehensive analyses that did not differentiate between species, treatments, or tissue types, showed that IDH (isocitrate dehydrogenase), SAND (SAND protein), ACT (actin), and A-Tub (α-tubulin) genes were the most stable. IDH was the most stable gene in all of the treatments.
Conclusion
Comparing these results with those of other studies on eucalyptus, we concluded that five genes are stable in different species and experimental conditions: IDH, SAND, ACT, A-Tub, and UBQ (ubiquitin). It is usually recommended a minimum of two reference genes is expression analysis; therefore, we propose that IDH and two others genes among the five identified genes in this study should be used as reference genes for a wide range of conditions in eucalyptus.
Keywords: Eucalyptus, Drought, Cold, Light, Reference genes
Background
The genus Eucalyptus belongs to the Myrtaceae family, which consists of approximately 600 species and subspecies, exhibits physiological plasticity with worldwide dispersion and successfully grows in different regions. Eucalyptus is a major source of cellulose for manufacturing paper [1] and globally, Brazil ranks first in the production of short fibre cellulose [2]. Due to its economic importance, the genome sequencing of the species Eucalyptus grandis was recently completed [3].
E. globulus and E. grandis are the two main species cultivated in the world [4]. Among the existing species, E. globulus is known for its resistance to cold temperatures and its lower content and type (greater amount of S units) of lignin, which results in a higher extraction yield of cellulose [5-7]. E. globulus also exhibits tolerance to cold weather, which allows it to be cultivated in areas where other species do not grow well. In Brazil, the hybrid E. urograndis (hybrid of E. urophylla x E. grandis) has been widely cultivated for its unique features, compared with other cultivated species [8,9]. It has been reported that more than 600,000 acres are cultivated with this hybrid species in Brazil, being the basis of the Brazilian silviculture for the production of cellulose and paper [10]. E. uroglobulus is a hybrid derived from E. urograndis x E. globulus, with a high cellulose yield [11].
One of the most widely used approaches for molecular characterisation is gene expression analysis, which aims to identify genes that can be used as molecular markers or for genetic manipulations. The most common technique for analysing gene expression is quantitative reverse transcription polymerase chain reaction (qPCR), which is advantageous because of its speed, sensitivity and specificity [12].
One of the most important aspects in gene expression analysis by qPCR is the determination of genes that have constitutive expression, which will serve as references in the analysis of plant material from different origins. Thus, the expression of these genes must be constant, regardless of the tissue under analysis comes from different parts of the same plant or plants from different origins. Such genes are called normalisers because it is possible to compare gene expression in various types of biological materials [13-15] based on the individual expression of these genes. However, genes that have previously been described as adequate normalisers for some plant species and/or experimental conditions are not always valid for other species or working conditions [16]. Typically, the selected endogenous genes are related to cellular maintenance processes that are common among different cell types [17]. The use of the appropriate normaliser genes is a limitation to the use of qPCR, and this justifies the considerable number of published studies aimed at identifying useful endogenous genes for the study of gene expression in different species, including rice [18], sugarcane [19], A. thaliana[20], potato [21], Brachypodium spp. [22], tomato [23], coffee [24], wheat [25], tobacco [26] and Eucalyptus spp. [27-30].
de Oliveira [27] performed an extensive search of 21,432 eucalyptus genes in E. grandis leaves and xylem and E. globulus xylem. The genes were evaluated using microarray hybridisation analysis, and 50 candidate genes were identified that showed little variation in their expression. These genes were evaluated in gene expression studies associated with the flower, leaf, and xylem of six species of eucalyptus. de Almeida et al. [30] aimed to identify endogenous genes for gene expression studies associated with adventitious rooting in E. globulus microcuttings because low rooting efficiency is a problem for the cloning and multiplication of this species. Fernández et al. [28] attempted to identify normaliser genes in E. globulus for gene expression studies related to cold acclimation. Similarly, Boava et al. [29] sought to identify normaliser genes for studies of Urograndis hybrid resistance to leaf rust caused by Puccinia psidii.
None of these studies identified normaliser genes in eucalyptus that could be widely used in gene expression studies related to cold and drought stress in species of significant economic importance, such as E. urograndis, E. uroglobulus, and E. globulus. Low temperature and limited water are stresses that limit the growth of eucalyptus [11,31-35]. In this study, we evaluated normaliser genes in E. urograndis, E. uroglobulus, and E. globulus plants exposed to low temperature and stressed with water shortage. The recovery of the water stressed plants after re-watering was also evaluated. Additionally, we also evaluated normaliser genes in the seedlings and plants of E. globulus grown in the presence and absence of light, which have been used in lignification studies in this species (Araújo and Mazzafera, unpublished). The validation of endogenous genes under different growth conditions could enable their use in a broad range of studies of gene expression in eucalyptus. In our evaluations we compared the bioinformatics programs NormFinder and GeNorm.
Results
Several comparisons were performed combining species, treatments, tissues and all data using both GeNorm and NormFinder. The results of these analyses are shown in Table 1, which was split into four sections to improve the presentation of the data.
Table 1.
Identification of the best pairs of genes for different treatments, species, and tissues
|
A | |||||||
|---|---|---|---|---|---|---|---|
| Species | Software | CTRL | DRT | REC | CTRL+DRT+REC Apex | CTRL+DRT+REC Base | All treatments |
|
E. globulus |
GeNorm |
A-Tub/UBQ |
A-Tub/H2B |
IDH/SAND |
UBQ/H2B |
H2B/A-Tub |
H2B/UBQ |
| NormFinder |
IDH/ACT |
ACT/A-Tub |
A-Tub/IDH |
IDH/A-Tub |
A-Tub/ACT |
A-Tub/ACT |
|
|
E. urograndis |
GeNorm |
SAND/A-Tub |
H2B/UBQ |
A-Tub/SAND |
UBQ/A-Tub |
IDH/ACT |
IDH/ACT |
| NormFinder |
ACT/SAND |
UBQ/H2B |
SAND/A-Tub |
ACT/SAND |
A-Tub/SAND |
ACT/SAND |
|
|
E. uroglobulus |
GeNorm |
H2B/UBQ |
EF1/IDH |
IDH/ACT |
SAND/IDH |
IDH/18S |
A-Tub/ACT |
| NormFinder |
A-Tub/EF1 |
SAND/ACT |
18S/H2B |
H2B/A-Tub |
A-Tub/ACT |
A-Tub/H2B |
|
|
All species |
GeNorm |
UBQ/H2B |
IDH/SAND |
IDH/ACT |
IDH/ACT |
IDH/ACT |
IDH/ACT |
| |
NormFinder |
IDH/SAND |
ACT/IDH |
SAND/IDH |
SAND/ACT |
SAND/IDH |
SAND/IDH |
|
B | |||||||
|
Species |
Software |
CTRL |
COLD |
CTRL+COLD Apex |
CTRL+COLD Base |
All treatments |
|
|
E. globulus |
GeNorm |
A-Tub/UBQ |
EF1/IDH |
EF1/IDH |
SAND/EF1 |
EF1/IDH |
|
| NormFinder |
ACT/UBQ |
SAND/UBQ |
ACT/A-Tub |
ACT/SAND |
ACT/A-Tub |
|
|
|
E. urograndis |
GeNorm |
IDH/UBQ |
A-Tub/UBQ |
UBQ/A-Tub |
UBQ/18S |
A-Tub/UBQ |
|
| NormFinder |
IDH/UBQ |
SAND/UBQ |
SAND/IDH |
ACT/IDH |
IDH/SAND |
|
|
|
E. uroglobulus |
GeNorm |
IDH/ACT |
UBQ/SAND |
SAND/UBQ |
B-tub/SAND |
UBQ/SAND |
|
| NormFinder |
A-Tub/H2B |
ACT/IDH |
H2B/UBQ |
A-Tub/ACT |
H2B/A-Tub |
|
|
|
All species |
GeNorm |
SAND/UBQ |
ACT/SAND |
UBQ/IDH |
IDH/SAND |
IDH/SAND |
|
| |
NormFinder |
A-Tub/EF1 |
IDH/ACT |
H2B/UBQ |
IDH/SAND |
IDH/SAND |
|
|
C | |||||||
|
Species |
Software |
DRT+COLD Apex |
DRT+COLD Base |
All treatments |
|
|
|
|
E. globulus |
GeNorm |
IDH/A-Tub |
IDH/SAND |
IDH/ACT |
|
|
|
| NormFinder |
UBQ/IDH |
UBQ/A-Tub |
UBQ/A-Tub |
|
|
|
|
|
E. urograndis |
GeNorm |
H2B/UBQ |
SAND/A-Tub |
IDH/ACT |
|
|
|
| NormFinder |
ACT/A-Tub |
SAND/A-Tub |
ACT/IDH |
|
|
|
|
|
E. uroglobulus |
GeNorm |
IDH/ACT |
EF1/IDH |
IDH/ACT |
|
|
|
| NormFinder |
H2B/A-Tub |
A-Tub/ACT |
A-Tub/H2B |
|
|
|
|
|
All species |
GeNorm |
ACT/IDH |
IDH/SAND |
IDH/ACT |
|
|
|
| |
NormFinder |
ACT/IDH |
SAND/IDH |
IDH/ACT |
|
|
|
|
D | |||||||
|
Species |
Software |
in vitro (dark and light) |
Greenhouse (dark and night) |
All treatments |
|
|
|
|
E. globulus |
GeNorm |
H2B/IDH |
EF1/ACT |
UBQ/IDH |
|
|
|
| NormFinder | UBQ/ACT | IDH/EF1 | ACT/IDH | ||||
A – Water stress experiment; B – Low-temperature experiment; C – Water stress + low-temperature experiments in tissues (apex and base), D – light experiments. CTRL = control; DRT – water stress; REC – recovery from water stress; COLD – low temperature.
GeNorm
In the water stress trial, the analyses using GeNorm showed a wide range of genes, depending on the combination of treatments, species, and types of tissues analysed (Table 1A). For E. globulus, the most stable pair of genes, regardless of treatment or stem position, was H2B/UBQ. IDH/ACT and A-Tub/ACT were the best pairs for E. urograndis and E. uroglobus, respectively. When analysing the best pair of genes among all of the situations (treatments, species, and types of tissues), the most stable pair was IDH/ACT. Among the nine genes analysed, only five were observed in the combinations using GeNorm: IDH, UBQ, ACT, A-Tub and H2B.
In the low-temperature experiment, a large variation between the gene pairs was observed, depending on the type of treatment, species and tissue combination (Table 1B). In the general analyses within each species IDH, UBQ, A-Tub, SAND and EF1 emerged as the most stable. From the general analysis, disregarding treatments, species, and types of tissues, the best gene pair was IDH/SAND.
Using GeNorm, the comparison of tissue type (Table 1C) without distinction between treatments, tissue types and species showed the gene pair IDH/ACT as the most stable. GeNorm indicated that UBQ/IDH was the best gene pair from the analyses with seedlings and plants of E. globulus with or without exposure to light (Table 1D).
NormFinder
When NormFinder was used, several pairs of genes were identified and like in the GeNorm analysis, they varied depending on the treatment, species, and plant tissue. In the water stress treatments (Table 1A), at least one of the genes identified was consistently different from those suggested using GeNorm. For example, in the evaluation of E. urograndis, the general analysis of the treatments and tissues resulted in the identification of IDH/ACT by GeNorm and ACT/SAND by NormFinder. There were cases that both genes of the pair were different, such as for E. globulus, where H2B/UBQ and A-Tub/ACT were identified by GeNorm and NormFinder, respectively. In the analysis grouping the data from the water stress experiments, which included all species, treatments (control, water stress and recovery) and tissue types (apex and base), the observed most stable gene pair was SAND/IDH. When the low-temperature data were analysed altogether (Table 1B), the gene pair IDH/SAND was identified as the best combination for NormFinder. Among the nine genes analysed, five genes were present among all of the possible pairs during the water stress and low-temperature treatments when the analysis was carried out with NormFinder: A-Tub, ACT, SAND, H2B and IDH.
When comparing the tissue types (Table 1C), the best combination of primers for all species was IDH/ACT, which was also the same pair of genes identified in the analysis of plants/seedlings of E. globulus subjected to light and dark treatments (Table 1D).
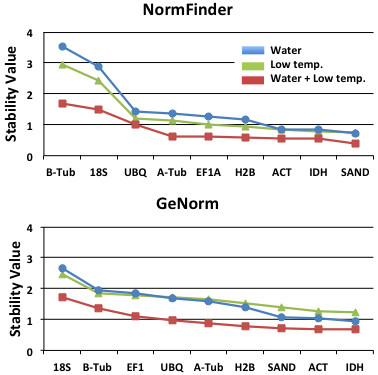
In the general analysis, i.e., all treatments, species and tissues types, SAND, IDH and ACT presented the lower stability values, indicating they were the most stable genes (Figure 1).
Figure 1.
The stability values of the genes calculated according to the two software programmes used in the analyses of the most stable genes. The graph shows the analyses differentiating the water stress treatments (control + water stress + water stress recovery), the temperature treatments (control + low temperature), and both treatments together, regardless of species or tissue types (apex and base).
Discussion
In this study, we used three Eucalyptus species in experiments where plants were exposed to different water regimes (sufficient, stressed, and recovery) and low temperature. For the analysis of plants under water stress, plant tissues were harvested for gene expression analysis only when they displayed the same wilting characteristics, i.e., lack of turgor recovery, in the youngest leaves. Consequently, the time required for wilting varied among plants. We previously used this same strategy to induce drought stress in coffee plants and observed that the water potential in the leaves, even among different species, was homogeneous, resulting in reduced variation in the gene expression analysis [36]. For the low-temperature treatments, the plants were placed in a cold chamber for 12 h daily for at least 6 days. We chose water stress as a treatment because it is the field condition that most limits plant growth [37]. The low-temperature treatment was selected because E. globulus is adapted and cultivated in cold regions [5-7].
We also used in our assays seedlings/plants of E. globulus subjected to light treatment. For these experiments, the seedlings were grown in either the light or the dark. In addition, the stem apex of six-month-old plants were covered with aluminium foil and grown under greenhouse conditions. The reason to include such treatments was because we have been using light in our studies on modulation of lignin biosynthesis in E. globulus (Araújo and Mazzafera, unpublished data).
The qPCR technique facilitated the discrimination of genes expressed during different treatments among plants of the same or different species [38,39]. However, to validate these comparisons, the analysis must consider genes that have constitutive expression to serve as references for the analysis of plant material from different sources [14,15]. The selection of these genes is used to evaluate the stability of gene expression in different tissues, using various bioinformatics tools. In the present study, two bioinformatics tools were used that differ in terms of data analysis. The software GeNorm analyses the data comprehensively, establishing a reference value or M-value (stability value). The candidates with the highest values are eliminated until only two candidates remain [40]. The tool NormFinder only analyses variations between samples and variations within each sample group [41]. Subsequently, the variances between and within groups are used to determine the stability value, for which smaller values are considered most stable. Although NormFinder provides a more detailed analysis as compared with GeNorm, both tools are widely used for the validation of normalisers.
In the evaluations performed in the present study using GeNorm and NormFinder, few consistencies were observed among the data generated using both methodologies, as also observed in other studies with eucalyptus [27]. This was mainly observed when data were compared separately. Some few exception were observed: for example, the water stress treatment (DRT column in Table 1A) showed that for both methodologies, the best endogenous gene pair in E. urograndis was H2B /UBQ. However, consistency appeared when more comprehensive analyses involving different species and treatments were carried out, what then highlighted the IDH gene. In the analyses related to the low-temperature treatment, the genes IDH and SAND were common in the analyses using GeNorm and NormFinder. However, in addition to IDH, ACT appeared to be stable in the analyses distinguishing the tissue type (apex or base) in the samples from for the stress and low-temperature treatments.
IDH was also the most common gene in the GeNorm and NormFinder analyses for the material subjected to different light regimens. It is interesting to observe that the gene UBQ was also identified in the E. globulus greenhouse material using GeNorm, while ACT was identified using the in vitro material with NormFinder. The results reflect differences in the experimental conditions and the developmental stages of the plants.
IDH, SAND and ACT were the genes identified here as the most frequent in the gene pairs indicated in the analyses using GeNorm and NormFinder. IDH, SAND and ACT perform different roles related to housekeeping functions in plant cells. SAND is a member of a large family of proteins that are found in many organisms, which was identified initially in Saccharomyces cerevisiae and subsequently in Schizosaccharomyces pombe, Caenorhabditis elegans, Drosophila melanogaster, mouse, human, and A. thaliana[42]. SAND protein is involved in membrane trafficking, enabling the fusion of the vesicle to the vacuole [43]. IDH is a mitochondrial enzyme involved in the oxidative decarboxylation of isocitrate in the Krebs cycle to produce α-ketoglutarate and CO2, a reaction coupled with the conversion of NAD+ to NADH. Five isoforms of IDH are found in A. thaliana. Four of these enzymes are expressed in all tissues, and only one is specific to the pollen grain [44]. ACT is an important plant cytoskeleton protein that has a pivotal role in several cellular processes involved in the regulation of cell growth and morphology [45].
de Oliveira et al. [27] evaluated the microarray data from 21,432 genes and selected the 50 most stable genes expressed in the leaves of E. grandis and xylem from E. grandis and E. globulus. Of these 50 genes, eight were chosen, together with seven other genes that were previously tested in the literature, for studies on the stability of gene expression in the tissues of six Eucalyptus species (leaf and xylem of E. grandis, E. dunnii, E. pellita, E. saligna and E. urophylla; xylem of E. globulus; and flowers of E. grandis). Overall, these 15 genes showed adequate expression stability in the tissues analysed, but the best results were obtained with the genes Eucons04, Eucons08, and Eucons21, which were selected from the microarray analyses. Eucons04 putatively encodes a protein similar to cyclin-dependent protein kinases (CDKs). Eucons08 encodes a transcription elongation factor SII (TFIIS). Eucons21 encodes a protein similar to aminoacyl-tRNA synthetases. All three genes showed similarities with genes from Ricinus communis. Among the genes used in our study, only H2B was included in the study of de Oliveira et al. [27], and it was moderately expressed. In our results, the H2B/A-Tub pair was consistently observed in the general analyses of drought and cold treatments in E. uroglobulus using the NormFinder software. Our data showed that the stability value places H2B as the fourth most stable gene, after ACT, IDH, and SAND. The differences observed between the data obtained in this study and that of de Oliveira et al. [27] could reflect differences in the species used and the treatments applied.
Other authors evaluated the stability of the genes used herein in eucalyptus. Almeida et al. [30] used the genes coding for 18S ribosomal RNA (18S), actin 2/7, histone H2B, NADP-isocitrate dehydrogenase, polyubiquitin, SAND protein, α-tubulin, TIP41-like protein, translation elongation factor 2, a expressed protein without determined function (33380), and a putative RNA-binding protein (EUC12) to study the formation of adventitious roots in E. globulus, of which the first eight genes were the same genes used in our study. de Almeida et al. [30] identified the best gene pairs as NADP-Isocitrate Dehydrogenase/SAND using GeNorm software, and Histone H2B/α-Tubulin using NormFinder. However, among the 13 genes, UBQ and IDH displayed the highest stability in two eucalyptus clones (rust-resistant and susceptible) exposed to biotic (P. psidii) and abiotic (acibenzolar-S-methyl) stresses [29]. In choosing the reference genes in studies on the acclimatisation of E. globulus to the cold, Fernández et al. [28] used the genes for elongation factor 1-a, actin, α-tubulin, protein phosphatase 1A, SAND, and ubiquitin C, concluding that the genes for ubiquitin, elongation factor 1-a and α-tubulin were the most stable.
Conclusions
The data of Boava et al. [29], Fernández et al. [28], de Almeida et al. [30], and the current study indicate that the most frequent stable genes in the various analyses performed in different eucalyptus species and under various conditions are ACT, UBQ, SAND, IDH, and A-Tub, with IDH as the most stable. Therefore, apparently these five genes could be used as reference genes in a wide range of treatments in eucalyptus research where gene expression is evaluated. Given that the use of more than one reference gene [46] or even several genes [47] have been recommended to minimise errors in the estimation of gene expression, we propose that IDH and at least two more of the five genes listed could be used in a wide range of studies of biotic and abiotic stresses in eucalyptus.
Methods
Plant material
The low-temperature and water stress experiments were performed with E. uroglobulus (E. urograndis x E. globulus Labill), E. globulus Labill and E. urograndis (E. urophylla S.T. Blake x E. grandis Hill ex-Maiden), which were kindly provided by the Fibria Cellulose SA (São Paulo, Brazil). Upon receipt, the seedlings were maintained in a greenhouse at room temperature until the low-temperature and drought trials were performed. The plants for the low-temperature trials were grown in 10 L pots containing a mixture of soil and sand (1:1, v/v). The plants for the water stress experiments were transplanted into 10 L pots containing a soil-sand mixture (2:1, v/v) and were maintained at three plants per pot. At the time of the experiments, the plants were 90 days old in the cold experiment and 120 days old in the water stress experiment.
Low temperature experiments
All the plants were maintained in a greenhouse at room temperature during the day. At the end of the day, half of the plants were transferred to cold and dark chambers, and the other half were transferred to chambers that were only dark and without temperature control. The plants were kept in these chambers for 12 h. The average values of the minimum and maximum temperature were, respectively, 3.2 and 5.7°C for the trial with E. urograndis, 6.1 and 9.1°C for E. globulus, and 5.7 and 8.8°C for E. uroglobulus. The cold experiments lasted seven (E. urograndis), eight (E. globulus), and ten (E. uroglobulus) days. In the control dark chamber, the average temperature ranged from 20 to 26°C. In the greenhouse, the temperature ranged from 22 to 30°C during the day.
Water stress experiments
In these trials, 120-day-old plants grown in a greenhouse with no temperature control were divided into three groups (control, water stress, and water stress recovery). The control plants were watered daily. For the plants subjected to water stress, the irrigation was interrupted. When each individual plant showed symptoms of wilting in the younger leaves at 12:00 pm on a given day, without recovery on the morning (8:00 am) next day, the plant tissues were collected. This strategy indicated that the plant had lost the capacity to recover water turgor during the night [36]. Because the plants were only collected when showing constant signs of wilting, the development of stress in each plant occurred at different times. In general, the experiments lasted an average of 13 (E. uroglobulus), 19 (E. globulus), and 21 (E. urograndis) days. In the third group, the plants that did not recover their water turgor in the morning were watered, and after recovery, which occurred at approximately 16 h after irrigation, the tissues were collected.
Light treatment
The seedling of E. globulus originating from seeds germinated in vitro were subjected to light treatments. The seeds were sterilised with 70% alcohol for 2 min, followed by four washes with 2% hypochlorite solution and five washes with sterile water. The seeds were subsequently placed in vials containing 2.5% (w/v) phytagel, prepared with 0.5X Murashige and Skoog media (MS; [48]). The vials were placed either under light (25°C, 12 h photoperiod, 120 μmols photons.m-2.s-1) or in the dark (25°C) for 15 days (Figure 2). After this period hypocotyls were collected for analysis. To compare the results obtained with seedlings hypocotyls with plants at a more advanced stage of development, the apexes of six-month-old E. globulus plants were either covered with aluminium foil or uncovered for 30 days. The plants were maintained in the greenhouse without temperature control.
Figure 2.
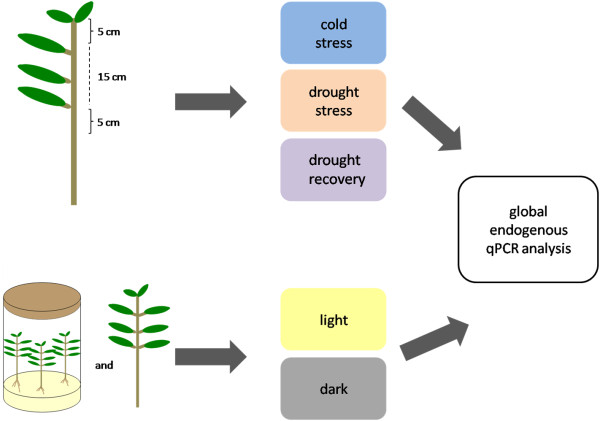
A schematic diagram of the treatments used in the characterisation of the pattern of endogenous gene expression in eucalyptus. Gene expression under low temperatures, drought, and recovery from drought treatments was analysed in E. globulus, E. uroglobulus, and E. Urograndis plants. Only E. globulus was used in the light and dark trial. The seedlings were cultivated in vitro, and the six-month plants were grown in the greenhouse.
Tissue collection
At the end of the water stress and low-temperature trials, the leaves were removed, and stem collection was performed. The first 5 cm from the apex (denominated apex) was collected, the following 15 cm were discarded, and the following 5 cm were collected (denominated base) (Figure 2). The apex represented the region where longitudinal cell growth is predominant, and the base represented the region where radial growth increases in intensity. The collected tissues were immediately frozen in liquid N2 and stored in a biofreezer (−80°C). Hypocotyls segments from seedlings grown in vitro were cut from the middle of the hypocotyl, frozen in liquid N2, and stored in a biofreezer (−80°C) (Figure 2).
In all experiments three biological replicates were used, each one composed by five plants (water and low temperature experiments) or ten seedlings (light experiment). For each replicate three technical replicates were made in the qPCR analysis.
Nucleic acid manipulation
Total RNA extraction from the stem segments of the plants under stress was performed according to Chang et al. [49], with slight modifications. Briefly, 750 μL of extraction buffer was heated to 65°C in 2 mL Eppendorf tubes (1 tube per extracted sample), and 7.5 μL of 2-mercaptoethanol was added to each tube. Approximately 100 mg of sample was macerated in N2 and added to each tube followed by intense agitation in a vortex and cooling at room temperature. Subsequently, the RNA was extracted once with 750 μL of chloroform:isoamyl alcohol (24:1), mixed for 2 min, and centrifuged at 10,000 rpm for 10 min at 4°C. The supernatant was collected, and after the addition of 188 μL of a solution of 10 M LiCl, the RNA was precipitated for 12 h at 4°C. Then, the samples were centrifuged at 10,000 rpm for 30 min at room temperature. The supernatant was then discarded, and the pellet was dissolved in 200 μL of SSTE (37°C). A total of 200 μL of chloroform:isoamyl alcohol (24:1, v/v) was added. After manual agitation by tube inversion, the mixture was centrifuged at 10,000 rpm for 15 min, and the supernatant was collected. Subsequently, 1 mL of absolute ethanol was added to the supernatant and incubated at −20°C for 2 h. The samples were centrifuged at 14,000 rpm for 20 min at 4°C, and the supernatants were discarded. The pellet was washed with 75% ethanol and centrifuged at 14,000 rpm for 20 min at 4°C. The final supernatant was discarded, and the pellet was dried at room temperature. The dried RNA pellet was dissolved in 20 μL of water treated with diethylpyrocarbonate. Total RNA was extracted from the seedling hypocotyls using TRIZOL reagent (Invitrogen, Carlsbad, CA, USA). All of the extracted RNA was treated with DNAse (turbo DNA-free, Ambion), and the cDNA synthesis was performed using the SuperScriptR VILOTM cDNA synthesis kit (Invitrogen). The quantification of the RNA concentration from each sample was performed in a Qubit fluorometer (Invitrogen), and the RNA quality was assessed on a 1% (m/v) agarose gel using ethidium bromide with subsequent visualisation under UV light.
Primer design
The sequences of the genes used were obtained from four public banks related to Eucalyptus ( http://bioinformatics.psb.ugent.be/webtools/bogas, http://www.phytopkmdzome.net/eucalyptus.php, http://eucalyptusdb.bi.up.ac.za, and http://www.polebio.scsv.ups-tlse.fr/eucatoul/). The primers were designed using the internet software Primer3 [50] according to the following parameters: 18 and 30 bp in length, 60°C melting temperature, 40-60% GC content, and a 50–150 bp amplified fragments length.
Nine genes were selected for the stability study (Table 2): elongation factor 1-alpha (EF1), ß-tubulin (B-Tub), actin (ACT), ubiquitin (UBQ), SAND protein (SAND), isocitrate dehydrogenase (IDH), histone (H2B), α-tubulin (A-Tub), and 18S ribosomal RNA (18S). The choice of these genes was based on previous studies with Eucalyptus[28,30]. A pair of primers was designed for each gene, and for the UBQ gene, one pair of primers was designed based on the sequence of E. globulus and another based on the sequence of E. urograndis (Table 3). For each primer pair, the amplification efficiency was determined based on the slope of the standard curve for each of the genes.
Table 2.
The gene identification numbers and names of Eucalyptus genes and their orthologous genes and functions in A. thaliana
| Gene ID | Gene name | A. thaliana gene | Function | Reference |
|---|---|---|---|---|
|
EF1a |
Translation elongation factor |
At1g18070 |
Translation factor activity, nucleic acid binding, GTP binding, GTPase activity; |
[51] |
|
B-Tub |
β-Tubulin |
At5g62690 |
GTPase activity, protein binding, structural molecule activity |
[52] |
|
ACT |
Actin |
At5g09810 |
Structural constituent of cytoskeleton |
[53] |
|
UBQ |
Ubiquitin |
At4g05050 |
Signalling complexes for protein degradation, translation control, DNA repair, endocytosis regulation, protein traffic |
[54] |
|
SAND |
SAND protein |
At2g28390 |
Intracellular vesicular transport, biogenesis and vacuole signalling |
[17] |
|
IDH |
Isocitrate dehydrogenase |
At1g54340 |
Isocitrate dehydrogenase (NADP+) activity, oxidoreductase activity, acting on the CH-OH group of donors, NAD or NADP as acceptor |
[55] |
|
H2B |
Histone H2B |
At5g22880 |
Structural constituent of the eukaryotic nucleosome core |
[51] |
|
A-Tub |
α-Tubulin |
At5g19780 |
Structural constituent of cytoskeleton, microtubule-based processes |
[56] |
| 18S | RNA ribosomal 18S | At3g41768 | Cytosolic small ribosomal subunit, translation | [57] |
Table 3.
Gene identification and primers sequences used in the qPCR analyses
| Gene | Primer | Primer sequence (5’-3’) | Amplicon (bp)* | Amplification efficiency (%)** |
|---|---|---|---|---|
| Elongation factor-1α |
Forward |
CCTGTCCTTGATTGTCACACTTCC |
130 |
110 |
| Reverse |
CCATTCCAGCATCACCGTTCTTC |
|||
| Ubiquitin (E. gobulus) |
Forward |
TCCGTCAAAAGCGAACAGA |
173 |
97 |
| Reverse |
CATTTCCCTCCAGATTACCC |
|||
| Ubiquitin (E. urograndis) |
Forward |
GGACTTTCGTTCGTTTTGGT |
107 |
97 |
| Reverse |
GTGATTTGGGGAGGGTTTG |
|||
| Actin |
Forward |
AGATGACCCAGATTATGTTTGAGACCTTC |
122 |
97 |
| Reverse |
ACCATCACCAGAATCCAACACAATACC |
|||
| SAND protein |
Forward |
TGGGTCACACAGGATTTTGA |
130 |
100 |
| Reverse |
CTCCCAGCAAAAAGATCTCG |
|||
| Isocitrate dehydrogenase-NADP |
Forward |
AGTTTGAGGCTGCTGGAATC |
100 |
103 |
| Reverse |
CTTGCATGCCCACACATAAC |
|||
| Histone H2B |
Forward |
AACAAGAAGCCCACCATCAC |
142 |
96 |
| Reverse |
ACAACTTCCTCCTCGCTCAC |
|||
| α-Tubulin |
Forward |
CCAGTGAACAAATGCCCTCT |
92 |
107 |
| Reverse |
TGATCAGCAACAACACAGCA |
|||
| Ribossomal 18S |
Forward |
CATGGCCGTTCTTAGTTGGT |
71 |
95 |
| Reverse |
TAGCAGGCTGAGGTCTCGTT |
|||
| β-Tubulin |
Forward |
GATGGGGACGCTATTGATTT |
225 |
100 |
| Reverse | CTTGGGTTGATGAGTTTCAGG |
* Melting temperature = 60°C for all primers. **E=10^(-1/slope)-1.
Real time quantitative RT-PCR (qPCR)
The qPCR reactions were performed using a iCycler iQ5 (Bio-Rad). The reactions contained a final volume of 10 μL and included 5 μl of QuantiFastTM SYBR Green – PCR Mix (Qiagen), 0.4 μM of primers, 3 μL of cDNA, and 1.6 μL of autoclaved MilliQ water. The thermocycling conditions were 95°C for 3 min, followed by 40 cycles of 95°C for 10 sec and 60°C for 30 sec and 71 cycles of 60°C for 30 seconds. The reactions were performed in triplicate, and for each biological replicate, technical triplicates were performed. Control reactions, which contained no DNA template, were conducted for all of the experiments (NTC – Non Template Control).
Data analysis
Determination of the best normaliser gene or best gene pair was performed using two software programmes: NormFinder[41] and GeNorm[40]. Given that NormFinder allows the selection of up to two genes and GeNorm allows the selection of more than two, we chose to work with the best combination of two genes. Expression levels were determined as the necessary number of cycles for the amplifications reach a threshold fixed in the exponential phase of PCR reaction (CT) [58]. The CTs were transformed into quantities using PCR efficiencies [40] in order to use GeNorm and NormFinder.
Abbreviations
EF1: Elongation factor 1-alpha; B-Tub: ß-tubulin; ACT: Actin; UBQ: Ubiquitin; SAND: SAND protein; IDH: Isocitrate dehydrogenase; H2B: Histone; A-Tub: α-tubulin; 18S: 18S RNA ribosomal; qPCR: Reverse-transcription followed by quantitative real- time Polymerase Chain Reaction; RNA: Ribonucleic Acid; PCR: Polymerase Chain Reaction; cDNA: Complementary Deoxyribonucleic Acid.
Competing interests
The authors declare no competing interest.
Authors’ contributions
JCMSM and PA contributed equally to this work, as they performed the experiments, analysed the data and helped to write the manuscript draft. URS helped to perform RNA extraction and some of the experiments. MSB analysed the data and helped to write the manuscript draft. JOFV produced the plants and gave information about their agronomical characteristics. PM designed and supervised the study, wrote the final version of the paper. All authors read and approved the last version of the manuscript.
Contributor Information
Jullyana Cristina Magalhães Silva Moura, Email: jucbio@gmail.com.
Pedro Araújo, Email: araujo.pedro@gmail.com.
Michael dos Santos Brito, Email: msantosbrito@yahoo.com.br.
Uiara Romero Souza, Email: uiarakrs@yahoo.com.br.
Julianade Oliveira Fernandes Viana, Email: juliana.viana@fibria.com.br.
Paulo Mazzafera, Email: pmazza@unicamp.br.
Acknowledgements
J.C.M.S.M., P.A. and M.S.B. thank CNPq-Brazil, CAPES-Brazil and FAPESP-São Paulo for doctoral (J.C.M.S.M. and P.A.) and post-doctoral fellowships. P.M. thanks CNPq for a research fellowship. The authors thank Cesar Augusto Valencise Bonine (Fibria Celulose S.A.) for continuous support.
References
- Stape JL, Binkley D, Ryan MG. Eucalyptus production and the supply, use and efficiency of use of water, light and nitrogen across a geographic gradient in Brazil. Forest Ecol Manag. 2004;193:17–31. doi: 10.1016/j.foreco.2004.01.020. [DOI] [Google Scholar]
- Mora AL, Garcia CH. The Eucalyptus culture in Brazil (A cultura do eucalipto no Brasil) São Paulo - Brazil: Sociedade Brasileira de Silvicultura-SBS; 2000. [Google Scholar]
- Grattapaglia D, Vaillancourt R, Shepherd M, Thumma B, Foley W, Külheim C, Potts B, Myburg A. Progress in Myrtaceae genetics and genomics: Eucalyptus as the pivotal genus. Tree Genet Genomes. 2012;8:463–508. doi: 10.1007/s11295-012-0491-x. [DOI] [Google Scholar]
- Cotterill PP, Brolin A. Improving Eucalyptus wood, pulp and paper quality by genetic selection. Salvador, Brazil: EMBRAPA-Brazil; 1997. pp. 1–14. (Proceedings of the IUFRO Conference on Silviculture and Improvement of Eucalypts). [Google Scholar]
- Pascoal-Neto C, Evtuguin D, Pinto P. Componentes macromoleculares das madeiras de Eucalyptus e de outras folhosas: estrutura e influencia na aptidão ao cozimento e branqueamento. O Papel - Aveiro. 2005;1:17–27. [Google Scholar]
- Silvestre AJD, Pascoal-Neto C, Freire CSR. Componentes lipofílicos da madeira de Eucalyptus globulus: composição e comportamento durante a produção de pasta de papel. O Papel - Aveiro. 2005;1:5–16. [Google Scholar]
- Chiang VL. From rags to riches. Nature Biotechnol. 2002;20:557–558. doi: 10.1038/nbt0602-557. [DOI] [PubMed] [Google Scholar]
- Hillman DC, Rooks A. Single species pulping: the worlds preferred market pulps. Solutions! 2002;85:28–30. [Google Scholar]
- Cruz MP, Barbosa LCA, Maltha CRA, Gomide JL, Milanez AF. Caracterização química do “pitch” em indústria de celulose e papel de Eucalyptus. Quim Nova. 2006;29:459–466. doi: 10.1590/S0100-40422006000300011. [DOI] [Google Scholar]
- Silvério F, Barbosa L, Silvestre A, Piló-Veloso D, Gomide J. Comparative study on the chemical composition of lipophilic fractions from three wood tissues of Eucalyptus species by gas chromatography–mass spectrometry analysis. J Wood Sci. 2007;53:533–540. doi: 10.1007/s10086-007-0901-0. [DOI] [Google Scholar]
- Moura-Sobczak J Souza U Mazzafera P Drought stress and changes in the lignin content and composition in Eucalyptus BMC Proc 20115Suppl 7P103. 10.1186/1753-6561-5-S7-P10322373057 [DOI] [Google Scholar]
- Gachon C, Mingam A, Charrier B. Real-time PCR: what relevance to plant studies? J Exp Bot. 2004;55:1445–1454. doi: 10.1093/jxb/erh181. [DOI] [PubMed] [Google Scholar]
- Thellin O, Zorzi W, Lakaye B, De Borman B, Coumans B, Hennen G, Grisar T, Igout A, Heinen E. Housekeeping genes as internal standards: use and limits. J Biotechnol. 1999;75:291–295. doi: 10.1016/S0168-1656(99)00163-7. [DOI] [PubMed] [Google Scholar]
- Dheda K, Huggett JF, Chang JS, Kim LU, Bustin SA, Johnson MA, Rook GAW, Zumla A. The implications of using an inappropriate reference gene for real-time reverse transcription PCR data normalization. Anal Biochem. 2005;344:141–143. doi: 10.1016/j.ab.2005.05.022. [DOI] [PubMed] [Google Scholar]
- Guenin S, Mauriat M, Pelloux J, Van Wuytswinkel O, Bellini C, Gutierrez L. Normalization of qRT-PCR data: the necessity of adopting a systematic, experimental conditions-specific, validation of references. J Exp Bot. 2009;60:487–493. doi: 10.1093/jxb/ern305. [DOI] [PubMed] [Google Scholar]
- Volkov RA, Panchuk II, Schöffl F. Heat‐stress‐dependency and developmental modulation of gene expression: the potential of house‐keeping genes as internal standards in mRNA expression profiling using real‐time RT‐PCR. J Exp Bot. 2003;54:2343–2349. doi: 10.1093/jxb/erg244. [DOI] [PubMed] [Google Scholar]
- Czechowski T, Stitt M, Altmann T, Udvardi MK, Scheible WR. Genome-wide identification and testing of superior reference genes for transcript normalization in Arabidopsis. Plant Physiol. 2005;139:5–17. doi: 10.1104/pp.105.063743. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jain M, Nijhawan A, Tyagi AK, Khurana JP. Validation of housekeeping genes as internal control for studying gene expression in rice by quantitative real-time PCR. Biochem Biophys Res Commun. 2006;345:646–651. doi: 10.1016/j.bbrc.2006.04.140. [DOI] [PubMed] [Google Scholar]
- Iskandar H, Simpson R, Casu R, Bonnett G, Maclean D, Manners J. Comparison of reference genes for quantitative real-time polymerase chain reaction analysis of gene expression in sugarcane. Plant Mol Biol Rep. 2004;22:325–337. doi: 10.1007/BF02772676. [DOI] [Google Scholar]
- Remans T, Smeets K, Opdenakker K, Mathijsen D, Vangronsveld J, Cuypers A. Normalisation of real-time RT-PCR gene expression measurements in Arabidopsis thaliana exposed to increased metal concentrations. Planta. 2008;227:1343–1349. doi: 10.1007/s00425-008-0706-4. [DOI] [PubMed] [Google Scholar]
- Nicot N, Hausman JF, Hoffmann L, Evers D. Housekeeping gene selection for real-time RT-PCR normalization in potato during biotic and abiotic stress. J Exp Bot. 2005;56:2907–2914. doi: 10.1093/jxb/eri285. [DOI] [PubMed] [Google Scholar]
- Hong S-Y, Seo P, Yang M-S, Xiang F, Park C-M. Exploring valid reference genes for gene expression studies in Brachypodium distachyon by real-time PCR. BMC Plant Biol. 2008;8:112. doi: 10.1186/1471-2229-8-112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Exposito-Rodriguez M, Borges AA, Borges-Perez A, Perez JA. Selection of internal control genes for quantitative real-time RT-PCR studies during tomato development process. BMC Plant Biol. 2008;8:131. doi: 10.1186/1471-2229-8-131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barsalobres-Cavallari CF, Severino FE, Maluf MP, Maia IG. Identification of suitable internal control genes for expression studies in Coffea arabica under different experimental conditions. BMC Mol Biol. 2009;10:1. doi: 10.1186/1471-2199-10-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Paolacci A, Tanzarella O, Porceddu E, Ciaffi M. Identification and validation of reference genes for quantitative RT-PCR normalization in wheat. BMC Mol Biol. 2009;10:11. doi: 10.1186/1471-2199-10-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schmidt GW, Delaney SK. Stable internal reference genes for normalization of real-time RT-PCR in tobacco (Nicotiana tabacum) during development and abiotic stress. Mol Genet Genom. 2010;283:233–241. doi: 10.1007/s00438-010-0511-1. [DOI] [PubMed] [Google Scholar]
- De Oliveira LA, Breton MC, Bastolla FM, Camargo SS, Margis R, Frazzon J, Pasquali G. Reference genes for the normalization of gene expression in Eucalyptus species. Plant Cell Physiol. 2012;53:405–422. doi: 10.1093/pcp/pcr187. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fernández M, Villarroel C, Balbontín C, Valenzuela S. Validation of reference genes for real-time qRT-PCR normalization during cold acclimation in Eucalyptus globulus. Trees - Struct Funct. 2010;24:1109–1116. doi: 10.1007/s00468-010-0483-0. [DOI] [Google Scholar]
- Boava LP, Laia ML, Jacob TR, Dabbas KM, Goncalves JF, Ferro JA, Ferro MIT, Furtado EL. Selection of endogenous genes for gene expression studies in Eucalyptus under biotic (Puccinia psidii) and abiotic (acibenzolar-S-methyl) stresses using RT-qPCR. BMC Res Notes. 2010;3:1–9. doi: 10.1186/1756-0500-3-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- de Almeida MR, Ruedell CM, Ricachenevsky FK, Sperotto RA, Pasquali G, Fett-Neto AG. Reference gene selection for quantitative reverse transcription-polymerase chain reaction normalization during in vitro adventitious rooting in Eucalyptus globulus Labill. BMC Mol Biol. 2010;11:73. doi: 10.1186/1471-2199-11-73. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Navarrete-Campos D, Bravo L, Rubilar R, Emhart V, Sanhueza R. Drought effects on water use efficiency, freezing tolerance and survival of Eucalyptus globulus and Eucalyptus globulus × nitens cuttings. New For. 2012. [DOI]
- White DA, Beadle CL, Worledge D. Leaf water relations of Eucalyptus globulus ssp. globulus and E. nitens: seasonal, drought and species effects. Tree Physiol. 1996;16:469–476. doi: 10.1093/treephys/16.5.469. [DOI] [PubMed] [Google Scholar]
- Silva FC, Shvaleva A, Maroco JP, Almeida MH, Chaves MM, Pereira JS. Responses to water stress in two Eucalyptus globulus clones differing in drought tolerance. Tree Physiol. 2004;24:1165–1172. doi: 10.1093/treephys/24.10.1165. [DOI] [PubMed] [Google Scholar]
- Silva FC, Shvaleva A, Broetto F, Ortuño MF, Rodrigues ML, Almeida MH, Chaves MM, Pereira JS. Acclimation to short-term low temperatures in two Eucalyptus globulus clones with contrasting drought resistance. Tree Physiol. 2009;29:77–86. doi: 10.1093/treephys/tpn002. [DOI] [PubMed] [Google Scholar]
- Li C, Berninger F, Koskela J, Sonninen E. Drought responses of Eucalyptus microtheca provenances depend on seasonality of rainfall in their place of origin. Funct Plant Biol. 2000;27:231–238. doi: 10.1071/PP99056. [DOI] [Google Scholar]
- Santos AB, Mazzafera P. Dehydrins are highly expressed in water-stressed plants of two coffee species. Trop Plant Biol. 2012. [DOI]
- Passioura JB. In: Drought tolerance in higher plants: genetical, physiological, and molecular biological analysis. Belhassen I, editor. Dordrecht: Kluwer Academic; 1997. Drought and drought tolerance; pp. 1–7. [Google Scholar]
- VanGuilder HD, Vrana KE, Freeman WM. Twenty-five years of quantitative PCR for gene expression analysis. Biotechniques. 2008;44:619–626. doi: 10.2144/000112776. [DOI] [PubMed] [Google Scholar]
- Ginzinger DG. Gene quantification using real-time quantitative PCR: An emerging technology hits the mainstream. Exp Hematol. 2002;30:503–512. doi: 10.1016/S0301-472X(02)00806-8. [DOI] [PubMed] [Google Scholar]
- Vandesompele J, De Preter K, Pattyn F, Poppe B, Van Roy N, De Paepe A, Speleman F. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 2002;3 doi: 10.1186/gb-2002-3-7-research0034. research0034.0031–0011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Andersen CL, Jensen JL, Ørntoft TF. Normalization of real-time quantitative reverse transcription-PCR data: a model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Cancer Res. 2004;64:5245–5250. doi: 10.1158/0008-5472.CAN-04-0496. [DOI] [PubMed] [Google Scholar]
- Cottage A, Mullan L, Portela MBD, Hellen E, Carver T, Patel S, Vavouri T, Elgar G, Edwards YJK. Molecular characterisation of the sand protein family: a study based on comparative genomics, structural bioinformatics and phylogeny. Cell Mol Biol Lett. 2004;9:739–753. [PubMed] [Google Scholar]
- Wang CW, Stromhaug PE, Kauffman EJ, Weisman LS, Klionsky DJ. Yeast homotypic vacuole fusion requires the Ccz1-Mon1 complex during the tethering/docking stage. J Cell Biol. 2003;163:973–985. doi: 10.1083/jcb.200308071. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lemaitre T, Hodges M. Expression analysis of Arabidopsis thaliana NAD-dependent isocitrate dehydrogenase genes shows the presence of a functional subunit that is mainly expressed in the pollen and absent from vegetative organs. Plant Cell Physiol. 2006;47:634–643. doi: 10.1093/pcp/pcj030. [DOI] [PubMed] [Google Scholar]
- Hussey PJ, Ketelaar T, Deeks MJ. Control of the actin cytoskeleton in plant cell growth. Annu Rev Plant Biol. 2006;57:109–125. doi: 10.1146/annurev.arplant.57.032905.105206. [DOI] [PubMed] [Google Scholar]
- Hellemans J, Mortier G, De Paepe A, Speleman F, Vandesompele J. qBase relative quantification framework and software for management and automated analysis of real-time quantitative PCR data. Genome Biol. 2007;8:R19. doi: 10.1186/gb-2007-8-2-r19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bustin SA, Benes V, Garson JA, Hellemans J, Huggett J, Kubista M, Mueller R, Nolan T, Pfaffl MW, Shipley GL. et al. The MIQE guidelines: minimum information for publication of quantitative real- time PCR experiments. Clin Chem. 2009;55:611–622. doi: 10.1373/clinchem.2008.112797. [DOI] [PubMed] [Google Scholar]
- Murashige T, Skoog F. A revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiol Plant. 1962;15:473–497. doi: 10.1111/j.1399-3054.1962.tb08052.x. [DOI] [Google Scholar]
- Chang S, Puryear J, Cairney J. A simple and efficient method for isolating RNA from pine trees. Plant Mol Biol Rep. 1993;11:113–116. doi: 10.1007/BF02670468. [DOI] [Google Scholar]
- Rozen S, Skaletsky H. Primer3 on the WWW for general users and for biologist programmers. Method Mol Biol. 2000;132:365–386. doi: 10.1385/1-59259-192-2:365. [DOI] [PubMed] [Google Scholar]
- Pendle AF, Clark GP, Boon R, Lewandowska D, Lam YW, Andersen J, Mann M, Lamond AI, Brown JWS, Shaw PJ. Proteomic analysis of the Arabidopsis nucleolus suggests novel nucleolar functions. Mol Biol Cell. 2005;16:260–269. doi: 10.1091/mbc.E04-09-0791. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Snustad DP, Haas NA, Kopczak SD, Silflow CD. The small genome of Arabidopsis contains at least nine expressed beta-tubulin genes. Plant Cell. 1992;4:549–556. doi: 10.1105/tpc.4.5.549. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McDowell JM, An YQ, Huang SR, McKinney EC, Meagher RB. The arabidopsis ACT7 actin gene is expressed in rapidly developing tissues and responds to several external stimuli. Plant Physiol. 1996;111:699–711. doi: 10.1104/pp.111.3.699. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Callis J, Carpenter T, Sun CW, Vierstra RD. Structure and evolution of genes encoding polyubiquitin and ubiquitin-like proteins in Arabidopsis thaliana ecotype Columbia. Genetics. 1995;139:921–939. doi: 10.1093/genetics/139.2.921. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Behal RH, Oliver DJ. NAD(+)-dependent isocitrate dehydrogenase from Arabidopsis thaliana. Characterization of two closely related subunits. Plant Mol Biol. 1998;36:691–698. doi: 10.1023/A:1005923410940. [DOI] [PubMed] [Google Scholar]
- Kopczak SD, Haas NA, Hussey PJ, Silflow CD, Snustad DP. The small genome of Arabidopsis contains at least six expressed alpha-tubulin genes. Plant Cell. 1992;4:539–547. doi: 10.1105/tpc.4.5.539. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Unfried I, Stocker U, Gruendler P. Nucleotide sequence of the 18S rRNA gene from Arabidopsis thaliana Col0. Nucl Acids Res. 1989;17:7513. doi: 10.1093/nar/17.18.7513. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Walker NJ. A technique whose time has come. Science. 2002;296:557–559. doi: 10.1126/science.296.5567.557. [DOI] [PubMed] [Google Scholar]