Abstract
Objective
To identify the signatures of miRNAs differentially expressed in HER2(+) versus HER2(−) breast cancers that accurately predict the HER2 status of breast cancer, and to provide further insight into breast cancer therapy.
Methods
By the methods of literature search, aberrant expressed miRNAs were collected. By target prediction algorithm of TargetScan and PicTar and the data enrichment analysis, target gene sets of miRNAs differentially expressed in HER2(+) versus HER2(−) breast cancers were built. Then, using the Database for Annotation, Visualization, and Integrated Discovery (DAVID) database, the function modules of Gene Ontology categories and Kyoto Encyclopedia of Genes and Genomes (KEGG) and BIOCARTA pathway, biological functions and signaling pathways that are probably regulated by miRNAs, were analyzed.
Results
We got five sets of miRNAs expressed in different HER2 status of breast cancers finally. The five sets of data contain 22; 32; 3; 38; and 62 miRNAs, respectively. After miRNAs target prediction and data enrichment, 5,734; 22,409; 1,142; 22,293; and 43,460 target genes of five miRNA sets were collected. Gene ontology analysis found these genes may be involved in transcription, protein transport, angiogenesis, and apoptosis. Moreover, certain KEGG and BIOCARTA signaling pathways related toHER2 status were found.
Conclusion
Using TargetScan and PicTar for data enrichment, and DAVID database, Gene Ontology categories, KEGG and BIOCARTA pathway for analysis of miRNAs different expression, we conducted a new method for biological interpretation of miRNA profiling data in HER2(+) versus HER2(−) breast cancers. It may improve understanding the regulatory roles of miRNAs in different molecular subtypes of breast cancers. Therefore, it is beneficial to improve the accuracy of experimental efforts to breast cancer and potential therapeutic targets.
Key words: bioinformatics, breast cancer, HER2, miRNAs
Introduction
MiRNAs, a small noncoding RNA of 21–22 nucleotides long, have recently been linked to cancer development.1 miRNAs have diverse functions, which include the regulation of cellular differentiation, proliferation, and apoptosis.2 Recently, altered miRNA expression has been reported in various cancers, and the profiles of tissue miRNAs exhibit great potential for an application in cancer diagnosis.3,4 Therefore, miRNAs may be regarded as novel noninvasive biomarkers for diagnosis of cancer and other diseases.
The available evidence has shown that miRNAs widely participate in the development or progression of many types of cancers, including breast cancer. Breast cancer is the most common cancer in women and the second most common cause of cancer-related deaths in women, with ∼230,480 new cases of invasive breast cancer and 39,520 deaths expected in US women in 2011.5 Among the genes that may be potentially affected by miRNA, HER2 is perhaps the best known. It is a member of the erbB gene family, which is associated with breast cancer.6 HER2 is expressed in embryonic development and plays an important role in the growth and development of a variety of tissues and organs in adult. This gene is expressed at a low level in normal human tissues. However, when it is overexpressed, it produces the malignant phenotype and leads to cell proliferation. Approximately 25% of human breast cancers overexpress the HER2 proto-oncogene, and these breast cancers have a more aggressive tumor phenotype and produce a poor prognosis in patients with this disease.7 Therefore, HER2 has been an important prognostic indictor of breast cancer. However, there is little knowledge regarding the precise regulation of these receptors. Because the role of miRNAs in breast cancer has been widely investigated, here we sought to identify miRNAs associated with the HER2 receptor. One study has found that HER2 may upregulate CXCR4 by inhibiting expression of miR-139 in gastric cancer cells at the epigenetic level.8
While thousands of mammalian genes are potentially targeted by miRNAs, the functions of miRNAs in the context of gene networks are not well understood. In the present study, we use the corresponding online database to make a full bioinformatics analysis of miRNA signatures for differently expressed in HER2(+) versus HER2(−) breast cancers. Our goal was to determine the functions of HER2-related miRNAs and provide further insight into breast cancer therapy.
Material and Methods
Literature reviews
The Gene Expression Omnibus (GEO) database (www.ncbi.nlm.nih.gov/geo)9 was established at the National Center for Biotechnology Information (NCBI) in 2003. It is a public repository that archives and freely distributes microarray and other forms of high-throughput functional genomic data submitted by the scientific community. Today, it stores over 20,000 microarray- and sequence-based functional genomics studies, and continues to handle the majority of direct high-throughput data submissions from the research community. ArrayExpress (www.ebi.ac.uk/arrayexpress),10 an international public archive, was launched in 2002. The repository contains data from over 6,000 experiments comprising ∼200,000 assays, and the database doubles in size every 15 months. The majority of the data are array based, including high-throughput sequencing transcriptomics and epigenetic data. We retrieved any article that was categorized by the word of miRNAs and breast cancer in the two databases. Then, we make a further analysis of these microarray data related to HER2 status in these literatures. We take the same microarray data of the same experiment as one miRNA data.
miRNA target predictions
TargetScan (http://genes.mit.edu/targetscan/)11 provides a significant increase of available miRNA targets derived from all contemporary experimental techniques. It hosts detailed information for each miRNA–gene interaction, ranging from miRNA- and gene-related facts to information specific to the experimental validation methodologies and their outcomes. The database will be regularly updated in 4-month intervals so as to capture the growing number of publications covering novel targets. The PicTar (http://pictar.bio.nyu.edu)12 is the first method that uses the criteria of coexpression in space and time of miRNAs and their targets. And the experimental validation of 7 out of 13 tested predicted targets demonstrates the efficiency of the algorithm. We predicted the target genes of miRNA by these two softwares, and merged the final target gene predictions of one miRNA and removed duplicate results. In the same way, we established the corresponding set of target genes for one miRNA data set.
Enrichment analysis and signaling analysis
Database for Annotation, Visualization, and Integrated Discovery (DAVID) Knowledgebase,13 a gene-centered database, not only provides the quick accessibility to a wide range of heterogeneous annotation data in a centralized location, but also enriches the level of biological information for an individual gene. It contains different files, such as gene ontology (GO) biological process (BP), GO molecular function (MF), Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway, and BIOCARTA pathway. And it intends to integrate and organize the high-quality, world-class bioinformatics databases into a centralized location in a gene-centric format. In this way, more than 40 publicly available functional annotation sources can be comprehensively integrated and centralized by the DAVID gene clusters.
For our first step, we turned redundant gene IDs into the DAVID gene clusters to improve the cross-referencing capability. With the entire human genome for the background control, we filtered out the specific gene expressed in breast cancer tissue via the use of the “target gene set in breast cancer tissue-specific expression enrichment organization” module of the DAVID database. Secondly, with the same human genome as the background control, we classified genetic features screening in the DAVID database by the modules of GO BP and GO MF analysis, KEGG pathway and BIOCARTA pathway, which specify a statistically significant threshold p≤0.05. In summary, the flow chart of our study is showed in the Figure 1.
FIG. 1.
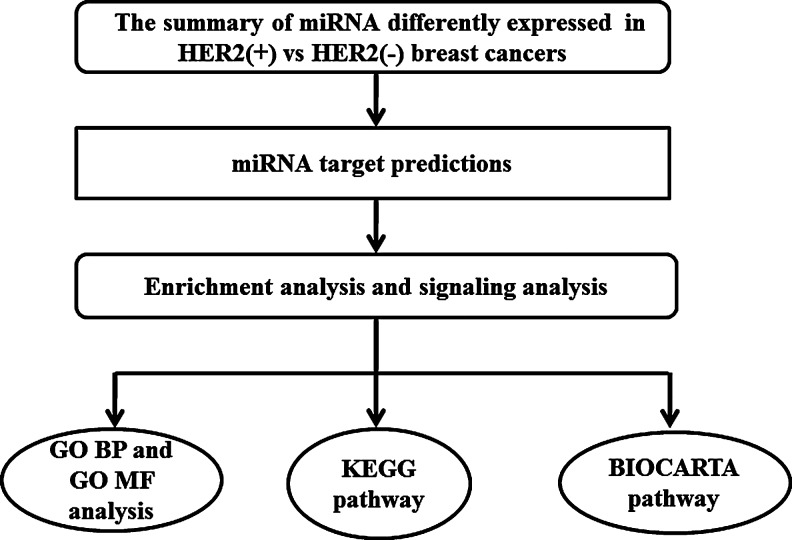
Flow chart of bioinformatics analysis of miRNAs differentially expressed in HER2(+) versus HER2(−) breast cancers. BP, biological process; GO, gene ontology; KEGG, Kyoto Encyclopedia of Genes and Genomes; MF, molecular function.
Results
Differentially expressed miRNAs in HER2(+) versus HER2(−) breast cancers
In the GEO and ArrayExpress database, we analyzed five microarray data sets. By combining the original literature reviews and taking the threshold p≤0.05, we filtered out five sets of miRNAs differentially expressed in HER2(+) versus HER2(−)breast cancers. These data named L1, L2, L3, L4, and L5 contained 22,14 32,15 3,16 38,17 and 6218 miRNAs, respectively (Table 1). This study data ranged from 2006 (L5) to 2012 (L1). The sizes of these five data sets ranged from 3 miRNA to 62 miRNAs.
Table 1.
Five Data Base of MiRNAs Differentially Expressed in HER2(+) Versus HER2(−) Breast Cancers
| Array | Differentially expressed miRNAs |
|---|---|
| L1 | mir-18, mir-21, mir-23a, mir-26a, mir-26b, mir-28-5p, mir-29b, mir-98, mir-100, mir-125a-3p, mir-129-5p, mir-149, mir-187, mir-195, mir-210, mir-425, mir-494, mir-663, mir-720, mir-1280, mir-1908 |
| L2 | mir-148a, mir-221, mir-222, mir-23a, mir-27a, mir-29a, mir-34a, mir-125b.mir-138, mir-141, mir-146a, mir-185, mir-190, mir-191, mir-196a, mir-200a.mir-200b, mir-200c, mir-203, mir-208, mir-224, mir-363, mir-367, mir-375, mir-377, mir-429, mir-497, mir-503, mir-529, mir-544, let-7b, let-7c |
| L3 | mir-181c, mir-302c, mir-376b |
| L4 | mir-10a, mir-10b, mir-15b, mir-17-5p, mir-18a, mir-20a, mir-25, mir-29b, mir-31, mir-93, mir-99a, mir-100, mir-103, mir-106a, mir-106b, mir-107, mir-126, mir-130a, mir-135b, mir-142-3p, mir-142-5p, mir-145, mir-149, mir-150, mir-152, mir-155, mir-181b, mir-199a, mir-199a*, mir-200a, mir-214, mir-342, mir-382, mir-505, mir-let-7a, mir-let-7c, mir-let-7f |
| L5 | mir-10b, mir-126*, mir-148a, mir-15, mir-16, mir-19a, mir-19b, mir-21, mir-221, mir-222, mir-22, mir-24, mir-25, mir-26a, mir-26b, mir-27a, mir-27b, mir-29a, mir-29c, mir-30a-5p, mir-30b, mir-30c, mir-30e, mir-30d, mir-34a, mir-34b, mir-92, mir-93, mir-95, mir-99a, mir-99b, mir-100, mir-106a, mir-106b, mir-107, mir-125a, mir-125b, mir-126, mir-141, mir-142-5p, mir-143, mir-145, mir-150, mir-154, mir-182, mir-191, mir-199a, mir-199a*, mir-199b, mir-200b, mir-205, mir-210, mir-224, mir-324-5p, mir-330, mir-342, mir-let-7a, mir-let-7b, mir-let-7c, mir-let-7f, mir-let-7g, mir-let-7i |
Indicated complementary strand of miRNA.
The set of miRNA target genes in breast cancer tissue
miRNAs exert their regulatory effects by binding to imperfect complementary sites within the 3′ untranslated regions (UTRs) of their mRNA targets, and repress target gene expression post-transcriptionally.1 Each may have hundreds and even more target genes, while one gene could be regulated by several miRNA. To avoid too many target genes, we filtered out the specific expression of target genes in breast cancer tissue with the use of the “tissue-specific expression enrichment organization” module in the DAVID database. Finally, the target genes of these five sets were 5,734; 22,409; 1,142; 22,293; and 43,460 in sequence. Here, we did not list every gene.
The GO Function and signaling analysis of target genes
Classification analysis of GO BP and GO MF
We used the GO BP and GO MF module of DAVID database to analyze these target genes. And the numbers of BPs using the GO BP classification in the five set were 1,724; 1,955; 899; 1,391; and 1,939. The numbers of MF using GO MF classification in the five sets were 347; 1,967; 160; 283; and 316. Comparing all the results of GO BP and GO MF analysis, we listed the major processes (Figs. 2 and 3). The target genes related to HER2 status may participate in important processes of GO BP classification, including angiogenesis, apoptosis, antiapoptosis, death, female gonad development, protein transport, and response to hormone stimulus. The GO MF classification related to HER2 status covers ATP binding, DNA binding, phosphate activity, protein binding, and transcription factor activity. This result of our study shows that HER2 may play an important role in these BPs.
FIG. 2.
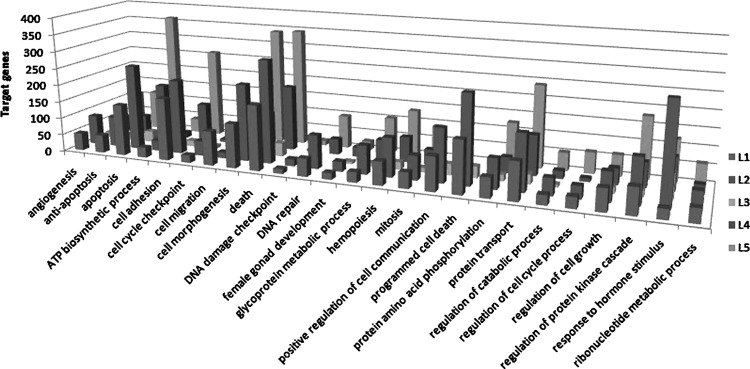
Parts of GO BP categories of different miRNA expressed in HER2 status.
FIG. 3.
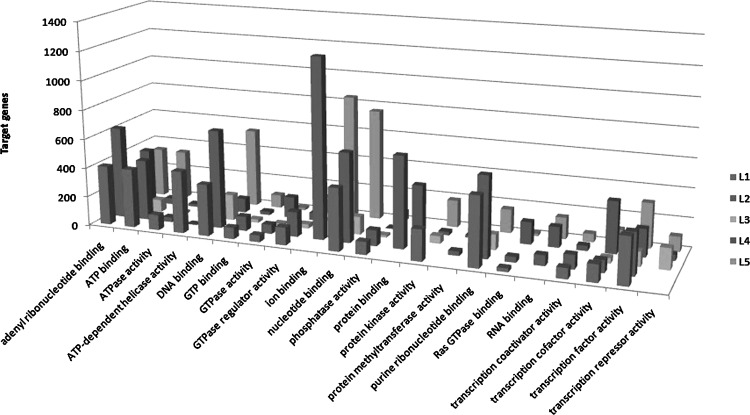
Parts of GO MF categories of different miRNA expressed in HER2 status.
Analysis of KEGG signaling pathways
We used the KEGG pathway to target gene function and conducted pathway analysis. We set the threshold at p≤0.05. The number of KEGG signaling pathways of L1, L2, L3, L4, and L5 were 38; 156; 51; 161; and 46; respectively. By merging these data, we identified 13 common signaling pathways (Table 2), such as Wnt signaling pathway, VEGF signaling pathway, natural killer cell-mediated cytotoxicity, T cell receptor signaling pathway, B cell receptor signaling pathway, colorectal cancer, endometrial cancer, prostate cancer, and nonsmall cell lung cancer.
Table 2.
The Major KEGG Pathways Related to MiRNAs Differentially Expressed in HER2(+) Versus HER2(−) Breast Cancers
| KEGG_pathways_term | KEGG number |
|---|---|
| Wnt signaling pathway | hsa04310 |
| VEGF signaling pathway | hsa04370 |
| Natural killer cell mediated cytotoxicity | hsa04650 |
| T cell receptor signaling pathway | hsa04660 |
| B cell receptor signaling pathway | hsa04662 |
| Colorectal cancer | hsa05210 |
| Endometrial cancer | hsa05213 |
| Prostate cancer | hsa05215 |
| Nonsmall cell lung cancer | hsa05223 |
| Hypertrophic cardiomyopathy | hsa05410 |
| Arrhythmogenic right ventricular cardiomyopathy | hsa05412 |
| Dilated cardiomyopathy | hsa05414 |
KEGG, Kyoto Encyclopedia of Genes and Genomes.
Analysis of BIOCARTA metabolic signaling pathways
We used the BIOCARTA pathway analysis to identify gene function. We set the threshold at p≤0.05. As shown in the Table 3, the number of BIOCARTA metabolic signaling pathways of L1, L2, L3, L4, and L5 sets were 99; 102; 7; 74; and 32; respectively. These metabolic signaling pathways contain PTEN-dependent cell cycle arrest and apoptosis, Angiotensin II mediated activation of JNK pathway via Pyk2-dependent signaling, role of ERBB2 in signal transduction and oncology, MAPKinase signaling pathway, AKT signaling pathway and Erk1/Erk2 MAPK signaling pathway and TNF/Stress-related signaling. These pathways have been shown in breast cancer.19,20
Table 3.
The Major BIOCARTA Pathways Related to MiRNAs Differentially Expressed in HER2(+) Versus HER2(−) Breast Cancers
| BIOCARTA_pathways_term | miRNAs data sets |
|---|---|
| PTEN-dependent cell cycle arrest and apoptosis | L1, L4, L5 |
| Angiotensin II mediated activation of JNK Pathway via Pyk2-dependent signaling | L2, L4 |
| Regulation of eIF4e and p70 S6 kinase | L1, L2 |
| Role of ERBB2 in signal transduction and oncology | L1, L2, L4, L5 |
| MAPKinase signaling pathway | L1, L4 |
| EGF signaling pathway | L1, L2, L4 |
| AKT signaling pathway | L1, L4, L5 |
| Erk1/Erk2 Mapk signaling pathway | L1, L2, L4, L5 |
| TNF/stress-related signaling | L1 |
Discussion
In this study, we summarized 5 sets of miRNAs specially expressed in breast cancer with the GEO and ArrayExpress databases. Then, by the target gene software of TargetScan and PicTar, we got target genes of those HER2-related miRNAs. Through the enrichment and function analysis of DAVID database, we got that these target genes of HER2 relevant miRNAs may participate in many important BPs, including angiogenesis, apoptosis, antiapoptosis, death, female gonad development, protein transport, and response to hormone stimulus. Furthermore, these target genes also take part in many important signaling pathways, such as the Wnt signaling pathway, VEGF signaling pathway, MAPKinase signaling pathway, EGF signaling pathway, and AKT signaling pathway. Although therapeutic targeted HER2 benefit numbers of HER2-overexpressed breast cancer patients, there is still a problem of drug resistance. With bioinformatics analysis of miRNAs signatures for differentially expressed in HER2(+) versus HER2(−) breast cancers, our study found miRNAs signatures related to HER2 status in breast cancers and some potential biological function of HER2. Then, it might provide further insight into breast cancer therapy.
HER2 is one member of the ErbB receptor family. The major pathways associated with ErbB signaling include the Ras/MAPK pathway, the phosphatidylinositol 3-kinase (PI3K/Akt) pathway, the Janus kinase (JAK)/signal transducers and activators of transcription factor (STAT) pathway, and the phospholipase Cγ pathway. These signaling pathways ultimately affect the important BP of cell proliferation, survival, motility, and adhesion.21 HER2 overexpression could lead to malignant transformation of cells and induce tumor necrosis resistance. Moreover, it is involved in the inhibition of apoptosis and promotes the survival of tumor cells.22 It is reported that HER2 overexpression may lead to ligand-independent receptor dimerization and subsequent upregulation of EGFR/HER2 signaling pathways.23 In this way, HER2 overexpression cause many cancers, including breast cancer. Hence, it is a matter of great urgency to make a comprehensive study of the biological functions of HER2.
miRNAs, noncoding single-stranded RNAs of 21∼22 nucleotides, constitute a novel class of gene regulators in both plants and animals. miRNAs exert their regulatory effects by binding to imperfect complementary sites within the 3′UTRs of their mRNA targets, and then repress target gene expression post-transcriptionally, Therefore, miRNAs have widespread impact on both the expression and evolution of protein-coding genes. In this study, we predict the miRNA differentially expressed in HER2(+) versus HER2(−) breast cancers so as to identify miRNAs associated with the HER2 receptor.
So far, the method of experimental verification cannot meet the comprehensive understanding of the function of miRNAs in the system level. There is a great need for further studies of miRNA to validate the target genes that targeted as much as possible. The developments of bioinformatics offer the possibility of faster miRNAs. By the bioinformatics method, the key points of the analysis of differentially expressed miRNAs lie in the sensitivity and specificity of miRNAs prediction software. Single target gene prediction software cannot achieve the desired prediction efficiency, and the performance of false-positive and false-negative predictions are more obvious.24 In this article, we make use of TargetScan and the PicTar, two recognized prediction softwares, to take the predicted results and to exclude false-negative predictions. The current predictions by TargetScan and PicTar have a high degree of overlap because they now all require stringent seed pairing. However, they are not 100% identical. Some reasons can be traced to alignment artifacts, the use of slightly different UTR databases, or the use of different miRNA sequences. Other reasons are intrinsic to the prediction algorithms themselves, such as the treatment of the target nucleotide opposite the first miRNA nucleotide.25 And they all have room for improvement. However, the accuracy of the computational prediction is increasingly accepted.
miRNAs have been shown to control a wide range of biological functions, such as cellular proliferation, differentiation, and apoptosis. Expression profiling of miRNAs has been an accurate method of classifying cancer subtypes than using the expression profiles of protein-coding genes.26 The differential expression of certain miRNAs in various tumors might become a powerful tool to aid in the diagnosis and treatment of cancer. Gene therapies that use miRNAs will be an effective approach to blocking tumor progression. In this study, we aim to summarize the signatures of miRNAs differentially expressed in HER2(+) versus HER2(−) breast cancers and find potential therapeutic targets. In the future, we will carry on the bioinformatics analysis and other computational methods to identify the regulatory roles of miRNAs in the HER2 status. Finally, we expect to use the miRNA to classify the HER2 status of the breast tumor samples and could identify novel targets for therapeutic manipulation.
Acknowledgments
This project is supported by grants from National Natural Science Foundation of China (Nos. 81272252, 81141094, and 81071261), Natural Science Foundation of Jiangsu Province (BK2011656). The authors thank Dr. Walter Pagel from the Department of Scientific Publications, MD Anderson Cancer Center for the language editing.
Disclosure Statement
No competing financial interests exist.
References
- 1.Mendell JT. Olson EN. MicroRNAs in stress signaling and human disease. Cell. 2012;148:1172. doi: 10.1016/j.cell.2012.02.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Ambros V. The functions of manimal miRNAs. Nature. 2004;431:350. doi: 10.1038/nature02871. [DOI] [PubMed] [Google Scholar]
- 3.Esquela-Kerscher A. Slack FJ. Oncomirs—miRNAs with a role in cancer. Nat Rev Cancer. 2006;6:259. doi: 10.1038/nrc1840. [DOI] [PubMed] [Google Scholar]
- 4.Calin GA. Croce CM. MiRNA signatures in human cancers. Nat Rev Cancer. 2006;6:857. doi: 10.1038/nrc1997. [DOI] [PubMed] [Google Scholar]
- 5.Siegel R. Ward E. Brawley O. Jemal A. The impact of eliminating socioeconomic and racial disparities on premature cancer deaths. CA Cancer J Clin. 2011;61:212. doi: 10.3322/caac.20121. [DOI] [PubMed] [Google Scholar]
- 6.Yarden Y. Baselga J. Miles D. Molecular approach to breast cancer treatment. Semin Oncol. 2004;315:6. doi: 10.1053/j.seminoncol.2004.07.016. [DOI] [PubMed] [Google Scholar]
- 7.Zhang ZJ. Ma SL. miRNAs in breast cancer tumorigenesis. Oncol Rep. 2012;27:903. doi: 10.3892/or.2011.1611. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Bao W. Fu HJ. Xie QS, et al. HER2 interacts with CD44 to up-regulate CXCR4 via epigenetic silencing of miRNA-139 in gastric cancer cells. Gastroenterology. 2011;141:2076. doi: 10.1053/j.gastro.2011.08.050. [DOI] [PubMed] [Google Scholar]
- 9.Barrett T. Troup DB. Wilhite SE, et al. NCBI GEO: Archive for functional genomics data sets? A10 years on. Nucleic Acids Res. 2011;39:1005. doi: 10.1093/nar/gkq1184. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Parkinson H. Kapushesky M. Kolesnikov N, et al. ArrayExpress update—from an archive of functional genomics experiments to the atlas of gene expression. Nucleic Acids Res. 2009;37:868. doi: 10.1093/nar/gkn889. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Vergoulis T. Vlachos IS. Alexiou P, et al. TarBase 6.0: Capturing the exponential growth of miRNA targets with experimental support. Nucleic Acids Res. 2012;40:222. doi: 10.1093/nar/gkr1161. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.John B. Sander C. Marks DS. Prediction of human microRNA targets. Methods Mol Biol. 2006;342:101. doi: 10.1385/1-59745-123-1:101. [DOI] [PubMed] [Google Scholar]
- 13.Sherman BT. Huang da W. Tan Q, et al. DAVID Knowledgebase: A gene-centered database integrating heterogeneous gene annotation resources to facilitate high-throughput gene functional analysis. BMC Bioinform. 2007;8:426. doi: 10.1186/1471-2105-8-426. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Ichikawa T. Sato F. Terasawa K, et al. Trastuzumab produces therapeutic actions by upregulating miR-26a and miR-30b in breast cancer cells. PLoS One. 2012;7:31422. doi: 10.1371/journal.pone.0031422. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Mackiewicz M. Huppi K. Pitt JJ, et al. Identification of the receptor tyrosine kinase AXL in breast cancer as a target for the human miR-34a miRNA. Breast Cancer Res Treat. 2011;130:1. doi: 10.1007/s10549-011-1690-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Lowery AJ. Miller N. Devaney A, et al. MiRNA signatures predict oestrogen receptor, progesterone receptor and HER2/neu receptor status in breast cancer. Breast Cancer Res. 2009;11:27. doi: 10.1186/bcr2257. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Blenkiron C. Goldstein LD. Thorne NP, et al. MiRNA expression profiling of human breast cancer identifies new markers of tumor subtype. Genome Biol. 2007;8:214. doi: 10.1186/gb-2007-8-10-r214. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Mattie MD. Benz CC. Bowers J, et al. Optimized high-throughput miRNA expression profiling provides novel biomarker assessment of clinical prostate and breast cancer biopsies. Mol Cancer. 2006;5:24. doi: 10.1186/1476-4598-5-24. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Cellurale C. Girnius N. Jiang F, et al. Role of JNK in mammary gland development and breast cancer. Cancer Res. 2012;72:472. doi: 10.1158/0008-5472.CAN-11-1628. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Chinnaiyan AM. Prasad U. Shankar S, et al. Combined effect of tumor necrosis factor-related apoptosis-inducing ligand and ionizing radiation in breast cancer therapy. Proc Natl Acad Sci U S A. 2000;97:1754. doi: 10.1073/pnas.030545097. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Saxena R. Dwivedi A. ErbB family receptor inhibitors as therapeutic agents in breast cancer: Current status and future clinical perspective. Med Res Rev. 2012;32:166. doi: 10.1002/med.20209. [DOI] [PubMed] [Google Scholar]
- 22.Sahin AA. Biologic and clinical significance of HER2/neu(cerbB 2)in breast cancer. Adv Anat Pathol. 2000;7:158. doi: 10.1097/00125480-200007030-00005. [DOI] [PubMed] [Google Scholar]
- 23.Lynch TJ. Bell DW. Sordella R, et al. Activating mutations in the epidermal growth factor receptor underlying responsiveness of non-small-cell lung cancer to Gefitinib. N Engl J Med. 2004;350:2129. doi: 10.1056/NEJMoa040938. [DOI] [PubMed] [Google Scholar]
- 24.Bartel DP. MiRNAs: Target recognition and regulatory functions. Cell. 2009;136:215. doi: 10.1016/j.cell.2009.01.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.David PB. MicroRNAs: Target recognition and regulatory functions. Cell. 2009;136:215. doi: 10.1016/j.cell.2009.01.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Aurora EK. Frank JS. Oncomirs—microRNAs with a role in cancer. Nature. 2006;6:259. doi: 10.1038/nrc1840. [DOI] [PubMed] [Google Scholar]


