Abstract
Costimulatory molecules of the CD28 family on T lymphocytes integrate cues from innate immune system sensors and modulate activation responses in conventional CD4+ T cells (Tconv) and their FoxP3+ regulatory counterparts (Treg). To better understand how costimulatory and coinhibitory signals might be integrated, we profiled the changes in gene expression elicited in the hours and days after engagement of Treg and Tconv by anti-CD3 and either anti-CD28, -CTLA4, -ICOS, -PD1, -BLA, or -CD80. In Tconv, a shared “main response” was induced by CD28, ICOS, and, surprisingly, BTLA and CD80, with very limited CD28-specific (primarily Il2) or ICOS-specific elements (including Th1 and Th2 but not the follicular T signature). CTLA4 and PD1 had a very subtle impact in this system, similarly inhibiting the response to anti-CD3. Treg responded to the same costimulatory hierarchy and to the same extent as Tconv, but inducing different clusters of genes. In this reductionist system, costimulatory or coinhibitory engagement mainly elicits generic responses, suggesting that the variability of their effects in vivo result from temporal or anatomical differences in their engagement, rather than from inherently different wiring.
Keywords: expression profiling, immunoregulation, lymphocyte differentiation
Costimulatory molecules of the CD28 family play a crucial role in the genesis of an effective and controlled response in T lymphocytes, complementing and modulating the interaction of TCR with MHC-peptide complex. They have attracted great interest because of their profound influence, and because they offer the possibility to therapeutically tune or deviate lymphocyte activation in autoimmune and tumor contexts (1, 2). Five molecules of the CD28 family are expressed at different stages of differentiation of T lymphocytes: CD28, CTLA4, ICOS, PD1, BTLA (3); in addition, CD80, the ligand for CD28 and CTLA4 normally present on APCs, is also expressed on T cells (4), as is PDL1. The surface expression of CD28 family molecules changes dynamically during the course of T-cell activation and effector cell differentiation, serving as feedback control on activation. In current views of costimulation, each of these molecules has a different personality:
CD28 is considered the primary costimulatory molecule (hereafter “costim” for short), through its interactions with CD80 and CD86 on APCs, complement TCR-mediated signals and promote T-cell activation, proliferation and survival. Initial activation of naïve T cells requires CD28 signal to enhance the signal derived by TCR and the production of IL2 (5) for the robust expansion of antigen-specific T cells. CD28 signaling is thought to proceed via the PI3K/Akt/mTOR pathway, hence activating the cell’s energetic and anabolic metabolism (6, 7).
CTLA4 also interacts with CD80 and CD86, but has an inhibitory effect, as evidenced by the extreme phenotype of CTLA4-deficient mice. The mechanism through which CTLA4 delivers coinhibitory signals is still debated, and several non-mutually-exclusive mechanisms have been proposed (7, 8). It may interfere with TCR or CD28 downstream signaling pathways, for instance by activating SHP2 and PP2A phosphatases, perturb the supramolecular organization of TCR-CD28 in microclusters, outcompete CD28 for CD80/86 ligands for which it has intrinsically higher affinity, or strip CD80/86 from APCs through transendocytosis. Adding complexity, CTLA4 affects both Treg and Tconv cells, which may balance each other. PD1, which binds PDL1 and PDL2, also behaves as an inhibitory molecule. Like CTLA4, it is induced upon T-cell activation (also in B and myeloid cells). Its inhibitory pathways have been proposed to be different from those of CTLA4 (9).
ICOS, through specific interaction with ICOSL, has more nuanced effects. Unlike CD28, ICOS does not impact Grb2 and NF-κB signaling, but it does activate the PI3k-Akt pathway and can deliver positive costimulatory signals. More specifically, ICOS promotes T-cell differentiation toward B-cell help, favoring expression of IL4 and the T “follicular helper” phenotype (10).
BTLA is expressed on activated T cells, especially Th1 cells (11). It is unusual in that its known ligand, HVEM, belongs not to the B7 but to the TNFR-family member (and itself engages in complex interactions with several ligands). BTLA is generally viewed as a coinhibitory molecule, in connection with the intracytoplasmic ITIM motif it shares with CTLA4 and PD1, which can recruit inhibitory SHP1/2. On the other hand, BTLA interacts and activates Grb2 and PI3K (12), indicating it may also have prosurvival activity (13).
An additional twist to this complexity came from the demonstration that CD80, the canonical costimulatory ligand expressed by antigen-presenting cells (APC), is also expressed in T cells, binding PD-L1 on APCs and stromal cells, delivering inhibitory signals and contributing to immune tolerance (14).
CD28 family molecules are also expressed on FoxP3+ Treg cells, key regulators of lymphoid homeostasis. CTLA4, ICOS, and PD1 are overexpressed on Tregs relative to conventional CD4+ T cells. CD28 and CTLA4 control Treg differentiation and homeostasis, and there is strong evidence that CTLA4 and PD1 are important players in Treg function (15). It is not clear, however, whether the function of costims on Tregs is merely a facet of their usual activity in Tconv, or whether they trigger distinct signals in Tregs.
It is reasonable to assume that these diverse influences should have distinct footprints in the T-cell transcriptome. Application of microarray profiling indicated that engagement of CD28 amplified the response elicited by CD3 alone (16, 17), that CD28 and ICOS induced comparable alterations in the transcriptome of Jurkat and primary CD4+ T cells, CTLA4 engagement blocking some of those changes. In another study, PD1 engagement blocked CD3/CD28-induced changes, as did CTLA4, but somewhat differently (18).
We revisit here these transcriptional analyses. We tested the transcriptional consequences in CD4+ Treg and Tconv cells of engaging every costim of the CD28 family (CD28, CTLA4, PD1, ICOS, BTLA and CD80), focusing on the initial events; this timeframe should better reflect direct signaling events than the integrated response that unfolds after a few days. Many of the costims generically elicited a positive amplification of a large fraction [but not all] of the response induced by TCR engagement, even for costims usually thought to be inhibitory like BTLA and CD80. The existence of direct but subtle inhibition by both CTLA4 and PD1 engagement was confirmed, and unique effects of ICOS engagement highlighted. Overall, the response differed markedly in Treg and Tconv cells.
Results
Our strategy was willfully reductionist. CD4+ T cells purified by magnetic negative selection were stimulated in vitro, to avoid indirect effects on APCs and other in vivo confounders. To analyze responses under identical conditions, we did not prefractionate Treg and Tconv cells but purified them at the end of the culture period, accepting the caveat that some of the late results may reflect in part the indirect influence of the other cell-type. Cognizant that affinities of specific antibodies differ from those of natural costim ligands, we nevertheless opted to use monoclonal antibodies (mAbs) immobilized on beads (“artificial APCs”) to observe pure effects of costim signaling, not confounded by multiply-reactive ligands. The amounts of anti-CD3 and anti-CD28 mAbs used for conjugation were picked as subsaturating in preliminary experiments (Fig. S1), a higher dose being used for mAbs to other costims.
We assessed the ability of these sets of artificial APCs, carrying anti-CD3 alone or together with anti-costim mAbs, to activate T cells to a full proliferative response. As expected, anti-CD3 alone elicited little proliferation, anti-CD28 was the most effective costimulator for both Tconv and Treg cells (Fig. 1). Only anti-ICOS had a significant costimulatory effect, and this only in Treg cells. In contrast, anti-CTLA4, -PD1, -BTLA, and -CD80 all seemed to have inhibitory activity relative to anti-CD3 alone, most clearly for Treg cells.
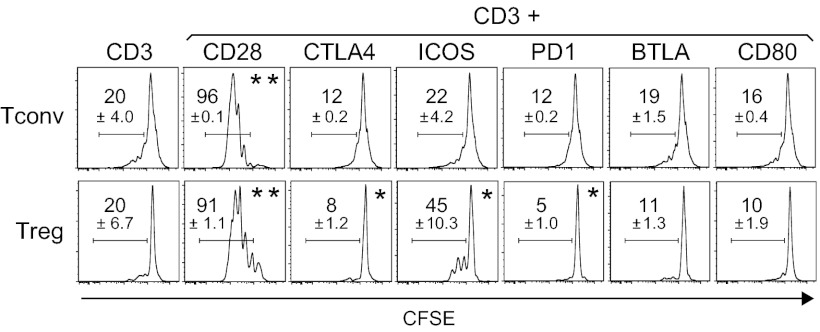
Fig. 1.
Impact of costim engagement on T-cell proliferation. CD4+ splenocytes were labeled with CFSE, and stimulated for 66 h in vitro with beads conjugated with anti-CD3 alone or with anti-costim mAbs, and cell division assessed by flow cytometry measure of residual CFSE in CD4+CD3+ cells. *P < 0.05; **P < 0.005 from anti-CD3 alone.
Overall Comparison of Costimulatory Molecules’ Footprints.
Gene expression profiles were then generated in three independent experiments (the experiments were highly concordant overall, and for simplicity the mean of all replicates are used). Cells were harvested after 1, 4, 20, and 48 h of stimulation; the 1- and 4-h lysates were pooled before RNA purification and profiling (“Early” pool), as were the 20- and 48-h samples (“Late”). The former encompassed immediate-early and early changes, the latter events in the amplified response (for Treg cells, only the 20-h sample was used because significant cell death occurred after 1 d of culture). As a first step in the analysis, we simply counted how many transcripts changed by a fold change > 2 (corresponding to an FDR of 0.05 or better in most cases) in Tconv or Treg cells (Table 1). The response to CD3 stimulation alone was truncated, greater at early than at late times, in both of Tconv and Treg. In contrast, engagement of activating costim expanded the response over time. Only apparent for CD28 and ICOS at early times (ICOS strongest), the response became much more prominent later for CD28, ICOS, and to a lesser extent BTLA and CD80. CTLA4 and PD1 seemed to have no effect, at least with this simple metric (see below).
Table 1.
Magnitude of changes elicited by triggering of CD3 and individual costims
| CD3+ |
|||||||
| CD3 | CD28 | CTLA4 | ICOS | PD1 | BTLA | CD80 | |
| Early | |||||||
| Tconv | 212/53 | 14/0 | 0/0 | 37/0 | 0/0 | 1/0 | 2/0 |
| Treg | 249/44 | 11/0 | 1/0 | 14/39 | 0/0 | 0/0 | 0/0 |
| Late | |||||||
| Tconv | 142/133 | 566/127 | 0/0 | 188/17 | 0/0 | 65/2 | 5/2 |
| Treg | 107/63 | 450/37 | 2/0 | 296/89 | 1/0 | 7/0 | 3/0 |
Changes were estimated by counting the number of transcripts changing after engagement of CD3 alone (left column) or with individual costims, in sorted Tconv or Treg cells, at early or late times. For each condition, the two numbers denote transcripts >twofold induced and repressed, respectively.
A general perspective of the results is presented in Fig. 2A, which displays changes effected by each costim relative to anti-CD3 alone (for genes affected in either Treg or Tconv cells; all changes listed in Datasets S1 and S2). Several conclusions are already apparent here: First, there is a shared “costimulation cluster” in Tconv, affected by both CD28 and ICOS and more weakly by BTLA and CD80. The relative impact of CD28 and ICOS is subtly different, however. As expected from Table 1, ICOS is more active early, whereas CD28 dominates at late times. Second, CTLA4 and PD1 again appear inert in this representation, reflected as quasi-uniform black streaks. Third, the response in Tregs partially overlaps that of Tconv, particularly at early times, but also contains many distinct elements (examined below, we first focus on the Tconv response).
Fig. 2.

Transcriptional impact of costim coengagement . CD4+ splenocytes from B6.Foxp3fgfp mice were stimulated in vitro with bead-conjugated anti-CD3 alone or with anti-costim antibodies, and CD4+CD3+GFP− Tconv and GFP+ Treg were sorted for gene expression profiling. (A) Heatmap representation of the ratio of expression levels for each costim coengagement relative to CD3 alone, for Tconv and Treg cells (for transcripts that change by > twofold with CD3 or any of the costims); order by hierarchical clustering. (B) Fold change/fold change (FC/FC) plot comparing the effect in Tconv cells of each costim (y axis) to that of CD28 (x axis).
The fold change/fold change plots of Fig. 2B present a more direct comparison of the effects in Tconv cells of CD28 vs. all other costims. Here, it becomes clear, unexpectedly with regards to prior literature, that BTLA and CD80 essentially behave as weaker surrogates of CD28, as indicated by the high degree of correlation with CD28’s effects, at both early and late times; the off-diagonal placement of the gene clouds indicates that their impact is weaker, in accord with the heatmap (Fig. 2A). This CD28-like effect is also true for ICOS, with perhaps more distinctions relative to CD28 (see below). Here again, CTLA4 and PD1 seem to have little or no impact on the transcripts costimulated by CD28.
Comparative Analysis of CD28 and ICOS.
In earlier studies of human T cells using printed arrays, Riley et al. (17) reported similar effects of ICOS and CD28, with only a few differently affected genes such as IL2 and IL9. Consistent with these observations, we found the effects of ICOS engagement to be comparable to those of CD28, the off-diagonal placement denoting a stronger effect of CD28. However, a number of transcripts responded differently, as particularly marked in the late stages (Fig. 3A; Fig. S2 for early time). Both sets included cell cycle and proliferation genes, but anti-ICOS preferentially enhanced transcripts associated with T-cell differentiation, including Th1-type transcripts such as Eomes and Ifng but also Th2-type molecules such as Gata3 and Ccr8 (but not Il4). On the other hand, genes that stood out as preferentially or uniquely activated by CD28 included factors more generically involved in T-cell proliferation and survival such as Il2 (as noted in ref. 17), the apoptosis/survival controllers Bnip3 or Anxa2, or metabolism control factors such as Hk2 or Ak3. Indeed, the lasting induction of Il2 was unique to CD28 (Table S1).
Fig. 3.
Subtle differences between CD28 and ICOS engagement on activated Tconv. (A). FC/FC plot comparing the late effect of CD28 (x axis) and ICOS (y axis) over CD3 engagement alone. Red and blue dots denote genes preferentially responding to CD28 or ICOS, respectively. (B) Same plot, highlighting genes over- or under-expressed in different Th signatures (numbers along the axes indicate the number of signature genes over- or underexpressed in response to the corresponding costim). Genes in the Th1/2/17 signatures per ref. 24; the Tfh signature per ref. 25.
Studies in knockout mice have shown that ICOS is required for optimal production of differentiated cytokines of the Th2 and Th1 types, and plays a key role in the differentiation of follicular T cells in which it is overexpressed (10). To investigate this point at the genomic level, we highlighted transcripts characteristic of the differentiated Th signatures onto the CD28/ICOS comparison plot (Fig. 3B). Th1 and Th2 signature genes were clearly enhanced by costimulation relative to anti-CD3 alone: Overexpressed genes in these signatures (red) were induced, suppressed genes (green) were down-regulated. For the most part, CD28 and ICOS seemed equivalent in this respect. On the other hand, transcripts typical of Th17 and “follicular Th” were not influenced by either costim, perhaps contrary to expectation.
Thus, engagement of positively acting costims thus impacts, in both shared and specific manner, on the transcriptional program of activated T cells.
Costim Effects on the CD3-Induced Response.
The analyses above focused on the effects of costims in addition to anti-CD3 alone. We next investigated the effect of costim engagement on the changes induced by anti-CD3 alone, asking whether costims act as a generic amplifier (or dampener) of TCR signaling. In Fig. 4A, transcripts were ranked according to their repression or induction by anti-CD3, and the FoldChange [relative to unstimulated] resulting from the additional engagement of each costims was plotted for the most CD3-responsive genes (blue dots). As expected, CD28 and ICOS superactivated many of these TCR-induced genes (as did, to a lesser extent, BTLA and CD80), but the effect was clearly not uniform; indeed, some CD3-induced genes were largely unaffected by additional costim signals, as more clearly visualized in Fig. 4B and Fig. S3. For instance, the induction of CD25 (Il2ra), 4.1BB (Tnfrsf9), or Irf4 was not further accentuated by CD28 costimulation (a similar pattern was observed with ICOS, right).
Fig. 4.
Costim effects on TCR-induced responses. (A) Transcripts most strongly induced or repressed by CD3 engagement in Tconv cells were selected and ranked according to this ratio (black dots), and their change (relative to Unstimulated) in response to anti-CD3 plus each costim were plotted on the same scale (blue dots). (B) FC/FC plots comparing the response to CD3 alone vs. 3+CD28 (Left) or 3+ICOS (Right). (C) Inhibitory effects of CTLA4 or PD1: The effect of coinibitory engagement was calculated as a ratio of Fold changes [as ((CD3+CTLA4)/Unstim)/(CD3/Unstim)] and significance estimated as a t test. Transcripts with P value <0.05 for either CTLA4 or PD1 are shown.
Although CTLA4 and PD1 seemed largely inert in the comparisons described above, subtle but significant effects were detected here: The response of aCD3-activated genes was dampened by engagement of CTLA4 and PD1 (Fig. 4A), indicated by a displacement of the blue relative to the black dots, which was particularly clear for the early response. These effects were quantitated by computing the ratio of fold changes (Fig. 4C, legend), with significance estimated by a one-sample t test. Significant effects were strongly shared between CTLA4 and PD1 at early times (Fig. 4C Left) and more divergent at later times. Thus, CTLA4 had a direct dampening effect on the earliest consequences of TCR-mediated activation and PD1 did essentially the same.
Distinct Costimulatory Effects in Tconv and Treg Cells.
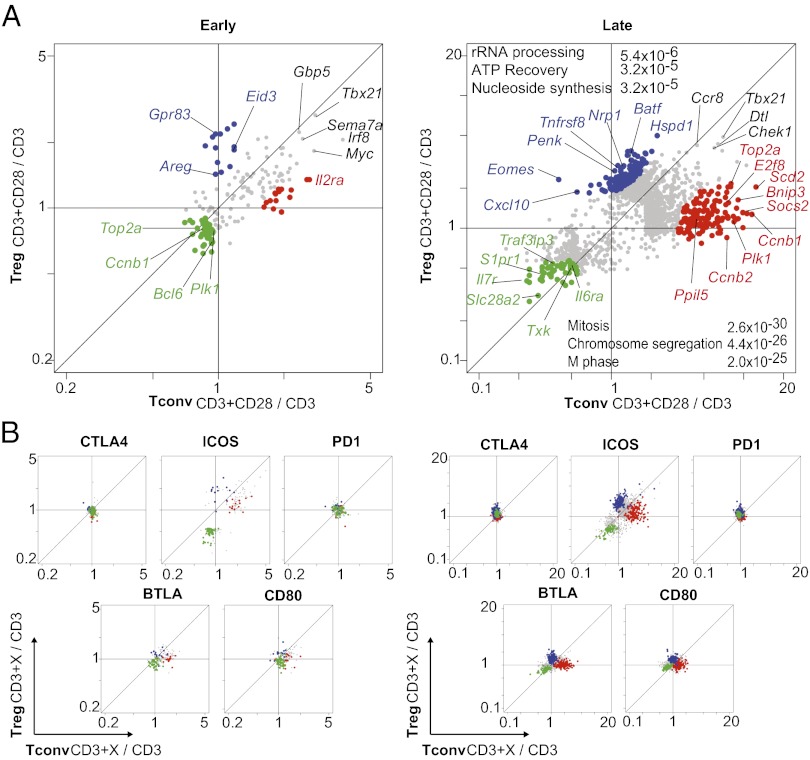
Do costimulatory molecules have the same footprints in Treg cells as in Tconv? The representations of Table 1 and Fig. 2A indicated some degree of sharing, but also distinct divergence. To directly address this question, we first compared the effects of CD28, which were the most robust. The displays of Fig. 5A compares the changes elicited by CD3+CD28 triggering relative to CD3 alone, in Tconv vs. Treg cells (x and y axes, respectively) at early and late times, and this for a set of genes selected as affected by any one of the costims. A clear demarcation was observed. Some of the response was shared by both cell types at early times (e.g., Tbx21 or Lif, gray dots), in particular for transcripts repressed in both cell types (green dots). However, and particularly as the response progressed at later times, many of the genes induced by CD28 showed a preferential response, quantitative or absolute, in one or the other cell type (blue or red dots, respectively). For instance, Gpr83, Areg, Tnfrsf8 (CD30), or Nrp1 were only enhanced in Treg. Conversely, many genes associated with cell cycle progression were preferentially induced late in Tconv (Ccnb2, Top2a). There was no particular overlap between these differential responses and the canonical “Treg signature,” genes that distinguish resting Treg and Tconv cells. Interestingly, the same differential activation in Treg and Tconv were largely reproduced by engagement of other positively acting costims, albeit in quantitatively different levels (ICOS, but also BTLA and CD80), as indicated by the placement of transcripts color-coded according to differential response to CD28 (Fig. 5B), which independently corroborate these distinctions. Although one might have expected stronger effects in light of their overexpression and function in Tregs, the effects of CTLA4 and PD1 were weak or inexistent there. We asked whether Treg cells would show a specific response to ICOS, as shown above for Tconv cells (Fig. 3A). Perhaps surprisingly, because ICOS is overexpressed in Tregs, ICOS-specific effects were fewer than earlier in Tconv (Fig. S4), although the characteristic preferential induction of Eomes, Eno3 and Xcl1 was also seen in Tregs.
Fig. 5.
Comparative effects of costims in Tconv and Treg cells. (A) Effect of CD28. The ratio of expression after CD3+CD28 engagement, relative to CD3 alone, was calculated in Tconv (x axis) and Treg (y axis) at early (Left) and late (Right) times. Genes induced specifically (>twofold differential) in Treg or Tconv are highlighted (blue and red, respectively); genes repressed in both are shown in green. (B) Disposition of the genes identified and color-coded in A, in plots that compare the effect of each costim in Tconv (x axis) vs. Treg (y axis) cells.
Discussion
Two dominant conclusions come forth from these data, along with several more specific or subtle subtexts: First, the existence of a dominant costimulation response that is shared by all positive costimulators (CD28, ICOS, CD80, and BTLA), but not CTLA4 or PD1. Second, a generic response is also observed in Tregs, but which is quantitatively quite different. The subtexts include the unexpected direction of CD80 and BTLA’s effects, that not all of the TCR-elicited response is susceptible to costimulation, and the subtle but significant inhibitory effects of direct CTLA4 and PD1 engagement.
At the onset, it is important to acknowledge the limitations of the present study. Using an in vitro activation system, activating cells with bead-bound mAbs without APCs, were motivated by the goal of measuring direct effects downstream of TCR and costim triggering to test basic differences in downstream wiring. Undoubtedly, using natural ligands of normal affinity and cross-reactivity, modulating responses with the interplay of several costimulatory effects in the context of a normal APC, and varying doses of triggering agents, would add additional layers of complexity to the responses. In addition, we could not perform dose–response studies (for obvious financial limitations) or use mAbs of graded affinity, such that conclusions concerning quantitative differences between effects of individual costims must be treated with some caution, potentially influenced by the affinity of the mAbs used.
Paradoxically, we observed effects from costims that are largely unrelated to their measurable levels at the cell surface. ICOS elicits a stronger early response in Tconv cells than CD28, where it is normally present at low levels and only induced secondarily; the subtle inhibitory effects of CTLA4 and PD1 are more marked at early times, even though these molecules are mainly expressed upon activation. This paradox may be related to the difficulty in evaluating the true “functional” presence of molecules on the cell surface, as opposed to the presence detectable by antibody staining and flow cytometry. With molecules that can have very rapid rates of endocytosis, such as the CD28 family, transient presence may be sufficient for activity.
There are several aspects to the “main response,” shared by all four positive costimulatory molecules: the same sets of genes are induced or repressed, at different times after activation (Fig. 2); the differential response in Treg and Tconv is reproducible with all four. This overall sharing was already observed in the early samples (1 and 4 h), thus unlikely to stem from indirect effects. The implication is that all four have similar wiring of downstream signals, and/or that they similarly impact on the molecular dynamics of the TCR in the signaling synapse. CD28 is recruited to the immunological synapse soon after initiation of activation, where it recruits Lck and PKCθ, thus enhancing TCR signals. Such physical interactions may also apply to ICOS, BTLA and CD80. Importantly, there was a distinct split in the TCR-induced transcriptional activation: Although most of the response to CD3 was further enhanced, a distinct gene cluster was refractory to additional costimulation (Fig. 4B). Might those TCR signals that result in induction of the Il2ra/Irf4 cluster be routed through a PKCθ independent pathway?
ICOS is known to be able to provide basic costimulatory support, but also has a unique input in vivo for the development of biased effector functions in CD4+ T cells into IL17-producing cells or Bcl6hiCXCR5+ T follicular helpers (19). Accordingly, in our analyses, ICOS was the only costim to have a signature clearly distinct from that of CD28, uniquely activating several transcripts and factors controlling differentiated effector functions (Ifng, Eomes, Gata3, Xcl1), albeit not with a marked skew on the corresponding signatures. IL21 was transiently induced in response to ICOS (Table S1). Thus, the full effect of ICOS may only be manifest when APCs or B cells are present to provide the natural ligand and/or to amplify the intrinsic signature of ICOS engagement.
We were surprised that BTLA4 and CD80, generally portrayed as coinhibitory molecules, actually activated the transcriptional main response. Both seemed to mimic CD28 very faithfully in all aspects, albeit at lower levels, with little or no specific effects as observed with ICOS. On the other hand, BTLA and CD80 did not activate proliferation in our assays (Fig. 1), and would appear inhibitory in proliferation-dependent contexts. This surprising transcriptional costimulation is unlikely to be an artifact from nonspecific cross-linking, because it was not observed with either CTLA4 or PD1. It may indicate a physical effect on the synapse and TCR microclusters mentioned above, but is also is consistent with the demonstrated binding of Grb2 and Pi3K to a membrane-proximal phosphotyrosine in the intracytoplasmic domain of BTLA, and some prosurvival effects of BTLA (12, 13). Activation of Pi3K may then dominate inhibitory signals elicited through the ITIM domains which BTLA shares with CTLA4 and PD1. In some contexts, blockade of the HVEM–BTLA axis did not have all of the negative consequences expected (11), and it may be that the coinhibitory classification of BTLA results from cross-competition, or coaggregation, with other HVEM ligands.
Many mechanisms have been proposed for the inhibitory actions of CTLA-4 and PD1, as presented above. Their dampening of the TCR response, in the minimalist experimental setting used here, were significant but very subtle. A difference from past studies in which stronger inhibitory effects were noted is that here they were tested alone, rather than in combination with CD28; hence, their major impact may be in dampening CD28’s action. There is probably some direct inhibition of TCR signals by CTLA4 and PD1 (from interference within the synapse and/or activation of inhibitory phosphatases via their ITIM motifs), but also grounds to believe that much of their action must involve other players (e.g., competition for, or down-regulation, of B7 family ligands on APCs). Importantly, and perhaps unexpectedly, PD1 and CTLA4 behaved very similarly in our assays.
Finally, another surprise was that the hierarchy of effects were similar in Tconv and Treg cells. Because the latter express significantly more CTLA4 and PD1, one might have expected a stronger response. A sizeable fraction of the responses were similar in the two cell-types, such as Tbx21 or Ccr8. On the other hand, the responses that unfolded were very distinct (Fig. 5). There is good evidence that TCR signaling pathways are activated in a quantitatively different manner in Treg cells, with lower recruitment of PKCθ (20–22), which must lead to the impressive differences in transcriptional induction observed here.
In conclusion, our observations both simplify the understanding of costimulatory molecules in T cells by highlighting generic cell-autonomous pathways, but also add additional complexity in several respects.
Materials and Methods
Mice.
Foxp3igfp knockin mice (23) were maintained in specific pathogen-free facilities at Harvard Medical School per Institutional Animal Care and Use Committee Protocol 02954.
Cell Activation and Purification.
Dynabeads M450 beads were coupled per manufacturer’s instructions with anti-CD3 (0.3 μg per 107 beads) alone or with anti-costimulatory molecules (0.3 μg of anti-CD28 clone 37.51; 1.2 μg for all others to maximize their effects: anti-CTLA4: 9H10; -PD1: 19G8; -ICOS: C398.4A; -BTLA: 6F7; -CD80: 1G10); all complemented to 2 μg of total antibody with nonbinding control IgG. CD4+ T cells from pooled spleen and lymph nodes of male mice were purified to >90% purity (Dynal CD4 Negative Isolation kit, Invitrogen), and 2 × 105 cells were cultured in 96-well flat-bottom plates in 200 μL of RPMI 1640, 10% (vol/vol) FCS, at a cell:bead ratio of 1:2. Cells were harvested at 1, 4, 20, or 48 h, Tconv and Treg were sorted as DAPI−CD45R−CD8a−CD11b/c− CD4+ and either GFP+ or GFP−, respectively, directly into TRIzol reagent. The 1- and 4-h, or 20- and 48-h, lysates were pooled as early or late time points, respectively (for Treg cells, the 20 h lysate only was used as the late sample). For proliferation assays, purified CD4+ T cells were labeled with 10 μM CFSE (Molecular Probes) at 107 cells per mL RPMI1640 for 20 min at 37 °C. After 66-h culture, proliferation was assessed by CFSE dilution.
Gene-Expression Analysis.
RNA was amplified, labeled, and hybridized to Mouse Gene 1.0 ST arrays with the data generation and quality control pipeline of the Immunological Genome Project (www.immgen.org), in biological triplicates (duplicates only for ICOS and CD80). Raw data were background-corrected and normalized using the RMA algorithm (Affymetrix PowerTools), and analyzed with GenePattern, S+, and Ingenuity Pathway Analysis software. Data were filtered for unannotated probes, for genes with no expression in any condition (EV < 120), and for probes giving discordant data between replicates (interreplicate CV > 0.7). Datasets have been deposited at Gene Expression Omnibus.
Supplementary Material
Acknowledgments
We thank Drs. A. Sharpe, G. Freeman, and V. Kuchroo for insightful discussions and K. Hattori and K. Rothamel for help with mice and microarrays. This work was supported by National Institutes of Health Grant P01-AI056299 (to D.M. and C.B.) and benefitted from data generated by the Immunological Genome Project.
Footnotes
The authors declare no conflict of interest.
Data deposition: The data reported in this paper have been deposited in the Gene Expression Omnibus (GEO) database, www.ncbi.nlm.nih.gov/geo (accession no. GSE42276).
This article contains supporting information online at www.pnas.org/lookup/suppl/doi:10.1073/pnas.1220688110/-/DCSupplemental.
References
- 1.Quezada SA, Peggs KS, Simpson TR, Allison JP. Shifting the equilibrium in cancer immunoediting: From tumor tolerance to eradication. Immunol Rev. 2011;241(1):104–118. doi: 10.1111/j.1600-065X.2011.01007.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Rosenblum MD, Gratz IK, Paw JS, Abbas AK. Treating human autoimmunity: Current practice and future prospects. Sci Transl Med. 2012;4:125sr1. doi: 10.1126/scitranslmed.3003504. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Sharpe AH, Freeman GJ. The B7-CD28 superfamily. Nat Rev Immunol. 2002;2(2):116–126. doi: 10.1038/nri727. [DOI] [PubMed] [Google Scholar]
- 4.Azuma M, Yssel H, Phillips JH, Spits H, Lanier LL. Functional expression of B7/BB1 on activated T lymphocytes. J Exp Med. 1993;177(3):845–850. doi: 10.1084/jem.177.3.845. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Thompson CB, et al. CD28 activation pathway regulates the production of multiple T-cell-derived lymphokines/cytokines. Proc Natl Acad Sci USA. 1989;86(4):1333–1337. doi: 10.1073/pnas.86.4.1333. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Pagès F, et al. Binding of phosphatidylinositol-3-OH kinase to CD28 is required for T-cell signalling. Nature. 1994;369(6478):327–329. doi: 10.1038/369327a0. [DOI] [PubMed] [Google Scholar]
- 7.Rudd CE, Taylor A, Schneider H. CD28 and CTLA-4 coreceptor expression and signal transduction. Immunol Rev. 2009;229(1):12–26. doi: 10.1111/j.1600-065X.2009.00770.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Walker LS, Sansom DM. The emerging role of CTLA4 as a cell-extrinsic regulator of T cell responses. Nat Rev Immunol. 2011;11(12):852–863. doi: 10.1038/nri3108. [DOI] [PubMed] [Google Scholar]
- 9.Riley JL. PD-1 signaling in primary T cells. Immunol Rev. 2009;229(1):114–125. doi: 10.1111/j.1600-065X.2009.00767.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Simpson TR, Quezada SA, Allison JP. Regulation of CD4 T cell activation and effector function by inducible costimulator (ICOS) Curr Opin Immunol. 2010;22(3):326–332. doi: 10.1016/j.coi.2010.01.001. [DOI] [PubMed] [Google Scholar]
- 11.Murphy TL, Murphy KM. Slow down and survive: Enigmatic immunoregulation by BTLA and HVEM. Annu Rev Immunol. 2010;28:389–411. doi: 10.1146/annurev-immunol-030409-101202. [DOI] [PubMed] [Google Scholar]
- 12.Gavrieli M, Murphy KM. Association of Grb-2 and PI3K p85 with phosphotyrosile peptides derived from BTLA. Biochem Biophys Res Commun. 2006;345(4):1440–1445. doi: 10.1016/j.bbrc.2006.05.036. [DOI] [PubMed] [Google Scholar]
- 13.Hurchla MA, Sedy JR, Murphy KM. Unexpected role of B and T lymphocyte attenuator in sustaining cell survival during chronic allostimulation. J Immunol. 2007;178(10):6073–6082. doi: 10.4049/jimmunol.178.10.6073. [DOI] [PubMed] [Google Scholar]
- 14.Butte MJ, Keir ME, Phamduy TB, Sharpe AH, Freeman GJ. Programmed death-1 ligand 1 interacts specifically with the B7-1 costimulatory molecule to inhibit T cell responses. Immunity. 2007;27(1):111–122. doi: 10.1016/j.immuni.2007.05.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Bour-Jordan H, Bluestone JA. Regulating the regulators: Costimulatory signals control the homeostasis and function of regulatory T cells. Immunol Rev. 2009;229(1):41–66. doi: 10.1111/j.1600-065X.2009.00775.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Diehn M, et al. Genomic expression programs and the integration of the CD28 costimulatory signal in T cell activation. Proc Natl Acad Sci USA. 2002;99(18):11796–11801. doi: 10.1073/pnas.092284399. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Riley JL, et al. Modulation of TCR-induced transcriptional profiles by ligation of CD28, ICOS, and CTLA-4 receptors. Proc Natl Acad Sci USA. 2002;99(18):11790–11795. doi: 10.1073/pnas.162359999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Parry RV, et al. CTLA-4 and PD-1 receptors inhibit T-cell activation by distinct mechanisms. Mol Cell Biol. 2005;25(21):9543–9553. doi: 10.1128/MCB.25.21.9543-9553.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Nurieva RI, Liu X, Dong C. Yin-Yang of costimulation: Crucial controls of immune tolerance and function. Immunol Rev. 2009;229(1):88–100. doi: 10.1111/j.1600-065X.2009.00769.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Hickman SP, Yang J, Thomas RM, Wells AD, Turka LA. Defective activation of protein kinase C and Ras-ERK pathways limits IL-2 production and proliferation by CD4+CD25+ regulatory T cells. J Immunol. 2006;177(4):2186–2194. doi: 10.4049/jimmunol.177.4.2186. [DOI] [PubMed] [Google Scholar]
- 21.Zanin-Zhorov A, et al. Protein kinase C-theta mediates negative feedback on regulatory T cell function. Science. 2010;328(5976):372–376. doi: 10.1126/science.1186068. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Tsang JY, et al. Altered proximal T cell receptor (TCR) signaling in human CD4+CD25+ regulatory T cells. J Leukoc Biol. 2006;80(1):145–151. doi: 10.1189/jlb.0605344. [DOI] [PubMed] [Google Scholar]
- 23.Bettelli E, et al. Reciprocal developmental pathways for the generation of pathogenic effector TH17 and regulatory T cells. Nature. 2006;441(7090):235–238. doi: 10.1038/nature04753. [DOI] [PubMed] [Google Scholar]
- 24.Wu HJ, et al. Gut-residing segmented filamentous bacteria drive autoimmune arthritis via T helper 17 cells. Immunity. 2010;32(6):815–827. doi: 10.1016/j.immuni.2010.06.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Yusuf I, et al. Germinal center T follicular helper cell IL-4 production is dependent on signaling lymphocytic activation molecule receptor (CD150) J Immunol. 2010;185(1):190–202. doi: 10.4049/jimmunol.0903505. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.