Abstract
Non-structural proteins (NSPs) based diagnostics are useful for large-scale sero-surveillance of foot-and-mouth disease (FMD) and to monitor viral activity as a follow up to the vaccination campaign in FMD endemic countries like India which aim at disease control through vaccination. These diagnostics are also handy in the serology of import/export of cloven-footed animals. In the present study, non-structural protein RNA polymerase (3D gene) of FMD virus (FMDV) was expressed using baculovirus expression system. Protein expression was analyzed by SDS-PAGE and confirmed by its immuno-reactivity with serum from a FMDV infected bovine, in the western blot. Recombinant 3D protein was purified and evaluated in the indirect ELISA with 1072 cattle serum samples. Diagnostic sensitivity and specificity of the assay were found to be 92 and 100 %, respectively, when tested with cattle sera of known FMD status. The 3D based ELISA developed here is useful for screening the animals as an adjunct to other NSP based diagnostics available for routine serosurveillance of FMD.
Keywords: Foot-and-mouth disease virus, Baculovirus, Non-structural protein, 3D, ELISA
Introduction
Foot-and-mouth disease (FMD) is considered a high priority disease in livestock around the world because of its economic implications. It is a highly contagious disease affecting cloven-hoofed animals comprising both domestic livestock and wild species. Though associated with limited fatality rates, the disease is notable because of its high rate of transmissibility and trade sanctions on the countries affected. The disease is controlled in endemic countries by means of effective and systematic vaccination programs, thorough sero-surveillance and vigorous stamping out policy, where possible.
The etiological agent, FMD virus (FMDV), belongs to the genus Aphthovirus in the family Picornaviridae. Its genome contains a single open reading frame and encodes a polyprotein, which is cleaved by viral protease 3C to yield four structural (VP1, VP2, VP3 and VP4) and ten nonstructural proteins (L, 2A, 2B, 2C, 3A, 3B1, 3B2, 3B3, 3C and 3D). The four structural proteins, VP1-4, are the major subunits of viral capsid, which induce neutralizing antibody response. The currently available inactivated vaccines mainly consist of capsid proteins [6] and the vaccinated animals are less likely to have antibodies to NSPs. Therefore, detection of antibodies to NSPs is the preferred diagnostic method to distinguish FMDV-infected animals from vaccinated animals. The most accurate testing protocol involves the identification of infected animals by a combination of sequential NSP detection assays. NSPs viz. 2C [13], 3A, 3B, 3AB, 3ABC and 3D [2] have been used by different investigators using either an enzyme-linked immunotransfer blot (EITB) [5, 4] or ELISA formats [5, 16, 23, 24]. However, there is still no international reference test (gold standard) for NSP based detection assays. There is a need to develop tests that may result in a higher sensitivity of detection assays for differentiation of infected from vaccinated animals. The OIE has proposed that Pan American Foot-and-Mouth Disease Center’s (PANAFTOSA) diagnostic system be adopted as the standard “reference test” for the diagnosis of FMD. This system combines the index screening system 3ABC ELISA with immunoblot test for antibodies against NSPs- 3A, 3B, 2C, 3D and 3ABC [3, 8, 25].
Among all NSPs, 3D protein is the most immunogenic and highly conserved. Antibodies to it persist for longer period of time in animal body after exposure to live virus and hence assays to detect antibodies to the 3D protein have been used for export/import testing, epidemiological surveys to determine the spread of FMDV in animal population and for identification of carrier animals [7, 11, 21, 22]. The present work was carried out with the objective of production of the purified recombinant 3D antigen of FMDV using baculovirus expression system and evaluation of its immuno-reactivity in indirect ELISA (iELISA).
Materials and Methods
Cells and Virus
BHK-21 cells were maintained in Glasgow’s modified Eagle’s Medium (SAFC Biosciences, USA) supplemented with 10 % fetal bovine serum. Asia1-IND63/72 strain of FMD vaccine virus was grown using BHK-21 cells. Sf-21 insect cells (Spodoptera frugiperda, IPLB-Sf21-AE clone, Invitrogen) were cultured in Sf-900 II SFM (Invitrogen) with added 10 % fetal bovine serum. High-five cells, TN5 (Trichoplusia ni, BTT-TN-5B1-4, Invitrogen) were cultured in Ex-Cell™ 405 SFM (SAFC Biosciences).
Primers and PCR
Virus was pelleted by centrifugation (Sorvall WX Ultra 80) of clarified virus at a speed of 100,000 g for 1 h over 10 % (w/v) sucrose cushion. Pellet was suspended in 100 μl of PBS from which viral RNA was extracted using RNeasy mini kit (Qiagen) for synthesis of cDNA. Oligonucleotide primers targeting a full-length 3D gene were designed based on FMDV Asia 1 sequence (GenBank accession AF207520). Sense primer contained the BamHI site followed by start codon and 6× histidine tag in frame with coding sequence, while antisense primer contained BamHI site alone on 5′end. (Sense primer) HDF5: TGGGATCCATGCATCACCATCACCATCACGGGTTGATCGTTGAC, (anti-sense primer) HDR6: TGGGATCCTTATGCGTCACCGCACAC. Reverse transcription (RT) of viral RNA was performed in 20 μl reaction volume using oligo (dT) primer and AMV reverse transcriptase with ThermoScript™ RT-PCR System (Invitrogen). The cDNA obtained was used as template for PCR amplification of 3D gene using pfu DNA polymerase (MBI, Fermentas Life Sciences, USA). PCR was carried out by initial denaturation of 95 °C for 2 min. followed by 30 cycles of 95 °C for 45 s, 55 °C for 30 s, 72 °C for 90 s.
Cloning of 3D PCR Product into Transfer Vector
The PCR product was purified from 1 % agarose electrophoresed gel using agarose gel extraction kit (Qiagen) and cloned into into BamHI site of pFastBac1 donor plasmid vector (Invitrogen). Sequence of the gene was determined to check the orientation and correctness of the reading frame. Transformation of the recombinant transfer vectors into MAX Efficiency®DH10Bac™ competent cells (Invitrogen) was carried out as per manufacturer’s instructions.
Transfection and Isolation of Recombinant Baculovirus
Recombinant bacmid DNA was isolated and transfected into Sf-21 insect cells by Cellfectin®II Reagent (Invitrogen). Following incubation period of 96 h, supernatant containing recombinant baculovirus was harvested and stored at 4 °C. Following one round of plaque purification, baculovirus stocks were amplified several folds by infecting Sf-21 insect cell monolayer culture at multiplicity of infection (m.o.i) of 0.01–0.1 and harvesting the virus 72 h post-infection. Recombinant baculovirus stocks were titrated by plaque assay and further used for infection of TN-5 cells for expression of 3D protein.
Analysis of Expression of FMDV-3D Polymerase
High five cells were infected with recombinant baculovirus at a m.o.i. of 5. Infected cell lysates prepared in 2× SDS sample buffer were subjected to 12 % polyacrylamide gel electrophoresis (SDS-PAGE) and specificity of the expressed protein was confirmed by western blotting using serum from FMDV type A (IND-40/00 strain), experimentally infected bull calf.
Preparation of Recombinant 3D Protein
TN5 cells (2 × 107 cells) were seeded in 225 cm2 flasks and infected with recombinant baculovirus for 96 h to prepare cell lysate. Cells were harvested and used for purification of 3D protein using ProBond™ Purification System (Invitrogen). Briefly, cells were homogenized using syringe in 8.0 ml of 1x native purification buffer containing protease inhibitor cocktail (Sigma). Cell lysate was clarified by centrifugation at 3,000 rpm for 10 min in Sorvall RT7 refrigerated centrifuge and supernatant subjected to purification as per manufacturer’s instructions. Eluted fractions of purified protein were dialyzed against PBS, analyzed by SDS-PAGE and stored at −80 °C after adding 25 % glycerol.
Serum Samples
For optimization of ELISA, serum samples collected from bull calves experimentally infected with FMDV and the samples of adult bovine sera from New Zealand and Australia origin were used as positive and negative control sera, respectively. Serum samples (N = 287) collected from herds with no history of exposure to FMD and having titres of <8 in serum neutralization and liquid phase blocking ELISA (LPBE), were used as negative samples to establish test specificity. Samples collected from bull calves experimentally infected with FMDV serotype O and A (7–56 days post infection) or from diary, which had experienced FMD outbreak about 6 months to 1 year back were used. All these samples were previously found to be positive for FMD by means of SNT and 3ABC-ELISA, thus were considered as known positive samples. Sequential serum samples (79 milch cows) collected from organized herds both pre-vaccination and 28-days post-vaccination with trivalent FMD vaccine, were screened for presence of anti-3D antibodies. Serum samples collected at 28–30 days after last vaccination of multiply vaccinated animals (n = 52) from a dairy herd were also screened for presence of anti-3D antibodies. These animals were vaccinated regularly at 4–6 months intervals for 5 times. Samples collected randomly from field (n = 450) were also assayed for presence of NSP antibodies.
Assay Optimization for 3D-iELISA
Optimization of the 3D-iELISA was carried out by checkerboard titration for all reagents. Immunoassay plate (Maxisorp, Nunc, Denmark) was coated with recombinant 3D antigen in coating buffer (Carbonate-bicarbonate buffer 0.05 M, pH 9.6) and incubated at 4 °C overnight. On the following day, after discarding plate contents unbound sites in the wells were saturated with blocking buffer containing, 4 % w/v skimmed milk powder and 1 % bovine serum albumin in washing buffer (sodium phosphate buffer 0.01 M containing 0.5 M NaCl and 0.1 % Tween 20) for 45 min at 37 °C with intermittent shaking. After three washings in wash buffer, test serum samples, a positive control and a negative control serum diluted (1:50) in blocking buffer were dispensed at 50 μl/well in duplicate. This dilution was determined based on absorbance values of serial dilutions of known positive and negative sera. Two wells containing only blocking buffer without serum served as background control. After incubation for 1 h at 37 °C the wells were washed five times. Anti cow-HRPO conjugate (Dako) diluted 1:3000 in blocking buffer was dispensed at 50 μl/well and plate incubated for 1 h at 37 °C. After five final washings, substrate solution containing ortho-phenylene diamine (OPD) and H2O2 was added at 50 μl/well. The plate was incubated in dark at room temperature for 15 min. Colour development was stopped with addition of 50 μl 1 M H2SO4 in each well. Absorbance was measured at 492 nm wavelength (with reference wavelength at 620 nm) in an ELISA reader. Samples that gave optical density (OD) values, more than 2 times the mean OD value of the negative serum samples were considered positive in the test system.
Results
Cloning of FMDV-3D Gene into pFastBac1 Donor Plasmid and Generation of Recombinant Virus
PCR amplification of the 3D gene was carried out using the cDNA template obtained from FMDV type Asia 1 with oligo(dT) primer. The PCR product of the expected size 1.45 kb was confirmed on agarose (1 %) gel electrophoresis. The 3D gene contained open reading frame (ORF) of 1434 bases including start codon followed by 6 × histidine codons at n-terminus. The ORF was cloned into pFastBac1 and confirmed by restriction digestion. Recombinant pFastBac1 donor plasmid was transformed into MAX Efficiency® DH10Bac™ competent cells to generate recombinant bacmid DNA containing 3D. Cells were then transfected with recombinant bacmid DNA. Transfected cultures showed cytopathic changes by day 3 post-transfection. The culture supernatants were harvested 4th day post transfection and subjected to one round of plaque purification. Viral titres were amplified up to 8.3 × 108 pfu/ml by passaging recombinant virus in Sf-21 cells. These virus stocks were stored at 4 °C for further use in infection of TN-5 cells for expression analysis.
Expression and Analysis of Non-Structural Protein 3D Polymerase of FMDV
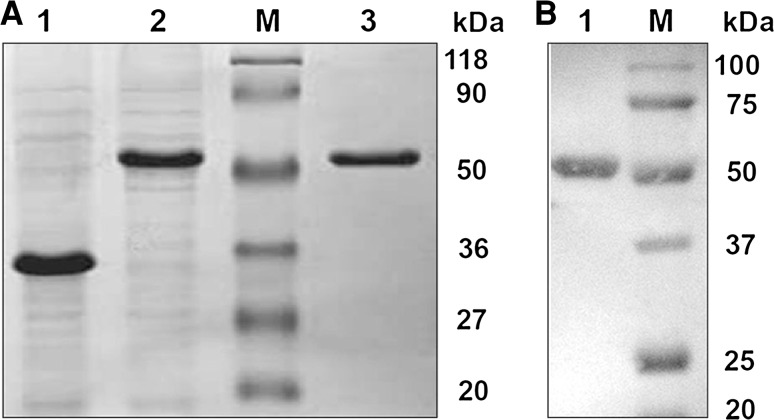
TN-5 cell monolayers were infected at a m.o.i. of 4–5 for expression studies. Cells were harvested at regular time intervals such as 72, 96 and 120 h and assayed for expression by SDS-PAGE and western blotting. High level protein expression was achieved at 96 h post infection of TN5 cells. SDS-PAGE of TN-5 cell lysate showed the over expressed protein band of molecular weight 53 kDa (Fig. 1). Protein specificity was confirmed by western blotting. A distinct band of 53 kDa was observed with the expressed protein reacting strongly and specifically with the infected bovine serum (Fig. 2). Antigenic specificity of the expressed protein was checked by reactivity with infected bovine serum in indirect ELISA by using crude lysates of TN5 infected with recombinant and wild type baculovirus (AcMNPV), respectively, the latter being a negative control.
Fig. 1.
Expression of 3D of FMDV in insect cells. A SDS-PAGE analysis of baculovirus expressed proteins in TN-5 cells. Lane 1 Polyhydrin protein (29 kDa) expressed by AcMNPV (wild type baculovirus). Lane 2 3D protein (53 kDa) of FMDV expressed by recombinant baculovirus in insect cells. Lane M Prestained protein molecular weight marker (MBI, fermentas life sciences). Lane 3 3D protein (53 kDa) fraction purified by nickel affinity chromatography. B Western blotting for expressed 3D protein. Lane 1 3D protein 3D protein (53 kDa) of FMDV expressed by recombinant baculovirus in TN-5 insect cells. Lane M Prestained molecular weight marker (Biorad)
Fig. 2.
Reactivity of the control sera at various dilutions to check the sensitivity of the test system
Protein was subsequently purified to homogeneity under native conditions using probond purification kit (Invitrogen, USA) as per manufacturer’s instructions. Purified protein yield was 1.04 mg ± 0.16 of every 10 million TN5 cells grown under stationary culture.
Screening of Serum Samples
Antigen concentration of 80 ng per well and serum dilutions of 1:50 were found to be optimum. By evaluation of bovine sera of known FMD status, a cut-off were established as follows: (a) 3D NSP reactivity positive: if T/N value is ≥2, (b) 3D NSP reactivity negative: if T/N value is <2.
The assay showed sensitivity of 92 % while the specificity was 100 % on given set of serum samples (Table 1). In experimentally infected animals, 3D antibodies were detectable by day 7 post infection. Frequency distribution of T/N values of all serum samples tested is shown in Fig. 3. Of the sequential cattle sera samples collected from two dairy farms, none of 79 vaccinated animals showed rise in antibody titres 1-month following vaccination. On the other hand, sera from animals that were multiple vaccinated at least five times in the past, 4/52 (7.7 %) animals showed positive reactivity.
Table 1.
Screening of bovine sera in 3D-iELISA
| Status of sera samples | Total number of samples | Positive reactors | Percent reactors |
|---|---|---|---|
| Naïve animals | 287 | 5 | 1.74 |
| Infected animalsb | 204 | 188 | 92.16 |
| 28 days post vaccinated | 79 | 0 | 0.00 |
| Multiple vaccinateda | 52 | 4 | 7.69 |
| Random samples | 450 | 88 | 19.6 |
All samples tested were from Indian cattle (Bos indicus) or from their cross breds, aged >6 months
aSamples were collected 28–30 days after last vaccination in these animals that received regular multiple vaccination every 4–6 months
bSamples comprise both experimentally infected and naturally infected animals collected 7–56 days post infection and 1 year after the disease outbreak, respectively
Fig. 3.
Frequency distribution of the T/N values of serum samples (n = 1072), collected from naïve, infected, vaccinated and field animals collected at random
Discussion
NSP based diagnostics were not given much importance in India until recently. With the launch of FMD control program by government of India through intensive mass vaccination campaigns, these diagnostics have been considered valuable tool for monitoring the incidence of the disease in the areas covered under this programme. Early identification of animals that have been infected with FMDV is of considerable importance in the sero-surveillance of FMD, during the control program. Several NSP based ELISAs have been developed, amongst which 3ABC based assays are considered more reliable and hence in common use. However use of more than one NSP antigens is suggested to improve the diagnostic sensitivity [25] as certain doubtful reactors always need to be retested to confirm the reactivity. It can also be valuable as complementary assay for DIVA in addition to 3ABC antigen [26]. Among all FMDV serotypes, 3D is most conserved and immunogenic protein, therefore a 3D-iELISA can be used to detect the infection in animals exposed to any of the serotypes at the earliest signs of an outbreak. Although NSP diagnostic is not a prescribed test for trade by the OIE, the assay can be valuable in a situation where the type specificity of the virus in a territory in question is not clear [3]. 3D based assays would be also useful in the screening of exotic animals for import restrictions, epidemiological surveys to determine the spread of FMDV in animal population and for identification of persistently infected animals [7, 11, 22, 21].
Although it is relatively easy to produce recombinant antigens in E.coli, the major disadvantage is high background reactivity of bacterially expressed protein in ELISA unless the protein is purified. In this report, 3D protein was produced in insect cells using baculovirus system. Several post-transnational modifications have been reported to occur in the baculovirus expression system, including N and O-linked glycosylation, phosphorylation, carboxymethylation, isoprenylation, signal peptide cleavage and proteolytic cleavage [12]. The sites where these modifications occur are often identical to those of the authentic protein in its native cellular environment. Therefore, activity of the recombinant protein is expected to be similar to its native counterpart, both structurally and functionally. Recombinant protein can be applied directly in ELISA obviating the need for purification as protein is generally over expressed in insect cells under polyhedrin promoter. However in this study, we optimized the purification of the protein employing Ni–NTA affinity column. Protein could be purified to homogeneity under native conditions.
The expressed 3D protein was confirmed by use of experimentally infected bovine sera against type A Ind-40/2000 FMDV strain. Western blotting showed a distinct protein band of 53 kDa size. To reduce inter-run variation due to difference in absolute absorbance between runs, final results for each test serum were expressed as the test to negative control ratio (T/N). The diagnostic sensitivity and specificity of the assay was 92 and 100 % respectively. Findings of this assay are in agreement with those in previous reports for 3D based immunoassay [5, 19]. Reduced sensitivity of the 3D-iELISA as compared to other NSP assays, may be ascribed to poor immunogencity of 3D in some animals. Variation in the 3D antibody response of individual animals after infection has been reported [1, 10].
3D-iELISA was used to assess the presence of 3D antibodies in few organized herds. In a herd with no previous exposure to FMD in the past, no detectable antibodies were found in these animals. In experimentally infected animals, our test system could clearly detect the infected animals 7 days post infection. Antibodies to 3D develop earlier and persist for longer period too in animal body after exposure to live virus. 3D antibodies are detectable following exposure as early as 5–6 days post infection and at later stages in persistently infected animals up to more than 700 days [4, 17]. Antibodies to nonstructural proteins, including 3D have been detected by EITB both at early and late phases of infection [4].
Sequential sera samples (n = 79) collected from same animals 28 days post vaccination (dpv) showed no detectable levels of 3D antibodies. Immune response to structural proteins following vaccination was confirmed by serum neutralization test (data not shown). However in the multiple vaccinated animals, 4 of 52 animals showed 3D antibodies. Antibodies to 3D post vaccination are detectable [19, 20]. O’Donnell et al. [19] have shown that 5 of 6 cattle immunized with FMD oil adjuvant vaccine at 30 dpv were positive in ELISA. At 60 dpv, only 2 sera remained positive, and at 120 dpv, none of the vaccinated cattle had antibodies detectable by the 3D-ELISA. Although some mature virions contain RNA polymerase [18], it is not demonstrated whether endogenous or exogenous contamination of virions with 3D is responsible for the induction of 3D antibodies [9].
NSP antibody titres are higher in the infected populations and persist for longer even up to 687–700 days [4, 6, 17]. The amount of antibodies to 3D, and to other NSP, that is detected in sera from vaccinated animals depends on the immunogenicity of the NSP concerned, the amount that is present in the vaccine, the number of vaccinations that the animal has received and the time since last vaccination [15]. This would be a possible explanation for the differences among vaccines in the 3D antibody response in vaccinated cattle, which is influenced by the methodology of antigen production, inclusion of purification steps, the antigen concentration, or the presence of adjuvants [4, 9, 14] (Alonso et al. 1988).
For epidemiological studies, sera sampling is usually carried out 4–6 months after the last vaccination in cattle. In India, routine vaccination is carried out twice in a year, at an interval of 6 months. Little or no reactivity of 3D antibodies in post vaccination or post multiple vaccination status could be explained by the fact that the antibodies to NSP don’t last longer than 3–4 months following vaccination. Thus, sera samples collected after 4–6 months following vaccination may or may not show 3D antibodies induced due to vaccine. Since vaccination may prevent the disease but not clinical infection, vaccinated animals can be infected, and might show antibodies against NSPs [6].
The guidelines for FMD vaccine production in the OIE Manual of Standards for Diagnostic Tests and Vaccines recommend the manufacturers to exclude NSPs from their products. Therefore, DIVA is more useful in context of the animal herds from organized farm where the regular vaccination is practiced. The test is easily adaptable in the large-scale serosurveillance of FMD before further serotyping is considered. In the efforts towards progressive control through vaccination, it is desirable that manufacturers produce the vaccines that are certified free from NSP antigens so that NSP serology can be increasingly adopted for routine sero-surviellance.
Acknowledgments
The authors thank the Director, Indian Veterinary Research Institute (IVRI), Izatnagar, for facilitating this work. RK acknowledges receipt of IVRI-JRF. Authors thank Prof. M. S. Shaila, IISc, Bangalore and Dr S. E. Hasnain, CDFD, Hyderabad for providing wild strains of baculovirus. Dr Suresh Basagoudanavar is acknowledged for critical review of the manuscript.
References
- 1.Alonso Fernandez A, Auge de Melo P, Gomes I, Rosemberg F. The use of virus infection associated antigen (VIA) on the detection of cattle exposed to FMDV. Bol Cent Panam Fiebre Aftosa. 1975;17–18:17–22. [Google Scholar]
- 2.Alonso A, Gomes MPD, Martins MA, Sondahl MS. Detection of foot-and-mouth disease virus infection-associated antigen antibodies: comparison of the enzyme-linked immunosorbent assay and agar gel immunodiffusion tests. Prev Vet Med. 1990;9:233–240. doi: 10.1016/0167-5877(90)90069-T. [DOI] [Google Scholar]
- 3.Anon (2004) Manual of standards for diagnostic tests and vaccines. Office Internationale des Epizooties (6th edition). Office Internationale des Epizooties, Paris, 1, p. 190–216.
- 4.Bergmann IE, Augé de Mello P, Neitzert E, Beck E, Gomes I. Diagnosis of persistent aphthovirus infection and its differentiation from vaccination response in cattle by use of enzyme-linked immunoelectrotransfer blot analysis with bioengineered nonstructural viral antigens. Am J Vet Res. 1993;6:825–831. [PubMed] [Google Scholar]
- 5.Bergmann I, Malirat V, Neitzert E, Beck E, Panizzutti N, Sanchez C. Improvement of a serodiagnostic strategy for foot-and-mouth Disease Virus surveillance in cattle under systematic vaccination: a combined system of an indirect ELISA-3ABC with an enzyme linked immunoelectrotransfer blot assay. Arch Virol. 2000;145:473–479. doi: 10.1007/s007050050040. [DOI] [PubMed] [Google Scholar]
- 6.Clavijo A, Wright P, Kitching P. Developments in diagnostic techniques for differentiating infection from vaccination in foot-and-mouth disease. Vet J. 2004;167:9–22. doi: 10.1016/S1090-0233(03)00087-X. [DOI] [PubMed] [Google Scholar]
- 7.Cowan KM, Graves JG. A third antigenic component associated with foot-and-mouth disease virus infection. Virol. 1996;30:528–540. doi: 10.1016/0042-6822(66)90128-0. [DOI] [PubMed] [Google Scholar]
- 8.Dekker A, Sammin D, Greiner M, Bergmann I, Paton D, Grazioli S, De Clercq K, Brocchi E. Use of continuous results to compare ELISAs for the detection of antibodies to non-structural proteins of foot-and-mouth disease virus. Vaccine. 2008;26:2723–2732. doi: 10.1016/j.vaccine.2008.03.052. [DOI] [PubMed] [Google Scholar]
- 9.Frolov VG, Duque H, Palmenberg AC. Quantification of endogenous viral polymerase, 3Dpol, in preparations of meningo and encepahlomyocarditis viruses. Virology. 1999;260:148–155. doi: 10.1006/viro.1999.9808. [DOI] [PubMed] [Google Scholar]
- 10.Gebauer F, La Torre JC, Gomes I, Mateu MG, Barahona H, Tiraboschi B, Bergmann I, de Mello PA, Domingo E. Rapid selection of genetic and antigenic variants of FMDV during persistence in cattle. J Virol. 1988;62:2041–2049. doi: 10.1128/jvi.62.6.2041-2049.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Hedger RS. Observations on the carrier state and related antibody titres during an outbreak of foot and mouth disease. J Hyg Camb. 1968;66:27–36. doi: 10.1017/S0022172400040912. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Hoss A, Moarefi I, Scheidtmann KH, Cisek LJ, Corden JL, Dornreiter I, Arthur AK, Fanning E. Altered phosphorylation pattern of simian virus 40 T antigen expressed in insect cells by using a baculovirus vector. J Virol. 1990;64:4799–4807. doi: 10.1128/jvi.64.10.4799-4807.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Lubroth J, Brown F. Identification of native foot-and-mouth disease virus non-structural protein 2C as a serological indicator to differentiate infected from vaccinated livestock. Res Vet Sci. 1995;59:70–78. doi: 10.1016/0034-5288(95)90034-9. [DOI] [PubMed] [Google Scholar]
- 14.Lubroth J, Lopez A, Ramalho AR, Meyer RF, Brown F, Darsie GC. Cattle response to foot-and-mouth disease virus non-structural proteins as antigens within vaccines produced using different concentrations. Vet Q. 1998;20:S13–S17. doi: 10.1080/01652176.1998.9694955. [DOI] [PubMed] [Google Scholar]
- 15.Mackay DKJ, Forsyth MA, Davies PR, Berlinzani A, Belsham GJ, Flint M, Ryan MD. Differentiating infection from vaccination in foot-and-mouth disease using a panel of recombinant, non-structural proteins in ELISA. Vaccine. 1998;16:446–459. doi: 10.1016/S0264-410X(97)00227-2. [DOI] [PubMed] [Google Scholar]
- 16.Mohapatra JK, Pandey LK, Sanyal A, Pattnaik B. Recombinant non-structural polyprotein 3AB-based serodiagnostic strategy for FMD surveillance in bovines irrespective of vaccination. J Virol Methods. 2011;177:184–192. doi: 10.1016/j.jviromet.2011.08.006. [DOI] [PubMed] [Google Scholar]
- 17.Neitzert E, Beck E, Augé de Mello P, Gomes I, Bergmann IE. Expression of the aphthovirus RNA polymerase gene in E. coli and its use together with other bioengineered non-structural antigens in detection of late persistent infections. Virology 1991; 184:799–804 [DOI] [PubMed]
- 18.Newman JFE, Piatti PG, Ryan M, Brown F. Functions of minor polypeptides in foot-and-mouth disease virus and poliovirus. Trends Microbiol. 1994;2:494–497. doi: 10.1016/0966-842X(94)90655-6. [DOI] [PubMed] [Google Scholar]
- 19.O’Donnell VK, Boyle DB, Sproat K, Fondevila NA, Forman A, Schudel AA, Smitsaart EN. Detection of antibodies against foot-and-mouth disease virus using a liquid-phase blocking sandwich ELISA (LPBE) with a bioengineered 3D protein. J Vet Diagn Invest. 1996;8:143–150. doi: 10.1177/104063879600800201. [DOI] [PubMed] [Google Scholar]
- 20.O’Donnell VK, Smitsaart E, Cetra B, Duffy S, Finelli J, Boyle D, Draghi G, Fondevila N, Schudel AA. Detection of virus infection-associated antigen and 3D antibodies in cattle vaccinated against foot-and-mouth disease. Rev Sci Tech Off Int Epiz. 1997;16:833–840. [PubMed] [Google Scholar]
- 21.Pinto AA, Garland AJM. Immune response to virus-infection-associated (VIA) antigen in cattle repeatedly vaccinated with foot-and-mouth disease virus inactivated by formalin or acetylethyleneimine. J Hyg. 1979;82:41–50. doi: 10.1017/S0022172400025456. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Pinto AA, Hedger RS. The detection of antibody to virus-infection-associated (VIA) antigen in various species of African wild life following natural and experimental infection with FMD virus. Arch Virol. 1978;57:307–314. doi: 10.1007/BF01320070. [DOI] [PubMed] [Google Scholar]
- 23.Rodriguez A, Dopazo J, Saiz JC, Sobrino F. Immunogenicity of non-structural proteins of Foot-and-Mouth Disease Virus differences between infected and vaccinated swine. Arch Virol. 1994;136:123–131. doi: 10.1007/BF01538822. [DOI] [PubMed] [Google Scholar]
- 24.Shen F, Chen PD, Walfield AM, Ye J, House J, Brown F. Differentiation of convalescent animals from vaccinated against foot-and- mouth disease by a peptide ELISA. Vaccine. 1999;17:3039–3049. doi: 10.1016/S0264-410X(99)00148-6. [DOI] [PubMed] [Google Scholar]
- 25.Uttenthal A, Parida S, Rasmussen TB, Paton DJ, Haas B, Dundon WG. Strategies for differentiating infection in vaccinated animals (DIVA) for foot-and-mouth disease, classical swine fever and avian influenza. Expert Rev Vac. 2010;9:73–87. doi: 10.1586/erv.09.130. [DOI] [PubMed] [Google Scholar]
- 26.Sorenson KJ, Madsen KG, Madsen ES, Salt JS, Nqindi J, Mackay DKJ. Differentiation of infection from vaccination in foot-and-mouth disease by the detection of antibodies to the non-structural proteins 3D, 3AB and 3ABC in ELISA using antigens expressed in baculovirus. Arch. Virol. 1998;143:1461–1476. [DOI] [PubMed]