Abstract
Realizing the potential of human embryonic stem cells (hESCs) has been hindered by the inefficiency and instability of generating desired cell types from pluripotent cells through multi-lineage differentiation. We recently reported that pluripotent hESCs maintained under a defined platform can be uniformly converted into a cardiac or neural lineage by small molecule induction, which enables lineage-specific differentiation direct from the pluripotent state of hESCs and opens the door to investigate human embryonic development using in vitro cellular model systems. To identify mechanisms of small molecule induced lineage-specification of pluripotent hESCs, in this study, we compared the expression and intracellular distribution patterns of a set of cardinal chromatin modifiers in pluripotent hESCs, nicotinamide (NAM)-induced cardiomesodermal cells, and retinoic acid (RA)-induced neuroectodermal cells. Further, genome-scale profiling of microRNA (miRNA) differential expression patterns was used to monitor the regulatory networks of the entire genome and identify the development-initiating miRNAs in hESC cardiac and neural lineage-specification. We found that NAM induced nuclear translocation of NAD-dependent histone deacetylase SIRT1 and global chromatin silencing, while RA induced silencing of pluripotence-associated hsa-miR-302 family and drastic up-regulation of neuroectodermal Hox miRNA hsa-miR-10 family to high levels. Genome-scale miRNA profiling indentified that a unique set of pluripotence-associated miRNAs was down-regulated, while novel sets of distinct cardiac- and neural-driving miRNAs were up-regulated upon the induction of lineage-specification direct from the pluripotent state of hESCs. These findings suggest that a predominant epigenetic mechanism via SIRT1-mediated global chromatin silencing governs NAM-induced hESC cardiac fate determination, while a predominant genetic mechanism via silencing of pluripotence-associated hsa-miR-302 family and drastic up-regulation of neuroectodermal Hox miRNA hsa-miR-10 family governs RA-induced hESC neural fate determination. This study provides critical insight into the earliest events in human embryogenesis as well as offers means for small molecule-mediated direct control and modulation of hESC pluripotent fate when deriving clinically-relevant lineages for regenerative therapies.
Keywords: MicroRNA, Chromatin modification, Chromatin remodeling, Small molecule, Retinoic acid, Nicotinamide, Human embryonic stem cell, Pluripotence, Neural differentiation, Cardiac differentiation, Lineage-specification, Neuroectoderm, Cardiomesoderm, Neuronal progenitor, Cardiac precursor, Neuron, Cardiomyocyte, Defined culture system, SIRT1, MiR-302, MiR-10
Introduction
The human embryonic stem cells (hESCs), derived from the inner cell mass (ICM) or epiblast of human blastocyst, are genetically stable with unlimited expansion ability and unrestricted plasticity, proffering a pluripotent reservoir for in vitro derivation of a large supply of disease-targeted human somatic cells that are restricted to the lineage in need of repair [1,2]. The pluripotence of hESCs implies such cells’ tremendous potential for tissue and function restoration, whereas how to channel the wide differentiation potential of human pluripotent cells efficiently and predictably to a desired phenotype has been a major challenge for both developmental study and clinical translation. Conventional approaches rely on multi-lineage inclination of pluripotent cells through spontaneous germ layer differentiation, which yields mixed populations of cell types that may reside in three embryonic germ layers and often makes desired differentiation not only inefficient, but uncontrollable and unreliable as well [1,2]. Following transplantation, these pluripotent-cell-derived grafts tend to display not only a low efficiency in generating the desired cell types necessary for reconstruction of the damaged structure, but also phenotypic heterogeneity and instability, hence, a high risk of tumorigenicity [2–6]. In view of growing interest in the use of human pluripotent cells, including artificially-reprogrammed human induced pluripotent stem cells (hiPS cells) [6–11], teratoma formation and the emergence of inappropriate cell types have become a constant concern following transplantation. Developing a more practical approach that permits to channel the wide differentiation potential of pluripotent cells efficiently and predictably to a desired phenotype is vital to harnessing the power of hESC biology for safe and effective clinical translation.
The hESC lines initially were derived and maintained in co-culture with growth-arrested mouse embryonic fibroblasts (MEFs) [1]. Although several human feeder, feeder-free, and chemically-formulated culture systems have been developed for hESCs, the elements necessary and sufficient for sustaining the self-renewal of human pluripotent cells remain unsolved [2,12]. These exogenous feeder cells and biological reagents help maintain the long-term stable growth of undifferentiated hESCs whereas mask the ability of pluripotent cells to respond to developmental signals. Maintaining undifferentiated hESCs in a defined biologics-free culture system that allows faithful expansion and controllable direct differentiation is one of the keys to their therapeutic utility and potential. A defined platform for the maintenance of pluripotent hESCs may overcome some of the major obstacles in translational biology, including de novo derivation of clinically-suitable hESCs and effectively directing such hESCs uniformly towards clinically-relevant lineages [2,12]. Substantial neural and cardiac differentiation appears to occur at relatively early stages in embryonic stem cell cultivation under conditions that induce differentiation, consistent with the early development of the central nervous system (CNS) and the heart from the cells of epiblast in embryogenesis [2,12]. In early embryogenesis, the epiblast is composed of more progressed pluripotent cells developed from the ICM, serving as the most immediate precursors of the early somatic lineages [13–15]. Therefore, a defined culture system for maintenance of hESCs might not only render specification of clinically-relevant early lineages directly from the pluripotent state without an intervening multi-lineage germ-layer stage, but also allow identify the signaling molecules necessary and sufficient for inducing the cascade of organogenesis in a process that may emulate the human embryonic development [2,12].
Previously, we found that small molecule nicotinamide (NAM) and retinoic acid (RA) induced respective cardiac- and neural-lineage specification direct from the pluripotent state of hESCs maintained under a defined culture that further progressed to beating cardiomyocytes and neurons with high efficiency [12,16,17]. Having achieved uniformly conversion of pluripotent hESCs to a cardiac or neural lineage with small molecule induction, in this study, we examined the expression and intracellular distribution patterns of a set of cardinal chromatin modifiers in hEAC cardiac or neural fate switch direct of pluripotence in order to uncover the epigenetic mechanism governing early lineage specification direct from the pluripotent stage. To uncover key regulators, genome-scale profiling of microRNA (miRNA) differential expression patterns was further used to identify novel sets of human development-initiating miRNAs upon small-molecule- induced neural and cardiac lineage specification direct from the pluripotent stage. NAM was found to induce nuclear translocation of NAD-dependent histone deacetylase SIRT1 and global chromatin silencing. A unique set of pluripotence-associated miRNAs was down-regulated, while novel sets of distinct cardiac- and neural-driving miRNAs were up-regulated upon the induction of lineage specific differentiation from the pluripotent state of hESCs, including silencing of the most prominent pluripotence-associated hsa-miR-302 family and a drastic expression increase of Hox miRNA hsa-miR-10 family upon RA exposure.
Materials and Methods
Culture of undifferentiated hESCs
The hESC lines WA01 and WA09 (H1 and H9 from WiCell Research Institute, passages 30–50) were used in this study. The defined culture systems consist of DMEM/F-12 or KO-DMEM (knockout-DMEM) (80%), Knockout Serum Replacement (KO) (20%), L-alanyl-L-gln or L-gln (2 mM), MEM nonessential amino acids (MNAA, 1X), β-Mercaptoethanol (β-ME, 100 μM) (all from Invitrogen), human purified laminin (Sigma) or laminin/collagen (growth factor reduced Matrigel, BD Bioscience) as the matrix protein, and bFGF (basic fibroblast growth factor, 20 ng/ml) (PeproTech Inc). The KO can be replaced with defined essential factors containing MEM essential amino acids (MEAA, 1X), human insulin (20 μg/ml) (Sigma), and ascorbic acid (50 μg/ml) (Sigma), in which activin A (50 ng/ml, Sigma), human albumin (10 mg/ml, Sigma), and human transferrin (8 μg/ml, Sigma) were added in order to increase cell survival and maintain normal shape and healthy colonies.
Cellular immunofluorescence
The cells were fixed with 4% paraformaldehyde and blocked in PBS containing 0.2% Triton X-100 and 2% BSA. The cells were incubated with the primary antibody in 0.1% Triton X-100 in PBS at 4°C overnight, and then with secondary antibody (Molecular Probe) in the same buffer at room temperature for 45 min. After staining with DAPI, cells were visualized under an immunofluorescence and deconvolution microscope.
Cardiac lineage-specification direct from the pluripotent state of hESCs induced by NAM
Undifferentiated hESCs maintained under the defined culture conditions were treated with NAM (10 mM) 3 days after seeding for 4–5 days to form cardiomesoderm, and then allowed to form floating cellular clusters (cardioblasts) in a suspension culture in the hESC media for 4–5 days. After permitting attachment to a tissue culture substrate, the cardiblasts were further treated with NAM (10 mM) for one more week and cultured in a differentiation media consisting of DMEM/F-12 (90%), defined FBS (Hyclone) (10%), and L-alanyl-L-gln or L-gln (2 mM). Beating cardiomyocytes were observed in about one week after withdrawal of NAM, increased in numbers with time.
Neural lineage-specification direct from the pluripotent state of hESCs induced by RA
hESC colonies maintained on purified laminin or laminin/collagen in the defined media supplemented with RA (10 μM) were allowed to grow to day 7 to form neuroectoderm. The differentiated hESCs were transferred to a serum-free suspension culture to allow floating cellular clusters (neuroblasts) to form in the hESC media for 4–5 days. The neuroblasts were then permitted to attach to a tissue culture plate or seeded in a laminin/collagen polymerized 3-dimensional matrix in a defined medium containing DMEM/F-12, N-2 supplement (1%), heparin (8 μg/ml), VEGF (vascular endothelial growth factor, 20 ng/ml), NT-3 (neurotrophin-3, 10 ng/ml), and BDNF (brain derived neurotrophic factor, 10 ng/ml). Extensively neurite-bearing cells and pigmented cells were observed within 2 weeks of continuous cultivation, increased in numbers with time.
MiRNA microarray analysis
Total RNAs, including miRNAs, were isolated from hESCs and their cardiac (treated with NAM) and neural (treated with RA) derivatives under the defined culture using the Qiagen miRNeasy kit (Qiagen, www.qiagen.com). The quality of purified total RNAs was verified by electrophoresis analysis. The human miRNA expression profiling was done and the microarray data analysis from two biological replicate sets (WA01 and WA09) was provided by LC Sciences (Huston, TX, www.lcsciences.com) as part of miRNA microarray service. Multiple redundant regions and control probes were included in the content of each chip. Each region further comprises a miRNA probe region, which detects miRNA transcripts listed in Sanger miRBase Release 15.0 (http://www.sanger.ac.uk/Software/Rfam/mirna/). Statistic tests and clustering analysis were provided by LC Sciences as part of miRNA microarray service. The signal values were derived by background subtraction and normalization. Detectable transcripts were subjected to data processing statistics and signal intensities were listed in average values of repeating spots.
Results and Discussion
NAM-induced cardiomesodermal cells have acquired a silenced chromatin while RA-induced neuroectodermal cells retain an embryonic acetylated chromatin
In order to achieve uniformly conversion of human pluripotent cells to a particular lineage, previously, we employed a defined culture system capable of insuring the proliferation of undifferentiated hESCs to identify conditions for well-controlled efficient induction of pluripotent cells exclusively to a particular clinically-relevant lineage by simple provision of small molecules [12,16,17]. After screening the differentiation inducing effect of a variety of small molecules and growth factors on pluripotent hESCs maintained under a defined culture, RA was identified sufficient to induce the specification of neuroectoderm direct from the pluripotent state of hESCs that yielded β-III-tubulin- and Map-2-expressing neurons efficiently by promoting nuclear translocation of the neuronal-specific transcription factor Nurr-1 [12,16]. Similarly, the defined platform renders nicotinamide (NAM) sufficient to induce the specification of cardiomesoderm direct from the pluripotent state of hESCs by promoting the expression of the earliest cardiac-specific transcription factor Csx/Nkx 2.5 and triggering progression to beating cardiomyocytes efficiently [12,17]. Such a system emulating human embryonic development serves as a platform for a well-controlled efficient approach to specify pluripotent hESC differentiation exclusively to a particular clinically-relevant lineage by small molecule induction, which not only opens the door to investigate human embryonic development using in vitro cellular model systems, but also enables well-controlled efficient derivation of an unlimited supply of human cardiac and neuronal cells across the spectrum of developmental stages for cell-based therapeutics.
The eukaryotic genome is packaged into chromatin, a nucleoprotein complex in which the DNA helix is wrapped around an octamer of core histone proteins, known as nucleosome, to form a nucleosomal DNA structure that is further folded into higher-order chromatin structures with the involvement of other chromosomal proteins [18,19]. Chromatin modifications serve as important epigenetic marks for active and inactive chromatin and have emerged as the principal epigenetic mechanism in early embryogenesis. In general, histone acetylation and histone H3 K4 methylation correlate with a transcriptionally active (open) chromatin state, whereas histone deacetylation and histone H3 K9 methylation correlate with a transcriptionally repressed (closed) chromatin state [18,19]. Previously, chromatin states have been used to characterize and compare the intricate plasticity and potential of stem cell populations [20–23]. However, without a practical strategy to convert pluripotent cells direct into a specific lineage, previous studies and profiling of pluripotent hESCs and their differentiating multi-lineage aggregates have generated compromised implications to molecular controls in human embryonic development [20–23].
Having achieved uniformly conversion of pluripotent hESCs to a cardiac or neural lineage with small molecule induction (Figure 1A), we further examined and compared the expression and intracellular distribution patterns of a set of cardinal chromatin modifiers in pluripotent hESCs, NAM-induced cardiomesodermal cells, and RA-induced neuroectodermal cells in order to uncover the epigenetic mechanism governing early lineage specification direct from the pluripotent stage (Figure 1B–D). We observed that RA-induced hESC-derived neuroectodermal cells retained high levels of expression and nuclear localization of active chromatin modifiers that have been associated with the chromatin states of embryonic stem cells and their derivatives [20–23], including acetylated histones H3 and H4 (acH3 and acH4), ATP-dependent active chromatin-remodeling factors (Brg-1 and hSNF2H), and the histone deacetylase HDAC1 (Figure 1B), suggesting an acetylated globally active chromatin state. Chromatin remodeling factors are ATP-utilizing motor proteins that mediate the interaction of proteins with nucleosomal DNA by DNA/nucleosome-translocation [24]. Brg-1 is a subunit of the Swi/Snf chromatin remodeling complex implicated in the regulation of cellular proliferation and as a tumor suppressor; while hSNF2H is a human homolog of the ISWI family of chromatin remodeling proteins [25,26]. HDAC1 is a general maintenance histone deacetylase that sustains global transcription at a basal level [27]. Consistent with this observation, several repressive chromatin remodeling factors regulating histone H3K9 methylation and mediating chromatin-silencing [18,28,29], including the NAD+-dependent histone deacetylase SIRT1, the histone methyltransferase (HMT) SUV39H1, K9 methylated histone H3 (meH3K9), and the chromatin remodeling factors Brm, were either expressed at low levels (MeH3 and Brm) or localized mostly to the cytoplasm (SIRT1 and SUV39H1) (Figure 1B–1C), suggesting that they remained inactive in RA-induced hESC-derived neuroectodermal cells, similar to the expression patterns in pluripotent hESCs. Our observations suggested that, in spite of neural lineage specification, RA-induced hESC-derived neuroectodermal cells retain an embryonic acetylated chromatin.
Figure 1. NAM-induced cardiomesodermal cells have acquired a silenced chromatin while RA-induced neuroectodermal cells retain an embryonic acetylated chromatin.
(A) NAM (cardiac) or RA (neural) treatment induced differentiation under the defined culture system, as indicated by the appearance of Oct-4 (red) negative cells within the colony. Untreated cells were used as the control.
(B) The strong expression and nuclear localization of a set of active chromatin modifiers in untreated pluripotent hESCs (control), including acetylated histones H3 (acH3, green), acetylated histones H4 (acH4, green), ATP-dependent active chromatin-remodeling factors Brg-1 (red) and hSNF2H (green), and HDAC1 (red), and cytoplasmic localization of histone methyltransferase SUV39H1 (red). NAM-induced cardiomesodermal cells (cardiac) displayed significantly decreased levels of histone H3 and H4 acetylation, and Brg-1 and HDAC1 expression, while RA-induced neuroectodermal cells (neural) retained the similar expression levels and localization patterns. All cells are indicated by DAPI staining of their nuclei (blue) in the insets.
(C) The weak or undetectable expression of repressive chromatin remodeling factor Brm (green) and K9 methylated histone H3 (red).
(D) NAM induces nuclear translocation of NAD-dependent HDAC SIRT1 (red). All cells are shown by DAPI staining (blue) of their nuclei. Scale bars: 10 μm.
By contrast, NAM induced histone deacetylation, significant down-regulation of the expression of Brg-1 and HDAC1, and nuclear translocation of the class III NAD-dependent histone deacetylase SIRT1 (Figure 1B–D). This observation suggests that NAM triggers the activation of SIRT1 and NAD-dependent histone deacetylation that lead to global chromatin silencing yet selective activation of a subset of cardiac-specific genes, and subsequently cardiac fate determination of pluripotent hESCs [12,17]. NAD-dependent SIRT1 is a critical epigenetic regulator previously implicated in cardiovascular and metabolic diseases as well as during embryogenesis [30,31]. NAM acts as a noncompetitive product inhibitor of the forward deacetylation reaction of NAD-dependent SIRT1 and is likely regulating SIRT1 activity in vivo [18].
Down-regulation of a unique set of pluripotence-associated miRNAs upon lineage specification of pluripotent hESCs by small molecule induction
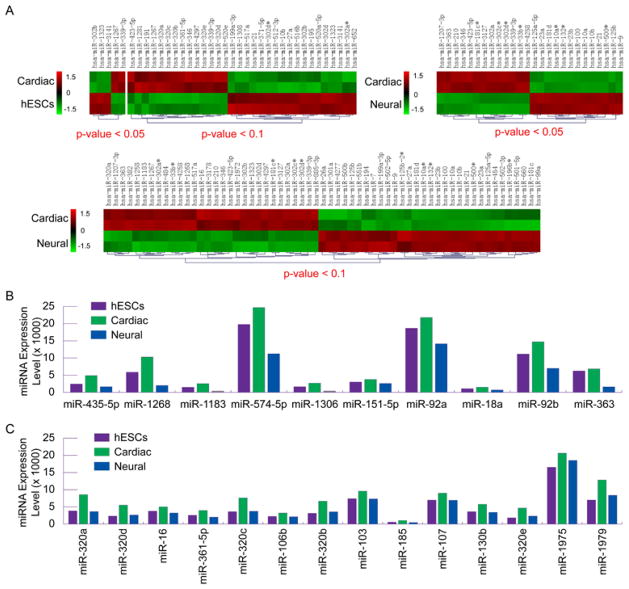
MiRNAs are emerging as important regulators of stem cell pluripotence and differentiation [32–35]. MiRNAs are small, evolutionarily conserved non-coding RNAs that modulate gene expression by inhibiting mRNA translation and promoting mRNA degradation. MiRNAs act as the governors of gene expression networks, thereby modify complex cellular phenotypes in development or disorders [36–38]. MiRNA expression profiling using microarrays is a powerful high-throughput tool capable of monitoring the regulatory networks of the entire genome and identifying functional elements of cellular phenotypes in development and disorders. To identify regulatory networks of small molecule induced lineage specification of pluripotent hESCs, we further performed miRNA microarray analysis (LC Sciences, www.lcsciences.com) to profile the differential expression patterns of miRNAs in hESC cardiac and neural fate choices (Figure 2A). We found that the expression of two most prominent clusters of pluripotence-associated miRNAs hsa-miR-302 and hsa-miR-371/372/373 was significantly suppressed upon lineage-induction of hESCs (Figures 2; 3A and 3C). The cluster of hsa-miR-302 family, which had a profile of the highest expression in pluripotent undifferentiated hESCs, was completely silenced upon neural induction by RA (average ~ 550-fold of down-regulation) (Figures 2A,3A and 3C). A novel group of abundant miRNA clusters in undifferentiated hESCs, including hsa-miR-1308, 3178, 4298, 3195, 1280, 3141, 221/221, and 720, was found to be significantly down-regulated upon small-molecule-induced lineage specification, albeit to less extents (Figures 2 and 3B). The clusters of hsa-miR-17 and hsa-miR- 20, which were strongly expressed in undifferentiated hESCs and which have near-identical seed sequences with hsa-miR-302 family that have been implicated in cell proliferation [33,34], were found to be significantly down-regulated upon RA-induced neural specification but not upon NAM-induced cardiac specification (Figures 2 and 3B). In most cases, significantly higher degrees of down-regulation of pluripotence-associated miRNAs were observed in RA induced neural specification in comparison with NAM-induced cardiac specification (Figures 2 and 3).
Figure 2. Genome-scale miRNA profiling of hESC cardiac and neural specification by small molecule induction.
(A) Hierarchal clustering of differentially expressed miRNAs in undifferentiated hESCs (hESC), cardiac-induced hESCs by NAM (cardiac), and neural-induced hESCs by RA (neural). Statistically significant p values and clustering analysis were provided by LC Sciences as part of miRNA microarray service.
(B) Pie charts showing decreased contributions of a set of pluripotence-associated miRNAs (purple) and increased contributions of distinct sets of cardiac- (green) and neural- (blue) driving miRNAs to the entire miRNA populations upon cardiac and neural induction of pluripotent hESCs by small molecules.
Figure 3. Down-regulation of a unique set of human pluripotence-associated miRNAs upon lineage-specific induction by small molecules.
(A) The expression of two most prominent clusters of pluripotence-associated miRNAs hsa-miR-302 and hsa-miR-371/372/373 was significantly suppressed upon lineage-induction of hESCs by small molecules.
(B) A novel group of miRNA clusters was found to be significantly down-regulated upon small-molecule-induced lineage differentiation, albeit to less extents.
(C) Log2 ratios of down-regulation of the human pluripotence-associated miRNAs. The levels of down-regulation: *: 5–10 fold, **: 10–200 fold, and ***: 200–1000 fold (green lines: cardiac induction by NAM, blue lines: neural induction by RA).
Human embryonic cardiac-driving miRNAs revealed in cardiac fate switch direct of pluripotence
A group of miRNAs displayed an expression pattern of up-regulation upon cardiac induction by NAM and down-regulation upon neural induction by RA (Figure 2,4A and 4B). Among this group of cardiac-driving miRNAs, the clusters of hsa-miR-1268, 574-5p, and 92 families contribute to the highest increased expression profile in NAM-induced cardiac specification of hESCs (Figures 2B and 4B). A second group of miRNAs had an expression pattern of up-regulation upon cardiac induction by NAM but was not significantly affected upon neural induction by RA (Figures 2,4A and 4C). Among the second group of cardiac-driving miRNAs, the clusters of hsa-miR-320 family, 1975, 1979, 103, and 107 contribute to the highest increased expression profile in NAM-induced cardiac specification of hESCs (Figures 2B and 4C). Recently, several groups identified miRNAs as the governors of gene expression in response to myocardial infarction (MI) and during post-MI remodeling of adult hearts [37,38]. Signature patterns of miRNAs identified that miR-1, 29, 30, 133, 150, and 320 were down-regulated, while miR-21, 23a, 125, 195, 199 and 214 were up-regulated during pathological cardiac remodeling in rodents and humans [37,38]. Gain- and loss-of-function studies in mice revealed miR-1 and miR-133 as key regulators in cardiac development and stress-dependent remodeling, miR-138 in control of cardiac patterning, miR-143/145 and miR126 in cardiovascular development and angiogenesis [37,38]. The miR-1 and miR-133 were previously shown to promote mesoderm and muscle differentiation from mouse and human ES cells by repressing nonmuscle gene expression [32]. Our study suggested that a novel set of miRNAs, many of which were not previously linked to cardiac development and function, contribute to the initiation of cardiac fate switch of pluripotent hESCs (Figures 2 and 4).
Figure 4. Up-regulation of a novel set of human embryonic cardiac-driving miRNAs upon cardiac induction by NAM.
(A) Hierarchal clustering of differentially expressed miRNAs in cardiac-induced hESCs by NAM, compared with pluripotent hESCs and neural-induced hESCs by RA. Statistically significant p values and clustering analysis were provided by LC Sciences as part of miRNA microarray service.
(B) A group of cardiac-driving miRNAs displayed an expression pattern of up-regulation upon cardiac induction and down-regulation upon neural induction.
(C) A group of cardiac-driving miRNAs had an expression pattern of up-regulation upon cardiac induction but was not significantly affected upon neural induction.
Human embryonic neural-driving miRNAs revealed in neural fate switch direct of pluripotence
A group of miRNAs displayed an expression pattern of up-regulation upon neural induction by RA and down-regulation upon cardiac induction by NAM (Figures 2,5A,5B, and 5D). Among this group of neural-driving miRNAs, the clusters of hsa-miR-10 family, let-7 family (let-7a, c, d, e, f, g), 21, 100, 125b, 23 family, and 4324 contribute to the highest increased expression profile in RA-induced neural specification of hESCs (Figures 2B,5B and 5D). Notably, the expression of hsa-miR-10 family was silenced in undifferentiated hESCs and displayed a drastic increase (~ 95-fold) upon neural induction of hESCs by RA (Figures 2B,5B and 5D). The miR-10 genes locate within the Hox clusters of developmental regulators [39]. In several species, miR-10 is coexpressed with a set of Hox genes and has been found to repress the translation of Hox transcripts [39]. The enhancer of the mouse Hoxb-1 gene, which controls the RA response and regulates expression predominantly in neuroectoderm, contains a RA response element (RARE) that is not only involved in the ectopic response to RA, but is also essential for establishing the early Hoxb-1 expression pattern [40]. The drastic expression increase of hsa-miR-10 upon exposure of hESCs to RA suggested that RA might induce the expression of Hox genes and co-expression of Hox miRNA hsa-miR- 10 to silence pluripotence-associated genes and miRNA hsa-miR- 302 to drive a neural fate switch of pluripotent hESCs, consistent with our observation of a neuroectodermal phenotype of RA-treated hESCs (Figures 2,3,5A,5B and 5D) [12,16]. The let-7 miRNAs silence the ESC self-renewal program in vivo and in culture, down-regulating pluripotence factors such as Myc and Lin28, an RNA binding protein that inhibits let-7 processing [35]. Let-7b is expressed in adult mammalian brains and exhibits increased expression during neural differentiation of adult CNS-derived neural stem cells [41].
Figure 5. Up-regulation of a novel set of human embryonic neural-driving miRNAs upon neural induction by RA.
(A) Hierarchal clustering of differentially expressed miRNAs in neural-induced hESCs by RA, compared with pluripotent hESCs. Statistically significant p values and clustering analysis were provided by LC Sciences as part of miRNA microarray service.
(B) A group of neural-driving miRNAs displayed an expression pattern of up-regulation upon neural induction and down-regulation upon cardiac induction.
(C) A group of neural-driving miRNAs had an expression pattern of up-regulation upon neural induction but was not significantly affected upon cardiac induction.
(D) Log2 ratios of up-regulation of the human embryonic neural-driving miRNAs. The levels of up-regulation: *: 5–10 fold, **: 10–50 fold, and ***: 50–200 fold.
A second group of miRNAs had an expression pattern of up-regulation upon neural induction by RA but was not significantly affected upon cardiac induction by NAM (Figures 2,5A,5C and 5D). Among the second group of neural-driving miRNAs, the clusters of hsa-miR-181 family, 9, 125a-5p, 99 family, 26 family, 30b, and 335 contribute to the highest increased expression profile in RA-induced neural specification of hESCs (Figures 2B,5C and 5D). The miR-9 is implicated in regulation of spinal motor neuron function previously [36]. Knockdown of miR-9 inhibits hESC-derived human neural progenitor cells (hNPCs) proliferation, delays maturation, and promotes migration in vitro and in vivo [42]. Our study suggested that a distinct set of miRNAs, many of which were not previously linked to neural development and function, contribute to the initiation of neural fate switch of pluripotent hESCs (Figures 2 and 5).
Conclusion
Distinct mechanisms governing small molecule induced cardiac and neural lineage specification of pluripotent hESCs
Due to the prevalence of cardiovascular disease worldwide and acute shortage of donor organs, there is intense interest in developing hESC-based therapies [43,44]. There is also a large unfulfilled need for a clinically-suitable human neuronal cell source for repair or regeneration of the damaged CNS structure and circuitry in today’s healthcare industry [2,12]. Lineage-specific differentiation direct from the pluripotent state of hESCs with small molecule induction not only enables well-controlled efficient derivation of an unlimited supply of human cardiac and neuronal cells across the spectrum of developmental stages for cell-based therapeutics, but also opens the door to investigate human embryonic development using in vitro cellular model systems [12,16,17]. To identify mechanisms of small molecule induced lineage specification of pluripotent hESCs, in this study, we compared the expression and intracellular distribution patterns of a set of cardinal chromatin modifiers. Further, genome-scale profiling of miRNA differential expression patterns was used to identify the regulatory networks in hESC cardiac and neural lineage specification. We found that NAM induced nuclear translocation of NAD-dependent histone deacetylase SIRT1 and global chromatin silencing, while RA induced silencing of pluripotence-associated hsa-miR- 302 family and drastic up-regulation of neuroectodermal Hox miRNA hsa-miR-10 family to high levels (Figures 1–5). Genome-scale miRNA profiling indentified that a unique set of pluripotence-associated miRNAs was down-regulated, while novel sets of distinct cardiac- and neural-driving miRNAs were up-regulated upon the induction of lineage specification direct from the pluripotent state of hESCs (Figures 2–5). In most cases, significantly higher degrees of up or down-regulation of miRNAs were observed in RA induced hESC neural lineage specification in comparison with NAM-induced hESC cardiac lineage specification (Figures 2–5). Taken together, these findings suggest that a predominant epigenetic mechanism via SIRT1-mediated global chromatin silencing governs NAM-induced hESC cardiac fate determination, while a predominant genetic mechanism via silencing of pluripotence-associated hsa-miR-302 family and drastic up-regulation of neuroectodermal Hox miRNA hsa-miR-10 family governs RA-induced hESC neural fate determination. Our study provides critical insight into the earliest events in human embryogenesis as well as offers means for small molecule-mediated direct control and modulation of hESC pluripotent fate when deriving clinically-relevant lineages for regenerative therapies. Finally, the results and conclusion we presented here are based on global profiling of chromatin modifications by immunofluorescence and deconvolution microscopy analysis and large-scale profiling of genomic regulatory networks by miRNA microarray analysis, which is limited by the current resolution of the techniques and which may vary at single gene regulatory level.
Acknowledgments
XHP has been supported by National Institute of Health (NIH) grants from National Institute on Aging (NIHK01AG024496) and The Eunice Kennedy Shriver National Institute of Child Health and Human Development (NIHR21HD056530).
Footnotes
This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
References
- 1.Thomson JA, Itskovitz-Eldor J, Shapiro SS, Waknitz MA, Swiergiel JJ, et al. Embryonic stem cell lines derived from human blastocysts. Science. 1998;282:1145–1147. doi: 10.1126/science.282.5391.1145. [DOI] [PubMed] [Google Scholar]
- 2.Parsons XH, Teng YD, Snyder EY. Important precautions when deriving patient-specific neural elements from pluripotent cells. Cytotherapy. 2009;11:815–824. doi: 10.3109/14653240903180092. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Aubry L, Bugi A, Lefort N, Rousseau F, Peschanski M, et al. Striatal progenitors derived from human ES cells mature into DARPP32 neurons in vitro and in quinolinic acid-lesioned rats. Proc Natl Acad Sci USA. 2008;105:16707–16712. doi: 10.1073/pnas.0808488105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Lee G, Kim H, Elkabetz Y, Al Shamy G, Panagiotakos G, et al. Isolation and directed differentiation of neural crest stem cells derived from human embryonic stem cells. Nat Biotechnol. 2007;25:1468–1475. doi: 10.1038/nbt1365. [DOI] [PubMed] [Google Scholar]
- 5.Roy NS, Cleren C, Singh SK, Yang L, Beal MF, et al. Functional engraftment of human ES cell-derived dopaminergic neurons enriched by coculture with telomerase-immortalized midbrain astrocytes. Nat Med. 2006;12:1259–1268. doi: 10.1038/nm1495. [DOI] [PubMed] [Google Scholar]
- 6.Wernig M, Zhao JP, Pruszak J, Hedlund E, Fu D, et al. Neurons derived from reprogrammed fibroblasts functionally integrate into the fetal brain and improve symptoms of rats with Parkinson’s disease. Proc Natl Acad Sci USA. 2008;105:5856–5861. doi: 10.1073/pnas.0801677105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Kim JB, Sebastiano V, Wu G, Araúzo-Bravo MJ, Sasse P, et al. Oct4- induced pluripotency in adult neural stem cells. Cell. 2009;136:411–419. doi: 10.1016/j.cell.2009.01.023. [DOI] [PubMed] [Google Scholar]
- 8.Park IH, Zhao R, West JA, Yabuuchi A, Huo H, et al. Reprogramming of human somatic cells to pluripotency with defined factors. Nature. 2008;451:141–146. doi: 10.1038/nature06534. [DOI] [PubMed] [Google Scholar]
- 9.Takahashi K, Tanabe K, Ohnuki M, Narita M, Ichisaka T, et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell. 2007;131:861–872. doi: 10.1016/j.cell.2007.11.019. [DOI] [PubMed] [Google Scholar]
- 10.Yu J, Vodyanik MA, Smuga-Otto K, Antosiewicz-Bourget J, Frane JL, et al. Induced pluripotent stem cell lines derived from human somatic cells. Science. 2007;318:1917–1920. doi: 10.1126/science.1151526. [DOI] [PubMed] [Google Scholar]
- 11.Kim JB, Sebastiano V, Wu G, Araúzo-Bravo MJ, Sasse P, et al. Oct4- induced pluripotency in adult neural stem cells. Cell. 2009;136:411–419. doi: 10.1016/j.cell.2009.01.023. [DOI] [PubMed] [Google Scholar]
- 12.Parsons XH, Teng YD, Moore DA, Snyder EY. Patents on technologies of human tissue and organ regeneration from pluripotent human embryonic stem cells. Recent Patents on Regenerative Medicine. 2011;1:142–163. doi: 10.2174/2210297311101020142. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Brons IG, Smithers LE, Trotter MW, Rugg-Gunn P, Sun B, et al. Derivation of pluripotent epiblast stem cells from mammalian embryos. Nature. 2007;448:191–195. doi: 10.1038/nature05950. [DOI] [PubMed] [Google Scholar]
- 14.Chazaud C, Yamanaka Y, Pawson T, Rossant J. Early lineage segregation between epiblast and primitive endoderm in mouse blastocysts through the Grb2-MAPK pathway. Dev Cell. 2006;10:615–624. doi: 10.1016/j.devcel.2006.02.020. [DOI] [PubMed] [Google Scholar]
- 15.Tesar PJ, Chenoweth JG, Brook FA, Davies TJ, Evans EP, et al. New cell lines from mouse epiblast share defining features with human embryonic stem cells. Nature. 2007;448:196–199. doi: 10.1038/nature05972. [DOI] [PubMed] [Google Scholar]
- 16.Parsons XH, Teng YD, Parsons JF, Snyder EY, Smotrich DB, et al. Efficient derivation of human neuronal progenitors and neurons from pluripotent human embryonic stem cells with small molecule induction. J Vis Exp. 2011:e3273. doi: 10.3791/3273. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Parsons XH, Teng YD, Parsons JF, Snyder EY, Smotrich DB, et al. Efficient derivation of human cardiac precursors and cardiomyocytes from pluripotent human embryonic stem cells with small molecule induction. J Vis Exp. 2011:e3274. doi: 10.3791/3274. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Parsons XH, Garcia SN, Pillus L, Kadonaga JT. Histone deacetylation by Sir2 generates a transcriptionally repressed nucleoprotein complex. Proc Natl Acad Sci USA. 2003;100:1609–1614. doi: 10.1073/pnas.0434064100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Horn PJ, Peterson CL. Molecular biology. Chromatin higher order folding--wrapping up transcription. Science. 2002;297:1824–1827. doi: 10.1126/science.1074200. [DOI] [PubMed] [Google Scholar]
- 20.Spivakov M, Fisher AG. Epigenetic signatures of stem-cell identity. Nat Rev Genet. 2007;8:263–271. doi: 10.1038/nrg2046. [DOI] [PubMed] [Google Scholar]
- 21.Azuara V, Perry P, Sauer S, Spivakov M, Jørgensen HF, et al. Chromatin signatures of pluripotent cell lines. Nat Cell Biol. 2006;8:532–538. doi: 10.1038/ncb1403. [DOI] [PubMed] [Google Scholar]
- 22.Bernstein BE, Mikkelsen TS, Xie X, Kamal M, Huebert DJ, et al. A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell. 2006;125:315–326. doi: 10.1016/j.cell.2006.02.041. [DOI] [PubMed] [Google Scholar]
- 23.Mikkelsen TS, Ku M, Jaffe DB, Issac B, Lieberman E, et al. Genome-wide maps of chromatin state in pluripotent and lineage-committed cells. Nature. 2007;448:553–560. doi: 10.1038/nature06008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Haushalter KA, Kadonaga JT. Chromatin assembly by DNA-translocating motors. Nat Rev Mol Cell Biol. 2003;4:613–620. doi: 10.1038/nrm1177. [DOI] [PubMed] [Google Scholar]
- 25.Hendricks KB, Shanahan F, Lees E. Role for BRG1 in cell cycle control and tumor suppression. Mol Cell Biol. 2004;24:362–376. doi: 10.1128/MCB.24.1.362-376.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Loyola A, Huang JY, LeRoy G, Hu S, Wang YH, et al. Functional analysis of the subunits of the chromatin assembly factor RSF. Mol Cell Biol. 2003;23:6759–6768. doi: 10.1128/MCB.23.19.6759-6768.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Huang X, Kadonaga JT. Biochemical analysis of transcriptional repression by Drosophila histone deacetylase 1. J Biol Chem. 2001;276:12497–12500. doi: 10.1074/jbc.C100034200. [DOI] [PubMed] [Google Scholar]
- 28.Harikrishnan KN, Chow MZ, Baker EK, Pal S, Bassal S, et al. Brahma links the SWI/SNF chromatin-remodeling complex with MeCP2-dependent transcriptional silencing. Nat Genet. 2005;37:254–264. doi: 10.1038/ng1516. [DOI] [PubMed] [Google Scholar]
- 29.Vaquero A, Scher M, Erdjument-Bromage H, Tempst P, Serrano L, et al. SIRT1 regulates the histone methyl-transferase SUV39H1 during heterochromatin formation. Nature. 2007;450:440–444. doi: 10.1038/nature06268. [DOI] [PubMed] [Google Scholar]
- 30.Pillarisetti S. A review of Sirt1 and Sirt1 modulators in cardiovascular and metabolic diseases. Recent Pat Cardiovasc Drug Discov. 2008;3:156–164. doi: 10.2174/157489008786263989. [DOI] [PubMed] [Google Scholar]
- 31.Borradaile NM, Pickering JG. NAD(+), sirtuins, and cardiovascular disease. Curr Pharm Des. 2009;15:110–117. doi: 10.2174/138161209787185742. [DOI] [PubMed] [Google Scholar]
- 32.Ivey KN, Muth A, Arnold J, King FW, Yeh RF, et al. MicroRNA regulation of cell lineages in mouse and human embryonic stem cells. Cell Stem Cell. 2008;2:219–229. doi: 10.1016/j.stem.2008.01.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Laurent LC, Chen J, Ulitsky I, Mueller FJ, Lu C, et al. Comprehensive microRNA profiling reveals a unique human embryonic stem cell signature dominated by a single seed sequence. Stem Cells. 2008;26:1506–1516. doi: 10.1634/stemcells.2007-1081. [DOI] [PubMed] [Google Scholar]
- 34.Marson A, Levine SS, Cole MF, Frampton GM, Brambrink T, et al. Connecting microRNA genes to the core transcriptional regulatory circuitry of embryonic stem cells. Cell. 2008;134:521–533. doi: 10.1016/j.cell.2008.07.020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Melton C, Judson RL, Blelloch R. Opposing microRNA families regulate self-renewal in mouse embryonic stem cells. Nature. 2010;463:621–626. doi: 10.1038/nature08725. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Haramati S, Chapnik E, Sztainberg Y, Eilam R, Zwang R, et al. miRNA malfunction causes spinal motor neuron disease. Proc Natl Acad Sci USA. 2010;107:13111–13116. doi: 10.1073/pnas.1006151107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Liu N, Olson EN. MicroRNA regulatory networks in cardiovascular development. Dev Cell. 2010;18:510–525. doi: 10.1016/j.devcel.2010.03.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Williams AH, Liu N, van Rooij E, Olson EN. MicroRNA control of muscle development and disease. Curr Opin Cell Biol. 2009;21:461–469. doi: 10.1016/j.ceb.2009.01.029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Yekta S, Tabin CJ, Bartel DP. MicroRNAs in the Hox network: an apparent link to posterior prevalence. Nat Rev Genet. 2008;9:789–796. doi: 10.1038/nrg2400. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Marshall H, Studer M, Pöpperl H, Aparicio S, Kuroiwa A, et al. A conserved retinoic acid response element required for early expression of the homeobox gene Hoxb-1. Nature. 1994;370:567–571. doi: 10.1038/370567a0. [DOI] [PubMed] [Google Scholar]
- 41.Zhao C, Sun G, Li S, Lang MF, Yang S, et al. MicroRNA let-7b regulates neural stem cell proliferation and differentiation by targeting nuclear receptor TLX signaling. Proc Natl Acad Sci USA. 2010;107:1876–1881. doi: 10.1073/pnas.0908750107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Delaloy C, Liu L, Lee JA, Su H, Shen F, et al. MicroRNA-9 coordinates proliferation and migration of human embryonic stem cell-derived neural progenitors. Cell Stem Cell. 2010;6:323–335. doi: 10.1016/j.stem.2010.02.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Passier R, van Laake LW, Mummery CL. Stem-cell-based therapy and lessons from the heart. Nature. 2008;453:322–329. doi: 10.1038/nature07040. [DOI] [PubMed] [Google Scholar]
- 44.Zhu WZ, Hauch KD, Xu C, Laflamme MA. Human embryonic stem cells and cardiac repair. Transplant Rev (Orlando) 2009;23:53–68. doi: 10.1016/j.trre.2008.05.005. [DOI] [PMC free article] [PubMed] [Google Scholar]