Abstract
Paenibacillus sp.Y412MC10 was one of a number of organisms isolated from Obsidian Hot Spring, Yellowstone National Park, Montana, USA under permit from the National Park Service. The isolate was initially classified as a Geobacillus sp. Y412MC10 based on its isolation conditions and similarity to other organisms isolated from hot springs at Yellowstone National Park. Comparison of 16 S rRNA sequences within the Bacillales indicated that Geobacillus sp.Y412MC10 clustered with Paenibacillus species, and the organism was most closely related to Paenibacillus lautus. Lucigen Corp. prepared genomic DNA and the genome was sequenced, assembled, and annotated by the DOE Joint Genome Institute. The genome sequence was deposited at the NCBI in October 2009 (NC_013406). The genome of Paenibacillus sp. Y412MC10 consists of one circular chromosome of 7,121,665 bp with an average G+C content of 51.2%. Comparison to other Paenibacillus species shows the organism lacks nitrogen fixation, antibiotic production and social interaction genes reported in other paenibacilli. The Y412MC10 genome shows a high level of synteny and homology to the draft sequence of Paenibacillus sp. HGF5, an organism from the Human Microbiome Project (HMP) Reference Genomes. This, combined with genomic CAZyme analysis, suggests an intestinal, rather than environmental origin for Y412MC10.
Keywords: Geobacillus sp. Y412MC10, Paenibacillus sp. Y412MC10, Obsidian Hot Spring
Introduction
Numerous novel microorganisms have been isolated from hot springs in Yellowstone National Park. Many of these organisms have been shown to possess enzymes with significant potential in biotechnological applications [1]. Among the organisms first isolated from Yellowstone hot springs are Thermus aquaticus [2,3], Thermus brockianus [4], Acidothermus cellulollyticus [5], and Synechococcus species [6]. As part of a project in conjunction with the Department of Energy Joint Genome Institute, Lucigen Corp. isolated, characterized, and sequenced a number of new isolates from Yellowstone hot springs. The bacterial isolate Y412MC10 was one of four microorganisms isolated from Obsidian Hot Spring, Yellowstone National Park, Montana, USA and submitted for whole genome sequencing. Y412MC10 was initially classified as a Geobacillus sp. based on its isolation conditions and morphological similarity to other organisms such as Geobacillus species Y412MC61 (GenBank 544556), Y412MC52 (GenBank 550542), and Geobacillus thermoglucosidasius C56-YS93 (GenBank 634956). The Geobacillus sp.Y412MC10 draft genome sequence was deposited at the NCBI in October 2009 (NC_013406) with the lineage entry indicating that it is a Geobacillus. Following assembly of the complete genome of Y412MC10, the 16S rRNA sequence and genome properties properly assigned the organism as a Paenibacillus sp. Y412MC10 represents the first Paenibacillus sp. isolated from a hot spring to have its genome completely sequenced.
Paenibacillus sp. were originally grouped in the genus Bacillus until 1993, when Ash et al. [7] proposed that members of "group 3" should be transferred to the genus Paenibacillus, and proposed Paenibacillus polymyxa as the type species. Paenibacillus sp. have been isolated from a wide range of environments including soil [8], the Antarctic [9], and the oral cavity of a dog [10]. Paenibacillus sp. are of interest for a number of reasons, including production of antibiotics [11-13], biopolymer-degrading enzymes [14-16], and their ability to fix nitrogen [17,18]. One species, P. vortex shows highly unusual organized growth morphologies on solid surfaces [19,20]; another species, P. dendritiformis, also shows unusual growth morphologies on solid surfaces [19,21,22]. Comparison of the genetic content of Y412MC10 with genomes of Paenibacillus sp. from other environments will give insights into the evolutionary adaptations that have occurred in the Paenibacillus. The organism also may be a source of novel polysaccharide degrading enzymes for use in biomass degradation.
Classification and features
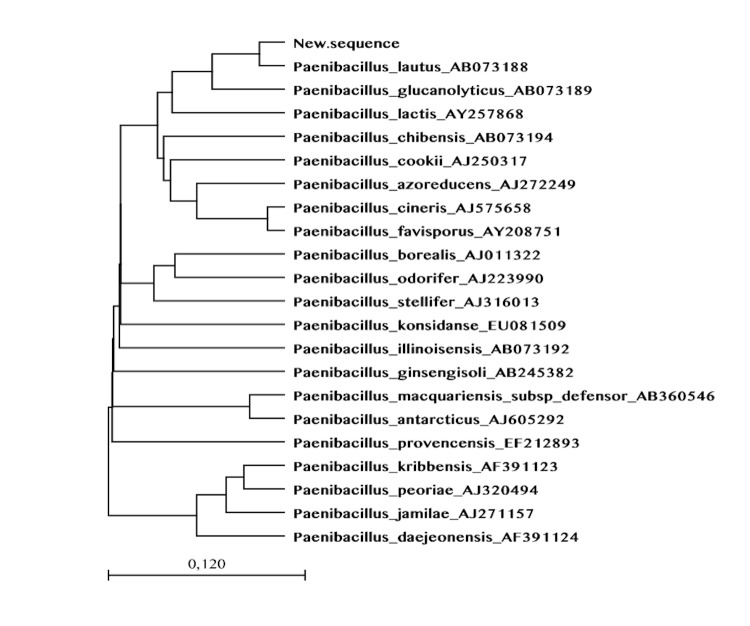
A phylogenetic tree was constructed to identify the family relationship of strain Y412MC10 (Figure 1). The tree was created using BLAST2Tree software [23]. The analysis was carried out using only type strains of validly-named organisms, and the analysis shows that Y412MC10 does not clade with known Geobacillus species. Rather, Y412MC10 clades in the Paenibacillus genus. Based on r16S analysis of validly-named organisms, Y412MC10 is most closely related to Paenibacillus lautus DSM 3055T (AB073188). The classification of the isolate was confirmed using the EzTaxon-e server [24], again on the basis of 16S rRNA sequence data. When compared to their entire r16S database, Y412MC10 was identified as being a strain of Paenibacillus lautus with 99.09% identity and 100% completeness to the r16S of the type strain, Paenibacillus lautus NRRL NRS-666 GenBank D78472.
Figure 1.
Phylogenetic tree highlighting the position of Paenibacillus lautus strain Y412MC10 and relative strains within the Bacillales. Strains used for the comparison (16S rRNA accession number) were (Paenibacillus Y412MC10); Paenibacillus lautus DSM 3035T (AB073188); Paenibacillus glucanolyticus DSM 5162T (AB073189); Paenibacillus illinoisensis DSM 11733T (AB073192); Paenibacillus chibensis DSM 11731T (AB073194); Paenibacillus ginsengisoli LMG 23406 T (AB245382); Paenibacillus macquariensis subsp defensor NCIB 14397 T (AB360546); Paenibacillus kribbensis KCTC 0766BP T (AF391123); Paenibacillus daejeonensis KCCM 41557 T (AF391124); Paenibacillus borealis DSM 13188 T (AJ011322); Paenibacillus odorifer DSM 15391 T (AJ223990); Paenibacillus cookie LMG 18419 T (AJ250317); Paenibacillus jamilae DSM 13815 T (AJ271157); Paenibacillus azoreducens DSM 13822 T (AJ272249); Paenibacillus stellifer DSM 14472 T (AJ316013); Paenibacillus peoriae DSM 8320 T (AJ320494); Paenibacillus cineris LMG 18349 T (AJ575658); Paenibacillus antarcticus LMG 22078 T (AJ605292); Paenibacillus favisporus LMG 20987 T (AY208751); Paenibacillus lactis DSM 15596 T (AY257868); Paenibacillus provencensis CIP 109358T (EF212893); Paenibacillus konsidanse KCTC 13165T (EU081509).
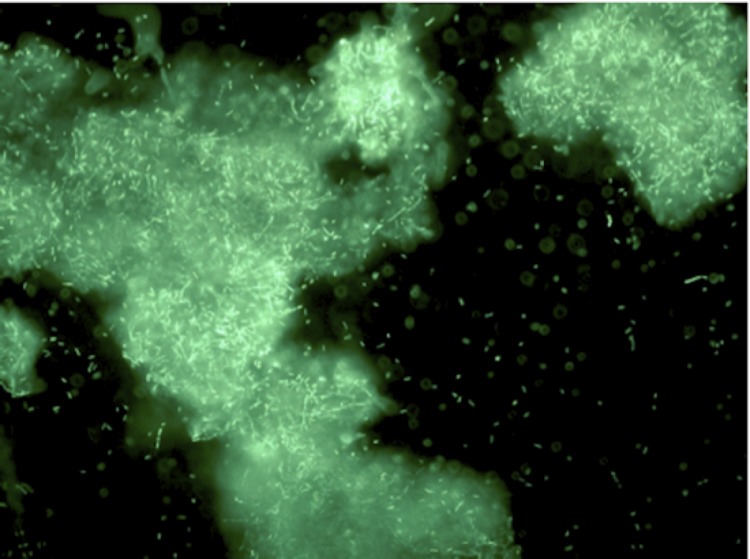
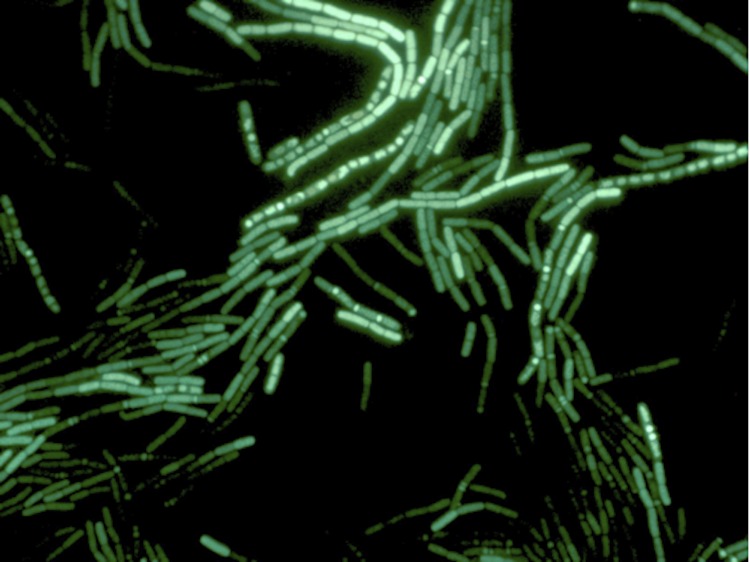
Paenibacillus sp. Y412MC10 was one of a number of organisms isolated from Obsidian Hot Spring, Yellowstone National Park, Montana, USA (44.6100594° latitude and -110.4388217° longitude) under a sampling permit from the National Park Service. The hot spring possesses a pH of 6.37 and a temperature range of 42-90°C. The organism was isolated from a sample of hot spring water by enrichment and plating on YTP-2 medium [25] (YTP-2 media contains (per liter) 2.0 g yeast extract, 2.0 g tryptone, 2.0 g sodium pyruvate, 1.0 g KCl, 2.0 g KNO3, 2.0 g Na2HPO4.7H2O, 0.1 g MgSO4, 0.03 g CaCl2, and 2.0 ml clarified tomato juice) at 50°C. Culture stocks were routinely maintained on YT (containing (per liter) 5.0 g yeast extract, 8.0 g tryptone, and 2.5 g NaCl) agar plates at 37°C. As part of the sequencing agreement with the Joint Genome Institute, the culture is available without restrictions from the authors. Lucigen, the National Park Service, and the Joint Genome Institute have placed no restrictions on the use of the culture or sequence data. Y412MC10 is a Gram-positive facultative anaerobe (Table 1) that grows well on a wide variety of standard lab media (YT, TB, LB). On plates, the organism grows as rods or chains of rods (Figure 2A). After growth for 6 days on plates, the cells still appear rod-shaped, but an extracellular matrix appears to surround and bind the individual cells together (light green background, Figure 2B). In liquid culture, the organism appears to also grow as a mixture of single cells and large clumps of cells surrounded by an extracellular matrix (Figure 2C). Prolonged growth on plates or in liquid culture results in sporulation of the culture; spores are subterminal with swollen sporangia.
Table 1. Classification and general features of Paenibacillus strain Y412MC10.
| MIGS ID | Property | Term | Evidence code |
|---|---|---|---|
| Current classification | Domain Bacteria | TAS [26] | |
| Phylum Firmicutes | TAS [27-29] | ||
| Class Bacilli | TAS [30,31] | ||
| Order Bacillales | TAS [32,33] | ||
| Family Bacillaceae | TAS [32,34] | ||
| Genus Paenibacillus | TAS [35-39] | ||
| Species Paenibacillus lautus | TAS [40] | ||
| Strain Y412MC10 | IDA | ||
| Gram stain | positive | IDA | |
| Cell shape | rods and chains of rods | IDA | |
| Motility | motile | IDA | |
| Sporulation | sporulating | IDA | |
| Temperature range | mesophilic | IDA | |
| Optimum temperature | 37°C | IDA | |
| MIGS-22 | Oxygen requirement | facultative anaerobe | IDA |
| Carbon source | Carbohydrate or protein | IDA | |
| Energy source | chemoorganotrophic | IDA | |
| Electron acceptor | Oxygen, nitrate | IDA | |
| MIGS-6 | Habitat | hot spring | IDA |
| MIGS-6.3 | Salinity | Grows in 3% NaCl | IDA |
| MIGS-15 | Biotic relationship | free-living | IDA |
| MIGS-14 | Pathogenicity | None, BSL1 | IDA |
| Isolation | Obsidian spring | IDA | |
| MIGS-4 | Geographic location | Yellowstone National Park | IDA |
| MIGS-5 | Sample collection time | September 2003 | IDA |
| MIGS-4.1 | Latitude | 44.6100594 | TAS [1] |
| MIGS-4.2 | Longitude | -110.4388217 | TAS [1] |
| MIGS-4.3 | Depth | Surface of spring | IDA |
| MIGS-4.4 | Altitude | 2416 m | TAS [1] |
Evidence codes - IDA: Inferred from Direct Assay; TAS: Traceable Author Statement (i.e., a direct report exists in the literature); NAS: Non-traceable Author Statement (i.e., not directly observed for the living, isolated sample, but based on a generally accepted property for the species, or anecdotal evidence). These evidence codes are from the Gene Ontology project [41].
Figure 2A.
Micrograph of Paenibacillus strain Y412MC10 cells showing individual cells and chains of cells. Cells were streaked on YT agar and incubated 18 hr. at 37°C. A colony was removed, re-suspended in sterile water and stained using a 5 μM solution of SYTO® 9 fluorescent stain in sterile water (Molecular Probes). Dark field fluorescence microscopy was performed using a Nikon Eclipse TE2000-S epifluorescence microscope at 2000× magnification using a high-pressure Hg light source and a 500 nm emission filter.
Figure 2B.
Micrograph of same plate of Paenibacillus strain Y412MC10 cells showing individual cells and chains of cells. Same plate as Figure 2A, but incubated 6 days at 37°C.
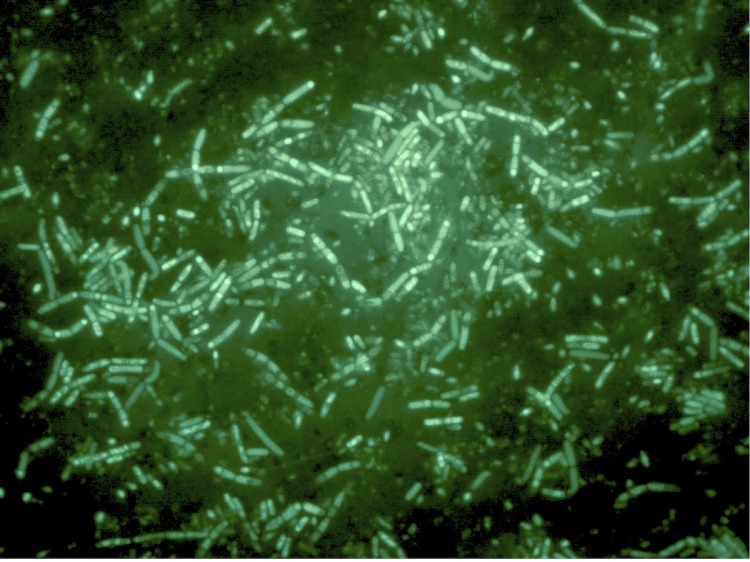
Figure 2C.
Micrograph of Paenibacillus strain Y412MC10 cells showing individual cells and clumps of cells. Cells were grown in YTP-2 media for 18 hours at 37°C and 200 rpm. An aliquot was removed and stained using a 5 μM solution of SYTO® 9 fluorescent stain in sterile water (Molecular Probes). Dark field fluorescence microscopy was performed using a Nikon Eclipse TE2000-S epifluorescence microscope at 200× magnification using a high-pressure Hg light source and 500 nm emission filter.
Genome sequencing and annotation
Genome project history
Paenibacillus sp.Y412MC10 was selected for sequencing on the basis of its biotechnological potential as part of the U.S. Department of Energy's Genomic Science program (formerly Genomics:GTL). The genome sequence is deposited in the Genomes On Line Database [42] (GOLD ID = Gc01127), and in GenBank (NCBI Reference Sequence = NC_013406). Sequencing, finishing and annotation were performed by the DOE Joint Genome Institute (JGI). A summary of the project information and its association with MIGS identifiers is shown in Table 2.
Table 2. Genome sequencing and project information.
| MIGS ID | Property | Term |
|---|---|---|
| MIGS-31 | Finishing quality | Finished |
| MIGS-28 | Libraries used | 6 kb and 34 kb |
| MIGS-29 | Sequencing platforms | ABI3730, 454 Titanium, Illumina GAii |
| MIGS-31.2 | Fold coverage | 5.8 |
| MIGS-30 | Assemblers | Phred/Phrap/Consed |
| MIGS-32 | Gene calling method | Prodigal, GenePRIMP |
| GenBank ID | CP001793.1 | |
| GenBank Date of Release | October 7, 2009 | |
| GOLD ID | Gi02010 | |
| Project relevance | Biotechnological |
Growth conditions and DNA Isolation
For preparation of genomic DNA, liter cultures of Y412MC10 were grown from a single colony in YTP-2 medium at 37°C in flasks agitated at 200 rpm and collected by centrifugation. The cell concentrate was lysed using a combination of SDS and proteinase K, and genomic DNA was isolated using a phenol/chloroform extraction [43]. The genomic DNA was precipitated, and treated with RNase to remove residual contaminating RNA.
Genome sequencing and assembly
The genome of Paenibacillus lautus Y412MC10 was sequenced at the Joint Genome Institute (JGI) [44] using Sanger sequencing with a combination of 6 kb and 34 kb DNA libraries and 454 FLX pyrosequencing done to a depth of 20× coverage [45]. Both libraries provided 5.8× coverage of the genome. Draft assemblies were based on 39,162 total reads. Solexa sequencing data was used to polish the assembly. All general aspects of library construction and sequencing performed at the JGI can be found at their website. The Phred/Phrap/Consed software package [46] was used to assemble 6-kb and fosmid libraries and to assess quality. Possible mis-assemblies were corrected; gaps between contigs were closed by 2,744 primer walks from sub-clones or 83 PCR end reads, 5 mini-libraries, and 10 PCR shatter libraries. The error rate of the completed genome sequence was 0.08, based on 49,558 total reads. Table 2 presents the project information and its association with MIGS version 2.0 compliance [47].
Genome annotation
Genes were identified using Prodigal [48] as part of the Oak Ridge National Laboratory genome annotation pipeline, followed by a round of manual curation using the JGI GenePRIMP pipeline [49]. The predicted CDSs were translated and used to search the National Center for Biotechnology Information (NCBI) nonredundant database, UniProt, TIGRFam, Pfam, PRIAM, KEGG, COG, and InterPro databases. These data sources were combined to assert a product description for each predicted protein. Non-coding genes and miscellaneous features were predicted using tRNAscan-SE [49], RNAMMer [50], Rfam [51], TMHMM [52], and signalP [52].
Genome properties
The genome of Paenibacillus lautus Y412MC10 consists of one circular chromosome of 7,121,665 bp with an average G+C content of 51.2% (Table 3 and Figure 3). There are 73 tRNA genes, 24 rRNA genes and 4 “other” identified RNA gene. There are 6,343 predicted protein-coding regions and 105 pseudogenes in the genome. A total of 4,651 genes (72.2%) have been assigned a predicted function while the rest have been designated as hypothetical proteins. The numbers of genes assigned to each COG functional category are listed in Table 4. About 20% of the annotated genes were either not assigned to a COG or have an unknown function.
Table 3. Genome statistics.
| Attribute | Value | % of totala |
|---|---|---|
| Genome size (bp) | 7,121,665 | 100.0 |
| DNA coding region (bp) | 6,141,611 | 86.2 |
| DNA G+C content (bp) | 3,649,102 | 51.2 |
| Number of replicons | ||
| Total genes | 6,444 | 100.0 |
| RNA genes | 101 | 1.6 |
| rRNA operons | ||
| Protein-coding genes | 6,343 | 98.4 |
| Pseudogenes | ||
| Genes in paralog clusters | 1,599 | 24.8 |
| Genes assigned to COGs | 4,651 | 72.2 |
| Genes with signal peptides | 1,506 | 23.4 |
| Genes with transmembrane helices | 1,921 | 29.8 |
| Paralogous groups | 532 |
a) The total is based on either the size of the genome in base pairs or the total number of protein coding genes in the annotated genome.
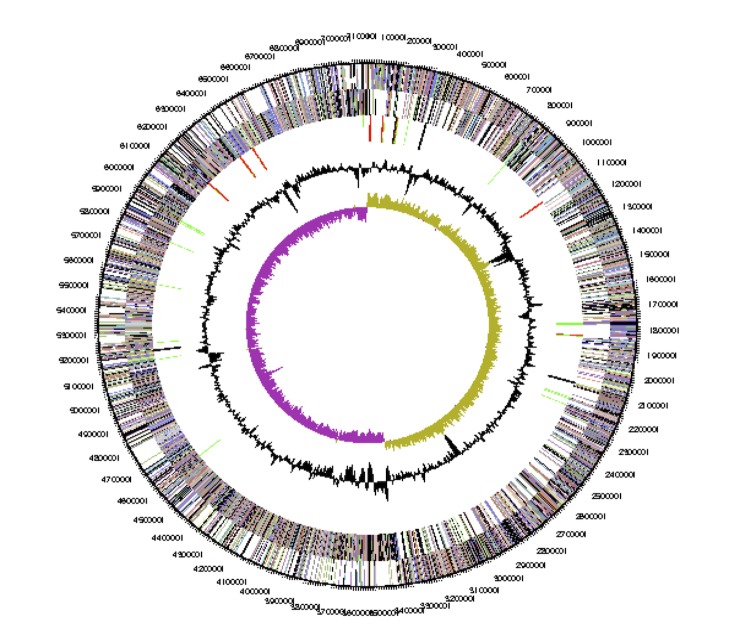
Figure 3.
Graphical circular map of the chromosome. From outside to the center: Genes on forward strand (color by COG categories), Genes on reverse strand (color by COG categories), RNA genes (tRNAs green, rRNAs red, other RNAs black), GC content, GC skew.
Table 4. Number of genes associated with the 25 general COG functional categories.
| Code | Value | %age | Description |
|---|---|---|---|
| J | 224 | 3.6 | Translation, ribosomal structure and biogenesis |
| A | RNA processing and modification | ||
| K | 663 | 10.6 | Transcription |
| L | 152 | 3.2 | Replication, recombination and repair |
| B | 2 | 0.03 | Chromatin structure and dynamics |
| D | 213 | 3.4 | Cell cycle control, cell division, chromosome partitioning |
| Y | Nuclear structure | ||
| V | 244 | 3.9 | Defense mechanisms |
| T | 480 | 7.7 | Signal transduction mechanisms |
| M | 440 | 7.0 | Cell wall/membrane/envelope biogenesis |
| N | 130 | 2.1 | Cell motility |
| Z | 5 | 0.1 | Cytoskeleton |
| W | 0 | 0.0 | Extracellular structures |
| U | 41 | 0.7 | Intracellular trafficking, secretion, and vesicular transport |
| O | 331 | 5.3 | Posttranslational modification, protein turnover, chaperones |
| C | 415 | 6.6 | Energy production and conversion |
| G | 1,030 | 16.5 | Carbohydrate transport and metabolism |
| E | 788 | 12.6 | Amino acid transport and metabolism |
| F | 244 | 3.9 | Nucleotide transport and metabolism |
| H | 373 | 6.0 | Coenzyme transport and metabolism |
| I | 88 | 1.4 | Lipid transport and metabolism |
| P | 559 | 9.0 | Inorganic ion transport and metabolism |
| Q | 283 | 4.5 | Secondary metabolites biosynthesis, transport and catabolism |
| R | 872 | 14.0 | General function prediction only |
| S | 371 | 6.0 | Function unknown |
| - | 0 | 0.0 | Not in COGs |
The total is based on the total number of protein coding genes in the annotated genome. Data from [53].
Insights from the genome sequence
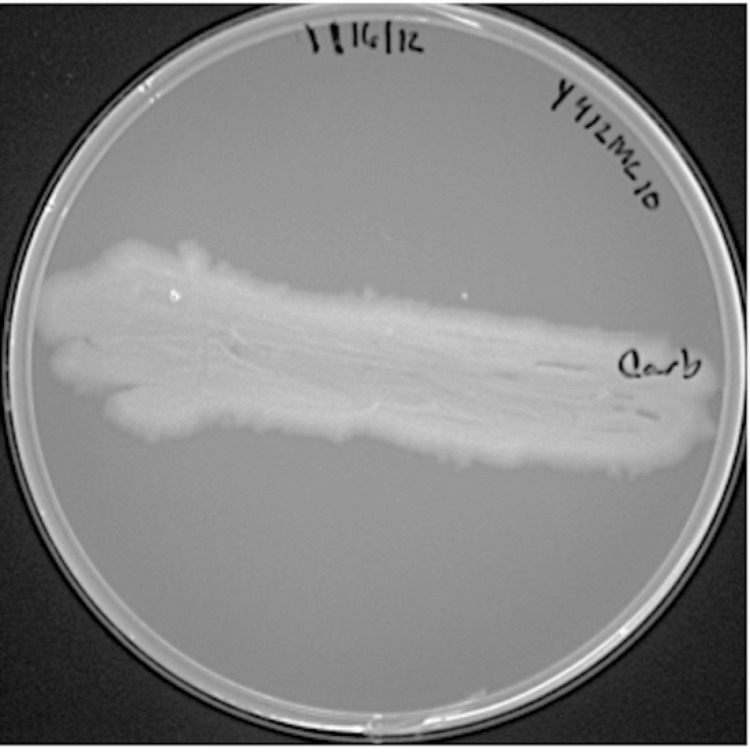
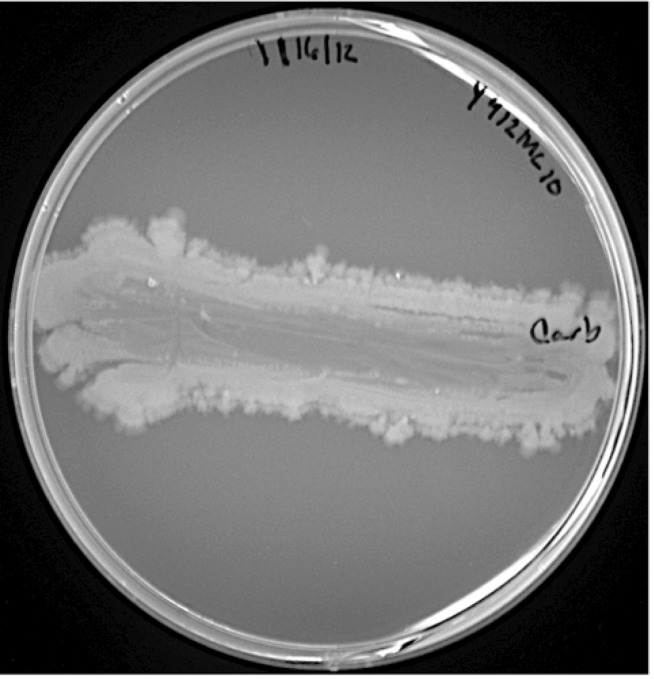
Motility of Paenibacillus cells on solid media has been observed with a number of species. P. lautus is reported to spread across plates 69]. P. vortex shows highly unusual organized growth morphologies on solid surfaces [19,20] forming complex patterns on the plate. Another species, P. dendritiformis, also shows unusual growth morphologies on solid surfaces [19,21,22]. Y412MC10 was evaluated for spreading behavior on plates; the results ((Figure 4A Figure 4B Figure 4C) show definite spreading behavior for Y412MC10. The spreading behavior does not, however, appear to be as complex as reported for P. vortex and P. dendritiformis.
Figure 4A.
Photograph of Paenibacillus sp. Y412MC10 streaked on YT agar containing 100 mg/l carbenicillin and incubated at 37°C for 18 hours.
Figure 4B.
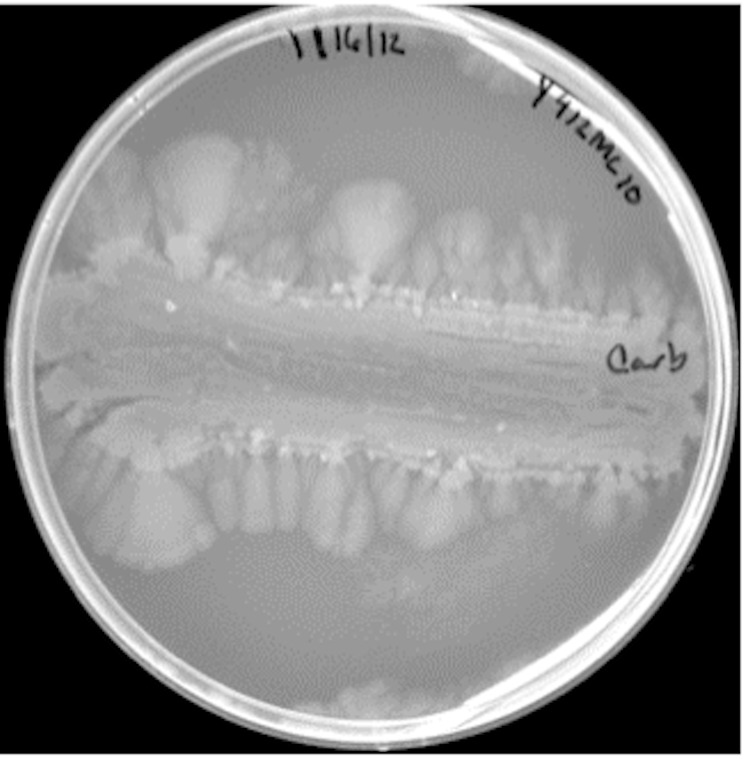
Photograph of Paenibacillus sp. Y412MC10 streaked on YT agar and incubated at 37°C for 50 hours. Note continued clearing of center area and significant spreading of outside edges of culture.
Figure 4C.
Photograph of Paenibacillus sp. Y412MC10 streaked on YT agar and incubated at 37°C for 168 hours. Note continued clearing of center area and significant spreading of outside edges of culture.
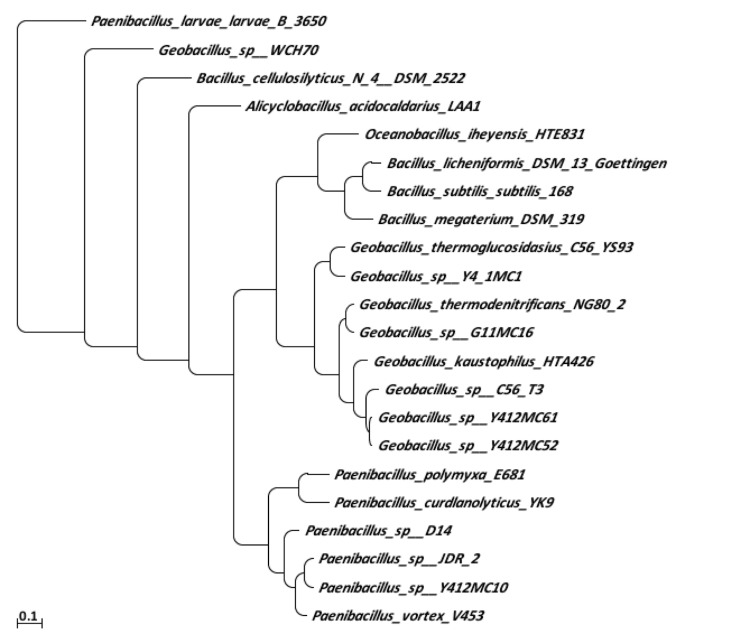
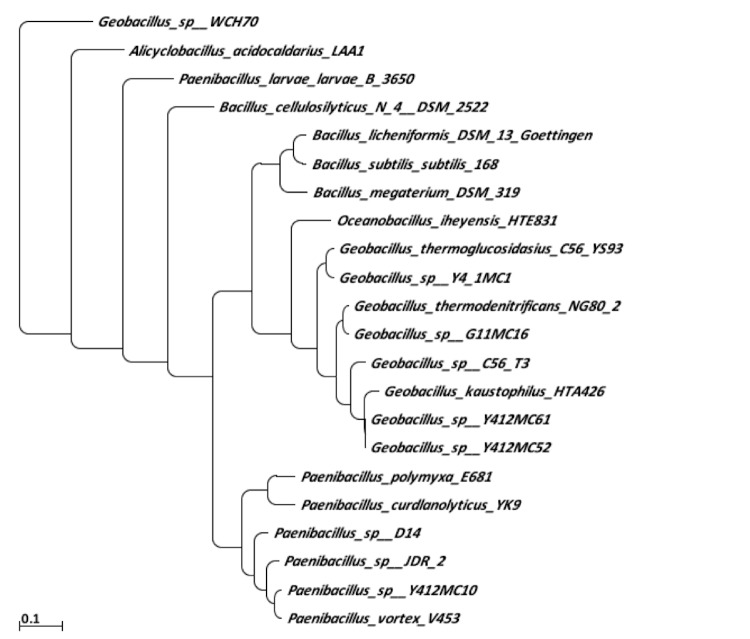
To further understand the phylogenetic relationships of Y412MC10, a separate phylogenetic tree was constructed of Paenibacillus sp. having either complete or draft genome. This was necessary because there is little or no overlap between validly-named Paenibacillus strains and the Paenibacillus strains submitted for whole genome sequencing. The tree was created on the IMG website [54] using the alignment of 16S genes based on the SILVA database and dnadist and neighbor tools from the Phylip package [55]. Paenibacillus sp. JDR-2, Paenibacillus curdlanolyticus YK9, Paenibacillus polymyxa E681, Paenibacillus polymyxa SC2, and Paenibacillus mucilaginosus KNP414 are all species isolated from soil or plant roots. Paenibacillus vortex V453, Paenibacillus sp. HGF5, Paenibacillus sp. HGF7, are oral or intestinal human isolates. Paenibacillus larvae subsp. larvae B-3650, 741161 is a honey bee pathogen that attacks bee larvae. The r16S analysis (Figure 5) shows Y412MC10 is most closely related to Paenibacillus sp. HGF5 (NCBI Taxon ID 908341, Gold ID Gi05716), an organism being sequenced as part of the Human Microbiome Project (HMP) Reference Genomes (http://www.hmpdacc.org/reference_genomes/reference_genomes.php).
Figure 5.
Phylogenetic tree highlighting the position of Paenibacillus sp. Y412MC10 and relative strains within the Bacillales. The strains and their corresponding GenBank taxonomy ID numbers are: Paenibacillus sp. JDR-2, 324057; Paenibacillus curdlanolyticus YK9, A717606; Paenibacillus sp. Y412MC10, 481743; Paenibacillus vortex V453, 715225; Paenibacillus polymyxa E681, 349520; Paenibacillus polymyxa SC2, 886882; Paenibacillus mucilaginosus KNP414, 1036673; Paenibacillus sp. HGF5, 908341; Paenibacillus sp. HGF7, 944559; Paenibacillus larvae subsp. larvae B-3650, 741161; Bacillus subtilis subtilis 168, 2243082.
COG [Figure 6] and TIGRfam [Figure 7] whole genome comparisons were carried out between Y412MC10 and draft and finished genomes of closely related organisms using IMG software [56]. The results of the COGs and TIGRfam whole genome comparisons place Y412MC10 clearly among the Paenibacillus species, in agreement with the results from 16S analysis. The 16S analysis shows Y412MC10 is most closely related to P. vortex and P. sp. HGF5, both human isolates, and then to the two P. polymyxa sp. In both whole genome analyses, Y412MC10 is again most closely related to Paenibacillus vortex and Paenibacillus sp. HGF5. In the COG comparison, the other human isolate, P. sp. HGF7, is not closely related to P. vortex and P. sp. HGF5; in the TIGRfam comparison, HGF7 clades closely with Y412MC10, P. vortex and P. sp. HGF5. These results also suggest a mammalian, rather than environmental, ecosystem as the home of Y412MC10.
Figure 6.
COGs whole genome comparison of selected strains. Comparison was performed as described in text; organisms and GenBank accession numbers are described in Figure 5.
Figure 7.
TIGRfam whole genome comparison of selected strains. Comparison was performed as described in text; organisms and GenBank accession numbers are described in Figure 5.
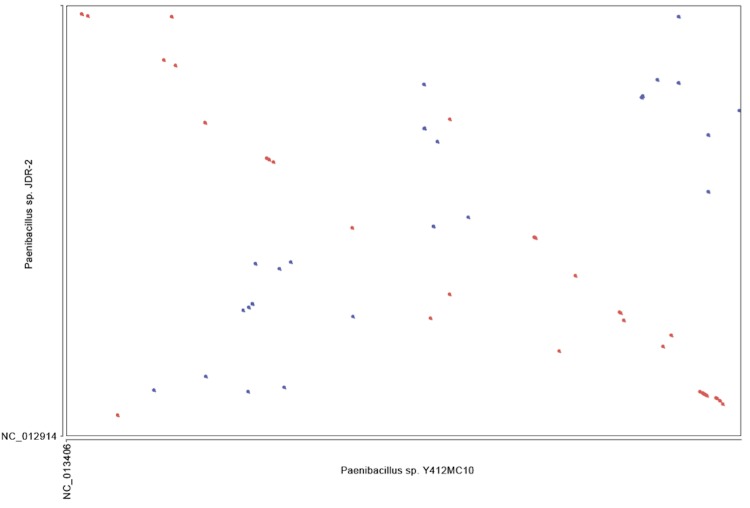
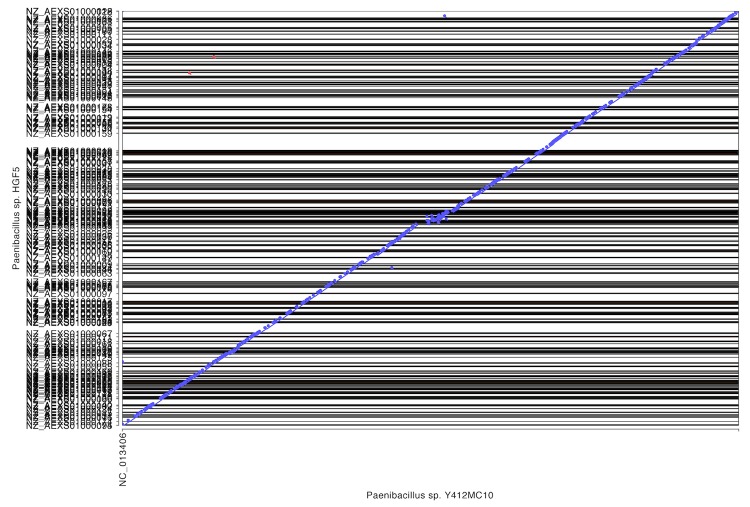
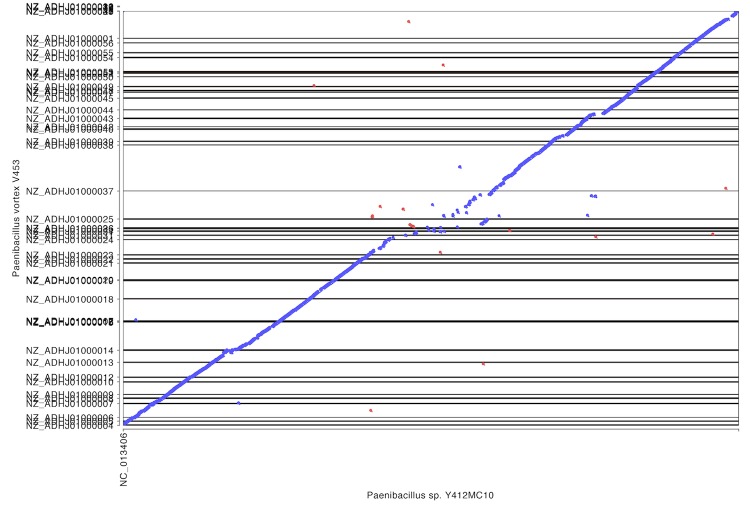
To further understand the relationship between these organisms, whole genome alignments were performed using Mummer software to generate dot plot diagrams comparing pairs of genomes on the IMG website [57] using input DNA sequences directly (NUCmer). The close relationship between the genome of Y412MC10 and the genomes of HGF5 and P. vortex is reflected in the high levels of homology and synteny seen (Figure 8, Figure 9) with these two human isolates.
Figure 8.
Dot plot comparison of Y412MC10 and HGF5 performed on IMG website. HGF5 draft genome contains 185 scaffolds.
Figure 9.
Dot plot comparison of Y412MC10 and P. vortex performed on IMG website. P. vortex draft genome contains 56 scaffolds.
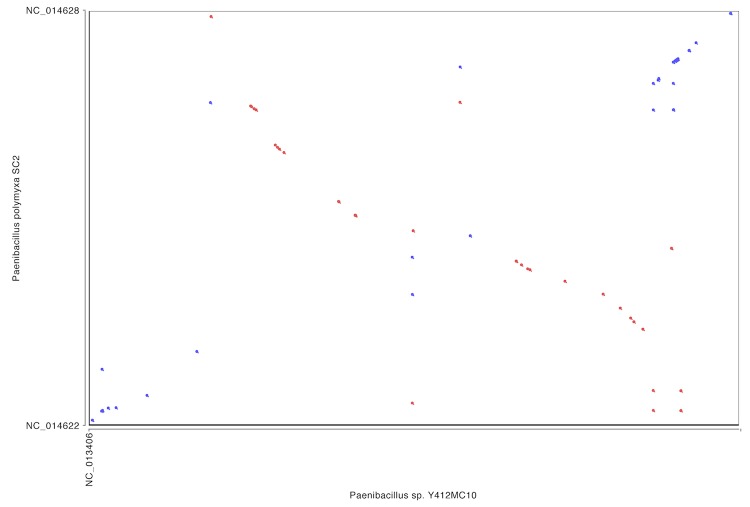
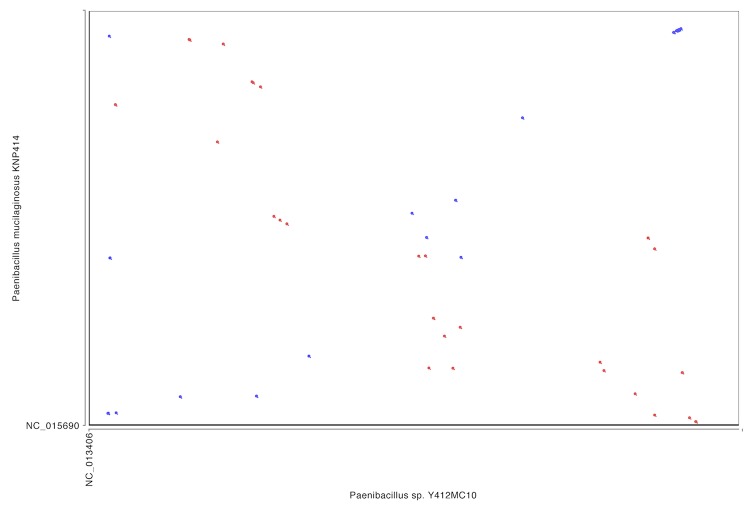
In comparison, whole genome alignment of Y412MC10 with the genomes of P. polymyxa and P. mucilaginosus show little homology or synteny between Y412MC10 and the two soil organisms (Figure 10, Figure11).
Figure 10.
Dot plot comparison of Y412MC10 and P. polymyxa performed on IMG website.
Figure 11.
Dot plot comparison of Y412MC10 and P mucilaginosus performed on IMG website.
The similarity between the r16S sequences of P. lautus and Y412MC10 led us to examine if biochemical evidence suggested a similar habit for both. Bacillus lautus was first isolated from the intestinal tract of children [58]; later, the identity of the organism was re-confirmed and the organism was reclassified and renamed Paenibacillus lautus. Examination of the genome of Y412MC10 lends support to the hypothesis that Y412MC10 also has an intestinal origin. An analysis of the carbohydrate active enzymes (CAZY [59]) shows very low levels of GH family 5, 6, 8, 9, 10, 11, and 48 as well as no CBM 2 or 3 members, suggesting an inability to significantly degrade cellulose and hemicellulose components of biomass. CAZy analysis shows a genome enriched in GH29 and GH95 α-fucosidases; the genome is also enriched in GH38 and GH125 α-mannosidases and GH78 α-L-rhamnosidases. All these enzyme groups attack carbohydrate sidechains attached to eukaryotic glycoproteins; such glycoproteins are found in abundance in intestinal cell walls. CAZy analysis also shows a genome enriched in GH18 chitinases, GH28 polygalacturonases, GH88 unsaturated glucuronyl hydrolases, GH105 unsaturated rhamnogalacturonyl hydrolases and pectate lyase (PL) family members. These enzymes attack dietary fiber components that would be resistant to digestion by most ruminant bacteria, allowing the organism to scavenge sugars from pre-digested dietary sources. The enzymes required for bacillibactin production appear to be present in the genome of Y412MC10; bacillibactin is involved in iron acquisition. Iron is in limited supply in intestinal environments, but is present in large excess (approximately 2 μM Fe2+) in Obsidian hot spring. This again argues for an intestinal origin for the organism. Y4112MC10 does not possess genes usually involved in detoxification of heavy metals and sulfide found in other hot springs organisms (unpublished results). The organism also lacks antibiotic production genes, indicating it comes from an environment with excess resources, typical of the intestine. The growth temperature range and optimum of Y412MC10 is an excellent match for intestinal conditions, but a poor fit for the conditions of Obsidian hot spring, where temperatures average 79±4°C. Nitrogen fixing Paenibacillus have been isolated from the rhizosphere, including Paenibacillus brasilensis [60], and Paenibacillus zanthoxyli [61]. Paenibacillus lautus Y412MC10 has no nitrogen-fixing genes; these would be of no advantage for a free-living organism in an intestinal environment.
Complex cooperative behaviors such as those seen with P. dendritiformis [62], and P. vortex [19] are not observed with Y412MC10; again, these behaviors may be unnecessary for survival in the intestine. Formation of external matrices in liquid and solid cultures may be beneficial to Y412MC10 for survival; the matrix may allow attachment of the bacteria to intestinal mucosa.
Conclusion
Paenibacillus sp. Y412MC10 is the first hot spring Paenibacillus sp. for which a whole genome sequence is available. Based on examination of the enzymes and biochemical pathways present in the organism, r16S comparison to other sequenced organisms and type strains, and whole genome comparisons, Y412MC10 appears to be of intestinal, rather than environmental origin.
The bison herds that are present around Obsidian hot spring may be the reservoir of this organism; on multiple collection trips, bison dung was seen in and around the pool. The upper growth temperature of 50°C and/or sporulation may have contributed to Y412MC10’s survival in this otherwise inhospitable environment.
A major need for understanding the relationships among the Paenibacilli is both genome sequence information on validly-named type strains and the naming of sequenced strains. The majority of sequenced strains have not been validly named, nor has significant genomic analysis been performed on type strains. The result is two, independent, phylogenetic trees that cannot be easily overlapped (compare Figure 1 and Figure 5). For both sets of data to be useful, a consensus should be reached on a system for incorporating both sets of data.
Acknowledgements
This work was performed under the auspices of the US Department of Energy Office of Science, Biological and Environmental Research Program, and by the University of California, Lawrence Berkeley National Laboratory under contract No. DE-AC02-05CH11231, Lawrence Livermore National Laboratory under Contract No. DE-AC52-07NA27344, and Los Alamos National Laboratory under contract No. DE-AC02-06NA25396. This work was funded in part by the DOE Great Lakes Bioenergy Research Center (DOE Office of Science BER DE-FC02-07ER64494).
Abbreviations:
- YT
yeast extract tryptone broth, LB, Luria broth, TB, Terrific Broth
References
- 1.Brock TD. The value of basic research: discovery of Thermus aquaticus and other extreme thermophiles. Genetics 1997; 146:1207-1210 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Brock TD, Freeze H. Thermus aquaticus gen. n. and sp. n., a nonsporulating extreme thermophile. J Bacteriol 1969; 98:289-297 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Brock TD, Edwards MR. Fine structure of Thermus aquaticus, an extreme thermophile. J Bacteriol 1970; 104:509-517 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Williams RA, Smith KE, Welch SG, Micallef J, Sharp RJ. DNA relatedness of Thermus strains, description of Thermus brockianus sp. nov., and proposal to reestablish Thermus thermophilus (Oshima and Imahori). Int J Syst Bacteriol 1995; 45:495-499 10.1099/00207713-45-3-495 [DOI] [PubMed] [Google Scholar]
- 5.Mohagheghi A, Grohmann K, Himmel M, Leighton L, Updegraff DM. Isolation and characterization of Acidothermus cellulolyticus gen. nov., sp. nov., a new genus of thermophilic, acidophilic, cellulolytic bacteria. Int J Syst Bacteriol 1986; 36:435-443 10.1099/00207713-36-3-435 [DOI] [Google Scholar]
- 6.Ramsing NB, Ferris MJ, Ward DM. Light-induced motility of thermophilic Synechococcus isolates from Octopus Spring, Yellowstone National Park. Appl Environ Microbiol 1997; 63:2347-2354 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Ash C, Priest FG, Collins MD. Molecular identification of rRNA group 3 bacilli (Ash, Farrow, Wallbanks and Collins) using a PCR probe test. Proposal for the creation of a new genus Paenibacillus. Antonie van Leeuwenhoek 1994; 64:253-260 10.1007/BF00873085 [DOI] [PubMed] [Google Scholar]
- 8.Khianngam S, Tanasupawat S, Akaracharanya A, Kim KK, Lee KC, Lee JS. Paenibacillus xylanisolvens sp. nov., a xylan-degrading bacterium from soil. Int J Syst Evol Microbiol 2011; 61:160-164 10.1099/ijs.0.022269-0 [DOI] [PubMed] [Google Scholar]
- 9.Montes MJ, Mercade E, Bozal N, Guinea J. Paenibacillus antarcticus sp. nov., a novel psychrotolerant organism from the Antarctic environment. Int J Syst Evol Microbiol 2004; 54:1521-1526 10.1099/ijs.0.63078-0 [DOI] [PubMed] [Google Scholar]
- 10.Fujihara M, Maeda K, Sasamori E, Matsushita M, Harasawa R (2009) Effects of chelating reagents on colonial appearance of Paenibacillus alvei isolated from canine oral cavity. The Journal of veterinary medical science / the Japanese Society of Veterinary Science 71: 147-153. [DOI] [PubMed]
- 11.Deng Y, Lu Z, Bi H, Lu F, Zhang C, Bie X. Isolation and characterization of peptide antibiotics LI-F04 and polymyxin B6 produced by Paenibacillus polymyxa strain JSa-9. Peptides 2011; 32:1917-1923 10.1016/j.peptides.2011.08.004 [DOI] [PubMed] [Google Scholar]
- 12.Ding R, Wu XC, Qian CD, Teng Y, Li O, Zhan ZJ, Zhao YH. Isolation and identification of lipopeptide antibiotics from Paenibacillus elgii B69 with inhibitory activity against methicillin-resistant Staphylococcus aureus. J Microbiol 2011; 49:942-949 10.1007/s12275-011-1153-7 [DOI] [PubMed] [Google Scholar]
- 13.Mageshwaran V, Walia S, Govindasamy V, Annapurna K. Antibacterial activity of metabolite produced by Paenibacillus polymyxa strain HKA-15 against Xanthomonas campestris pv. phaseoli. Indian J Exp Biol 2011; 49:229-233 [PubMed] [Google Scholar]
- 14.Singh AK, Chhatpar HS. Purification and characterization of chitinase from Paenibacillus sp. D1. Appl Biochem Biotechnol 2011; 164:77-88 10.1007/s12010-010-9116-8 [DOI] [PubMed] [Google Scholar]
- 15.Shinoda S, Kanamasa S, Arai M. Cloning of an endoglycanase gene from Paenibacillus cookii and characterization of the recombinant enzyme. Biotechnol Lett 2012; 34:281-286 10.1007/s10529-011-0759-5 [DOI] [PubMed] [Google Scholar]
- 16.Zhao Y, Meng K, Luo H, Yang P, Shi P, Huang H, Bai Y, Yao B. Cloning, expression, and characterization of a new xylanase from alkalophilic Paenibacillus sp. 12-11. J Microbiol Biotechnol 2011; 21:861-868 10.4014/jmb.1102.02024 [DOI] [PubMed] [Google Scholar]
- 17.Niu B, Rueckert C, Blom J, Wang Q, Borriss R. The genome of the plant growth-promoting rhizobacterium Paenibacillus polymyxa M-1 contains nine sites dedicated to nonribosomal synthesis of lipopeptides and polyketides. J Bacteriol 2011; 193:5862-5863 10.1128/JB.05806-11 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Khan N, Mishra A, Chauhan PS, Sharma YK, Nautiyal CS (2011) Paenibacillus lentimorbus enhances growth of chickpea (Cicer arietinum L.) in chromium-amended soil. Antonie Van Leeuwenhoek. [DOI] [PubMed]
- 19.Ingham CJ, Ben Jacob E. Swarming and complex pattern formation in Paenibacillus vortex studied by imaging and tracking cells. BMC Microbiol 2008; 8:36. 10.1186/1471-2180-8-36 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Sirota-Madi A, Olender T, Helman Y, Ingham C, Brainis I, Roth D, Hagi E, Brodsky L, Leshkowitz D, Galatenko V. Genome sequence of the pattern forming Paenibacillus vortex bacterium reveals potential for thriving in complex environments. BMC Genomics 2010; 11:710. 10.1186/1471-2164-11-710 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Tcherpakov M, Ben-Jacob E, Gutnick DL. Paenibacillus dendritiformis sp. nov., proposal for a new pattern-forming species and its localization within a phylogenetic cluster. Int J Syst Bacteriol 1999; 49:239-246 10.1099/00207713-49-1-239 [DOI] [PubMed] [Google Scholar]
- 22.Be'er A, Florin EL, Fisher CR, Swinney HL, Payne SM. Surviving bacterial sibling rivalry: inducible and reversible phenotypic switching in Paenibacillus dendritiformis. MBio 2011; 2:e00069-e00011 10.1128/mBio.00069-11 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Croce O, Chevenet F, Christen R: A New Web Server for the Rapid Identification of Microorganisms. Journal of microbiological and Biochemical Technology 2010, 2:84-88.
- 24.Kim OS, Cho YJ, Lee K, Yoon SH, Kim M, Na H, Park SC, Jeon YS, Lee JH, Yi H, et al. Introducing EzTaxon-e: A Prokaryotic 16S rRNA Gene Sequence Database with Phylotypes that Represent Uncultured Species. Int J Syst Evol Microbiol 2011; (In press). [DOI] [PubMed] [Google Scholar]
- 25.Gao D, Chundawat S, Liu T, Hermanson S, Gowda K, Brumm P, Dale B, Balan V. Strategy for Identification of Novel Fungal and Bacterial Glycosyl Hydrolase Hybrid Mixtures that can Efficiently Saccharify Pretreated Lignocellulosic Biomass. BioEnergy Research 2010; 3:67-81 10.1007/s12155-009-9066-6 [DOI] [Google Scholar]
- 26.Woese CR, Kandler O, Wheelis ML. Towards a natural system of organisms: proposal for the domains Archaea, Bacteria, and Eucarya. Proc Natl Acad Sci USA 1990; 87:4576-4579 10.1073/pnas.87.12.4576 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Gibbons NE, Murray RGE. Proposals Concerning the Higher Taxa of Bacteria. Int J Syst Bacteriol 1978; 28:1-6 10.1099/00207713-28-1-1 [DOI] [Google Scholar]
- 28.Garrity GM, Holt JG. The Road Map to the Manual. In: Garrity GM, Boone DR, Castenholz RW (eds), Bergey's Manual of Systematic Bacteriology, Second Edition, Volume 1, Springer, New York, 2001, p. 119-169. [Google Scholar]
- 29.Murray RGE. The Higher Taxa, or, a Place for Everything...? In: Holt JG (ed), Bergey's Manual of Systematic Bacteriology, First Edition, Volume 1, The Williams and Wilkins Co., Baltimore, 1984, p. 31-34. [Google Scholar]
- 30.List Editor List of new names and new combinations previously effectively, but not validly, published. List no. 132. Int J Syst Evol Microbiol 2010; 60:469-472 10.1099/ijs.0.022855-0 [DOI] [PubMed] [Google Scholar]
- 31.Ludwig W, Schleifer KH, Whitman WB. Class I. Bacilli class nov. In: De Vos P, Garrity G, Jones D, Krieg NR, Ludwig W, Rainey FA, Schleifer KH, Whitman WB (eds), Bergey's Manual of Systematic Bacteriology, Second Edition, Volume 3, Springer-Verlag, New York, 2009, p. 19-20. [Google Scholar]
- 32.Skerman VBD, McGowan V, Sneath PHA. Approved Lists of Bacterial Names. Int J Syst Bacteriol 1980; 30:225-420 10.1099/00207713-30-1-225 [DOI] [PubMed] [Google Scholar]
- 33.Prévot AR. In: Hauderoy P, Ehringer G, Guillot G, Magrou. J., Prévot AR, Rosset D, Urbain A (eds), Dictionnaire des Bactéries Pathogènes, Second Edition, Masson et Cie, Paris, 1953, p. 1-692. [Google Scholar]
- 34.Fischer A. Untersuchungen über bakterien. Jahrbücher für Wissenschaftliche Botanik 1895; 27:1-163 [Google Scholar]
- 35.List Editor Validation List no. 51. Validation of the publication of new names and new combinations previously effectively published outside the IJSB. Int J Syst Bacteriol 1994; 44:852. 10.1099/00207713-44-4-852 [DOI] [Google Scholar]
- 36.Ash C, Priest FG, Collins MD. Molecular identification of rRNA group 3 bacilli (Ash, Farrow, Wallbanks and Collins) using a PCR probe test. Proposal for the creation of a new genus Paenibacillus. Antonie van Leeuwenhoek 1994; 64:253-260 10.1007/BF00873085 [DOI] [PubMed] [Google Scholar]
- 37.Shida O, Takagi H, Kadowaki K, Nakamura LK, Komagata K. Transfer of Bacillus alginolyticus, Bacillus chondroitinus, Bacillus curdlanolyticus, Bacillus glucanolyticus, Bacillus kobensis, and Bacillus thiaminolyticus to the genus Paenibacillus and emended description of the genus Paenibacillus. Int J Syst Bacteriol 1997; 47:289-298 10.1099/00207713-47-2-289 [DOI] [PubMed] [Google Scholar]
- 38.Behrendt U, Schumann P, Stieglmeier M, Pukall R, Augustin J, Spröer C, Schwendner P, Moissl-Eichinger C, Ulrich A. Characterization of heterotrophic nitrifying bacteria with respiratory ammonification and denitrification activity – Description of Paenibacillus uliginis sp. nov., an inhabitant of fen peat soil and Paenibacillus purispatii sp. nov., isolated from a spacecraft assembly clean room. Syst Appl Microbiol 2010; 33:328-336 10.1016/j.syapm.2010.07.004 [DOI] [PubMed] [Google Scholar]
- 39.Judicial Commission of the International Committee on Systematics of Prokaryotes The type species of the genus Paenibacillus Ash et al. 1994 is Paenibacillus polymyxa. Opinion 77. Int J Syst Evol Microbiol 2005; 55:513. 10.1099/ijs.0.63546-0 [DOI] [PubMed] [Google Scholar]
- 40.Heyndrickx M, Vandemeulebroecke K, Scheldeman P, Kersters K, de Vos P, Logan NA, Aziz AM, Ali N, Berkeley RC. A Polyphasic Reassessment of the Genus Paenibacillus, Reclassification of Bacillus lautus (Nakamura 1984) as Paenibacillus lautus comb. nov. and of Bacillus peoriae (Montefusco et al. 1993) as Paenibacillus peoriae comb. nov., and Emended Descriptions of P. lautus and of P. peoriae. Int J Syst Bacteriol 1996; 46:988-1003 10.1099/00207713-46-4-988 [DOI] [PubMed] [Google Scholar]
- 41.Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet 2000; 25:25-29 10.1038/75556 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Liolios K, Chen IM, Mavromatis K, Tavernarakis N, Hugenholtz P, Markowitz VM, Kyrpides NC. The Genomes On Line Database (GOLD) in 2009: status of genomic and metagenomic projects and their associated metadata. Nucleic Acids Res 2010; 38:D346-D354 10.1093/nar/gkp848 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Sambrook J, Fritsch EF, Maniatis T. (1989) Molecular Cloning: A Laboratory Manual. NY: Cold Spring Harbor Laboratory Press. [Google Scholar]
- 44.DOE Joint Genome Institute http://www.jgi.doe.gov
- 45.Margulies M, Egholm M, Altman WE, Attiya S, Bader JS, Bemben LA, Berka J, Braverman MS, Chen YJ, Chen Z. Genome sequencing in microfabricated high-density picolitre reactors. Nature 2005; 437:376-380 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.The Phred/Phrap/Consed software package. http://www.phrap.com
- 47.Field D, Garrity G, Gray T, Morrison N, Selengut J, Sterk P, Tatusova T, Thomson N, Allen MJ, Angiuoli SV, Ashburner M. The minimum information about a genome sequence (MIGS) specification. Nat Biotechnol 2008; 26:541-547 10.1038/nbt1360 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Hyatt D, Chen GL, Locascio PF, Land ML, Larimer FW, Hauser LJ. Prodigal Prokaryotic Dynamic Programming Genefinding Algorithm. BMC Bioinformatics 2010; 11:19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Lowe TM, Eddy SR. tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res 1997; 25:955-964 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Lagesen K, Hallin P, Rodland EA, Staerfeldt HH, Rognes T, Usser DW. RNAmmer: consistent and rapid annotation of ribosomal RNA genes. Nucleic Acids Res 2007; 35:3100-3108 10.1093/nar/gkm160 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Griffiths-Jones S, Bateman A, Marshall M, Khanna A, Eddy SR. Rfam: an RNA family database. Nucleic Acids Res 2003; 31:439-441 10.1093/nar/gkg006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Krogh A, Larsson B, von Heijne G, Sonnhammer EL. Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J Mol Biol 2001; 305:567-580 10.1006/jmbi.2000.4315 [DOI] [PubMed] [Google Scholar]
- 53.Computational biology at ORNL. http://genome.ornl.gov/microbial/geob_y412
- 54.DOE Joint Genome Institute Integrated Microbioal Genomics http://img.jgi.doe.gov/
- 55.PHYLIP http://evolution.genetics.washington.edu/phylip/general.html
- 56.IMG software http://img.jgi.doe.gov/cgi-bin/w/main.cgi?section=EgtCluster&page=topPage
- 57.IMG website. http://img.jgi.doe.gov/cgi-bin/w/main.cgi?section=DotPlot&page=plot
- 58.Batchelor MD. Aerobic spore-bearing bacteria in the intestinal tract of children. J Bacteriol 1919; 4:23-34 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Carbohydrate-Active enZYmes Database. http://www.cazy.org/
- 60.von der Weid I, Duarte GF, van Elsas JD, Seldin L. Paenibacillus brasilensis sp. nov., a novel nitrogen-fixing species isolated from the maize rhizosphere in Brazil. Int J Syst Evol Microbiol 2002; 52:2147-2153 10.1099/ijs.0.02272-0 [DOI] [PubMed] [Google Scholar]
- 61.Ma Y, Zhang J, Chen S. Paenibacillus zanthoxyli sp. nov., a novel nitrogen-fixing species isolated from the rhizosphere of Zanthoxylum simulans. Int J Syst Evol Microbiol 2007; 57:873-877 10.1099/ijs.0.64652-0 [DOI] [PubMed] [Google Scholar]
- 62.Be'er A, Ariel G, Kalisman O, Helman Y, Sirota-Madi A, Zhang HP, Florin EL, Payne SM, Ben-Jacob E, Swinney HL. Lethal protein produced in response to competition between sibling bacterial colonies. Proc Natl Acad Sci USA 2010; 107:6258-6263 10.1073/pnas.1001062107 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.https://www.gem.re.kr/idbPPE/index.php/PPE/Structure
- 64.Computational biology at ORNL. http://genome.ornl.gov/microbial/paen
- 65.Darling AE, Mau B, Perna NT. progressiveMauve: Multiple Genome Alignment with Gene Gain, Loss and Rearrangement. PLoS ONE 2010; 5:e11147. 10.1371/journal.pone.0011147 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Aziz RK, Bartels D, Best AA, DeJongh M, Disz T, Edwards RA, Formsma K, Gerdes S, Glass EM, Kubal M. The RAST Server: Rapid Annotations using Subsystems Technology. BMC Genomics 2008; 9:75. 10.1186/1471-2164-9-75 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.RAST genome annotation. http://rast.nmpdr.org/
- 68.Overbeek R, Begley T, Butler RM, Choudhuri JV, Chuang HY, Cohoon M, de Crécy-Lagard V, Diaz N, Disz T, Edwards R. The subsystems approach to genome annotation and its use in the project to annotate 1,000 genomes. Nucleic Acids Res 2005; 33:5691-5702 10.1093/nar/gki866 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Heyndrickx M, Vandemeulebroecke K, Scheldeman P, Kersters K, De Vos P, Logan NA, Aziz AM, Ali N, Berkeley RCW. A Polyphasic Reassessment of the Genus Paenibacillus, Reclassification of Bacillus lautus (Nakamura 1984) as Paenibacillus lautus comb. nov. and of Bacillus peoriae (Montefusco et al. 1993) as Paenibacillus peoriae comb. nov., and Emended Descriptions of P. lautus and of P. peoriae. Int J Syst Bacteriol 1996; 46:988-1003 10.1099/00207713-46-4-988 [DOI] [PubMed] [Google Scholar]