Abstract
Antimicrobial peptides are secreted by small intestinal Paneth cells as components of innate immunity. To investigate the role of α-defensins in enteric host defenses in nonhuman primates, α-defensin cDNAs were isolated, α-defensin peptides were purified from rhesus macaque small bowel, and the bactericidal activities of the peptides were measured. Six rhesus enteric α-defensin (RED) cDNAs, RED-1 to RED-6, were identified in a jejunum cDNA library; the deduced RED peptides exhibited extensive diversity relative to the primary structures of rhesus myeloid α-defensins. RED-4 was purified from monkey jejunum, and N-terminal peptide sequencing of putative RED-4 peptides identified two N termini, RTCYCRTGR… and TCYCRTGRC…; these corresponded to alternative N termini for the RED-4 molecules, as deduced from their molecular masses and RED cDNAs. In situ hybridization experiments localized RED mRNAs exclusively to small intestinal Paneth cells. Recombinant RED-1 to RED-4 were purified to homogeneity and shown to be microbicidal in the low micromolar range (≤10 μg/ml) against gram-positive and gram-negative bacteria, with individual peptides exhibiting variable target cell specificities. Thus, compared to myeloid α-defensins from rhesus macaques, enteric α-defensin peptides are highly variable in both primary structure and activity. These studies should facilitate further analyses of the role of α-defensins in primate enteric immunity.
In mammals, antimicrobial proteins (AMPs) and peptide genes are expressed by differentiated cell lineages, including epithelial cells and phagocytes (13). The general features of antimicrobial peptide biology have been reviewed comprehensively (36). In cells of myeloid origin, AMPs are stored in granules and mediate nonoxidative bacterial cell killing after phagocytosis and phagolysosomal fusion (7, 14-16, 25, 29, 31). In the airway, skin, oropharynx, gingival crevices, urogenital epithelium, and gastrointestinal tract, epithelial cells release AMPs onto mucosal surfaces as mediators of innate immunity (12, 13, 28). Of the two large mammalian AMP families, cathelicidins and defensins, the defensins comprise three families of cationic peptides characterized by six cysteine residues paired in unique tridisulfide arrays that define separate subfamilies (6).
The α-defensins are major constituents of neutrophil azurophilic granules (7), and they also are released as abundant secretory granule components of Paneth cells, an enteric epithelial cell lineage that is specific to mammalian small intestinal crypts. α-Defensins occur at millimolar concentrations in vivo both in phagolysosomes (21) and at the point of secretion into the crypt lumen (2), yet they are highly bactericidal in the low micromolar range (≤10 μg/ml) in vitro. Accounting for 5% to 18% of total cellular protein in neutrophils, α-defensins are estimated to reach concentrations of ∼10 mg/ml as granules dissolve in phagolysosomes following the ingestion of microorganisms (7), and Paneth cells secrete lineage-specific isoforms at concentrations of 25 to 100 mg/ml (2, 8).
The contribution of Paneth cell α-defensins to enteric mucosal immunity is evident from the phenotype of mice transgenic for the human Paneth cell α-defensin HD5 (26). These mice faithfully express the human minigene specifically in Paneth cells, and they are immune to oral infection by Salmonella enterica serovar Typhimurium, even when challenged orally with inocula 104 times the 50% lethal dose. Perhaps the presence of HD5 in mouse Paneth cell secretions modifies the composition of the microflora by unknown selective means, conferring greater resistance to serovar Typhimurium (C. L. Bevins and N. Salzman, personal communication).
Here, we report on six highly divergent α-defensin mRNAs in rhesus macaque small bowel and on the bactericidal activities of four of these peptides produced by recombinant means. The enteric peptides are bactericidal in the low micromolar range and, compared to myeloid α-defensins from rhesus macaques (33), enteric α-defensin peptides exhibit greater variability in primary structure and activity.
MATERIALS AND METHODS
Analysis of rhesus enteric α-defensin (RED) cDNAs.
Total small intestinal RNA was extracted from a sample of rhesus macaque jejunum (provided by Christopher J. Miller under the auspices of the California National Primate Research Center Immunology Core Laboratory) by using methods described for mouse small bowel (19, 30). Rhesus small intestinal cDNAs were reverse transcribed, prepared for cloning by using a SMART PCR cDNA library construction kit (Clontech Laboratories, Inc., Palo Alto, Calif.), and ligated directionally and packaged into the Lambda Zap II phagemid vector (Stratagene Cloning Systems, Inc., La Jolla, Calif.).
To identify putative α-defensin-coding cDNA clones, the library was screened in duplicate at a density of 2,000 PFU/dish by hybridization with a mixed rhesus α-defensin probe. Reasoning that the prepro-coding regions of human and rhesus α-defensins were likely to be conserved, human α-defensin HD5-based primers were used to amplify putative α-defensins from rhesus small intestinal RNA. Total RNA from rhesus jejunum was amplified by using sense-strand primer RhHD5-F (5′-GACCC CAGCC ATGAG GACCA TC), corresponding to nucleotides 29 to 51 in HD5, and anti-sense-strand primer RhHD5-R (5′-GGAAG CTCAG CGACA GCAGA G), corresponding to the reverse complement of HD5 cDNA nucleotides 309 to 330. Amplification products were cloned in the TA cloning vector (Invitrogen Life Technologies, Carlsbad, Calif.), and several clones were sequenced to identify three different α-defensin-coding sequences. The inserts from these three clones were combined to prepare a mixed rhesus α-defensin probe, labeled with [32P]dCTP by using an RTS prime DNA labeling system (Invitrogen), and hybridized to the phagemid library in 50% formamide- 10% dextran sulfate- 1 M NaCl- 1% sodium dodecyl sulfate. Double-positive clones were plaque purified by using standard methods (34) and excised with helper phage, and plasmids were sequenced with an ABI Prism 310 genetic analyzer instrument (Applied Biosystems, Foster City, Calif.).
Purification of RED-4 from rhesus macaque jejunum.
Small intestinal proteins were extracted from a sample of rhesus macaque jejunum homogenized in 30% acetic acid and clarified by ultracentrifugation at 25,000 rpm for 2 h in an SW28 rotor (Beckman Instruments, Fullerton, Calif.) at 4°C (1, 20, 32). Supernatants were separated by Bio-Gel P-30 gel permeation chromatography (Bio-Rad, Hercules, Calif.) with 5% acetic acid, and 0.5-μl samples of column fractions were mixed with 0.5 μl of α-cyano-4-hydroxycinnamic acid (10 mg/ml) on an Applied Biosystems V700666 sample plate and analyzed by matrix-assisted laser desorption ionization- time-of-flight mass spectrometry (MALDI-TOF MS) with a Voyager-DE STR instrument (Applied Biosystems) in the Physical Sciences Mass Spectrometry Facility at the University of California, Irvine.
Column fractions that were shown to contain 3- to 4-kDa peptides were combined and purified by C18 reverse-phase (RP) high-performance liquid chromatography (HPLC) with aqueous 0.1% trifluoroacetic acid and a 15 to 45% acetonitrile gradient developed over 150 min. To identify putative defensins, HPLC fractions with peptides containing six cysteine residues were identified by MALDI-TOF MS after fraction samples were reduced in 5 mM dithiothreitol at 97°C for 15 min and alkylated by the addition of 6.5 μl of 100 mM iodoacetamide for 60 min in the dark. N termini of putative α-defensins were sequenced for nine cycles by automated Edman degradation following S pyridylethylation of cysteine residues.
Preparation of recombinant RED and RMAD peptides.
Recombinant peptides of RED-1 to RED-4 and rhesus myeloid α-defensin (RMAD) 3 (RMAD-3) and RMAD-4 were prepared by using a pET28a expression system (Novagen, Madison, Wis.) as previously reported for mouse α-defensins (27, 32). Forward primers included a 5′ EcoRI site and a Met codon at the peptide N terminus; reverse primers contained consecutive stop codons and a 5′ SalI site. RED-1 to RED-4 and RMAD-3 and RMAD-4 (33) cDNAs were amplified from the corresponding cDNA clones or genomic DNA by using the following primer sets: RED-1, pET-RED1-F (5′-ATATAT GAATTC ATGCGC ACCTGC CGTTGC) and pET-RED4-R (5′-ACACAC GTCGAC TCATCA GCGACA GCAGAG TTT); RED-2, pET-RED2-F (5′-ATATAT GAATTC ATGTTC ACCTGC CATTGC) and pET-RED2-R (5′-ACACAC GTCGAC TCATCA GCGACA GCAGAG GTT); RED-3, pET-RED3-F (5′-ATATAT GAATTC ATGCAC ACCTGC TATTGC) and pET-RED4-R; RED-4, pET-RED4-F (5′-ATATAT GAATTC ATGCGC ACCTGC TATTGC) and pET-RED4-R; RMAD-3, pET-RMAD3F (5′-ACACAC GAATTC ATGGCC TGCTAT TGCAGA) and pET-RMAD3-R (5′-ACACAC GTCGAC TCATCA GCAGCA GAATGC CCA); and RMAD-4, pET-RMAD4-F (5′-ACACAC GAATTC ATGAGA CGCACC TGCCGT) and pET-RMAD4-R (5′-ACACAC GTCGAC TCATCA GCGACA GCAGAG ACT). Because the natural RED-3-coding sequence contains an EcoRI restriction site, primers pET-RED3-intF (5′-ACCCCT GAATTT CATGCC GG) and pET-RED3-intR (5′-CCGGCA TGAAAT TCAGGG GT) were used to mutagenize the EcoRI site without altering coding function (32). Following PCR amplification, samples (25 μl) from individual reactions were gel purified with 2% agarose gels and extracted with QIAEX II (Qiagen Inc., Valencia, Calif.). In most cases, amplification products were cloned in pCR2.1-TOPO, sequenced, and digested sequentially with SalI and EcoRI, and gel-purified inserts were ligated into EcoRI- and SalI-digested pET28a plasmid DNA and transformed into both Escherichia coli XL-2 Blue and BL21(DE3) Codon Plus cells (Stratagene).
His-tagged fusion proteins induced at 16°C for 18 h (RED-1 to RED-4) or at 37°C for 6 h (RMAD-3 and RMAD-4) were purified by nickel-nitrilotriacetic acid resin affinity chromatography (Ni-NTA Superflow; Qiagen) with 100 or 200 mM imidazole in 6 M guanidine-HCl- 100 mM Tris-HCl (pH 8.0) (27, 32). Eluted fusion proteins were cleaved with 5 mg of CNBr/ml in 6 M guanidine- 0.1 N HCl for 24 h at room temperature in the dark. The samples were diluted, acidified with 0.01% trifluoroacetic acid, and purified by C18 RP HPLC with a 15 to 45% acetonitrile gradient developed over 90 min (27, 32). Fractions with RED or RMAD peptides identified by MALDI-TOF MS were rechromatographed with a shallower acetonitrile gradient to obtain homogeneous peptides, as assessed by acid urea-polyacrylamide gel electrophoresis (PAGE). Peptide concentrations were determined by measuring the absorbance at 280 nm with extinction coefficients calculated by ProtParam at the EXPASY website (http://us.expasy.org/tools/).
Detection and localization of α-defensin mRNAs in rhesus macaque jejunum by in situ hybridization.
A 202-bp cDNA probe corresponding to the prepro-coding region of RED-4 in BlueScript II KS(+) was generously provided by Alison Quayle (Department of Microbiology, Immunology, and Parasitology, Health Sciences Center, Louisiana State University, New Orleans) for preparation of a digoxigenin (DIG)-labeled riboprobe for the detection of α-defensin mRNA. The α-defensin cDNA fragment corresponded to nucleotides 70 to 272 in the RED-4 cDNA sequence (not shown). The phagemid was linearized with either HindIII or EcoRI (Lofstrand Ltd., Gaithersburg, Md.) for the production of strand-specific DIG-labeled sense-strand and anti-sense-strand RED riboprobes (Lofstrand Ltd.).
α-Defensin mRNAs were localized in rhesus jejunum by in situ hybridization with the DIG-labeled, anti-sense-strand RED riboprobe. A sample of jejunum preserved in 10% formalin was embedded in paraffin, and 5-μm-thick sections were cut and placed on Superfrost-Plus slides. Sections were deparaffinized in xylene, rehydrated with graded ethanol washes and phosphate-buffered saline, and fixed with 4% paraformaldehyde in phosphate-buffered saline for 30 min. Sections were digested with proteinase XXIV (Sigma-Aldrich Corp., St. Louis, Mo.) for 10 min at 37°C and then preincubated with a hybridization solution consisting of 50% formamide, 1 μg of tRNA/μl, 10% dextran sulfate, and 50% Sigma hybridization solution for 15 min at 60°C. Anti-sense-strand or sense-strand riboprobes were denatured in hybridization solution at 80°C for 5 min and cooled, and tissue sections were hybridized with 50 μl of the riboprobe solution for 18 h at 60°C. Following hybridization, unreacted probe was removed by digestion with RNases A and T1. Endogenous alkaline phosphatase activity was blocked with serum-free blocking solution (Dakocytomation, Inc., Carpinteria, Calif.) containing levamisole prior to the application of alkaline phosphatase-labeled, sheep-anti-DIG antibody (Boehringer) containing 1% normal horse serum. Hybridized α-defensin mRNAs were visualized after application of the precipitating chromogenic substrate for alkaline phosphatase, nitroblue tetrazolium chloride- 5-bromo-4-chloro-3-indolylphosphate (Roche Applied Science, Indianapolis, Ind.). Sections were counterstained with nuclear fast red stain and dehydrated with graded ethanol solutions and xylene, and coverslips were applied with Permount.
Bactericidal peptide activity assays.
E. coli ML35, Staphylococcus aureus 710a, Listeria monocytogenes 10403S, and Vibrio cholerae 0395 were target organisms for bactericidal assays. Individual peptides were incubated with 106 CFU of log-phase bacterial cells/ml in 50 μl of buffer (5 × 104 the actual CFU in 50 μl) consisting of 10 mM piperazine-N,N′-bis(2-ethanesulfonic acid) (pH 7.4) (PIPES) supplemented with 1% (vol/vol) Trypticase soy broth at 37°C for 1 h. Samples from incubation mixtures diluted 100-fold were plated on semisolid media by using an Autoplater 4000 (Spiral Biotech, Bethesda, Md.), and surviving bacterial cells were determined as bacterial CFU by counting after overnight growth.
RESULTS
RED cDNAs.
RED-coding sequences were deduced from clones isolated from a rhesus macaque jejunum cDNA library (see Materials and Methods). α-Defensin cDNA clones from the library were identified by hybridization at moderate stringency by using an amplified cDNA probe for the human α-defensin HD5 prepro-coding region, and the coding functions of individual sequences were determined by DNA sequencing. Six distinct cDNA sequences, RED-1 to RED-6, were identified from the characterization of 64 isolates (Fig. 1). Alignment of RED-1 to RED-6 cDNA sequences with CLUSTALW showed that nucleotide sequences coding for signal peptides and prosegments are conserved but that all α-defensin-coding nucleotide sequences and deduced mature RED peptides diverge extensively (Fig. 1 and 2).
FIG. 1.
Alignment of RED cDNA sequences. α-Defensin clones from a rhesus macaque small bowel cDNA library were identified by hybridization with an amplified HD5 cDNA probe and sequenced (see Materials and Methods). Six obtained sequences were aligned with CLUSTALW. Dots in aligned sequences denote identities with RED-6; dashes denote spaces introduced to optimize the alignment.
FIG. 2.
Alignment of deduced RED-1 to RED-6 precursor proteins. RED precursor sequences were predicted from corresponding cDNAs (Fig. 1). N termini of RED-1 to RED-6 were deduced from N termini identified for the natural RED-4 peptide (see Fig. 3), which is conserved in the six deduced pro-RED sequences. Shaded residues denote conserved arginines that may serve as canonical trypsin-processing sites (8). Signal peptides and prosegments are highly conserved, and mature RED peptides show extensive variations between sequences.
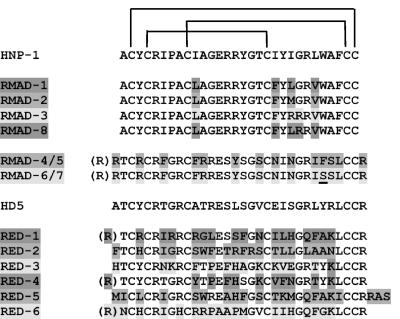
Rhesus macaque Paneth cells express at least six α-defensin-coding sequences that are distinct from the coding sequences for human α-defensins HD5 (Fig. 3) and HD6 (data not shown). Pairwise BLASTP alignments showed that RED-3 and RED-4 were the most closely related rhesus α-defensins, sharing 77% amino acid sequence identity. Also, RED-3 and RED-4 were the peptides most closely related to HD5, with 58 and 67% amino acid sequence identities, respectively, and neither peptide had meaningful similarity to HD6, human neutrophil α-defensins (HNPs), or RMAD-4. When the nine amino acid residue positions that are conserved in nearly every α-defensin, i.e., the six Cys positions, Arg68, Glu76, and Gly80, are excluded from the peptide comparisons, differences between RED peptides are more apparent. For example, outside the nine consensus positions, RED-4 and HD5 are identical at 15 positions, but RED-1, RED-2, RED-5, and RED-6 exhibit identity with HD5 at only 5 or 6 of the nonconserved sites (Fig. 3). This finding is in contrast to the similarity of most mouse Paneth cell α-defensins (19) and of RMAD-1 to RMAD-3 and RMAD-8 to HNP-1 to HNP-3 (29).
FIG. 3.
Alignment of myeloid and enteric α-defensins from rhesus macaques. The primary structures of RMADs and REDs are shown aligned with human myeloid α-defensin HNP-1 and human Paneth cell α-defensin HD5. RMAD-1 to RMAD-3 and RMAD-8 residue positions that differ from those of HNP-1 are shaded. Similarly, RMAD-4 to RMAD-7 and RED residue positions that differ from those of HD5 also are shaded. N-terminal residues within parentheses represent alternative N termini determined here and by Selsted et al. (29). Note that RED peptides are markedly more varied than RMAD peptides.
Purification of rhesus macaque RED-4.
To ascertain the biochemical properties of RED peptides in rhesus macaque small bowel, protein extracts were prepared from a sample of monkey jejunum by extraction with 30% acetic acid (30). Samples of acid-soluble jejunum protein were subjected to Bio-Gel P-30 separation (19, 30), and fractions containing peptides of ≤10 kDa, as identified by MALDI-TOF MS, were resolved by C18 RP HPLC (see Materials and Methods). MALDI-TOF MS comparisons of samples from HPLC fractions before and after reduction and alkylation identified fractions containing putative defensins as ∼4-kDa peptides with six Cys residues that increased in mass by ∼347 atomic mass units as a result of alkylation with iodoacetamide. Peptides in the putative defensin-containing fractions were purified further by C18 RP HPLC.
One peptide corresponded to a molecule predicted from the RED-4 cDNA sequence, shifting from 3804.6 to 4150.8 atomic mass units after reduction and alkylation. N-terminal peptide sequencing by Edman degradation detected the N termini RTCYCRTGR… and TCYCRTGRC…, which were identified as RED-4 isoforms on the basis of the peptide masses determined by MALDI-TOF MS. Thus, the purified peptide actually consisted of two coeluting RED-4 molecules that correspond to alternative N-terminal processing variants of pro-RED-4 that are produced during activation (Fig. 2 and 3). The demonstration that RED-4 has two N-termini, Arg63 and Thr64 (Fig. 3; see above as well), suggests that alternative cleavage events occur in RED-4 activation: at Arg62↓Arg63 and at Arg63↓Thr64 (Fig. 2 and 3). All deduced pro-RED sequences contain Arg, a canonical trypsin cleavage site (8), at residue position 62 (Fig. 2), suggesting that trypsin may process pro-α-defensins in monkey Paneth cells as it does in human Paneth cells (8). In contrast to the diversity noted among RED-coding regions, RMAD-1 to RMAD-3 and RMAD-8 are very similar to each other and to HNP-1, and RMAD-4 to RMAD-7 differ by only a single nucleotide substitution and correspond to alternative N-terminal processing variants (33).
Detection and localization of α-defensin mRNA in rhesus macaque jejunum by in situ hybridization.
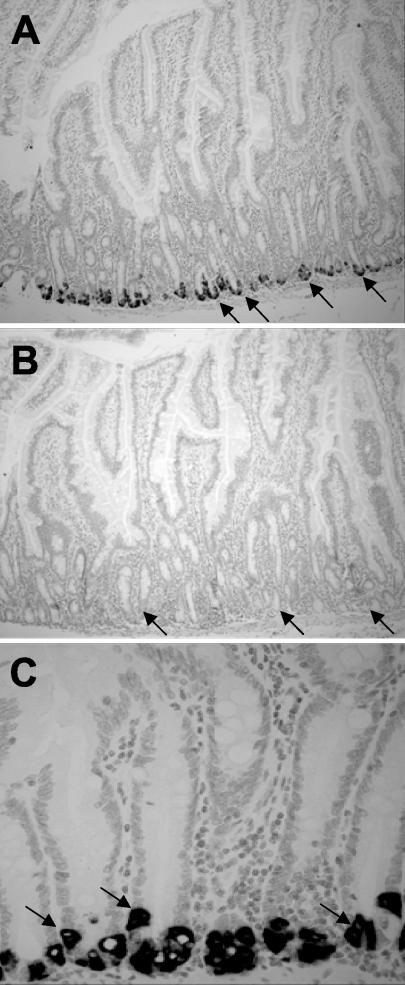
α-Defensin-coding mRNAs were localized exclusively to small intestinal Paneth cells by in situ hybridization with RED riboprobes corresponding to the conserved prepro-coding region (Fig. 4). The Paneth cell content of α-defensin mRNAs was evident from the dark blue-violet precipitate from nitroblue tetrazolium chloride-5-bromo-4-chloro-3-indolylphosphate in the alkaline phosphatase reaction at the base of every crypt for sections hybridized with the anti-sense-strand riboprobe (Fig. 4A and C). In contrast, adjacent sections of jejunum were negative with the control sense-strand riboprobe (Fig. 4B). Consistent with findings in mice, humans, and rats, the only crypt cells expressing α-defensin mRNAs were Paneth cells, recognized by their characteristic eosinophilic secretory granules evident in adjacent tissue sections by staining with eosin and hematoxylin (data not shown) (23). Also consistent with previous studies, no α-defensin mRNA expression was detected in villus enterocytes, goblet cells, or cells populating the lamina propria or submucosa (Fig. 4C).
FIG. 4.
Localization of RED mRNAs in Paneth cells of rhesus macaque jejunum. Sections of monkey jejunum were hybridized to a RED-4 prosegment cDNA probe as described in Materials and Methods. (A and B) Hybridization with anti-sense-strand and sense-strand riboprobes, respectively, shown at a magnification of ×95. Arrows indicate the base of jejunum crypts occupied by Paneth cells. Note the absence of hybridization with the negative control sense-strand probe in panel B. (C) Like panel A but shown at a magnification of ×380. Arrows indicate strongly positive individual Paneth cells.
Purification of recombinant enteric α-defensins.
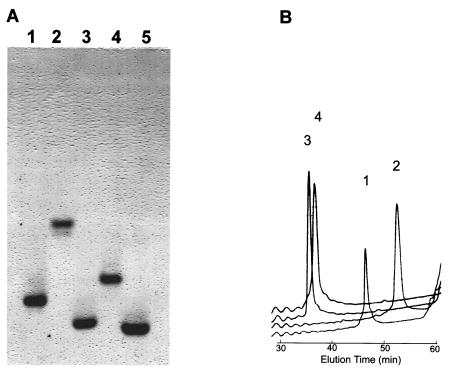
To evaluate the bactericidal activities of rhesus Paneth cell α-defensins, recombinant RED-1 to RED-4, RMAD-3, RMAD-4, and mouse cryptdin-4 (Crp4) were expressed and purified. Although the N termini of RED-1 to RED-3 have not been determined, we reasoned that the canonical trypsin cleavage site (8), at residue position 62 in all pro-RED sequences, may constitute a consensus activation site for all pro-REDs, as apparent from the purification of natural RED-4 peptides. RED peptides were expressed as N-terminal six-His-tagged fusion proteins in E. coli by using expression vector pET28a (1, 27, 32). Peptide homogeneity was apparent from analyses by acid urea-PAGE (Fig. 5A) and C18 RP HPLC (Fig. 5B). In addition, the molecular masses of all recombinant peptides matched their respective theoretical masses (Fig. 6).
FIG. 5.
Recombinant RED peptides. (A) Samples (1 μg) of purified recombinant RED-1 to RED-4 were separated by acid urea-PAGE and stained with Coomassie blue. Lanes: 1, RED-1; 2, RED-2; 3, RED-3; 4, RED-4; 5, Crp4. (B) Peptide samples (4 μg) were analyzed separately on an analytical C18 RP HPLC column and eluted with a gradient of 0 to 40% acetonitrile for 60 min. Numerals 1 to 4 above the peaks of the tracings denote the relative elution positions for RED-1 to RED-4, respectively.
FIG. 6.
Biochemical features of recombinant RED and RMAD peptides. Recombinant RED and RMAD amino acid sequences and their theoretical (theor.) molecular weights (M.W.) are shown along with their experimental M.W. observed by MALDI-TOF MS. Net charges were calculated from the following values: Arg, +1; Lys, +1; His, +0.5; Asp, −1; and Glu, −1.
Bactericidal activities of RED peptides.
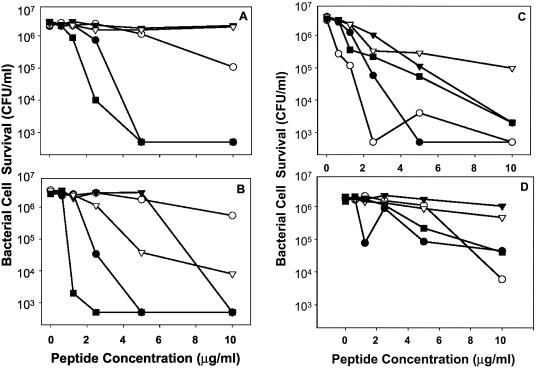
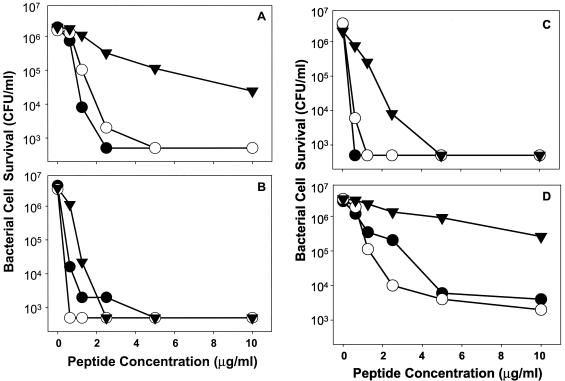
The four RED peptides tested had overall bactericidal activities in the low micromolar range, similar to those of Crp4 and RMADs under the conditions of these assays. Consistent with the properties of most AMPs (36), bacterial species varied in their susceptibility to individual RED peptides (Fig. 7) at 0 to 10 μg of peptide/ml, low concentrations relative to the levels found in vivo in humans and mice (2, 8, 36). For example, RED-2 showed little activity against E. coli and V. cholerae but was highly bactericidal against L. monocytogenes (Fig. 7). Conversely, RED-1 was active against gram-negative species but not against S. aureus (Fig. 7). In contrast to the variability among RED peptides, RMAD-3 and RMAD-4 were highly bactericidal against all species tested (Fig. 8) (33). The differences in peptide activity cannot be attributed to charge alone, because RMAD-3 and RMAD-4 have charges of +5 and +8, respectively, yet have very similar activities. With a charge of +4.5, RED-2 is the least cationic of the four RED peptides tested, but it is the most active RED against L. monocytogenes (Fig. 7C). Similarly, RED-1 is highly potent against V. cholerae, but RED-3 is not (Fig. 7A), even though the two peptides differ by only 0.5 unit of electropositive charge (Fig. 6). These findings suggest that cationic properties may contribute to bactericidal activity but only within the context of a particular primary structure.
FIG. 7.
Bactericidal peptide activities of recombinant RED peptides. Individual peptides at the concentrations shown were incubated with 106 CFU of log-phase bacterial cells/ml in 50 μl for 1 h and then plated on semisolid media (see Materials and Methods). Surviving bacterial cells were determined as bacterial CFU by counting after overnight growth. The results shown are representative of duplicate or triplicate experiments performed on separate days, and individual data points denote single determinations. (A) V. cholerae. (B) E. coli. (C) L. monocytogenes. (D) S. aureus. Symbols: •, RED-1; ○, RED-2; ▾, RED-3; ▿, RED-4; ▪, Crp4.
FIG. 8.
Bactericidal peptide activities of recombinant RMAD peptides. Peptides were assayed for bactericidal activities as described in the legend to Fig. 7. Again, the results shown are representative of duplicate or triplicate experiments performed on separate days, and individual data points denote single determinations. (A) V. cholerae. (B) E. coli. (C) L. monocytogenes. (D) S. aureus. Symbols: •, RMAD-3; ○, RMAD-4; ▾, RED-4.
DISCUSSION
The role of α-defensins in enteric host defense of nonhuman primates has been investigated by characterizing α-defensin cDNA peptides from rhesus macaque small intestine. Six RED cDNAs, RED-1 to RED-6, were identified, and the deduced RED peptides are diverse compared to the RMAD peptides. As predicted from studies with humans, mice, and rats (23), RED mRNAs were found exclusively in small intestinal Paneth cells by localization with in situ hybridization. RED-1 to RED-4 prepared by recombinant means were microbicidal against gram-positive and gram-negative bacteria in the low micromolar range. Thus, compared to RMAD peptides, RED peptides are highly variable in both primary structure and activity. Also, relative to the two α-defensins expressed by human Paneth cells (3, 10, 11), the number of α-defensins is larger in this primate. Because transgenic expression of human HD5 in mice confers immunity to oral infection by S. enterica serovar Typhimurium (26), it may be of interest to test whether mice transgenic for macaque α-defensins are comparably protected.
Evidence suggests that trypsin may process pro-α-defensins after secretion by monkey Paneth cells, as in humans (8). For example, all RED precursors deduced from cDNA sequences contain Arg, a canonical trypsin cleavage site (8), at residue position 62 (Fig. 2). The quantity of activated RED peptides recovered from the sample of macaque small intestine analyzed was very low relative to the level of activated Crp peptides routinely isolated from mouse small intestine, suggesting that pro-REDs may be activated after secretion in monkeys. Comparisons of RED precursor sequences deduced from RED cDNAs suggest that, as shown for human Paneth cell α-defensins (8), trypsin may be the pro-RED-activating enzyme. On the basis of the conservation of Arg62 in all pro-REDs, the fact that trypsin processes human pro-α-defensins (8), and the absence of matrix metalloproteinase 7 (the mouse pro-α-defensin-activating enzyme) from rhesus macaque small bowel (data not shown), it is reasonable to predict that pro-REDs will be activated by trypsin-mediated proteolysis between Arg62 and position 63 in rhesus Paneth cells. Because RED-4 has alternative N termini, Arg63 and Thr64 (Fig. 3), we infer that activating cleavage steps could occur at Arg62↓Arg63 and at Arg63↓Thr64 in pro-RED-1 and pro-RED-4 and at Arg63↓Asn64 in pro-RED-6 (Fig. 3). Interestingly, apparently N-terminally processed forms of RMAD-4 and RMAD-5 and of RMAD-6 and RMAD-7 also have been described, but the enzyme responsible for activation is not known (33).
Enteric α-defensin transcripts and peptides have been localized exclusively to Paneth cells or Paneth cell secretory granules (4, 5, 18, 20, 24, 30). In contrast, myeloid α-defensin mRNAs are expressed almost exclusively in the bone marrow, where they are found at the highest levels in promyelocytes and at lower levels in myeloblasts and myelocytes (35). Also, β-defensins, widely expressed on many mucosal surfaces, have not been detected in normal small intestine. Myeloid α-defensin genes consist of three exons with a unique first intron that interrupts the 5′ untranslated region and is absent from all known Paneth cell α-defensin genes. All known Paneth cell α-defensin genes contain two exons separated by a single intron (17, 22). To our knowledge, exceptions to this lineage specificity of α-defensin gene expression have not been reported. This consistent, lineage-specific expression of α-defensin gene subsets implies that specific cis-acting gene elements or lineage-specific transcription factors are involved in regulating their transcription. The rhesus myeloid and Paneth cell α-defensin genes have not been mapped or characterized structurally, but that information may provide comparative insights.
The markedly greater diversity of enteric α-defensin primary structures than of myeloid α-defensin primary structures suggests that Paneth cell α-defensins may be subject to higher rates of amplification and diversification than genes expressed in bone marrow. Perhaps because REDs are released into the lumen of the small bowel, a conduit for the many ingested microbes, evolving a diverse population of Paneth cell α-defensins may confer selective advantages. Certainly, the multiplicity of mouse enteric α-defensins (9, 19) is consistent with this notion. Also, cryptdin cDNAs cloned from C57BL/6 small intestine are remarkably numerous, but they also diverge from those in C3H/HeJ, BALB/cJ, and outbred Swiss mice, suggesting that the genes are subject to rapid evolution as well (H. Tanabe and A. J. Ouellette, unpublished data; GenBank accession numbers BG069980, AV070777, AV062546, AV069220, AV067046, AV069888, AV071421, AV062870, AV055939, AV068170, AV067388, AV058312, AV049476, AV064285, AV070188, AV071100, AV070571, AV066826, AV069820, AV069763, AV067626, AV069817, AV067447, AV064683, AV070855, AV065200, and AV070313). We speculate that evolutionary pressure for greater Paneth cell α-defensin gene diversity may relate to the fact that myeloid α-defensin function is complemented by oxidative microbicidal mechanisms, whereas enteric α-defensins act in the extracellular milieu under conditions of progressive dilution after secretion.
Acknowledgments
This work was supported by NIH CNPRC base grant RR00169 and NIH grants DK43183 (to S.D.), AI22931 (to M.E.S.), and DK44632 (to A.J.O.).
We thank Alison Quayle for providing the RED-4 prepro-coding region cDNA construct for use as an in situ hybridization probe. Xiaoqing Qu and Victoria Rojo provided excellent technical assistance.
Editor: F. C. Fang
REFERENCES
- 1.Ayabe, T., D. P. Satchell, P. Pesendorfer, H. Tanabe, C. L. Wilson, S. J. Hagen, and A. J. Ouellette. 2002. Activation of Paneth cell alpha-defensins in mouse small intestine. J. Biol. Chem. 277:5219-5228. [DOI] [PubMed] [Google Scholar]
- 2.Ayabe, T., D. P. Satchell, C. L. Wilson, W. C. Parks, M. E. Selsted, and A. J. Ouellette. 2000. Secretion of microbicidal alpha-defensins by intestinal Paneth cells in response to bacteria. Nat. Immunol. 1:113-118. [DOI] [PubMed] [Google Scholar]
- 3.Bevins, C. L., D. E. Jones, A. Dutra, J. Schaffzin, and M. Muenke. 1996. Human enteric defensin genes: chromosomal map position and a model for possible evolutionary relationships. Genomics 31:95-106. [DOI] [PubMed] [Google Scholar]
- 4.Condon, M. R., A. Viera, M. D'Alessio, and G. Diamond. 1999. Induction of a rat enteric defensin gene by hemorrhagic shock. Infect. Immun. 67:4787-4793. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Cunliffe, R. N., F. R. Rose, J. Keyte, L. Abberley, W. C. Chan, and Y. R. Mahida. 2001. Human defensin 5 is stored in precursor form in normal Paneth cells and is expressed by some villous epithelial cells and by metaplastic Paneth cells in the colon in inflammatory bowel disease. Gut 48:176-185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Ganz, T. 2003. Defensins: antimicrobial peptides of innate immunity. Nat. Rev. Immunol. 3:710-720. [DOI] [PubMed] [Google Scholar]
- 7.Ganz, T., M. E. Selsted, D. Szklarek, S. S. Harwig, K. Daher, D. F. Bainton, and R. I. Lehrer. 1985. Defensins. Natural peptide antibiotics of human neutrophils. J. Clin. Investig. 76:1427-1435. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Ghosh, D., E. Porter, B. Shen, S. K. Lee, D. Wilk, J. Drazba, S. P. Yadav, J. W. Crabb, T. Ganz, and C. L. Bevins. 2002. Paneth cell trypsin is the processing enzyme for human defensin-5. Nat. Immunol. 3:583-590. [DOI] [PubMed] [Google Scholar]
- 9.Huttner, K. M., M. E. Selsted, and A. J. Ouellette. 1994. Structure and diversity of the murine cryptdin gene family. Genomics 19:448-453. [DOI] [PubMed] [Google Scholar]
- 10.Jones, D. E., and C. L. Bevins. 1993. Defensin-6 mRNA in human Paneth cells: implications for antimicrobial peptides in host defense of the human bowel. FEBS Lett. 315:187-192. [DOI] [PubMed] [Google Scholar]
- 11.Jones, D. E., and C. L. Bevins. 1992. Paneth cells of the human small intestine express an antimicrobial peptide gene. J. Biol. Chem. 267:23216-23225. [PubMed] [Google Scholar]
- 12.Lehrer, R. I., and T. Ganz. 2002. Cathelicidins: a family of endogenous antimicrobial peptides. Curr. Opin. Hematol. 9:18-22. [DOI] [PubMed] [Google Scholar]
- 13.Lehrer, R. I., and T. Ganz. 2002. Defensins of vertebrate animals. Curr. Opin. Immunol. 14:96-102. [DOI] [PubMed] [Google Scholar]
- 14.Lehrer, R. I., T. Ganz, and M. E. Selsted. 1988. Oxygen-independent bactericidal systems. Mechanisms and disorders. Hematol. Oncol. Clin. North Am. 2:159-169. [PubMed] [Google Scholar]
- 15.Mambula, S. S., E. R. Simons, R. Hastey, M. E. Selsted, and S. M. Levitz. 2000. Human neutrophil-mediated nonoxidative antifungal activity against Cryptococcus neoformans. Infect. Immun. 68:6257-6264. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Newman, S. L., L. Gootee, J. E. Gabay, and M. E. Selsted. 2000. Identification of constituents of human neutrophil azurophil granules that mediate fungistasis against Histoplasma capsulatum. Infect. Immun. 68:5668-5672. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Ouellette, A. J., and C. L. Bevins. 2001. Paneth cell defensins and innate immunity of the small bowel. Inflamm. Bowel Dis. 7:43-50. [DOI] [PubMed] [Google Scholar]
- 18.Ouellette, A. J., D. Darmoul, D. Tran, K. M. Huttner, J. Yuan, and M. E. Selsted. 1999. Peptide localization and gene structure of cryptdin 4, a differentially expressed mouse Paneth cell α-defensin. Infect. Immun. 67:6643-6651. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Ouellette, A. J., M. M. Hsieh, M. T. Nosek, D. F. Cano-Gauci, K. M. Huttner, R. N. Buick, and M. E. Selsted. 1994. Mouse Paneth cell defensins: primary structures and antibacterial activities of numerous cryptdin isoforms. Infect. Immun. 62:5040-5047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Ouellette, A. J., D. P. Satchell, M. M. Hsieh, S. J. Hagen, and M. E. Selsted. 2000. Characterization of luminal Paneth cell alpha-defensins in mouse small intestine. Attenuated antimicrobial activities of peptides with truncated amino termini. J. Biol. Chem. 275:33969-33973. [DOI] [PubMed] [Google Scholar]
- 21.Ouellette, A. J., and M. E. Selsted. 1995. Defensins in granules of phagocytic and nonphagocytic cells. Trends Cell Biol. 5:114-119. [DOI] [PubMed] [Google Scholar]
- 22.Ouellette, A. J., and M. E. Selsted. 1996. Paneth cell defensins: endogenous peptide components of intestinal host defense. FASEB J. 10:1280-1289. [DOI] [PubMed] [Google Scholar]
- 23.Porter, E. M., C. L. Bevins, D. Ghosh, and T. Ganz. 2002. The multifaceted Paneth cell. Cell. Mol. Life Sci. 59:156-170. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Porter, E. M., L. Liu, A. Oren, P. A. Anton, and T. Ganz. 1997. Localization of human intestinal defensin 5 in Paneth cell granules. Infect. Immun. 65:2389-2395. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Rice, W. G., T. Ganz, J. M. Kinkade, Jr., M. E. Selsted, R. I. Lehrer, and R. T. Parmley. 1987. Defensin-rich dense granules of human neutrophils. Blood 70:757-765. [PubMed] [Google Scholar]
- 26.Salzman, N. H., D. Ghosh, K. M. Huttner, Y. Paterson, and C. L. Bevins. 2003. Protection against enteric salmonellosis in transgenic mice expressing a human intestinal defensin. Nature 422:522-526. [DOI] [PubMed] [Google Scholar]
- 27.Satchell, D. P., T. Sheynis, Y. Shirafuji, S. Kolusheva, A. J. Ouellette, and R. Jelinek. 2003. Interactions of mouse Paneth cell alpha-defensins and alpha-defensin precursors with membranes: prosegment inhibition of peptide association with biomimetic membranes. J. Biol. Chem. 278:13838-13846. [DOI] [PubMed] [Google Scholar]
- 28.Schutte, B. C., and P. B. McCray, Jr. 2002. Beta-defensins in lung host defense. Annu. Rev. Physiol. 64:709-748. [DOI] [PubMed] [Google Scholar]
- 29.Selsted, M. E., S. S. Harwig, T. Ganz, J. W. Schilling, and R. I. Lehrer. 1985. Primary structures of three human neutrophil defensins. J. Clin. Investig. 76:1436-1439. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Selsted, M. E., S. I. Miller, A. H. Henschen, and A. J. Ouellette. 1992. Enteric defensins: antibiotic peptide components of intestinal host defense. J. Cell Biol. 118:929-936. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Selsted, M. E., Y. Q. Tang, W. L. Morris, P. A. McGuire, M. J. Novotny, W. Smith, A. H. Henschen, and J. S. Cullor. 1996. Purification, primary structures, and antibacterial activities of beta-defensins, a new family of antimicrobial peptides from bovine neutrophils. J. Biol. Chem. 271:16430. [DOI] [PubMed] [Google Scholar]
- 32.Shirafuji, Y., H. Tanabe, D. P. Satchell, A. Henschen-Edman, C. L. Wilson, and A. J. Ouellette. 2003. Structural determinants of procryptdin recognition and cleavage by matrix metalloproteinase-7. J. Biol. Chem. 278:7910-7919. [DOI] [PubMed] [Google Scholar]
- 33.Tang, Y. Q., J. Yuan, C. J. Miller, and M. E. Selsted. 1999. Isolation, characterization, cDNA cloning, and antimicrobial properties of two distinct subfamilies of α-defensins from rhesus macaque leukocytes. Infect. Immun. 67:6139-6144. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Tang, Y. Q., J. Yuan, G. Osapay, K. Osapay, D. Tran, C. J. Miller, A. J. Ouellette, and M. E. Selsted. 1999. A cyclic antimicrobial peptide produced in primate leukocytes by the ligation of two truncated alpha-defensins. Science 286:498-502. [DOI] [PubMed] [Google Scholar]
- 35.Yount, N. Y., M. S. Wang, J. Yuan, N. Banaiee, A. J. Ouellette, and M. E. Selsted. 1995. Rat neutrophil defensins. Precursor structures and expression during neutrophilic myelopoiesis. J. Immunol. 155:4476-4484. [PubMed] [Google Scholar]
- 36.Zasloff, M. 2002. Antimicrobial peptides of multicellular organisms. Nature 415:389-395. [DOI] [PubMed] [Google Scholar]