Summary: Normal tomato ripening is influenced by alterations on both cytosolic phosphoenolpyruvate carboxykinase and plastidic NADP-malic enzyme. This study provides compelling evidence of their roles in starch biosynthesis, respiration rates, and tricarboxylic acid cycle flux.
Abstract
The aim of this work was to investigate the effect of decreased cytosolic phosphoenolpyruvate carboxykinase (PEPCK) and plastidic NADP-dependent malic enzyme (ME) on tomato (Solanum lycopersicum) ripening. Transgenic tomato plants with strongly reduced levels of PEPCK and plastidic NADP-ME were generated by RNA interference gene silencing under the control of a ripening-specific E8 promoter. While these genetic modifications had relatively little effect on the total fruit yield and size, they had strong effects on fruit metabolism. Both transformants were characterized by lower levels of starch at breaker stage. Analysis of the activation state of ADP-glucose pyrophosphorylase correlated with the decrease of starch in both transformants, which suggests that it is due to an altered cellular redox status. Moreover, metabolic profiling and feeding experiments involving positionally labeled glucoses of fruits lacking in plastidic NADP-ME and cytosolic PEPCK activities revealed differential changes in overall respiration rates and tricarboxylic acid (TCA) cycle flux. Inactivation of cytosolic PEPCK affected the respiration rate, which suggests that an excess of oxaloacetate is converted to aspartate and reintroduced in the TCA cycle via 2-oxoglutarate/glutamate. On the other hand, the plastidic NADP-ME antisense lines were characterized by no changes in respiration rates and TCA cycle flux, which together with increases of pyruvate kinase and phosphoenolpyruvate carboxylase activities indicate that pyruvate is supplied through these enzymes to the TCA cycle. These results are discussed in the context of current models of the importance of malate during tomato fruit ripening.
Fruit ripening and development is well established to be under tight genetic control (Matas et al., 2009; Karlova et al., 2011; Klee and Giovannoni, 2011). A growing body of evidence suggests that, in parallel, large metabolic shifts additionally occur during this process (Carrari and Fernie, 2006; Osorio et al., 2011). Insights into the genetic mechanisms that mediate fruit ripening-related processes, such as cell wall metabolism, pigment synthesis, and sugar metabolism, have resulted from studies that collectively span a wide range of plant species and indicate that they are broadly conserved (Seymour et al., 2008; Moing et al., 2011; Zhang et al., 2011). However, certain species-specific differences exist in the dynamics of other metabolite pools across ripening, with, for example, grape (Vitis vinifera), strawberry (Fragaria × ananassa), prune (Prunus domestica), and pepper (Capsicum annuum) displaying slightly different metabolic programs from tomato (Solanum lycopersicum; Carrari et al., 2006; Zamboni et al., 2010; Lombardo et al., 2011; Osorio et al., 2011, 2012; Zhang et al., 2011). This fact notwithstanding, tomato has become the primary experimental model in which to study the development and ripening of fleshy fruits (Giovannoni, 2004; Fernandez et al., 2009). Furthermore, it is arguably also the best characterized fruit at the biochemical level (Rose and Bennett, 1999; Carrari et al., 2006; Fraser and Bickmore, 2007; Moco et al., 2007).
One interesting observation is that the levels of several organic acids correlate strongly with transcriptional programs of tomato fruit development (Carrari et al., 2006). This finding led to follow-up work demonstrating that the level of malate played an important role in tomato fruit ripening influencing starch accumulation, total soluble solid levels at ripening, and postharvest properties (Centeno et al., 2011), when taken alongside the results of many recent studies on the control of stomatal aperture by malate (Nunes-Nesi et al., 2007; Lee et al., 2008; Araújo et al., 2011; Penfield et al., 2012), thus expanding the documented biological roles for this acid beyond those previously documented (Fernie and Martinoia, 2009; Meyer et al., 2010). This finding thus suggests that malate metabolism may exert a key influence on the normal ripening and metabolism of tomato fruit. Many of our previous studies were targeted at mitochondrial reactions of the tricarboxylic acid (TCA) cycle (for review, see Sweetlove et al., 2010; Nunes-Nesi et al., 2011); here, however, we took RNA interference (RNAi) strategies to silence either the cytosolic phosphoenolpyruvate carboxykinase (PEPCK) or the plastidic NADP-dependent malic enzyme (ME) under the control of the ripening-specific E8 promoter. In plants, PEPCK is a cytosolic enzyme that catalyzes the ATP-dependent decarboxylation of oxaloacetate (OAA) to phosphoenolpyruvate (PEP; Chollet et al., 1996). In tomato, PEPCK abundance increases at the start of ripening, during which there is a decrease in malate content (Bahrami et al., 2001). This has led to the suggestion that PEPCK is involved in this dissimilation and that part of this dissimilated malate may be used in gluconeogenesis (Leegood and Walker, 2003; Famiani et al., 2005, 2009). In contrast, NADP-ME isoforms function in chloroplasts and the cytosol (Drincovich et al., 2001; Gerrard Wheeler et al., 2005) and catalyze the reversible conversion of malate and pyruvate (Farineau, 1977; Drincovich et al., 2001). In tomato, NADP-ME has been implicated in respiration during ripening, providing pyruvate and/or NADPH as a substrate for respiration (Farineau, 1977; Edwards and Andreo, 1992).
In this study, the two generated tomato lines (cv MicroTom) with reduced cytosolic PEPCK and plastidic NADP-ME gene expression were characterized at morphological and biochemical levels, including comprehensive metabolite profiling and an evaluation of the effect of these genetic interventions on cellular fluxes. The results are discussed with respect to current models of energy metabolism within the fruit and with respect to the roles of carboxylic acids during fruit ripening and development.
RESULTS
Generation of Tomato Plants with Fruit-Specific Inhibition of PEPCK and NADP-ME
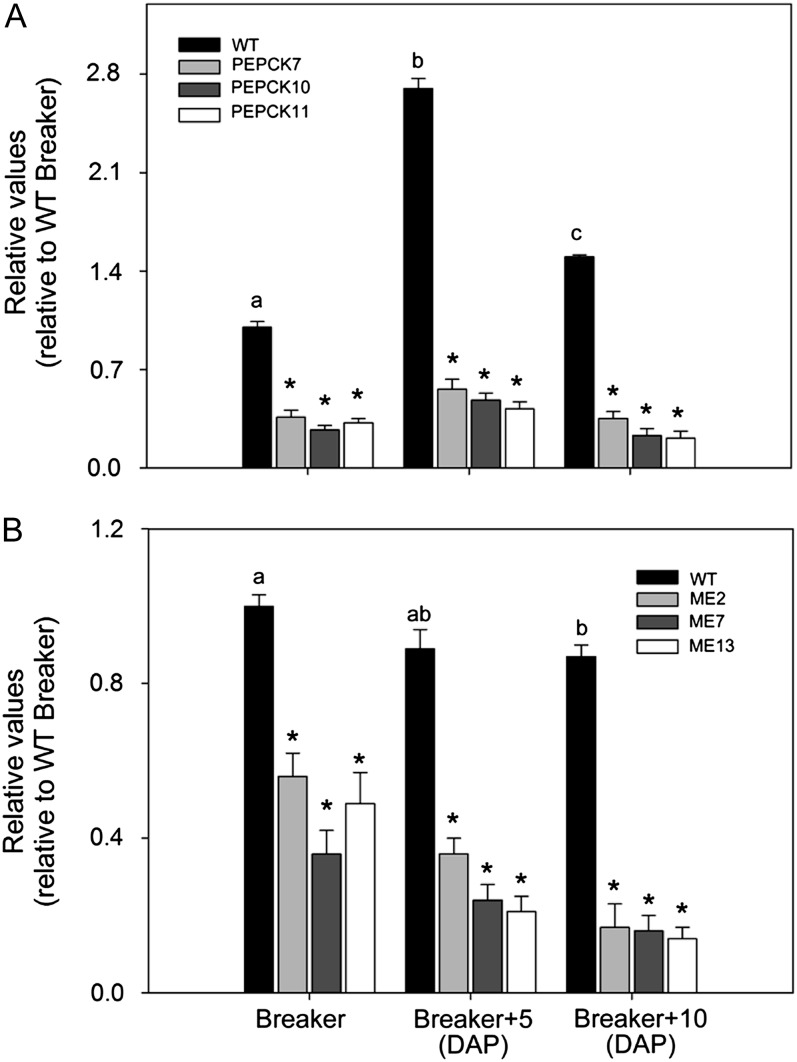
An 560-bp fragment of the complementary DNA (cDNA) encoding cytosolic PEPCK (accession no. AY007226) and a 601-bp fragment of the cDNA encoding a tomato plastidic NADP-ME (accession no. AF001269) were independently cloned using RNAi design into the pBINplus under the control of the ripening promoter E8. Plants (cv Micro-Tom) obtained by an Agrobacterium tumefaciens-mediated transformation were grown in the greenhouse, and seeds were collected. Eight lines for RNAi-PEPCK and nine lines for RNAi-ME were selected on the basis of PEPCK and NADP-ME1 gene expression by quantitative real-time (qRT)-PCR (Fig. 1, A and B). Three lines per construct were chosen: PEPCK7, PEPCK10, and PEPCK11 (PEPCK lines) and ME2, ME7, and ME13 (ME lines). Eight T2 plants per line were subsequently grown in the greenhouse, and fruits at breaker, breaker + 5 d after pollination (DAP), and breaker + 10 DAP were harvested. Assays of PEPCK and NADP-ME activities revealed that the lines displayed considerable reduction in activity during ripening (Tables I and II), and qRT-PCR analysis confirmed that they additionally displayed reduced expression of the PEPCK and NADP-ME1 genes, respectively (Fig. 2). We additionally observed an increase in PEPCK level during normal tomato ripening, while the NADP-ME1 level declined gradually at the onset of ripening as described previously (Bahrami et al., 2001; Tomato Genome Consortium, 2012; Fig. 2).
Figure 1.
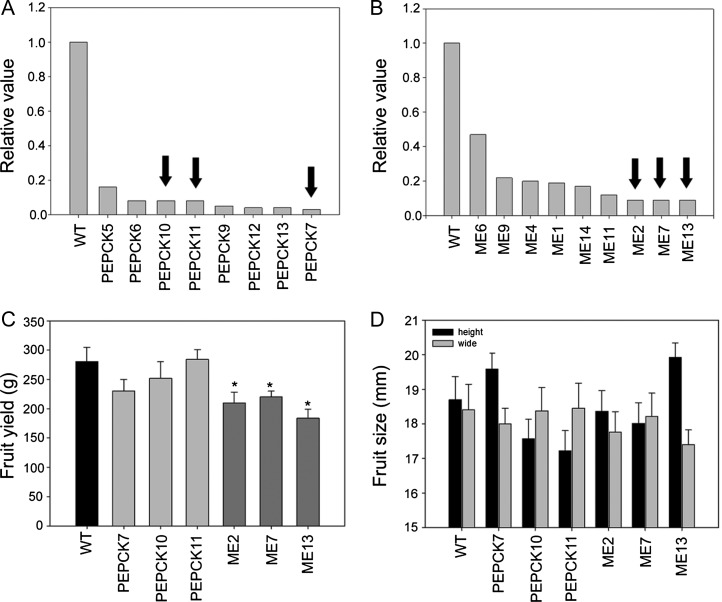
Characterization and expression of tomato cytosolic PEPCK and NADP-ME. A and B, Screening by qRT-PCR in primary PEPCK (A) and ME (B) transformants in extracts of tissue sampled at breaker + 10 DAP. Black arrows indicate lines selected for further analysis. C and D, Total fruit yield (C) and fruit size (D) of the PEPCK and ME lines. For all parameters, values are presented as means ± se of eight biological replicates (one biological replicate is represented by one individual plant). Asterisks indicate values that were determined by Student’s t test to be significantly different (P < 0.05) from the wild type (WT).
Table I. PEPCK activity during tomato fruit ripening of PEPCK lines.
Values (μmol min−1 g−1 fresh weight) are presented as means ± se of six biological determinations. Values that are significantly different by Student’s t test from the wild type are set in boldface (P < 0.05).
Table II. NADP-ME activity during tomato fruit ripening of ME lines.
Values (μmol min−1 g−1 fresh weight) are presented as means ± se of six biological determinations. Values that are significantly different by Student’s t test from the wild type are set in boldface (P < 0.05).
Figure 2.
PEPCK and ME expression during the ripening of PEPCK and ME lines. The abundance of PEPCK (accession no. AY007226; A) and NADPH-ME1 (accession no. AF001269; B) mRNAs was measured by qRT-PCR. The ripening stages were as follows: breaker, breaker + 5 DAP, and breaker + 10 DAP. Values are presented as means ± se of six individual plants per line. Asterisks indicate significant differences of transgenic lines against the wild type (WT) for each stage. Different letters indicate significant differences within the wild-type line in the different ripening stages using ANOVA and the Tukey honestly significant difference test adjusted to 95% significance level.
Specific Expression of SlNADP-ME Genes
The Arabidopsis (Arabidopsis thaliana) genome is known to contain four genes encoding functional NADP-ME isoforms (Gerrard Wheeler et al., 2005). NADP-ME4 is localized to plastids, while NADP-ME1, NADP-ME2, and NADP-ME3 are cytosolic isoforms (Gerrard Wheeler et al., 2005, 2009). The genome of tomato possesses three more genes as part of NADP-ME1 encoding putative NADP-MEs: cytosolic NADP-ME2 (AF001270; Knee and Finger, 1992), plastidic NADP-ME3 (Solyc03g120990; Tomato Genome Consortium, 2012), and a putative NAD(P)-ME4 (Tomato Genome Consortium, 2012). We compared the abundance of all three NADP-MEs and NADP-ME1 transcripts using published tomato (cv Heinz) RNAseq data from three ripening stages (green, breaker, and breaker + 10 DAP; Tomato Genome Consortium, 2012). The expression levels of NADP-ME2 and NADP-ME1 were similar, while the expression of NADP-ME3 and NADP-ME4 accounted for less than 100 times that of NADP-ME1 and NADP-ME2 (Tomato Genome Consortium, 2012). Interestingly, NADP-ME2 and NADP-ME3 expression was decreased during ripening, while NADP-ME4 expression was unaltered during fruit ripening (Supplemental Fig. S1; Tomato Genome Consortium, 2012). Importantly, qRT-PCR evaluation of the expression levels of cytosolic NADP-ME2, plastidic NADP-ME3, and NADP-ME4 during ripening in the transgenic ME lines revealed that the plastidic NADP-ME3 was decreased in all transgenic lines and in all three ripening stages in comparison with the wild type (Supplemental Fig. S1). However, the levels of the transcripts of NADP-ME2 and NADP-ME4 were unaltered in the transformant lines (Supplemental Fig. S1).
Fruit Size and Yield
Fruit size and total fruit yield were determined in fully mature fruit (breaker + 10 DAP). The ME transformants exhibited a significantly lower fruit yield consistent across all three transgenic lines (Fig. 1C). Given that fruit size was essentially unaltered (Fig. 1D), it is apparent that this was a result of a minor decrease in total fruit number in the ME transformants (data not shown). By contrast, PEPCK transformants exhibited unaltered fruit size and yield (Fig. 1, C and D).
Metabolic Profiling of Breaker Fruits of PEPCK and NADP-ME Lines
In order to gain a deeper comprehension of the effects of reducing the expression of these two enzymes, we next determined metabolite levels in the pericarp tissue of breaker-stage fruit using an established gas chromatography-mass spectrometry (GC-MS)-based metabolite profiling method (Schauer et al., 2006). In the PEPCK lines, we observed a reduction in the levels of the amino acids Asp (all three lines), Ile (lines PEPCK7 and PEPCK11), and Thr (lines PEPCK7 and PEPCK10; Fig. 3). Similarly, a decrease in the levels of citric acid (all three lines), Fru and Glc (all three lines), as well as putrescine (lines PEPCK7 and PEPCK11) was observed (Fig. 3). Additionally, PEPCK lines showed an increase in the levels of Glu (all three lines) and galactinol (PEPCK10 and PEPCK11; Fig. 3). In the ME lines, a smaller number of changes were observed. These lines were characterized by increases of some amino acids, such as Asp (lines ME7 and ME13), Ser (lines ME7 and ME13), and Val (lines ME7 and ME13; Fig. 4), sugars and sugar derivatives such as Fuc (lines ME2 and ME7) and myoinositol (lines ME2 and ME13; Fig. 4), as well as putrescine (lines ME2 and ME13).
Figure 3.
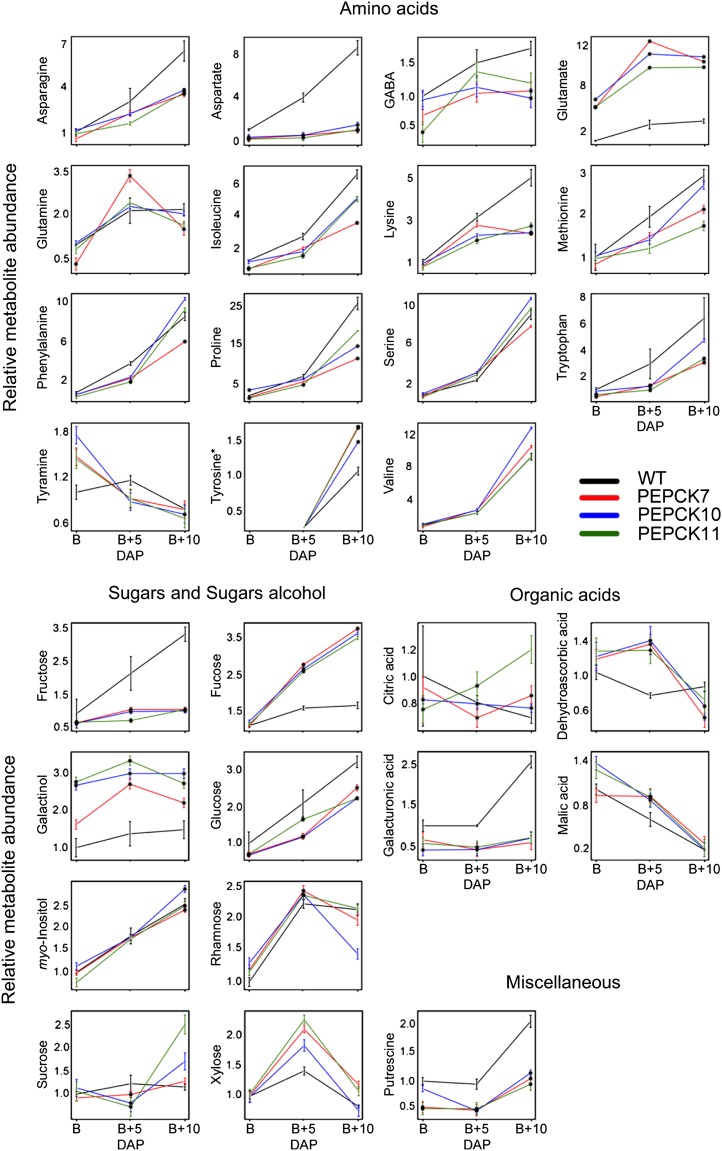
Primary metabolite levels in the receptacle of the wild type (WT) and PEPCK lines at three ripening stages (breaker [B], breaker + 5 DAP [B+5 DAP], and breaker + 10 DAP [B+10 DAP]). Data are normalized to mean values of the wild type at the breaker stage. Values are means ± se of three replicates. Black dots indicate significant differences by Student’s t test (P < 0.05) of the transgenic lines compared with the wild type at the same developmental stage.
Figure 4.
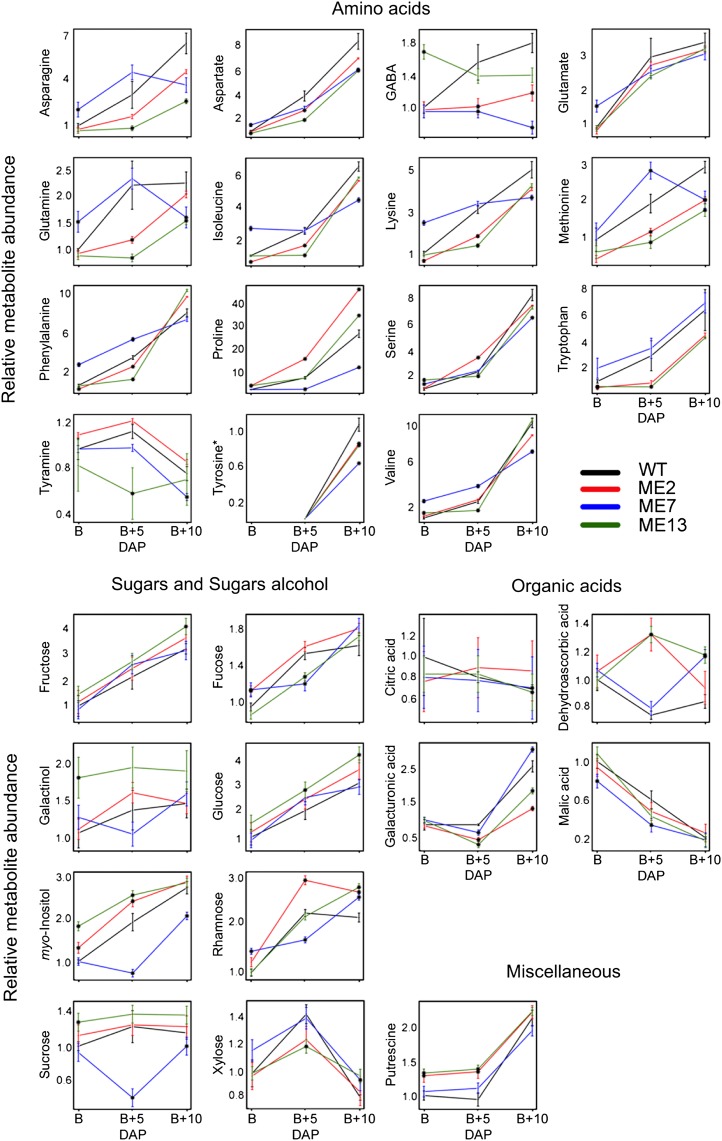
Primary metabolite levels in the receptacle of the wild type (WT) and ME lines at three ripening stages (breaker [B], breaker + 5 DAP [B+5 DAP], and breaker + 10 DAP [B+10 DAP]). Data are normalized to mean values of the wild type at the breaker stage. Values are means ± se of three replicates. Black dots indicate significant differences by Student’s t test (P < 0.05) of the transgenic lines compared with the wild type at the same developmental stage.
Metabolic Profiling during Ripening of PEPCK and NADP-ME Fruits
Given that changes in the content of primary metabolites during the ripening process are highly dynamic (Carrari et al., 2006; Fraser et al., 2007; Osorio et al., 2011), we next evaluated metabolite contents during ripening, breaker + 5 DAP, and breaker + 10 DAP stages. In the PEPCK lines, many metabolites at breaker + 5 DAP and breaker + 10 DAP stages displayed similar patterns of change to those observed at breaker stage. This behavior was observed for amino acids such as Asp (all three lines) and Glu (all three lines) as well as for sugars such as Fru (all three lines), galactinol (all three lines), Glc (all three lines), Fuc (all three lines for breaker + 5 DAP and lines PEPCK7 and PEPCK10 for breaker + 10 DAP), and putrescine (all three lines; Fig. 3). Other changes of note in the metabolite profiles at breaker + 5 DAP stage were the significant increases in dehydroascorbic acid (all three lines), malic acid (all three lines), and Rha (all three lines; Fig. 3) as well as decreases in Trp (all three lines) and GalA (all three lines; Fig. 3). Notably, at breaker + 10 DAP stage, Asn (all three lines), Met (lines PEPCK7 and PEPCK11), Pro (lines PEPCK7 and PEPCK10), γ-aminobutyrate (GABA; lines PEPCK7 and PEPCK10), and myoinositol (lines PEPCK7 and PEPCK11) were significantly decreased, while Tyr was significantly increased (all three lines; Fig. 3). By contrast, evaluation of the primary metabolite composition of ME lines revealed that the pattern of changes in metabolites was more conserved in the comparison between breaker + 5 DAP and breaker + 10 DAP stages than with the breaker stage (Fig. 4). Only a reduction in Asp was conserved between the three studied fruit stages. In addition, other conserved changes between breaker + 5 DAP and breaker + 10 DAP stages were noted, such as decreases in the levels of some amino acids, such as GABA, Met, and Tyr, as well as in the organic acid GalA (Fig. 4). Increases of dehydroascorbic acid and Rha were observed in the transformants at these developmental stages (Fig. 4).
Deficiency in PEPCK and NADP-ME Leads to Alteration in Nucleotide and Starch Metabolism
To further characterize these lines, we evaluated one of the major products of the TCA cycle, namely pyrimidine-reducing equivalents. This analysis revealed a decrease in NADPH levels in breaker stage of the PEPCK lines (lines PEPCK7 and PEPCK11) but an increase in NAD+ (lines PEPCK10 and PEPCK11). Thus, the NADPH/NADP+ and NADH/NAD+ ratios slightly decreased and increased in PEPCK11, respectively (Table III).
Table III. Pyridine nucleotide levels in breaker fruits in PEPCK antisense lines.
Values (nmol g−1 fresh weight) are presented as means ± se of six biological determinations. Values that are significantly different by Student’s t test from the wild type are set in boldface (P < 0.05).
| Pyridine Nucleotide | Wild Type | PEPCK7 | PEPCK10 | PEPCK11 |
|---|---|---|---|---|
| NADPH | 43.32 ± 1.73 | 33.05 ± 0.94 | 40.93 ± 1.10 | 31.62 ± 1.43 |
| NADP+ | 10.37 ± 0.84 | 8.34 ± 0.34 | 9.76 ± 1.02 | 10.34 ± 0.74 |
| NADP(H) | 53.69 ± 2.81 | 41.39 ± 1.1 | 50.69 ± 2.18 | 41.96 ± 2.2 |
| NADPH/NADP+ | 4.18 ± 0.51 | 3.96 ± 0.27 | 4.19 ± 0.55 | 3.06 ± 0.36 |
| NADH | 74.27 ± 2.45 | 71.37 ± 1.67 | 78.22 ± 3.02 | 75.2 ± 2.89 |
| NAD+ | 33.47 ± 1.39 | 29.48 ± 2.04 | 37.48 ± 1.34 | 28.77 ± 0.94 |
| NAD(H) | 107.74 ± 5.58 | 100.85 ± 5.49 | 115.7 ± 6.61 | 103.97 ± 4.90 |
| NADH/NAD+ | 2.22 ± 0.17 | 2.42 ± 0.22 | 2.09 ± 0.16 | 2.61 ± 0.19 |
By contrast, the pattern of change of pyrimidine levels in the ME lines was markedly different. At breaker stage, the levels of NADP+ increased in one line (ME13). As a consequence, the NADPH/NADP+ ratio decreased (Table IV).
Table IV. Pyridine nucleotide levels in breaker fruits in ME antisense lines.
Values (nmol g−1 fresh weight) are presented as means ± se of six biological determinations. Values that are significantly different by Student’s t test from the wild type are set in boldface (P < 0.05).
| Pyridine Nucleotide | Wild Type | ME2 | ME7 | ME13 |
|---|---|---|---|---|
| NADPH | 43.32 ± 1.73 | 40.48 ± 0.83 | 41.38 ± 1.34 | 44.27 ± 1.44 |
| NADP+ | 10.37 ± 0.84 | 9.36 ± 0.57 | 12.36 ± 1.48 | 15.37 ± 1.23 |
| NADP(H) | 53.69 ± 2.81 | 49.84 ± 1.21 | 53.74 ± 3.44 | 59.64 ± 3.14 |
| NADPH/NADP+ | 4.18 ± 0.51 | 4.32 ± 0.35 | 3.35 ± 0.51 | 2.88 ± 0.32 |
| NADH | 74.27 ± 2.45 | 75.27 ± 1.45 | 72.45 ± 1.72 | 73.11 ± 0.95 |
| NAD+ | 33.47 ± 1.39 | 35.40 ± 0.93 | 31.23 ± 1.10 | 33.13 ± 0.86 |
| NAD(H) | 107.74 ± 5.58 | 110.67 ± 2.57 | 103.68 ± 3.37 | 106.24 ± 1.73 |
| NADH/NAD+ | 2.22 ± 0.17 | 2.13 ± 0.10 | 2.32 ± 0.14 | 2.21 ± 0.09 |
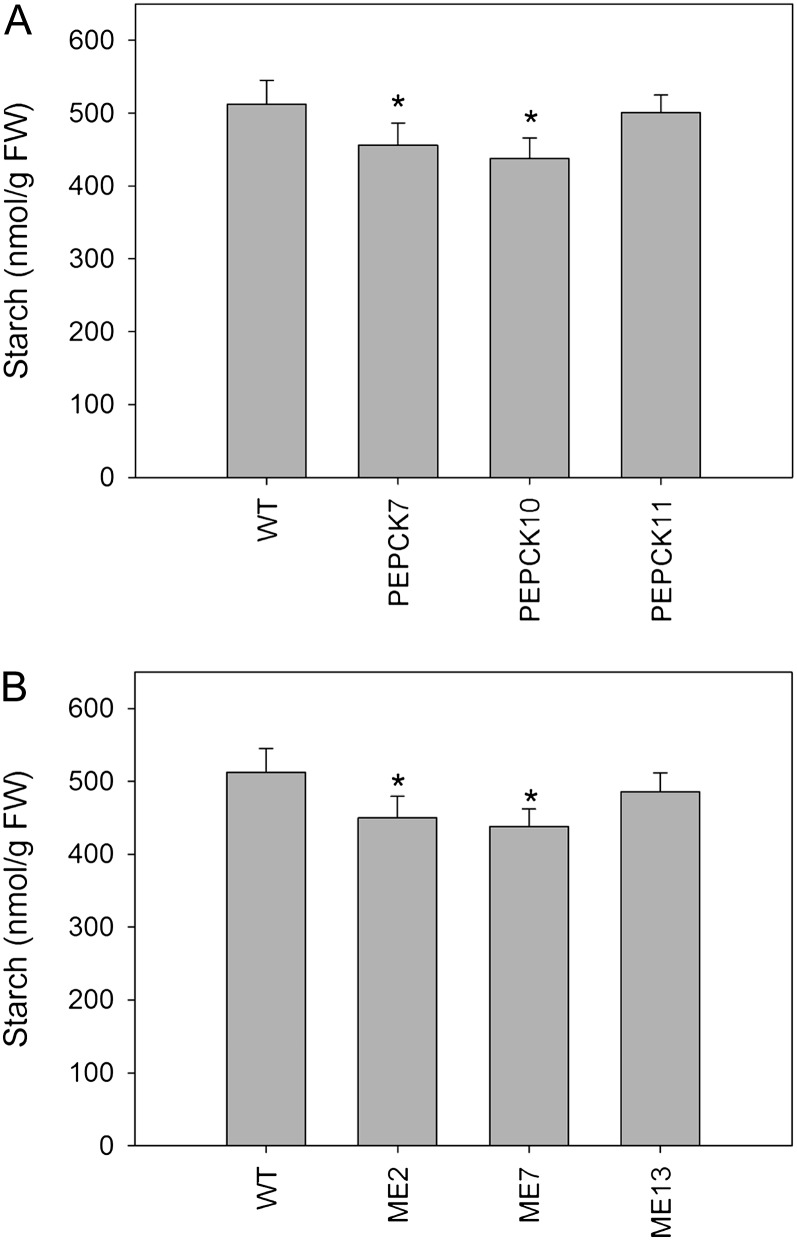
Since pyruvate and PEP metabolites are transformed via the gluconeogenic pathway to sugar phosphates, which can subsequently be converted to starch (Lara et al., 2004), and the fact that previous studies demonstrated that phosphoenolpyruvate carboxylase (PEPC) activity plays an important role within potato (Solanum tuberosum) starch metabolism (Rademacher et al., 2002), we next evaluated the starch content of both the wild type and transformants at breaker stage (Fig. 5). A significant decrease in starch levels was observed in both transgenic sets, PEPCK and ME lines, suggesting that the rate of transient starch degradation in the fruit is, at least partially, dependent on the total reserves of starch available (Fig. 5).
Figure 5.
Starch content of PEPCK and ME lines. Starch content is shown in the PEPCK (A) and ME (B) lines at breaker stage. Values are presented as means ± se of six individual plants per line. Asterisks indicate values that were determined by Student’s t test to be significantly different (P < 0.05) from the wild type (WT). FW, Fresh weight.
Enzymes Involved in Starch and Organic Acid Metabolism
To better understand the above-described metabolic alterations, we next surveyed the activities of a broad range of enzymes from central metabolism (Tables V and VI). In the PEPCK lines, these activities were generally invariant from the wild type, with the exception of the decrease in ADP-glucose pyrophosphorylase (AGPase) and the total NADP-malate dehydrogenase (MDH) activities in two of the lines (PEPCK7 and PEPCK10) at breaker. In addition, a significant reduction was also observed in the level (or activity) of pyruvate kinase (PK) in line PEPCK11 and in NAD-ME and NADP-ME in line PEPCK7 in the same ripening stage (Table V).
Table V. Enzymatic activities in breaker fruits in the PEPCK lines.
Values are presented as means ± se of six biological determinations. Values that are significantly different by Student’s t test from the wild type are set in boldface (P < 0.05). Vsel, Absence of dithiothreitol; Vred, presence of dithiothreitol.
| Enzymatic Activities | Wild Type | PEPCK7 | PEPCK10 | PEPCK11 |
|---|---|---|---|---|
| AGPase + 3PGA (nmol min−1 g−1 fresh wt) | 15.67 ± 1.37 | 11.23 ±1.20 | 10.98 ± 0.98 | 13.34 ± 1.17 |
| AGPase − 3PGA (nmol min−1 g−1 fresh wt) | 11.37 ± 1.02 | 9.34 ± 0.89 | 8.99 ± 0.94 | 10.37 ± 0.85 |
| AGPase activation state (Vsel/Vred) | 0.82 ± 0.04 | 0.71 ± 0.03 | 0.73 ± 0.02 | 0.73 ± 0.01 |
| NADP-MDH total (nmol min−1 g−1 fresh wt) | 0.53 ± 0.07 | 0.37 ± 0.03 | 0.41 ± 0.02 | 0.38 ± 0.07 |
| NADP-MDH initial (nmol min−1 g−1 fresh wt) | 0.15 ± 0.04 | 0.14 ± 0.07 | 0.15 ± 0.05 | 0.12 ± 0.06 |
| NADP-MDH activation stage (nmol min−1 g−1 fresh wt) | 0.82 ± 0.05 | 0.78 ± 0.07 | 0.8 ± 0.03 | 0.75 ± 0.07 |
| NAD-MDH (μmol min−1 g−1 fresh wt) | 4.52 ± 0.32 | 4.33 ± 0.41 | 4.83 ± 0.27 | 4.99 ± 0.37 |
| PK (nmol min−1 g−1 fresh wt) | 0.83 ± 0.09 | 0.75 ± 0.07 | 0.88 ± 0.05 | 0.69 ± 0.07 |
| PEPC (μmol min−1 g−1 fresh wt) | 0.29 ± 0.06 | 0.26 ± 0.04 | 0.27 ± 0.07 | 0.24 ± 0.03 |
| NAD-ME (μmol min−1 g−1 fresh wt) | 0.61 ± 0.05 | 0.50 ± 0.04 | 0.59 ± 0.05 | 0.52 ± 0.06 |
| NADP-ME (μmol min−1 g−1 fresh wt) | 1.23 ± 0.08 | 0.87 ± 0.05 | 1.15 ± 0.09 | 1.26 ± 0.08 |
Table VI. Enzymatic activities in breaker fruits in the ME lines.
Values are presented as means ± se of six biological determinations. Values that are significantly different by Student’s t test from the wild type are set in boldface (P < 0.05). Vsel, Absence of dithiothreitol; Vred, presence of dithiothreitol.
| Enzymatic Activities | Wild Type | ME2 | ME7 | ME13 |
|---|---|---|---|---|
| AGPase + 3PGA (nmol min−1 g−1 fresh wt) | 15.67 ± 1.37 | 12.73 ± 1.03 | 11.38 ± 1.12 | 16.73 ± 2.03 |
| AGPase − 3PGA (nmol min−1 g−1 fresh wt) | 11.37 ± 1.02 | 8.36 ± 0.93 | 7.23 ± 1.02 | 12.21 ± 0.93 |
| AGPase activation state (Vsel/Vred) | 0.82 ± 0.04 | 0.69 ± 0.04 | 0.73 ± 0.02 | 0.80 ± 0.04 |
| NADP-MDH total (nmol min−1 g−1 fresh wt) | 0.53 ± 0.07 | 0.37 ± 0.05 | 0.38 ± 0.06 | 0.56 ± 0.04 |
| NADP-MDH initial (nmol min−1 g−1 fresh wt) | 0.15 ± 0.04 | 0.11 ± 0.03 | 0.18 ± 0.04 | 0.18 ± 0.05 |
| NADP-MDH activation stage (nmol min−1 g−1 fresh wt) | 0.82 ± 0.05 | 0.74 ± 0.04 | 0.85 ± 0.05 | 0.88 ± 0.05 |
| NAD-MDH (μmol min−1 g−1 fresh wt) | 4.52 ± 0.32 | 4.01 ± 0.42 | 4.78 ± 0.27 | 3.98 ± 0.34 |
| PK (nmol min−1 g−1 fresh wt) | 0.83 ± 0.09 | 1.21 ± 0.09 | 1.43 ± 0.08 | 0.93 ± 0.07 |
| PEPC (μmol min−1 g−1 fresh wt) | 0.29 ± 0.06 | 0.42 ± 0.05 | 0.37 ± 0.06 | 0.40 ± 0.06 |
| NAD-ME (μmol min−1 g−1 fresh wt) | 0.61 ± 0.05 | 0.68 ± 0.05 | 0.58 ± 0.04 | 0.55 ± 0.06 |
| PEPCK (μmol min−1 g−1 fresh wt; ×10) | 0.36 ± 0.03 | 0.42 ± 0.04 | 0.39 ± 0.03 | 0.32 ± 0.04 |
Similar to the situation in the PEPCK lines, analysis of the ME lines at breaker stage revealed that the activities of both AGPase and NADP-MDH decreased in the lines ME2 and ME7 (Table VI). Moreover, a significant increase in PEPC (all three lines) and PK (lines ME2 and ME7) activities was also observed (Table VI).
Respiration in the Transgenic Fruits
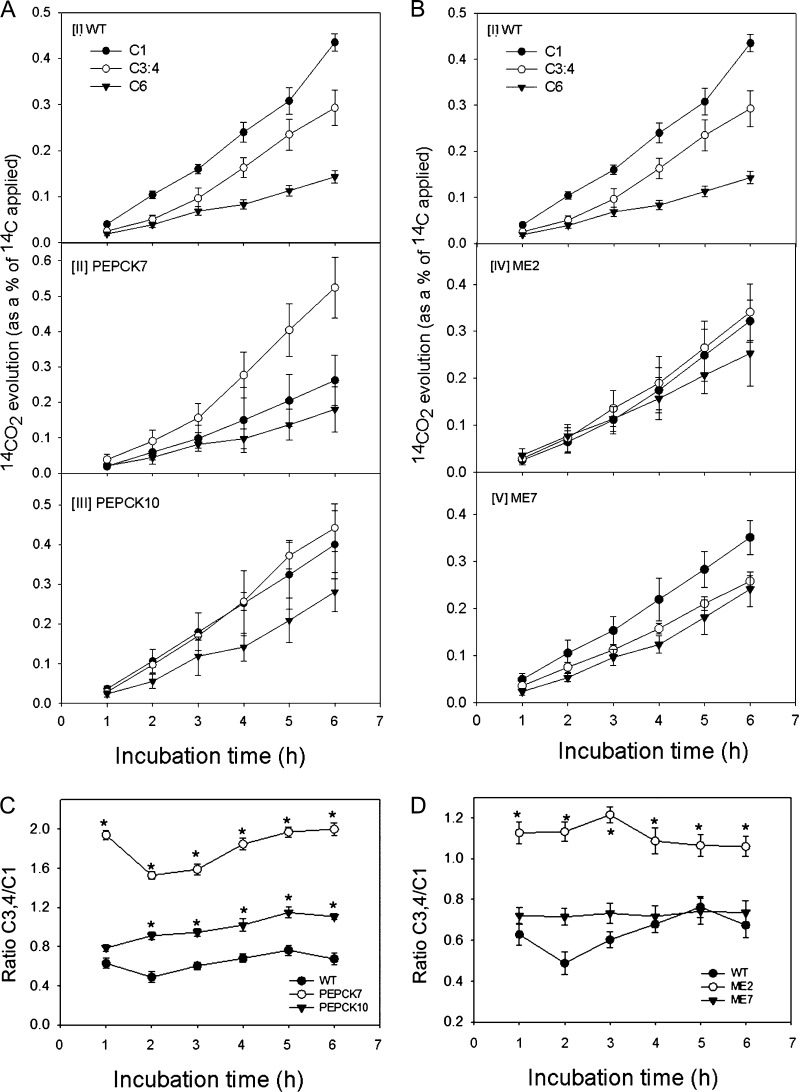
In order to evaluate the effect of deficiencies in the expression of these enzymes on the respiration rate, we next evaluated the relative rate of flux through the major pathways of carbohydrate oxidation at breaker + 5 DAP. For this purpose, we incubated pericarp discs taken from fruits in the light and supplied them with [1-14C]Glc, [3:4-14C]Glc, or [6-14C]Glc for a period of 6 h. During this period, we collected the 14CO2 evolved at hourly intervals. Carbon dioxide can be released from position C1 by the action of enzymes that are not associated with mitochondrial respiration, but carbon dioxide released from the C3,4 positions of Glc cannot (Nunes-Nesi et al., 2005). Thus, the ratio of carbon dioxide evolution from C1 to C3,4 of Glc provides an indication of the relative rate of the TCA cycle in comparison with the other routes of carbohydrate oxidation.
When the relative 14CO2 release of the transgenic and wild-type lines was compared, an interesting pattern emerges, with a relatively unaltered carbon dioxide evolution in the ME lines but the PEPCK lines displaying considerably altered carbohydrate oxidation. Both lines PEPCK7 and PEPCK10 displayed a greater relative release of 14CO2 from C3,4 positions than the wild type, indicating a greater flux through the reactions catalyzed by pyruvate dehydrogenase and/or ME (Fig. 6). The difference in release from the C6 position was less marked, however, suggesting that there was no alteration in fluxes through pentan synthesis (Stitt and ap Rees, 1979). Despite there being little change in the absolute release of 14CO2 from C3,4 in the MADP-ME lines, the relative release from the C3,4 positions was considerably higher in line ME2.
Figure 6.
Respiratory parameters in breaker + 10 DAP fruits of PEPCK and NADP-ME lines. A and B, Evolution of 14CO2 from breaker + 10 DAP pericarp discs of PEPCK (A) and ME (B) lines. C and D, Ratio of dioxide evolution from C3,4/C1 positions of Glc in breaker + 10 DAP pericarp discs of PEPCK (C) and ME (D) lines. WT, Wild type.
Effect of Decreased PEPCK and NADP-ME Activities on Metabolic Fluxes
The effect of reduced PEPCK and NADP-ME activities on carbon fluxes was determined by incubating pericarp discs at breaker + 10 DAP stage from the wild type and two lines of the PEPCK (PEPCK7 and PEPCK10) and ME (ME2 and ME7) transformants with [U-14C]Glc for 2 h. Following this incubation, the discs were washed and frozen prior by fractionation of the labeled material to determine the label redistribution (Table VII). Neither line from the PEPCK or the ME transformants displayed differences from the wild type in the total label uptake or in the metabolized label incorporated into cell wall, amino acids, or Suc (Table VII). However, there was decreased label incorporation into starch, protein, and organic acids in lines of both transformants (Table VII). Moreover, PEPCK7 and PEPCK10 lines showed significant decreases of label into the pool of hexose phosphates (Table VII). Determination of the specific activity of the hexose phosphate pools reveals, as expected, that the PEPCK transformants had proportionally lower label in this metabolic pool. On the calculation of the metabolic fluxes, it emerged that both transformants, PEPCK and ME, displayed dramatic decreases in the rate of starch synthesis (Table VII). A similar pattern was found for the fluxes to organic acids and protein, although these changes are considerably less marked than for starch.
Table VII. Metabolism of [U-14C]Glc in pericarp discs from PEPCK and ME lines in breaker + 10 DAP fruits.
Pericarp discs were cut from six separate plants per genotype. After 2 h of incubation, the discs were extracted and fractioned. The specific activity of the hexose phosphate pool was estimated by dividing the label retained in the phosphate ester pool by the summed carbon of the hexose phosphates and used to calculate absolute fluxes. Values represent means ± se of six biological determinations. Values that are significantly different by Student’s t test from the wild type are set in boldface (P < 0.05).
| Parameter | Wild Type | PEPCK7 | PEPCK10 | ME2 | ME7 |
|---|---|---|---|---|---|
| Label incorporated (Bq) | |||||
| Total uptake | 287.2 ± 16.5 | 235.2 ± 12.2 | 208.3 ± 14.8 | 260.6 ± 19.2 | 269.5 ± 5.4 |
| Insoluble | |||||
| Starch | 10.2 ± 0.2 | 5.5 ± 0.6 | 6.3 ± 1.0 | 4.8 ± 0.6 | 5.1 ± 0.2 |
| Cell wall | 5.1 ± 0.7 | 5.2 ± 0.5 | 6.4 ± 1.0 | 4.1 ± 0.5 | 6.0 ± 0.7 |
| Protein | 3.0 ± 0.3 | 1.7 ± 0.1 | 1.4 ± 0.1 | 0.9 ± 0.1 | 1.7 ± 0.2 |
| Soluble | |||||
| Organic acids | 130.7 ± 8.6 | 96.9 ± 11.3 | 65.0 ± 3.8 | 76.7 ± 6.4 | 86.7 ± 6.6 |
| Total hexose phosphates | 11.3 ± 2.2 | 6.1 ± 2.7 | 6.4 ± 2.1 | 10.2 ± 3.5 | 12.3 ± 2.8 |
| Amino acids | 104.3 ± 2.2 | 84.5 ± 7.7 | 85.4 ± 7.0 | 125.1 ± 9.0 | 114.9 ± 16.1 |
| Suc | 31.1 ± 4.4 | 27.3 ± 1.1 | 27.7 ± 2.4 | 30.8 ± 1.7 | 31.4 ± 2.7 |
| Specific activity (dpm nmol−1 g−1 fresh wt) | |||||
| Hexose phosphate pool | 19.3 ± 0.8 | 13.6 ± 0.7 | 12.9 ± 0.5 | 19.3 ± 0.5 | 18.3 ± 0.4 |
| Metabolic flux (nmol−1 g−1 fresh wt) | |||||
| Starch | 621.0 ± 94.3 | 434.2 ± 75.4 | 376.4 ± 112.3 | 587.3 ± 84.4 | 393.0 ± 77.5 |
| Cell wall | 84.4 ± 21.3 | 94.3 ± 17.4 | 103.3 ± 17.3 | 91.8 ± 20.3 | 103.3 ± 19.4 |
| Organic acids | 193.4 ± 22.5 | 134.4 ± 14.4 | 153.5 ± 15.5 | 89.5 ± 17.4 | 149.4 ± 23.5 |
| Amino acids | 224.5 ± 17.5 | 204.4 ± 17.2 | 194.4 ± 20.4 | 248.4 ± 16.4 | 235.4 ± 21.4 |
| Protein | 54.4 ± 9.4 | 39.4 ± 8.4 | 37.2 ± 10.8 | 28.4 ± 8.2 | 68.3 ± 6.4 |
| Suc | 76.3 ± 7.4 | 86.3 ± 6.4 | 79.3 ± 5.6 | 80.3 ± 9.4 | 70.4 ± 7.8 |
Redistribution of 13C Label following [13C]Pyruvic Acid Feeding of Transgenic Fruits
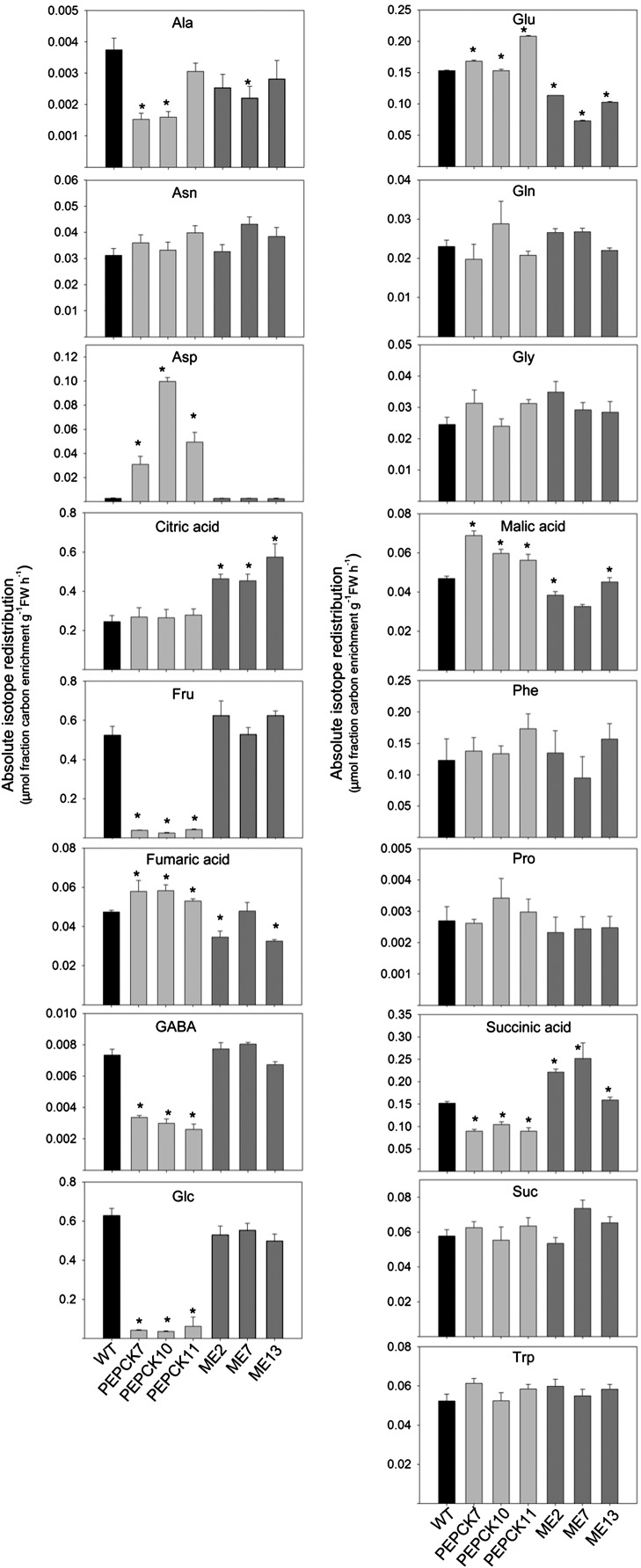
To further investigate changes in primary metabolism, we fed isotopically labeled [13C]pyruvic acid (Pyr) to breaker + 10 DAP fruits from PEPCK and ME lines. The absolute redistribution of 13C was measured by assessing the incorporation of the isotope into each intermediate within the relevant pathways (Fig. 7) in metabolites closely associated with Pyr using an adapted GC-MS protocol that facilitates isotope tracing (Roessner-Tunali et al., 2004). Interestingly, the changes in redistribution of isotope were conserved across the transformants.
Figure 7.
Redistribution of 13C label following the incubation of transgenic lines (PEPCK and ME) and wild-type (WT) fruits (breaker + 10 DAP stage). The absolute isotope redistribution (μmol fraction carbon enrichment g−1 fresh weight [FW] h−1) is shown after an incubation period of 6 h of [U-13C]Pyr. Values are means ± se of determinations on four independent samples; asterisks indicate values that were determined by Student’s t test to be significantly different (P < 0.05) from the wild type.
The PEPCK lines were characterized by elevated label incorporation in fumaric acid, malic acid, and Glu in addition to dramatic elevation in label incorporation in Asp. Moreover, they displayed reduced incorporation in Ala, GABA, Fru, Glc, and succinic acid (Fig. 7). However, the pattern of changes in label accumulation in the ME lines was different. This suggested that there were significant increases in label incorporation to citric acid and succinic acid. In contrast, decreased label incorporation in Ala, Glu, fumaric acid, and malic acid was observed (Fig. 7).
DISCUSSION
During the ripening of tomato, there is a large decrease in the organic acid content of the pericarp (Knee and Finger, 1992; Carrari et al., 2006). This decrease in organic acids could be brought about either by a restriction of their synthesis or an enhanced degradation potentially initially catalyzed by NAD-ME, NADP-ME, or PEPCK. Malic acid has long been implicated in the physiology of fruit ripening (Carrari and Fernie, 2006; Sweetman et al., 2009). It has recently been demonstrated that the level of malic acid plays an important role in the regulation of starch biosynthesis and thereby the accumulation of total soluble solid during tomato ripening. Detailed biochemical characterization revealed that this was due to regulation of the activation state of the first committed step of starch synthesis that is catalyzed by AGPase (Centeno et al., 2011). Intriguingly, however, this is not the first study to implicate reactions beyond the direct pathway of starch biosynthesis, since the mitochondrial NAD-ME (Jenner et al., 2001), the plastidial adenylate kinase (Regierer et al., 2002), and the cytosolic UMP synthase (Geigenberger et al., 2005) have been demonstrated to be important in determining the levels of starch in storage organs. However, the exact mechanisms of these effects of starch synthesis are essentially unknown, although in the case of the adenylate kinase manipulation both the substrate levels and activation status of the AGPase reaction were affected (Oliver et al., 2008). In addition, the manipulation of malate metabolism in tomato has been demonstrated to affect stomatal opening (Nunes-Nesi et al., 2007; Lee et al., 2008; Araújo et al., 2011; Penfield et al., 2012) and normal root development (van der Merwe et al., 2009). In our previous studies, we focused on mitochondrial reactions of the TCA cycle, and here, we evaluated the effect of modifying malate-pyruvate balances at different subcellular locations in a fruit ripening-specific manner by silencing either cytosolic PEPCK or the plastidic NADP-ME.
Effects of Cytosolic PEPCK and Plastidic NADP-ME on Starch Biosynthesis
Different studies concerning starch metabolism in potato and tomato have suggested that AGPase activity plays an important role in regulation (Geigenberger et al., 1999; Sweetlove et al., 1999; Geigenberger, 2011). AGPase activity is known to be modulated via several different mechanisms. AGPase is sensitive to allosteric regulation, being inhibited by inorganic phosphate and activated by 3-phosphoglycerate (3PGA; Sowokinos, 1981; Sowokinos and Preiss, 1982). Additionally, it has been demonstrated to be transcriptionally regulated by sugars, nitrate, phosphate, and trehalose-6-phosphate (Müller-Röber et al., 1990; Nielsen et al., 1998; Kolbe et al., 2005; Michalska et al., 2009). Moreover, it has been described that AGPase is also redox regulated (Tiessen et al., 2002; Centeno et al., 2011), with malic acid potentially being a key component in this process at least in photosynthetically active tissues (Szecowka et al., 2012).
In tomato, as previously mentioned, a differential regulation of starch biosynthesis was observed in transgenic lines exhibiting different fruit malic acid levels in which the activation stage of AGPase correlated with the accumulation of transitory starch (Centeno et al., 2011). Similar, in this context, was the fact that the down-regulation of cytosolic PEPCK during ripening resulted in a significantly higher malic acid level at the breaker + 5 DAP stage. However, similar down-regulation of the plastidic NADP-ME during ripening displayed no significant changes in the malic acid level at steady state.
The observed decrease in starch content in the cytosolic PEPCK antisense lines at the breaker stage seems to result from an alteration in the malic acid level, as observed previously in the TCA cycle transformants (Centeno et al., 2011). However, this correlation was not apparent in plastidic NADP-ME antisense lines, which also revealed decreased starch content. Interestingly, however, we observed a significant decrease of NADPH in the cytosolic PEPCK lines, which leads to a slight decrease of the NADPH/NADP+ ratio. On the other hand, only one line showed changes in the plastidial redox state in the plastidic NADP-ME antisense lines. Consistent with these changes in the plastidial redox status were the alterations in the activities and activation state of AGPase (in PEPCK and ME lines) and plastidial malate dehydrogenase (in ME lines), which are well documented diagnostics of alteration in redox status (Scheible and Stitt, 1988; Tiessen et al., 2002; Nashilevitz et al., 2010). The lack of absolute starch accumulation in both lines was also corroborated with a decrease in the starch biosynthesis flux. Thus, the observations described here demonstrate that, as we postulated previously, starch metabolism is strongly affected by changes in plastidial redox state but seems not to alter malate levels per se.
When other areas of metabolism are considered, some interesting observations are apparent. It is suggested that in fruit ripening, NADP-ME may produce the respiratory substrates pyruvate and NADPH from malate and NADP+ (Ruffner, 1982; Drincovich et al., 2001) in cytosol and maybe thus partake in malate degradation during ripening (Ruffner et al., 1984; Franke and Adams, 1995). Here, feeding experiments involving the incubation of positionally labeled Glcs showed that the strong decrease in plastidic NADP-ME levels did not lead to substantial changes in overall respiration rates and TCA cycle flux in tomato fruits. However, it is important to note that one line (ME2) was characterized by an increase in the C3,4-C1 ratio, also suggestive of an increased flux through pyruvate dehydrogenase (albeit one not mediated by an elevated enzymatic capacity of the NAD-ME). It is conceivable that the remaining activity of plastidic NADP-ME might still be high enough to supply the TCA cycle with sufficient pyruvate to maintain the overall respiration rate or that there are other alternative sources that may be able to continue to supply the mitochondria with pyruvate or some other carbon source, such as cytosolic NADP-ME, PK, and PEPC. In light of these observations, it is interesting that cytosolic NADP-ME expression was not modified in the ME lines. Interestingly, the activities of PK and PEPC enzymes displayed a clear increase in the lines with reduced plastidic NADP-ME activity. These observations indicate that the lack of plastidic NADP-ME activity increased the flow through these enzymes, which suggests that plastidic NADP-ME activity is likely an important source of pyruvate synthesis in ripening tomato fruits.
Intriguingly, the large decrease in the organic acid content frequently documented as occurring in the pericarp on ripening (Knee and Finger, 1992; Carrari et al., 2006) correlates with a large increase in PEPCK (Bahrami et al., 2001). This prompts the question, what is the contribution of this enzyme to flux through the TCA cycle? If the TCA cycle is drained via OAA, PEPCK activity requires support from neither PEPC nor PK nor from any part of the TCA cycle. However, we can exclude an increase of PK and PEPC activities in compensation for the decrease in cytosolic PEPCK. Moreover, in the PEPCK lines, uniformly labeled Glc feeding experiments also revealed a reduced flux to protein. Furthermore, feeding experiments involving the incubation of positionally labeled Glcs suggested that the PEPCK lines displayed higher flux through the reactions catalyzed by pyruvate dehydrogenase, ME, or both. This can be concluded from the increased C3,4-C1 ratios displayed by PEPCK lines, since the carbon release from the C3,4 positions occurs via decarboxylation catalyzed by these enzymes. Both the mitochondrial NAD-ME and plastidial NADP-ME activities showed no differences in the PEPCK lines, suggesting that these fruits showed high flux through the TCA cycle via pyruvate dehydrogenase.
PEP required for gluconeogenesis in fruit may originate from malate through the activities of MDH and PEPCK or potentially MEs (Ruffner, 1982). Evidence from radiolabeling work in tomato fruit (Farineau, 1977; Halinska and Frenkel, 1991) suggested that gluconeogenesis does occur during ripening stages, when sugars are accumulating rapidly. Here, we observed that when cytosolic PEPCK was reduced (PEPCK lines), this led to reduced levels of the major sugars, Glc and Fru, during fruit ripening, accompanied by an accumulation of malate, which provides further evidence for gluconeogenesis from organic acids (Halinska and Frenkel, 1991) and the important role of PEPCK enzymes in this process.
In order to address the extent to which PEPCK activity contributes to the regulation of the TCA cycle, it is first necessary to consider how individual carbon fluxes around the complete TCA cycle differ depending on the particular biosynthetic output. When biosynthesis is absent, the net flux of carbon through each TCA cycle step will be equal, likewise if the TCA cycle’s biosynthetic output is exclusively from OAA to Asp. By contrast, if the biosynthetic output of the TCA cycle is exclusively from 2-oxoglutarate to Glu, then the flux from malate to 2-oxoglutarate will be proportionately higher than the flux from 2-oxoglutarate to malate. Bearing these considerations in mind, it is interesting that we observed that during ripening, the level of Asp derived from OAA was highly reduced in fruits deficient in PEPCK activity compared with the wild type. By contrast, the steady-state level of Glu was higher than in the wild type during ripening. However, 13C feeding experiments revealed a higher redistribution of the label in both metabolites (Asp and Glu). Interestingly, a higher redistribution of 13C label in malic and fumaric acids as well as lower label in succinic acid were also displayed in the PEPCK lines. Although with these data, the flux of carbon through the TCA cycle from malate to 2-oxoglutarate and from 2-oxoglutarate to malate cannot be calculated, these data indicate that when carbon in the form of OAA cannot be converted into PEP through the activity of PEPCK and thus cannot completely flow through the TCA cycle, the flux from TCA intermediates toward the synthesis of Asp is increased and reintroduced in the TCA cycle via 2-oxoglutarate/Glu.
In conclusion, we present here compelling evidence of the importance of both cytosolic PEPCK and plastidic NADP-ME in normal tomato ripening metabolism. Both enzymes exhibited considerable influence on starch biosynthesis metabolism. Their functions in other areas of metabolism, however, are somewhat divergent with our results, suggesting that the lack of accumulation of transitory starch was reflected by a decrease of major sugars in the PEPCK lines. This observation was not seen in plastidic NADP-ME. Moreover, the inactivation of PEPCK affected the respiration rate, suggesting that excess OAA was converted to Asp and reintroduced in the TCA cycle via 2-oxoglutarate/Glu. These results indicate that the cytosolic PEPCK plays a role in carbon and nitrogen remobilization pathways during fruit ripening. However, further studies will be required to investigate these processes, and detailed analysis of starch metabolism revealed that this was impacted by the effect of alterations in the plastid redox status on AGPase. As such, these data provide direct support for our earlier model (Centeno et al., 2011), suggesting that malate-mediated effects on starch metabolism are caused by cellular redox changes, AGPase, and pyrimidine nucleotide levels (in PEPCK lines). Rather unexpected, in this context, was the fact that the down-regulation of plastidic NADP-ME had an effect on starch and AGPase activation state but no effect was observed on malate levels. This observation demonstrates that changes in plastidial redox state may be totally independent of changes in the malate levels. Another explanation is that reduction in plastidic NADP-ME activity provokes a small reduction in plastidial malate that was not observed in the global malate pool, but it was enough to display a change in redox state to affect the activation state of AGPase. Increasing plastidial malate dehydrogenase corroborated the change in redox state in these plants (Scheible and Stitt, 1988; Tiessen et al., 2002; Nashilevitz et al., 2010).
Additionally, we also found that the negligible impact of NADPH-ME activity on respiratory metabolism can be explained by increasing PK and PEPC activities, which can compensate for the reduction in pyruvate derived from malate via plastidic NADP-ME.
Intriguingly, the changes in redox state observed here are not as severe as those described previously for TCA cycle manipulations, despite the fact that the inhibition of the enzymes modified here is at least as efficient as that in our previous study. For example, the lack of apparent changes in pigmentation or postharvest characteristics indicates that the plastidic NADP-ME and cytosolic PEPCK reactions are quantitatively less important for malate-mediated ripening control than those catalyzed by the mitochondrial malate dehydrogenase and fumarase enzymes. There have been recent advances in our understanding of the regulation of starch synthesis in response to metabolic signals. However, our knowledge of the signal transduction cascades has not been elucidated, although this work highlights the complex interactions that exist between different metabolic pathways and suggests novel approaches for the manipulation of starch synthesis.
MATERIALS AND METHODS
Generation of Transgenic Plants
A 560-bp fragment of SlPEPCK (accession no. AY007226) and a 601-bp fragment of SlNADP-ME (accession no. AF001269) were independently cloned using the RNAi approach into the vector PBINplus, between the E8 promoter and the octopine synthase gene terminator. These constructs were subsequently introduced into tomato (Solanum lycopersicum) plants by the Agrobacterium tumefaciens-mediated transformation protocol. Plants were selected and maintained as described in the literature (Tauberger et al., 2000). For initial screening, eight RNAi-PEPCK lines and nine RNAi-ME lines were selected on the basis of PEPCK and NADP-ME1 gene expression by qRT-PCR. This screening allowed the identification of three lines per construct that were taken to the next generation.
Analysis of PEPCK and NADP-ME mRNA Expression by qRT-PCR
Total RNA was extracted according to Bugos et al. (1995) with minor modifications. The integrity of the extracted RNA was checked by electrophoresis under denaturing conditions after treating the RNA with RNase-free DNaseI (Roche). First-strand cDNA synthesis of 1 mg of RNA in a final volume of 20 mL was performed with Moloney murine leukemia virus reverse transcriptase, Point Mutant RNase H Minus (Promega), according to the supplier’s protocol using oligo(dT) T19 primer.
Expression of the cytosolic PEPCK and plastidic NADP-ME genes was analyzed by qRT-PCR using the fluorescent intercalating dye SYBR Green in an iCycler detection system (Bio-Rad; http://www.bio-rad.com/). Relative quantification of the target expression level was performed using the comparative cycle threshold method. The following primers were used: for analysis of PEPCK transcript levels (GenBank accession no. AY007226), forward, 5′-AGACGAAACCACTGAGGACGA-3′, reverse, 5′-CATTCACAAACACCTTCTCCAA-3′; for NADP-ME1 (GenBank accession no. AF001269), forward, 5′-CGGTTGAGATGGAGGTTAAATC-3′, reverse, 5′-TTACGCAACAATGAGAAACCAC-3′; for NADP-ME2 (GenBank accession no. AF001270), forward, 5′-TTATTGTTAGTGGTCGCAAGGA-3′, reverse, 5′-CGTAAATGTTCTTCCAGTTCCAG-3′; for NADP-ME3 (GenBank accession no. Solyc03g120990), forward, 5′-CTATGGTGGTGGTCTCTGTGAA-3′, reverse, 5′-CTGGAGCATTTTCCTTCATCTC-3′; and for NADP-ME4 (GenBank accession no. Solyc08g066290), forward, 5′-AAGAGAACTTGGAGAAGGGATT-3′, reverse, 5′-TTATGCGGTTGAGGTAGACGAG-3′. To normalize gene expression for differences in the efficiency of cDNA synthesis, transcript levels of the constitutively expressed elongation factor1a of tomato (GenBank accession no. X14449) were measured using the following primers: forward, 5′-ACCACGAAGCTCTCCAGGAG-3′; reverse, 5′-CATTGAACCCAACATTGTCACC-3′ (Zanor et al., 2009).
Metabolite Profiling
Metabolite extraction, derivatization, standard addition, and sample injection for GC-MS were performed according to Schauer et al. (2006). Both chromatograms and mass spectra were evaluated using TAGFINDER (Luedemann et al., 2008). The level of starch in the tissue was determined exactly as described previously (Fernie et al., 2001). NAD+, NADP+, NADH, and NADPH were determined as described previously (Schippers et al., 2008).
Enzyme Assays
NAD-MDH and PK
These proteins were extracted and assayed as described by Centeno et al. (2011).
AGPase
AGPase activity was measured in the pyrophosphorolysis direction with a spectrophotometric assay as described (Tiessen et al., 2002, 2003). Frozen tissues were homogenized in liquid nitrogen, and approximately 100 mg was extracted rapidly (1 min) with 1 mL of extraction buffer (50 mm HEPES-KOH, pH 7.8, and 5 mm MgCl2) at 4°C. After centrifugation (30 s at 13,000g at 4°C), 10 µL of the supernatant was used for the AGPase assay. The reaction was performed in a total volume of 200 µL containing 50 mm HEPES-KOH, pH 7.8, 5 mm MgCl2, 10 µm Glc-1,6-bisP, 0.6 mm NADP+, 2.5 mm sodium-inorganic pyrophosphate, 1 unit mL−1 phosphoglucomutase, 2.5 units mL−1 Glc-6-P dehydrogenase, and a range of concentrations of ADP-Glc (0.4–1 mm) in the absence of inorganic phosphate and 3PGA, with or without dithiothreitol (10 mm) for activation assay. Reactions were followed online at 340 nm and were linear up to 30 min. AGPase activity was measured in the ADP-Glc-cleaving direction in the absence (Vsel) or presence of dithiothreitol (Vred). The ratio between the activities in these two assays (Vsel/Vred) is termed “activation state”.
PEPCK
Protein extraction was performed as described by Bahrami et al. (2001). The activity of PEPCK was measured in the carboxylation direction by following oxidation of NADH at 340 nm at room temperature (Walker et al., 1999).
NAD-ME and PEPC
Protein extraction and enzyme activity were analyzed as described (Borsani et al., 2009).
NADH-ME
A total of 100 mg of material was suspended in 100 mm Tris-HCl, pH 8.0, 5 mm MgCl2, 2 mm EDTA, 10% (v/v) glycerol, and 10 mm 2-mercaptoethanol. The supernatants were collected after centrifugation (10 min at 13,000g). NADP-ME activity was determined at 340 nm. The reaction mix contains 50 mm Tris-HCl, 10 mm MgCl2, 0.5 nm NADP, and 10 mm malate. The reaction was started by the addition of malate (Voll et al., 2012). The optimal pH of the NADP-ME reaction was determined using different buffers for various pH ranges: 100 mm MES, pH 5.5, 100 mm Tricine-MOPS, pH 7.0, 100 mm Tris-HCl, pH 7.5, 100 mm Tris-HCl, pH 8.0, 100 mm Tris-HCl, pH 8.5, as described by Detarsio et al. (2003). We observed that the optimal pH for the NADP-ME reaction in tomato fruits was at pH 8.0 (Supplemental Fig. S2).
Analysis of [13C]Pyr-Labeled Samples
Discs (10-mm-diameter latitudinal core) from tomato fruit pericarp at breaker + 10 DAP stage from both transgenic lines (PEPCK and ME lines) and the wild type were fed with 34 mm [13C]Pyr (Cambridge Isotope Laboratories) by incubation in buffered solution (10 mm MES-KOH, pH 6.5) for a period of 6 h. At the end of the incubation, discs were snap frozen in liquid nitrogen. They were subsequently extracted in 100% methanol and processed exactly as described (Timm et al., 2008). The metabolic fate of these substrates was subsequently assessed exactly as described previously (Tieman et al., 2006). A total of four biological replicates were used.
Incubation of Plant Material with [U-14C]Glc
Discs (10 mm-diameter latitudinal core) from tomato fruit pericarp at breaker + 10 DAP stage from both transgenic lines (PEPCK and ME lines) and the wild type were taken. Discs were washed three times in fresh incubation medium (10 mm MES-KOH, pH 6.5), and then eight discs were incubated in 5 mL of incubation medium containing 1.00 μCi of [U-14C]Glc (specific activity of 8.11 MBq mmol–1). Samples were then incubated for 2 h before washing again three times in unlabeled incubation medium and freezing in liquid nitrogen until further analysis. All incubations were performed in a sealed 100-mL flask at 25°C and shaken at 150 rpm. The evolved 14CO2 was collected (in hourly intervals) in 0.5 mL of 10% (w/v) KOH, and the amount of radiolabel was subsequently quantified by liquid scintillation counting. After this step, the discs were harvested, washed three times in buffer (eight discs per 100 mL), and frozen in liquid nitrogen to enable further analysis. A total of four biological replicates were used.
Fractionation of 14C-Labeled Material
Tissue was fractionated exactly as described by Fernie et al. (2001), with the exception that hexoses were fractionated enzymatically rather than utilizing thin-layer chromatography as described by Carrari et al. (2006). Labeled Suc levels were determined after 4 h of incubation of 200 μL of total neutral fraction with 4 units mL−1 hexokinase in 50 mm Tris-HCl, pH 8.0, containing 13.3 mm MgCl2 and 3.0 mm ATP at 25°C. For labeled Glc and Fru levels, 200 μL of neutral fraction was incubated with 1 unit mL−1 Glc oxidase and 32 units mL−1 peroxidase in 0.1 m potassium phosphate buffer, pH 6, for a period of 6 h at 25°C. After the incubation time, all reactions were stopped by heating at 95°C for 5 min. The label was separated by ion-exchange chromatography as described by Fernie et al. (2001). The reliability of these fractionation techniques has been thoroughly documented previously (Runquist and Kruger, 1999; Fernie et al., 2001; Carrari et al., 2006). Fluxes were calculated as described by Fernie et al. (2001), following the assumptions detailed by Geigenberger et al. (1997, 2000).
Supplemental Data
The following materials are available in the online version of this article.
Supplemental Figure S1. Expression of SlNADP-ME genes during ripening.
Supplemental Figure S2. NADP-ME activity of tomato fruit at different pH ranges.
Acknowledgments
We acknowledge the excellent care of the plants by Helga Kulka (Max-Planck-Institut für Molekulare Pflanzenphysiologie) and Hanna Levanony (Weizmann Institute of Science).
Glossary
- PEPCK
phosphoenolpyruvate carboxykinase
- ME
malic enzyme
- OAA
oxaloacetate
- PEP
phosphoenolpyruvate
- TCA
tricarboxylic acid
- RNAi
RNA interference
- qRT
quantitative real-time
- DAP
days after pollination
- GC-MS
gas chromatography-mass spectrometry
- GABA
γ-aminobutyrate
- PEPC
phosphoenolpyruvate carboxylase
- MDH
malate dehydrogenase
- PK
pyruvate kinase
- AGPase
ADP-glucose pyrophosphorylase
- 3PGA
3-phosphoglycerate
- cDNA
complementary DNA
References
- Araújo WL, Nunes-Nesi A, Osorio S, Usadel B, Fuentes D, Nagy R, Balbo I, Lehmann M, Studart-Witkowski C, Tohge T, et al. (2011) Antisense inhibition of the iron-sulphur subunit of succinate dehydrogenase enhances photosynthesis and growth in tomato via an organic acid-mediated effect on stomatal aperture. Plant Cell 23: 600–627 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bahrami AR, Chen ZH, Walker RP, Leegood RC, Gray JE. (2001) Ripening-related occurrence of phosphoenolpyruvate carboxykinase in tomato fruit. Plant Mol Biol 47: 499–506 [DOI] [PubMed] [Google Scholar]
- Borsani J, Budde CO, Porrini L, Lauxmann MA, Lombardo VA, Murray R, Andreo CS, Drincovich MF, Lara MV. (2009) Carbon metabolism of peach fruit after harvest: changes in enzymes involved in organic acid and sugar level modifications. J Exp Bot 60: 1823–1837 [DOI] [PubMed] [Google Scholar]
- Bugos RC, Chiang VL, Zhang XH, Campbell ER, Podila GK, Campbell WH. (1995) RNA isolation from plant tissues recalcitrant to extraction in guanidine. Biotechniques 19: 734–737 [PubMed] [Google Scholar]
- Carrari F, Baxter C, Usadel B, Urbanczyk-Wochniak E, Zanor MI, Nunes-Nesi A, Nikiforova V, Centero D, Ratzka A, Pauly M, et al. (2006) Integrated analysis of metabolite and transcript levels reveals the metabolic shifts that underlie tomato fruit development and highlight regulatory aspects of metabolic network behavior. Plant Physiol 142: 1380–1396 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carrari F, Fernie AR. (2006) Metabolic regulation underlying tomato fruit development. J Exp Bot 57: 1883–1897 [DOI] [PubMed] [Google Scholar]
- Centeno DC, Osorio S, Nunes-Nesi A, Bertolo AL, Carneiro RT, Araújo WL, Steinhauser MC, Michalska J, Rohrmann J, Geigenberger P, et al. (2011) Malate plays a crucial role in starch metabolism, ripening, and soluble solid content of tomato fruit and affects postharvest softening. Plant Cell 23: 162–184 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chollet R, Vidal J, O’Leary MH. (1996) Phosphoenolpyruvate carboxylase: a ubiquitous, highly regulated enzyme in plants. Annu Rev Plant Physiol Plant Mol Biol 47: 273–298 [DOI] [PubMed] [Google Scholar]
- Detarsio E, Wheeler MC, Campos Bermúdez VA, Andreo CS, Drincovich MF. (2003) Maize C4 NADP-malic enzyme: expression in Escherichia coli and characterization of site-directed mutants at the putative nucleoside-binding sites. J Biol Chem 278: 13757–13764 [DOI] [PubMed] [Google Scholar]
- Drincovich MF, Casati P, Andreo CS. (2001) NADP-malic enzyme from plants: a ubiquitous enzyme involved in different metabolic pathways. FEBS Lett 490: 1–6 [DOI] [PubMed] [Google Scholar]
- Edwards GE, Andreo CS. (1992) NADP-malic enzyme from plants. Phytochemistry 31: 1845–1857 [DOI] [PubMed] [Google Scholar]
- Famiani F, Baldicchi A, Battistelli A, Moscatello S, Walker RP. (2009) Soluble sugar and organic acid contents and the occurrence and potential role of phosphoenolpyruvate carboxykinase (PEPCK) in gooseberry (Ribes grossularia L.). J Hortic Sci Biotechnol 84: 249–254 [Google Scholar]
- Famiani F, Cultrera NG, Battistelli A, Casulli V, Proietti P, Standardi A, Chen ZH, Leegood RC, Walker RP. (2005) Phosphoenolpyruvate carboxykinase and its potential role in the catabolism of organic acids in the flesh of soft fruit during ripening. J Exp Bot 56: 2959–2969 [DOI] [PubMed] [Google Scholar]
- Farineau J. (1977) Light versus dark carbon metabolism in cherry tomato fruits. II. Relationship between malate metabolism and photosynthetic activity. Plant Physiol 60: 877–880 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fernandez AI, Viron N, Alhagdow M, Karimi M, Jones M, Amsellem Z, Sicard A, Czerednik A, Angenent G, Grierson D, et al. (2009) Flexible tools for gene expression and silencing in tomato. Plant Physiol 151: 1729–1740 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fernie AR, Martinoia E. (2009) Malate: jack of all trades or master of a few? Phytochemistry 70: 828–832 [DOI] [PubMed] [Google Scholar]
- Fernie AR, Roscher A, Ratcliffe RG, Kruger NJ. (2001) Fructose 2,6-bisphosphate activates pyrophosphate:fructose-6-phosphate 1-phosphotransferase and increases triose phosphate to hexose phosphate cycling in heterotrophic cells. Planta 212: 250–263 [DOI] [PubMed] [Google Scholar]
- Franke KE, Adams DO. (1995) Cloning of a full-length cDNA for malic enzyme (EC 1.1.1.40) from grape berries. Plant Physiol 107: 1009–1010 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fraser P, Bickmore W. (2007) Nuclear organization of the genome and the potential for gene regulation. Nature 447: 413–417 [DOI] [PubMed] [Google Scholar]
- Fraser PD, Enfissi EM, Halket JM, Truesdale MR, Yu D, Gerrish C, Bramley PM. (2007) Manipulation of phytoene levels in tomato fruit: effects on isoprenoids, plastids, and intermediary metabolism. Plant Cell 19: 3194–3211 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Geigenberger P. (2011) Regulation of starch biosynthesis in response to a fluctuating environment. Plant Physiol 155: 1566–1577 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Geigenberger P, Fernie AR, Gibon Y, Christ M, Stitt M. (2000) Metabolic activity decreases as an adaptive response to low internal oxygen in growing potato tubers. Biol Chem 381: 723–740 [DOI] [PubMed] [Google Scholar]
- Geigenberger P, Muller-Rober B, Stitt M. (1999) Contribution of adenosine 5′-diphosphoglucose pyrophosphorylase to the control of starch synthesis is decreased by water stress in growing potato tubers. Planta 209: 338–345 [DOI] [PubMed] [Google Scholar]
- Geigenberger P, Regierer B, Nunes-Nesi A, Leisse A, Urbanczyk-Wochniak E, Springer F, van Dongen JT, Kossmann J, Fernie AR. (2005) Inhibition of de novo pyrimidine synthesis in growing potato tubers leads to a compensatory stimulation of the pyrimidine salvage pathway and a subsequent increase in biosynthetic performance. Plant Cell 17: 2077–2088 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Geigenberger P, Reimholz R, Geiger M, Merlo L, Canale V, Stitt M. (1997) Regulation of sucrose and starch metabolism in potato tubers in response to short-term water deficit. Planta 201: 502–518 [Google Scholar]
- Gerrard Wheeler M, Arias CL, Maurino VG, Andreo CS, Drincovich MF. (2009) Identification of domains involved in the allosteric regulation of cytosolic Arabidopsis thaliana NADP-malic enzyme. FEBS Lett 276: 5665–5677 [DOI] [PubMed] [Google Scholar]
- Gerrard Wheeler MC, Tronconi MA, Drincovich MF, Andreo CS, Flügge UI, Maurino VG. (2005) A comprehensive analysis of the NADP-malic enzyme gene family of Arabidopsis. Plant Physiol 139: 39–51 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Giovannoni JJ. (2004) Genetic regulation of fruit development and ripening. Plant Cell (Suppl) 16: S170–S180 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Halinska A, Frenkel C. (1991) Acetaldehyde stimulation of net gluconeogenic carbon movement from applied malic acid in tomato fruit pericarp tissue. Plant Physiol 95: 954–960 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jenner HL, Winning BM, Millar AH, Tomlinson KL, Leaver CJ, Hill SA. (2001) NAD malic enzyme and the control of carbohydrate metabolism in potato tubers. Plant Physiol 126: 1139–1149 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Karlova R, Rosin FM, Busscher-Lange J, Parapunova V, Do PT, Fernie AR, Fraser PD, Baxter C, Angenent GC, de Maagd RA. (2011) Transcriptome and metabolite profiling show that APETALA2a is a major regulator of tomato fruit ripening. Plant Cell 23: 923–941 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Klee HJ, Giovannoni JJ. (2011) Genetics and control of tomato fruit ripening and quality attributes. Annu Rev Genet 45: 41–59 [DOI] [PubMed] [Google Scholar]
- Knee M, Finger FL. (1992) NADP+ malic enzyme and organic acid levels in developing tomato fruits. J Am Soc Hortic Sci 117: 799–801 [Google Scholar]
- Kolbe A, Tiessen A, Schluepmann H, Paul M, Ulrich S, Geigenberger P. (2005) Trehalose 6-phosphate regulates starch synthesis via posttranslational redox activation of ADP-glucose pyrophosphorylase. Proc Natl Acad Sci USA 102: 11118–11123 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lara MV, Drincovich MF, Andreo CS. (2004) Induction of a Crassulacean acid-like metabolism in the C(4) succulent plant, Portulaca oleracea L: study of enzymes involved in carbon fixation and carbohydrate metabolism. Plant Cell Physiol 45: 618–626 [DOI] [PubMed] [Google Scholar]
- Lee M, Choi Y, Burla B, Kim YY, Jeon B, Maeshima M, Yoo JY, Martinoia E, Lee Y. (2008) The ABC transporter AtABCB14 is a malate importer and modulates stomatal response to CO2. Nat Cell Biol 10: 1217–1223 [DOI] [PubMed] [Google Scholar]
- Leegood RC, Walker RP. (2003) Regulation and roles of phosphoenolpyruvate carboxykinase in plants. Arch Biochem Biophys 414: 204–210 [DOI] [PubMed] [Google Scholar]
- Lombardo VA, Osorio S, Borsani J, Lauxmann MA, Bustamante CA, Budde CO, Andreo CS, Lara MV, Fernie AR, Drincovich MF. (2011) Metabolic profiling during peach fruit development and ripening reveals the metabolic networks that underpin each developmental stage. Plant Physiol 157: 1696–1710 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Luedemann A, Strassburg K, Erban A, Kopka J. (2008) TagFinder for the quantitative analysis of gas chromatography-mass spectrometry (GC-MS)-based metabolite profiling experiments. Bioinformatics 24: 732–737 [DOI] [PubMed] [Google Scholar]
- Matas AJ, Gapper NE, Chung MY, Giovannoni JJ, Rose JK. (2009) Biology and genetic engineering of fruit maturation for enhanced quality and shelf-life. Curr Opin Biotechnol 20: 197–203 [DOI] [PubMed] [Google Scholar]
- Meyer S, De Angeli A, Fernie AR, Martinoia E. (2010) Intra- and extra-cellular excretion of carboxylates. Trends Plant Sci 15: 40–47 [DOI] [PubMed] [Google Scholar]
- Michalska J, Zauber H, Buchanan BB, Cejudo FJ, Geigenberger P. (2009) NTRC links built-in thioredoxin to light and sucrose in regulating starch synthesis in chloroplasts and amyloplasts. Proc Natl Acad Sci USA 106: 9908–9913 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moco S, Capanoglu E, Tikunov Y, Bino RJ, Boyacioglu D, Hall RD, Vervoort J, De Vos RC. (2007) Tissue specialization at the metabolite level is perceived during the development of tomato fruit. J Exp Bot 58: 4131–4146 [DOI] [PubMed] [Google Scholar]
- Moing A, Aharoni A, Biais B, Rogachev I, Meir S, Brodsky L, Allwood JW, Erban A, Dunn WB, Kay L, et al. (2011) Extensive metabolic cross-talk in melon fruit revealed by spatial and developmental combinatorial metabolomics. New Phytol 190: 683–696 [DOI] [PubMed] [Google Scholar]
- Müller-Röber BT, Kossmann J, Hannah LC, Willmitzer L, Sonnewald U. (1990) One of two different ADP-glucose pyrophosphorylase genes from potato responds strongly to elevated levels of sucrose. Mol Gen Genet 224: 136–146 [DOI] [PubMed] [Google Scholar]
- Nashilevitz S, Melamed-Bessudo C, Izkovich Y, Rogachev I, Osorio S, Itkin M, Adato A, Pankratov I, Hirschberg J, Fernie AR, et al. (2010) An orange ripening mutant links plastid NAD(P)H dehydrogenase complex activity to central and specialized metabolism during tomato fruit maturation. Plant Cell 22: 1977–1997 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nielsen TH, Krapp A, Roper-Schwarz U, Stitt M. (1998) The sugar-mediated regulation of genes encoding the small subunit of Rubisco and the regulatory subunit of ADP glucose pyrophosphorylase is modified by phosphate and nitrogen. Plant Cell Environ 21: 443–454 [Google Scholar]
- Nunes-Nesi A, Araújo WL, Fernie AR. (2011) Targeting mitochondrial metabolism and machinery as a means to enhance photosynthesis. Plant Physiol 155: 101–107 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nunes-Nesi A, Carrari F, Gibon Y, Sulpice R, Lytovchenko A, Fisahn J, Graham J, Ratcliffe RG, Sweetlove LJ, Fernie AR. (2007) Deficiency of mitochondrial fumarase activity in tomato plants impairs photosynthesis via an effect on stomatal function. Plant J 50: 1093–1106 [DOI] [PubMed] [Google Scholar]
- Nunes-Nesi A, Carrari F, Lytovchenko A, Smith AM, Loureiro ME, Ratcliffe RG, Sweetlove LJ, Fernie AR. (2005) Enhanced photosynthetic performance and growth as a consequence of decreasing mitochondrial malate dehydrogenase activity in transgenic tomato plants. Plant Physiol 137: 611–622 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oliver SN, Tiessen A, Fernie AR, Geigenberger P. (2008) Decreased expression of plastidial adenylate kinase in potato tubers results in an enhanced rate of respiration and a stimulation of starch synthesis that is attributable to post-translational redox-activation of ADP-glucose pyrophosphorylase. J Exp Bot 59: 315–325 [DOI] [PubMed] [Google Scholar]
- Osorio S, Alba R, Damasceno CM, Lopez-Casado G, Lohse M, Zanor MI, Tohge T, Usadel B, Rose JK, Fei Z, et al. (2011) Systems biology of tomato fruit development: combined transcript, protein, and metabolite analysis of tomato transcription factor (nor, rin) and ethylene receptor (Nr) mutants reveals novel regulatory interactions. Plant Physiol 157: 405–425 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Osorio S, Alba R, Nikoloski Z, Kochevenko A, Fernie AR, Giovannoni JJ. (2012) Integrative comparative analyses of transcript and metabolite profiles from pepper and tomato ripening and development stages uncovers species-specific patterns of network regulatory behavior. Plant Physiol 159: 1713–1729 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Penfield S, Clements S, Bailey KJ, Gilday AD, Leegood RC, Gray JE, Graham IA. (2012) Expression and manipulation of phosphoenolpyruvate carboxykinase 1 identifies a role for malate metabolism in stomatal closure. Plant J 69: 679–688 [DOI] [PubMed] [Google Scholar]
- Rademacher T, Häusler RE, Hirsch HJ, Zhang L, Lipka V, Weier D, Kreuzaler F, Peterhänsel C. (2002) An engineered phosphoenolpyruvate carboxylase redirects carbon and nitrogen flow in transgenic potato plants. Plant J 32: 25–39 [DOI] [PubMed] [Google Scholar]
- Regierer B, Fernie AR, Springer F, Perez-Melis A, Leisse A, Koehl K, Willmitzer L, Geigenberger P, Kossmann J. (2002) Starch content and yield increase as a result of altering adenylate pools in transgenic plants. Nat Biotechnol 20: 1256–1260 [DOI] [PubMed] [Google Scholar]
- Roessner-Tunali U, Liu J, Leisse A, Balbo I, Perez-Melis A, Willmitzer L, Fernie AR. (2004) Kinetics of labelling of organic and amino acids in potato tubers by gas chromatography-mass spectrometry following incubation in (13)C labelled isotopes. Plant J 39: 668–679 [DOI] [PubMed] [Google Scholar]
- Rose JK, Bennett AB. (1999) Cooperative disassembly of the cellulose-xyloglucan network of plant cell walls: parallels between cell expansion and fruit ripening. Trends Plant Sci 4: 176–183 [DOI] [PubMed] [Google Scholar]
- Ruffner HP. (1982) Metabolism of tartaric and malic acids in Vitis: a review. Part A. Vitis 21: 346–358 [Google Scholar]
- Ruffner HP, Possner D, Brem S, Rast DM. (1984) The physiological role of malic enzyme in grape ripening. Planta 160: 444–448 [DOI] [PubMed] [Google Scholar]
- Runquist M, Kruger NJ. (1999) Control of gluconeogenesis by isocitrate lyase in endosperm of germinating castor bean seedlings. Plant J 19: 423–431 [DOI] [PubMed] [Google Scholar]
- Schauer N, Semel Y, Roessner U, Gur A, Balbo I, Carrari F, Pleban T, Perez-Melis A, Bruedigam C, Kopka J, et al. (2006) Comprehensive metabolic profiling and phenotyping of interspecific introgression lines for tomato improvement. Nat Biotechnol 24: 447–454 [DOI] [PubMed] [Google Scholar]
- Scheible R, Stitt M. (1988) Comparison of NADP-malate dehydrogenase activation QA reduction and O2 evolution in spinach leaves. Plant Physiol Biochem 26: 473–481 [Google Scholar]
- Schippers JH, Nunes-Nesi A, Apetrei R, Hille J, Fernie AR, Dijkwel PP. (2008) The Arabidopsis onset of leaf death5 mutation of quinolinate synthase affects nicotinamide adenine dinucleotide biosynthesis and causes early ageing. Plant Cell 20: 2909–2925 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Seymour G, Poole M, Manning K, King GJ. (2008) Genetics and epigenetics of fruit development and ripening. Curr Opin Plant Biol 11: 58–63 [DOI] [PubMed] [Google Scholar]
- Sowokinos JR. (1981) Pyrophosphorylases in Solanum tuberosum. II. Catalytic properties and regulation of ADP-glucose and UDP-glucose pyrophosphorylase activities in potatoes. Plant Physiol 68: 924–929 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sowokinos JR, Preiss J. (1982) Pyrophosphorylases in Solanum tuberosum. III. Purification, physical, and catalytic properties of ADPglucose pyrophosphorylase in potatoes. Plant Physiol 69: 1459–1466 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stitt M, ap Rees T. (1979) Capacities of pea chloroplasts to catalyse the oxidative pentose phosphate pathway and glycolysis. Phytochemistry 18: 1905–1911 [Google Scholar]
- Sweetlove LJ, Beard KFM, Nunes-Nesi A, Fernie AR, Ratcliffe RG. (2010) Not just a circle: flux modes in the plant TCA cycle. Trends Plant Sci 15: 462–470 [DOI] [PubMed] [Google Scholar]
- Sweetlove LJ, Muller-Rober B, Willmitzer L, Hill SA. (1999) The contribution of adenosine 5′-diphosphoglucose pyrophosphorylase to the control of starch synthesis in potato tubers. Planta 209: 330–337 [DOI] [PubMed] [Google Scholar]
- Sweetman C, Deluc LG, Cramer GR, Ford CM, Soole KL. (2009) Regulation of malate metabolism in grape berry and other developing fruits. Phytochemistry 70: 1329–1344 [DOI] [PubMed] [Google Scholar]
- Szecowka M, Osorio S, Obata T, Araújo WL, Rohrmann J, Nunes-Nesi A, Fernie AR. (2012) Decreasing the mitochondrial synthesis of malate in potato tubers does not affect plastidial starch synthesis, suggesting that the physiological regulation of ADPglucose pyrophosphorylase is context dependent. Plant Physiol 160: 2227–2238 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tauberger E, Fernie AR, Emmermann M, Renz A, Kossmann J, Willmitzer L, Trethewey RN. (2000) Antisense inhibition of plastidial phosphoglucomutase provides compelling evidence that potato tuber amyloplasts import carbon from the cytosol in the form of glucose-6-phosphate. Plant J 23: 43–53 [DOI] [PubMed] [Google Scholar]
- Tieman D, Taylor M, Schauer N, Fernie AR, Hanson AD, Klee HJ. (2006) Tomato aromatic amino acid decarboxylases participate in synthesis of the flavor volatiles 2-phenylethanol and 2-phenylacetaldehyde. Proc Natl Acad Sci USA 103: 8287–8292 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tiessen A, Hendriks JH, Stitt M, Branscheid A, Gibon Y, Farré EM, Geigenberger P. (2002) Starch synthesis in potato tubers is regulated by post-translational redox modification of ADP-glucose pyrophosphorylase: a novel regulatory mechanism linking starch synthesis to the sucrose supply. Plant Cell 14: 2191–2213 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tiessen A, Prescha K, Branscheid A, Palacios N, McKibbin R, Halford NG, Geigenberger P. (2003) Evidence that SNF1-related kinase and hexokinase are involved in separate sugar-signalling pathways modulating post-translational redox activation of ADP-glucose pyrophosphorylase in potato tubers. Plant J 35: 490–500 [DOI] [PubMed] [Google Scholar]
- Timm S, Nunes-Nesi A, Pärnik T, Morgenthal K, Wienkoop S, Keerberg O, Weckwerth W, Kleczkowski LA, Fernie AR, Bauwe H. (2008) A cytosolic pathway for the conversion of hydroxypyruvate to glycerate during photorespiration in Arabidopsis. Plant Cell 20: 2848–2859 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tomato Genome Consortium (2012) The tomato genome sequence provides insights into fleshy fruit evolution. Nature 485: 635–641 [DOI] [PMC free article] [PubMed] [Google Scholar]
- van der Merwe MJ, Osorio S, Moritz T, Nunes-Nesi A, Fernie AR. (2009) Decreased mitochondrial activities of malate dehydrogenase and fumarase in tomato lead to altered root growth and architecture via diverse mechanisms. Plant Physiol 149: 653–669 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Voll LM, Zell MB, Engelsdorf T, Saur A, Wheeler MG, Drincovich MF, Weber AP, Maurino VG. (2012) Loss of cytosolic NADP-malic enzyme 2 in Arabidopsis thaliana is associated with enhanced susceptibility to Colletotrichum higginsianum. New Phytol 195: 189–202 [DOI] [PubMed] [Google Scholar]
- Walker RP, Chen ZH, Tecsi LI, Famiani F, Lea PJ, Leegood RC. (1999) Phosphoenolpyruvate carboxykinase plays a role in interactions of carbon and nitrogen metabolism during grape seed development. Planta 210: 9–18 [DOI] [PubMed] [Google Scholar]
- Zamboni A, Di Carli M, Guzzo F, Stocchero M, Zenoni S, Ferrarini A, Tononi P, Toffali K, Desiderio A, Lilley KS, et al. (2010) Identification of putative stage-specific grapevine berry biomarkers and omics data integration into networks. Plant Physiol 154: 1439–1459 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zanor MI, Osorio S, Nunes-Nesi A, Carrari F, Lohse M, Usadel B, Kühn C, Bleiss W, Giavalisco P, Willmitzer L, et al. (2009) RNA interference of LIN5 in tomato confirms its role in controlling Brix content, uncovers the influence of sugars on the levels of fruit hormones, and demonstrates the importance of sucrose cleavage for normal fruit development and fertility. Plant Physiol 150: 1204–1218 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang J, Wang X, Yu O, Tang J, Gu X, Wan X, Fang C. (2011) Metabolic profiling of strawberry (Fragaria × ananassa Duch.) during fruit development and maturation. J Exp Bot 62: 1103–1118 [DOI] [PubMed] [Google Scholar]