Abstract
Proteus mirabilis is a urinary tract pathogen and well known for its ability to move over agar surfaces by flagellum-dependent swarming motility. When P. mirabilis encounters a highly viscous environment, e.g., an agar surface, it differentiates from short rods with few flagella to elongated, highly flagellated cells that lack septa and contain multiple nucleoids. The bacteria detect a surface by monitoring the rotation of their flagellar motors. This process involves an enigmatic flagellar protein called FliL, the first gene in an operon (fliLMNOPQR) that encodes proteins of the flagellar rotor switch complex and flagellar export apparatus. We used a fliL knockout mutant to gain further insight into the function of FliL. Loss of FliL results in cells that cannot swarm (Swr−) but do swim (Swm+) and produces cells that look like wild-type swarmer cells, termed “pseudoswarmer cells,” that are elongated, contain multiple nucleoids, and lack septa. Unlike swarmer cells, pseudoswarmer cells are not hyperflagellated due to reduced expression of flaA (the gene encoding flagellin), despite an increased transcription of both flhD and fliA, two positive regulators of flagellar gene expression. We found that defects in fliL prevent viscosity-dependent sensing of a surface and viscosity-dependent induction of flaA transcription. Studies with fliL cells unexpectedly revealed that the fliL promoter, fliL coding region, and a portion of fliM DNA are needed to complement the Swr− phenotype. The data support a dual role for FliL as a critical link in sensing a surface and in the maintenance of flagellar rod integrity.
INTRODUCTION
Proteus mirabilis is a Gram-negative enteric gammaproteobacterium that is well known for its ability to move over nutrient agars by flagellum-dependent swarming motility and for its role as an opportunistic pathogen in urinary tract infections (UTIs) (1, 2). P. mirabilis is dimorphic and exhibits two cell morphotypes whose expression is dependent on the surrounding environment. In liquid environments, i.e., nutrient broths, P. mirabilis exists as a short vegetative swimmer cell that is 1.5 to 2.0 μm in length and possesses between 4 and 10 flagella per cell. When a swimmer cell encounters a viscous environment or a surface, e.g., nutrient agars, the swimmer cell differentiates into a swarmer cell 10 to 80 μm in length, with multiple evenly spaced nucleoids within its cytoplasm, that is propelled by hundreds or thousands of flagella per cell (1, 3, 4). Swarmer cells are uniquely adapted to life on surfaces (3, 5–8), and their ability to sense and respond to the host cell surface environment is thought to be critical in the regulation of several important virulence factors (1, 9–11).
Expression of the genes associated with swarmer cell differentiation is induced when a swimmer cell contacts a surface, and this effect is termed surface sensing (1, 4, 5). More than 50 genes, including flagellar and virulence genes, are involved in P. mirabilis swarmer cell differentiation (4, 12, 13). Expression of the swarmer cell regulon is controlled in large measure by FlhD4C2, which is well-known as the master regulator of flagellar biosynthesis, acting to control a three-tiered hierarchical cascade of transcription (14–18). In this hierarchy, the flhDC operon (the sole member of class I) encodes the flagellar master regulator that in turn controls the expression of class II genes. Class II gene products include proteins for the assembly of the flagellar hook-basal body (HBB) complex, the type III protein export apparatus, and the regulators FliA and FlgM, which in turn control expression of class III genes (19, 20). Class III genes encode proteins required for later steps in flagellar assembly, including motor and chemotaxis proteins, flagellar filament-associated proteins, and flagellin (FlaA), the major protein of the flagellum (18, 20, 21). When P. mirabilis differentiates from a swimmer to a swarmer cell, transcription at all levels of the flagellar regulon is upregulated to produce the hundreds of newly synthesized flagella on the swarmer cell (12, 13, 21). Inhibition of cell division and septation is also necessary to form polyploid swarmer cells and is controlled (in part) by FlhD4C2 (22).
Of the class II proteins, the function of FliL remains obscure. fliL is ubiquitous among the genomes of flagellated bacteria and is often the first gene in an operon (fliLMNOPQR) that encodes proteins of the cytoplasm-facing rotor switch complex known as the C-ring (FliM and FliN) and of the flagellar export apparatus (FliO, FliP, FliQ, and FliR) (23). FliL localizes to the inner membrane, in close proximity to the flagellar basal body (24–26). FliL is ca. 18 kDa, with a single transmembrane (TM) domain at its N terminus, suggesting that the N-terminal end of FliL resides in the cytoplasm while the rest of the protein is periplasmic.
Unlike other flagellar proteins, whose functions are conserved across motile bacterial species, fliL defects produce species-specific phenotypic changes. Several examples illustrate this. In the spirochete Borrelia burgdorferi, fliL mutants were observed to have defects in motility coupled with a reversal in the orientation of their periplasmic flagella (26). In alphaproteobacterial species, e.g., Caulobacter crescentus, Silicibacter sp. TM1040, and Rhodobacter sphaeroides, fliL null mutations result in a nonmotile phenotype (27–29), and fliL defects in C. crescentus result in the release of flagella (28). In enteric bacteria, e.g., Escherichia coli and Salmonella enterica serovar Typhimurium (Salmonella herein), the loss of FliL does not affect swimming, but fliL cells have dramatic defects in swarming (24, 30). Similar to results in C. crescentus, fliL defects in Salmonella also result in detachment of flagella (24, 28), with breakage occurring between the proximal and distal rod proteins (FlgF and FlgG, respectively). Breakage of flagella in fliL cells has been hypothesized to be due to an increase in the torsional force on the basal body that results from the high viscosity associated with the surface of nutrient agar, which suggests that FliL functions in maintaining the integrity and stability of the flagellar rod (24, 31).
Belas and Suvanasuthi (32) reported that conditions that caused a stalling of P. mirabilis flagellar motors, e.g., by increasing the external viscosity or by addition of anti-FlaA antiserum to broths, induced swarmer cell differentiation and virulence gene expression (32). This result led to the hypothesis that the surface signal and inhibition of flagellar rotation are linked; however, the mechanism for signal propagation from the stalled flagellar motor to the cytoplasm is currently unknown.
More recently, we found that defects in fliL affect swarmer cell differentiation and result in the inappropriate production of elongated cells reminiscent of swarmer cells under noninducing conditions (nutrient broth growth). We refer to these elongated fliL cells as pseudoswarmers (12, 32), and their presence suggests that defects in fliL cause errors in sensing the surface signal that are manifested in transcriptomic changes (12).
Here, we used a fliL knockout mutant to gain further insight into the function of FliL. We show that loss of FliL results in cells that cannot swarm on agar surfaces (Swr− phenotype) but swim in broth cultures (Swm+) and that pseudoswarmer cells present in liquid are elongated (Elo+), contain multiple, evenly spaced nucleoids, and lack septa, characteristics shared with swarmer cells. Unlike swarmer cells, pseudoswarmer cells are not hyperflagellated due to reduced expression of flaA (the gene encoding flagellin), despite an increased transcription of both flhD and fliA, two positive regulators of flagellar gene expression. Defects in fliL prevent viscosity-dependent sensing of a surface and viscosity-dependent induction of flaA transcription. Results from experiments designed to complement fliL and its Swr− phenotype reveal a requirement of the fliL promoter, fliL coding region, and a portion of fliM DNA. The data support a dual role for FliL as a critical link in surface sensing and in the maintenance of flagellar rod integrity.
MATERIALS AND METHODS
Bacterial strains, plasmids, and growth conditions.
Bacterial strains and plasmids used in this study are listed in Table 1. P. mirabilis BB2000 served as the wild type and the parent of all P. mirabilis mutants used in this study. E. coli XL1-Blue (Stratagene) was used for plasmid manipulations. Plasmids pACYC184 (37), pTrc99a (35), pBAD30, and pBAD33 (36) were used to express fliL in trans. E. coli and P. mirabilis were maintained in Luria-Bertani (LB) broth (10 g liter−1 Bacto tryptone, 5 g liter−1 yeast extract, 10 g liter−1 sodium chloride) (39) or LB agar (LB medium containing 15 g liter−1 Bacto agar) at 37°C. When isolated colonies of P. mirabilis were required, LSW− agar (10 g liter−1 Bacto tryptone, 5 g liter−1 yeast extract, 0.4 g liter−1 NaCl, 5 ml liter−1 glycerol, 20 g liter−1 Bacto agar) (33) was used to prevent swarming. Swimming motility was screened using Mot agar (10 g liter−1 Bacto tryptone, 5 g liter−1 NaCl, 3 g liter−1 Bacto agar). As required, media were supplemented with 100 μg ml−1 ampicillin, 40 μg ml−1 chloramphenicol, and 50 μg ml−1 kanamycin, respectively.
Table 1.
Bacterial strains and plasmids used in this study
| Strain or plasmid | Relevant characteristicsa | Source or reference |
|---|---|---|
| Strains | ||
| P. mirabilis | ||
| BB2000 | Wild type; spontaneous Rfr from PRM1 | 33 |
| YL1003 | fliL::kan-nt30; BB2000 with group II intron insertion in the nt 30 of fliL, Kmr | This study |
| YL1005 | PMI3101::kan-nt166, BB2000 with group II intron insertion in the nt 166 of PMI3101; Kmr | This study |
| E. coli | ||
| XL1-Blue | recA1 endA1 gyrA96 thi-1 hsdR17 supE44 relA1 lac [F′ proAB lacIqZΔM15Tn10 (Tcr)] | Stratagene |
| Plasmids | ||
| pACD4K-C | Intron expression vector; p15A ori, Cmr | Sigma-Aldrich |
| pYL25 | pACD4K-C containing retargeting intron specific to nt 30 of P. mirabilis fliL gene; Cmr | This study |
| pYL31 | pACD4K-C containing retargeting intron specific to nt 166 of PMI3101; Cmr | This study |
| pAR1219 | T7 RNA polymerase expression vector; pBR322 ori, Apr | 34 |
| pTrc99a | Ptrc expression vector; pBR322 ori, Apr | 35 |
| pBAD30 | PBAD expression vector; p15A ori, Apr | 36 |
| pBAD33 | PBAD expression vector; p15A ori, Cmr | 36 |
| pACYC184 | Cloning vector; p15A ori, Cmr, Tcr | 37 |
| pYL30 | pACYC184 containing a 699-bp DNA fragment containing PfliL (200-bp upstream to the start codon), the codon region and 16 bp downstream fliL; Cmr | This study |
| pYL35 | pTrc99a containing fliL coding region, Apr | This study |
| pYL36 | pBAD30 containing the Shine-Dalgarno sequence and the codon region of fliL from pYL30, Apr | This study |
| pYL37 | pBAD33 containing the Shine-Dalgarno sequence and the codon region of fliL from pYL30, Cmr | This study |
| pYL47 | pACYC184 containing PfliL (487 bp upstream to the start codon of fliL) and codon region of the complete fliLMNOPQR operon, Cmr | This study |
| pYL60 | pYL47 with deletion of HindIII fragment containing the fliQR codon region and 14 bp in the 3′ end of the flip codon region, Cmr | This study |
| pYL61 | pYL47 with deletion of the fliPQR codon region generated by PCR using pACYC184-HindIII_r and fliO-HindIII_f primers, Cmr | This study |
| pYL62 | pYL47 with deletion of the fliOPQR codon region generated by PCR using pACYC184-HindIII_r and fliN-HindIII_r primers; Cmr | This study |
| pYL63 | pYL47 with deletion of the fliNOPQR codon region generated by PCR using pACYC184-HindIII_r and fliM-HindIII_r primers, Cmr | This study |
| pYL64 | pYL47 with deletion of the fliPQR codon region generated by PCR using pACYC184-HindIII_r and fliL-HindIII_r primers, Cmr | This study |
| pYL65 | pTrc99a containing the codon region of fliLM, Apr | This study |
| pYL66 | pBAD30 containing the Shine-Dalgarno sequence and the codon region of fliLM; Apr | This study |
| pYL67 | pBAD33 containing the Shine-Dalgarno sequence and the codon region of fliLM, Cmr | This study |
| pGP704 | Suicide vector; R6K ori, Apr | 38 |
| pYL42 | pGP704 containing 1.5kb of PMI3099'-3101 immediately followed by fliLp::fliL from pYL30 | This study |
| pYL70 | pACYC184 containing PflaA::lacZ fusion, Cmr | This study |
Apr, Cmr, Kmr, Rfr, and Tcr indicate ampicillin, chloramphenicol, kanamycin, rifampin, and tetracycline resistance, respectively.
Construction of null mutations in BB2000.
fliL null mutations in BB2000 were constructed by group II intron insertion (40) using a TargeTron gene knockout system (Sigma-Aldrich) following the manufacturer's instructions. Plasmid pACD4K-C was used to construct the reprogrammed intron containing a kanamycin-resistant cassette. The intron was reprogrammed to insert between nucleotides (nt) 30 and 31 in fliL, producing plasmid pYL25. pYL25 and the intron expressing helper plasmid pAR1219 (34) were sequentially transformed into BB2000. Intron expression was induced by addition of isopropyl β-d-1-thiogalactopyranoside (IPTG), and isolates with the correct insertion of the intron were selected by kanamycin resistance. Plasmids pYL25 and pAR1219 were cured by passage of cells to nonselective medium several times followed by electroporation (41) and then by screening for loss of ampicillin and chloramphenicol resistance. The resulting null mutant was called YL1003 (fliL::kan-nt30) (where nt30 indicates the location of the intron insertion).
PMI3101, encoding the sole GGDEF-containing protein in the genome of P. mirabilis, was similarly knocked out at nt 166/167 using the same method.
The efficacy of the group II intron insertions was determined by PCR amplification and nucleotide sequencing. Primers used to reprogram the group II intron are listed in Table 2.
Table 2.
Oligonucleotide primers used in this study
| Primer | Sequence (5′→3′)a | Construct(s) or use |
|---|---|---|
| fliL30aEBS1d | CAGATTGTACAAATGTGGTGATAACAGATAAGTCGCTATAGCTAACTTACCTTTCTTTGT | YL1003 |
| fliL30aEBS2 | TGAACGCAAGTTTCTAATTTCGATTTAATTTCGATAGAGGAAAGTGTCT | YL1003 |
| fliL30aIBS | AAAAAAGCTTATAATTATCCTTAAATTACGCTATAGTGCGCCCAGATAGGGTG | YL1003 |
| 3101-166aEBS1d | CAGATTGTACAAATGTGGTGATAACAGATAAGTCATCTTTTATAACTTACCTTTCTTTGT | YL1005 |
| 3101-166aEBS2 | TGAACGCAAGTTTCTAATTTCGATTTAACTTCGATAGAGGAAAGTGTCT | YL1005 |
| 3101-166aIBS | AAAAAAGCTTATAATTATCCTTAAGTTACATCTTTGTGCGCCCAGATAGGGTG | YL1005 |
| fliLpF | GGATCCGTGGTGTCGATATTT | Sequencing |
| fliLR | AGCGTAACGTGATCCCTATG | Sequencing |
| fliLp-BamHI_f | CTTGGATCCGTGGTGTCGATATTTTTG | pYL30 |
| fliL-HindIII_r | CGGAAGCTTATCGCTCATAATGATTATC | pYL30, pYL35, pYL64 |
| fliLup-BglI_f2 | AATGCCTCCTCGGCTCAATTACATCTTGCCTCTC | pYL47 |
| fliR-SalI_r | CACGTCGACTCAAGAAACTAATTGCTGCAA | pYL47 |
| pACYC184-HindIII_r | CTACCGCATTAAAGCTTATCGATG | pYL61, pYL62, pYL63, pYL64 |
| fliO-HindIII_r | ACTAAGCTTTTATGAGTTGCCTTTATCTCGC | pYL61 |
| fliN-HindIII_r | ATTAAGCTTTCAACGACTCAGACGGCG | pYL62 |
| fliM-HindIII_r | ATTAAGCTTTCACTCATTGGTTTGTTCCTC | pYL63 |
| fliL-NcoI_f | ATCCCATGGCTAATTACAGCAATGAACGTAA | pYL35 |
| SacI-SD-fliL_f | CAAGAGCTCCACTACAAAACAGGAATTT | pYL36, pYL37 |
| fliL-KpnI_r | CTTGGTACCTTATCGCTCATAATGATTATC | pYL36, pYL37 |
| 3101CO-SacI_f | CGGGAGCTCGGGTGATTATGCTGAAGATAATATG | pYL42 |
| 3101CO-SmaI_r | CCGCCCGGGAAATATAGTGTTATTTAATACTCTG | pYL42 |
| fliLp-SmaI_f | TTTCCCGGGGTGGTGTCGATATTTTTGCC | pYL42 |
| fliL-EcoRI_r | CCGGAATTCTTATCGCTCATAATGATTATC | pYL42 |
| flaApF | GGATCCACTGGTCTCCTTTCT | pYL70 |
| flaApEcoRI_r | AACGAATTCAGCTAGATTCCTTTAAAA | pYL70 |
| lacZ-HindIII_r | TGATAAGCTTTTATTTTTGACACCAGACCAACTGG | pYL70 |
| fliL_r | GAGCATCTATCTCTGCCTGTGAAAGG-3' | qRT-PCR |
| flhDF | AAGGCTTCCGCAATGTTTAGAC | qRT-PCR |
| flhDR | GTTGCAAATCATCCACTCTGGA | qRT-PCR |
| fliAF | CCTGCGAGTGTTGAATTGGA | qRT-PCR |
| fliAR | GGATTGTGTCACTTCTCTTGC | qRT-PCR |
| fliMF | GTCAATTCCGTATGGGGCTG | qRT-PCR |
| fliMR | AACCAGATTCGGCTCAAAGG | qRT-PCR |
| fliOF | CAACCTTTACCCGTTGGACA | qRT-PCR |
| fliOR | CGACAACACGCTCTTTGGTA | qRT-PCR |
| flaAF | CAACTGAAGGTGCATTGAAC | qRT-PCR |
| flaAR | TGATTTCTCACCGCTCAGTA | qRT-PCR |
Underlining indicates restriction sites for cloning.
Complementation of fliL defects.
Plasmids expressing fliL under the control of its native promoter (PfliL) or inducible promoters (Ptrc and PBAD) were constructed as follows. PfliL::fliL, including 200 bp upstream of the fliL start codon and going to the stop codon of fliL, was generated by PCR and ligated into the BamHI and HindIII sites in pACYC184. The resulting plasmid was called pYL30. For construction of Ptrc::fliL and PBAD::fliL expression plasmids, the same fliL codon region was generated by PCR and cloned into the NcoI and HindIII sites downstream of the trc promoter in pTrc99a or into the SacI and KpnI sites downstream of the araBAD promoter in pBAD36 or pBAD37, respectively. A plasmid containing the complete fliL operon was constructed by PCR amplification of a DNA fragment from 499 bp upstream of the start codon of fliL through the stop codon of fliR, followed by ligation into the BglI and SalI sites in pACYC184, resulting in plasmid pYL47.
Truncations of the fliL operon were made from pYL47 to remove fliM-fliR genes one after another, starting from the 3′ end. Removal of fliQ and fliR genes in pYL47 was done by HindIII digestion, and the new plasmid was called pYL60 (see Fig. 6C). Removal of fliP-fliR, fliO-fliR, fliN-fliR, and fliM-fliR in pYL47 was achieved by reverse PCR with Phusion High-Fidelity DNA Polymerase (NEB), using one primer (pACYC184-HindIII_r) downstream of the fliR gene and the other (fliO-HindIII_r, fliN-HindIII_r, fliM-HindIII_r, and fliL-HindIII_r) on the 3′ end of the fliO, fliN, fliM, and fliL genes, respectively. The PCR-generated DNA fragments were ligated following HindIII digestion to create pYL61, pYL62, pYL63, and pYL64 with deletions of fliP-fliR, fliO-fliR, fliN-fliR, and fliM-fliR, respectively (see Fig. 6C). Primers used for cloning are listed in Table 2. Each deletion in the fliL operon was confirmed by sequencing, after which the respective plasmids were separately transformed into YL1003, and the resulting swimming and swarming phenotypes were measured.
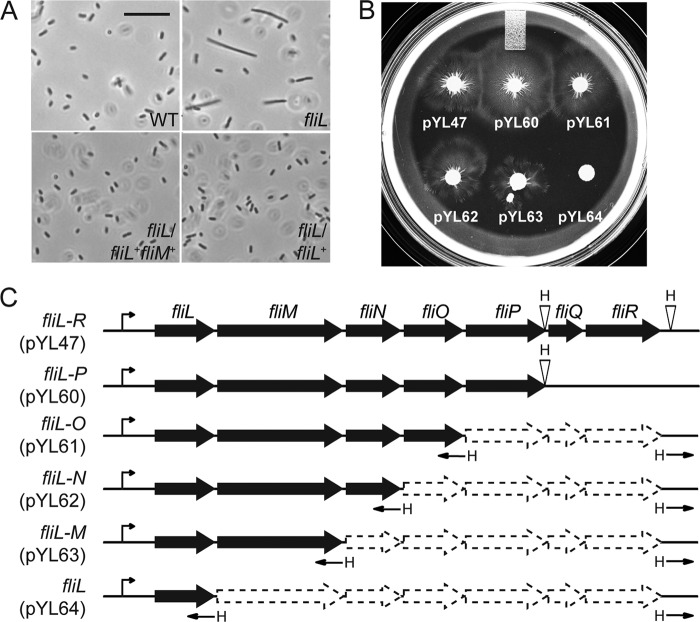
Fig 6.
Complementation of fliL cells by plasmids bearing deletions of the fliL operon. (A) Phase-contrast microscopy to detect pseudoswarmer cells. Scale bar, 20 μm. (B) Swarming motility on LB agar. (C) Sequential deletion of the fliL operon resulted in six plasmids, each harboring one less gene starting from the 3′ end of the operon. This yielded pYL47 (fliL to fliR), pYL60 (fliL to fliP), pYL61 (fliL to fliO), pYL62 (fliL to fliN), pYL63 (fliL to fliM), and pYL64 (fliL). All plasmids contain the native fliL promoter (PfliL; arrows in front of fliL). HindIII sites are indicated by H.
Ectopic single-copy expression of PfliL::fliL.
A method was developed to use Campbell-style single-crossover integration of a suicide plasmid containing PfliL::fliL into an ectopic site on the P. mirabilis genome. A 2,183-bp fragment containing 1.5 kb of the PMI3101 gene (see Results), plus upstream sequence (PMI3099′-PMI3100-PMI3101) followed by PfliL::fliL, was ligated into the SacI and EcoRI sites in the suicide vector pGP704 (38), resulting in pYL42. pYL42 was transferred to YL1003 via biparental mating between E. coli S17-1 λpir/pYL42 and YL1003, as previously described (33). The exconjugants were selected on LSW− agar containing kanamycin and ampicillin. Complementation of the Swr− defect in YL1003 was determined by measuring swarming motility on LB agar without antibiotic selection.
Swimming and swarming motility assay.
P. mirabilis cells were grown overnight in 2 ml of LB broth from a single colony. For bacterial cells prepared from broth, the overnight culture was inoculated into fresh LB broth at a 1:100 dilution and incubated at 37°C with shaking (200 rpm) for 2.75 h, when fliL strains produce the most pseudoswarmer cells in a population (12). For bacterial cells prepared from surfaces, 100 μl of an overnight culture was spread on the surface of LB agar in a 100-mm by 15-mm petri dish and incubated at 37°C for 4.5 h, when the wild-type strain BB2000 is actively swarming and produces the most swarmer cells in a population (data not shown). Cells were washed from the LB agar surface with 2 ml of 1× phosphate-buffered saline (PBS) (8 g liter−1 NaCl, 0.2 g liter−1 KCl, 1.44 g liter−1 Na2HPO4, 0.24 g liter−1 KH2PO4).
Swimming motility was measured as the outward migration of the cells following point inoculation of an overnight culture in the center of a petri dish containing semisolid Mot agar and LB agar (0.3% agar). Swarming motility was measured as bacterial migration over the surface of LB agar. Swarming plates were made by dispensing 20 ml of sterilized molten LB agar medium in each 100-mm by 15-mm petri dish placed on a bench in a single layer. The agar was allowed to gel at room temperature for ca. 20 h before use. Swarming was assessed following inoculation of a 5-μl sample of bacteria from overnight cultures grown in LB broth. Swimming and swarming assays were conducted in an environmental chamber at 37°C with a constant humidity of 46%.
RNA extraction, cDNA synthesis, and quantitative RT-PCR.
Total RNA was extracted and purified from bacterial samples using Ribopure (Ambion), following the manufacturer's instructions, with a 1-h DNase treatment. RNA was converted to cDNA using a high-capacity cDNA reverse transcription kit and RNase inhibitor (Applied Biosystems), with a thermocycling regime of 25°C for 10 min, 37°C for 2 h, and 85°C for 5 min. Quantitative reverse transcription-PCR (qRT-PCR) was performed using an ABI 7500 Fast real-time PCR system with the following thermocycling conditions: 95°C for 20 s, followed by 40 cycles of 95°C for 3 s and 60°C for 30 s. The change in target mRNA abundance was calculated using the 2−ΔΔCT formula (where CT is threshold cycle) (42), with rpoA (RNA polymerase subunit alpha) serving as a control.
PflaA::lacZ transcriptional fusion.
PCR was used to amplify the flaA promoter (386 bp 5′ of the start of flaA). The resulting DNA was digested with BamHI and EcoRI and ligated into the BamHI and EcoRI sites 5′ to a promoterless lacZ on pRS528 (43). The DNA fragment containing PflaA::lacZ transcriptional fusion was generated by PCR and subcloned into the BamHI and HindIII sites of pACYC184, resulting in pYL70. pYL70 was transformed into BB2000 or YL1003 by electroporation. Transcription of flaA, as measured by β-galactosidase activity, was determined using the Miller assay (44) as described previously (45), with cells harvested from the surface of LB agar after 4 h of incubation.
Protein sample preparation.
Protein was obtained from 5 ml of 2.75-h broth cultures or 2-ml cell suspensions prepared from agar surfaces (described above). In both cases the suspended bacteria were centrifuged at 6,000 rpm for 10 min to obtain supernatant and cell pellet fractions. The pellets were washed with 1× PBS and then resuspended in 1 ml of 1× PBS. As required, wide-bore pipette tips were used to prevent shearing of cell-attached flagella. Whole-cell homogenates of the pellets were prepared by six cycles of sonication for 10 s followed by a pause on ice for 30 s. The protein concentration of all fractions was measured using a bicinchoninic acid (BCA) protein assay (Pierce).
Western blotting.
The method described by Belas et al. (4) was used with Hybond-P polyvinylidene difluoride (PVDF) membranes (Amersham) and an ECL Plus system (GE Health Sciences). Flagellin (FlaA) was detected using rabbit polyvalent anti-FlaA antiserum (antiserum 497) (4) at a dilution of 1:5,000. FliM was detected with rabbit FliM antiserum (a gift from David Blair, University of Utah), using a 1:1,000 dilution of the stock. Horseradish peroxidase (HRP)-conjugated goat anti-rabbit IgG (Cell Signaling Technology) was used as the secondary antiserum at a titer of 1:50,000 for FlaA and 1:10,000 for FliM antibodies. ImageJ (National Institutes of Health) software was used to determine protein abundance on the developed Western blots.
Fluorescent staining and microscopy.
Bacterial cells were harvested from 2.75-h broth or 4.5-h plate cultures and resuspended in 500 μl of 1× PBS using wide-bore tips to prevent shearing of the flagella. Bacterial cells were fixed by the addition of 3% (vol/vol; final concentration) paraformaldehyde (Sigma-Aldrich) to the cell suspension and incubation at room temperature for 10 min with gentle rocking. Fixed bacteria were stored at 4°C until use, at which time 50 μl was placed on a poly-l-lysine-coated coverslip and left undisturbed at room temperature for 30 min. Nonadherent and loosely bound cells were removed by washing 10 times with 1× PBS. Cells bound to the coverslip were sequentially treated in 1-h steps with 50 μl of 2% bovine serum albumin (BSA) in 1× PBS, followed by rabbit anti-FlaA antiserum (4) (1:100 in 2% BSA) and Alexa Fluor 568-conjugated goat anti-rabbit IgG (Invitrogen) (1:100 in 2% BSA). The coverslips were kept in a humidified chamber in the dark during the entire process. Each reagent was removed by copious washing with 1× PBS. FM1-43 (15 μg ml−1; Invitrogen) was used to label membranes, while bacterial nucleoids were stained using 4′,6′-diamidino-2-phenylindole (DAPI) (2 μg ml−1; Invitrogen). Stained specimens were mounted in ProLong antifade reagent (Invitrogen), imaged using wide-field fluorescence microscopy with a Zeiss Axio Observer Z1 microscope and a Hamamatsu Orca-R2 CCD camera, and analyzed with Volocity, version 6.1.1, software (PerkinElmer).
Production of pseudoswarmers in 2.75-h broth cultures in a wet-mount sample of living cells was also examined using phase-contrast light microscopy with an Olympus BX60. Phase-contrast images were captured with a QImaging QCam charge-coupled-device (CCD) camera and analyzed with Volocity, version 6.1.1, software.
As previously described (12), a cell length of >7 μm was used as the cutoff to define a swarmer cell. At least five fields (>500 cells) of phase-contrast images and more than 100 fluorescently stained cells of each sample were examined. The mean and standard deviation of the cell length of each cell type were calculated using Prism, version 4.0, software (GraphPad).
RESULTS
Phenotype of fliL knockout strains.
Swimming and swarming motility were measured in a fliL knockout strain (YL1003; fliL::kan-nt30). Due to the nature of the mutation, YL1003 has the potential to encode no more than a highly truncated peptide, MSNYSNERKSVK, lacking the TM domain of FliL and making it highly unlikely that the FliL peptide is functional and equally unlikely that it localizes to the inner membrane or periplasm, unlike wild-type FliL.
Analysis of the swarming and swimming motility of YL1003 demonstrated that this fliL knockout strain was unable to swarm (Fig. 1A) but was able to swim in semisolid Mot agar (Fig. 1B); i.e., it has an Swr− Swm+ phenotype although the swimming motility is substantially reduced (migration distances are 33.5 mm for the wild type and 21.0 mm for fliL cells in Fig. 1B). These results demonstrate that FliL is essential for P. mirabilis swarming but not for its swimming. When grown in broth, YL1003 produces pseudoswarmer cells (Elo+) (Fig. 1C, right panel), with a mean length of 9.49 ± 2.33 μm. When grown on nutrient agar, consistent with our previous observation (32), the fliL mutant produces elongated cells with a mean cell length equal to or greater than that of the wild-type swarmer cells (data not shown). Maximal production of YL1003 pseudoswarmer cells occurs between 2.5 and 3 h, and these elongated cells comprise a small part (2.1% by number) of the population. Both observations are consistent with previous reports describing P. mirabilis fliL mutations (22). It should be remembered that pseudoswarmer cells are longer than wild-type swimmer cells; therefore, in terms of swimmer cell equivalents, fliL pseudoswarmers represent ca. 10 to 25% of the population of swimmer cells.
Fig 1.
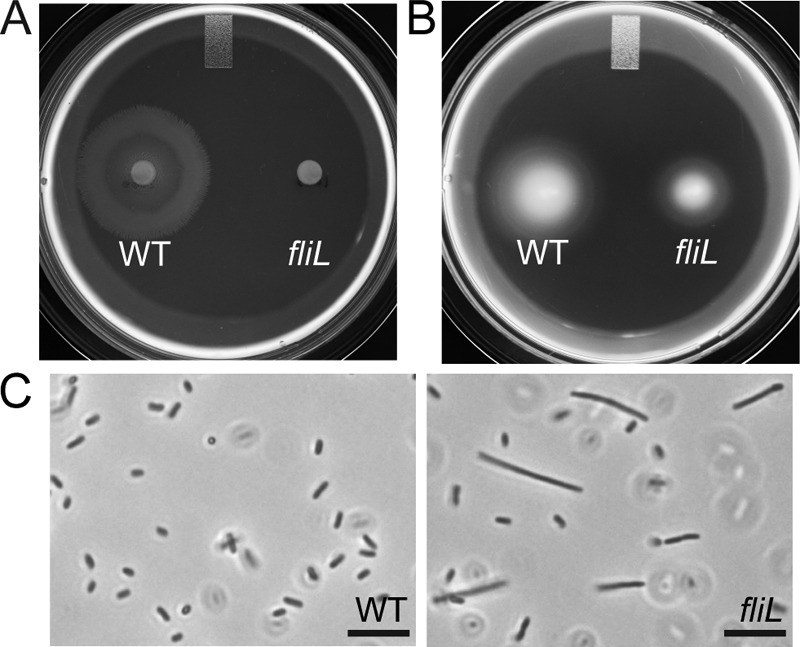
Phenotypic characterization of fliL strain YL1003. (A) Swarming motility on LB agar. (B). Swimming motility through semisolid Mot agar. (C) Phase-contrast micrographs of cells obtained from LB broth after 2.75 h of incubation. Scale bar, 10 μm. WT, wild type.
While pseudoswarmer cells are similar to wild-type swarmer cells in length, we do not know if they possess other hallmarks of swarmer cells, specifically, multiple nucleoids, lack of septa, and many flagella. To better characterize the pseudoswarmer cells, immunofluorescence microscopy was used to observe the cell membrane, nucleoids, and flagella, comparing these characteristics to wild-type swimmer and swarmer cells. As shown in Fig. 2, wild-type swimmer cells are short rods (average length, 2.26 ± 0.6 μm) with a single nucleoid per cell (Fig. 2A) and have distinct septa during division (data not shown). In contrast, wild-type swarmer cells (Fig. 2B) are elongated (average length, 9.79 ± 3.16 μm) cells, possess multiple, evenly spaced nucleoids along the length of the cytoplasm, lack septa, and are covered with many flagella. fliL pseudoswarmer cells (Fig. 2C) resemble wild-type swarmer cells in that they are elongated (average length, 9.49 ± 2.33 μm), have multiple nucleoids, and lack septa. However, fliL pseudoswarmer cells have far fewer flagella than wild-type swarmer cells.
Fig 2.
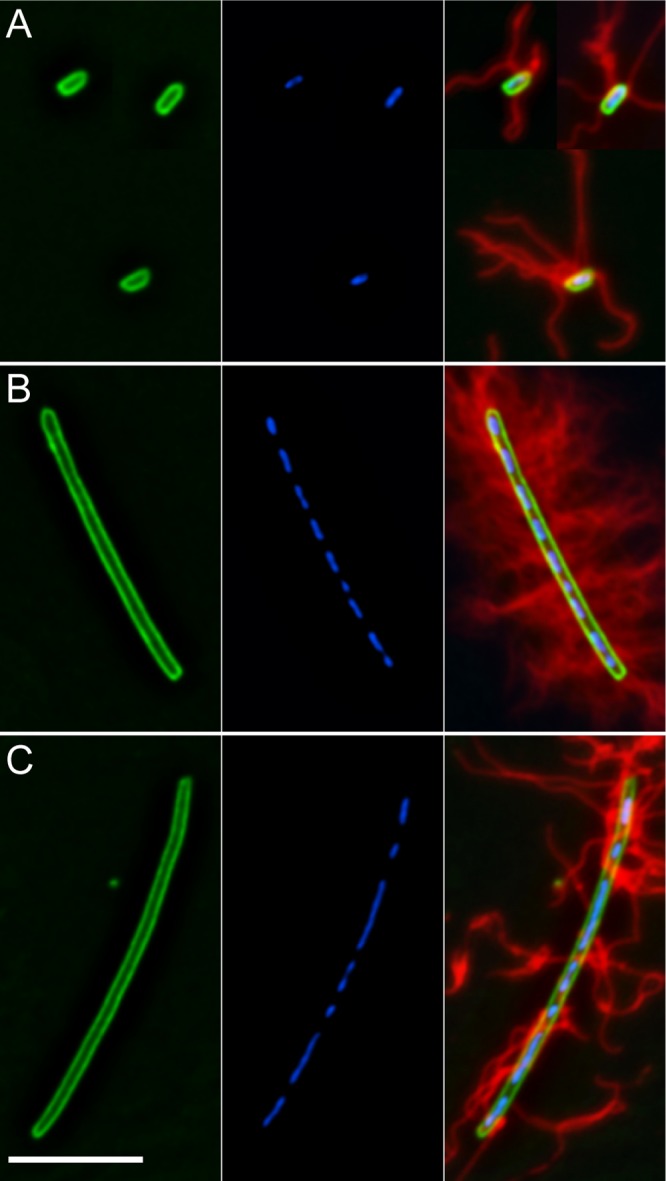
Comparison of fliL pseudoswarmer cells to wild-type swimmer and swarmer cells. (A) BB2000 swimmer cells. (B) BB2000 swarmer cell. (C) YL1003 pseudoswarmer cell. From left to right, membrane (FM1-43; green), nucleoids (DAPI; blue), and composite with flagellum immunostaining in red. Scale bar, 7 μm.
We found it difficult to accurately count the flagella on swarmer and pseudoswarmer cells due to multiple overlapping and clumping filaments. To overcome this problem, the relative amount of flagellin per microgram of total cellular protein was determined using immunoblotting against FlaA (flagellin). Measuring differences in band intensity, we observed that wild-type swarmer cells possess 5.25 times more FlaA than swimmer cells (Fig. 3). In contrast, YL1003 pseudoswarmer cells have the same amount of FlaA per total cell protein as swimmer cells. Growth on nutrient agar increased FlaA levels in YL1003 cells; however, the increase was much less than the 5.25-fold increase seen in wild-type cells. We conclude that fliL pseudoswarmer cells are similar but not identical to swarmer cells, with the lack of the hyperflagellated phenotype being the most notable difference.
Fig 3.
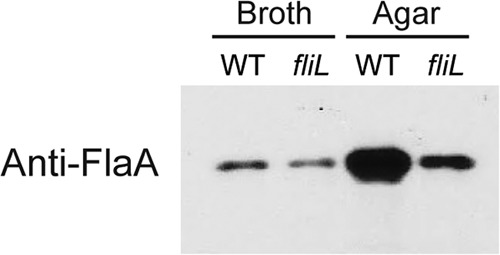
Immunoblot with polyvalent antibodies to flagellin (FlaA). BB2000 (WT) and YL1003 (fliL) cells were harvested from LB broth or agar.
The expression of representative flagellar genes from each class in the regulatory hierarchy was measured to determine if a reduction in transcription was responsible for the lack of flagella on fliL pseudoswarmer cells (Fig. 4). As expected, the flaA expression level decreased in fliL pseudoswarmer cells compared to the level in the wild type, especially when measured from cells grown on nutrient agar, where a 5-fold decrease in flaA levels in pseudoswarmer cells was noted. Much less of a change in flaA transcription was observed (0.9-fold less than in wild-type swimmer cells) in broth-grown fliL cells. Thus, a decrease in flaA transcription helps to explain why pseudoswarmer cells have few flagella.
Fig 4.
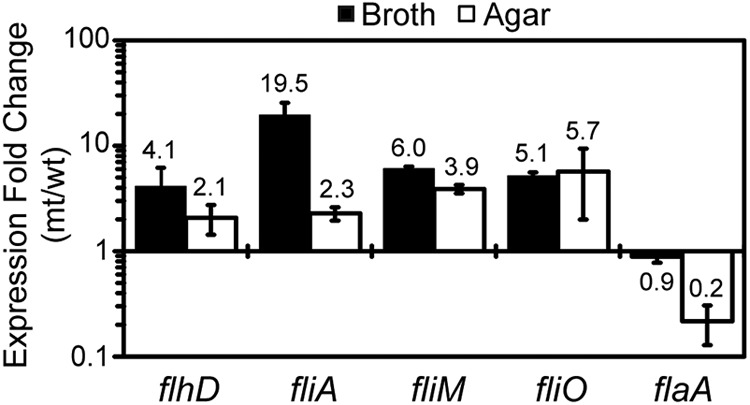
Expression of flagellar genes in fliL cells. Values are a comparison of expression of the target gene in fliL cells (mt) to the same gene in wild-type (wt) cells. A value of >1-fold change indicates that expression of the gene is greater in the fliL cells than in the wild-type cells, while a value of <1 means that expression decreases in fliL cells. Growth media are as indicated on the figure. Means and standard deviations for three independent measurements from three biological samples are shown.
While flaA expression decreased in fliL cells, transcription of other flagellar genes, including flhD, fliA, fliM, and fliO, increased significantly, compared to wild-type expression (Fig. 4). Although flaA transcription decreased, expression of flhD and fliA, encoding the flagellar master transcriptional activator and sigma factor, respectively, increased in YL1003 compared to wild-type levels (2.1- to 4.1-fold for flhD; 2.3- to 19.5-fold for fliA). Similar increases in expression of fliM and fliO (both class II genes) are also evident in pseudoswarmer cells taken from either broth or agar culture (Fig. 4). One interpretation of these results is that fliL defects somehow repress or inhibit the positive transcriptional activation of flaA by FlhD4C2 and FliA while not affecting transcription of other flagellar biosynthetic genes (see Discussion).
FliL functions to enhance motility in viscous environments.
The current model of FliL function suggests that it is required to maintain flagellar structural integrity when torsional stress increases due to external conditions that prevent rotation of the flagella. We tested the hypothesis that fliL defects would prevent P. mirabilis from responding to the surface signal (in this case, the viscosity of the medium). This was done by measuring swimming and swarming motility of wild-type and YL1003 cells in nutrient broths with different concentrations of agar. The results are shown in Fig. 5A and Table S1 in the supplemental material. As predicted, wild-type cells showed proficient swarming, which was maximal on 1.0% agar (44.7 ± 6.1 mm) and declined as the concentration of agar increased. Wild-type P. mirabilis did not swarm at agar concentrations of 0.6% or lower; instead, as the concentration decreased, the cells transitioned to swimming through the agar matrix. In contrast, YL1003 cells did not swarm at any agar concentration and were less able to swim through 0.3% agar (Fig. 5A; see also Table S1). These results show that P. mirabilis swarms in a viscosity-dependent manner and demonstrate a link between viscosity and FliL function.
Fig 5.
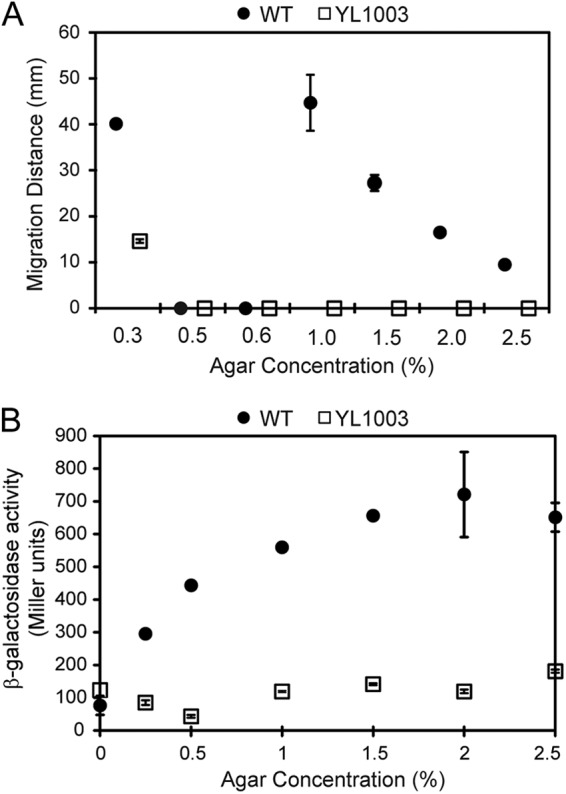
Motility and flaA expression in fliL cells (YL1003) and wild-type (WT) cells (BB2000) as a function of the viscosity of the medium. (A) Swimming (0.3% agar) or swarming (1.0 to 2.5% agar) as the distance (mm) the cells moved after 5 h. (B) flaA (flagellin) expression, measured as β-galactosidase activity from a PflaA::lacZ transcriptional fusion. Means ± standard deviations are shown.
In parallel experiments, flaA expression was measured in wild-type and YL1003 cells over the same concentrations of agar. While expression of flaA rose in wild-type cells as the concentration of agar in the medium increased, fliL cells showed no change in flaA transcription (Fig. 5B). Thus, loss of FliL function prevents increased flaA expression that would normally result from the surface-sensing events.
Complementation of the fliL knockout requires the fliL coding sequence, the fliL promoter region, and sequences in the fliM coding region.
The fliL phenotype has two characteristics: inability to swarm (Swr− phenotype) and pseudoswarmer formation (Elo+). We attempted to complement the fliL phenotype of YL1003 by introducing plasmids carrying the fliL coding region transcribed from either the fliL promoter (PfliL::fliL+), an IPTG-inducible promoter (Ptrc::fliL+), or an arabinose-inducible promoter (PBAD::fliL+). With all three plasmids, the Elo+ phenotype was complemented, returning YL1003 to swimmer cell morphology (Fig. 6A, compare the upper right with the bottom right panel). However, none of these plasmids was able to complement the Swr− phenotype, and the cells remained nonswarming. Our results show that the elongation and swarming phenotypes in the fliL mutant are separable, based on the complementation data. fliL transcripts were detectable from all three promoters (data not shown). Thus, the failure to complement the complete fliL phenotype of YL1003 is not due to a failure of fliL transcription in trans.
Motelab et al. suggested that Borrelia FliL functionality is sensitive to fliL copy number (26), which led us to test whether complementation of YL1003 could be achieved by expressing PfliL::fliL from an ectopic chromosomal site. In an unrelated study, we discovered that a knockout of PMI3101 (encoding the sole GGDEF protein in P. mirabilis) did not affect any measureable phenotype (data not shown), making this gene an excellent site for integration of PfliL::fliL. A single-crossover, Campbell-style integration was performed, and the exconjugants were screened for swarming (see Materials and Methods). Six Swr+ exconjugants were found, but none had PfliL::fliL integrated at PMI3101. Instead, in all six exconjugants, single-crossover recombination events occurred at the native fliL codon region, leading to a resurrection of a wild-type fliLMNOPQR operon and resulting in Swr+ bacteria. This was unexpected since statistical probability would be predicted to favor a crossover at the PMI3101 locus due to the much larger target (1.5 kb) it represents than the fliL target (<500 bp).
Although we do not fully understand why the recombination at fliL occurred, we inferred from this result that complementation of the Swr− phenotype in YL1003 required DNA beyond the fliL promoter and coding region. To test this hypothesis, a DNA fragment containing the fliL operon (fliLMNOPQR) transcribed from its native fliL promoter (PfliL) was placed on a plasmid, and the resulting plasmid pYL47 was moved into YL1003. In agreement with the hypothesis, pYL47 containing the fliL operon restored swarming of YL1003 (Fig. 6B).
Complementation of the Swr− phenotype in YL1003 is not due to loss of transcription of genes downstream from fliL. This was demonstrated in the results shown in Fig. 4 (expression of fliM and fliO). The abundance of FliM was also measured to verify that YL1003 was able to translate the proteins of the fliL operon, without FliL (Fig. 7). Indeed, the amount of FliM in YL1003 is identical to that of wild-type cells when they are grown in broth and decreased slightly when fliL cells are grown on agar medium (Fig. 7). These results suggested to us that complementation of the Swr− phenotype of YL1003 required fliL operon DNA beyond the fliL promoter and coding region.
Fig 7.
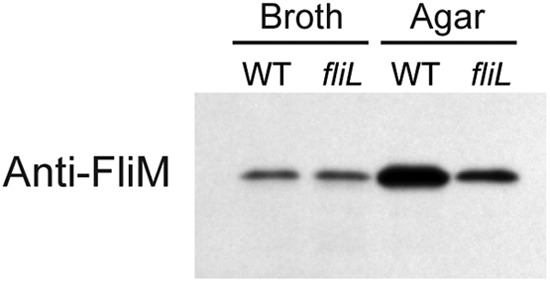
Immunoblotting with polyvalent antibodies to FliM. BB2000 (WT) and YL1003 (fliL) cells were harvested from LB broth or agar. The antiserum used was directed against E. coli FliM and cross-reacts with P. mirabilis FliM.
A set of plasmids bearing deletions in the fliL operon was made (Fig. 6C) and used to complement YL1003. Plasmids bearing fliL and fliM (pYL60, pYL61, pYL62, and pYL63 in Fig. 6B) returned YL1003 to an Swr+ phenotype, but loss of fliM DNA (pYL64) could not restore swarming to the fliL cells. As shown in Fig. 6B, the swarming diameters of YL1003 containing pYL47, pYL60, pYL61, pYL62, and pYL63 are, respectively, 20.7 mm, 29.7 mm, 22.5 mm, 20.7 mm, and 18.9 mm. As expected, both pYL63 (fliL+ filM+) and pYL64 (fliL+) complemented the Elo+ phenotype of YL1003 (Fig. 6A). Therefore, complementation of fliL requires three things: the fliL promoter region, the fliL coding region, and DNA within the fliM coding region.
Expressing fliLM in trans improves fliL motility.
The complementation results predict that PfliL::fliLM cells would regain viscosity-dependent swarming. We measured swarming over various agar concentrations (as previously performed) and found that fliL+ fliM+, but not fliL+ alone, restored swarming of YL1003 on 1.0 and 1.5% agar surfaces in a viscosity-dependent manner (see Table S1 in the supplemental material). YL1003 (fliL+ fliM+) is not proficient in swarming (i.e., longer lag and lower rate than wild-type cells), but its overall response to viscosity is the same (see Table S1). Both have maximal swarming in 1.0% agar, the maximal swarming rates for each are as fast as the swimming rate (in 0.3% agar), and their swarming rates both decrease in the same degree (0.4- to 0.5-fold) when the agar concentration increases from 1.0% to 1.5%. Prolonged incubation of the complemented YL1003 cells did not restore swarming.
The fliL+ fliM+ complementation results predict that YL1003/pYL63 (fliL+ fliM+) has regained wild-type flagellin synthesis, and this was measured by Western blotting of cells obtained after growth on nutrient agar (Fig. 8). As predicted, complementation with fliL+ fliM+ increased the overall production of flagellin, compared to fliL cells or YL1003/pYL64 (fliL+). There is a noticeable difference in attached (pellet fraction) and unattached (supernatant) flagella in these cells. In addition to increasing the overall abundance of FlaA, complementation of YL1003 with fliL+ fliM+ also increases the number of attached flagella per cell (compare fliL/fliL+ fliM+ with fliL/fliL+), which is reasonable since FliL is required to maintain the integrity of the flagellar rod. The surprise is that FliL function also requires fliM DNA.
Fig 8.
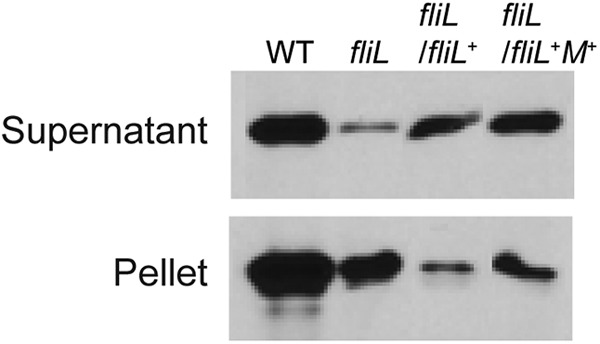
Immunoblot measuring the abundance of detached and attached flagella in fliL cells complemented with fliL+ or fliL+ fliM+. Wild-type and fliL cells were harvested after 4.5 h of incubation on LB agar. Rabbit polyvalent antiserum to FlaA was used, as described in the legend of Fig. 3.
DISCUSSION
Most, if not all, bacteria are able to live either as independent planktonic cells or as members of organized surface-anchored communities called biofilms (46). Biofilm formation is a developmental process that begins when a motile bacterium attaches to a surface and requires bacterial processes. One such process is flagellum-mediated motility, which facilitates attachment to and colonization of surfaces. However, while it is generally agreed that motility and biofilm development are mutually exclusive, the molecular mechanisms that underlie the swim-or-stick lifestyle switch remain virtually unknown for most bacterial species.
The long-term goal of our laboratory is to understand the molecular mechanism involved in how P. mirabilis senses a surface. The results from the current study further implicate FliL as a major player in the swim-or-stick mechanism of the swarming bacterium P. mirabilis. Our previous studies used a pseudoswarmer-producing strain containing a transposon insertion (fliL::Tn5) at nucleotide 440 in fliL (32). This strain has an Swm− Swr− phenotype due to polar effects of the transposon on downstream genes in the fliL operon, which in turn decreases expression of flagellar class III genes, e.g., flaA, and prevents flagellar biosynthesis (12). Furthermore, due to the transposon, this strain has a frameshift in the 3′ end of fliL such that the mutant retains most (>90%) of the wild-type fliL sequence. Both the polar effects on the fliL operon and the frameshift of fliL are disadvantageous for studying FliL function. To overcome these drawbacks, in the present study, we constructed a nonpolar fliL null mutation by site-specific TargeTron insertion at nucleotide 30, a site chosen to result in a highly truncated FliL, thereby eliminating the problems associated with the use of the fliL::Tn5 strain. As we along with others have shown, defects in fliL abolish swarming and reduce swimming motility. Decreased swimming of fliL::kan-nt30 cells is due to decreased flagella/flagellin production (compared to wild-type cells) when cells are grown in semisolid nutrient agar (Fig. 5B). This does not suggest that broth-grown FliL− cells lack a phenotype because their morphology is highly similar to a wild-type swarmer cell. These pseudoswarmer cells, as we call them, are elongated, nonseptated rods, with multiple nucleoids (Fig. 2). Using only these criteria, there is no difference between a wild-type swarmer cell and a fliL pseudoswarmer from broth culture. This leads us to conclude that loss of FliL inappropriately flips the swim-or-stick switch such that the bacterium responds as though it is on a surface, despite being in a noninducing environment, i.e., nutrient broth. However, it is not as simple as that.
Unlike swarmer cells, pseudoswarmer cells are not hyperflagellated, with roughly the same number of flagella per μm2 as wild-type swimmer cells (Fig. 2 and 3), indicating a major difference between fliL pseudoswarmer cells and wild-type swarmer cells. This difference occurs at the level of flaA expression that translates to a smaller amount of FlaA per cell. Oddly, while flaA (a flagellar class III gene) transcription decreases in pseudoswarmer cells, expression of flhD (class I) and fliA (class II) increases. Although the mechanism is not immediately apparent, an increase in flhD expression makes sense as it parallels upregulation of flhD transcription observed when swimmer cells differentiate into swarmer cells (21). The disparity in flhD and fliM expression levels on agar between the fliL::kan-nt30 and fliL::Tn5 mutants is certainly due to the aforementioned polar effect and fliL frameshift.
Why does fliA expression increase? Since FliA, the flagellar gene sigma factor, is required for flaA expression (47), an increase in FliA should result in an increase in FlaA, yet the opposite occurs. An increase in FlhD4C2 is expected to increase fliA expression, offering a partial explanation for these results. While a complete explanation for this reproducible event is not immediately evident, it is obvious that loss of FliL, a structural gene, results in an increase in the transcription of two flagellar regulatory genes. This could occur if loss of FliL somehow damaged the flagellar type III secretion system, which would prevent export of FlgM, the flagellar anti-sigma factor, inhibit the activity of FliA, and result in a decrease in class III gene expression. This hypothesis predicts that the expression of other class III flagellar genes is also low in pseudoswarmers, and experiments are under way to determine if this is true.
P. mirabilis senses a surface by monitoring the rotation of its swimmer cell flagella (32). As flagellum movement becomes inhibited by conditions that prevent rotation, e.g., viscosity of the medium, a signal is sent that activates transcription of a set of genes that produce swarmer cells (3, 12, 13, 32, 48, 49). Based on this knowledge, we predicted that fliL cells would be blind to changes in the viscosity of the medium and would be unable to sense (or respond to) the stimulus and unable to swarm (even on low concentrations of agar). Indeed, this is exactly what was seen. YL1003 did not swarm on any agar medium, while wild-type cells swarmed on concentrations of agar up to 2.5%, the highest we tested (Fig. 5A). Furthermore, both wild-type and fliL cells swam through semisolid agar matrices, albeit fliL cells migrated through the agar at less than half the rate of wild-type cells. As predicted, expression of flaA increased in wild-type cells roughly in step with increased agar concentration (Fig. 5B) and reached a plateau around 2.0% agar, while transcription of flaA in YL1003 remained at a basal level throughout the agar concentration series. The pseudoswarmer phenotype, but not the swarming defect, of fliL cells is rescued by fliL alone (Fig. 6A); therefore, fliL cells reveal that cell elongation and the swarming phenotypes are separable characteristics. The fliL mutant constitutively produced elongated pseudoswarmer cells under noninducing conditions, demonstrating that the loss of FliL affects not only swarming, due to the defect in flaA expression, but also the ability of the cell to detect surfaces, as the mutant “thinks” it is on a surface when it is not. Thus, defects in fliL affect the ability of the cells to sense surfaces. We believe that this hints at two functions for FliL: maintenance of flagellum integrity and involvement in surface signaling.
Figure 5A also reveals another interesting result: wild-type P. mirabilis is incapable of either swimming or swarming at agar concentrations between 0.5 and 0.6%. One reasonable interpretation is that these concentrations of agar are too high for swimming and too low to induce swarmer cell differentiation, presumably because they do not produce adequate torsional forces on the flagellar motor. This is supported by observations that swarmer cells are not observed on either 0.5 or 0.6% agar (data not shown).
Breakage of flagella occurs in both wild-type and fliL cells (Fig. 8), emphasizing that movement over agar increases the probability of flagellar damage. We also observed that when fliL cells are complemented by fliL+ fliM+ but not by fliL+ alone, the ratio of attached to unattached flagella decreased, and more flagella were attached to the fliL+ fliM+ cells. While this observation underscores the requirement of FliL in maintaining flagellar integrity, it also highlights one of the most interesting of our results that an extragenic nucleotide sequence within the fliM coding region is required for maximal FliL activity (Fig. 6 and 8).
It comes as a surprise that complementation of the Swr− phenotype in fliL cells requires the fliL promoter (including the 5′ untranslated region [UTR]), fliL coding region, and a portion of fliM DNA (Fig. 6). The results rule out the possibility that requirement of fliM is due to defects in fliM expression (transcription or translation) in YL1003 (Fig. 4 and 7). We also have preliminary evidence that full FliL function requires <200 bp of the 5′ end of fliM (unpublished data). It is tempting to speculate that a cis-acting factor located in fliM is acting to enhance or control FliL function, perhaps working in a similar manner to the antisense RNA that overlaps the fliL operon of Salmonella (50). Studies are in progress to determine the molecular mechanism behind the fliM DNA requirement.
Supplementary Material
ACKNOWLEDGMENTS
We thank David Blair for the gift of FliM antibody, Brian Youchak for assisting with the swarming assay, Rayford Payne for comments on the manuscript, and members of Belas Laboratory for helpful discussion.
J.P. is supported by the Meyerhoff Scholars Program at University of Maryland Baltimore County. This work was supported by award MCB-0919820 from the National Science Foundation.
Footnotes
Published ahead of print 7 December 2012
Supplemental material for this article may be found at http://dx.doi.org/10.1128/JB.02024-12.
REFERENCES
- 1. Mobley HL, Belas R. 1995. Swarming and pathogenicity of Proteus mirabilis in the urinary tract. Trends Microbiol. 3:280–284 [DOI] [PubMed] [Google Scholar]
- 2. Rozalski A, Sidorczyk Z, Kotelko K. 1997. Potential virulence factors of Proteus bacilli. Microbiol. Mol. Biol. Rev. 61:65–89 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Alavi M, Belas R. 2001. Surface sensing, swarmer cell differentiation, and biofilm development. Methods Enzymol. 336:29–40 [DOI] [PubMed] [Google Scholar]
- 4. Belas R, Erskine D, Flaherty D. 1991. Proteus mirabilis mutants defective in swarmer cell differentiation and multicellular behavior. J. Bacteriol. 173:6279–6288 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Allison C, Lai HC, Hughes C. 1992. Co-ordinate expression of virulence genes during swarm-cell differentiation and population migration of Proteus mirabilis. Mol. Microbiol. 6:1583–1591 [DOI] [PubMed] [Google Scholar]
- 6. Henrichsen J. 1972. Bacterial surface translocation: a survey and a classification. Bacteriol. Rev. 36:478–503 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Hoeniger JF. 1966. Cellular changes accompanying the swarming of Proteus mirabilis. II. Observations of stained organisms. Can. J. Microbiol. 12:113–123 [DOI] [PubMed] [Google Scholar]
- 8. Kearns DB. 2010. A field guide to bacterial swarming motility. Nat. Rev. Microbiol. 8:634–644 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Allison C, Coleman N, Jones PL, Hughes C. 1992. Ability of Proteus mirabilis to invade human urothelial cells is coupled to motility and swarming differentiation. Infect. Immun. 60:4740–4746 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Coker C, Poore CA, Li X, Mobley HL. 2000. Pathogenesis of Proteus mirabilis urinary tract infection. Microbes Infect. 2:1497–1505 [DOI] [PubMed] [Google Scholar]
- 11. Jacobsen SM, Stickler DJ, Mobley HL, Shirtliff ME. 2008. Complicated catheter-associated urinary tract infections due to Escherichia coli and Proteus mirabilis. Clin. Microbiol. Rev. 21:26–59 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Cusick K, Lee YY, Youchak B, Belas R. 2012. Perturbation of FliL interferes with Proteus mirabilis swarmer cell gene expression and differentiation. J. Bacteriol. 194:437–447 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Pearson MM, Rasko DA, Smith SN, Mobley HL. 2010. Transcriptome of swarming Proteus mirabilis. Infect. Immun. 78:2834–2845 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Apel D, Surette MG. 2008. Bringing order to a complex molecular machine: the assembly of the bacterial flagella. Biochim. Biophys. Acta 1778:1851–1858 [DOI] [PubMed] [Google Scholar]
- 15. Chevance FF, Hughes KT. 2008. Coordinating assembly of a bacterial macromolecular machine. Nat. Rev. Microbiol. 6:455–465 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Komeda Y. 1982. Fusions of flagellar operons to lactose genes on a mu lac bacteriophage. J. Bacteriol. 150:16–26 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Kutsukake K, Ohya Y, Iino T. 1990. Transcriptional analysis of the flagellar regulon of Salmonella typhimurium. J. Bacteriol. 172:741–747 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Macnab RM. 1996. Flagella and motility, p 123–145 In Neidhardt FC, Ingraham JL, Low KB, Magasanik B, Schaechter M, Umbarger HE. (ed), Escherichia coli and Salmonella: cellular and molecular biology. American Society for Microbiology Press, Washington, DC [Google Scholar]
- 19. Liu X, Matsumura P. 1996. Differential regulation of multiple overlapping promoters in flagellar class II operons in Escherichia coli. Mol. Microbiol. 21:613–620 [DOI] [PubMed] [Google Scholar]
- 20. Liu X, Matsumura P. 1994. The FlhD/FlhC complex, a transcriptional activator of the Escherichia coli flagellar class II operons. J. Bacteriol. 176:7345–7351 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Furness RB, Fraser GM, Hay NA, Hughes C. 1997. Negative feedback from a Proteus class II flagellum export defect to the flhDC master operon controlling cell division and flagellum assembly. J. Bacteriol. 179:5585–5588 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Pruss BM, Matsumura P. 1996. A regulator of the flagellar regulon of Escherichia coli, flhD, also affects cell division. J. Bacteriol. 178:668–674 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Kihara M, Homma M, Kutsukake K, Macnab RM. 1989. Flagellar switch of Salmonella typhimurium: gene sequences and deduced protein sequences. J. Bacteriol. 171:3247–3257 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Attmannspacher U, Scharf BE, Harshey RM. 2008. FliL is essential for swarming: motor rotation in absence of FliL fractures the flagellar rod in swarmer cells of Salmonella enterica. Mol. Microbiol. 68:328–341 [DOI] [PubMed] [Google Scholar]
- 25. Schoenhals GJ, Macnab RM. 1999. FliL is a membrane-associated component of the flagellar basal body of Salmonella. Microbiology 145:1769–1775 [DOI] [PubMed] [Google Scholar]
- 26. Motaleb MA, Pitzer JE, Sultan SZ, Liu J. 2011. A novel gene inactivation system reveals altered periplasmic flagellar orientation in a Borrelia burgdorferi fliL mutant. J. Bacteriol. 193:3324–3331 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Belas R, Horikawa E, Aizawa S, Suvanasuthi R. 2009. Genetic determinants of Silicibacter sp. TM1040 motility. J. Bacteriol. 191:4502–4512 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Jenal U, White J, Shapiro L. 1994. Caulobacter flagellar function, but not assembly, requires FliL, a non-polarly localized membrane protein present in all cell types. J. Mol. Biol. 243:227–244 [DOI] [PubMed] [Google Scholar]
- 29. Suaste-Olmos F, Domenzain C, Mireles-Rodriguez JC, Poggio S, Osorio A, Dreyfus G, Camarena L. 2010. The flagellar protein FliL is essential for swimming in Rhodobacter sphaeroides. J. Bacteriol. 192:6230–6239 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Raha M, Sockett H, Macnab RM. 1994. Characterization of the fliL gene in the flagellar regulon of Escherichia coli and Salmonella typhimurium. J. Bacteriol. 176:2308–2311 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Berg HC, Turner L. 1993. Torque generated by the flagellar motor of Escherichia coli. Biophys. J. 65:2201–2216 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Belas R, Suvanasuthi R. 2005. The ability of Proteus mirabilis to sense surfaces and regulate virulence gene expression involves FliL, a flagellar basal body protein. J. Bacteriol. 187:6789–6803 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Belas R, Erskine D, Flaherty D. 1991. Transposon mutagenesis in Proteus mirabilis. J. Bacteriol. 173:6289–6293 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Davanloo P, Rosenberg AH, Dunn JJ, Studier FW. 1984. Cloning and expression of the gene for bacteriophage T7 RNA polymerase. Proc. Natl. Acad. Sci. U. S. A. 81:2035–2039 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Amann E, Ochs B, Abel KJ. 1988. Tightly regulated tac promoter vectors useful for the expression of unfused and fused proteins in Escherichia coli. Gene 69:301–315 [DOI] [PubMed] [Google Scholar]
- 36. Guzman LM, Belin D, Carson MJ, Beckwith J. 1995. Tight regulation, modulation, and high-level expression by vectors containing the arabinose PBAD promoter. J. Bacteriol. 177:4121–4130 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Chang AC, Cohen SN. 1978. Construction and characterization of amplifiable multicopy DNA cloning vehicles derived from the P15A cryptic miniplasmid. J. Bacteriol. 134:1141–1156 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Miller VL, Mekalanos JJ. 1988. A novel suicide vector and its use in construction of insertion mutations: osmoregulation of outer membrane proteins and virulence determinants in Vibrio cholerae requires toxR. J. Bacteriol. 170:2575–2583 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Sambrook J, Russell DW. 2001. Molecular cloning: a laboratory manual, 3rd ed Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY [Google Scholar]
- 40. Zhong J, Karberg M, Lambowitz AM. 2003. Targeted and random bacterial gene disruption using a group II intron (TargeTron) vector containing a retrotransposition-activated selectable marker. Nucleic Acids Res. 31:1656–1664 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Heery DM, Powell R, Gannon F, Dunican LK. 1989. Curing of a plasmid from E. coli using high-voltage electroporation. Nucleic Acids Res. 17:10131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Livak KJ, Schmittgen TD. 2001. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25:402–408 [DOI] [PubMed] [Google Scholar]
- 43. Simons RW, Houman F, Kleckner N. 1987. Improved single and multicopy lac-based cloning vectors for protein and operon fusions. Gene 53:85–96 [DOI] [PubMed] [Google Scholar]
- 44. Miller JH. 1992. A short course in bacterial genetics. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY [Google Scholar]
- 45. Lee YY, Barker CS, Matsumura P, Belas R. 2011. Refining the binding of the Escherichia coli flagellar master regulator, FlhD4C2, on a base-specific level. J. Bacteriol. 193:4057–4068 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Hall-Stoodley L, Stoodley P. 2005. Biofilm formation and dispersal and the transmission of human pathogens. Trends Microbiol. 13:7–10 [DOI] [PubMed] [Google Scholar]
- 47. Belas R, Flaherty D. 1994. Sequence and genetic analysis of multiple flagellin-encoding genes from Proteus mirabilis. Gene 148:33–41 [DOI] [PubMed] [Google Scholar]
- 48. Rather PN. 2005. Swarmer cell differentiation in Proteus mirabilis. Environ. Microbiol. 7:1065–1073 [DOI] [PubMed] [Google Scholar]
- 49. Morgenstein RM, Szostek B, Rather PN. 2010. Regulation of gene expression during swarmer cell differentiation in Proteus mirabilis. FEMS Microbiol. Rev. 34:753–763 [DOI] [PubMed] [Google Scholar]
- 50. Wang Q, Harshey RM. 2009. Rcs signalling-activated transcription of rcsA induces strong anti-sense transcription of upstream fliPQR flagellar genes from a weak intergenic promoter: regulatory roles for the anti-sense transcript in virulence and motility. Mol. Microbiol. 74:71–84 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.