Significance
Microbial communities regulate the cycling of energy and matter in the environment, yet how they respond to environmental change is not well-known. We describe here a day in the life of wild planktonic microbial species using robotic sampling coupled with genome-wide gene expression analysis. Our results showed that closely related populations, as well as very different bacterial and archaeal species, displayed remarkably similar time-variable synchronous patterns of gene expression over 2 d. Our results suggest that specific environmental cues may elicit cross-species coordination of gene expression among diverse microbial groups, potentially enabling multispecies coupling of metabolic activity.
Keywords: autonomous sampling, environmental microbiology, metatranscriptomics, microbial oceanography, community ecology
Abstract
Planktonic marine microbes live in dynamic habitats that demand rapid sensing and response to periodic as well as stochastic environmental change. The kinetics, regularity, and specificity of microbial responses in situ, however, are not well-described. We report here simultaneous multitaxon genome-wide transcriptome profiling in a naturally occurring picoplankton community. An in situ robotic sampler using a Lagrangian sampling strategy enabled continuous tracking and repeated sampling of coherent microbial populations over 2 d. Subsequent RNA sequencing analyses yielded genome-wide transcriptome profiles of eukaryotic (Ostreococcus) and bacterial (Synechococcus) photosynthetic picoplankton as well as proteorhodopsin-containing heterotrophs, including Pelagibacter, SAR86-cluster Gammaproteobacteria, and marine Euryarchaea. The photosynthetic picoplankton exhibited strong diel rhythms over thousands of gene transcripts that were remarkably consistent with diel cycling observed in laboratory pure cultures. In contrast, the heterotrophs did not cycle diurnally. Instead, heterotrophic picoplankton populations exhibited cross-species synchronous, tightly regulated, temporally variable patterns of gene expression for many genes, particularly those genes associated with growth and nutrient acquisition. This multitaxon, population-wide gene regulation seemed to reflect sporadic, short-term, reversible responses to high-frequency environmental variability. Although the timing of the environmental responses among different heterotrophic species seemed synchronous, the specific metabolic genes that were expressed varied from taxon to taxon. In aggregate, these results provide insights into the kinetics, diversity, and functional patterns of microbial community response to environmental change. Our results also suggest a means by which complex multispecies metabolic processes could be coordinated, facilitating the regulation of matter and energy processing in a dynamically changing environment.
Planktonic microbial communities are characterized by high productivity and rapid turnover. As a result, they respond rapidly to environmental fluctuations, and minor perturbations have the potential to trigger large and rapid shifts in ecosystem properties and function (1). Characterizing the dynamics of natural microbial communities on relevant temporal and spatial scales however, remains challenging. Consequently, few data are available on the timing, magnitude, and specific biological details of response and regulation among diverse microbial taxa experiencing environmental fluctuations on short time scales in situ.
Acquiring accurate and detailed assessments of microbial dynamics in natural ecosystems poses several challenges. One challenge is the paucity of methods available for estimating the disparate activities and environmental responses of diverse microbes inhabiting complex communities. A recent approach that addresses this challenge involves the use of community RNA sequencing (e.g., metatranscriptomics) to facilitate simultaneous transcriptional profiling of co-occurring taxa within a microbial assemblage (2–5). Initial studies using this approach focused on changes in overall community metabolic profiles. Recent advances in sequencing technologies, combined with the growing number of environmentally relevant reference genomes, now enable elucidation of genome-wide transcription profiles of abundant microbial groups represented in metatranscriptomic datasets (6–8). In situ sampling of discrete, coherent microbial populations over time represents another formidable challenge. This problem is especially apparent in aquatic environments, where, because of hydrodynamic processes, samples collected at a fixed location tend to convolute temporal variability with spatial heterogeneity (6, 9). To overcome these challenges and better assess microbial community environmental responses and dynamics in situ, we combined an automated Lagrangian sampling approach with microbial community transcriptome analyses to generate a high-resolution 2-d time series of transcriptional activity among sympatric marine picoplankton populations.
Results and Discussion
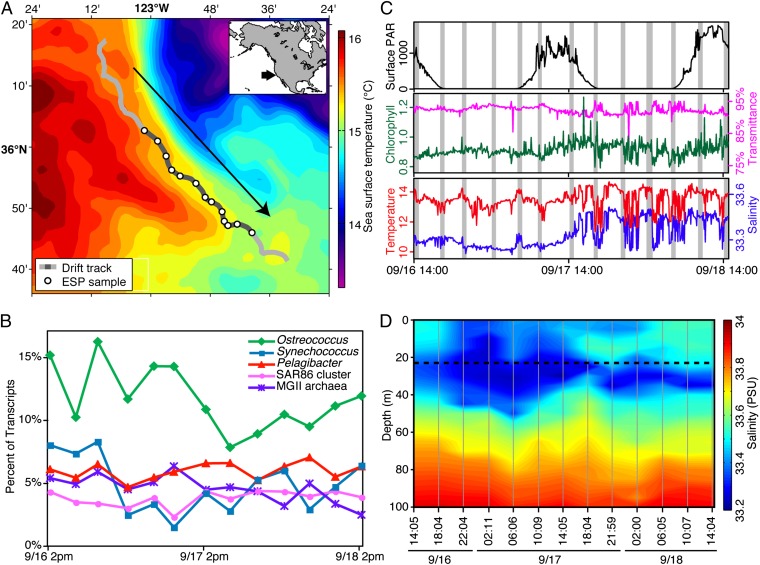
Marine surface water microbes were collected and preserved by a robotic system (10) suspended beneath a free-drifting buoy deployed off the coast of northern California (Fig. 1). Over a 2-d sampling period, the instrument drifted 50.3 km along the warm side of a front that was generated by coastal upwelling to the east (Fig. 1). Samples for community RNA sequencing were collected and preserved every 4 h. Portions of the sampling track were marked by strong vertical gradients in salinity and temperature (Fig. 1 C and D). However, current velocity data suggest that the water sampled for microbial transcriptomic analysis was relatively stable horizontally (SI Appendix, Fig. S1 and Table S1). The taxonomic representation in the metatranscriptome sampled over this time also remained relatively constant (Fig. 1). Of the over 2.4 million sequences that were assigned to 8,117 unique National Center for Biotechnology Information (NCBI) Taxonomy IDs, ∼98% matched taxa that were detected in all 13 samples. Additionally, our samples showed greater overall similarity in taxonomic composition to one another across all of the time points than did samples collected over a 24-h period at a fixed spatial location in Monterey Bay (6) (SI Appendix, Fig. S2).
Fig. 1.
Sampling locations and sample characteristics. (A) The ESP drift track imposed over a map showing average sea surface temperature [Polar orbiting environment satellites, advanced very high resolution radiometer, local area coverage. Western United States, day/night, 5 × 5-pixel (5 × 5-km) median-filtered composite from September 21, 2010; cloud cover precluded satellite observation during the sampling period]. Inset shows the sampling location relative to western North America. (B) Transcript abundances of major taxa represented as percent of sequences with matches in the NCBInr peptide database. (C) Environmental conditions in the immediate vicinity of the ESP (measurements taken by ESP-mounted instruments). Grey bars represent sample collection times. (D) Integrated depth profile showing salinity gradients surrounding the sampling location (dotted line). Measurements were taken near the ESP by a ship-deployed instrument. The arrow in A indicates the direction of the drift.
Our analyses focused on transcriptional dynamics among five abundant microbial populations in the sampled community, including Ostreococcus, Synechococcus, Pelagibacter, SAR86 cluster Gammaproteobacteria (SAR86), and marine group II Euryarchaea (MGII) (Table 1). These populations represent widely distributed and ecologically important clades of marine picoplankton (11–15). Transcripts mapping to each of these groups were identified and annotated using a newly developed computational workflow (SI Appendix, Fig. S3) that assigned sequences to specific taxon bins based on best-scoring matches within the full NCBI database. Within each taxon bin, transcript counts for genes shared between multiple reference genomes of the same taxa were combined, and analyses of transcriptional dynamics focused on changes in relative transcript abundance within each specific taxonomic population, independent of fluctuations in its abundance relative to the total community transcriptome. (Table 1 and SI Appendix, Figs. S3–S11 and Tables S2 and S3 have details of transcript mapping and annotation.)
Table 1.
Assignment of sequences to taxon bins
| Sample | CDS |
Ostreococcus |
Synechococcus |
Pelagibacter |
SAR86 cluster |
MGII Archaea |
|||||
| Reads | Orthologs | Reads | Orthologs | Reads | Orthologs | Reads | Orthologs | Reads | Orthologs | ||
| 9/16 2:00 PM | 234,860 | 35,590 | 4,661 | 18,777 | 2,535 | 14,262 | 1,278 | 10,053 | 1,499 | 12,600 | 1,195 |
| 9/16 6:00 PM | 208,667 | 21,292 | 4,148 | 15,286 | 2,437 | 11,275 | 1,199 | 7,207 | 1,383 | 10,185 | 1,097 |
| 9/16 10:00 PM | 198,687 | 32,218 | 5,112 | 16,412 | 2,282 | 12,850 | 1,235 | 6,665 | 1,173 | 11,635 | 1,229 |
| 9/17 2:00 AM | 269,745 | 31,412 | 4,908 | 6,551 | 1,749 | 12,508 | 1,253 | 7,990 | 1,438 | 12,072 | 1,160 |
| 9/17 6:00 AM | 209,305 | 29,903 | 4,402 | 6,908 | 1,618 | 11,339 | 1,238 | 8,003 | 1,441 | 10,574 | 1,233 |
| 9/17 10:00 AM | 384,870 | 54,840 | 4,271 | 5,579 | 1,279 | 22,572 | 1,377 | 8,698 | 1,341 | 24,340 | 1,384 |
| 9/17 2:00 PM | 209,634 | 22,695 | 4,037 | 8,714 | 1,955 | 13,707 | 1,304 | 9,057 | 1,490 | 9,309 | 1,127 |
| 9/17 6:00 PM | 183,482 | 14,357 | 3,443 | 5,046 | 1,406 | 12,056 | 1,192 | 6,804 | 1,268 | 8,512 | 1,105 |
| 9/17 10:00 PM | 176,778 | 15,692 | 4,231 | 9,261 | 2,070 | 9,161 | 1,180 | 7,678 | 1,480 | 7,643 | 1,142 |
| 9/18 2:00 AM | 220,569 | 23,025 | 4,501 | 13,138 | 2,035 | 13,859 | 1,284 | 9,456 | 1,527 | 6,922 | 1,067 |
| 9/18 6:00 AM | 222,828 | 21,064 | 3,239 | 6,384 | 1,306 | 15,609 | 1,217 | 8,717 | 1,309 | 11,038 | 1,240 |
| 9/18 10:00 AM | 208,782 | 23,215 | 3,641 | 9,730 | 1,818 | 11,393 | 1,195 | 9,010 | 1,521 | 6,954 | 1,040 |
| 9/18 2:00 PM | 232,718 | 27,704 | 3,699 | 14,811 | 2,216 | 14,639 | 1,098 | 8,973 | 1,451 | 5,692 | 925 |
| Total | 2,960,925 | 353,007 | 7,491 | 136,597 | 3,950 | 175,230 | 1,810 | 108,311 | 2,226 | 137,476 | 1,691 |
The total number of putative coding sequences (unique nonrRNA sequence reads with at least one hit in the NCBInr database with bit score >50) identified in each sample is listed. For each taxon bin, the number of sequence reads assigned to that group and the number of ortholog clusters with at least one assigned sequence within each sample are listed. CDS, coding sequence.
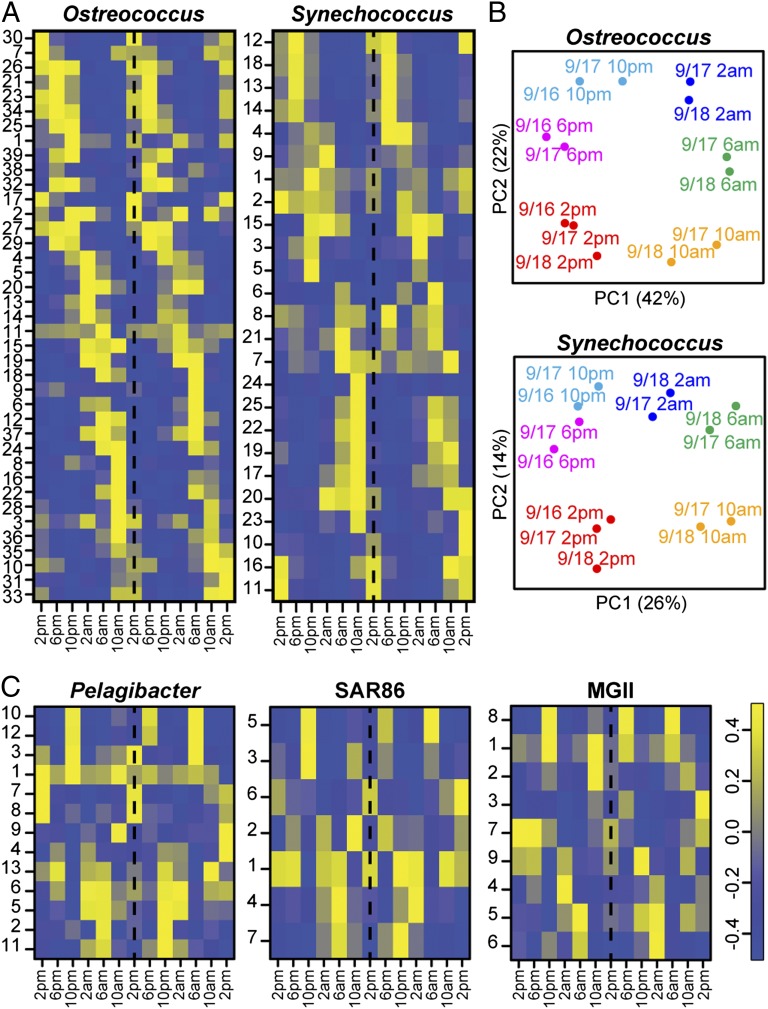
To confirm that complex transcriptional dynamics could be observed within transcriptional profiles extracted from community-wide gene expression datasets, we used cluster analysis to assess global transcriptional patterns among the five most highly represented taxonomic groups (Fig. 2 and SI Appendix, Figs. S12–S17). Within each taxon, we identified groups of genes that shared similar transcriptional profiles using the geneARMA software package (16), which uses an autoregressive moving average model, ARMA(p,q), for the longitudinal covariance structure and Fourier series functions to model gene expression patterns. As anticipated, a large number of coexpressed genes with apparent 24-h periodicity were identified among Ostreococcus and Synechococcus populations. Additionally, principal components analysis clearly separated the transcriptome profiles of these taxa based on time of day (Fig. 2B). Pure cultures of both Ostreococcus and Synechococcus are known to have fully functional circadian clocks that coordinate large-scale transcriptional dynamics (17, 18), and those rhythms were readily apparent in the transcriptome profiles of wild populations as well as the behavior of individual transcripts (see below). In contrast, cluster analysis of transcription among the three proteorhodopsin-expressing heterotrophic populations did not exhibit evidence of significant diel regulation of gene expression (Fig. 2C). Nevertheless, transcripts recovered from these populations, particularly transcripts from Pelagibacter, did reveal variable and coordinated regulatory patterns among a large variety of different gene suites and metabolic pathways (Fig. 2 and SI Appendix, Figs. S15–S17).
Fig. 2.
Global transcriptional profiles from phototrophic and heterotrophic taxon bins. Heat maps depict third-order Fourier series models from geneARMA clustered transcripts within phototrophic (A) and heterotrophic (C) taxa. Heat maps show model amplitude at the 13 time points (columns) for each of the geneARMA clusters (rows). GeneARMA cluster models were normalized to an amplitude of one. Membership information and individual gene transcript traces are shown in SI Appendix, Figs. S13–S17. (B) Principal components analysis of Ostreococcus and Synechococcus transcriptomes. Axes represent the first and second principal components and are labeled with the percent of total variance explained.
Diel Rhythms in Gene Expression Among Ostreococcus and Synechococcus.
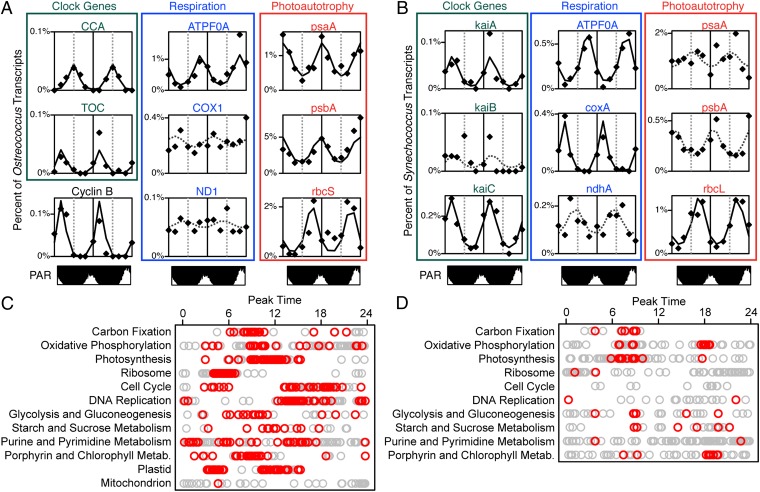
To further explore diel transcriptional rhythms, we used harmonic regression-based analyses to identify individual transcripts that followed a sinusoidal curve with 24-h periodicity (Fig. 3). Using this approach, 2,097 of 7,491 Ostreococcus transcripts and 130 of 3,950 Synechococcus transcripts were identified as significantly periodic. None of the transcripts from the three heterotrophic populations were identified as significantly periodic using this method.
Fig. 3.
Periodic gene expression in Ostreococcus- and Synechococcus-assigned transcripts. (A and B) 48-h time series of observed (points) and fitted (lines) transcript abundances is shown for selected transcripts from Ostreococcus (A) and Synechococcus (B) populations. Fitted values with solid lines represent transcripts with significantly periodic expression, whereas dotted lines represent best-fit curves for transcripts not passing significance cutoffs. For reference, plots of relative light levels are shown. (C and D) Plots showing peak expression times for all orthologs (grey circles) and significantly periodic orthologs (red circles) assigned to major cellular functions in Ostreococcus (C) and Synechococcus (D). KEGG pathways for photosynthesis proteins and antenna proteins were combined for the purposes of this plot along with purine and pyrimidine metabolism pathways. Ostreococcus (OC) and Synechococcus (SC) ortholog cluster designations for transcripts in A and B: ATPF0A, ATP synthase subunit A, OC 9555 (plastid-encoded), SC 1180; Circadian Clock Associated 1 (CCA1) and Timing of Cab expression 1 (TOC1), Ostreococcus clock genes OC 3107 and 7575; COX1, coxA, cytochrome c oxidase subunit I, OC 9595 (mitochondrial), SC 1503; Cyclin B, mitotic cyclin B, OC 658; kaiA, -B, and -C, Synechococcus clock genes SC 332, 3370, and 334; ND1, ndhA, NADH dehydrogenase I subunit 1, OC 9600 (mitochondrial), SC 210; PAR, photosynthetically available radiation; psaA, PSI apoprotein A1, OC 9562 (plastid-encoded), SC 2040; psbA, PSII reaction center D1, OC 9541 (plastid-encoded), SC 1091; rbcS, rbcL, RuBisCo large and small subunits, OC 6808, SC 130.
Ostreococcus population transcripts showing strong periodic trends in transcript abundance included the master clock genes Circadian Clock Associated 1 and Timing of Cab expression 1 and genes associated with major metabolic functions (Fig. 3). Ribosomal protein gene expression peaked in the early morning followed by an increase in gene transcripts associated in carbon fixation, a subsequent maximum in photosynthesis gene expression around midday, and finally, cell cycle and DNA replication gene expression, which reached a maximum at the end of the day. Although only one mitochondrial gene (NADH dehydrogenase I subunit 6) was identified as significantly periodic, 49 of 61 total plastid genes exhibited 24-h periodicity, with a predawn peak of biosynthetic genes (ribosomal proteins and RNA polymerase) and a midafternoon peak of photosynthesis genes. Interestingly, whereas most genes involved in carbon fixation were identified as significantly periodic with peak expression around 8:00 AM, the large subunit of RuBisCO (the only carbon fixation gene still encoded by the Ostreococcus plastid genome) did not show cyclical trends in transcript abundance.
Among Synechococcus population transcripts, two of three kaiABC clock genes as well as genes involved in oxidative phosphorylation, photosynthesis, respiration, and carbon fixation (Fig. 3) exhibited periodic trends in transcript abundance. The third clock gene, kaiB, was not identified as significantly periodic, but its relatively low coverage level (zero to five copies per library) may have precluded accurate quantification. ATP synthase, carboxysome, and Calvin cycle transcripts largely exhibited a morning peak in expression, whereas cytochrome oxidase transcripts peaked in the evening. In contrast to observed patterns among Ostreococcus, only one photosystem gene (Cluster 8842 psaK) and relatively few antenna proteins (4 of 25) were identified as significantly periodic in our dataset. Genes in these categories did, however, show high-amplitude and highly variable expression, suggesting that they may be differentially regulated based on environmental factors. Also unlike Ostreococcus, evidence for diel periodicity in growth and division among this Synechococcus population was relatively weak, with no cell cycle or DNA replication transcripts showing significantly periodic expression. However, given the complicated relationship between nutrient status and the timing of DNA replication and cell division in Synechococcus (19, 20), the unknown growth state of this population, and the apparent presence in our dataset of two ecologically distinct Synechococcus clades (SI Appendix, Fig. S8), a lack of a clear periodic signal in population-averaged transcript abundance for individual cell cycle and DNA replication genes is perhaps not surprising.
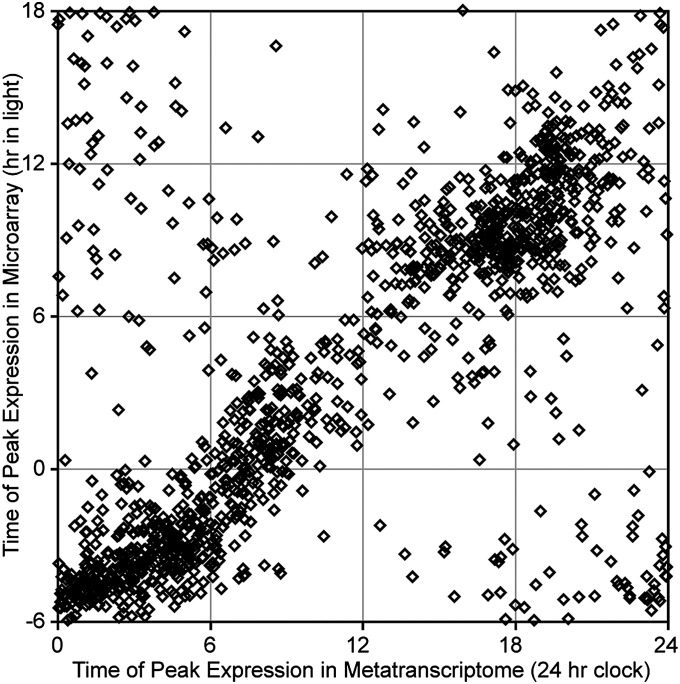
Both the Ostreococcus and Synechococcus data reflected broad trends previously observed in laboratory monocultures. The diurnal timing of expression of transcripts in wild Ostreococcus populations in particular was remarkably consistent with gene expression patterns observed in microarray-based laboratory studies of O. tauri cultures grown in 12:12-h light:dark cycles (18) (Fig. 4). However, there were also a number of differences between results obtained for our natural populations and those results observed in laboratory analyses of pure cultures.
Fig. 4.
Comparison of peak expression times for periodically expressed Ostreococcus orthologs in field populations vs. a laboratory pure culture. Each point represents 1 of 1,290 transcripts detected as significantly periodic in our field study reported here and a previous microarray study of O. tauri (18). For this comparison, microarray data (as reported in Gene Expression Omnibus accession no. GSE16422) were reprocessed using our harmonic regression method with a Gaussian error model.
A direct comparison of our field study with any previously published laboratory studies of Synechococcus was problematic, because most existing datasets (17) focus on S. elongatus, a freshwater species, and were performed under constant light illumination. It is worth noting, however, that our study identified fewer Synechococcus transcripts that exhibited periodic trends in expression compared with laboratory studies of the freshwater Synechococcus species. However, the orthologs identified in our field populations do not simply represent a high-amplitude subset of the periodically expressed transcripts detected in laboratory studies. Of 69 periodically expressed Synechococcus orthologs in our field study that could be mapped to probes in the laboratory-based microarray study, 24 orthologs were not identified as significantly periodic under laboratory conditions.
For Ostreococcus, the populations reported here and a previous laboratory study (18) identified ∼2,000 periodically expressed Ostreococcus transcripts. However, of 1,683 significantly periodic orthologs in our field populations that could be mapped to probes in the O. tauri microarray, 881 orthologs were not identified as significantly periodic in the laboratory study. Reprocessing the laboratory microarray data using our regression-based approach (with a Gaussian error model) increased the overlap in significantly periodic genes, but this analysis still yielded 393 genes identified as periodic in our field data but not in the laboratory study. We did not identify any obvious biological trends among Ostreococcus transcripts that were identified as periodically expressed in the field but not the laboratory. Although some of these differences in gene expression patterns may be the result of methodological differences, many are likely to represent responses to cues present in the natural environment but not within the relatively static laboratory environment.
In sum, these analyses validate our approach and confirm that complex transcriptional patterns within distinct populations can be resolved within bulk community RNA profiles. Although previous studies have suggested single-time point day/night differences in the overall transcriptional profiles of marine microbial communities, our analyses here provide a much higher-resolution picture of genome-wide diel transcriptional dynamics among different microbial populations in a natural microbial community in situ.
Transcriptional Dynamics Within Pelagibacter Population Transcripts.
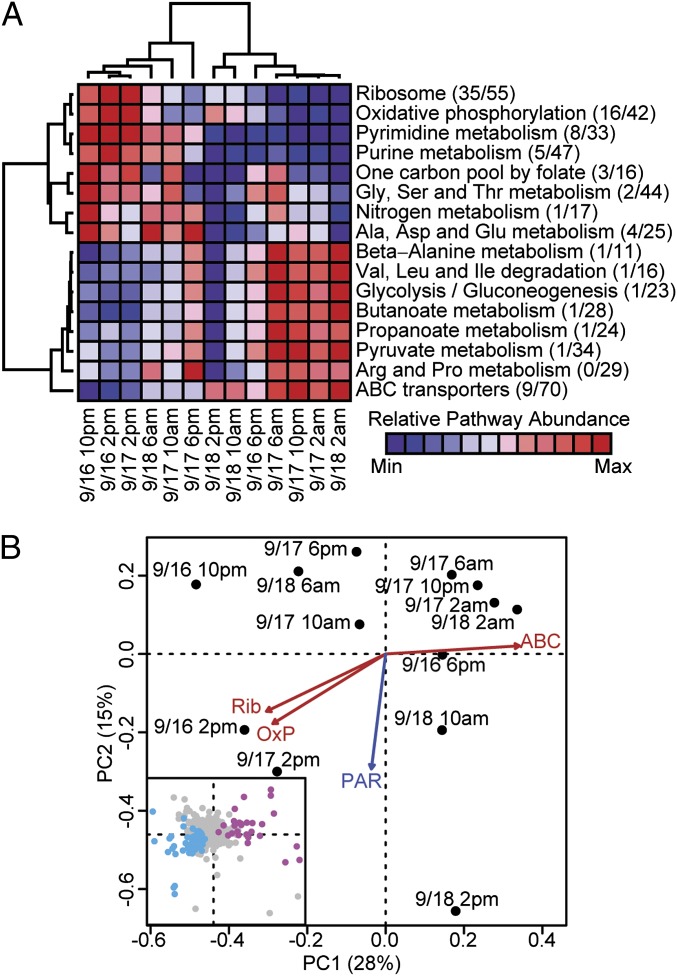
The naturally occurring Pelagibacter populations that we sampled did not exhibit strong circadian rhythms of gene expression. We did however observe evidence for well-orchestrated, genome-wide transcriptional regulation within this group. Hierarchical clustering of samples and pathways showed a large degree of covariance between some major metabolic pathways (Fig. 5A). In particular, the pathway-level signal for ribosomal proteins and oxidative phosphorylation showed strong positive correlation with one another (correlation coefficient = 0.98, P value = 1 × 10−8) and were negatively correlated with many transport gene transcripts, including the ATP binding cassette (ABC) transporter family (correlation coefficient = −0.88, P value = 1 × 10−5 for ribosomal proteins vs. ABC). Principal components analysis suggested that these metabolic signals explain more of the variability observed in wild Pelagibacter transcript profiles than any of the measured environmental parameters (Fig. 5B). Furthermore, a Poisson regression-based analysis found that 101 of 1,810 observed Pelagibacter transcripts were significantly correlated with pathway-level signals for either ribosome biosynthesis or ABC transport (SI Appendix, Tables S4 and S5). A total of 74 of these transcripts was identified as up-regulated in tandem with the ribosome synthesis pathway (and down-regulated with transport up-regulation), including not only transcripts coding for ribosomal proteins but also genes associated with C1 metabolism (21), secretion, ATP synthase (six of nine subunits), and proton-translocating pyrophosphatase. The 27 transcripts that followed the opposite trend (up-regulated with transporters and down-regulated with ribosome synthesis) seemed to represent a generalized transport signal, encompassing not only ABC transporters of amino acids, polyamines, and phosphonates but also TRAP (Tripartite ATP-independent Periplasmic) transporters of carboxylic acids, an ammonium transporter, and an Na+/solute symporter.
Fig. 5.
Analysis of Pelagibacter transcriptional profiles. (A) Heat map showing relative abundance of major metabolic pathways among Pelagibacter-assigned sequences. Hierarchical clustering of samples and pathways used average-linkage clustering based on Pearson correlation coefficients. For each pathway, the fraction of transcripts assigned to each pathway that is significantly correlated (based on Poisson regression) with the overall pathway-level signal is listed. (B) Principal components analysis of Pelagibacter transcriptional profiles. Axes represent the first two components and are labeled with the proportion of variance explained by each. Vector fits for selected KEGG pathways (ABC, ABC transporters; OxP, oxidative phosphorylation; Rib, ribosomal proteins) were highly significant (P < 0.0001) and are shown in red. Of the environmental data collected, only surface PAR (blue; P = 0.003) was significantly correlated (P < 0.05) with the ordination. (Inset) Loadings of Pelagibacter transcripts on the principal component axes. Transcripts significantly correlated with either ribosome or ABC transport pathways are colored based on their relationship with those pathways (cyan for orthologs positively correlated with ribosome and/or negatively correlated with ABC transport; magenta for the opposite relationship).
In many microbial species, the abundance of ribosomal proteins and their transcripts is tightly regulated with respect to cellular growth rate (22–24), and the relative abundance of ribosomal protein transcripts for a given taxon has been proposed as a metric for assessing in situ growth rates (8). The trends in Pelagibacter gene expression that we report here however, suggest a very dynamic and rapidly fluctuating reallocation of cellular resources between growth and nutrient acquisition. In this context, cell populations exhibiting decreased ribosomal protein synthesis and increased transporter activity are most likely indicators of sporadically limiting substrate availability in the ambient environment. The broad range of transporters that were expressed when ribosomal synthesis was down-regulated does not suggest limitation by any single nutrient. Additionally, neither set of genes seems to reflect stationary-phase responses previously reported in laboratory cultures of Candidatus P. ubique (25) (SI Appendix, Tables S4 and S5), suggesting that none of these populations have entered a starvation state. Instead, these trends reflect highly dynamic and variable transcriptional responses (and potentially, metabolic and growth rate variability) over short time scales, that seem to be dictated by surrounding environmental and nutrient variability.
Synchronous Transcriptional Dynamics Within Pelagibacter, SAR86, and MGII.
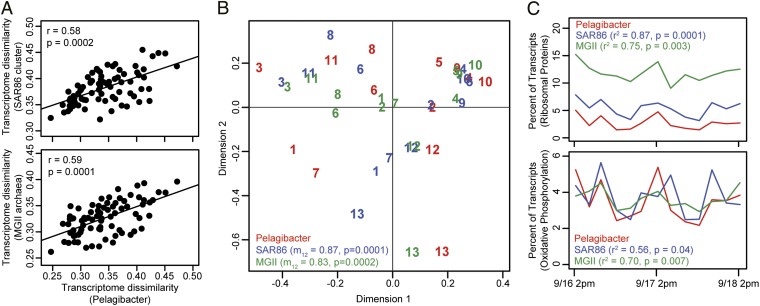
Although Pelagibacter exhibited the strongest and most coherent patterns in transcript abundance among the three proteorhodopsin-bearing heterotrophic populations examined, SAR86 and MGII populations also exhibited transcriptional changes over the 2-d time series. Interestingly, these patterns seemed to reflect synchronous responses to the same cryptic environmental changes that appeared to be driving Pelagibacter transcription dynamics. The degree of similarity between samples for the three organisms was significantly related (Fig. 6A), suggesting that the overall transcriptional profiles of these groups were changing simultaneously, or nearly so. This relationship was even more evident when Procrustes tests were used to compare only the first two axes of population-specific principal components analyses (Fig. 6B), restricting the comparison to the strongest trends in sample-to-sample variability. Furthermore, the relative abundance of transcripts involved in ribosome biosynthesis and oxidative phosphorylation was positively correlated across the 13 time points for the three groups (Fig. 6C and SI Appendix, Table S6). Similarly, independent geneARMA analyses of the three taxa identified gene cluster models exhibiting similar trends in transcript abundance across the three datasets (SI Appendix, Fig. S18). Altogether, these analyses suggest that all three populations were responding to a common environmental signal, exhibiting global, synchronous changes in taxon-level taxonomic profiles.
Fig. 6.
Synchronous transcriptional dynamics among three heterotrophic populations. (A) Mantel test showing a significant relationship in transcriptome dissimilarity for Pelagibacter vs. SAR86 (Upper) and MGII (Lower) populations. Comparisons used pairwise Euclidean distances (square root of the sum of squared differences in abundance for all transcripts). (B) Procrustes analysis revealing a large degree of congruence in sample clustering patterns for the three heterotrophic populations. In Procrustes tests, the results of principal components analyses are rotated and scaled to identify similarities in clustering patterns while maintaining relationships between samples. A smaller distance between points corresponding to a single sample reflects a more similar clustering pattern. Rotated and scaled SAR86 (blue) and MGII (green) analyses are overlaid on the Pelagibacter (red) results from Fig. 5B. Samples are labeled according to position in the time series (9/16 2:00 PM is sample 1). Procrustes correlation (m12 ) and permutation-based significances are shown for each comparison. (C) The relative abundance of transcripts for ribosomal proteins (Upper) and genes associated with oxidative phosphorylation (Lower) within each taxon bin at each time point. Pearson correlation coefficient and P value are listed for relative abundances of these pathways within the Pelagibacter vs. their abundances in SAR86 and MGII transcriptomes.
Among heterotrophic marine picoplankton, Pelagibacter, SAR86, and MGII have been hypothesized to catabolize different types of carbon molecules (14, 15, 21, 26). Pelagibacter seems to use simple peptides, amino acids, osmolytes, and single carbon compounds, whereas the SAR86 and MGII groups have been hypothesized to specialize in the consumption of larger, more complex polymers, such as proteins, polysaccharides, and lipids. Consistent with these predictions, the most abundant transcripts for each of the taxon bins reflected very different metabolic profiles (SI Appendix, Fig. S19). Pelagibacter expressed ABC transporters for small peptides, osmolytes, and dicarboxylic acids at high levels, consistent with genomic analyses (26). SAR86 transcripts included a large number of TonB-dependent receptors, previously hypothesized to mediate uptake and metabolism of large polysaccharides and lipids in this organism (14). Finally, the MGII transcriptome was dominated by large cell surface proteins and amino acid transporters, consistent with a hypothesized ability to metabolize large proteins (15). Therefore, although these three very different populations exhibited similar trends in expression of pathways involved in growth and energy metabolism, they did not show similar trends in most metabolic pathways (SI Appendix, Table S6). This trend suggests that the synchronous transcriptional dynamics observed for these groups may reflect bulk changes in the availability of a broad range of carbon-based substrates rather than responses to the availability of a single common nutrient or substrate limitation.
Overall, the Pelagibacter population showed a stronger global transcriptional response, involving a larger number of transcripts, than the transcriptional responses observed for the SAR86 and MGII populations. Although some of this difference may be due to the higher sequence coverage of Pelagibacter (because of its smaller genome and greater overall abundance), SAR86 and MGII showed less sample-to-sample variation between transcriptional profiles than did Pelagibacter, even when all datasets were resampled to represent an even coverage level. Pelagibacter species have been shown to have highly streamlined genomes with reduced regulatory machinery (26). As a result, these α-proteobacteria may exhibit a smaller range of transcriptional dynamics in response to complex environmental cues, resulting in strong global transcriptional dynamics. In contrast, SAR86 and MGII, with larger genomes and corresponding increased metabolic and regulatory versatility, might be expected to exhibit more complex time- and location-specific behaviors not as easily distinguished using cluster- and correlation-based approaches. Alternatively, the SAR86 and MGII populations may simply respond less strongly to the environmental cues that elicited strong Pelagibacter responses. It is also possible that higher-molecular weight, more-complex substrates preferred by SAR6 and MGII had a more patchy distribution than the simple peptides and osmolytes used by Pelagibacter. As a result, gene expression in SAR86 and MGII populations might be expected exhibit a larger degree of cell-to-cell variability, resulting in weaker or less synchronized transcriptional dynamics when averaged at the temporal (∼40 min) and spatial (∼1 L) scale of our sample collections.
Notably, although many pathway-level signals from SAR86 and MGII populations were individually correlated with pathway-level signals from Pelagibacter, these populations showed significantly less congruence to each other (SI Appendix, Table S6). The metabolic profiles of SAR86 and MGII, although indicative of different substrate preferences, share the commonality of the binding and hydrolysis of large polymeric substrates, such as cell wall and membrane components. The overall abundances of SAR86 and MGII transcripts in the metatranscriptomes are negatively correlated (correlation coefficient = −0.55, P value = 0.047), which may suggest some degree of niche overlap and competition between these organisms. In contrast, Pelagibacter specializes in a very different fraction of the substrate pool, including low-molecular weight monomers, such as carboxylic acids and amino acids, that are likely to be generated as a byproduct of both SAR86 and MGII metabolic activities.
Altogether, our results revealed unexpected interspecies synchronicity in the regulation of some pathways, as well as a surprising degree of heterogeneity in the transcriptional profiles among photoheterotrophic picoplankton populations. Transcripts encoding ribosomal proteins and genes involved in oxidative phosphorylation were previously identified as highly variable between samples collected at distant geographic locations (5). Notably, we observed as much variability in transcript abundance in these pathways over only a few hours time and in the same water mass, as had been previously reported in transoceanic metagenomic surveys. These data suggest that activity levels and respiration rates for heterotrophic populations may be spatially and temporally patchy in marine surface waters, potentially due to episodic substrate releases, such as small-scale lytic events and other stochastic environmental processes. Additional exploration of these behavioral patterns and the environmental cues that control them is likely to provide significant insight into niche specialization of key microbial groups in the planktonic environment.
Implications.
Episodic environmental variation and subsequent microbial responses play significant roles in shaping marine biogeochemical cycles (1). To better understand and predict microbial community responses to such events, it is critical to observe them on relevant temporal and spatial scales in situ. Here, we show that Lagrangian sampling combined with microbial community transcriptome analyses can resolve microbial dynamics on time scales of hours to days, yielding robust transcriptional expression patterns. Within a given taxon, the presence of reproducible temporal patterns in the genome-wide transcription profile indicated that a large fraction of individual cells within a given population was responding synchronously, at least within the temporal resolution of our measurements. Furthermore, disparate heterotrophic taxa within the community also seemed to be simultaneously responding to similar environmental cues but expressed different functional gene suites in response to them, suggesting a potential means by which multispecies metabolic and biogeochemical processing might be coordinated.
The specific environmental factors that influence the observed synchronized transcriptional regulation of diverse heterotrophic microbial species are unknown at present. It may be that each species population is responding independently to the same (or simultaneously occurring) physicochemical environmental cues. However, we cannot rule out that these transcriptional patterns are partly influenced by species-to-species communication and signaling cascade events. It is well-known that specific autoinducer molecules can elicit complex regulatory responses within and between disparate bacterial species (27). It, therefore, seems possible that specific physicochemical environmental cues might be sensed initially by only one or a few species. These cues might then be indirectly broadcast to other species by small-molecule signaling, thereby transmitting the response to other community members. Future work, using higher-frequency sampling of microbial community transcriptional profiles, may provide the temporal resolution necessary to distinguish between these different cross-community sensing and response modalities. Regardless of the specific mechanism(s) of multipopulation environmental sensing and response, both the above possibilities could elicit the multispecies transcriptional events that we observed. This temporal entrainment could conceivably serve to coordinate downstream biogeochemical processing and nutrient regeneration. For example, hydrolysis of higher-molecular weight organic compounds by SAR86 and GII Euryarchaea could produce monomers that were subsequently processed by Pelagibacter.
Very little is known about the actual metabolic rates of specific heterotrophic picoplankton species in the environment. Most in situ growth estimates have been derived from bulk averaged radioisotope incorporation into DNA or protein across entire (heterogeneous) assemblages. Complex predator–prey dynamics involving phages and protists further complicate the measurement of species-specific growth rates in situ. The transcriptional profiles of heterotrophic marine picoplankton that we observed suggested that disparate species were responding rapidly to environmental variability with frequent and synchronized up- or down-regulation of transcripts in many pathways. In particular, gene transcript abundance in growth-related pathways, like ribosome biosynthesis and oxidative phosphorylation, varied significantly over the 2-d sampling period in these populations. Notably, we did not observe gene expression patterns that would suggest transition into the stationary phase over the 2-d sampling period. In total, these data suggest that frequent periods of metabolic acceleration and deceleration, even over the time span of only one doubling, may be a common modality in heterotrophic marine picoplankton species in situ.
The kinetics and regularity as well as quantitative and qualitative attributes of microbial response dynamics in situ have implications beyond biogeochemical considerations. Short time-scale microbe–environment and microbe–microbe interactions ultimately give rise to microbial population variation, functional variability, microbial community succession, and large-scale taxonomic shifts over days, weeks, and months and across seasons. A better understanding of short time-scale ecological microbial community processes should therefore provide a new perspective on longer-term community assembly, structure, and functional patterns. Future studies using the approaches we describe here have potential to yield deeper insight into microbial environmental interactions and their ecological consequences in a dynamic and constantly changing environment.
Methods
Sample Collection.
Seawater samples (1 L) were collected along the central California coast in September of 2010 using an Environmental Sample Processor (ESP) (6, 10) suspended beneath a free-drifting surface float at 23-m depth. Microbes in the 0.22- to 5-μm size fraction were collected and preserved as previously described (6) but with a reduced incubation time in RNALater (Ambion) of 2 min per wash, which yields RNA of similar integrity. The instrument was recovered on September 19, 2010, and sample filters were moved to individual vials for long-term storage at −80 °C within 36 h.
Library Preparation and Sequencing.
Approximately one-half of each filter was used for extraction of total community RNA and subtractive hybridization of rRNAs as previously described (28). Synthesis of antisense rRNA probes used DNA extracted from 5.8- to 7.1-L seawater samples collected using a rosette sampler at 23-m depth near the ESP at 10:00 AM on September 15, 17, and 19. PCR products from the three dates were pooled for use as templates for synthesis of bacterial, archaeal, and eukaryotic large- and small-subunit rRNA probes. Approximately 150 ng total community RNA were hybridized with 300 ng each bacterial, 100 ng each archaeal, and 150 ng each eukaryotic small- and large-subunit probes. Probe removal used two successive 5-min incubations with 75 μL washed Streptavidin beads (NEB) in a final volume of 50 μL. Purified and concentrated message was linearly amplified and converted to cDNA as described previously (2).
A GS FLX Titanium system (Roche) was used to sequence cDNA. Library preparation followed the Titanium Rapid Library Preparation protocol. To improve the retention of smaller cDNA molecules, adaptor-ligated libraries were not diluted before size selection with AMPure XP beads. Libraries were quantified using the Titanium Slingshot kit (Fluidigm) and added to emulsion PCR reactions at 0.1 molecules per bead. Sequencing and quality control followed the manufacturer’s recommendations.
Sequence Analysis and Annotation.
Our analytical pipeline for sequence annotation is summarized in SI Appendix, Fig. S3. Metatranscriptomic sequence libraries were screened for rRNA-derived transcripts and duplicates as previously described (28). Putative coding sequences with bit scores ≥ 50 were initially identified by BLASTX against the NCBI nonredundant peptide database as downloaded on May 31, 2010. After this initial analysis, additional reference sequences became available for SAR86 cluster Gammaproteobacteria and MGII Euryarchaea. All unique nonrRNA sequences were again compared by BLASTX with these newly released genome sequences, retaining those sequences with bit scores ≥50 that were greater than or equal to their best match in the previous NCBInr database search (SI Appendix, Fig. S3). Sequence classification and annotation used the highest-scoring database match and followed the NCBI taxonomy (with the exception of α-proteobacterium HIMB114, which we included within the Pelagibacter). For sequences matching equally well to multiple genes within the database, all matches were required to fall within the Chlorophyta for assignment to Ostreococcus, the Cyanobacteria for Synechococcus, and the SAR11 cluster for Pelagibacter. All top-scoring matches were required to fall within the SAR86 cluster or the MGII Euryarchaea for assignment to those taxon bins. Sequences were mapped to a single reference gene for annotation purposes, with preference given to references that were abundant in the dataset and references derived from sequenced genomes. Data files containing all taxon-specific transcript sequences for Ostreococcus, Synechococcus, Pelagibacter, SAR86, and Group II Euryarchaea are available from the authors on request.
Within each major taxonomic bin, sequence counts for genes present in multiple reference genomes were compiled to generate ortholog cluster-based transcript abundances. This approach was implemented to avoid artificial division of transcript pools from environmental organisms among multiple imperfectly matched reference sequences. Pairwise reciprocal best BLAST hits between translated coding sequences of reference genomes were compiled to generate ortholog cluster assignments. Identification of shared genes in Ostreococcus used the previously described e-value–based significance cutoff of 10−8 (29), whereas Synechococcus, Pelagibacter, and MGII comparisons used an e-value cutoff of 10−5 and required 30% alignment identity over 80% of the longer sequence; SAR86 comparisons used the 10−5 e-value and 30% identity cutoffs, but (because of the highly fragmented nature of the SAR86 assemblies) only 50% of the longer sequence was required to align. Functional annotation of ortholog clusters used the Kyoto Encyclopedia of Genes and Genomes (KEGG) (30) annotations where available. Key species lacking curated annotations were analyzed using the KEGG automated annotation pipeline (31). In some cases, metatranscriptomic sequences were mapped to reference genes that were not derived from sequenced genomes (i.e., environmental clones). Where possible, these references were assigned to ortholog clusters based on single-directional peptide BLAST (significance cutoffs as above). Full lists of ortholog cluster membership, annotation, and results of statistical analyses are available in Datasets S1, S2, and S3.
Functional Clustering of Transcriptional Profiles.
Cluster-based analyses were used to examine global patterns of transcription within and across taxon bins. Because many of these analyses assume normally distributed data, a variance-stabilizing transformation (32) was applied before analysis:
 |
where c is the count of each transcript and N is the library size at each time point. Mantel tests, principal components analyses, and Procrustes tests used variance-stabilized transcript abundances and were carried out using the vegan software package (33).
The geneARMA (16) package was used to identify global patterns of coregulation among Ostreococus, Synechococcus, Pelagibacter SAR86, and MGII transcripts. This algorithm performs model-based soft clustering of transcriptional profiles using an autoregressive moving average model, ARMA(p,q), for the longitudinal covariance structure and Fourier series functions to model gene expression patterns. Data were filtered before analysis such that the maximum count for a row (ortholog time series) was greater than 5 and the sum of the row was greater than 10. For each dataset, the algorithm was run multiple times (100–400 iterations) from random initializations for P = 1–2 autocorrelation terms, q = 0–1 moving average terms, K = 1–3 Fourier function terms, and J = 1–40 clusters. The iteration possessing the highest likelihood for each parameter combination was used for final inference, and the Akaike Information Criterion was used to assess model complexity. The optimal model configuration, as identified by Akaike Information Criterion, for all five datasets was an ARMA(1,1) covariance structure and a three-term Fourier series mean function, and it included 39 clusters for Ostreococcus, 25 clusters for Synechococcus, 13 clusters for Pelagibacter, 7 clusters for SAR86, and 9 clusters for MGII (SI Appendix, Fig. S12).
Regression Tests for Count Data.
Gene-by-gene tests to identify transcripts exhibiting sinusoidal periodicity or covariance with pathway-level functions used Poisson log-linear regression as implemented in the R software package (34). Library size offsets were based on the total number of transcripts assigned to a given taxon at each time point. For periodicity tests, the sinusoidal function  , where A represents the amplitude, ω is the phase, and t is the midpoint of the sampling time in hours, was reduced to the linear equation
, where A represents the amplitude, ω is the phase, and t is the midpoint of the sampling time in hours, was reduced to the linear equation  , where
, where  and
and  .
.
The significance of each model fit was assessed using both a χ2 test (as implemented in the anova.glm function) and a permutation test. Permutation P values were calculated as the fraction of randomized datasets with a model fit (evaluated using the difference between the null and residual deviance) as good or better than model fits of the actual experimental data. To optimize computational resources, permutations continued until at least 10 randomized datasets with likelihood ratios equal to or exceeding the observed data had been identified (500–50,000 permutations). False discovery rate-corrected (35) P values of at least 0.1 from both tests were required for a relationship to be considered significant.
Supplementary Material
Acknowledgments
This manuscript is dedicated to the memory of Carl R. Woese, who profoundly and forever changed our understanding of the evolution and nature of life on Earth. We thank the Environmental Sample Processor team for their help in planning and implementing this experiment, especially Gene Massion, Brent Roman, Scott Jensen, Roman Marin III, Chris Preston, Jim Birch, and Julie Robidart. We also thank the engineering technicians and machinists at Monterey Bay Aquarium Research Institute (MBARI) for their help and dedication to instrument development, and the crew of the R/V Western Flyer for their support and expertise during field operations. This work is a contribution to the Center for Microbial Ecology: Research and Education (C-MORE). Development and application of the Environmental Sample Processor has been supported by National Science Foundation Grant OCE-0314222 (to C.A.S.), National Aeronautics and Space Administration Astrobiology Grants NNG06GB34G and NNX09AB78G (to C.A.S.), the Gordon and Betty Moore Foundation (C.A.S.), and the David and Lucile Packard Foundation. This work was supported by National Science Foundation Science and Technology Center Award EF0424599 (to C.A.S. and E.F.D.), a grant from the Gordon and Betty Moore Foundation (to E.F.D.), and a gift from the Agouron Institute (to E.F.D.).
Footnotes
The authors declare no conflict of interest.
Data deposition: The sequences reported in this paper have been deposited in the GenBank database (accession no. SRA062433), and the CAMERA (http://camera.calit2.net) database repository (accession no. CAM_P_0001026).
This article contains supporting information online at www.pnas.org/lookup/suppl/doi:10.1073/pnas.1222099110/-/DCSupplemental.
References
- 1.Karl DM. Nutrient dynamics in the deep blue sea. Trends Microbiol. 2002;10(9):410–418. doi: 10.1016/s0966-842x(02)02430-7. [DOI] [PubMed] [Google Scholar]
- 2.Frias-Lopez J, et al. Microbial community gene expression in ocean surface waters. Proc Natl Acad Sci USA. 2008;105(10):3805–3810. doi: 10.1073/pnas.0708897105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Gilbert JA, et al. Detection of large numbers of novel sequences in the metatranscriptomes of complex marine microbial communities. PLoS One. 2008;3(8):e3042. doi: 10.1371/journal.pone.0003042. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Poretsky RS, et al. Comparative day/night metatranscriptomic analysis of microbial communities in the North Pacific subtropical gyre. Environ Microbiol. 2009;11(6):1358–1375. doi: 10.1111/j.1462-2920.2008.01863.x. [DOI] [PubMed] [Google Scholar]
- 5.Hewson I, Poretsky RS, Tripp HJ, Montoya JP, Zehr JP. Spatial patterns and light-driven variation of microbial population gene expression in surface waters of the oligotrophic open ocean. Environ Microbiol. 2010;12(7):1940–1956. doi: 10.1111/j.1462-2920.2010.02198.x. [DOI] [PubMed] [Google Scholar]
- 6.Ottesen EA, et al. Metatranscriptomic analysis of autonomously collected and preserved marine bacterioplankton. ISME J. 2011;5(12):1881–1895. doi: 10.1038/ismej.2011.70. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Marchetti A, et al. Comparative metatranscriptomics identifies molecular bases for the physiological responses of phytoplankton to varying iron availability. Proc Natl Acad Sci USA. 2012;109(6):E317–E325. doi: 10.1073/pnas.1118408109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Gifford SM, Sharma S, Booth M, Moran MA. Expression patterns reveal niche diversification in a marine microbial assemblage. ISME J. 2012 doi: 10.1038/ismej.2012.96. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Martin AP, Zubkov MV, Burkill PH, Holland RJ. Extreme spatial variability in marine picoplankton and its consequences for interpreting Eulerian time-series. Biol Lett. 2005;1(3):366–369. doi: 10.1098/rsbl.2005.0316. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Preston CM, et al. Underwater application of quantitative PCR on an ocean mooring. PLoS One. 2011;6(8):e22522. doi: 10.1371/journal.pone.0022522. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Morris RM, et al. SAR11 clade dominates ocean surface bacterioplankton communities. Nature. 2002;420(6917):806–810. doi: 10.1038/nature01240. [DOI] [PubMed] [Google Scholar]
- 12.Scanlan DJ, et al. Ecological genomics of marine picocyanobacteria. Microbiol Mol Biol Rev. 2009;73(2):249–299. doi: 10.1128/MMBR.00035-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Demir-Hilton E, et al. Global distribution patterns of distinct clades of the photosynthetic picoeukaryote Ostreococcus. ISME J. 2011;5(7):1095–1107. doi: 10.1038/ismej.2010.209. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Dupont CL, et al. Genomic insights to SAR86, an abundant and uncultivated marine bacterial lineage. ISME J. 2012;6(6):1186–1199. doi: 10.1038/ismej.2011.189. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Iverson V, et al. Untangling genomes from metagenomes: Revealing an uncultured class of marine Euryarchaeota. Science. 2012;335(6068):587–590. doi: 10.1126/science.1212665. [DOI] [PubMed] [Google Scholar]
- 16.Li N, et al. Functional clustering of periodic transcriptional profiles through ARMA(p,q) PLoS One. 2010;5(4):e9894. doi: 10.1371/journal.pone.0009894. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Ito H, et al. Cyanobacterial daily life with Kai-based circadian and diurnal genome-wide transcriptional control in Synechococcus elongatus. Proc Natl Acad Sci USA. 2009;106(33):14168–14173. doi: 10.1073/pnas.0902587106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Monnier A, et al. Orchestrated transcription of biological processes in the marine picoeukaryote Ostreococcus exposed to light/dark cycles. BMC Genomics. 2010;11:192. doi: 10.1186/1471-2164-11-192. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Binder BJ, Chisholm SW. Cell cycle regulation in marine Synechococcus sp. strains. Appl Environ Microbiol. 1995;61(2):708–717. doi: 10.1128/aem.61.2.708-717.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Binder B. Cell cycle regulation and the timing of chromosome replication in a marine Synechococcus (cyanobacteria) during light- and nitrogen-limited growth. J Phycol. 2000;36(1):120–126. [Google Scholar]
- 21.Sun J, et al. One carbon metabolism in SAR11 pelagic marine bacteria. PLoS One. 2011;6(8):e23973. doi: 10.1371/journal.pone.0023973. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Keener J, Nomura M. Regulation of ribosome synthesis. In: Neidhardt FC, et al., editors. Escherichia coli and Salmonella: Cellular and Molecular Biology. 2nd Ed. Washington, DC: ASM Press; 1996. [Google Scholar]
- 23.Fazio A, et al. Transcription factor control of growth rate dependent genes in Saccharomyces cerevisiae: A three factor design. BMC Genomics. 2008;9:341. doi: 10.1186/1471-2164-9-341. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Hendrickson EL, et al. Global responses of Methanococcus maripaludis to specific nutrient limitations and growth rate. J Bacteriol. 2008;190(6):2198–2205. doi: 10.1128/JB.01805-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Smith DP, et al. Transcriptional and translational regulatory responses to iron limitation in the globally distributed marine bacterium Candidatus pelagibacter ubique. PLoS One. 2010;5(5):e10487. doi: 10.1371/journal.pone.0010487. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Giovannoni SJ, et al. Genome streamlining in a cosmopolitan oceanic bacterium. Science. 2005;309(5738):1242–1245. doi: 10.1126/science.1114057. [DOI] [PubMed] [Google Scholar]
- 27.Ng WL, Bassler BL. Bacterial quorum-sensing network architectures. Annu Rev Genet. 2009;43:197–222. doi: 10.1146/annurev-genet-102108-134304. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Stewart FJ, Ottesen EA, DeLong EF. Development and quantitative analyses of a universal rRNA-subtraction protocol for microbial metatranscriptomics. ISME J. 2010;4(7):896–907. doi: 10.1038/ismej.2010.18. [DOI] [PubMed] [Google Scholar]
- 29.Palenik B, et al. The tiny eukaryote Ostreococcus provides genomic insights into the paradox of plankton speciation. Proc Natl Acad Sci USA. 2007;104(18):7705–7710. doi: 10.1073/pnas.0611046104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Kanehisa M, Goto S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 2000;28(1):27–30. doi: 10.1093/nar/28.1.27. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Moriya Y, Itoh M, Okuda S, Yoshizawa AC, Kanehisa M. KAAS: An automatic genome annotation and pathway reconstruction server. Nucleic Acids Res. 2007;35(Web Server issue):W182–W185. doi: 10.1093/nar/gkm321. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Liu Z, Hsiao W, Cantarel BL, Drábek EF, Fraser-Liggett C. Sparse distance-based learning for simultaneous multiclass classification and feature selection of metagenomic data. Bioinformatics. 2011;27(23):3242–3249. doi: 10.1093/bioinformatics/btr547. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Okansen J, et al. Vegan: Community Ecology Package, R Package Version 2.0-2. 2011. Available at http://cran.r-project.org/web//packages/vegan/index.html.
- 34.Team RDC. R: A Language and Environment for Statistical Computing. Vienna: R Foundation for Statistical Computing; 2012. [Google Scholar]
- 35.Benjamini Y, Hochberg Y. Controlling the false discovery rate—a practical and powerful approach to multiple testing. J R Stat Soc Series B Stat Methodol. 1995;57(1):289–300. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.